Abstract
1,4-Dioxane, a probable human carcinogen, is a co-contaminant at many chlorinated solvent-contaminated sites. Although numerous 1,4-dioxane-degrading aerobic bacteria have been isolated, almost no information exists on the microorganisms able to degrade this chemical under anaerobic conditions. Here, the potential for 1,4-dioxane biodegradation was examined using multiple inocula and electron acceptor amendments. The inocula included uncontaminated agricultural soils and river sediments as well as sediments from two 1,4-dioxane contaminated sites. Five separate experiments involved the examination of triplicate live microcosms and abiotic controls for approximately 1 year. Compound-specific isotope analysis (CSIA) was used to further investigate biodegradation in a subset of the microcosms. Also, DNA was extracted from microcosms exhibiting 1,4-dioxane biodegradation for microbial community analysis using 16S rRNA gene amplicon high-throughput sequencing. Given the long incubation periods, it is likely that electron acceptor depletion occurred and methanogenic conditions eventually dominated. The iron/EDTA/humic acid or sulfate amendments did not result in 1,4-dioxane biodegradation in the majority of cases. 1,4-dioxane biodegradation was most commonly observed in the nitrate amended and no electron acceptor treatments. Notably, both contaminated site sediments illustrated removal in the samples compared to the abiotic controls in the no electron acceptor treatment. However, it is important to note that the degradation was slow (with concentration reductions occurring over approximately 1 year). In two of the three cases examined, CSIA provided additional evidence for 1,4-dioxane biodegradation. In one case, the reduction in 1,4-dioxane in the samples comparing the controls was likely too low for the method to detect a significant 13C/12C enrichment. Further research is required to determine the value of measuring 2H/1H for generating evidence for the biodegradation of this chemical. The microbial community analysis indicated that the phylotypes unclassified Comamonadaceae and 3 genus incertae sedis were more abundant in 1,4-dioxane-degrading microcosms compared to the live controls (no 1,4-dioxane) in microcosms inoculated with contaminated and uncontaminated sediment, respectively. The relative abundance of known 1,4-dioxane degraders was also investigated at the genus level. The soil microcosms were dominated primarily by Rhodanobacter with lower relative abundance values for Pseudomonas, Mycobacterium, and Acinetobacter. The sediment communities were dominated by Pseudomonas and Rhodanobacter. Overall, the current study indicates 1,4-dioxane biodegradation under anaerobic and, likely methanogenic conditions, is feasible. Therefore, natural attenuation may be an appropriate cleanup technology at sites where time is not a limitation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
There is a critical need to develop remediation strategies for the contaminant 1,4-dioxane due to its widespread occurrence (Adamson et al. 2015) and its classification as a probable human carcinogen (DeRosa et al. 1996). Historically, 1,4-dioxane was used as a stabilizer in 1,1,1-trichloroethane (1,1,1-TCA) formulations and is now frequently detected at chlorinated solvent-contaminated sites (Adamson et al. 2015; Adamson et al. 2014; Mohr et al. 2010). As 1,4-dioxane was typically not on the EPA’s target compound lists, it is likely that closed sites will require reopening to address contamination. In fact, a multisite survey aimed at examining the extent of the 1,4-dioxane problem, indicated a primary risk is the large number of sites where this chemical is likely to be present but has yet to be identified (Adamson et al. 2014).
The remediation of 1,4-dioxane contaminated sites is problematic because of chemical characteristics that result in movement and persistence (Adamson et al. 2015; Mohr et al. 2010). Traditional methods such as air stripping or activated carbon can be ineffective because of the chemical’s low organic carbon partition coefficient (log KOC = 1.23) and low Henry’s Law Constant (5 × 10−6 atm m3mol−1) (Mahendra and Alvarez-Cohen 2006; Steffan et al. 2007; Zenker et al. 2003). Ex situ oxidation methods such as ozone and hydrogen peroxide (Adams et al. 1994) or hydrogen peroxide and ultraviolet light (Stefan and Bolton 1998) have been commercially applied, although at high concentrations these methods can be cost prohibitive (Steffan et al. 2007). Given the limitations associated with traditional remediation methods, interest has turned to the use of microorganisms to biodegrade this problematic contaminant (Chu et al. 2009; Li et al. 2010; Lippincott et al. 2015).
Numerous bacteria have been associated with the biodegradation (co-metabolic or metabolic) of 1,4-dioxane. Those linked to aerobic metabolic biodegradation classify within the genera Pseudonocardia (Kampfer and Kroppenstedt 2004; Mahendra and Alvarez-Cohen 2005; Mahendra and Alvarez-Cohen 2006; Matsui et al. 2016; Parales et al. 1994; Sei et al. 2013a; Yamamoto et al. 2018), Mycobacterium (Kim et al. 2009; Sei et al. 2013a), Afipia (Isaka et al. 2016; Sei et al. 2013a), Xanthobacter (Chen et al. 2016), Acinetobacter (Huang et al. 2014), Rhodococcus (Bernhardt and Diekmann 1991; Inoue et al. 2016; Inoue et al. 2018), and Rhodanobacter (Pugazhendi et al. 2015).
Genera associated with aerobic co-metabolic 1,4-dioxane degradation include Pseudonocardia (Kohlweyer et al. 2000; Mahendra and Alvarez-Cohen 2006; Vainberg et al. 2006)], Mycobacterium (Lan et al. 2013; Masuda 2009) and Rhodococcus (Hand et al. 2015; Lippincott et al. 2015; Mahendra and Alvarez-Cohen 2006; Sei et al. 2013b; Steffan et al. 1997; Stringfellow and Alvarez-Cohen 1999), Flavobacterium (Sun et al. 2011), Burkholderia (Mahendra and Alvarez-Cohen 2006), Nocardia (Masuda 2009), Ralstonia (Mahendra and Alvarez-Cohen 2006), Pseudomonas (Mahendra and Alvarez-Cohen 2006), Methylosinus (Mahendra and Alvarez-Cohen 2006; Whittenbury et al. 1970) and Azoarcus (Deng et al. 2018). However, it is unlikely that these aerobic 1,4-dioxane degraders will be effective at chlorinated solvent sites, as these sites are typically highly reducing. Under such conditions, tetrachloroethene and trichloroethene undergo sequential reductive dechlorination to cis-1,2-dichloroethene and vinyl chloride, finally forming the nontoxic end product, ethene (Freedman and Gossett 1989). Reduction is commonly associated with microbial taxa such as Dehalococcoides mccartyi and Dehalobacter (Cupples 2008; Cupples et al. 2003; He et al. 2003; Holliger et al. 1998; Löffler et al. 2013; Maymó-Gatell et al. 1997; Sung et al. 2006). Commercially available reductive dechlorinating mixed cultures containing such strains are frequently used for bioaugmenting contaminated groundwater aquifers (Major et al. 2002; Steffan and Vainberg 2013; Vainberg et al. 2009). The lack of information on the susceptibility of the chlorinated solvent co-contaminant 1,4-dioxane to biodegradation under such highly reducing conditions is a significant knowledge gap.
No anaerobic 1,4-dioxane-degrading isolates have been identified, and limited research has addressed 1,4-dioxane biodegradation under anaerobic conditions. One project investigated 1,4-dioxane degradation over a range of redox conditions (aerobic, nitrate reducing, iron reducing, sulfate reducing, and methanogenic) (Steffan 2007). The work involved microcosm experiments with soil and groundwater from a site heavily contaminated with the chlorinated solvents and 1,4-dioxane. In these tests, nitrate, nitrite, sulfate, and ferric iron were added as electron acceptors. In another set of experiments, samples from across a vegetable oil biobarrier were investigated, without the addition of electron acceptors, as it was expected that the biobarrier had resulted in a range of redox conditions. Notably, 1,4-dioxane was not degraded in any of the anaerobic microcosms during > 400 days.
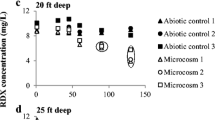
Another study produced more promising results, documenting 1,4-dioxane biodegradation under iron-reducing conditions using an enrichment originating from wastewater treatment plant sludge (Shen et al. 2008). The researchers found that when Fe(III) was supplied as Fe(III)-EDTA, biodegradation was stimulated. Also, they reported that humic acids stimulated the activity of the Fe(III) reducing bacteria and 1,4-dioxane biodegradation. The authors hypothesized that humic acids promoted electron shuttling to Fe(III) in a catalytic manner. Others have also reported stimulation of bacterial growth and biodegradation (cyclic nitroamines RDX and HMX) in the presence of humic acids and Fe(III) (Bhushan et al. 2006).
The overall objective of the current study was to determine the susceptibility of 1,4-dioxane to biodegradation under anaerobic conditions using inocula from contaminated and uncontaminated sites. For this, microcosms were established with different inocula (soils or sediments) and various electron acceptor amendments (nitrate, iron-EDTA/humic acid, sulfate, and no amendment). Additionally, compound-specific isotope analysis (CSIA) was used to determine changes in 13C/12C ratios in a subset of the samples. The work is novel as it is the first to document the frequency of 1,4-dioxane biodegradation over a range redox conditions, and inocula types and provides critical evidence for the feasibility of anaerobic 1,4-dioxane bioremediation.
Methods
Chemicals and inocula
1,4-Dioxane was purchased from Alfa Aesar (MA, USA, 99+ % purity) and Sigma-Aldrich (MO, USA, 99.8% purity). Ethylenediaminetetraacetic acid (EDTA), iron (III) sodium salt, sodium sulfate, sodium nitrate, and humic acid were purchased from Sigma-Aldrich (MO, USA). All stock solutions and dilutions were prepared using DI water. The uncontaminated agricultural soil samples were collected from two locations on the campus of Michigan State University (MSU), East Lansing, MI (herein called Soils E, F, and G) and two locations at the Kellogg Biological Station (MSU), Hickory Corners, MI (KBS Soils 1, 2, and 3). The uncontaminated river sediment samples (Sediments H and J) were collected from Red Cedar River, Okemos, MI, and from a river leading to Lake Lansing in Haslett, MI. The contaminated site samples were sent to MSU from California (contaminated with trichloroethene, 1,1-dichloroethene, and 1,4-dioxane) and Maine (contaminated with traces of 1,4-dioxane). All of the samples were stored in the dark at 6 °C until use.
Experimental setup
Five sets of experiments were performed to examine the susceptibility of 1,4-dioxane to biodegradation under different potential electron acceptors (Table 1). All five experiments contained triplicates of live samples and abiotic controls (autoclaved daily for three consecutive days). The final set of experiments (experiment 5) also included live control microcosms (no 1,4-dioxane) to enable comparisons to the 1,4-dioxane-degrading microbial communities.
The first set of experiments was designed based on the positive results generated previously for 1,4-dioxane under iron-reducing conditions (Shen et al. 2008). For this, 18 microcosms (70-mL serum bottles) were each inoculated with one of three agricultural soils (Soils E, F and G, 20 g wet weight) and 55 mL of a solution consisting of EDTA iron (III) sodium salt (10 mM), sodium lactate (5 mM), and humic acid (0.5 g/L). The triplicate microcosms and abiotic controls were amended with approximately 5 mg/L of 1,4-dioxane (Alfa Aesar).
The second set of microcosms contained the same three agricultural soils (Soils E, F, and G) under four redox conditions (nitrate, iron, sulfate amended, and no amendment). Here, 72 microcosms were established (3 soils, 4 treatments) containing soil (5 g wet weight), 10 mL of a solution with either NaNO3 (10 mM), NaSO4 (10 mM), EDTA iron(III) sodium salt (10 mM) or DI water (no amendment) in 30 mL glass serum bottles. The microcosms and abiotic controls were amended with approximately 5 mg/L of 1,4-dioxane (Alfa Aesar). The third set of microcosms contained each of the uncontaminated river sediment samples (Sediments H and J) using the same experimental setup (four redox conditions). Sodium lactate was not added for both sets of experiments to encourage the metabolic degradation of 1,4-dioxane.
The fourth set of microcosms (48 bottles including 2 sediments, 4 treatments) contained each of the contaminated sediment samples (from CA and MN) also under four redox conditions (nitrate, iron, sulfate amended, and no amendment). In this case, a media solution was added containing NH4Cl (1.5 g/L), NaH2PO4 (0.6 g/L), CaCl2.2H2O (0.1 g/L), KCl (0.1 g/L), MgCl2.6H2O (0.002 g/L), and sodium lactate (5 mM). The electron acceptor and 1,4-dioxane amendments were as described above, and 1,4-dioxane from Sigma Aldrich was used.
The final set of microcosms contained no electron acceptor amendment with the aim of creating methanogenic conditions (based on positive results from some of the microcosms described above). The 72 microcosms included 8 different inocula (Soils E, F, and G; KBS Soils 1, 2, and 3; Sediment H; and the CA contaminated sediment) with triplicates of sample microcosms and abiotic controls. Additionally, triplicate live control microcosms were included, and these were treated in the same manner as the sample microcosms except no 1,4-dioxane was added. As stated above, this treatment was included to enable comparisons to the microbial communities exposed to 1,4-dioxane. The microcosms contained 5 g (wet weight) of soil or sediment and 25 mL of a solution containing sodium lactate (5 mM), NH4Cl (1.5 g/L), NaH2PO4 (0.6 g/L), CaCl2.2H2O (0.1 g/L), KCl (0.1 g/L), and MgCl2.6H2O (0.002 g/L) in 30-mL amber glass serum bottles. 1,4-dioxane (Sigma Aldrich, between 5 and 14 mg/L) was added the sample microcosms and abiotic controls.
All solutions were purged under a stream of nitrogen before being introduced into the anaerobic chamber. The solutions and soils/sediments were added to each serum bottle in the anaerobic chamber, and, after approximately 3 days, 1,4-dioxane was added. The microcosms, closed with septa, were incubated in the anaerobic chamber at 20 °C. The anaerobic chamber was maintained with gaseous mix of approximately 5% H2, 90% N2, and 5% CO2. The vials were sealed using BiMetal vial crimp with PTFE/silicone septas to maintain the cultures at anaerobic conditions. All microcosms were transferred on a shaker at 200 rpm and maintained at 20 °C. The nitrate-amended microcosms were tested for methane after 200 days of incubation using a GC (Hewlett Packard 5890).
1,4-Dioxane analysis
A GC/MS with Agilent 5975 GC/single quadrupole MS (Agilent Technologies, CA, USA) equipped with a CTC Combi Pal autosampler was used to determine 1,4-dioxane concentrations. Sterile syringes (1 mL) and needles (22 Ga 1.5 in.) were used to collect samples (0.7 mL) from each microcosm. The samples were filtered (0.22 μm nylon filter) before being injected into an amber glass vial (40 mL) for GC/MS analysis. A method was developed to analyze 1,4-dioxane using solid-phase micro-extraction (SPME). The SPME fiber was inserted in the headspace of the vial and exposed to the analyte for 1 min before being injected into the GC for thermal desorption. The fiber coating can adsorb the analytes in the vapor phase. Splitless injection was executed and the vials were maintained at 40 °C. The SPME fiber assembly involved 50/30 μm divinylbenzene/carboxen/polydimethylsiloxane (DVB/CAR/PDMS) and 24 Ga needle for injection. The initial oven temperature was 35 °C and was programmed to increase at a rate of 20 to 120 °C/min. Once it reached 120 °C, it increased at a rate of 40 to 250 °C/min, which was maintained for 3 min. A VF5MS column was used with helium as the carrier gas in constant flow mode at a flow rate of 1 ml/min. The conditioning of the SPME fiber was at 270 °C for 60 min at the beginning of each sequence.
Compound-specific isotope analysis (CSIA)
At the last sampling point, three sets of samples and abiotic controls were submitted for CSIA analysis, including Soil F (nitrate amended, experiment two), and the two contaminated site samples with no electron acceptor amendment (experiment four). CSIA was performed at the University of Waterloo Environmental Isotope Laboratory (UWEIL), Ontario, Canada. The ratios of 13C/12C and 2H/1H were measured using a recently developed method. For this, the dilute 1,4-dioxane samples were concentrated onto a sorbent and subjected to thermal desorption in a GC coupled with an isotope ratio mass spectrometer (Bennett et al. 2018). All three sets of samples were submitted to 13C/12C analysis; however, only Soil F was subject to 2H/1H.
DNA extraction and high-throughput amplicon sequencing
DNA was extracted (2 mL liquid and 0.6 g soil/sediment, QIAGEN DNeasy PowerSoil kit) from the sample microcosms and corresponding live control microcosms (no electron acceptor amendment, experiment five) that illustrated 1,4-dioxane degradation (Soil F, CA Contaminated Site, Sediment H and KBS Soil 3, 24 extracts). Additionally, DNA was extracted from 31 other microcosms that illustrated 1,4-dioxane biodegradation (from the other four experiments). The concentration of DNA was determined using QUBIT dsDNA HS kit. The DNA extracts were submitted for high-throughput 16S rRNA gene amplicon sequencing following previously described protocols (Caporaso et al. 2012; Caporaso et al. 2011) at the Research Technology Support Facility at Michigan State University. Specifically, 55 samples of microbial metagenomic DNA were submitted for amplicon library preparation and sequencing. The V4 hypervariable region of the 16S rRNA gene was amplified using the Illumina compatible, dual indexed primers 515f/806r. The primers and the library construction protocol developed in Patrick Schloss’ lab were previously described (Kozich et al. 2013). PCR products were batch normalized using Invitrogen SequalPrep DNA Normalization plates and product recovered from the plate pooled. This pool was cleaned up and concentrated using AmpureXP magnetic beads. The pool was QC’d and quantified using a combination of Qubit dsDNA HS, Agilent 4200 TapeStation HS DNA1000, and Kapa Illumina Library Quantification qPCR assays. The pool was then loaded onto an Illumina MiSeq v2 Standard flow cell and sequencing was performed in a 2x250bp paired end format using a MiSeq v2 500 cycle reagent kit. Custom sequencing and index primers complementary to the 515f/806r oligos were added to appropriate wells of the reagent cartridge as previously described (Kozich et al. 2013). Base calling was performed by Illumina Real-Time Analysis (RTA) v1.18.54, and the output of RTA was demultiplexed and converted to FastQ format with Illumina Bcl2fastq v2.19.1.
Analysis of sequencing data
The amplicon sequencing data in the fastq file format was analyzed with mothur (version 1.41.3) from Patrick D. Schloss Laboratory (Schloss 2009) using the MiSeq standard operating procedure (Schloss 2013). The steps involved the removal of barcode information, and contiguous sequences were created using the forward and reverse reads. These were analyzed for errors and then classified. This involved checking the samples for the proper read length (< 275 bp), ambiguous bases, and homopolymer length greater than 8 to eliminate such sequences. The sequences were then aligned with the SILVA bacteria database (Release 132) (Pruesse et al. 2007) for the V4 region. Chimeras and mitochondrial and chloroplast lineage sequences were removed, and then the sequences were classified into OTUs. The summary file created by mothur was analyzed with Microsoft Excel 2016 and STAMP (Statistical Analyses of Metagenomic Profiles, software version 2.1.3.) (Parks et al. 2014). STAMP was used to detect differences in the relative proportions of the taxonomic profiles between the live controls (no 1,4-dioxane) and the samples for each of the four inocula in the fifth set of experiments (Soil F, Contaminated Site 10A, Sediment H, and KBS Soil 3). This analysis included Welch’s two-sided t-test for two groups (samples and live controls) (p < 0.05) to generate extended error bar figures for each soil. The sequencing data was submitted to NCBI under Bioproject PRJNA590578 (accession numbers SAMN13335577 to SAMN13335631, Supplementary Table 1).
Statistical analysis
1,4-Dioxane concentrations in the samples and abiotic controls were compared using the Student’s t test for independent variables with equal variance. Normality of the 1,4-dioxane concentration data was confirmed using the Shapiro-Wilk test (p > 0.05) and the assumption of equal variance was confirmed using Levene’s tests (p > 0.05). The same tests were used to confirm equal variance and normality for the CSIA carbon data. The CSIA hydrogen data were not normal; therefore the Mann-Whitney U test was used. These statistical tests were performed in XLSTAT software for statistical analysis in Excel (2019, http://www.xlstat.com).
Results
1,4-Dioxane removal trends
1,4-Dioxane concentrations in samples and abiotic controls were monitored for approximately 1 year in each of the five sets of experiments. Given the long incubation period, it is likely that the electron acceptors (nitrate, iron, sulfate) were depleted for each treatment. This was confirmed in one case, by the production of methane in the nitrate-amended, agricultural soil-inoculated microcosms. The first set of microcosms (inoculated with three agricultural soils and an EDTA iron/humic acid amendment) illustrated no significant difference (p < 0.05) between the samples and abiotic controls for the majority of sampling times, suggesting no biological 1,4-dioxane degradation occurred with this treatment for these soils (Fig. 1).
1,4-Dioxane concentrations in sample and abiotic control microcosms of three soils amended with EDTA/iron and humic acid. The values and bars represent averages and standard deviations from triplicates. The circles represent a significant difference (two-tailed t-test, p < 0.05) between the samples and abiotic controls (when abiotic controls > samples)
The second set of experiments (agricultural soils and four electron acceptor treatments) produced a variety of trends between the soils and treatments (Fig. 2). Comparing the treatments, the most consistent removal was in the nitrate-amended microcosms (Fig. 2a). For all three soils, there was a statistically significant difference (p < 0.05) between the samples and abiotic controls at the last sampling point (day 295 to 316). The Soil F nitrate-amended sample microcosms and abiotic controls were submitted for CSIA (as discussed later). Similar to the first set of microcosms, no sustained significant degradation was noted in the samples compared to the abiotic controls in the iron-amended microcosms (Fig. 2b).
1,4-Dioxane concentrations in sample and abiotic control microcosms of three soils with different electron acceptor amendments (a–d). The values and bars represent averages and standard deviations from triplicates (except Soil G, n = 2 for three cases). The circles represent a significant different (two-tailed t-test, p < 0.05) between the samples and abiotic controls (when controls > samples). The arrows indicate the samples subject to DNA extraction. CSIA was performed on the samples and abiotic controls from the Soil F, nitrate-amended treatments
In the sulfate-amended microcosms, only Soil E resulted in a significant difference between the samples and abiotic controls over several time points (Fig. 2c). When no electron acceptor was added, only Soils E and F illustrated consistent reduced concentrations in the samples compared to the abiotic controls (Fig. 2d). Overall, the three soils illustrated a range of 1,4-dioxane biodegradation abilities, depending on the electron acceptor amendment. Specifically, Soil E most commonly produced significant differences between the samples and abiotic controls, followed by Soil F and then Soil G (Table 2).
The third set of experiments involved Sediment H and Sediment J (both collected from uncontaminated rivers) as inocula. No significant decreases in 1,4-dioxane concentrations were seen in nitrate-amended, sulfate-amended or iron-amended samples compared to the abiotic controls for both sediments (Fig. 3a, b, c). Among the samples with no amendments, only Sediment H indicated a significant decrease in 1,4-dioxane concentrations compared to the abiotic controls (Fig. 3d). This was one of the most dramatic reductions in 1,4-dioxane in the live sample microcosms compared to the abiotic controls over all treatments and inocula types.
1,4-Dioxane concentrations in Sediments J and H sample and abiotic control microcosms over time with different amendments. The points illustrate average values from triplicates (except nitrate-amended controls, Sediment H, n = 1), and the bars illustrate standard deviations. The circles represent a significant different (two-tailed t-test, p < 0.05) between the samples and abiotic controls. The arrow indicates the samples subject to DNA extraction
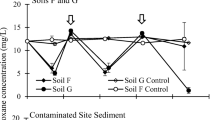
The fourth set of experiments (inoculated with sediments from two contaminated sites over four treatments) illustrated similar trends as those from the agricultural soils (Fig. 4). Specifically, in one case for each of the nitrate-amended and sulfate-amended microcosms, there was a significant difference between the samples and abiotic controls (CA and MN, respectively) (Fig. 4a, c). Further, no differences were observed between the samples and abiotic controls under the iron-amended treatment (Fig. 4b). Finally, in both cases, a substantial decrease (~ 17% and 50% for CA and MN) was observed in the samples compared to the abiotic controls when no electron acceptor was added (Fig. 4d). The sample microcosms and abiotic control microcosms from the no amendment treatment with both sediments were submitted for CSIA analysis.
1,4-Dioxane concentrations in Maine and California sample and abiotic control microcosms over time with different amendments. The points illustrate average values from triplicates (except sulfate-amended Maine sample, n = 2, and sulfate-amended CA control, n = 2) and the bars illustrate standard deviations. The circles represent a significant different (two-tailed t-test, p < 0.05) between the samples and abiotic controls (controls > samples). The arrows indicate the samples subject to DNA extraction. CSIA was performed on the samples and abiotic controls from both of the no electron acceptor treatment (D)
A summary of the results described in the four sets of experiments is provided, focusing only on the last sampling point for each (Table 2). In the nitrate-amended treatments, four of the six inocula types (or 57%) produced a significant reduction in 1,4-dioxane in the samples compared to the abiotic controls. In contrast, no differences were noted between the samples and abiotic controls under any of the iron amendment treatments. Only two (Soil E and the MN contaminated site sediments) of the seven cases (or 29%) illustrated biodegradation under the sulfate amendment treatment. Finally, the most frequent (five of the seven inocula types or 71%) reduction of 1,4-dioxane was noted under the no amendment treatment. The average removal (again, only at the last time point) was greatest under the no amendment treatment (22 ± 19%), followed by the nitrate-amended samples (16 ± 16%) and the sulfate-amended samples (7 ± 12%). Overall, it appears that likelihood of 1,4-dioxane biodegradation is the greatest with no amendments, when the microbial populations are likely under methanogenic redox conditions.
The fifth set of experiments were established to compare the microbial communities between the live microcosms and the live control microcosms (no 1,4-dioxane amendment) and focused only on the no amendment treatment. For this, six agricultural soils, one contaminated (CA) and one uncontaminated sediment (Sediment H) sample were used as inocula (Fig. 5). 1,4-Dioxane concentrations were monitored in the samples and abiotic controls for more than 400 days. A significant reduction in 1,4-dioxane in the samples compared to the abiotic controls was only noted for two of six agricultural soils (Fig. 5b, h). Consistent with the results described above (third and fourth set of experiments), a significant reduction in 1,4-dioxane was noted in the samples compared to the abiotic controls in Sediment H (Fig. 5e) and the contaminated samples from CA (Fig. 5d). In this set of experiments, only 50% of the inocula types illustrated detectable levels of 1,4-dioxane biodegradation.
1,4-Dioxane concentrations in sample and abiotic control microcosms over time with different inocula. The points illustrate average values from triplicates (except Soil E and KBS Soil 2 samples, which are replicates) and the bars illustrate standard deviations. The circles represent a significant different (two-tailed t-test, p < 0.05) between the samples and abiotic controls. The arrows indicate the samples subject to DNA extraction
Compound-specific isotope analysis (CSIA)
CSIA was used to further investigate biodegradation in three sets of the anaerobic 1,4-dioxane-degrading microcosms. For this, subsamples were collected from sample microcosms and abiotic controls from the Soil F/nitrate treatment (second set of experiments) and from the CA and MN contaminated site sediments/no electron acceptor treatment (fourth set of experiments) and these were sent to UWEIL for CSIA. This laboratory has already developed the methodology to measure δ13C and δ2H values for 1,4-dioxane (Bennett et al. 2018). All three sets of samples (six for each treatment, eighteen in total) were subject to 13C/12C analysis, but only the six Soil F/nitrate microcosms were subject to 2H/1H analysis. 1,4-Dioxane degradation should result in more positive 13C/12C and 2H/1H ratios (or more positive δ13C and δ2H values) because bonds involving heavier isotopes are more difficult to break, and so bonds consisting of lighter isotopes are preferentially degraded, causing the residual, non-degraded contaminant to be heavy isotope enriched. In all cases, more positive δ13C and δ2H values were found in the live samples compared to the abiotic controls (Fig. 6). However, the differences in δ13C values were only statistically significantly different between the samples and abiotic controls for Soil F/nitrate and contaminated site MN/no electron acceptor. It is likely that the decrease in 1,4-dioxane concentrations was too low for this method to detect a difference in δ13C values for the contaminated site CA/no electron acceptor. Specifically, for this treatment the average reduction in 1,4-dioxane concentration in the samples compared to the controls was 17% compared to 25% and 50% reductions in the other two treatments. The δ2H values were not significantly different between the samples and controls for Soil F (Fig. 6b).
Comparison of δ13C (a) and δ2H (h) values in samples and abiotic controls of a subset of the experimental microcosms. The error bars represent standard deviations from triplicate samples and control microcosms, and the p values were derived from the Student’s t test (two tailed, independent variables, equal variance)
Putative 1,4-dioxane degraders
To determine if any microorganism could obtain a growth benefit from 1,4-dioxane degradation, sample microcosms were supplied with media and 1,4-dioxane, and the live control microcosms were supplied with the same media, but no 1,4-dioxane (fifth set of microcosms). Therefore, an increase in the relative abundance of any microorganism between the samples and live controls could be attributed to the presence of 1,4-dioxane. From this, a reasonable hypothesis would be that the enriched microorganisms are being exposed to growth supporting substrates from 1,4-dioxane degradation. This comparison focused on the four inocula that illustrated 1,4-dioxane degradation (Fig. 5b, d, e, h). Significant phylotype enrichments (p < 0.05) were noted for the microcosms inoculated with CA contaminated site sediments, Sediment F and KBS Soil 3 (Fig. 7). Only one phylotype, unclassified Comamonadaceae, (phylum Proteobacteria, class Betaproteobacteria, order Burkholderiales), was enriched in the CA contaminated site microcosms (Fig. 7a). Nine phylotypes were enriched in the uncontaminated site (Sediment H) microcosms, with 3 genus incertae sedis (phylum Verrucomicrobia, class Subdivision3) being dominant (Fig. 7b). There was a small (yet significant) enrichment of one phylotype, Pseudoxanthomonas (phylum Proteobacteria, class Gammaproteobacteria, order Xanthomonadales, family Xanthomonadaceae) in the KBS Soil 3 microcosms (Fig. 7c) and no enrichment of any phylotypes in the Soil F microcosms.
Phylotypes with a statistically significant difference (Welch’s t-test, two-sided, p < 0.05) between the 1,4-dioxane-degrading samples (n = 3) compared to the live controls (n = 3). The comparisons are shown for the Contaminated Site (CA) microcosms (a), the Sediment H microcosms (b), and KBS Soil 3 microcosms (c). No differences were noted for Soil F. The data points to the left of the dashed line indicate phylotypes more abundant in the 1,4-dioxane-degrading samples compared to the controls and those to the right indicate the reverse. For each, the x-axis has different scales
The sequencing data sets were also analyzed to determine the relative abundance of previously identified 1,4-dioxane degraders. It is important to note that the analysis could only be performed at the genus level; therefore, it is not possible to determine if particular species or strains are present. The 55 DNA extracts examined were from the fifth set of microcosms that illustrated 1,4-dioxane degradation (Soil F, Contaminated Site 10A, Sediment H and KBS Soil 3) and from 31 other microcosms that illustrated 1,4-dioxane biodegradation (from the other four experiments) (Table 1). Interestingly, all four soil-inoculated microcosms contained a high relative abundance (~5–45%) of the genus Rhodanobacter (Fig. 8, inserts). All four soils contained the genera Pseudomonas, Mycobacterium, and Acinetobacter, although the relative abundance values were much lower (< 0.15%) than Rhodanobacter (Fig. 8). The microcosms inoculated with KBS Soil 3 illustrated the largest number of previously identified 1,4-dioxane degraders (11 from 15 genera) (Fig. 8a). The Soils E, F, and G inoculated microcosms primarily contained 4, 8, and 5 of these genera. The well-studied Pseudonocardia and Rhodococcus genera were only common in KBS Soil 3 microcosms (Fig. 8a). The microcosms inoculated with contaminated site (CA) and uncontaminated site (Sediment H) sediments contained lower relative abundance values (< 0.04%) for Rhodanobacter (Fig. 9a, b). The CA contaminated site microcosms illustrated high levels of Pseudomonas (~ 20–70%) and lower levels of Rhodanobacter and Acinetobacter (Fig. 9a). Flavobacterium and Rhodococcus were only present in two from the twelve CA microcosms examined. The Sediment H microcosms contained lower levels (< 0.011%) of Pseudomonas, along with Rhodanobacter and four other genera (Fig. 9b). The KBS Soil 3 microcosms illustrated higher values for Rhodanobacter (in one case, Pseudomonas) and lower levels of seven other genera (Fig. 9c). Rhodococcus was present at low levels with all three inocula but only in a small number of microcosms. Pseudonocardia was absent from the CA microcosms, present at low levels in both a subset of the Sediment H microcosms and in all of the KBS Soil 3 microcosms. The average relative abundance values of the two enriched phylotypes discussed above (unclassified Comamonadaceae at 4.9% and 3 genus incertae sedis at 2.7%) were markedly higher than all of the previously identified dioxane degraders, except for Rhodanobacter and Pseudomonas.
Relative abundance (%) of genera previously associated with 1,4-dioxane biodegradation in replicates of 1,4-dioxane-degrading microcosms or live controls (c no 1,4-dioxane) inoculated with different soils. The inserts have a different axis to better illustrate the relative abundance of Rhodanobacter
Relative abundance (%) of genera previously associated with 1,4-dioxane biodegradation in replicates of 1,4-dioxane-degrading microcosms or live controls (c no 1,4-dioxane) inoculated with different sediments (a–c). The inserts have a different axis to better illustrate the relative abundance of Pseudomonas
Discussion
Although much is known about the microorganisms and functional genes associated with 1,4-dioxane degradation under aerobic conditions, limited research has addressed the susceptibility of 1,4-dioxane to biodegradation under anaerobic conditions (Rodriguez 2016; Shen et al. 2008; Steffan 2007). This research area is particularly important because 1,4-dioxane is a frequent co-contaminant at chlorinated solvent sites, which are commonly subject to remediation under highly reducing conditions (i.e., sulfate reducing or methanogenic). The current study examined the occurrence of 1,4-dioxane biodegradation using numerous inocula (soils/sediments from contaminated and uncontaminated sites) and a range of electron acceptor treatments (nitrate, iron/humic acid, sulfate, and no amendment). Additionally, the phylotypes enriched during 1,4-dioxane biodegradation were investigated.
As discussed above, previous research reported no 1,4-dioxane biodegradation over a range of redox conditions (Steffan 2007) with biodegradation occurring under iron-reducing conditions (Rodriguez 2016; Shen et al. 2008). Shen, Chen, and Pan (2008) observed anaerobic 1,4-dioxane biodegradation using sludge from a wastewater treatment plant. Three cases were highlighted using samples which were amended with (a) Fe(III) oxide (b) Fe(III)-EDTA, and (c) Fe(III)-EDTA and humic acid. The reductions in 1,4-dioxane concentrations increased from case (a) through (c) within the same 40-day period (Shen et al. 2008). However, the microorganisms responsible for 1,4-dioxane removal were not identified. Rodriguez (2016) reported that 1,4-dioxane appeared to degrade under iron-reducing conditions in the first 40 days. However, the concentration decreases could not be attributed to anaerobic biodegradation due to oxygen intrusion leading to oxidation of reduced iron (Rodriguez 2016). In the current work, the iron/EDTA/humic acid-amended microcosms did not result in any significant removal in the live samples compared to the abiotic controls. One hypothesis for the conflicting results (compared to Shen, Chan, and Pan 2008) could be the different sources of inocula (anaerobic wastewater sludge compared to soils and sediments in the current study) for the two studies, resulting in microbial communities with varying degradative abilities. It is important to note that in the current study, the contaminated site microcosms did not illustrate any degradation under these conditions (iron/EDTA/humic acid); therefore, it is unlikely that in situ iron amendments would be a successful 1,4-dioxane remediation approach.
An important limitation to the current study concerns the lack of data on electron acceptor concentrations. Given the long incubation periods, it is likely that electron acceptor depletion occurred and methanogenic conditions eventually dominated (which was confirmed in one case). Therefore, caution should be taken interpreting the impact of the addition of the various electron acceptors. However, it is reasonable to conclude that iron/EDTA/humic acid or sulfate amendments did not facilitate 1,4-dioxane biodegradation in the majority of cases and therefore these treatments would be ineffective in situ remediation techniques.
1,4-Dioxane biodegradation most commonly occurred in the nitrate-amended treatments and no electron acceptor treatments. Four and five (from seven) inocula sources examined in the first four sets of experiments produced significant reductions in 1,4-dioxane concentrations in the samples compared to the abiotic controls in the nitrate and no electron acceptor treatments, respectively. Further, both contaminated site sediments illustrated significant removal in the samples compared to the controls in the no electron acceptor treatment. However, the biodegradation rates were slow, with concentration reductions occurring over approximately 1 year. Taken together, the data indicate in situ 1,4-dioxane bioremediation under methanogenic conditions may be a feasible remediation approach if sufficient time is permissible for site cleanup.
CSIA has been previously applied to examine aerobic 1,4-dioxane biodegradation (Bennett et al. 2018; Gedalanga et al. 2016; Pornwongthong 2014). Recently, CSIA enabled the discrimination between the activities of the bacteria Rhodococcus rhodochrous ATCC 21198 and Pseudonocardia tetrahydrofuranoxidans K1 during 1,4-dioxane biodegradation (Bennett et al. 2018). Another study examined 1,4-dioxane biodegradation using CSIA in both pure and mixed cultures and found more positive δ13C and the δ2H values after biodegradation (Pornwongthong 2014). Also, CSIA provided additional evidence of 1,4-dioxane biodegradation in a groundwater plume that contained both metabolic and co-metabolic 1,4-dioxane-degrading bacteria (Gedalanga et al. 2016). Similar to the current work, CSIA has been adopted to characterize biodegradation of many organic compounds including chlorinated solvents, aromatic petroleum hydrocarbons, and fuel oxygenates (Hunkeler et al. 2008; Pooley et al. 2009; Qian et al. 2019; Rosell et al. 2007; Wang et al. 2004). The applicability of CSIA has also been explored in in situ biodegradation of organic contaminants such as hexachlorocyclohexane in contaminated soils (Qian et al. 2019).
To our knowledge, this is the first application of CSIA to investigate anaerobic 1,4-dioxane biodegradation. In two of the three cases examined, CSIA provided additional evidence for 1,4-dioxane biodegradation. In one case, the reduction in 1,4-dioxane in the samples comparing the controls was likely too low for the method to detect a significant 13C/12C enrichment. The changes in 13C/12C ratios in the current study were less than those reported previously for aerobic 1,4-dioxane biodegradation (Bennett et al. 2018); however, the changes in 1,4-dioxane concentrations were also less in the current study. The preliminary data set generated here indicates the value and limitations for 13C/12C analysis for documenting anaerobic 1,4-dioxane biodegradation. Further research is required to determine the value of measuring 2H/1H for generating evidence for the biodegradation of this chemical.
The greatest 1,4-dioxane decreases were observed in the microcosms inoculated with river sediment (Sediment H) and the contaminated sediment from Maine. Therefore, future research should examine the capabilities of other sediments to biodegrade this contaminant. The current research included experiments both with and without the addition of sodium lactate as an electron donor. Unfortunately, no clear trend was apparent as to the value of adding this substrate. Many chlorinated solvent sites are amended with an electron donor (e.g., vegetable soil, hydrogen release compound), so at least this particular substrate did not have any negative impact on 1,4-dioxane biodegradation. Additional research will be needed before any conclusions can be reached on the value of adding lactate to enhance biodegradation.
The analysis of the microbial communities from 1,4-dioxane-degrading microcosms indicated unclassified Comamonadaceae were obtaining a growth benefit in microcosms inoculated with sediment from the CA contaminated site. Comamonadaceae is a large and diverse family (over 100 species in 29 genera) that illustrates an impressive level of phenotypic diversity, including aerobic organotrophs, anaerobic denitrifiers, iron-reducing bacteria, hydrogen oxidizers, photoautotrophic and photoheterotrophic bacteria, and fermentative bacteria (Willems 2014). Additional research will be needed to determine which particular species within this family were enriched during 1,4-dioxane degradation.
For the uncontaminated sediment (Sediment H), nine phylotypes were enriched in the 1,4-dioxane-degrading microcosms compared to the controls with 3 genus incertae sedis (phylum Verrucomicrobia, class Subdivision3) being more highly enriched compared to the others (average 2.7% relative abundance). This phylotype also exhibited a high relative abundance (average 2.9%) in the MN contaminated sediment microcosms that illustrated 1,4-dioxane degradation. Verrucomicrobia is a rarely cultivated phylum that is generally poorly studied (Naumoff and Dedysh 2018; Tanaka et al. 2017). Again, additional research is needed to confirm the role of these phylotypes in 1,4-dioxane biodegradation.
One genus, Pseudoxanthomonas (phylum Proteobacteria, class Gammaproteobacteria, order Xanthomonadales, family Xanthomonadaceae) was enriched following 1,4-dioxane biodegradation in KBS Soil 3.This genus was recently linked to the biodegradation of anti-inflammatory drugs in the environment (Lu et al. 2019). The 1,4-dioxane-degrading communities also contained sequences that classified in the genera of previously reported 1,4-dioxane degraders. The soil microcosms were dominated primarily by Rhodanobacter with lower abundance values for Pseudomonas, Mycobacterium, and Acinetobacter. The sediment communities were dominated by Pseudomonas and Rhodanobacter. However, as stated above, it is important to note that the classification is only to the genus level; therefore, it is unknown if specific strains or species are present. Again, additional work is needed to determine the importance of these genera in 1,4-dioxane degradation under these experimental conditions.
Overall, the current study indicates 1,4-dioxane biodegradation under anaerobic and, likely highly reducing conditions, is feasible. Therefore, natural attenuation may be an appropriate cleanup technology at sites where time is not a constraint. This work also provides novel insights on microbial community obtaining a growth benefit from 1,4-dioxane under anaerobic conditions. Future research should target sediments from other contaminated sites along with additional CSIA measurements to confirm biodegradation.
References
Adams CD, Scanlan PA, Secrist ND (1994) Oxidation and biodegradability enhancement of 1,4-dioxane using hydrogen peroxide and ozone. Environ Sci Technol 28(11):1812–1818
Adamson DT, Mahendra S, Walker KL, Rauch SR, Sengupta S, Newell CJ (2014) A multisite survey to identify the scale of the 1,4-dioxane problem at contaminated groundwater sites. Environ Sci Tech Let 1(5):254–258
Adamson DT, Anderson RH, Mahendra S, Newell CJ (2015) Evidence of 1,4-dioxane attenuation at groundwater sites contaminated with chlorinated solvents and 1,4-dioxane. Environ Sci Technol 49(11):6510–6518. https://doi.org/10.1021/acs.est.5b00964
Bennett P, Hyman M, Smith C, El Mugammar H, Chu MY, Nickelsen M, Aravena R (2018) Enrichment with carbon-13 and deuterium during monooxygenase-mediated biodegradation of 1,4-dioxane. Environ Sci Tech Let 5(3):148–153. https://doi.org/10.1021/acs.estlett.7b00565
Bernhardt D, Diekmann H (1991) Degradation of dioxane, tetrahydrofuran and other cyclic ethers by an environmental Rhodococcus strain. Appl Microbiol Biot 36(1):120–123
Bhushan B, Halasz A, Hawari J (2006) Effect of iron(III), humic acids and anthraquinone-2,6-disulfonate on biodegradation of cyclic nitramines by Clostridium sp. EDB2. J Appl Microbiol 100(3):555–563. https://doi.org/10.1111/j.1365-2672.2005.02819.x
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. P Natl Acad Sci USA 108:4516–4522
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M, Gormley N, Gilbert JA, Smith G, Knight R (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6(8):1621–1624
Chen DZ, Jin XJ, Chen J, Ye JX, Jiang NX, Chen JM (2016) Intermediates and substrate interaction of 1,4-dioxane degradation by the effective metabolizer Xanthobacter flavus DT8. Int Biodeterior Biodegradation 106:133–140. https://doi.org/10.1016/j.ibiod.2015.09.018
Chu MYJ, Bennett P, Dolan M, Hyman M, Anderson R, Bodour A, Peacock A (2009) Using aerobic cometabolic biodegradation and groundwater recirculation to treat 1,4-dioxane and co-contaminants in a dilute plume. Paper presented at the tenth international conference on the remediation of chlorinated and recalcitrant compounds. Baltimore, Maryland
Cupples AM (2008) Real-time PCR quantification of Dehalococcoides populations: Methods and applications. J Microbiol Methods 72(1):1–11. https://doi.org/10.1016/j.mimet.2007.11.005
Cupples AM, Spormann AM, McCarty PL (2003) Growth of a Dehalococcoides-like microorganism on vinyl chloride and cis-dichloroethene as electron acceptors as determined by competitive PCR. Appl Environ Microbiol 69(2):953–959
Deng DY, Li F, Wu C, Li MY (2018) Synchronic biotransformation of 1,4-dioxane and 1,1-dichloroethylene by a gram-negative propanotroph Azoarcus sp DD4. Environ Sci Tech Let 5(8):526. https://doi.org/10.1021/acs.estlett.8b00312
DeRosa CT, Wilbur S, Holler J, Richter P, Stevens YW (1996) Health evaluation of 1,4-dioxane. Toxicol & Indust Health 12(1):1–43
Freedman DL, Gossett JM (1989) Biological reductive dechlorination of tetrachloroethylene and trichloroethylene to ethylene under methanogenic conditions. Appl Environ Microbiol 55(9):2144–2151
Gedalanga P, Madison A, Miao Y, Richards T, Hatton J, DiGuiseppi WH, Wilson J, Mahendra S (2016) A multiple lines of evidence framework to evaluate intrinsic biodegradation of 1,4-dioxane. Remediation 27(1):93–114. https://doi.org/10.1002/rem.21499
Hand S, Wang BX, Chu KH (2015) Biodegradation of 1,4-dioxane: effects of enzyme inducers and trichloroethylene. Sci Total Environ 520:154–159
He J, Ritalahti KM, Yang K-L, Koeningsberg SS, Löffler FE (2003) Detoxification of vinyl chloride to ethene coupled to growth of an anaerobic bacterium. Nature 424:62–65
Holliger C, Wohlfarth G, Diekert G (1998) Reductive dechlorination in the energy metabolism of anaerobic bacteria. FEMS Microbiol Rev 22(5):383–398
Huang HL, Shen DS, Li N, Shan D, Shentu JL, Zhou YY (2014) Biodegradation of 1,4-dioxane by a novel strain and its biodegradation pathway. Water Air Soil Poll 225:9
Hunkeler D, Meckenstock RU, Lollar BS, Schmidt TC, Wilson JT (2008) A guide for assessing biodegradation and source identification of organic ground water contaminants using compound specific isotope analysis (CSIA). USEPA, Oklahoma
Inoue D, Tsunoda T, Sawada K, Yamamoto N, Saito Y, Sei K, Ike M (2016) 1,4-Dioxane degradation potential of members of the genera Pseudonocardia and Rhodococcus. Biodegrad 27(4–6):277–286. https://doi.org/10.1007/s10532-016-9772-7
Inoue D, Tsunoda T, Yamamoto N, Ike M, Sei K (2018) 1,4-Dioxane degradation characteristics of Rhodococcus aetherivorans JCM 14343. Biodegrad 29(3):301–310. https://doi.org/10.1007/s10532-018-9832-2
Isaka K, Udagawa M, Sei K, Ike M (2016) Pilot test of biological removal of 1,4-dioxane from a chemical factory wastewater by gel carrier entrapping Afipia sp strain D1. J Hazard Mater 304:251–258. https://doi.org/10.1016/j.jhazmat.2015.10.066
Kampfer P, Kroppenstedt RM (2004) Pseudonocardia benzenivorans sp nov. Int J Syst Evol Micr 54:749–751. https://doi.org/10.1099/ijs.0.02825-0
Kim YM, Jeon JR, Murugesan K, Kim EJ, Chang YS (2009) Biodegradation of 1,4-dioxane and transformation of related cyclic compounds by a newly isolated Mycobacterium sp PH-06. Biodegrad 20(4):511–519
Kohlweyer U, Thiemer B, Schrader T, Andreesen JR (2000) Tetrahydrofuran degradation by a newly isolated culture of Pseudonocardia sp strain K1. Fems Microbiol Lett 186(2):301–306. https://doi.org/10.1111/j.1574-6968.2000.tb09121.x
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microb 79(17):5112–5120. https://doi.org/10.1128/Aem.01043-13
Lan RS, Smith CA, Hyman MR (2013) Oxidation of cyclic ethers by alkane-grown Mycobacterium vaccae JOB5. Remediation 23(4):23–42. https://doi.org/10.1002/rem.21364
Li MY, Fiorenza S, Chatham JR, Mahendra S, Alvarez PJJ (2010) 1,4-Dioxane biodegradation at low temperatures in Arctic groundwater samples. Water Res 44(9):2894–2900. https://doi.org/10.1016/j.watres.2010.02.007
Lippincott D, Streger SH, Schaefer CE, Hinkle J, Stormo J, Steffan RJ (2015) Bioaugmentation and propane biosparging for in situ biodegradation of 1,4-dioxane. Ground Water Monit R 35(2):81–92
Löffler FE, Yan J, Ritalahti KM, Adrian L, Edwards EA, Konstantinidis KT, Müller JA, Fullerton H, Zinder SH, Spormann AM (2013) Dehalococcoides mccartyi gen. Nov., sp. nov., obligately organohalide-respiring anaerobic bacteria relevant to halogen cycling and bioremediation, belong to a novel bacterial class, Dehalococcoidia classis nov., order Dehalococcoidales Ord. Nov. and family Dehalococcoidaceae fam. Nov., within the phylum Chloroflexi. Int J Syst Evol Microbiol 63(Pt 2):625–635
Lu ZD, Sun WJ, Li C, Ao XW, Yang C, Li SM (2019) Bioremoval of non-steroidal anti-inflammatory drugs by Pseudoxanthomonas sp. DIN-3 isolated from biological activated carbon process. Water Res 161:459–472. https://doi.org/10.1016/j.watres.2019.05.065
Mahendra S, Alvarez-Cohen L (2005) Pseudonocardia dioxanivorans sp nov., a novel actinomycete that grows on 1,4-dioxane. Int J Syst Evol Micr 55:593–598. https://doi.org/10.1099/ijs.0.63085-0
Mahendra S, Alvarez-Cohen L (2006) Kinetics of 1,4-dioxane biodegradation by monooxygenase-expressing bacteria. Environ Sci Technol 40(17):5435–5442. https://doi.org/10.1021/es060714v
Major DW, McMaster ML, Cox EE, Edwards EA, Dworatzek SM, Hendrickson ER, Starr MG, Payne JA, Buonamici LW (2002) Field demonstration of successful bioaugmentation to achieve dechlorination of tetrachloroethene to ethene. Environ Sci Technol 36(23):5106–5116
Masuda H (2009) Identification and characterization of monooxygenase enzymes involved in 1,4-dioxane degradation in Pseudonocardia sp. strain ENV478, Mycobacterium sp. strain ENV421, and Nocardia sp. strain ENV425. Rutgers the State University of new Jersey and University of Medicine and dentistry of New Jersey
Matsui R, Takagi K, Sakakibara F, Abe T, Shiiba K (2016) Identification and characterization of 1,4-dioxane-degrading microbe separated from surface seawater by the seawater-charcoal perfusion apparatus. Biodegrad 27(2–3):155–163. https://doi.org/10.1007/s10532-016-9763-8
Maymó-Gatell X, Chien Y-T, Gossett JM, Zinder SH (1997) Isolation of a bacterium that reductively dechlorinates tetrachloroethene to ethene. Science 276(6):1568–1571
Mohr T, Stickney J, DiGuiseppi W (2010) Environmental investigation and remediation: 1,4-Dioxane and other solvent stabilizers. CRC Press, Boca Raton
Naumoff DG, Dedysh SN (2018) Bacteria from poorly studied phyla as a potential source of new enzymes: beta-galactosidases from Planctomycetes and Verrucomicrobia. Microbiology 87(6):796–805. https://doi.org/10.1134/S0026261718060127
Parales RE, Adamus JE, White N, May HD (1994) Degradation of 1,4-dioxane by an Actinomycete in pure culture. Appl Environ Microb 60(12):4527–4530
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30(21):3123–3124
Pooley KE, Blessing M, Schmidt TC, Haderlein SB, MacQuarrie KTB, Prommer H (2009) Aerobic biodegradation of chlorinated ethenes in a fractured bedrock aquifer: quantitative assessment by compound-specific isotope analysis (CSIA) and reactive transport modeling. Environ Sci Technol 43(19):7458–7464. https://doi.org/10.1021/es900658n
Pornwongthong P (2014) Stable isotopic and molecular biological tools to validate biodegradation of 1,4-dioxane. University of California, Los Angeles
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J, Gloeckner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nuc Acids Res 35(21):7188–7196. https://doi.org/10.1093/nar/gkm864
Pugazhendi A, Banu JR, Dhavamani J, Yeom IT (2015) Biodegradation of 1,4-dioxane by Rhodanobacter AYS5 and the role of additional substrates. Ann Microbiol 65(4):2201–2208. https://doi.org/10.1007/s13213-015-1060-y
Qian Y, Chen K, Liu Y, Li J (2019) Assessment of hexachlorcyclohexane biodegradation in contaminated soil by compound-specific stable isotope analysis. Environ Pollut 254:113008
Rodriguez FJB (2016) Evaluation of 1,4-dioxane biodegradation under aerobic and anaerobic conditions. Clemson University
Rosell M, Häggblom MM, Richnow H-H (2007) Compound-specific isotope analysis (CSIA) to characterise degradation pathways and to quantify in-situ degradation of fuel oxygenates and other fuel-derived contaminants. In: Barceló D (ed) Fuel Oxygenates. Springer, Berlin Heidelberg, Berlin, Heidelberg, pp 99–119
Schloss PD (2009) A high-throughput DNA sequence aligner for microbial ecology studies. PLoS One 4(12). https://doi.org/10.1371/journal.pone.0008230
Schloss PD (2013) MiSeq SOP. PUblisher. http://www.mothur.org/wiki/MiSeq_SOP Accessed 15th Sept 2013
Sei K, Miyagaki K, Kakinoki T, Fukugasako K, Inoue D, Ike M (2013a) Isolation and characterization of bacterial strains that have high ability to degrade 1,4-dioxane as a sole carbon and energy source. Biodegrad 24(5):665–674
Sei K, Oyama M, Kakinoki T, Inoue D, Ike M (2013b) Isolation and characterization of tetrahydrofuran- degrading bacteria for 1,4-dioxane-containing wastewater treatment by co-metabolic degradation. J Water Environ Technol 11:11–19. https://doi.org/10.2965/jwet.2013.11
Shen W, Chen H, Pan S (2008) Anaerobic biodegradation of 1,4-dioxane by sludge enriched with iron-reducing microorganisms. Bioresour Technol 99(7):2483–2487. https://doi.org/10.1016/j.biortech.2007.04.054
Stefan MI, Bolton JR (1998) Mechanism of the degradation of 1,4-dioxane in dilute aqueous solution using the UV hydrogen peroxide process. Environ Sci Technol 32(11):1588–1595
Steffan RJ (2007) ER-1422: biodegradation of 1,4-dioxane. vol Final Report. SERDP and ESTCP
Steffan RJ, Vainberg S (2013) Production and handling of Dehalococcodies bioaugmentation cultures. In: Stroo HF, Leeson A, Ward CW (eds) Bioaugmentation for groundwater remediation. SERDP and ESTCP Remediation Technology. Springer, New York
Steffan RJ, McClay K, Vainberg S, Condee CW, Zhang DL (1997) Biodegradation of the gasoline oxygenates methyl tert-butyl ether, ethyl tert-butyl ether, and tert-amyl methyl ether by propane-oxidizing bacteria. Appl Environ Microb 63(11):4216–4222
Steffan RJ, McClay KR, Masuda H, Zylstra GJ (2007) Biodegradatdion of 1,4-dioxane. Strategic environmental Research and Development program: ER-1422 final report
Stringfellow WT, Alvarez-Cohen L (1999) Evaluating the relationship between the sorption of PAHs to bacterial biomass and biodegradation. Water res 33(11):2535–2544. https://doi.org/10.1016/S0043-1354(98)00497-7
Sun BZ, Ko K, Ramsay JA (2011) Biodegradation of 1,4-dioxane by a Flavobacterium. Biodegrad 22(3):651–659. https://doi.org/10.1007/s10532-010-9438-9
Sung Y, Ritalahti KM, Apkarian RP, Loffler FE (2006) Quantitative PCR confirms purity of strain GT, a novel trichloroethene-to-ethene-respiring Dehalococcoides isolate. Appl Environ Microbiol 72(3):1980–1987
Tanaka Y, Matsuzawa H, Tamaki H, Tagawa M, Toyama T, Kamagata Y, Mori K (2017) Isolation of novel bacteria including rarely cultivated phyla, Acidobacteria and Verrucomicrobia, from the roots of emergent plants by simple culturing method. Microbes Environ 32(3):288–292. https://doi.org/10.1264/jsme2.ME17027
Vainberg S, McClay K, Masuda H, Root D, Condee C, Zylstra GJ, Steffan RJ (2006) Biodegradation of ether pollutants by Pseudonocardia sp strain ENV478. Appl Environ Microb 72(8):5218–5224
Vainberg S, Condee CW, Steffan RJ (2009) Large-scale production of bacterial consortia for remediation of chlorinated solvent-contaminated groundwater. J Ind Microbiol Biot 36(9):1189–1197. https://doi.org/10.1007/S10295-009-0600-5
Wang Y, Huang Y, Huckins JN, Petty JD (2004) Compound-specific carbon and hydrogen isotope analysis of sub-parts per billion level waterborne petroleum hydrocarbons. Environ Sci Technol 38(13):3689–3697. https://doi.org/10.1021/es035470i
Whittenbury R, Phillips KC, Jf W (1970) Enrichment, isolation and some properties of methane-utilizing bacteria. J Gen Microbiol 61:–205, 18. https://doi.org/10.1099/00221287-61-2-205
Willems A (2014) The Family Comamonadaceae. In: Eugene Rosenberg (Editor-in-Chief) EFD, Stephen Lory, Erko Stackebrandt and Fabiano Thompson (Eds.) (ed) The prokaryotes: Alphaproteobacteria and betaproteobacteria. 4th edn. Springer-Verlag Berlin Heidelberg
Yamamoto N, Saito Y, Inoue D, Sei K, Ike M (2018) Characterization of newly isolated Pseudonocardia sp N23 with high 1,4-dioxane-degrading ability. J Biosci Bioeng 125(5):552–558. https://doi.org/10.1016/j.jbiosc.2017.12.005
Zenker MJ, Borden RC, Barlaz MA (2003) Occurrence and treatment of 1,4-dioxane in aqueous environments. Env Eng Sci 20(5):423–432
Acknowledgments
We would like to thank Dr. Dan Jones and Dr. Scott Smith at the Mass Spectrometry Laboratory at the Research Technology Support Facility (MSU) for 1,4-dioxane analytical method support and also to Dr. Anthony Danko (Naval Facilities Engineering Command) for providing the contaminated site sediments.
Funding
Support for this research was also provided by the NSF Long-term Ecological Research Program (DEB 1832042) at the Kellogg Biological Station and by Michigan State University AgBioResearch.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare no conflict of interest.
Ethical statement
This work is supported by a grant awarded to Dr. Cupples from Strategic Environmental Research and Development Program (SERDP). Contract Number: W912HQ-17-C-0006.
Human and animal rights and informed consent
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 30 kb)
Rights and permissions
About this article
Cite this article
Ramalingam, V., Cupples, A.M. Anaerobic 1,4-dioxane biodegradation and microbial community analysis in microcosms inoculated with soils or sediments and different electron acceptors. Appl Microbiol Biotechnol 104, 4155–4170 (2020). https://doi.org/10.1007/s00253-020-10512-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10512-3