Abstract
The conversion of phenylglycinonitrile (2-aminophenylacetonitrile) by Escherichia coli strains was studied, which recombinantly expressed the arylacetonitrilase (NitA) from Pseudomonas fluorescens EBC191 and different nitrilase variants with altered reaction specificities. The whole-cell catalysts which formed the wild-type nitrilase converted (R,S)-phenylglycinonitrile preferentially to (S)-phenylglycine with a low degree of enantioselectivity. A recombinant strain which formed a variant of NitA produced mainly (S)-phenylglycine amide from (R,S)-phenylglycinonitrile and a second variant showed an almost complete enantioconversion and produced (R)-phenylglycine and left (S)-phenylglycinonitrile. The microbial-produced (S)-phenylglycinonitrile was used to study the chemical racemisation of (S)-phenylglycinonitrile at alkaline pH values in order to establish a dynamic kinetic resolution of the substrate. Subsequently, the conversion of (R,S)-phenylglycinonitrile by the whole-cell catalysts was studied at a pH of 10.8 which allowed a sufficient racemisation rate of phenylglycinonitrile. Surprisingly, under these conditions, strongly increased amounts of (S)-phenylglycine were formed by the recombinant E. coli cells expressing the amide-forming nitrilase variant. The aminopeptidase PepA from E. coli was identified by the construction of a deletion mutant and subsequent complementation as responsible amidase activity, which converted (S)-phenylglycine amide to (S)-phenylglycine.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Organic nitriles with the general formula R-CN can be enzymatically converted by nitrilases or nitrile hydratases. Nitrilases are hydrolytic enzymes found in many bacteria, fungi and plants which convert nitriles to the corresponding carboxylic acids and ammonia. Nitrile hydratases are almost exclusively found in bacteria and hydrate nitriles to amides. There is a considerable interest in chemistry in the utilisation of nitrilases and nitrile hydratases for chemo-, regio- or enantioselective biotransformation reactions (Bhalla et al. 2018; Gong et al. 2017; Martinková and Křen 2010; Prasad and Bhalla 2010; Singh et al. 2006).
A very interesting group of nitriles for biotransformation reactions are α-aminonitriles because they can be easily prepared by the Strecker reaction (or related syntheses) from aldehydes, ammonia and cyanide. The classical Strecker synthesis couples the formation of the aminonitriles in aqueous media with a subsequent acid hydrolysis of the aminonitriles to the corresponding acids. This is one of the most efficient and straightforward methods for the synthesis of various amino acids. The disadvantage of the Strecker reaction lies in the fact that the reaction results in the formation of racemic products. Therefore, significant interest exists in the development of asymmetric variations of the Strecker synthesis and organo- and metal catalysts have been described which induce enantioselectivity into the Strecker reaction (often using N-substituted aminonitriles as substrates) (Gröger 2003; Kouznetsov and Galvis 2018; Ooi et al. 2006; Surendra et al. 2006; Wang et al. 2011; Zuend et al. 2009).
There are some reports which describe the enantioselective conversion of racemic α-aminonitriles by nitrile hydratases and nitrilases (Wang and Lin 2001; Wegman et al. 2000, 2001; Hensel et al. 2002; Ewert et al. 2008). In the majority of these studies, the conversion of (substituted) phenylglycinonitrile(s) [2-aminophenylacetonitrile(s)] was studied as (substituted) (R)-phenylglycine(s) and (R)-phenylglycine amide(s) are used as building blocks in the synthesis of various β-lactam antibiotics and are therefore important industrial products (Bruggink 2001). Thus, different nitrile hydratases forming Rhodococcus, Pantoea and Klebsiella strains were found which convert racemic phenylglycinonitrile to (S)-phenylglycine with rather high enantioselectivities. The conversion of phenylglycinonitrile was caused in all analysed cases by the action of nitrile hydratases demonstrating only a low degree of enantioselectivity in combination with highly (S) specific amidases. Thus, these processes can lead to highly enantioenriched (R)-(D)-phenylglycine amide and (S)-(L)(+)-phenylglycine (Wang and Lin 2001; Wegman et al. 2000, 2001; Hensel et al. 2002; Ewert et al. 2008).
More recently, a coupling of a nitrile hydratase with two different amidases with opposite enantiopreferences together with an α-amino-ε-caprolactam racemase has been described which allowed the formation of almost enantiopure small aliphatic (R)- or (S)-amino acids via dynamic kinetic resolution processes (Yasukawa et al. 2011).
There are also some reports about the biotransformation of α-aminonitriles by nitrilases. Initially, it was found that nitrilases from different Rhodococcus, Acinetobacter and Aspergillus strains could convert aliphatic 2-aminonitriles and also phenylglycinonitrile with some enantioselectivity to the corresponding amino acids (Bhalla et al. 1992; Choi and Goo 1986; Macadam and Knowles 1985). Later, Chaplin et al. (2004) described the screening of a large set of recombinant nitrilases for the conversion of (R,S)-phenylglycinonitrile and identified one nitrilase which could be used for the synthesis of (R)-phenylglycine with a fairly good ee-value (91%). Recently, the application of this enzyme for the conversion of (R,S)-phenylglycinonitrile has been further optimised (Kawahara and Asano 2018). Furthermore, the application of a recombinant E. coli strain was reported, which expressed an enantioselective nitrilase from Sphingomonas wittichii and hydrolysed (R,S)-phenylglycinonitrile to (R)-phenylglycine with ee-values up to 97% (Qiu et al. 2014).
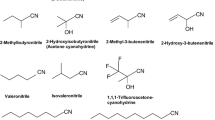
We have analysed throughout the last years a nitrilase from Pseudomonas fluorescens EBC191 (Baum et al. 2012; Brunner et al. 2018; Fernandes et al. 2006; Heinemann et al. 2003; Kiziak et al. 2005, 2007; Kiziak and Stolz 2009; Layh et al. 1992; Sosedov et al. 2010; Sosedov and Stolz 2014, 2015). This enzyme converts various phenylacetonitriles, such as 2-hydroxyphenylacetonitrile (mandelonitrile), 2-acetoxyphenylacetonitrile (O-acetylmandelonitrile), 2-methylphenylacetonitrile (2-phenylpropionitrile) and also α,α-disubstituted phenylacetonitriles with moderate enantioselectivities to the corresponding α-(or α,α-)(di)substituted carboxylic acids and varying amounts of the corresponding amides. Also several heterocyclic and aliphatic nitriles are converted by this enzyme (Baum et al. 2012; Brunner et al. 2018; Fernandes et al. 2006; Heinemann et al. 2003; Kiziak et al. 2005; Layh et al. 1992). In the course of the initial screening of the range of substrates which are converted by the nitrilase from P. fluorescens EBC191, it was found that the enzyme also converted phenylglycinonitrile (Kiziak et al. 2005). During the last years, several variants of this nitrilase have been generated which demonstrate significantly changed reaction- and enantiospecificities with mandelonitrile, 2-phenylpropionitrile and also fumarodinitrile (Brunner et al. 2018; Kiziak et al. 2007; Kiziak and Stolz 2009; Sosedov et al. 2010; Sosedov and Stolz 2014, 2015). There is still significant interest to find an efficient way to convert racemic phenylglycinonitriles produced by the Strecker synthesis to stoichiometric amounts of enantiopure phenylglycines or phenylglycine amides (this has been once claimed as one of the “holy grails of biocatalysis”; Sheldon and van Rantwijk 2004). Therefore, in the present study, different nitrilase variants were analysed for the conversion of (R,S)-phenylglycinonitrile.
Materials and methods
Bacterial strains, plasmids and culture conditions
Escherichia coli JM109 was used as standard host strain. The construction of plasmid pIK9, which encodes for the wild-type nitrilase (NitA) from Pseudomonas fluorescens EBC191, has been described previously (Kiziak et al. 2005). Plasmids pIK9/Ala165Phe and pIK9/Trp188Lys were generated from plasmid pIK9 by site-directed mutagenesis (Kiziak and Stolz 2009; Sosedov and Stolz 2015). The nitrilase genes were expressed in these constructs under the control of a rhamnose-inducible promoter (Stumpp et al. 2000).
The cultivation of the recombinant strains and the conditions for the expression of NitA and its variants have been described before (Kiziak et al. 2005).
Biotransformations
The conversion of (R,S)-phenylglycinonitrile by resting cells was routinely determined in reaction mixtures (1 ml each) containing 100 mM of the relevant buffers, 10 mM nitrile and an appropriate amount of cells (~optical density (OD600nm) 0.5–5). The stock solutions of (R,S)-phenylglycinonitrile (100 mM) were prepared in methanol. The reaction mixtures were incubated in a thermomixer at 750 rpm at 23 °C. After different time intervals, samples (100 μl each) were taken and the reactions were stopped by the addition of 1 M HCl (10 μl). The samples were centrifuged at 15,000g for 2 min and the supernatants were analysed using high-pressure liquid chromatography (HPLC).
Preparation of (S)-phenylglycinonitrile
Resting cells of E. coli JM109(pIK9/Ala165Phe) were suspended in 9 ml K-phosphate buffer (100 mM, pH 6.0) to an OD600nm of about 5. The cell suspension was transferred to a 250-ml Erlenmeyer flask and the reaction started by adding 1 ml of 100 mM (R,S)-phenylglycinonitrile (in methanol). The reaction mixture was incubated at room temperature and stirred at 500 rpm. After 30 min, the reaction was stopped by acidification with 1 ml of 1 M HCl. The cells were removed by centrifugation (20,000g, 2 min) and the supernatant lyophilised overnight. The remaining powder was resuspended directly before the experiments in 1 ml of water to give a solution of (S)-phenylglycinonitrile.
Racemisation of (S)-phenylglycinonitrile under alkaline conditions
A preparation of (S)-phenylglycinonitrile (prepared as described above) was mixed with 100 mM Na-carbonate buffer (pH 10.8), the pH adjusted with 1 M NaOH and the reaction mixtures (150 μl each) were placed in 300 μl HPLC vials. The vials were placed in a HPLC autosampler at 23 °C. Aliquots (5 μl each) were withdrawn every 30 min using the HPLC injector and analysed directly by chiral HPLC.
Analytical methods
The synthesis and turn over of (R,S)-phenylglycinonitrile was routinely quantified by HPLC. Two different HPLC methods were applied.
For the achiral HPLC analysis, a solvent system consisting of 5% (v/v) methanol, 1 g/l hexane-1-sulfonic acid sodium salt and 0.1% (v/v) formic acid in H2O was applied. A Pro C18 AQ column (Trentec Analysetechnik, Rutesheim, Germany) at a flow rate of 1 ml/min was used. The average retention times (Rt) under these conditions were as follows: benzaldehyde Rt = 11.5 min, mandelonitrile Rt = 8.8 min, phenylglycinonitrile Rt = 15.1 min, phenylglycine Rt = 3.0 min and phenylglycine amide Rt = 6.5 min.
The separation of the enantiomers of phenylglycinonitrile, phenylglycine amide and phenylglycine was achieved on a Crownpak CR(+) column (Daicel). The mobile phase (flow rate 1 ml/min) consisted of 16.3 g/l of 70% perchloric acid in water. The column was at 35 °C. For the enantiomers of phenylglycine amide and phenylglycine, stable purchasable reference compounds were available. In the described chromatographic system, the average retention times were for (R)- and (S)-phenylglycine amide Rt = 3.2 and 4.4 min, and for (R)- and (S)-phenylglycine Rt = 3.7 and 5.6 min, respectively.
The commercially available racemic (R,S)-phenylglycinonitrile showed under the same conditions two signals at Rt = 6.7 and 7.3 min. It was deduced that the signal with the shorter retention time was due to (R)-phenylglycinonitrile because in the cases of phenylglycine amide and phenylglycine the respective (R)-enantiomers, both showed the shorter retention times. This interpretation was confirmed during the biotransformation experiments as the preferred formation of the (R)- or (S)-enantiomers of phenylglycine amide and/or phenylglycine could always be correlated in an enantioconservative way with the preferred disappearance of the signal for the respective enantiomer of phenylglycinonitrile.
The separated compounds were detected at a detector wavelength of 210 nm.
Construction of a ΔpepA variant of E. coli JM109
The deletion of pepA was conducted by the method of Datsenko and Wanner (2000). The sequence coding for a kanamycin resistance and flanked by FRT sites was amplified from plasmid pC01kanFRTa using the primers pepA-DW-fw (5′CTATCTGTAGCCACCGCCGTTGTCTTTAAGATTCAGGAGCGTAGTGCT TGTGTAGGCTGGAGCTGCTTCG-3′) and pepA-DW-rev (5′-GATAAGGCGTTCACG CCGCATCCGGCAATAACAGCCTTGCCTGACGCAACATATGAATATCCTCCTTAGTTC-3′). These primers added about 50-bp regions lying up- and downstream of the pepA sequence (underlined above). The PCR product was digested by DpnI and gel-purified. Then E. coli JM109(pKD46) was grown for 5 h at 30 °C in LB medium supplemented with ampicillin (100 μg/ml) and arabinose (0.2%). After 5 h, 5 ml of cells were harvested and washed twice with ice-cold glycerin (10% (v/v) and finally resuspended in ice-cold glycerin. Subsequently, in a final volume of 100 μl, 500 ng of the gel-purified PCR product was added and the cells transformed by electroporation. Next, 1 ml of LB medium was added to the cell suspension and the cells were incubated for 2 h at 37 °C. Finally, 250 μl of the cell suspension was plated on LB agar supplemented with kanamycin (50 μg/ml) and incubated overnight at 37 °C.
Colonies growing on LB-kanamycin agar were tested for the deletion of pepA by PCR using primers pepA-fw (5′-CACAGAAGGACGTGCATTAC-3′) and pepA-rev (5′-CATGATGGTTTGCTGGAAGG-3′) followed by digestion of the PCR product by EcoRV. Positive colonies (showing the correct EcoRV digestion pattern) were further tested for the ability to grow on LB-ampicillin agar plates. Positive E. coli clones, which were ΔpepA and grew on LB-kanamycin but not on LB-ampicillin (thus no longer bearing pKD46), were purified on LB-kanamycin agar plates. Single colonies of E. coli JM109 ΔpepA::kan were grown in liquid LB medium with kanamycin for 4 h at 37 °C and 180 rpm. The cells were harvested, washed twice with ice-cold glycerin and electroporated with pCP20 (Cherepanov and Wackernagel 1995), resuspended in LB and incubated at 30 °C for 2 h. Then the cells were plated onto LB agar plates with ampicillin and the plates incubated at 30 °C overnight. The colonies were streaked on LB agar and incubated at 42 °C overnight in order to cure pCP20. The resulting colonies were tested for growth on ampicillin and kanamycin (at 30 °C) and the mutation verified by PCR using primers pepA-fw and pepA-rev.
Cloning of the gene coding for aminopeptidase A
The coding sequence of pepA was amplified from E. coli JM109 using primers pepA-NdeI-fw (AAAAACATATGGAGTTTAGTGTAAAAAG) and pepA-BamHI-rv (AAAAAGGATCCTTACTCTTCGCCGTTAAAC), which introduced NdeI and BamHI restriction sites up- and downstream of the pepA sequence (restriction sites underlined). The PCR product and pJOE2775 (Stumpp et al. 2000) were cut with NdeI and BamHI, the fragments gel-purified and finally ligated, yielding plasmid pEN15.
Chemicals
(R, S)-phenylglycinonitrile, (R)- and (S)-phenylglycine amide and (R)- and (S)-phenylglycine were supplied by Sigma-Aldrich, Fluorochem and Fluka, respectively.
Results
Conversion of (R,S)-phenylglycinonitrile at pH 6 by variants of the nitrilase from Pseudomonas fluorescens EBC191
It was previously shown that resting cells of E. coli JM109(pIK9), which synthesize the wild-type nitrilase (NitA) from P. fluorescens EBC191, converted racemic phenylglycinonitrile to phenylglycine and phenylglycine amide and that preferentially the (S)-enantiomers of the products were formed (Kiziak et al. 2005). During the last years, several variants of NitA have been described, which convert mandelonitrile (2-hydroxyphenylacetonitrile) and 2-phenylpropionitrile (2-methylphenylacetonitrile) with significantly changed enantioselectivities and/or degrees of amide formation (Kiziak et al. 2007; Kiziak and Stolz 2009; Sosedov et al. 2010; Sosedov and Stolz 2014, 2015). Therefore, it was analysed if these variants also demonstrated changes in the reaction- and enantiospecificity during the conversion of phenylglycinonitrile. The enzyme variants NitA(Ala165Phe) and NitA(Trp188Lys) were tested which converted (R,S)-mandelonitrile either to increased amounts of (R)-mandelic acid or to increased amounts of (S)-mandeloamide, respectively (Kiziak and Stolz 2009; Sosedov and Stolz 2015).
Resting cells of E. coli JM109(pIK9), E. coli JM109(pIK9/Ala165Phe) and E. coli JM109(pIK9/Trp188Lys) were incubated in 100 mM K-phosphate buffer (pH 6.0) with (R,S)-phenylglycinonitrile (10 mM) and the reactions analysed by HPLC. Thus, it was found that the resting cells of E. coli JM109(pIK9), E. coli JM109(pIK9/Ala165Phe) and E. coli JM109(pIK9/Trp188Lys) converted (R,S)-phenylglycinonitrile with specific activities of 0.88, 0.34 and 0.32 U/mg of protein, respectively. The analyses of the reactions by chiral HPLC demonstrated that E. coli JM109(pIK9) and E. coli JM109(pIK9/Trp188Lys) preferentially converted the (S)-enantiomer of phenylglycinonitrile. In contrast, E. coli JM109(pIK9/Ala165Phe) preferred the (R)-enantiomer of this substrate.
In order to compare the reaction- and enantiospecificities of the three types of NitA, the reactions were each analysed after about 30% conversion of the racemic substrate. Thus, for the wild-type enzyme, the previous results were confirmed and it was found that phenylglycinonitrile was mainly converted to (S)-phenylglycine with a low degree of enantioselectivity (ee = 42%). In addition, about 8% of phenylglycine amide with a surplus of the (S)-enantiomer (ee = 75%) was formed (Table 1).
The variant E. coli JM109(pIK9/Trp188Lys) produced almost exclusively phenylglycine amide from (R,S)-phenylglycinonitrile. In comparison to the wild type, there was an increase in the ee-value of the formed (S)-phenylglycine amide observed (Table 1).
The variant E. coli JM109(pIK9/Ala165Phe) demonstrated a decreased degree of amide formation compared to the wild type and formed about 3% phenylglycine amide from (R,S)-phenylglycinonitrile. In contrast to the wild-type enzyme, the variant NitA(Ala165Phe) almost exclusively formed (R)-phenylglycine (Fig. 1). Thus, this point mutation resulted in an almost complete stereoinversion of the reaction (Table 1).
Conversion of (R,S)-phenylglycinonitrile by resting cells of E. coli JM109(pIK9/Ala165Phe). The reaction mixture contained in 10 ml resting cells E. coli JM109(pIK9/Ala165Phe) (OD600nm = 5) in 100 mM K-phosphate buffer (pH 6). The reaction was started by the addition of (R,S)-phenylglycinonitrile (10 mM). The concentrations of (R)-phenylglycinonitrile (▲), (S)-phenylglycinonitrile (△), (R)-phenylglycine amide (▼), (S)-phenylglycine amide (▽), (R)-phenylglycine (■) and (S)-phenylglycine (□) were monitored by chiral HPLC
Generation and racemisation of (S)-phenylglycinonitrile
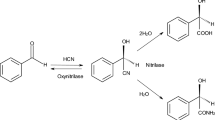
It is well known that α-aminonitriles, such as phenylglycinonitrile, are in a chemical equilibrium with the corresponding aldehyde, cyanide and ammonia. This has several implications for the intended biotransformation. On the one hand, racemic phenylglycinonitrile (and other α-aminonitriles) can be easily prepared by the Strecker reaction from an aldehyde, cyanide and ammonia. On the other hand, α-aminonitriles are notoriously unstable in aqueous systems. This might be contraproductive as it results in the decomposition of the substrate (Fig. 2). In the case of chiral substrates, the reversibility of the reaction also offers the potential to racemise the aminonitriles. This could allow a dynamic kinetic resolution and thus a complete conversion of a racemic α-aminonitrile to a single enantiomer of a product.
The racemisation of (R,S)-phenylglycinonitrile in aqueous solutions has been studied in some detail by Chaplin et al. (2004). They demonstrated that phenylglycinonitrile racemises under alkaline conditions and that at a pH of 10.5, a solution of (S)-phenylglycinonitrile completely racemised within 60 min. In contrast, at pH 8 and pH 9.5, the respective values were 13 h and 4 h, respectively.
In order to define the pH value required for a combination of a chemical racemisation with an enzymatic reaction in some more detail, a sample of (S)-phenylglycinonitrile was obtained by conversion of (R,S)-phenylglycinonitrile with E. coli JM109(pIK9/Ala165Phe). The formation of highly enriched (R)-phenylglycine by this strain correlated with the intermediate accumulation of (S)-phenylglycinonitrile (Fig. 2) and this process could be used to obtain enantioenriched preparations of (S)-phenylglycinonitrile (see “Materials and methods”). The (S)-phenylglycinonitrile (1.4 mM) obtained was incubated at pH 9.5, 10.0, 10.5 pH 9.5, 10.0, and 10.5 in 100 mM Na2CO3 buffer and the racemisation of (S)-phenylglycinonitrile analysed by chiral HPLC. Thus, at pH 9.5, 10.0 and 10.5 for (S)-phenylglycinonitrile, half-lives were determined of approximately 360 min, 180 min and 130 min, respectively (Fig. 3). Furthermore, it became evident that in addition to the racemisation of (S)-phenylglycinonitrile to the (R)-enantiomer, also a decomposition of phenylglycinonitrile to benzaldehyde took place and that the increase in the pH value also resulted in an increased formation of benzaldehyde.
Racemisation of (S)-phenylglycinonitrile at different pH values. The (S)-phenylglycinonitrile (PGN) was prepared with resting cells of E. coli JM109 (pIK9/Ala165Phe) as described in the “Materials and methods” section. This preparation was mixed (1:1, v:v) with 100 mM Na2CO3/NaHCO3 buffer (pH 10.8) and brought to the indicated pH values by the addition of NaOH. The changes in the ee-values of (S)-phenylglycinonitrile were determined at pH, 9.5 (●), pH 10.0 (■) and pH 10.5 (▲) by HPLC
Conversion of (R,S)-phenylglycinonitrile under alkaline conditions by E. coli JM109(pIK9)
The experiments described above suggested that in order to achieve a dynamic kinetic resolution, it is necessary to perform the biotransformation of (R,S)-phenylglycinonitrile at pH values > 10. Therefore, resting cells of E. coli JM109(pIK9) were incubated with (R,S)-phenylglycinonitrile at pH 6, 9, 10 and 10.8 and the decrease in the substrate concentration analysed by HPLC. This demonstrated that the nitrilase activities in the cells were still active at pH values > 10, but that the activities significantly decreased at increasing pH values. Thus, at a pH of 10.8, about 30% of the activity at pH 6 were recovered. The two variants E. coli JM109(pIK9/Ala165Phe) and E. coli JM109(pIK9/Trp188Lys) showed similar pH-dependent decreases in the relative reaction rates (Fig. 4).
Conversion of (R,S)-phenylglycinonitrile by resting cells of E. coli JM109(pIK9), E. coli JM109 (pIK9/Ala165Phe) and E. coli JM109 (pIK9/Trp188Lys) at different pH values. The reaction mixtures contained in a final volume of 1 ml resting cells of E. coli JM109 (pIK9) (OD600nm = 0.1 for reactions at pH 6.0 and OD600nm = 0.25 at pH 10.8), E. coli JM109(pIK9/Ala165Phe) or E. coli JM109(pIK9/Trp188Lys) (OD600nm = 1.0 at pH = 6.0 and OD600nm = 2.5 at pH 10.8) in 100 mM of K-phosphate- (pH 6.0) or 100 mM Na-carbonate buffer (pH 10.8). The reactions were started by the addition of 0.1 ml of 100 mM (R,S)-phenylglycinonitrile (in methanol). The concentrations of (R)-phenylglycinonitrile (▲), (S)-phenylglycinonitrile (Δ), (R)-phenylglycine (■), (S)-phenylglycine (□), (R)-phenylglycine amide (▼) and (S)-phenylglycine amide (∇) were determined by chiral HPLC
The incubation of the amide-forming strain E. coli JM109(pIK9/Trp188Lys) with (R,S)-phenylglycinonitrile at alkaline pH values showed a peculiar increase in the (S)-phenylglycine concentration at alkaline conditions. No (S)-phenylglycine could be detected under slightly acidic (pH 6.0) conditions. In contrast, at pH 10.8, significantly increased amounts of (S)-phenylglycine amide and (S)-phenylglycine were formed (Fig. 4).
Subsequently, the stability of the whole-cell catalysts in the relevant buffer systems was tested. Therefore, resting cells of E. coli JM109(pIK9) were incubated without substrate in the buffers (K-phosphate buffer pH 7, Na-carbonate buffer pH 10 and pH 10.8, all 100 mM) for 0, 30, 60 and 90 min before the substrate was added and the reaction rates were determined. This indicated that the nitrilase activity in the cells was at pH values between pH 7 and pH 10 stable for several hours, but showed at pH 10.8 only a half-life of approximately 2 h.
Conversion of phenylglycine amide by E. coli JM109
The experiments with E. coli JM109(pIK9/Trp188Lys) demonstrated that at pH 6 from (R,S)-phenylglycinonitrile, almost stoichiometric amounts of phenylglycine amide were formed. In contrast, at alkaline pH values, significant amounts of phenylglycine were detected (Fig. 4). This could be due either to a chemical hydrolysis of phenylglycine amide under alkaline conditions or to a biological activity. The analysis of the biotransformation by chiral HPLC demonstrated that the phenylglycine which was formed under alkaline conditions was almost pure (S)-phenylglycine. Furthermore, it was found that only (S)-phenylglycine amide disappeared and the small amount of (R)-phenylglycine amide that was formed by the nitrilase variant was not converted. This indicated the presence of an enantioselective amidase activity in the whole-cell catalyst. Therefore, resting cells of E. coli JM109 and E. coli JM109(pJOE2775) (“vector control”; Stumpp et al. 2000) were incubated with (R,S)-phenylglycine amide in different buffer systems. These experiments demonstrated that the reference strains converted (R,S)-phenylglycine amide to (S)-phenylglycine and that the reaction rates increased with increasing pH values (Fig. 5).
Enantioselective hydrolysis of (R)- and (S)-phenylglycine amide by resting cells of E. coli JM109. The reaction mixtures contained in 10 ml resting cells of E. coli JM109 (OD600nm = 10) in a 100 mM K-phosphate buffer (pH 6) or b 100 mM Na2CO3-buffer (pH 10.8). The reactions were started by the addition of (R)- and (S)-phenylglycine amide (5 mM each). The concentrations of (R)-phenylglycine amide (▼), (S)-phenylglycine amide (∇∇), (R)-phenylglycine (■) and (S)-phenylglycine (□) were measured by chiral HPLC
Enantioselective hydrolysis of (R,S)-phenylglycine amide by the aminopeptidase PepA from E. coli
The hydrolysis of (R,S)-phenylglycine amide to (S)-phenylglycine by enantioselective amidases has been previously described for other bacteria, e.g. Pseudomonas putida, Ochrobactrum anthropi or Mycobacterium neoaurum (Hermes et al. 1993a, 1994; Sonke et al. 2005). In the case of P. putida ATCC 12633, this enzyme has been identified as an l-aminopeptidase which enantioselectively hydrolyses several dipeptides and l-amino acid amides (Hermes et al. 1993b). Three major aminopeptidases have been described for E. coli, peptidase A (PepA), peptidase B (PepB) and peptidase N (PepN) (Vogt 1970; Bhosale et al. 2010). Therefore, a BLAST search was performed with the sequence of the amidase/aminopeptidase from P. putida ATCC 12633 (CAA09054.1) against genomic sequences from E. coli. Thus, it was found that the highest scores were found with PepA (synonymously named leucyl aminopeptidase) from E. coli (53% sequence identity). Therefore, a ΔpepA variant of E. coli JM109 was generated by site-directed mutagenesis using the method described by Datsenko and Wanner (2000) as described in the “Materials and methods” section.
The variant E. coli JM109ΔpepA was transformed with plasmid pJOE2775 (=“vector control”) and plasmid pEN15 which carried the pepA gene cloned into pJOE2775. In the following, the turn over of (R,S)-phenylglycine amide by both strains was analysed. Thus, it was found that E. coli JM109ΔpepA(pJOE2775) showed only some residual activity for (S)-phenylglycine amide. In contrast, the complemented variant E. coli JM109ΔpepA(pEN15) converted (S)-phenylglycine amide with much higher reaction rates (Fig. 6). These experiments clearly demonstrated that PepA from E. coli is able to hydrolyze (S)-phenylglycine amide.
Conversion of (R,S)-phenylglycine amide by resting cells of E. coli JM109 ΔpepA(pJOE2775) and E. coli JM109 ΔpepA(pEN15). The reaction mixtures contained in 1 ml resting cells of aE. coli JM109 Δ pepA (pJOE2775) (OD600nm = 5.0) and bE. coli JM109 ΔpepA(pEN15) (OD600nm = 1.0) in 100 mM Na-carbonate buffer (pH 10.8). The reactions were started by the addition of 5 mM (R)-phenylglycine amide plus 5 mM (S)-phenylglycine amide. The concentrations of (R)-phenylglycine (■), (S)-phenylglycine (□), (R)-phenylglycine amide (▼) and of (S)-phenylglycine amide (∇∇) were measured by chiral HPLC
Discussion
The previous studies with the nitrilase from P. fluorescens EBC191 showed that by simple point mutations, variants could be generated which demonstrated with mandelonitrile and 2-phenylpropionitrile significantly changed reaction- and enantiospecificities (Kiziak et al. 2007; Kiziak and Stolz 2009; Sosedov et al. 2010; Sosedov and Stolz 2015). In the present study, it was shown that the observations previously made with nitriles which carry in the α-position a hydroxyl-substituent (as in mandelonitrile) or a methyl-substituent (as in 2-phenylpropionitrile) can also be applied to substrates carrying an amino-function at the respective position. Thus, the nitrilase variant Ala165Phe (which carries in contrast to the wild-type enzyme a large substituent in direct neighbourhood to the catalytical active cysteine residue) demonstrated in comparison to the wild-type enzyme with mandelonitrile, a significantly enhanced (R)-selectivity and with 2-phenylpropionitrile and phenylglycinonitrile even an enantioconversion and formed instead of the (S)-enantiomers the respective (R)-enantiomers of the acids. Similarly, it was previously shown that the nitrilase variant Trp188Lys formed significantly increased amounts of amides from mandelonitrile and 2-phenylpropionitrile and in the present study it was established that this variant also forms a large surplus of phenylglycine amide from phenylglycinonitrile. The similar behaviour of the variants during the conversion of substrates carrying substituents with rather different polarity at the chiral centre suggested that the reaction- and enantiospecificity of these variants (and also of the nitrilase wild type) are largely governed by the size of the substituent.
The conversion of phenylglycine amide to (S)-phenylglycine has been found before with some Gram-positive and Gram-negative bacteria, such as Mycobacterium neoaurum, Ochrobactrum anthropi and Pseudomonas putida (Hermes et al. 1993a, 1993b, 1994; Sonke et al. 2005), but to the best of our knowledge, this reaction has not been previously described for E. coli. In the present study, it was clearly shown by the construction of deletion mutants and the subsequent complementation of these mutants that the aminopeptidase PepA from E. coli is able to hydrolyse (R,S)-phenylglycine amide to (S)-phenylglycine with a high degree of enantioselectivity. This aminopeptidase has been previously purified and characterised by Vogt (1970). It was found that the enzyme hydrolysed several di- and tripeptides and that tripeptides such as Met-Leu-Gly or Met-Gln-Gly were the preferred substrates. Furthermore, the aminopeptidase was optimally active in the pH range from pH 9–11 and almost inactive at pH 7. The extraordinary pH optimum of the aminopeptidase PepA added an interesting new twist to the conversion of (R,S)-phenylglycine amide by genetically modified E. coli strains with the ability to convert (R,S)-phenylglycinonitrile to phenylglycine amide. Thus, it was possible to produce with E. coli JM109(pIK9/Trp188Lys) at pH 6 (S)-phenylglycine amide with ee-values > 90%, but the simple change in the pH to pH> 10 allows the synthesis of (S)-phenylglycine even with a higher enantiomeric excess as the aminopeptidase activity with its pronounced (S)-selectivity further “polishes” the enantiocomposition of the product.
References
Baum S, Williamson DS, Sewell T, Stolz A (2012) Conversion of sterically demanding α,α-disubstituted phenylacetonitriles by the arylacetonitrilase from Pseudomonas fluorescens EBC191. Appl Environ Microbiol 78:48–57
Bhalla TC, Miura A, Wakamoto A, Ohba Y, Furuhashi K (1992) Asymmetric hydrolysis of α-aminonitriles to optically active amino acids by a nitrilase of Rhodococcus rhodochrous PA-34. Appl Microbiol Biotechnol 37:184–190
Bhalla TC, Kumar V, Kumar V, Thakur N, Savitri (2018) Nitrile metabolizing enzymes in biocatalysis and biotransformation. Appl Biochem Biotechnol 185:925–946
Bhosale M, Pande S, Kumar A, Kairamkonda S, Nandi D (2010) Characterization of two M17 family members in Escherichia coli, peptidase A and peptidase B. Biochem Biophys Res Comm 395:76–81
Bruggink A (2001) Synthesis of β-lactam antibiotics: chemistry, biocatalysis & process integration. Kluwer Academic Publishers, Dordrecht
Brunner S, Eppinger E, Fischer S, Gröning J, Stolz A (2018) Conversion of aliphatic nitriles by the arylacetonitrilase from Pseudomonas fluorescens EBC191. World J Microb Biotech 34:91
Chaplin JA, Levin MD, Morgan B, Farid N, Li J, Zhu Z, McQuaid J, Nicholson LW, Rand CA, Burk MJ (2004) Chemoenzymatic approaches to the dynamic kinetic asymmetric synthesis of aromatic amino acids. Tetrahedron Asym 15:2793–2796
Cherepanov PP, Wackernagel W (1995) Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene 158:9–14
Choi YY, Goo YM (1986) Hydrolysis of the nitrile group in α-aminophenylacetonitrile by nitrilase; development of a new biotechnology for stereospecific production of S-α-phenylglycine. Arch Pharm Res 9:45–47
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645
Ewert C, Lutz-Wahl S, Fischer L (2008) Enantioselective conversion of α-arylnitriles by Klebsiella oxytoca. Tetrahedron Asym 19:2573–2578
Fernandes BCM, Mateo C, Kiziak C, Wacker J, Chmura A, van Rantwijk F, Stolz A, Sheldon RA (2006) Nitrile hydratase activity of a recombinant nitrilase. Adv Synth Catal 348:2597–2603
Gong J-S, Shi J-S, Lu Z-M, Li H, Zhou Z-M, Xu ZH (2017) Nitrile-converting enzymes as a tool to improve biocatalysis in organic synthesis: recent insights and promises. Crit Rev Biotechnol 37:69–81
Gröger H (2003) Catalytic enantioselective Strecker reactions and analogous syntheses. Chem Rev 103:2795–2827
Heinemann U, Kiziak C, Zibek S, Layh N, Schmidt M, Griengl H, Stolz A (2003) Conversion of aliphatic 2-acetoxynitriles by nitrile-hydrolysing bacteria. Appl Microbiol Biotechnol 63:274–281
Hensel M, Lutz-Wahl S, Fischer L (2002) Stereoselective hydration of (RS)-phenylglycine nitrile by new whole cell biocatalysts. Tetrahedron Asym 13:2629–2633
Hermes HFM, Croes LM, Peeters WPH, Peters PJH, Dijkhuizen L (1993a) Metabolism of amino acid amides in Pseudomonas putida ATCC 12633. Appl Microbiol Biotechnol 40:519–525
Hermes HFM, Sonke T, Peters PJH, van Balken JAM, Kamphuis J, Dijkhuizen L, Meijer EM (1993b) Purification and characterization of an L-aminopeptidase from Pseudomonas putida ATCC 12633. Appl Environ Microbiol 59:4330–4334
Hermes HFM, Tandler RF, Sonke T, Dijkhuizen L, Meijer EM (1994) Purification and characterization of an L-amino amidase from Mycobacterium neoaurum ATCC 25795. Appl Environ Microbiol 60:153–159
Kawahara N, Asano Y (2018) Chemoenzymatic method for enantioselective synthesis of (R)-2-phenylglycine and (R)-2-phenylglycine amide from benzaldehyde and KCN using difference of enzyme affinity to the enantiomers. ChemCatChem 10:5000–5006
Kiziak C, Stolz A (2009) Identification of amino acid residues which are responsible for the enantioselectivity and amide formation capacity of the arylacetonitrilase from Pseudomonas fluorescens EBC191. Appl Environ Microbiol 75:5592–5599
Kiziak C, Conradt D, Stolz A, Mattes R, Klein J (2005) Nitrilase from Pseudomonas fluorescens EBC191: cloning and heterologous expression of the gene and biochemical characterization of the recombinant enzyme. Microbiology 151:3639–3648
Kiziak C, Klein J, Stolz A (2007) Influence of different carboxyterminal mutations on the substrate-, reaction-, and enantiospecifity of the arylacetonitrilase from Pseudomonas fluorescens EBC191. Prot Eng Design Sel 20:385–396
Kouznetsov VV, Galvis CEP (2018) Strecker reaction and α-amino nitriles: recent advances in their chemistry, synthesis, and biological properties. Tetrahedron 74:773–810
Layh N, Stolz A, Förster S, Effenberger F, Knackmuss H-J (1992) Enantioselective hydrolysis of O-acetylmandelonitrile to O-acetylmandelic acid by bacterial nitrilases. Arch Microbiol 158:405–411
Macadam AM, Knowles CJ (1985) The stereospecific bioconversion of α-aminopropionitrile to L-alanine by an immobilised bacterium isolated from soil. Biotechnol Lett 7:865–870
Martinková L, Křen V (2010) Biotransformations with nitrilases. Curr Opin Chem Biol 14:130–137
Ooi T, Uematsu Y, Maruoka K (2006) Asymmetric Strecker reaction of aldimines using aqueous potassium cyanide by phase-transfer catalysis of chiral quaternary ammonium salts with a tetranaphthyl backbone. J Am Chem Soc 128:2548–2549
Prasad S, Bhalla TC (2010) Nitrile hydratases (NHases): at the interface of academia and industry. Biotechnol Adv 28:725–741
Qiu J, Su E, Wang W, Wei D (2014) High yield synthesis of D-phenylglycine and its derivatives by nitrilase mediated dynamic kinetic resolution in aqueous 1-octanol biphasic system. Tetrahedron Lett 55:1448–1451
Sheldon RA, van Rantwijk F (2004) Biocatalysis for sustainable organic synthesis. Aust J Chem 57:281–289
Singh R, Sharma R, Teewari N, Geetanjali RDS (2006) Nitrilase and its application as a “green” catalyst. Chem Biodivers 3:1279–1287
Sonke T, Ernste S, Tandler RF, Kaptein B, Peeters WPH, van Assema FBJ, Wubbolts MG, Schoemaker HE (2005) L-Selective amidase with extremely broad substrate specificity from Ochrobactrum anthropi NCIMB 40321. Appl Environ Microbiol 71:7961–7973
Sosedov O, Stolz A (2014) Random mutagenesis of the arylacetonitrilase from Pseudomonas fluorescens EBC191 and identification of variants which form increased amounts of mandeloamide from mandelonitrile. Appl Microbiol Biotechnol 98:1595–1607
Sosedov O, Stolz A (2015) Improvement of the amides forming capacity of the arylacetonitrilase from Pseudomonas fluorescens EBC191 by site-directed mutagenesis. Appl Microbiol Biotechnol 99:2623–2635
Sosedov O, Baum S, Bürger S, Matzer K, Kiziak C, Stolz A (2010) Construction and application of variants of the arylacetonitrilase from Pseudomonas fluorescens EBC191 which form increased amounts of acids or amides. Appl Environ Microbiol 76:3668–3674
Stumpp T, Wilms B, Altenbuchner J (2000) Ein neues L-Rhamnose-induzierbares Expressionssystem für Escherichia coli. Biospektrum 1:33–36
Surendra K, Krishnaveni NS, Mahesh A, Rao KR (2006) Supramolecular catalysis of Strecker reaction in water under neutral conditions in the presence of α-cyclodextrin. J Org Chem 71:2532–2534
Vogt VM (1970) Purification and properties of an aminopeptidase from Escherichia coli. J Biol Chem 245:4760–4769
Wang M-X, Lin S-J (2001) Highly efficient and enantioselective synthesis of L-arylglycines and D-arylglycine amides from biotransformations of nitriles. Tetrahedron Lett 42:6925–6927
Wang J, Liu X, Feng X (2011) Asymmetric Strecker reactions. Chem Rev 111:6947–6983
Wegman MA, Heinemann U, Stolz A, van Randwijk F, Sheldon RA (2000) Stereoretentive nitrile hydratase catalysed hydration of D-phenylglycine nitrile. Org Process Res Develop 4:318–322
Wegman MA, Heinemann U, van Randwijk F, Stolz A, Sheldon RA (2001) Hydrolysis of D,L-phenylglycine nitrile by new bacterial cultures. J Mol Catal B 11:249–253
Yasukawa K, Hasemi R, Asano Y (2011) Dynamic kinetic resolution of α-aminonitriles to form chiral α-amino acids. Adv Synth Catal 353:2328–2332
Zuend SJ, Coughlin MP, Lalonde MP, Jacobsen EN (2009) Scaleable catalytic asymmetric Strecker syntheses of unnatural α-amino acids. Nature 461:968–971
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Eppinger, E., Stolz, A. Conversion of phenylglycinonitrile by recombinant Escherichia coli cells synthesizing variants of the arylacetonitrilase from Pseudomonas fluorescens EBC191. Appl Microbiol Biotechnol 103, 6737–6746 (2019). https://doi.org/10.1007/s00253-019-09957-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09957-y