Abstract
Fidaxomicin, an 18-membered macrolide antibiotic, is highly active against Clostridium difficile, the most common cause of diarrhea in hospitalized patients. Though the biosynthetic mechanism of fidaxomicin has been well studied, little is known about its regulatory mechanism. Here, we reported that FadR1, a LAL family transcriptional regulator in the fidaxomicin cluster of Actinoplanes deccanensis Yp-1, acts as an activator for fidaxomicin biosynthesis. The disruption of fadR1 abolished the ability to synthesize fidaxomicin, and production could be restored by reintegrating a single copy of fadR1. Overexpression of fadR1 resulted in an approximately 400 % improvement in fidaxomicin production. Electrophoretic mobility shift assays indicated that fidaxomicin biosynthesis is under the control of FadR1 through its binding to the promoter regions of fadM, fadA1-fadP2, fadS2-fadC, and fadE-fadF, respectively. And the conserved binding sites of FadR1 within the four promoter regions were determined by footprinting experiment. All results indicated that fadR1 encodes a pathway-specific positive regulator of fidaxomicin biosynthesis and upregulates the transcription levels of most of genes by binding to the four above intergenic regions. In summary, we not only clearly elucidate the regulatory mechanism of FadR1 but also provide strategies for the construction of industrial high-yield strain of fidaxomicin.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Actinomycetes are of particular interest as producers of a variety of antibiotics and other valuable secondary metabolites, such as antibiotics, anti-tumor agents, immunosuppressors, and enzyme inhibitors (Newman and Cragg 2012; Harvey 2008; Hopwood 2007). The regulatory network of these secondary metabolites is highly complex and is pivotal for responding to the changes in physiological and environmental conditions (Martin and Liras 2010; Liu et al. 2013; van Wezel and McDowall 2011). Typically, the biosynthesis of natural product is determined by a large gene cluster, within which there is always one or more pathway-specific regulatory genes (Malpartida and Hopwood 1986; Nett et al. 2009). Analysis of these regulatory genes is important to understand the molecular mechanisms of regulation, and this would be of great value in the pharmaceutical industry.
A number of regulators responsible for initiating secondary metabolite production have already been identified. For instance, the Streptomyces antibiotic regulatory proteins (SARPs) containing a DNA-binding domain at the N-terminus are well-characterized pathway-specific regulators and usually act as transcriptional activators (Wietzorrek and Bibb 1997; Arias et al. 1999; Narva and Feitelson 1990; Yang et al. 2015). The LAL regulators (large ATP-binding regulators of the LuxR family) are another important pathway-specific regulatory family, most of which have been reported to play positive roles in antibiotic production (Martín and Liras 2012). The LAL family contains two functional domains, an N-terminal ATP/GTP-binding domain with conserved Walker A motif and a C-terminal DNA-binding domain (DBD) with a conserved helix-turn-helix (HTH) motif (Walker et al. 1982; Chen and Xie 2011). So far, several LAL family regulators have been identified, including FkbN of tacrolimus gene cluster in Streptomyces tsukubaensis L19 (Zhang et al. 2016a, b), AveR of avermectin pathway in Streptomyces avermitilis (Guo et al. 2010), and AstG1 of ansatrienins gene cluster in Streptomyces sp. XZQH13 (Xie et al. 2015). In most cases, overexpression of these pathway-specific regulators resulted in an increased production of corresponding antibiotics (Zhang et al. 2016a, b; Qu et al. 2015), so it is very necessary to investigate the regulatory mechanisms of these regulators for accelerating the process of industrialization of value-added drugs.
Fidaxomicin, also known as tiacumicin B, lipiarmycin A3, PAR-101, OPT-80, and difimicin, is an 18-membered macrolide glycosidic polyketide with a halogenated moiety. It can act as an RNA polymerase inhibitor and is highly active against various gram-positive pathogenic bacteria (Gualtieri et al. 2006). It was approved by the FDA for the treatment of Clostridium difficile infection (CDI) which has become a significant problem in health care (Sullivan and Spooner 2010). Due to its importance and expensiveness in clinical medicine, it is necessary to focus more research on the biosynthesis of fidaxomicin to enhance the production titers and decrease the cost of fidaxomicin production. To date, several strains have been reported as fidaxomicin producers like Dactylosporangium aurantiacum NRRL 18085 (Hochlowski et al. 1987), Micromonospora echinospora subsp. Armeniaca (Omura et al. 1986), Actinoplanes deccanensis ATCC 21983 (Arnone et al. 1987), and Catellatospora sp. Bp3323-81(Kurabachew et al. 2008). The biosynthetic gene cluster of fidaxomicin has been described in D. aurantiacum NRRL 18085 and there are two regulatory genes, tiaR1 and tiaR2, which have been deduced to encode putative regulators homologous to LuxR-type transcriptional activator and ArsR-type repressor, respectively. However, no biological experiments have yet been performed to verify their functions in the production of fidaxomicin (Xiao et al. 2011; Niu et al. 2011).
In this work, we sequenced and annotated the whole genome of the A. deccanensis Yp-1 from China General Microbiological Culture Collection Center (CGMCC 4.2098). One gene cluster responsible for fidaxomicin biosynthesis was located in the genome and the coding gene for the putative regulator involved in fidaxomicin biosynthesis was annotated as fadR1 which was highly homologous to tiaR1 gene; however, no homolog of tiaR2 was found in the chromosome. Here, we provided genetic and biochemical evidences which suggested that FadR1 is a pathway-specific transcriptional activator for fidaxomicin biosynthesis. We also investigated the regulatory mechanism in which FadR1 upregulates the transcription levels of most of key genes by binding to four intergenic regions. Furthermore, we increased the yield of fidaxomicin by 400% through genetically engineering the fadR1 gene which will lay a good foundation for industrialization of fidaxomicin.
Materials and methods
Media, plasmids, strains, and growth conditions
All bacterial strains and plasmids used in this study were listed in Table 1. The manipulation of nucleic acids and bacterial growth was carried out according to the standard protocols as previously described (Bierman et al. 1992). Escherichia coli TG1 (Novagen, Madison, WI, USA) was the general cloning host. E. coli BL21 (DE3) (Novagen, Madison, WI, USA) was used for protein expression. Vectors used were pSET152, pKC1139, pIJ8630, and pTA2. A. deccanensis Yp-1 strain (CGMCC 4.2098) was cultured at 30 °C on ISP4 agar for sporulation and at 30 °C in YEME medium [3 g/L yeast extract, 3 g/L malt extract, 5 g/L tryptone, 10 g/L glucose] for fidaxomicin production.
In-frame deletion and complementation of fadR1
Disruption of fadR1 was performed by gene replacement according to the previously described method (Yuan et al. 2016). The upstream and downstream regions of fadR1 were amplified with the primer pairs fadR1-A-F and fadR1-A-R as well as fadR1-B-F and fadR1-B-R (Table S1). The PCR products were purified and ligated into pKC1139 using a ClonExpress II One Step Cloning Kit (Vazyme Biotech Co, Nanjing, Jiangsu, China) to generate the disruption plasmid pdel-fadR1. The resulting plasmid pdel-fadR1 was then conjugated by E. coli ET12567/pUZ8002 into A. deccanensis Yp-1. The mutants were selected by replica plating for apramycin-sensitive colonies and confirmed by PCR amplification using primer pair P1 and P2 (Table S1).
fadR1 with the fadR1 promoter fragment amplified using primers fadR1-HB-F and fadR1-HB-R (Table S1) was cloned into pSET152 to generate the complementation plasmid com-R1, which was integrated into the ΔfadR1 mutant to obtain the complementation strain.
Overexpression of fadR1
The fadR1 ORF was amplified using primers fadR1-GP-F and fadR1-GP-R (Table S1). This PCR product was then ligated into the pIJ8630 containing the ermE* promoter by the ClonExpress II One Step Cloning Kit (Vazyme Biotech Co, Nanjing, Jiangsu, China). The resulting plasmid OE-R1 was then introduced into A. deccanensis Yp-1.
Electrophoretic mobility shift assays
The primer pair pex-FadR1-F and pex-FadR1-R (Table S1) was used to amplify fadR1. The fadR1 fragment was cloned into pET28a to get the expression plasmid pex-fadR1, which was then introduced into E. coli BL21 (DE3). The resulting strain was grown at 37 °C to an optical density (OD600) of 0.6 and then induced with 0.1 mM isopropyl-thiogalactopyranoside (IPTG) at 16 °C for 8 h. The soluble histidine–tagged FadR1 was purified with Ni2+-nitrilotriacetic acid (NTA) resin (Qiagen) according to the manufacturer’s instructions.
The electrophoretic mobility shift assay (EMSA) DNA probes P1 (236bp), P2 (277bp), P3 (432bp), P4 (278bp), P5 (191bp), P6 (570bp), P7 (304bp), and P8 (336bp) were amplified by PCR using the primer pairs as listed in Table S1. The PCR products were firstly cloned into a pTA2-vector (TOYOBO, Kita-ku, Osaka, Japan). Then, the 5′-FAM-labeled probes were made using 5′-FAM-labeled M13 universal primers by PCR amplification. The probes PA (64 bp), PAmI (64 bp), PAmII (64 bp), PAmI-II (64 bp), PB (60 bp), PBmI (60 bp), PBmII (60 bp), PBmI-II (60 bp), PC (50 bp), PCmI (50 bp), PCmII (50 bp), PCmI-II (50 bp), PD (60 bp), PD(1)mII (60 bp), PD(2)mII (60 bp), PDmII (60 bp), PE (44 bp), and PEmII (44 bp) were prepared following the above method. The EMSAs assay were carried out as previously described with a little modification (Yu et al. 2014). In the EMSAs assay, 80 ng of the probes was incubated with varying quantities of FadR1, at 30 °C for 30 min in the buffer (20 mM Tris, pH 7.5, 5% glycerol, 0.01% BSA, 50 μg mL−1 sheared sperm DNA). The reactions were displayed on 5% acrylamide gels for separation in a 0.5 × TBE buffer. The shift bands were then detected using the ImageQuant LAS 4000 (GE Healthcare Life Sciences, Piscataway, NJ, USA).
DNase I footprinting assay
The DNase I footprinting assays were carried out as previously described (Yu et al. 2014). In brief, 5′-FAM-labeled M13 universal primers were used to amplify FAM-labeled probes. Next, the probes were purified by AxyPrep DNA Gel Extraction Kit (Axygen Biosciences, Union City, CA, USA) and were incubated with different quantities of FadR1 protein at 30 °C for 30 min. Then, 0.01 U of DNase I (Promega, Madison WI, USA) with 10 mM MgCl2 and 1 mM CaCl2 was added for digesting exactly 1 min at 30 °C. The reaction mixtures were stopped by adding 50 μL of 0.1 M EDTA (pH 8.0). After extraction with phenol/chloroform and precipitation with 0.75 M NH4Ac, 40 μg of glycogen, and 70% ethanol, DNAs mixed with Liz-500 DNA marker (MCLAB, South San Francisco, CA, USA) were loaded into an ABI 3130 sequencer and electropherograms were analyzed by Genemapper v4.0 software (Applied Biosystems, Foster City, CA, USA) to align and determine the protected region. The DNA sequencing ladder was prepared according to Thermo Sequence Dye Primer Manual Cycle Sequencing Kit (Affemetrix, Santa Clara, CA, USA). The alignment and consensus of the binding sequences was analyzed by DNAMAN software (Lynnon, San Ramon, CA, USA).
RNA isolation, reverse transcriptase PCR and quantitative real-time PCR
In order to prepare genomic DNA-free RNA, RNA isolation was performed as follows. In brief, A. deccanensis Yp-1 incubated in YEME medium at 30 °C, 220 rpm. Mycelia were collected and washed twice with RNase-free TE buffer. Total RNAs were prepared from mycelia using EASYspin Plus bacteria RNA extract kit (Aidlab Biotech, Beijing, China) according to the manufacturer’s instructions. In order to confirm the absence of genomic DNA contamination, the RNase-free DNase I (Takara, Shiga, Japan)–treated RNA samples were amplified by PCR with three different primer pairs. Then, the concentration of genomic DNA-free RNA was measured by Thermo Scientific™ NanoDrop™. The cDNA was prepared using PrimeScript™ 1st Strand cDNA Synthesis Kit (Takara, Shiga, Japan) following the manufacturer’s instructions.
To analyze transcription units in fidaxomicin biosynthesis gene cluster, the cDNA of A. deccanensis Yp-1 was used as templates for reverse transcriptase PCR (RT-PCR) by rTaq polymerase with primer pairs (Table S1). The co-transcription assay primers were designed based on the following principle: the upstream primers were selected from the end of genes and the downstream primers were designed at the middle of the adjacent genes. And the genomic DNA was also performed PCR amplification as a positive control. Subsequently, the PCR products were analyzed by 1% agarose gel electrophoresis after 31 cycles of amplification. These analyses were carried out at least three times for each primer pair.
To analyze the transcription level of the fidaxomicin gene cluster in A. deccanensis Yp-1, ΔfadR1, and OE-R1/WT, quantitative real-time PCR (RT-qPCR) was carried out on Roche LightCycler 480 (Roche, Mannheim, Germany) with the SYBR Premix Ex Taq (Takara, Shiga, Japan) in a 20-μL volume following the manufacturer’s instructions. The sequence-specific primers used for each gene are listed in Table S1. And the expression level of hrdB was used as the internal reference. The relative expression level was calculated by normalizing to the expression level of hrdB. Each experiment was performed in triplicate.
Determination of fidaxomicin production by HPLC analysis
Fidaxomicin production was confirmed by HPLC analysis using the Agilent 1100 HPLC system (Agilent Technologies, Palo Alto, CA, USA). A HC-C18 column (5 μm, 4.6 by 250 mm) was used with UV detector set at 254 nm. The mobile phase and gradient elution process were as previously described (Xiao et al. 2011). All experiments were performed in quadruplicate.
Accession number of nucleic acid sequence
The sequence of the fidaxomicin cluster in the A. deccanensis Yp-1 has been deposited in Genbank (accession number MG972807).
Results
Identification and analysis of the fidaxomicin gene cluster
A fidaxomicin-producing strain was purchased from China General Microbiological Culture Collection Center (CGMCC). 16S rRNA sequence analysis showed that the sequence shared 99%, 98%, and 97% nucleotide identity with the strain A. deccanensis IFO 13994, A. deccanensis JCM9916, and A. deccanensis Y16, respectively. We therefore designated this strain as A. deccanensis Yp-1 (CGMCC 4.2098). Illumina & PacBio sequencing techniques were used to determine the complete nucleotide sequence of the A. deccanensis Yp-1 genome at Beijing Genomics Institute (BGI) in Shenzhen, China. The size of the A. deccanensis Yp-1 draft genome was determined to be approximately 10.5 Mb. The genome was submitted to RAST Annotation Server (Aziz et al. 2008) to analyze putative protein-coding sequences (CDSs) and their functions which revealed about 9730 CDSs in the genome. After carrying out antiSMASH analysis (Blin et al. 2017), the fidaxomicin gene cluster was revealed, showing a 90% DNA sequence identity to that of D. aurantiacum NRRL 18085 (GenBank accession number HQ011923). And the sequence of the fidaxomicin cluster in the A. deccanensis Yp-1 has been deposited in GenBank (accession number MG972807). The cluster contained 33 ORFs, spanning 84,962 bp of DNA sequence, was shown in Fig. 1. The deduced functions of the ORFs in the fidaxomicin cluster from A. deccanensis Yp-1 and D. aurantiacum NRRL 18085 are presented in Table S2.
Genetic organization of the fidaxomicin cluster. The genetic organization of the 33 ORFs plus 5 upstream ORFs and 4 downstream ORFs of the fidaxomicin cluster. Arrows indicate deduced transcriptional units. Purple lines above the ORFs indicate DNA fragments for probes (P1–P8) in the EMSA experiments. Red probes (P2, P3, P4, and P5) showed retarded bands, and purple probes (P1, P6, P7, and P8) indicate that probes did not show retarded bands in the EMSA experiments
As compared with D. aurantiacum NRRL 18085, the fidaxomicin cluster in A. deccanensis Yp-1 has five more genes, including fadW1, fadW2, fadW3, fadY1, and fadY2. The deduced products of fadW1, fadW2, and fadW3 are methylmalonyl-CoA mutase–related enzymes, which catalyze the reversible isomerization of methylmalonyl-CoA to succinyl-CoA. The macrocyclic aglycone of fidaxomicin was reported to be biosynthesized from one propionyl-CoA (starter unit), 4 malonyl-CoAs, 4 methylmalonyl-CoAs, and one ethylmalonyl-CoA by TiaA1-TiaA4 (Xiao et al. 2011). So, it seems that succinyl-CoA does not participate in the process of fidaxomicin biosynthesis. This raises the question about the function of methylmalonyl-CoA mutase in the fidaxomicin cluster. fadY1 and fadY2 have been shown to encode 2-oxoacid oxidoreductase–related enzymes. The functions of these enzymes were suggested to use ferredoxin as an electron acceptor in the formation of a free radical intermediate (Gibson et al. 2016; Pierce et al. 2010). However, the fidaxomicin cluster of A. deccanensis Yp-1 did not contain homologous genes corresponding to tiaS3, tiaT3, and tiaT4 in D. aurantiacum NRRL 18085 (Table S2). tiaS3 was reported to encode a GDP-D-mannose 4,6-dehydratase, which converts GDP-D-mannose to D-rhamnose. TiaT3 and TiaT4 are ABC transporter–related proteins and may be involved in the transport of fidaxomicin and its analogues. It has been suggested that the functions of these enzymes may be replaced by other proteins or that they are just not involved in the fidaxomicin biosynthesis. It is also worth noting that the ArsR-type repressor, TiaR2, is not present in the fidaxomicin cluster or anywhere else in the chromosome of A. deccanensis Yp-1, indicating a different regulatory mechanism of fidaxomicin between A. deccanensis Yp-1 and D. aurantiacum NRRL 18085.
FadR1 is a putative pathway-specific transcriptional regulator
In the fidaxomicin gene cluster of D. aurantiacum NRRL 18085, there are two pathway-specific regulators, TiaR1 and TiaR2. In contrast, only one pathway-specific regulatory gene, fadR1, was found in the corresponding gene cluster of A. deccanensis Yp-1. Sequence analysis of the fadR1 gene product (873aa) showed a strikingly high sequence identity (88%) to TiaR1. FadR1 contained C-terminal DNA-binding domain of LuxR-like proteins with a helix-turn-helix motif and an N-terminal AAA ATPase domain with a P-loop motif involved in binding ATP, which suggested that FadR1 belongs to LAL family of transcription factors. To investigate the contribution of fadR1 to fidaxomicin biosynthesis in A. deccanensis Yp-1, partial coding region of fadR1 was deleted via homologous recombination as described in the “Materials and methods” section. The disruption mutant ΔfadR1 was confirmed by PCR analysis (see Fig. S1) and further verified by DNA sequencing (data not shown). Meanwhile, the complementary strain was constructed by introducing Com-R1, containing fadR1 and its own promoter into the deleted mutant ΔfadR1. High-performance liquid chromatography (HPLC) analysis (Fig. 2) showed that deletion of fadR1 totally blocked fidaxomicin production, while the complementary strain Com-R1/ΔfadR1 was found to restore the production of fidaxomicin to a similar level compared with that of the WT strain (Fig. 2). Nevertheless, there were no obvious differences in the growth rates and morphological differentiation among these three strains (Fig. S1). These results indicated that FadR1 is a pivotal positive regulator in fidaxomicin biosynthesis.
The effect of fadR1 disruption and overexpression on production of fidaxomicin. The strains were grown in YEME medium for 120 h. HPLC analysis of fermentation filtrates from the WT strain, the fadR1 deletion mutant, the fadR1-complementary mutant, and the fadR1 overexpression mutant. The peak of fidaxomicin is marked by an arrow. mAU milli-absorbance units
Overexpression of fadR1 results in an increased fidaxomicin production
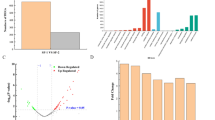
In order to further evaluate the effect of increased expression of fadR1 to the yield of fidaxomicin, a fadR1 overexpression plasmid OE-R1 was constructed, in which fadR1 was cloned into pIJ8630 under the control of the strong constitutive ermE* promoter. This plasmid was then introduced into the wild-type strain by conjugation, resulting in fadR1 overexpression strain OE-R1/WT. The overexpression of fadR1 was confirmed by RT-qPCR analysis, and its effects on cell growth, aerial mycelium formation, and fidaxomicin productivity in the fermentation medium were also determined. As seen in Fig. 3, the presence of an extra copy of fadR1 under ermE*p (OE-R1/WT) led to the increase of fidaxomicin production by approximately 400 % compared with that of the wild-type strain. And no apparent difference in the cell growth and morphology between the overexpression strain and wild type was observed (Fig. S1). These results further demonstrated the positive regulatory roles of FadR1 in the pathway.
Effects of FadR1 on the transcription levels of fidaxomicin biosynthetic genes
To test whether FadR1 positively regulates fidaxomicin production by tuning the transcription levels of associated biosynthetic genes, we performed RT-qPCR analysis. The results suggested that the transcription levels of most fidaxomicin-associated biosynthetic genes exhibited an obvious increase in the fadR1 overexpression strain. Firstly, the expressions of precursor-associated genes like fadJ, fadN, and fadK involved in synthesis of EM-CoA extender units and fadE encoding thioesterase responsible for eliminating abnormal precursors were upregulated by 5–20-fold. The transcription levels of fadB, fadF, and fadM involved in the biosynthesis and modification of the aromatic moiety were increased by 20–30-fold. Secondly, we can see that the transcription levels of four PKS genes fadA1–fadA4 that encode modular polyketide synthase responsible to assemble precursor units like propionyl-CoA to form fidaxomicin aglycone were increased by 10–40-fold. Thirdly, the transcription levels of fadS1-S6 encoding a series of modifying enzymes that were involved in the biosynthesis of deoxygenated sugars and fadG1 and fadG2 that modified these deoxygenated sugars at C-11 and C-20 were increased by 8–30-fold. Finally, FadR1 also significantly improved the expression of other genes such as fadW1, fadW2, and fadW3 (methylmalonyl-CoA mutase–related enzymes), fadP2 (cytochrome P450) fadT1 (transport protein), fadC (E1 alpha subunit), and fadD (E1 beta subunit). The increase of expression levels of these precursors or skeleton modification steps often helps to improve the yield of antibiotics (Matthews and Wurtzel 2000; Hemmerlin 2013). However, the transcription levels of all fidaxomicin-associated biosynthetic genes were decreased by at least 50% and some genes like fadS4, fadE, fadS2, fadS1, and fadJ were even barely expressed in the fadR1 deletion mutant (ΔfadR1) (Fig. 4). Accordingly, we can conclude that FadR1 positively regulates the transcription levels of most of genes in the fidaxomicin gene cluster and its overexpression resulted in the increase of fidaxomicin production. These results further confirmed that FadR1 is a pathway-specific positive transcriptional regulator of fidaxomicin biosynthesis. And in view of the regulatory mechanisms of most of pathway-specific regulators (Bibb 2005; Chen et al. 2010), we proposed that the binding sites of FadR1 were distributed in the promoter regions of fidaxomicin-associated biosynthetic genes.
Effect of fadR1 deletion and overexpression on the transcription level of fidaxomicin cluster genes. Compared with wild-type strain, the relative expression of fidaxomicin biosynthetic genes in fadR1 deletion mutant was shown in a and in fadR1 overexpression strain was shown in b. The RNA samples were obtained from cultures grown in YEME medium. The expression levels of fidaxomicin cluster genes are presented relative to the levels of corresponding genes in the wild-type sample, which were arbitrarily assigned a value of 1. The transcription level of hrdB was assayed as an internal control, and error bars were calculated by measuring the standard deviations among data from three replicates of each sample
Identification of FadR1 binding sites in the fidaxomicin gene cluster
In order to investigate the detailed regulatory mechanism of FadR1, we firstly characterized the organization of transcription units in the fidaxomicin cluster by RT-PCR (Materials and methods). cDNA samples were prepared from mycelia of A. deccanensis Yp-1 after being incubated in YEME medium at 30 °C, and primers covering the intergenic regions between two genes were designed (Table S1). As shown in Fig. 5, an amplified cDNA band was observed using the primers covering the intergenic region between fadK and fadL. However, no transcript was detected between fadJ and fadK. The results indicated that fadJ and fadK genes had their own promoters while fadK and fadL were co-transcribed as an operon. However, an overlapping sequence was observed between fadK and fadJ according to the nucleotide sequence of fidaxomicin gene cluster. So, in order to confirm that fadJ and fadK genes were transcribed separately, another two primer pairs were used to amplify the cDNA of A. deccanensis Yp-1 and still no transcripts were observed (data not shown). Finally, we concluded that fadK had its own promoter, and the promoter may be located at the inner of fadJ, and fadK and fadL were co-transcribed as an operon. Besides, based on the RT-PCR results, fadA1, fadA2, fadA3, fadA4, fadN, fadY1, fadY2, fadW1, fadW2, fadW3, and fadT2 were deduced to encode a polycistronic transcript, while fadE, fadP1, fadP2, and fadJ were transcribed separately. And the transcriptional features of other genes in the fidaxomicin cluster were also characterized. In view of the chromosomal arrangement, the fidaxomicin gene cluster was proposed to have twelve transcription units (Fig. 1).
To identify the specific binding sites of FadR1 in the fidaxomicin gene cluster, we carried out EMSAs with eight 5′-FAM-labeled probes (P1-P8), which covered all possible promoter regions according to the transcriptional organization of fidaxomicin gene cluster. As shown in Fig. 6, when FadR1 was incubated with probes P2 (intergenic region of fadE-fadF), P3 (intergenic region of fadS2-fadC), P4 (intergenic region of fadA1-fadP2), and P5 (promoter region of fadM), retarded bands were observed which suggested that FadR1 could directly bind to these promoter regions. However, no retarded bands appeared (Fig. S2) when FadR1 was incubated with other probes, indicating that FadR1 could not directly interact with these promoter regions. Besides, the pTA2 vector probe was used as a negative control which can confirm the binding specificity of FadR1 to the above four promoter regions.
To further define the specific binding sites of FadR1 in each promoter region, DNase I footprinting assays were carried out. The experiments revealed that there are two binding sites in the intergenic region of fadS2-fadC, while there is one binding site in the intergenic regions of fadE-fadF and fadA1-fadP2 and the promoter of fadM (Fig. 7a, c, e, g). The nucleotide sequences of the promoter regions of fadS2-fadC, fadE-fadF, fadA1-fadP2, and fadM, and the FadR1 binding sites were showed in Fig. 7b, d, f, and h. These results further confirmed the direct binding of FadR1 to the multiple binding sites of the fidaxomicin cluster.
DNase I footprinting assay for determination of the FadR1-binding sites. 5′-FAM-labeled P2 covering the promoter region of fadE–fadF (a), P3 covering the promoter region of fadS2–fadC (c), P4 covering the promoter region of fadA1–fadP2 (e), and P5 covering the promoter region of fadM (g) were used in the DNase I footprinting assays, with or without purified FadR1, respectively. The protected regions are underlined. Nucleotide sequences of the promoter regions of fadE–fadF, fadS2–fadC, fadA1–fadP2, and fadM were shown in b, d, f, and h, respectively. In site D, two consensus AGGG motsif were found, named site D(1) and site D(2). The FadR1-binding sites are overlined and the initiation codons are marked with arrows. Sequences of the determined FadR1-binding sites and the consensus FadR1-binding sequence (in boldface type) were shown in g
Based on the results from EMSA and footprinting, we can see that the binding sites of FadR1 were located at the intergenic regions of fadE-fadF, fadS2-fadC, and fadA1-fadP2 and the promoter of fadM. Combined with the analysis of transcription units, we can know that the promoters located at the above four intergenic regions were important for controlling the most of key genes of fidaxomicin biosynthesis like PKS skeleton-associated genes, genes for precursor supply, and genes for glycosyl synthesis and modification. And this result is consistent with the changes of transcription level learned from RT-qPCR. Finally, we concluded that fadR1 positively regulates the transcription of most of genes in the fidaxomicin gene cluster directly or indirectly by binding to the four intergenic regions, which resulted in the improvement of fidaxomicin production. According to the above results, we proposed the regulatory mode of FadR1 controlling the biosynthesis of fidaxomicin (Fig. 8).
Characterization of the consensus FadR1-binding sequence
By aligning the sequences within the protected regions of the five FadR1-binding regions, a consensus inverted repeat (IR) 5′-CCCTNxAGGG-3′ was identified in the intergenic regions of fadE-fadF and fadS2-fadC, and a consensus direct repeat (DR) 5′-AGGGNxAGGG-3′ was observed in the intergenic region of fadA1-fadP2, while only a consensus sequence 5′-AGGG-3′ was revealed in the promoter of fadM (Fig. 7i). To further study the roles of these consensus sequences for their FadR1-binding ability, EMSAs were carried out using the probes containing either the sequences of wild-type binding sites or the mutated sites (Fig. S3). When one of the consensus sequences in site A, site B, site C, or site D were mutated, binding shifts were still detected (Fig. 9b, c, f, g, j, k, n, o). However, when we carried out mutation in both consensus sequences, there were no evident banding shifts (Fig. 9d, h, l, p). On the other hand, there was only one consensus sequence in the site E, and the mutation of this sequence could completely abolish FadR1 complex formation with the labeled probe (Fig. 9r). The above data validated the FadR1 consensus binding sequences and confirmed that these consensus sequences are essential for the binding activity of FadR1. Based on these results, we proposed that FadR1 probably recognized the structures derived from the consensus sequence and then recruited RNA polymerase by binding to the target promoter regions to enhance the expression of fidaxomicin biosynthetic genes.
Mutational analysis of consensus sequences in the FadR1-binding sites. EMSAs for the determination of FadR1 binding to mutated sequences. Probes containing the FadR1-binding sequences of wild-type and mutants are as shown in Fig. S3.PA: Probe A, PAm: Probe A mutant, PB: Probe B, PBm: Probe B mutant, PC: Probe C, PCm: Probe C mutant, PD: Probe D, PDm: Probe D mutant, PE: Probe E, PEm: Probe E mutant. Lanes 1–3 are labeled fragment with 0 μg, 0.05 μg, and 0.2 μg of purified FadR1, respectively. The probe-FadR1 complex is marked by an arrow
Discussion
It has been reported that many pathway-specific positive regulators from polyene macrolide biosynthesis gene clusters belong to the LAL (large ATP-binding regulators of the LuxR) family, such as PimM, NysR-RIII, AmphRI-RIII, FscRI, PikD, and AveR. And they always share similar binding sites and functional conservation (Santos-Aberturas et al. 2011). However, only several LAL family regulators have been well-characterized, which is not enough for us to fully comprehend the significant roles of this protein family (Chen and Xie 2011).
In our study, complete genome sequencing and bioinformatic analysis reveal a typical LAL family regulator named FadR1 in fidaxomicin gene cluster from A. deccanensis Yp-1. And we have proven that FadR1 can bind to four intergenic regions of several key genes, thereby activating the transcription of most of genes in the fidaxomicin gene cluster directly or indirectly and finally leading to an increase in the production of fidaxomicin. In addition, the analysis of binding sites reveals a conserved sequence 5′-AGGG-3′ which is common and important for lots of LuxR regulators (Santos-Aberturas et al. 2011). Overall, the elucidation of the regulatory role of FadR1 contributes to the understanding of the entire fidaxomicin biosynthetic process and the investigation of important function of analogical LAL family regulators in the gene clusters of other macrolide antibiotics. However, according to our results, there are still some protein-probe complexes displaying a relatively smeared pattern although we have optimized several conditions for EMSA. One of the main possible reasons is that the binding of FadR1 to conserved sites occurred but the protein-probe complexes may be unstable in vitro, resulting in dissociation during electrophoresis. Similar phenomena are also observed for other various regulators like the LuxR transcriptional regulator PimM from Streptomyces natalensis (Santos-Aberturas et al. 2011), the SARP family regulator AfsR from S. coelicolor A3(2) (Tanaka et al. 2007), the osmotic regulator OmpR from E. coli (Harlocker et al. 1995), the global response regulator CtrA from Caulobacter crescentus (Spencer et al. 2009), the quorum-sensing regulator RsaL from Pseudomonas aeruginosa (Rampioni et al. 2007), and the developmental regulator BldD from Saccharopolyspora erythraea (Chng et al. 2008). Besides, as shown in Fig. 9, we also notice that when the conserved sites are mutated, the FadR1-probe complexes become more smeared, such as Fig. 9 c-3 or f-3. Similar phenomenon is also observed in the Pseudomonas quorum-sensing regulator RsaL (Rampioni et al. 2007). So, we suppose that the secondary structure of FadR1 binding sites formed by inverted repeat (IR) or direct repeat (DR) sites may also be required for the formation of stable complexes between FadR1 and its target DNAs. Certainly, further studies are needed to investigate the underlying causes.
Furthermore, it is particularly interesting to note that the regulatory mechanism of FadR1 is very similar to other LuxR regulators like PimM, AmphRIV, NysRIV, and PteF from polyene macrolide gene clusters. A previous study by Santos-Aberturas et al. (2011) proves that heterologous LuxR regulators like AmphRIV, NysRIV, and PteF can complement the function of PimM in S. natalensis ΔpimM, and the yield of pimaricin displays a marked difference, which suggests that these functionally conserved regulators are fully exchangeable and may display different regulatory intensities in different hosts. Therefore, we suppose that replacing FadR1 with other LuxR regulators may further improve the production of fidaxomicin and we will consider performing more relevant experiments in the future. Besides, Martínez-Burgo et al. (2019)’s research proves that heterologous expression of pimM from S. natalensis in S. clavuligerus can activate cryptic gene clusters. Similarly, Yushchuk’s (Yushchuk et al. 2018) study suggests that heterologous AdpA transcription factors of different origins can activate landomycin biosynthesis in S. cyanogenus S136 harboring a nonfunctional AdpA. Thus, we suppose that some gene clusters are silent in native strains but are active in heterologous hosts which may be due to the influence of regulatory networks. Since FadR1 as a pathway-specific positive regulator displays functional conservation with other LuxR transcriptional activators, it also has great potential to activate cryptic gene clusters in heterologous strains for drug discovery. Also, further researches are needed to prove this hypothesis. For instance, according to the conserved binding sites CTVGGGAWWTCCCBAG of some LuxR regulators, we can perform whole genome scanning to locate the conserved sites by the MEME suite (Santos-Aberturas et al. 2011; Bailey et al. 2015) and predict the target cryptic gene clusters. Then, FadR1 regulator can be expressed in the corresponding heterologous host to activate silent gene clusters, which may accelerate the excavation of novel natural products.
In summary, we have characterized FadR1 as a LuxR-type pathway-specific activator in controlling fidaxomicin biosynthesis, analyzed its regulatory mechanism, and proposed the action mode of FadR1 regulator in fidaxomicin biosynthesis. Our future research goals are to determine how FadR1 responds to environmental signals to initiate fidaxomicin biosynthesis and to look into whether there is a higher level regulatory network (Liu et al. 2013) directly involved in the regulation of FadR1. Also, we will try to discover more efficient LuxR regulators to replace FadR1 for further enhancing the production of fidaxomicin as well as to activate cryptic gene clusters by heterologous expression of FadR1. This study not only sets the stage for an increased understanding of the genetic control of fidaxomicin biosynthesis which would have general implications about the regulatory mechanism of other macrolide antibiotics harboring similar LAL family regulator in the gene clusters but also will provide an effective strategy to improve the yield of fidaxomicin.
References
Arias P, Fernandez-Moreno MA, Malpartida F (1999) Characterization of the pathway-specific positive transcriptional regulator for actinorhodin biosynthesis in Streptomyces coelicolor A3(2) as a DNA-binding protein. J Bacteriol 181:6958–6968
Arnone A, Nasini G, Cavalleri B (1987) Structure elucidation of the macrocyclic antibiotic lipiarmycin. J Chem Soc Perkin Trans 1:1353–1359. https://doi.org/10.1039/P19870001353
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:75. https://doi.org/10.1186/1471-2164-9-75
Bailey TL, Johnson J, Grant CE, Noble WS (2015) The MEME suite. Nucleic Acids Res 43(W1):W39–W49. https://doi.org/10.1093/nar/gkv416
Bibb MJ (2005) Regulation of secondary metabolism in streptomycetes. Curr Opin Microbiol 8:208–215. https://doi.org/10.1016/j.mib.2005.02.016
Bierman M, Logan R, Obrien K, Seno ET, Rao RN, Schoner BE (1992) Plasmid cloning vectors for the conjugal transfer of DNA from Escherichia coli to Streptomyces Spp. Gene 116:43–49. https://doi.org/10.1016/0378-1119(92)90627-2
Blin K, Wolf T, Chevrette MG, Lu XW, Schwalen CJ, Kautsar SA, Duran HS, Santos EL, Kim HU, Nave M, Dickschat JS, Mitchell DA, Shelest E, Breitling R, Takano E, Lee SY, Weber T, Medema MH (2017) antiSMASH 4.0—improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res W1:W36–W41. https://doi.org/10.1093/nar/gkx319
Chen J, Xie J (2011) Role and regulation of bacterial LuxR-like regulators. J Cell Biochem 10:2694–2702. https://doi.org/10.1002/jcb.23219
Chen YH, Smanski MJ, Shen B (2010) Improvement of secondary metabolite production in Streptomyces by manipulating pathway regulation. Appl Microbiol Biotechnol 86:19–25. https://doi.org/10.1007/s00253-009-2428-3
Chng C, Lum AM, Vroom JA, Kao CM (2008) A key developmental regulator controls the synthesis of the antibiotic erythromycin in Saccharopolyspora erythraea. Proc Natl Acad Sci USA 105(32):11346–11351. https://doi.org/10.1073/pnas.0803622105
Gibson MI, Chen PY, Drennan CL (2016) A structural phylogeny for understanding 2-oxoacid oxidoreductase function. Curr Opin Struct Biol 41:54–61. https://doi.org/10.1016/j.sbi.2016.05.011
Gualtieri M, Villain-Guillot P, Latouche J, Leonetti JP, Bastide L (2006) Mutation in the Bacillus subtilis RNA polymerase β′ subunit confers resistance to lipiarmycin. Antimicrob Agents Chemother 50:401–402. https://doi.org/10.1128/AAC.50.1.401-402.2006
Guo J, Zhao J, Li L, Chen Z, Wen Y, Li J (2010) The pathway-specific regulator AveR from Streptomyces avermitilis positively regulates avermectin production while it negatively affects oligomycin biosynthesis. Mol Gen Genomics 283:123–133. https://doi.org/10.1007/s00438-009-0502-2
Harlocker SL, Bergstrom L, Inouye M (1995) Tandem binding of six OmpR proteins to the ompF upstream regulatory sequence of Escherichia coli. J Biol Chem 270(45):26849–26856. https://doi.org/10.1074/jbc.270.45.26849
Harvey AL (2008) Natural products in drug discovery. Drug Discov Today 13:894–901. https://doi.org/10.1016/j.drudis.2008.07.004
Hemmerlin A (2013) Post-translational events and modifications regulating plant enzymes involved in isoprenoid precursor biosynthesis. Plant Sci 203:41–54. https://doi.org/10.1016/j.plantsci.2012.12.008
Hochlowski JE, Swanson SJ, Ranfranz LM, Whittern DN, Buko AM, McAlpine JB (1987) Tiacumicins, a novel complex of 18-membered macrolides. II. Isolation and structure determination. J Antibiot 40:575–588. https://doi.org/10.7164/antibiotics.40.575
Hopwood DA (2007) Streptomyces in nature and medicine: the antibiotic makers. Oxford University, New York
Kurabachew M, Lu SH, Krastel P, Schmitt EK, Suresh BL, Goh A, Cynamon M (2008) Lipiarmycin targets RNA polymerase and has good activity against multidrug-resistant strains of Mycobacterium tuberculosis. J Antimicrob Chemother 62(4):713–719. https://doi.org/10.1093/jac/dkn269
Liu G, Chater KF, Chandra G, Niu G, Tan H (2013) Molecular regulation of antibiotic biosynthesis in Streptomyces. Microbiol Mol Biol Rev 77:112–143. https://doi.org/10.1128/MMBR.00054-12
MacNeil DJ, Klapko LM (1987) Transformation of Streptomyces avermitilis by plasmid DNA. J Ind Microbiol 2:209–218. https://doi.org/10.1007/BF01569542
Malpartida F, Hopwood DA (1986) Physical and genetic characterisation of the gene cluster for the antibiotic actinorhodin in Streptomyces coelicolor A3(2). Mol Gen Genet 205:66–73. https://doi.org/10.1007/BF02428033
Martin JF, Liras P (2010) Engineering of regulatory cascades and networks controlling antibiotic biosynthesis in Streptomyces. Curr Opin Microbiol 13:263–273. https://doi.org/10.1016/j.mib.2010.02.008
Martín JF, Liras P (2012) Cascades and networks of regulatory genes that control antibiotic biosynthesis. Subcell Biochem 64:115–138. https://doi.org/10.1007/978-94-007-5055-5_6
Martinez-Burgo Y, Santos-Aberturas J, Rodriguez-Garcia A, Barreales EG, Tormo JR, Truman AW, Reyes F, Aparicio JF, Liras P (2019) Activation of secondary metabolite gene clusters in Streptomyces clavuligerus by the PimM regulator of Streptomyces natalensis. Front Microbiol 10. https://doi.org/10.3389/fmicb.2019.00580
Matthews PD, Wurtzel ET (2000) Metabolic engineering of carotenoid accumulation in Escherichia coli by modulation of the isoprenoid precursor pool with expression of deoxyxylulose phosphate synthase. Appl Microbiol Biotechnol 53:396–400. https://doi.org/10.1007/s002530051632
Narva KE, Feitelson JS (1990) Nucelotide sequence and transcriptional analysis of the redD locus of Streptomyces coelicolor A3(2). J Bacteriol 172:326–333. https://doi.org/10.1128/jb.172.1.326-333.1990
Nett M, Ikeda H, Moore BS (2009) Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat Prod Rep 26:1362–1384. https://doi.org/10.1039/B817069J
Newman DJ, Cragg GM (2012) Natural products as sources of new drugs over the 30 years from 1981 to 2010. J Nat Prod 75:311–335. https://doi.org/10.1021/np200906s
Niu S, Hu T, Li S, Xiao Y, Ma L, Zhang G, Zhang H, Yang X, Ju J, Zhang C (2011) Characterization of a sugar-O-methyltransferase TiaS5 affords new tiacumicin analogues with improved antibacterial properties and reveals substrate promiscuity. Chembiochem 12:1740–1748. https://doi.org/10.1002/cbic.201100129
Omura S, Imamura N, Oiwa R, Kuga H, Iwata R, Masuma R, Iwai YJ (1986) Clostomicins, new antibiotics produced by Micromonospora echinospora subsp. armeniaca subsp. nov. I. Production, isolation, and physico-chemical and biological properties. J Antibiot 39:1407–1412. https://doi.org/10.7164/antibiotics.39.1407
Pierce E, Becker DF, Ragsdale SW (2010) Identification and characterization of oxalate oxidoreductase, a novel thiamine pyrophosphate-dependent 2-oxoacid oxidoreductase that enables anaerobic growth on oxalate. J Biol Chem. 285:40515–40524. https://doi.org/10.1074/jbc.M110.155739
Qu S, Kang Q, Wu H, Wang L, Bai L (2015) Positive and negative regulation of GlnR in validamycin A biosynthesis by binding to different loci in promoter region. Appl Microbiol Biotechnol 99:4771–4783. https://doi.org/10.1007/s00253-015-6437-0
Rampioni G, Polticelli F, Bertani I, Righetti K, Venturi V, Zennaro E, Leoni L (2007) The Pseudomonas quorum-sensing regulator RsaL belongs to the tetrahelical superclass of H-T-H proteins. J Bacteriol 189(5):1922–1930. https://doi.org/10.1128/Jb.01552-06
Santos-Aberturas J, Payero TD, Vicente CM, Guerra SM, Canibano C, Martin JF, Aparicio JF (2011) Functional conservation of PAS-LuxR transcriptional regulators in polyene macrolide biosynthesis. Metab Eng 13:756–767. https://doi.org/10.1016/j.ymben.2011.09.011
Spencer W, Siam R, Ouimet MC, Bastedo DP, Marczynski GT (2009) CtrA, a global response regulator, uses a distinct second category of weak DNA binding sites for cell cycle transcription control in Caulobacter crescentus. J Bacteriol 191(17):5458–5470. https://doi.org/10.1128/Jb.00355-09
Sullivan KM, Spooner LM (2010) Fidaxomicin: a macrocyclic antibiotic for the management of Clostridium difficile infection. Ann. Pharmacother 44:352–359. https://doi.org/10.1345/aph.1M351
Sun J, Kelemen GH, Fernandez-Abalos JM, Bibb MJ (1999) Green fluorescent protein as a reporter for spatial and temporal gene expression in Streptomyces coelicolor A3(2). Microbiology 145:2221–2227. https://doi.org/10.1099/00221287-145-9-2221
Tanaka A, Takano Y, Ohnishi Y, Horinouchi S (2007) AfsR recruits RNA polymerase to the afsS promoter: a model for transcriptional activation by SARPs. J Mol Biol 369(2):322–333. https://doi.org/10.1016/j.jmb.2007.02.096
van Wezel GP, McDowall KJ (2011) The regulation of the secondary metabolism of Streptomyces: new links and experimental advances. Nat Prod Rep 28:1311–1333. https://doi.org/10.1039/C1NP00003A
Walker JE, Saraste M, Runswick MJ, Gay NJ (1982) Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J 1:945–951. https://doi.org/10.1002/j.1460-2075.1982.tb01276.x
Wietzorrek A, Bibb M (1997) A novel family of proteins that regulates antibiotic production in stretomycetes appears to contain an OmpR-like DNA-binding fold. Mol Microbiol 25:1177–1184. https://doi.org/10.1046/j.1365-2958.1997.5421903.x
Xiao Y, Li S, Niu S, Ma L, Zhang G, Zhang H, Zhang G, Ju J, Zhang C (2011) Characterization of tiacumicin B biosynthetic gene cluster affording diversified tiacumicin analogues and revealing a tailoring dihalogenase. J Am Chem Soc 133:1092–1105. https://doi.org/10.1021/ja109445q
Xie C, Deng JJ, Wang HX (2015) Identification of AstG1, A LAL family regulator that positively controls ansatrienins production in Streptomyces sp. XZQH13. Curr Microbiol 70:859–864. https://doi.org/10.1007/s00284-015-0798-6
Yang K, Qi LH, Zhang M, Hou XF, Pan HX, Tang GL, Wang W, Yuan H (2015) The SARP family regulator Txn9 and two-component response regulator Txn11 are key activators for trioxacarcin Biosynthesis in Streptomyces bottropensis. Curr Microbiol 71:458–464. https://doi.org/10.1007/s00284-015-0868-9
Yu P, Liu SP, Bu QT, Zhou ZX, Zhu ZH, Huang FL, Li YQ (2014) WblAch, a pivotal activator of natamycin biosynthesis and morphological differentiation in Streptomyces chattanoogensis L10, is positively regulated by AdpAch. Appl Environ Microbiol 80:6879–6887. https://doi.org/10.1128/AEM.01849-14
Yuan PH, Zhou RC, Chen X, Luo S, Wang F, Mao XM, Li YQ (2016) DepR1, a TetR family transcriptional regulator, positively regulates daptomycin production in an industrial producer, Streptomyces roseosporus SW0702. Appl Environ Microbiol 82:1898–1905. https://doi.org/10.1128/AEM.03002-15
Yushchuk O, Ostash I, Vlasiuk I, Gren T, Luzhetskyy A, Kalinowski J, Fedorenko V, Ostash B (2018) Heterologous AdpA transcription factors enhance landomycin production in Streptomyces cyanogenus S136 under a broad range of growth conditions. Appl Microbiol Biotechnol 102(19):8419–8428. https://doi.org/10.1007/s00253-018-9249-1
Zhang Y, He H, Liu H, Wang H, Wang X, Xiang W (2016a) Characterization of a pathway-specific activator of milbemycin biosynthesis and improved milbemycin production by its overexpression in Streptomyces bingchenggensis. Microb Cell Fact 15:152. https://doi.org/10.1186/s12934-016-0552-1
Zhang XS, Luo HD, Tao Y, Wang YY, Jiang XH, Jiang H, Li YQ (2016b) FkbN and Tcs7 are pathway-specific regulators of the FK506 biosynthetic gene cluster in Streptomyces tsukubaensis L19. J Ind Microbiol Biotechnol 43:1693–1703. https://doi.org/10.1007/s10295-016-1849-0
Acknowledgments
We gratefully thank Dr. Chris Wood, a native English biologist, for his critical reading of this manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (No 31730002 and No 31520103901) and the National Key Research and Development Program (2016YFD0400805).
Author information
Authors and Affiliations
Contributions
Yue-Ping Li and Pin Yu performed the experiments and wrote the paper, and these authors contributed equally to this work; Ji-Feng Li, Yi-Li Tang, and Qing-Ting Bu assisted with the primary data analysis; Yong-Quan Li and Xu-Ming Mao conceived and designed the project; Yong-Quan Li supervised the project and revised the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 644 kb)
Rights and permissions
About this article
Cite this article
Li, YP., Yu, P., Li, JF. et al. FadR1, a pathway-specific activator of fidaxomicin biosynthesis in Actinoplanes deccanensis Yp-1. Appl Microbiol Biotechnol 103, 7583–7596 (2019). https://doi.org/10.1007/s00253-019-09949-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09949-y