Abstract
Strategies to enhance process performance of anaerobic digestion remain of key importance to promote wider usage of this technology for integrated resource recovery from organic waste streams. Continuous inoculation of the microbial community in the digester via the feedstock could be such a cost-effective strategy. Here, anaerobic digestion of fresh waste activated sludge (WAS) was compared with sterilized WAS in response to two common process disturbances, i.e. organic overloading and increasing levels of salts, to determine the importance of feedstock inoculation. A pulse in the organic loading rate severely impacted process stability of the digesters fed sterile WAS, with a 92 ± 45% decrease in methane production, compared to a 42 ± 31% increase in the digesters fed fresh WAS, relative to methane production before the pulse. Increasing salt pulses did not show a clear difference in process stability between the digesters fed fresh and sterile WAS, and process recovery was obtained even at the highest salt pulse of 25 g Na+ L−1. Feedstock sterilization through thermal pretreatment strongly impacted the microbial community in the digesters. In conclusion, feedstock thermal pretreatment strongly impacted anaerobic digestion process stability, due to feedstock inoculation and compositional modification.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The increasing environmental pollution and energy insecurity are pressing issues in our present society, which makes it important to look for integrated strategies that provide a solution to both issues. The fossil resources around the world are being depleted at a tremendous velocity, i.e. reaching a global total energy use of 1.64 × 105 TWh in 2017 to which renewable sources only contributed 25% (Enerdate 2018). This makes a transition towards sustainable resources for materials and energy of key importance to halt the increase in CO2 and other greenhouse gases emissions and related climate change (De Meester et al. 2012; Hagos et al. 2017). Anaerobic digestion (AD) is a microbial process that can be one of the possible solutions for this problem. The number of full-scale AD plants is still increasing nowadays, even though it already exists for decades (Scarlat et al. 2018). The success of AD lies in the fact that it does not only allow the stabilization of organic waste streams, but it is also a key technology for the recovery of energy (Demitry 2016). The methanogenic archaea are the most critical microorganisms in the AD process, because they are responsible for the production of the energy-dense CH4, which can be used for electricity and heat production via a combined heat and power unit (Holm-Nielsen et al. 2009). However, these methanogens are most sensitive to suboptimal process conditions, e.g. the presence of potential toxic compounds and organic overloading, and hence, are susceptible to process failure (Appels et al. 2008; Eftaxias et al. 2018). This sensitivity can in some cases prevent implementation of this technology (Demitry 2016).
One of the most common solution for preventing process failure is anaerobic co-digestion. Co-digestion can improve process stability, by (1) diluting potential inhibitory substances, such as ammonia, and (2) optimising the nutrient balance (Eftaxias et al. 2019; Mata-Alvarez et al. 2011; Mata-Alvarez et al. 2000). Nevertheless, process stabilization and optimization remains difficult (Hubenov et al. 2015; Kacprzak et al. 2010), and the unbalanced availability of suitable feedstocks, due to seasonal variation in supply and geographical reasons, is another problem that hampers an efficient co-digestion process (Hagos et al. 2017).
Another strategy to improve process stability is bioaugmentation, whereby a specific consortium, either enriched or isolated from similar systems or obtained from other ecosystems, is added to enhance the desired activity (De Vrieze and Verstraete 2016; Schauer-Gimenez et al. 2010). These microorganisms can improve the start-up of new digesters (Saravanane et al. 2001a; Saravanane et al. 2001b), reduce odour emissions (Duran et al. 2006; Tepe et al. 2008) and/or facilitate recovery of the reactors after an organic overload (Lynch et al. 1987). The disadvantage of this method is that it requires a certain volume of the biomass in the digester itself, 10% or more, to be replaced by a suitable consortium, whereby it is often not cost-efficient (De Vrieze and Verstraete 2016; Fotidis et al. 2014). Usually, only a temporal increase in CH4 production can be observed, due to wash-out of the bioaugmented microorganism and/or possible competition with the indigenous microorganisms (Schauer-Gimenez et al. 2010).
The addition of cations is also an alternative to solve the problem of process stability. The influent often contains a suboptimal ratio of the most common cations, i.e. Ca2+, Na+, K+ and Mg2+ (Anukam et al. 2019; Kugelman and McCarty 1965). An imbalance of this ratio can cause inhibition of the methanogens, which leads to process failure. To prevent this occurrence, the addition of other cations can restore the optimal balance in the feedstock, and this should result in optimal conditions for the microorganisms (Appels et al. 2008; Kugelman and McCarty 1965). The downside of this approach is that these cations are costly, i.e. the bulk market price of CaCl2 is around € 150–250 ton−1, while the bulk market price of MgCl2 ranges between € 250–300 ton−1 (www.icis.com, consulted June 2019), and their addition to the feedstock will result in an increase in the conductivity, which can have an overall negative impact on the microbial community. Hence, preventing AD process failure via an economically feasible approach is a key aspect that requires further investigation before such a prevention strategy can be applied at the full scale. Such a strategy will further solidify the use of AD, which will result in a higher sustainability in the use and recovery of the energy, combined with an environmental friendly way to treat organic waste streams (Chen et al. 2008).
The aim of this research was to tackle process failure in a cost-efficient manner by considering the importance of continuous inoculation or bioaugmentation through microorganisms in the feedstock. Two different process disturbances, i.e. organic overloading and increasing levels of salts, were considered, given the potential different impact of feedstock inoculation through thermal pretreatment. We hypothesized that the microorganisms present in the feedstock can (1) support processes resistance against disturbances, and (2) enhance recovery following process inhibition. The change in feedstock composition, due to the thermal pretreatment, was hypothesized to affect process stability, depending on the applied disturbance.
Materials and methods
Inoculum and feedstock
The inoculum for the operation of the lab-scale anaerobic digesters was obtained from the full-scale sludge digester of the Ossemeersen wastewater treatment plant (Ghent, Belgium) (Table 1). The waste activated sludge (WAS) that was used as feedstock was also obtained from the Ossemeersen in two separate batches that were used in the first and second stage of the experiment, respectively (Table 2). The WAS was stored at 4 °C to limit biodegradation during storage until use.
Experimental design and operation
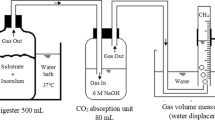
Six glass Schott bottles with a total volume of 1 L and a working volume of 800 mL were operated as lab-scale anaerobic digesters. The digesters were sealed with air-tight rubber stoppers, and connected to a water displacement system via gas-tight PVC tubing to monitor biogas production. The liquid in the water displacement system had a pH < 4.3 to avoid the CO2 in the biogas from dissolving. Gas samples were collected via a Laboport® vacuum pump (KNF Group International, Aartselaar, Belgium) and glass sampling tube of 250 mL (Glasgerätebau Ochs, Lenglern, Germany) for further analysis.
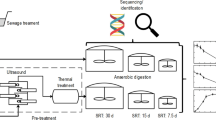
The digesters were operated in a semi-continuous stirred tank reactor mode, at mesophilic conditions in a temperature-controlled room at 34 ± 1 °C. The reactors were initially filled to a total volume of 800 mL with inoculum, which was diluted with tap water to obtain an initial VSS (volatile suspended solids) concentration of 10 g L−1. The digesters were fed with WAS manually three times per week by briefly opening the digesters. First, digestate was removed, after which new feedstock was added. Three digesters (biological replicates) were fed fresh WAS, while the three other digesters (also biological replicates) were fed WAS that was autoclaved (30 min at 121 °C, no stirring) twice (Table S1), using a Raypa autoclave (Beun - De Ronde Serlabo, Drogenbos, Belgium). This double autoclaving step, with an incubation period of 6–12 h at room temperature between the two autoclavation steps, was included to ensure complete sterilization of the sludge, including spore-forming bacteria.
During the start-up phase (day 0–26), the organic loading rate (OLR) was slowly increased from 0.94 to 3.75 g COD L−1 d−1 (chemical oxygen demand), and the hydraulic retention time was decreased from 40 to 10 days (Table 3). From day 27–126 on, an OLR of 3.75 g COD L−1 d−1 was maintained (WAS1), and between day 127–207, an OLR of 4.47 g COD L−1 d−1was used (WAS2). The WAS1 was diluted with tap water in a 1:1 ratio to avoid overloading, while the WAS2 was used as such, because of the lower VS content. Chemical-grade glycerol (86% glycerol, 14% water, Carl Roth GmbH, Karlsruhe, Germany) was added on day 48 as a single additional pulse of 1 g COD L−1 d−1 to provoke organic overloading. Different pulses of NaCl were added on day 118 (6.25 g Na+ L−1), day 160 (12.5 g Na+ L−1) and day 188 (25 g Na+ L−1).
The biogas production and composition were monitored three times a week, together with the pH and conductivity of each digester. Biogas production values were reported at standard temperature (273 K) and pressure (101,325 Pa) conditions (STP). The sulphate, phosphate, sodium, total ammonium and volatile fatty acids (VFA) concentrations were measured on weekly basis. Samples for microbial community analysis were taken on day 0 (inoculum and WAS), before addition of glycerol (day 48), and salt (day118, 160, 188), and at the end of the experiment (day 207) from each digester. The samples were stored at − 20 °C until DNA extraction was performed.
Microbial community analysis
The DNA extraction was carried out with the ZymoBIOMICS™ DNA Miniprep Kit (Zymo Research, Irvine, CA, USA), using a PowerLyzer® 24 Bench Top Bead-Based Homogenizer (MO BIO Laboratories, Inc., Carlsbad, CA, USA), in accordance with the instructions of the manufacturer. Agarose gel electrophoresis and PCR analysis were used to determine the quality of the DNA extracts. The PCR was carried out using the bacterial primers 341F (5′- CCTACGGGNGGCWGCAG) and 785Rmod (5′- GACTACHVGGGTATCTAAKCC), targeting the V3-V4 region of the 16S rRNA gene (Klindworth et al. 2013), following an in-house PCR protocol (Boon et al. 2002). After quality validation, the DNA extracts were sent to BaseClear B.V., Leiden, The Netherlands, for Illumina amplicon sequencing of the bacterial community, using the abovementioned primers, on the MiSeq platform with V3 chemistry. Amplicon sequencing and data processing are described in detail in SI (S2). Real-time PCR analysis was carried out to quantify total bacteria, the methanogenic orders Methanobacteriales and Methanomicrobiales and the methanogenic families Methanosaetaceae and Methanosarcinaceae, as described in SI (S3 and Table S2).
Statistical analyses
Following processing of the amplicon sequencing data, a table was generated with the relative abundances of the different OTUs (operational taxonomic units) and their taxonomic assignment (Supplementary file 2) of each sample. Statistical analyses were carried out in R version 3.3.1 (http://www.r-project.org) (R Development Core Team 2013). The similarity of the bacterial community in biological replicates was statistically validated (P < 0.05) using a repeated measures analysis of variance (ANOVA, aov function) (Connelly et al. 2017). Rescaling of the samples was carried out via the “common-scale” approach, by taking the proportions of all OTUs, multiplying them with the minimum sample size and rounding them to the nearest integer (McMurdie and Holmes 2014). Rarefaction curves (Fig. S1) were generated to estimate the degree of “coverage” of the bacterial community (Sanders 1968). The R packages vegan (Oksanen et al. 2016) and phyloseq (McMurdie and Holmes 2013) were used for bacterial community analysis.
Heatmaps were created at the phylum, order, class, family and OTU level using the pheatmap function (pheatmap package) for which the biological replicates were collated as described earlier (Connelly et al. 2017). Significant differences in microbial community composition between the digesters fed fresh and sterile WAS were identified by means of pair-wise permutational ANOVA (PERMANOVA) with Bonferroni correction, using the adonis function (vegan). The order based Hill’s numbers were used to evaluate the α-diversity in the different digesters. These Hill’s numbers represent richness (number of OTUs, H0), the exponential of the Shannon diversity index (H1) and the inverse Simpson index (H2). Significant differences in α-diversity between the digesters fed fresh and sterile WAS were determined with the Kruskal-Wallis rank sum test (kruskal.test function) (Hill 1973). Correlations between the sodium concentration and Hill’s numbers were determined using the Kendall’s tau correlation (cor.test function). The OTUs with a significant difference (P < 0.05) in relative abundance between the digesters with fresh or sterilized WAS as feedstock were determined with the DESeqDataSetFromMatrix function from the DESeq2 package (Love et al. 2014).
Analytical techniques
Total solids (TS), total suspended solids (TSS), volatile suspended solids (VSS), volatile solids (VS), Kjeldahl nitrogen (TKN) and total COD were measured according to standard methods (Greenberg et al. 1992). Soluble COD was measured using Nanocolor COD1500 or 15,000 test kits (Machery-Nagel, Düren, Germany), according to the manufacturer’s instructions. The concentrations of NH4+, Na+ and K+ were measured on a 761 Compact Ion Chromatograph (Metrohm, Herisau, Switzerland), which was equipped with a Metrosep C6-250/4.0 main column, a Metrosep C4 Guard/4.0 guard column and a conductivity detector. The eluent contained 1.7 mM HNO3 and 1.7 mM dipicolinic acid. Samples were centrifuged at 3000g for 3 min with a Labofuge 400 Heraeus centrifuge (Thermo Fisher Scientific Inc., Merelbeke, Belgium), filtered over a 0.22-μm filter (type PA-45/25, Macherey-Nagel, Germany) and diluted with Milli-Q water to reach the desired concentration range for quantification between 1 and 100 mg L−1. The pH was measured with a C532 pH meter, and conductivity was determined with a C833 conductivity meter (Consort, Turnhout, Belgium). The biogas composition was measured using a Compact Gas Chromatograph (Global Analyser Solutions, Breda, The Netherlands) (S5). The different VFA (C2-C8) were measured with a GC-2014 gas chromatograph (Shimadzu®, The Netherlands) (S6).
Data submission
The raw fastq files that served as a basis for the bacterial community analysis were deposited in the National Center for Biotechnology Information (NCBI) database (Accession number PRJNA540741).
Results
Digester performance
The impact of an organic pulse
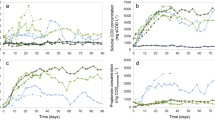
The initial start-up period during which the OLR was steadily increased and hydraulic retention time decreased (Table 3) over the first 27 days showed a steady increase in performance with increasing methane production rates (Fig. 1). For the next 21 days, steady-state conditions were maintained in both digesters from day 27–47 (Fig. 1), reflected in a stable methane production of 157 ± 33 and 316 ± 30 mL CH4 L−1 d−1 in the digesters fed fresh and sterile WAS, respectively. Thus, methane production in the digesters fed sterile WAS was, on average, a factor 2 higher than in the digesters fed fresh WAS during steady-state conditions. This higher methane production in the digesters fed sterile WAS can be related to their higher soluble COD content, which was a factor 7.3 ± 1.5 and 3.1 ± 1.5 higher for the sterile WAS1 and WAS2, respectively, compared to the fresh WAS samples (Table S1). A stable pH of 7.19 ± 0.08 and 7.38 ± 0.08 was observed in the digesters fed fresh and sterile WAS, respectively, from day 27–47. This relates with the residual VFA remaining below detection limit in the digesters fed fresh WAS and below 400 mg COD L−1 in the digesters fed sterile WAS (Fig. 2), confirming steady-state conditions. The addition of glycerol caused a differential effect in the digesters fed fresh and sterile WAS. A methane production rate of 224 ± 10 mL CH4 L−1 d−1 was observed in the digesters fed fresh WAS on day 53, which is a 42 ± 31% increase compared to the previous 21 days, because of the additional carbon source. In contrast, methane production decreased to only 25 ± 12 mL CH4 L−1 d−1 in the digesters fed sterile WAS, reflecting a 92 ± 45% decrease compared to the previous 21 days. This was also reflected in a pH value of 7.16 ± 0.03 for the digesters fed fresh WAS, while the pH in the digesters fed sterile WAS decreased to 6.68 ± 0.00. The recovery period was much shorter for the digesters fed fresh WAS in comparison with the digesters fed sterile WAS. Methane production and pH in the digesters fed sterile WAS reached the same values as prior to the glycerol pulse, only on day 71, while for the digesters fed fresh WAS, complete recovery was already the case on day 53 (Figs. 1 and 2). The residual VFA concentration also showed an increase following the glycerol pulse (Fig. 2b). The increase in VFA production was a factor two higher for the digesters fed sterile WAS than for the digesters fed fresh WAS, i.e. maximum concentrations of 4.15 ± 0.50 g COD L−1 (day 57) and 1.96 ± 0.13 g COD L−1 (day 50), respectively, were obtained (Fig. 2b). Overall, more time was needed for the recovery of the digesters fed sterile WAS than for the digesters fed fresh WAS.
Methane production in function of time in the digesters fed fresh and sterile waste activated sludge. The pulses of NaCl were added on day 118 (6.25 g Na+ L−1), day 160 (12.5 g Na+ L−1) and day 188 (25 g Na+ L−1). Average values of the biological replicates (n = 3) are presented, and the error bars represent standard deviations. The green line represents the organic loading rate (OLR)
pH (a) and total volatile fatty acid (VFA) concentration (b) in function of time in the digesters fed fresh and sterile waste activated sludge. The pulses of NaCl were added on day 118 (6.25 g Na+ L−1), day 160 (12.5 g Na+ L−1) and day 188 (25 g Na+ L−1). Average values of the biological replicates (n = 3) are presented, and the error bars represent standard deviations
The impact of increasing salt pulses
After the addition of glycerol and stabilization of the digesters to a new steady-state, NaCl was added as a novel disturbance. The increase in salt caused the increase in conductivity (Fig. S2). The first pulse of 6.25 g Na+ L−1 on day 118 had only a limited, though clear influence on the methane production, pH and VFA, as a limited decrease in methane production could be observed in both digesters from day 120 on (Fig. 1). A slight decrease in pH below the optimal value of 7.0 could be observed from day 120 on, but only in the digester fed fresh WAS (Fig. 2a). In contrast, the residual VFA reached higher values in the digesters fed sterile WAS (Fig. 2b), yet total VFA did not exceed 1 g COD L−1.
A second pulse of 12.5 g Na+ L−1 on day 160 resulted in a strong inhibition of the process, as reflected in a strong decrease in methane production and pH, and an increase in residual VFA. The methane production reached similar low values of 52 ± 5 and 58 ± 6 mL CH4 L−1 d−1 on day 162 for the digesters fed fresh and sterile WAS, respectively (Fig. 1). However, the relative decrease in methane production was higher for the digesters fed sterile WAS (84 ± 8%) compared to the digesters fed fresh WAS (76 ± 8%), due to the higher initial methane production in the digesters fed sterile WAS. The decrease in pH was stronger for the digesters fed fresh WAS (lowest value of 6.50 ± 0.02 on day 165) than for the digester fed sterile WAS (lowest value of 6.77 ± 0.07 on day 165), but the difference in pH between the steady-state values and the lowest value was the same for both digesters, i.e. about 0.5 units (Fig. 2a). The increase in residual VFA was similar for both digesters, with maximum values of 2.59 ± 0.01 and 2.48 ± 0.56 g COD L−1 in the digesters fed fresh and sterile WAS, respectively (Fig. 2b). Complete recovery of both digesters was prior to the addition of the final pulse on day 188.
The decrease in methane production and pH showed a similar trend as in response to the pulse of 12.5 g Na+ L−1 (Figs. 1 and 2a). The decrease in pH was again stronger for the digesters fed fresh WAS than for the digesters fed sterile WAS (Fig. 2a).The increase in residual VFA was in this case slightly higher in the digester fed sterile WAS (3.27 ± 0.44 g COD L−1 on day 195) than in the digester fed fresh WAS (2.52 ± 0.03 g COD L−1 on day 195) (Fig. 2b). However, similar to the previous NaCl pulses, process recovery took place, as can be observed in the increasing biogas and pH values (Figs. 1 and 2a) and decreasing residual VFA (Fig. 2b) towards the end of the experiment.
Microbial community composition and organization
Bacterial community
An average of 13,774 ± 2770 reads across all samples, representing 1488 ± 520 OTUs, were obtained per sample (including singletons) following amplicon sequencing. Removal of singletons and normalisation according to the common-scale approach resulted in an average of 9182 ± 327 reads and 648 ± 240 OTUs per sample. No significant differences (repeated measures ANOVA, P < 0.0001) could be detected in the bacterial community composition between the biological replicates.
The bacterial community composition strongly differed between the digesters fed fresh and sterile WAS (PERMANOVA, P = 0.0001), with 941 OTUs (21.7% of all OTUs, reflecting 86.6 ± 6.1% of the total relative abundance) showing a significant difference (DESeqDataSetFromMatrix, P < 0.05) in relative abundance, irrespective of the salt concentration or time point. This difference is reflected in the four main phyla, with the Actinobacteria (14.7 ± 2.0 vs. 4.7 ± 5.3%) and Proteobacteria (30.5 ± 1.0 vs. 2.6 ± 2.5%) showing a higher relative abundance in the digesters fed fresh WAS, and the Bacteroidetes (25.9 ± 5.6 vs. 16.3 ± 2.7%) and Firmicutes (44.0 ± 7.1 vs. 14.6 ± 4.5%) showing a higher relative abundance in the digesters fed sterile WAS (Fig. 3). This difference in bacterial community composition was also clear on the class, order, family and OTU level (Fig. S3-S6). The α-diversity analysis revealed a significantly higher richness H0 (P = 0.0025), and overall diversity H1 (P = 0.0015) and H2 (P = 0.0015) in the digesters fed fresh WAS, compared to the digesters fed sterile WAS (Fig. 4).
Heatmap showing the relative abundance of the bacterial community at the phylum level in the inoculum, two batches waste activated sludge (WAS1 and 2) and in the digesters fed fresh and sterile waste activated sludge on day 48, 118, 160, 188 and 207. Weighted average values of the biological replicates (n = 3) are presented. Only those phyla with at least 1% relative in one of the samples are included. The colour scale ranges from 0 (white) to 60% (red) relative abundance
Alpha diversity of the inoculum and in the digesters fed fresh and sterile waste activated sludge on day 48, 118, 160, 188 and 207. The three Hill order diversity numbers (a) H0 (richness, number of OTUs), (b) H1 (exponential value of the Shannon index) and (c) H2 (inverse Simpson index) were calculated based at the OTU level. Error bars represent standard deviations of the biological replicates (n = 3)
Even though the addition of glycerol and sodium impacted the overall methane production process, its direct effect on the bacterial community was limited. In total, 336 OTUs (7.7% of all OTUs) showed a significant (DESeqDataSetFromMatrix, P < 0.05) increase or decrease in function of the increasing sodium doses. Although the shift in relative abundance in response to the increased salinity could be detected for several dominant OTUs, such as OTU00004 (unclassified Rikenellaceae) and OTU00007 (unclassified Bacterium) (Fig. S6), this shift was not observed on the different phylogenetic levels (Fig. 3 and Figs. S3-S5). A differential impact of the increased sodium concentration on α-diversity could be observed between the digesters fed fresh and sterile WAS (Fig. 4). For the digesters fed fresh WAS, a significant negative correlation was observed between the sodium concentration and H0 (τ = − 0.47, P = 0.021), H1 (τ = − 0.69, P = 0.0006) and H2 (τ = − 0.67, P = 0.0010) diversity. In contrast, the increasing sodium concentration did not seem to impact the H0 ((τ = − 0.06 P = 0.77), H1 (τ = 0.11, P = 0.55) and H2 (τ = 0.31, P = 0.11) diversity in the digesters fed sterile WAS.
Methanogenic community
Real-time PCR analysis of the total bacteria and different methanogenic populations revealed a similar methanogens:bacteria ratio of 0.42 ± 0.23% across all digester samples, excluding the WAS samples. The WAS samples were characterized by a methanogens:bacteria ratio of 0.10 ± 0.01% for WAS1 and 0.07 ± 0.00% for WAS2. This indicates an overall strong dominance of the bacteria over the methanogens (at least a factor 100 higher absolute abundance) in the microbial community. No clear effect could be observed related to the feedstock sterilization, although the methanogens:bacteria ratio appeared to be higher in the digesters fed sterile WAS, especially at higher salt concentrations, though this does not necessarily imply a causal relation (Fig. S7). The methanogenic community in the two WAS batches was similar, with a dominance of the Methanosaetaceae, and this was also the case for the Inoculum (Figs. 5 and S8).
Relative abundance (%) of the Methanosaetaceae (blue,  ), Methanosarcinaceae (orange,
), Methanosarcinaceae (orange,  ), Methanobacteriales (grey,
), Methanobacteriales (grey,  ) and Methanomicrobiales (yellow,
) and Methanomicrobiales (yellow,  ) in the methanogenic community of the two batches waste activated sludge (WAS1 and 2), the inoculum and in the digesters fed fresh and sterile waste activated sludge on day 48, 118, 160, 188 and 207. Average values of the biological replicates (n = 3) are presented, and the error bars represent standard deviations
) in the methanogenic community of the two batches waste activated sludge (WAS1 and 2), the inoculum and in the digesters fed fresh and sterile waste activated sludge on day 48, 118, 160, 188 and 207. Average values of the biological replicates (n = 3) are presented, and the error bars represent standard deviations
Overall, a significant difference (PERMANOVA, P = 0.0003) in the methanogenic community profile could be observed between the digesters fed fresh and sterile WAS. After 48 days, prior to the addition of glycerol or sodium, a first divergence between the digesters with fresh and sterile WAS could be observed, with an increase in relative and absolute abundance of the Methanosarcinaceae in the digesters fed fresh WAS (13.6 ± 1.9%), compared with the digesters fed sterile WAS (2.6 ± 1.0%) (Fig. 5). The first and only pulse of glycerol, prior to the addition of sodium (day 118), did not provoke a clear effect on the methanogenic community in the digesters fed fresh WAS. In contrast, the Methanosarcinaceae showed a clear increase in relative abundance (88.0 ± 6.7%) on day 118 in the digesters fed sterile WAS, which was mainly due to the decrease in absolute abundance of the other methanogenic populations (Fig. S8). This increase in relative and absolute abundance of the Methanosarcinaceae was maintained in the digesters fed sterile WAS in response to the increasing sodium pulses (day 160, 188 and 207). In contrast, even though the Methanosarcinaceae also increased in relative and absolute abundance in the digesters fed fresh WAS in response to the increasing sodium pulses, the Methanomicrobiales became dominant on day 188 and especially day 207, reaching a relative abundance of 50.7 ± 2.6%. The Methanomicrobiales became also more dominant in the digesters fed sterile WAS, though the Methanosarcinaceae remained their dominancy.
Discussion
Thermal pretreatment of the feedstock, in this case waste activated sludge, resulted in a differentiating impact with respect to resistance to disturbances in anaerobic digestion. This can be attributed to (1) sterilization of the feedstock (no active microbial community) and (2) a change in composition. The microbial community in the feedstock positively contributed to resistance to organic overloading, in contrast to applying a sterile feedstock with increased soluble COD content, due to the pretreatment. However, a collapse in methane production in response to increasing salt pulses could not be prevented through feedstock inoculation, and process recovery took place irrespective of an active microbial community in and compositional modification of the feedstock. Feedstock thermal pretreatment strongly impacted the microbial community in the digesters in terms of composition and organization.
Different disturbances have a differentiating effect on process stability in relation to feedstock inoculation
The addition of glycerol and salt pulses showed a different impact on the activity of the microbial community. Glycerol that was added as an extra carbon source (Ma et al. 2008) can have two different effects on the process, which depends on the concentration, i.e. (1) an increase in methane production or (2) process failure due to overloading (Fountoulakis et al. 2010; González Arias et al. 2018; Holm-Nielsen et al. 2008). The addition of salt will normally inhibit microbial activity (Appels et al. 2008), but in this research, only a temporal effect on microbial activity and process stability could be observed, which can be attributed to the gradual decrease in salinity and subsequent reduction in residual VFA. When glycerol was added into the digesters, the recovery of the digester fed fresh WAS was faster than for the digesters fed sterile WAS. The reason behind this difference could be related to the presence of an active microbial community in the WAS (Lebiocka et al. 2018; Li et al. 2018), while this was not the case for the digesters fed sterile WAS. The presence of the active community in the fresh WAS, as observed in this research, could have contributed to the degradation of the organics, which led to an elevated methane production (Fountoulakis et al. 2010; González Arias et al. 2018). Additionally, the lower availability of readily biodegradable COD in the fresh WAS, in contrast to the sterile WAS, reduced the potential for VFA accumulation, indicating it was most likely a combined effect. Instead of an elevated methane production, a decrease was observed in the digesters fed sterile WAS, caused by the incremental organic loading because of glycerol addition. The microorganisms present in the digesters will quickly convert these organics into VFA, whereby the degradation of glycerol into VFA, mainly propionate and acetate, is faster than their subsequent conversion to methane. Hence, the methanogens could not keep up with the conversion rate of glycerol to VFA, which resulted in the accumulation of VFA (González Arias et al. 2018; Holm-Nielsen et al. 2008). This accumulation was confirmed by the pH decrease. Not only the addition of glycerol caused the accumulation of VFA, but also the sterilization of the feed, through thermal pretreatment, had a direct impact on the formation of VFA. The sterilization of the influent resulted in an increase in the soluble COD fraction, as also reported earlier (Papadimitriou 2010), mainly due to the increase in VFA. The COD that normally was not biodegradable in the influent was partially converted into readily biodegradable COD, mainly VFA, because of the sterilization process (Papadimitriou 2010; Tampio et al. 2014). Combined with the addition of glycerol, this resulted in an elevated organic loading, which led to the accumulation of VFA, and eventually to the inhibition of the microorganisms (Shi et al. 2017). Hence, the differential effect can be attributed to the presence of microbial community with a higher degree of activity in the fresh WAS and/or the increase in readily biodegradable COD in the sterile WAS.
Process recovery takes place irrespective of increasing salt pulses
With the addition of salt, no extra COD was introduced into the digesters, in contrast to the glycerol pulse, but salt is known as a common inhibitor of microbial activity in AD (Chen et al. 2008; Rinzema et al. 1988; Zhang et al. 2017a). The addition of salt commonly has a stronger impact, by causing a higher osmotic imbalance, on the methanogenic archaea, in comparison to the bacterial community (Rinzema et al. 1988; Wang et al. 2017; Zhang et al. 2017b). Despite elevated concentrations of NaCl, the bacterial community remained active, whereby they still converted the organics into VFA. In contrast, the salt concentration, especially at 12.5 and 25.0 g Na+ L−1, temporarily ceased methanogenic activity, resulting into an accumulation of residual VFA (Appels et al. 2008; Chen et al. 2008). The inhibition was only temporarily, because the methanogens managed to recover following the decrease in salt (Feijoo et al. 1995; Hierholtzer and Akunna 2014; Ismail et al. 2010), which is reflected in the decrease of VFA, and increases in pH and methane production. Because of the rather low organic loading rate and limited biodegradability of the WAS, the VFA accumulation potential was low, and residual VFA increase and pH decrease remained limited in both cases. Therefore, based on the results of both disturbances, i.e. glycerol and NaCl, it could be hypothesized that long-term inhibition in AD is not caused by the cations themselves, which only temporarily inhibit methanogenesis, but rather by the subsequent accumulation of VFA that permanently inhibit methanogenesis at higher organic loading rates (Mischopoulou et al. 2017; Zhang et al. 2017a). The similarity of both treatments proves that the microorganisms present in the feedstock and the increased soluble COD content due to the thermal pretreatment did not influence salts inhibition. This indicates that process inhibition in AD is a consequence of a tilting balance of VFA accumulation, which is due to inhibition of methanogenesis by another disturbance (Kugelman and McCarty 1965). The resulting pH decrease then further accelerates methanogenesis inhibition. This hypothesis remains to be confirmed with other disturbances.
Feedstock thermal pretreatment determines microbial community composition and organization
The microbial community in AD, containing representatives from both the bacterial and archaeal domains of life, is determined by different parameters central to the performance of the process, including total and free ammonia, temperature, salinity and pH (De Vrieze et al. 2018; De Vrieze et al. 2015b; Garcia and Angenent 2009; Westerholm and Schnürer 2019). This was also reflected in our study, as especially the addition of multiple salt pulses affected the bacterial and archaeal microbial community. However, this effect was clearly overshadowed by the differential impact of using fresh vs. sterile WAS as feedstock. This observation emphasises the importance of the feedstock composition, in this case related to the thermal pretreatment, with respect to the microbial community on two levels. First, even though certain operational parameters, such as temperature and solids retention time, can be set by the operator, the feedstock composition strongly affects the microbiome (Sundberg et al. 2013; Zhang et al. 2014), related to the abovementioned parameters. In our study, the impact of sterilization of the feed, which also changed the WAS chemical composition, on the process itself at stable conditions was obvious, related to the increase in methane production, pH and residual VFA at steady-state conditions. Second, the influx of microorganisms can be a key factor that influences or even steers the AD process (Kirkegaard et al. 2017; Shin et al. 2019), as observed in our study in response to a glycerol pulse. Sterilization through thermal pretreatment (autoclaving), in addition to killing of all microorganisms, also should degrade DNA molecules into small fragments (20–30 base pairs), though some larger fragments can remain behind (Esser et al. 2006), especially related to the complex WAS matrix. The WAS contains a living active microbial community with both bacteria and methanogenic archaea, as demonstrated earlier (De Vrieze et al. 2015a), but also dead/inactive microorganisms and/or free DNA could be present, thus influencing the microbial community profile (Kirkegaard et al. 2017). However, given (1) the presence of anaerobic sites in activated sludge units, reflected in diffuse methane emissions (Daelman et al. 2012), and (2) the importance of immigration in microbial community shaping (Sloan et al. 2006), the contribution of microorganisms in the WAS feedstock to process stability in AD is apparent, as confirmed by our results.
The impact of feedstock thermal pretreatment on the microbial community can be considered on different levels and across the domains of life. The strong difference in bacterial community composition already at the phylum level can be partially contributed directly to the WAS. For example, the digesters fed fresh WAS contained a higher relative abundance of Proteobacteria, thus reflecting the dominance of this phylum in the WAS itself. This was also reflected on the other phylogenetic levels. A similar result was obtained for the methanogenic community, because even though the absolute abundance of the methanogens in the WAS was up to a factor 10–100 lower than in the digesters, the digesters fed fresh WAS showed a similar profile as the WAS itself during the first sampling points. The strong dominance of the Methanosarcinaceae in the digesters fed sterile WAS could be due to their high growth rate at higher residual VFA (Conklin et al. 2006) as r-strategist in a more “open” niche (De Vrieze et al. 2017; Pianka 1970) in contrast to the digesters fed fresh WAS, where there was a continuous inflow of methanogens. The dominance of fewer taxa in the more open niche of the digesters fed sterile WAS was also reflected in their significantly lower diversity, compared to the digesters fed fresh WAS. As higher overall diversity can be linked to a higher functional redundancy (Briones and Raskin 2003; Langer et al. 2015; McMahon et al. 2007; Venkiteshwaran et al. 2016), this, at least partially, explains the higher resistance to the glycerol pulse of the digesters fed fresh WAS, compared to the digesters fed sterile WAS.
In conclusion, we demonstrated the importance of feedstock thermal pretreatment with respect to process performance and resistance towards disturbances in anaerobic digestion. A differential effect of feedstock thermal pretreatment was observed, as the microorganisms in the non-pretreated feedstock contributed to process stability in response to a glycerol pulse, while this was not the case for increasing salt pulses. Process recovery following the salt pulses took place irrespective of feedstock thermal pretreatment. Feedstock pretreatment strongly determined the bacterial and archaeal community composition and organization in the digesters, providing additional process stability security. This increased stability can be attributed to (1) feedstock inoculation, and (2) lower soluble COD concentrations than in the pretreated feedstock of which the composition was modified through the pretreatment. Overall, this opens the potential for feedstock inoculation through co-feedstocks rich in microorganisms, such as waste activated sludge and manure, for full-scale anaerobic digestion processes with limited feedstock inoculation (e.g. energy crops and crude glycerol) as a “right-of-the-shelve” strategy to enhance process stability.
References
Anthonisen AC, Loehr RC, Prakasam TBS, Srinath EG (1976) Inhibition of nitrification by ammonia and nitrous acid. J Water Pollut Control Fed 48(5):835–852
Anukam A, Mohammadi A, Naqvi M, Granström K (2019) A review of the chemistry of anaerobic digestion: methods of accelerating and optimizing process efficiency. Processes 7(8):504
Appels L, Baeyens J, Degrève J, Dewil R (2008) Principles and potential of the anaerobic digestion of waste-activated sludge. Prog Energy Combust 34(6):755–781. https://doi.org/10.1016/j.pecs.2008.06.002
Boon N, De Windt W, Verstraete W, Top EM (2002) Evaluation of nested PCR-DGGE (denaturing gradient gel electrophoresis) with group-specific 16S rRNA primers for the analysis of bacterial communities from different wastewater treatment plants. FEMS Microbiol Ecol 39(2):101–112. https://doi.org/10.1111/j.1574-6941.2002.tb00911.x
Briones A, Raskin L (2003) Diversity and dynamics of microbial communities in engineered environments and their implications for process stability. Curr Opin Biotechnol 14(3):270–276. https://doi.org/10.1016/s0958-1669(03)00065-x
Chen Y, Cheng JJ, Creamer KS (2008) Inhibition of anaerobic digestion process: a review. Bioresour Technol 99(10):4044–4064. https://doi.org/10.1016/j.biortech.2007.01.057
Conklin A, Stensel HD, Ferguson J (2006) Growth kinetics and competition between Methanosarcina and Methanosaeta in mesophilic anaerobic digestion. Water Environ Res 78(5):486–496. https://doi.org/10.2175/106143006x95393
Connelly S, Shin SG, Dillon RJ, Ijaz UZ, Quince C, Sloan WT, Collins G (2017) Bioreactor scalability: laboratory-scale bioreactor design influences performance, ecology, and community physiology in expanded granular sludge bed bioreactors. Front Microbiol 8:15. https://doi.org/10.3389/fmicb.2017.00664
Daelman MRJ, van Voorthuizen EM, van Dongen UGJM, Volcke EIP, van Loosdrecht MCM (2012) Methane emission during municipal wastewater treatment. Water Res 46(11):3657–3670. https://doi.org/10.1016/j.watres.2012.04.024
De Meester S, Demeyer J, Velghe F, Peene A, Van Langenhove H, Dewulf J (2012) The environmental sustainability of anaerobic digestion as a biomass valorization technology. Bioresour Technol 121:396–403. https://doi.org/10.1016/j.biortech.2012.06.109
De Vrieze J, Verstraete W (2016) Perspectives for microbial community composition in anaerobic digestion: from abundance and activity to connectivity. Environ Microbiol 18(9):2797–2809. https://doi.org/10.1111/1462-2920.13437
De Vrieze J, Raport L, Willems B, Verbrugge S, Volcke E, Meers E, Angenent LT, Boon N (2015a) Inoculum selection influences the biochemical methane potential of agro-industrial substrates. Microb Biotechnol 8(5):776–786. https://doi.org/10.1111/1751-7915.12268
De Vrieze J, Saunders AM, He Y, Fang J, Nielsen PH, Verstraete W, Boon N (2015b) Ammonia and temperature determine potential clustering in the anaerobic digestion microbiome. Water Res 75(0):312–323. https://doi.org/10.1016/j.watres.2015.02.025
De Vrieze J, Christiaens MER, Verstraete W (2017) The microbiome as engineering tool: manufacturing and trading between microorganisms. New Biotechnol 39(Part B):206–214. https://doi.org/10.1016/j.nbt.2017.07.001
De Vrieze J, Pinto AJ, Sloan WT, Ijaz UZ (2018) The active microbial community more accurately reflects the anaerobic digestion process: 16S rRNA (gene) sequencing as a predictive tool. Microbiome 6:13–13. https://doi.org/10.1186/s40168-018-0449-9
Demitry ME (2016) Anaerobic digestion process stability and the extension of the ADM1 for municipal sludge co- digested with bakery waste. All Graduate Theses and Dissertations. 4945. https://digitalcommons.usu.edu/etd/4945
Duran M, Tepe N, Yurtsever D, Punzi VL, Bruno C, Mehta RJ (2006) Bioaugmenting anaerobic digestion of biosolids with selected strains of Bacillus, Pseudomonas, and Actinomycetes species for increased methanogenesis and odor control. Appl Microbiol Biotechnol 73(4):960–966. https://doi.org/10.1007/s00253-006-0548-6
Eftaxias A, Diamantis V, Aivasidis A (2018) Anaerobic digestion of thermal pre-treated emulsified slaughterhouse wastes (TESW): effect of trace element limitation on process efficiency and sludge metabolic properties. Waste Manag 76:357–363. https://doi.org/10.1016/j.wasman.2018.02.032
Eftaxias A, Gkalimanis P, Koskinari MD, Koumara A, Diamantis V, Aivasidis A (2019) Use of diluted poultry manure as a low-cost emulsifier for anaerobic digestion of used cooking oil. Water Pract Technol 14(4):937–945. https://doi.org/10.2166/wpt.2019.074
Enerdate (2018) Global Energy Statistical Yearbook 2018
Esser DK-H, Marx DWH, Lisowsky PDT (2006) DNA decontamination: novel DNA-ExitusPlus™ in comparison with conventional reagents. BioTechniques 40(2):238–239. https://doi.org/10.2144/06402af01
Feijoo G, Soto M, Méndez R, Lema JM (1995) Sodium inhibition in the anaerobic digestion process: antagonism and adaptation phenomena. Enzym Microb Technol 17(2):180–188. https://doi.org/10.1016/0141-0229(94)00011-F
Fotidis IA, Wang H, Fiedel NR, Luo G, Karakashev DB, Angelidaki I (2014) Bioaugmentation as a solution to increase methane production from an ammonia-rich substrate. Environ Sci Technol 48(13):7669–7676. https://doi.org/10.1021/es5017075
Fountoulakis MS, Petousi I, Manios T (2010) Co-digestion of sewage sludge with glycerol to boost biogas production. Waste Manag 30(10):1849–1853. https://doi.org/10.1016/j.wasman.2010.04.011
Garcia ML, Angenent LT (2009) Interaction between temperature and ammonia in mesophilic digesters for animal waste treatment. Water Res 43(9):2373–2382. https://doi.org/10.1016/j.watres.2009.02.036
González Arias J, Sánchez ME, Gómez X (2018) Enhancing anaerobic digestion: the effect of carbon conductive materials, vol 4
Greenberg AE, Clesceri LS, Eaton AD (1992) Standard methods for the examination of water and wastewater. American Public Health Association Publications, Washington
Hagos K, Zong J, Li D, Liu C, Lu X (2017) Anaerobic co-digestion process for biogas production: progress, challenges and perspectives. Renew Sust Energ Rev 76:1485–1496. https://doi.org/10.1016/j.rser.2016.11.184
Hierholtzer A, Akunna JC (2014) Modelling start-up performance of anaerobic digestion of saline-rich macro-algae. Water Sci Technol 69(10):2059–2065. https://doi.org/10.2166/wst.2014.100
Hill MO (1973) Diversity and evenness: a unifying notation and its consequences. Ecology 54(2):427–432. https://doi.org/10.2307/1934352
Holm-Nielsen JB, Lomborg CJ, Oleskowicz-Popiel P, Esbensen KH (2008) On-line near infrared monitoring of glycerol-boosted anaerobic digestion processes: evaluation of process analytical technologies. Biotechnol Bioeng 99(2):302–313. https://doi.org/10.1002/bit.21571
Holm-Nielsen JB, Al Seadi T, Oleskowicz-Popiel P (2009) The future of anaerobic digestion and biogas utilization. Bioresour Technol 100(22):5478–5484. https://doi.org/10.1016/j.biortech.2008.12.046
Hubenov V, Mihaylova SN, Simeonov I (2015) Anaerobic co-digestion of waste fruits and vegetables and swine manure in a pilot-scale bioreactor. Bulg Chem Commun 47(3):788–792
Ismail SB, de La Parra CJ, Temmink H, van Lier JB (2010) Extracellular polymeric substances (EPS) in upflow anaerobic sludge blanket (UASB) reactors operated under high salinity conditions. Water Res 44(6):1909–1917. https://doi.org/10.1016/j.watres.2009.11.039
Kacprzak A, Krzystek L, Ledakowicz S (2010) Co-digestion of agricultural and industrial wastes. Chem Pap 64(2):127–131. https://doi.org/10.2478/s11696-009-0108-5
Kirkegaard RH, McIlroy SJ, Kristensen JM, Nierychlo M, Karst SM, Dueholm MS, Albertsen M, Nielsen PH (2017) The impact of immigration on microbial community composition in full-scale anaerobic digesters. Sci Rep 7:11. https://doi.org/10.1038/s41598-017-09303-0
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glockner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41(1):11. https://doi.org/10.1093/nar/gks808
Kugelman IJ, McCarty PL (1965) Cation toxicity and stimulation in anaerobic waste treatment. J Water Pollut Control Fed 37(1):97–116
Langer SG, Ahmed S, Einfalt D, Bengelsdorf FR, Kazda M (2015) Functionally redundant but dissimilar microbial communities within biogas reactors treating maize silage in co-fermentation with sugar beet silage. Microb Biotechnol 8(5):828–836. https://doi.org/10.1111/1751-7915.12308
Lebiocka M, Montusiewicz A, Cydzik-Kwiatkowska A (2018) Effect of bioaugmentation on biogas yields and kinetics in anaerobic digestion of sewage sludge. Int J Environ Res Public Health 15(8):1717. https://doi.org/10.3390/ijerph15081717
Li Y, Li L, Sun Y, Yuan Z (2018) Bioaugmentation strategy for enhancing anaerobic digestion of high C/N ratio feedstock with methanogenic enrichment culture. Bioresour Technol 261:188–195. https://doi.org/10.1016/j.biortech.2018.02.069
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550–550. https://doi.org/10.1186/s13059-014-0550-8
Lynch N, Daniels L, Parkin GF (1987) Bioaugmentation of stressed anaerobic filters with methanogenic enrichment cultures. In: Proceedings of the 42nd Industrial Waste Conference. (pp. 285-296). West Lafayette, Indiana: Purdue University
Ma J, Van Wambeke M, Carballa M, Verstraete W (2008) Improvement of the anaerobic treatment of potato processing wastewater in a UASB reactor by co-digestion with glycerol. Biotechnol Lett 30(5):861–867. https://doi.org/10.1007/s10529-007-9617-x
Mata-Alvarez J, Mace S, Llabres P (2000) Anaerobic digestion of organic solid wastes. An overview of research achievements and perspectives. Bioresour Technol 74(1):3–16. https://doi.org/10.1016/s0960-8524(00)00023-7
Mata-Alvarez J, Dosta J, Macé S, Astals S (2011) Codigestion of solid wastes: a review of its uses and perspectives including modeling. Crit Rev Biotechnol 31(2):99–111. https://doi.org/10.3109/07388551.2010.525496
McMahon KD, Martin HG, Hugenholtz P (2007) Integrating ecology into biotechnology. Curr Opin Biotechnol 18(3):287–292. https://doi.org/10.1016/j.copbio.2007.04.007
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8(4):e61217. https://doi.org/10.1371/journal.pone.0061217
McMurdie PJ, Holmes S (2014) Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput Biol 10(4):12. https://doi.org/10.1371/journal.pcbi.1003531
Mischopoulou M, Kalamaras SD, Naidis P, Kotsopoulos TA, Samaras P (2017) Start-up and steady-state results of a UASB reactor treating high pH baker's yeast molasses wastewater for methane production. J Chem Technol Biotechnol 92(4):775–780. https://doi.org/10.1002/jctb.5055
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara R.B., Simpson GL, Solymos P, Stevens MHH, Wagner H (2016) Vegan: community ecology package. R package version 2.3–4
Papadimitriou EK (2010) Hydrolysis of organic matter during autoclaving of commingled household waste. Waste Manag 30(4):572–582. https://doi.org/10.1016/j.wasman.2009.11.019
Pianka ER (1970) On r- and K-Selection. Am Nat 104(940):592–597
R Development Core Team (2013) R: a language and environment for statistical computing, 3.0 edn. R Foundation for Statistical Computing, Vienna
Rinzema A, van Lier J, Lettinga G (1988) Sodium inhibition of acetoclastic methanogens in granular sludge from a UASB reactor. Enzym Microb Technol 10(1):24–32. https://doi.org/10.1016/0141-0229(88)90094-4
Sanders HL (1968) Marine benthic diversity: a comparative study. Am Nat 102(925):243–282. https://doi.org/10.1086/282541
Saravanane R, Murthy DVS, Krishnaiah K (2001a) Bioaugmentation and anaerobic treatment of pharmaceutical effluent in fluidized bed reactor. J Environ Sci Health, Part A 36(5):779–791. https://doi.org/10.1081/ESE-100103760
Saravanane R, Murthy DVS, Krishnaiah K (2001b) Bioaugmentation and treatment of cephalexin drug-based pharmaceutical effluent in an upflow anaerobic fluidized bed system. Bioresour Technol 76(3):279–281. https://doi.org/10.1016/S0960-8524(00)00121-8
Scarlat N, Dallemand J-F, Fahl F (2018) Biogas: developments and perspectives in Europe. Renew Energy 129:457–472. https://doi.org/10.1016/j.renene.2018.03.006
Schauer-Gimenez AE, Zitomer DH, Maki JS, Struble CA (2010) Bioaugmentation for improved recovery of anaerobic digesters after toxicant exposure. Water Res 44(12):3555–3564. https://doi.org/10.1016/j.watres.2010.03.037
Shi X, Lin J, Zuo J, Li P, Li X, Guo X (2017) Effects of free ammonia on volatile fatty acid accumulation and process performance in the anaerobic digestion of two typical bio-wastes. J Environ Sci 55:49–57. https://doi.org/10.1016/j.jes.2016.07.006
Shin J, Cho S-K, Lee J, Hwang K, Chung JW, Jang H-N, Shin SG (2019) Performance and microbial community dynamics in anaerobic digestion of waste activated sludge: impact of immigration. Energies 12(3):573
Sloan WT, Lunn M, Woodcock S, Head IM, Nee S, Curtis TP (2006) Quantifying the roles of immigration and chance in shaping prokaryote community structure. Environ Microbiol 8(4):732–740. https://doi.org/10.1111/j.1462-2920.2005.00956.x
Sundberg C, Al-Soud WA, Larsson M, Alm E, Yekta SS, Svensson BH, Sørensen SJ, Karlsson A (2013) 454 pyrosequencing analyses of bacterial and archaeal richness in 21 full-scale biogas digesters. FEMS Microbiol Ecol 85(3):612–626. https://doi.org/10.1111/1574-6941.12148
Tampio E, Ervasti S, Paavola T, Heaven S, Banks C, Rintala J (2014) Anaerobic digestion of autoclaved and untreated food waste. Waste Manag 34(2):370–377. https://doi.org/10.1016/j.wasman.2013.10.024
Tepe N, Yurtsever D, Duran M, Mehta RJ, Bruno C, Punzi VL (2008) Odor control during post-digestion processing of biosolids through bioaugmentation of anaerobic digestion. Water Sci Technol 57(4):589–594. https://doi.org/10.2166/wst.2008.008
Venkiteshwaran K, Bocher B, Maki J, Zitomer D (2016) Relating anaerobic digestion microbial community and process function. Microbiol Insights 8(Suppl 2):37–44. https://doi.org/10.4137/MBI.S33593
Wang S, Hou X, Su H (2017) Exploration of the relationship between biogas production and microbial community under high salinity conditions. Sci Rep 7:1149. https://doi.org/10.1038/s41598-017-01298-y
Westerholm M, Schnürer A (2019) Microbial responses to different operating practices for biogas production systems biogas [working title]. IntechOpen
Zhang W, Werner JJ, Agler MT, Angenent LT (2014) Substrate type drives variation in reactor microbiomes of anaerobic digesters. Bioresour Technol 151:397–401. https://doi.org/10.1016/j.biortech.2013.10.004
Zhang Y, Alam MA, Kong X, Wang Z, Li L, Sun Y, Qi W (2017a) Effect of salinity on the microbial community and performance on anaerobic digestion of marine macroalgae. J Chem Technol Biotechnol 92(9):2392–2399. https://doi.org/10.1002/jctb.5246
Zhang Y, Li L, Kong X, Zhen F, Wang Z, Sun Y, Dong P, Lv P (2017b) Inhibition effect of sodium concentrations on the anaerobic digestion performance of Sargassum species. Energy Fuel 31(7):7101–7109. https://doi.org/10.1021/acs.energyfuels.7b00557
Acknowledgements
The authors would like to thank Tim Lacoere for his contribution to the molecular analyses and Aquafin for their assistance with sample collection. We thank Pieter Candry and Karel Folens for critically reading the manuscript.
Funding
Cindy Ka Y Law received support from the ELECTRA project “Electricity Driven Low Energy and Chemical Input Technology for Accelerated Bioremediation” financed by the H2020 of the European Commission under Grant number GA 826244. Jo De Vrieze is supported as a postdoctoral fellow by the Research Foundation Flanders (FWO-Vlaanderen).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Law, C., De Henau, R. & De Vrieze, J. Feedstock thermal pretreatment selectively steers process stability during the anaerobic digestion of waste activated sludge. Appl Microbiol Biotechnol 104, 3675–3686 (2020). https://doi.org/10.1007/s00253-020-10472-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10472-8