Abstract
Gut microbiota play a key role in the regulation of obesity and associated metabolic disorders. To study the relationship between them, antibiotics have been widely used to generate pseudo-germ-free rodents as control models. However, it is not clear whether antibiotics impact an animal’s metabolic phenotype. Therefore, the effect of antibiotics-induced gut microbial perturbations on metabolic phenotypes in high-fat diet (HFD) fed mice was investigated. The results showed that antibiotics perturbed gut microbial composition and structure. Community diversity and richness were reduced, and the phyla Firmicutes/Bacteroidetes (F/B) ratio was decreased by antibiotics. Visualization of Unifrac distance data using principal component analysis (PCA) and unweighted pair-group method with arithmetic mean (UPGAM) demonstrated that fecal samples of HFD-fed mice separated from those of chow diet (CD) fed mice. Fecal samples from antibiotics-treated and non-treated mice were clustered into two different microbial populations. Moreover, antibiotics suppressed HFD-induced metabolic features, including body weight gain (BWG), liver weight (LW), epididymal fat weight (EFW), and serum levels of total cholesterol (TC), low-density lipoprotein cholesterol (LDL-C), high-density lipoprotein cholesterol (HDL-C), alanine aminotransferase (ALT), fasting blood glucose (FBG), and insulin (INS) significantly (P < 0.05). Lachnospiraceae, Ruminiclostridium and Helicobacter, biomarkers of mouse gut microbiota before treatment by antibiotics, were positively correlated with obesity phenotypes significantly (P < 0.05) and were decreased by (92.95 ± 5.09) %, (97.73 ± 2.09) % and (99.48 ± 0.21) % respectively after 30 days of treatment by antibiotics. However, Bacteroidia were enriched in HFD-fed antibiotics-treated mice and were negatively correlated with obesity phenotypes significantly (P < 0.05). We suggested that the antibiotics-induced depletion of Lachnospiraceae, Ruminiclostridium, and Helicobacter, and the decrease in F/B ratio in gut microbiota played a role in the prevention of HFD-induced obesity in mice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Obesity is a global disease that severely impairs physical health (Al-Assal et al. 2018). Roughly, a third of the world’s population is affected by overweight or obesity. Indeed, over 1.9 billion people are overweight worldwide, of which ~ 600 million people are obese, which leads to considerable economic costs and public health challenges (Sara-Assar 2017). Obesity predisposes an individual to a variety of diseases, including metabolic syndrome, nonalcoholic fatty liver disease, several immune-related disorders, and even cancer (Donohoe et al. 2017; Portune et al. 2017; Rastelli et al. 2018). These disorders generate a spectrum of overlapping phenotypes, such as overweight, fat accumulation, dyslipidemia, and liver dysfunction that arise from a complex set of interactions between genetic factors and environmental influences (Marginean et al. 2018; Pearl et al. 2018). Recent studies in both humans and mammal have indicated that gut microbiota are major contributors to obesity and metabolic disorders (Al-Assal et al. 2018; Bianchi et al. 2018; Bluemel et al. 2016; Gerard 2016; Meijnikman et al. 2018; Sun et al. 2018; Ussar et al. 2015). In general, the intestinal microbial diversity of obese people is less when compared with that of their lean counterparts (Ridaura et al. 2013; Sun et al. 2018). In both humans and animals obesity and metabolic phenotypes have been altered through modification of gut microbiota by antibiotic treatment, prebiotic or probiotic supplements, co-housing ,or other treatments (Bianchi et al. 2018; Ciciliot et al. 2018; Dahiya et al. 2017; Hu et al. 2017). Omics technologies including genomics, metagenomics, metabolomics, transcriptomics, and proteomics revealed the detailed physiological and genomic features of the complex gut ecosystem, which expanded our knowledge in understanding the molecular mechanism of interaction between these dietary substances and the gut environment (Yadav et al. 2018a,b; Yadav and Shukla 2017;). In the intestine, probiotic bacteria are in a competitive or symbiotic relationship with other bacteria. The mutualism of probiotics such as Lactobacillus, Bifidobacterium and Bacteroides, can prevent the colonization of pathogenic bacterias such as Enterobacteriaceae and Enterococcus spp. and slow their growth in the gut. Prebiotics such as lactulose, raffinose, xylitol, inulin, and oligofructose, which are good substrates for probiotics, can promote the growth of probiotics, and stimulate the production of enzymes or bioactive substances to enhance the ecological balance of intestinal flora (Yadav et al. 2016, 2018b). Probiotics provide various health benefits, such as immune system modification, metabolism regulation, and colonization resistance (Yadav and Shukla 2017). Research outcomes have indicated that gut microbiota might regulate obesity and related diseases by affecting energy acquisition (Damms-Machado et al. 2015) and influencing fatty acid oxidation (Dahiya et al. 2017), appetite regulation (De Vadder et al. 2014; Duraffourd et al. 2012), cholesterol, bile acid, and choline metabolism adjustment (Just 2017), host gene regulation (Scheithauer et al. 2016; Sun et al. 2018), intestinal permeability (Mokkala et al. 2016), immune system mediation (Winer et al. 2016), metabolic endotoxinemia, and the endocannabinoid system regulation (Dahiya et al. 2017).
To study the relationship between intestinal microorganisms and disease, germ-free (GF), and Ob/ob mice on C57Bl/6 background have previously been used in establishing HFD-induced obesity models (Parseus et al. 2017; Schweiger et al. 2017). The principle advantage of the GF animal model is their use in proof-of-principle studies, microbe–microbe interaction studies, and that a complete microbiota or defined consortiums of bacteria can be introduced at various developmental stages (Luczynski et al. 2016). However, the altered physiology seen in GF mice raises issues. Grover and Kashyap (2014) stated that the enlarged cecum, reduced smooth muscle tone, decreased sodium and chloride ion concentrations in cecal contents, and impaired water absorption in the colon of GF mice were strikingly different when compared to that of conventionally raised mice. Lomasney et al. (2014) further highlighted that the colonic secretomotor function is preserved in GF mice. Luczynski et al. (2016) indicated that permanent neurodevelopmental deficits of GF mouse may deem the model unsuitable for specific scientific queries (Luczynski et al. 2016). Some hypogenesis of GF mice such as lymphatic dysplasia, low immune function, abnormal sensitivity to microorganisms, and long metabolic cycle may be doubtful when studying the interaction of gut microbiota and obesity (Luo and Jin 2014; Principi and Esposito 2016; Tannock 2005; Zarrinpar et al. 2018). Therefore, alternatives and complementary strategies to the GF model are warranted. To establish control animal models with similar physiologies but lacking intestinal flora, in many studies, broad-spectrum antibiotics were used to deplete gut commensal microflora to successfully establish pseudo-germ-free rodents that allow for analyzing the relationships between intestinal microorganisms and a wide range of diseases, including obesity (Membrez et al. 2008; Suarez-Zamorano et al. 2015), tumorigenesis (Chen et al. 2008; Grivennikov et al. 2012), serotonin biosynthesis (Ge et al. 2017; Josefsdottir et al. 2017), hepatic fibrosis (Seki et al. 2007), and intestinal homeostasis (Rakoff-Nahoum et al. 2004). Janssen et al. (2017) suppressed gut bacteria of mice by using antibiotics and discovered that angiopoietin-like 4 promoted bile acid absorption via a mechanism dependent on the gut microbiota. Moreover, Hu et al. (2015) depleted gut microbiota of rats by ampicillin, neomycin and metronidazole, and fund that antibiotics-induced imbalances in gut microbiota aggravated cholesterol accumulation and liver injuries in rats that were fed a high-cholesterol diet. Indeed, in previous studies, it has been shown that metabolic phenotypes of mice that were treated with high-dose of antibiotics were similar to that of GF mice (Cox et al. 2014; Grover and Kashyap 2014; Lee et al. 2012; Suarez-Zamorano et al. 2015). GF mice were protected from developing obesity when fed a high-fat, sugar-rich diet (Le Roy et al. 2019). Suarez-Zamorano et al. (2015) showed that treatment with broad-spectrum antibiotics could promote browning of white adipose tissue and reduced obesity in mice. The metabolic responses of antibiotic-treated mice to HFD were similar to that of GF mice. Zarrinpar et al. (2018) successfully depleted the gut microbiome of mice by oral gavage with ampicillin, vancomycin, neomycin, metronidazole, and amphotericin B. The results showed that microbiome depletion changed the metabolism of mice and altered glucose homeostasis by potentially shifting colonocyte energy utilization from short chain fatty acids (SCFAs) to glucose. The lower blood glucose levels, increased insulin sensitivity, lower oral glucose tolerance test (oGTT), and decreased luminal SCFAs of antibiotic-treated mice were similar to those of GF mice. Ellekilde et al. (2014) successfully depleted gut microbiota of mice by ampicillin treatment and transferred donors’ microbial compositions to antibiotic-treated mice instead of GF mice. They found out that the obese phenotypes of donors were transferred to recipients. It is speculated that the transferred microbiota may permanently modulate host functions (Ellekilde et al. 2014). Reikvam et al. (2011) presented a robust protocol for depleting cultivatable intestinal microbiota of mice by gavage with antibiotics and showed that the biological effect of this depletion phenocopied physiological characteristics of GF mice. In fact, the primary advantage of gut microbiota depletion by antibiotics over working with GF animals in isolators is that antibiotics can be introduced and terminated at pre-determined time points(Hansen et al. 2015). Furthermore, pseudo-germ-free rodents are less expensive than GF mice. The present study was designed to evaluate whether depletion of gut microbiota using antibiotics could influence host metabolic phenotypes that are associated with obesity, and whether, this approach was suitable for generating animal models that can be used for studying the effects of gut microbiota on obesity.
Materials and methods
Animals and antibiotics treatment
Male C57Bl/6J mice (5 weeks old, 18 ± 1 g) were purchased from the Hunan SJA Laboratory Animal Co.; Ltd., (Changsha, China). Mice (8 mice per cage) were kept in a specific pathogen–free (SPF) facility at 23 ± 2 °C under 12 h/day and night cycles until the end of the experiment. Mice were fed a CD containing 46.74% (w/w) carbohydrate, 24.3% (w/w) protein, 7.5% (w/w) fat, 3.5% (w/w) fiber, and 8.6% ash (w/w) during 1 week of acclimation. Subsequently, mice were randomly assigned to 8 groups (n = 8 per group) and labeled as ATGH, ATDH, PGH, PDH, ATGC, ATDC, PGC, and PDC, respectively. Groups ATGH, ATDH, PGH, and PDH were fed a HFD containing 80.0% (w/w) chow diet supplemented with 10% (w/w) lard oil and 10% (w/w) yolk powder, whereas groups ATGC, ATDC, PGC, and PDC were fed CD for 4 weeks. Mice in groups ATGH and ATGC received non-absorbable broad-spectrum antibiotics by gavage at a dose of 20 μg/g•bw neomycin, 10 μg/g•bw vancomycin, 10 μg/g•bw imipenem, 20 μg/g•bw metronidazole, 10 μg/ g•bw streptomycin, and 20 U/ g•bw penicillin (Sangon Biotech, Shanghai, China). Mice in groups PGH and PGC served as control groups and were gavaged with equivalent volumes of distilled water. Mice in groups ATDH and ATDC were given an antibiotic cocktail supplement in the drinking water, containing 100 μg/mL neomycin, 50 μg/mL vancomycin, 50 μg/mL imipenem, 100 μg/mL metronidazole, 50 μg/mL streptomycin, and 100 U/mL penicillin, according to the method described by Suarez-Zamorano et al. (2015). Mice in groups PDH and PDC were given a distilled water cocktail supplement without antibiotics in volumes equivalent to the control groups. Water bottles were changed twice weekly to supply fresh antibiotics. Body weight, food intake, and water intake of all mice were recorded weekly. Feces of each group (n = 8) were collected and combined into one sample in a sterile microtube at day 0, 15, and 30 after starting antibiotics treatment, frozen immediately in liquid nitrogen and stored at − 80 °C until analysis. At the end of the experiment, mice were anesthetized with pentobarbital sodium and sacrificed after 5 h of fasting. Heart blood was collected and stored at − 80 °C for further analysis. Liver, epididymal adipose, and small intestine tissues were excised and fixed with 10% neutral formalin immediately at room temperature. The animal protocol used in this study was approved by the Animal Care Committee at Hunan Agricultural University and performed in accordance with the Guide for the Care and Use of Laboratory Animals published by the Ministry of Health, People’s Republic of China.
Analysis of gut microbiota
Total DNA of feces was isolated using a FastDNA® Spin Kit for Soil (Mpbio, Santa Ana, USA). The relative abundance of bacteria was determined using 16S rRNA analysis, which was performed at the laboratory of Novogene (Beijing, China). 16S rRNA genes of V3–V4 hypervariable region were amplified used specific primer 338F-806R (Primer F 5′- ACTCCTACGGGAGGCAGCAG-3′ and Primer R 5′- GGACTACHVGGGTWTCTAAT-3′). The fragment library was constructed using Ion Plus Fragment Library Kit (Thermo Scientific, Shanghai, China) 48 runs, followed by DNA sequencing on Ion S5TMXL (Thermo Scientific, Shanghai, China) platform to generate 400 bp/600 bp single-end reads. The remaining unique reads were clustered into operational taxonomic units (OTUs) based on the Silva database (SILVA) by UPARSE with 97% similarity cutoff after the raw reads were quality filtered and merged by fastx, Cutadapt, Usearch, and FLASH. Mothur was used to calculate rarefaction analysis, the community richness index and alpha diversities including Shannon and Simpson indexes. Principal component analysis (PCA) and hierarchical cluster analysis by Bray–Curtis distance matrix and average method were performed according to the abundances of OTUs by QIIME (Version1.7.0) and R software (Version 2.15.3). The 16S rRNA data were accessible in the European Nucleotide Archive (ENA) database under accession ID PRJEB29982.
Histological analysis and morphometry
Formalin-fixed liver, adipose tissue, and small intestine samples from three mice per group were paraffin embedded, sectioned at 3–6 μm, and stained with hematoxylin and eosin for histological analysis. Images were taken using a camera-equipped light microscope (Leica Ltd., Wetzlar, Germany). Sections were analyzed in a blinded manner by a trained histopathologist and the size of epididymal adipocytes was evaluated by Image-Pro Plus software (Chen et al. 2018; Suarez-Zamorano et al. 2015).
Measurement of serum parameter
Serum was prepared from blood samples from eight mice per group by centrifugation at 3000 × g for 10 min in room temperature, then stored at − 80 °C. Levels of serum total cholesterol (TC), triglyceride (TG), high-density lipoprotein cholesterol (HDL-C), low-density lipoprotein cholesterol (LDL-C), aspartate aminotransferase (AST), alanine aminotransferase (ALT), fasting blood glucose (FBG), and insulin (INS) were measured using commercially available kits (Nanjing Jiancheng Bioengineering Institute, Nanjing, China).
Statistical analysis
Data were expressed as the mean ± standard error (SEM) or box-and-whisker plots. Data were subjected to two-way ANOVA analysis and graphics presentation using GraphPad Prism software 6.0 (La Jolla, CA, USA). Differences in relative abundances of OTUs were calculated using Tukey’s honest significant difference (HSD) test by QIIME (Version1.7.0) and R software (Version 2.15.3). The correlations between relative abundances of genus and host parameters were analyzed by Spearman’s correlation (R software Version 2.15.3). Statistical significances were set as follows: *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
Results
Overall structural changes of gut microbiota in response to high-fat diet and antibiotics
To determine the importance of commensal microbiota in obesity, mice that were fed an HFD or CD were subjected to oral administration of a combination of antibiotics by gavage or cocktail supplement for four consecutive weeks. Treatment with antibiotics resulted in changes in the composition of commensal bacteria as determined by 16S rRNA analysis.
The decrease in the Shannon and Chao indexes indicated that antibiotics treatment decreased microbiota community diversity and richness when compared with the baseline values. The differences in these two indexes between mice that were fed an HFD and CD continued to exist after the treatment with antibiotics (Fig. 1a–b). The antibiotics-induced depletion effects of gut microbiota were confirmed by PCA and UPGAM analysis, showing a clear separation between antibiotics-treated and untreated groups (Fig. 1c–d).
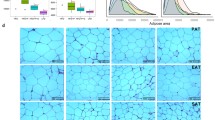
Effects of antibiotics on the high-fat diet-disrupted gut microbiota composition. Comparison of alpha diversity accessed by Chao (a) and Shannon (b) indexes; principal component analysis (PCA) of gut microbiota based on OUT relative abundance (c); relative abundance in phylum level (d); cladogram (e), generated from LEfSe analysis, represent taxa enriched in groups before antibiotics treatment (blue) and after antibiotics treatment with chow diet (CD) (red) or high-fat diet (HFD) (green). The central point represents the root of the tree (bacteria), and each ring represents the next lower taxonomic level (phylum through genus). The diameter of each circle represents the relative abundance of the taxon. When full identification was not possible, g_, or s_ alone was used for genus or species, respectively
PC1, which explained the greatest variance, segregated samples by antibiotics treatment, while PC2 segregated samples by intervention time and diet. The fecal samples of HFD-fed mice clustered together, whereas the samples from CD-fed mice comprised another group. Furthermore, after antibiotics treatment, samples in HFD groups were driven closer to those of CD groups and untreated groups in the PC2 level. These results indicated that HFD can induce striking differences in gut microbiota and that antibiotics can reduce the changes. UPGAM analysis based on unweighted unifrac distance showed that the bacterial communities of all samples were clustered into four different groups as follows: group before dietary intervention, antibiotic-treated group with HFD, antibiotic-treated group with CD, and the control group. The microbial community composition of the antibiotic-treated and non-treated groups was strikingly different, illustrating that treatment with antibiotics resulted in significant changes in the phylogenetic composition of gut microbiota. The fact that the intestinal microflora of mice under similar dietary conditions were more similar to each other indicated that HFD significantly influenced the gut microbiota of mice.
The changes of gut microbiota after antibiotics treatment were also observed in operational taxonomic unit levels. In brief, treatment with antibiotics gavage for a duration of 2 weeks decreased the OTU (optical transform unit) from 347 to 228 and 328 to 147 in mice that were fed HFD and CD, respectively. Four weeks later, the OTU were reduced to 209 and 194, respectively. Treatment with a cocktail of antibiotics decreased OTU from 327 to 146 and 360 to 183 in mice that were fed HFD and CD, respectively. Then, after 4 weeks of treatment, the OTU further reduced to 196 and 169, respectively. On the contrary, the OTU of mice that were given water without antibiotics (PDH and PDC groups) did not change.
Taxonomically, after antibiotics treatments, the abundance of fecal microbiota strikingly decreased at all levels as shown in Fig. 2b–f. At the phylum level, Bacteroidetes, Firmicutes, and Proteobacteria dominated gut microbiota and accounted for more than 90% of all bacteria, which was similar to the findings reported previously (Al-Assal et al. 2018). After supplementation with a HFD, decreased relative abundances of Bacteroidetes and Chloroflexi and increased relative abundances of Firmicutes, Proteobacteria, Verrucomicrobia, Deferribacteres, Cyanobacteria, and Saccharibacteria were observed when compared with mice in CD groups (Fig. 2b). In both HFD and CD groups, levels of Bacteroidetes increased twofold whereas Firmicutes decreased to less than half the level after 4 weeks of antibiotics treatment when compared with baseline levels. After antibiotics gavage supplemented with HFD, relative abundances of Clostridiaceae 1, Helicobacteraceae, Anaeroplasmataceae, Lachnospiraceae, and Peptostreptococcaceae decreased, whereas Veillonellaceae, Fusobacteriaceae, Peptostreptococcaceae, Bacteroidaceae, Erysipelotrichaceae, and Mycoplasmataceae increased when compared to baseline at the family level. In other antibiotics-treated groups, relative abundances of Bacteroidaceae and Verrucomicrobiaceae increased, while Lachnospiraceae and Ruminococcaceae decreased when compared to baseline levels (Fig. 3a).
Schematic overview of the experimental design, study groups, and time points of fecal sampling (a), and relative abundance of AGH0, AGH15, AGH30, ADH0, ADH15, ADH30, AGC0, AGC15, AGC30, ADC0, ADC15, ADC30, PDH15, PDC15, PDH30, and PDC30 in phylum level (b), class level (c), order level (d), family level (e), and genus level (f)
After antibiotics treatment, significant differences were observed in the LEfSe evolutionary branch diagram. Deferribacterales, Lachnospiraceae, Ruminococcaceae, Clostridia, Helicobacteraceae, Campylobacterales, and Epsilonproteobacteria could be used as biomarkers of gut microbiota in mice prior to treatment with antibiotics, whereas Bacteroidia could be a biomarker of gut microbiota in mice after antibiotic treatment with HFD and Erysiplotrichia, Enterobacteriales, Gammaproteobacteria, and Verrucomicrobiae with CD (Fig. 1e).
Antibiotics attenuated features of high-fat diet-induced metabolism
Obesity is the result of interactions between hosts, gut microbiota, and environmental factors, including diet. The composition of gut microbiota was established and developed by several physiological and environmental changes, which in turn influenced features of the host metabolism (Rastelli et al. 2018). After 4 weeks of HFD or CD, antibiotics-induced microbial perturbations expressed metabolic phenotypes. When compared with mice in CD groups, 4 weeks of HFD treatment resulted in significant increases in BWG, LW, EFW, and epididymal adipocyte size (P < 0.001) (Figs. 4, 6). Histopathological examination revealed hepatocellular microvesicular steatosis, inflammatory cell infiltration, and proliferation of intestinal mucosal epithelial cells in mice in HFD groups. These symptoms were obviously alleviated, and the number of smaller adipocytes increased after exposure to antibiotics (Figs. 5a, 6).
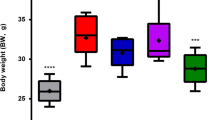
Effects of diet and antibiotics-induced microbiota perturbations on mouse food intake (a), water intake (b), body weight gain (c), liver weight (d), epididymal fat weight (e), and cell-size profiling of epididymal adipocytes (f). All values in a–e were presented as the mean ± SEM. (n = 8 per group), and points in f showed mean of pooled fractions from each animal ± SEM (n = 3 per group). Significance was calculated using two-way ANOVA (treatment and diet) with Bonferroni post hoc test. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001
Effects of diet and antibiotics-induced microbiota perturbations on serum levels of TC (a), TG (b), LDL-C (c), HDL-C (d), ALT (e), AST (f), fasting blood glucose (g), and insulin (h). All values in a–h were presented as the mean ± SEM. (n = 8 per group). Significance was calculated using two-way ANOVA (treatment and diet) with Bonferroni post hoc test. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001
As expected, in HFD-fed and CD-fed mice, antibiotics gavage significantly prevented the indicators mentioned above. However, an antibiotic cocktail supplement only prevented these indicators in mice in CD groups and reduced LW and EFW in mice in HFD groups.
Furthermore, when compared with mice in CD groups, serum levels of TC, TG, LDL-C, and ALT were significantly elevated after HFD intervention (P < 0.01) (Fig. 5a–b, e). In addition, the level of HDL-C was improved by HFD (Fig. 5d). In HFD-fed mice, gavage with antibiotics attenuated levels of TC, LDL-C, and ALT as well as HDL-C (Fig. 5a, c–e), whereas in CD-fed mice, antibiotics cocktail only decreased levels of TC and HDL-C (Fig. 5a, d). An antibiotics cocktail supplement reduced TC in both HFD-fed and CD-fed mice (Fig. 5a). Also, both antibiotic gavage and cocktail supplements led to an obvious decrease in the levels of FBG and INS, thereby indicating that microbial perturbation might improve INS sensitivity (Fig. 5g–h).
The results clearly demonstrated that intervention by antibiotics attenuated HFD-induced metabolic features even though the food intake of HFD-fed mice was less than that of CD-fed mice (Fig. 4a). Obviously, the method of gavage was more effective when compared with the use of an antibiotics cocktail supplement. This could be due to reduced water intake in mice in the antibiotics cocktail supplement group when compared to that in other groups (Fig. 4b). In addition, poor palatability of water containing antibiotics may have led to insufficient drug intake.
Correlation between gut microbiota abundance and mouse phenotype
To evaluate the correlation between the relative abundance of gut microbiota at the genus level and metabolic parameters, Spearman’s correlation analysis was employed to identify the contribution of antibiotics-induced perturbations in gut microbial diversity on metabolic phenotypes. As showed in Fig. 3b, 20 genera of microorganisms either negatively or positively associated with metabolic parameters including BWG, EFW, TC, ALT, and LDL-C. Specifically, Lachnospiraceae, Bifidobacterium, Helicobacter, Prevotellaceae_UCG.001, Anaerotruncus, Desulfovibrio, Mucispirillum, and X.Eubacterium._ xylanophilum_group positively correlated with BWG and TC significantly (P < 0.05), whereas Bacteroides showed a negative correlation. In addition, Ruminiclostridium, Alloprevotella, and Lachnospiraceae_UCG.001 positively correlated with BWG but not significantly with TC (P < 0.05). Lactobacillus, in contrast, positively correlated with TC only. Klebsiella, Escherichia, and Shigella negatively and significantly correlated with TC (P < 0.05). A significant correlation was also observed between the Rikenellaceae RC9_gut group and Faecalibaculum with EFW and LDL-C, and between the Rikenellaceae RC9 gut group and Ruminococcaceae with LDL-C (P < 0.05).
In this study, Lachnospiraceae, Ruminiclostridium, and Helicobacter were identified as biomarkers of gut microbiota in mice before antibiotics treatment and Bacteroidia as biomarker in antibiotics-treated mice (Fig. 1e). In particular, 15 days of antibiotics treatment decreased Lachnospiraceae, Ruminiclostridium, and Helicobacter by 87.29 ± 0.25%, 98.85 ± 0.99%, and 99.84 ± 0.25% respectively. Thirty days of antibiotics treatment decreased these levels by 92.95 ± 5.09%, 97.73 ± 2.09%, and 99.48 ± 0.21% respectively. However, the relative abundance of Bacteroides increased to more than twofold the original levers after antibiotics treatment (Fig. 1b, d–e).
These results indicated that HFD-induced metabolic phenotypes in mice were affected by gut microbiota, especially Lachnospiraceae, Ruminiclostridium, Helicobacter, and Bacteroidia. Yet, the mechanisms that underlie these effects need to be further elucidated.
Discussion
Gut microbiota is one of the most important factors affecting HFD-induced obesity (Al-Assal et al. 2018; Gerard 2016). In previous studies, HFD feeding has been shown to affect gut morphology and metabolic phenotypes and promote bacterial dysfunction (Roopchand 2015; Velikonja et al. 2018; Wang et al. 2014; Zhang et al. 2018a). Mice gained significantly more weight, which was associated with increased fasting glucose, insulin levels and impaired glucose and insulin tolerance when fed a HFD for 10 weeks in Parséus’s study (Parseus et al. 2017). HFD also stimulated body lipid accumulation and enlarged adipose cells (Parseus et al. 2017; Suarez-Zamorano et al. 2015). Similar to their studies, 4 weeks of HFD induced mice to be obese in this study. BWG, LW, EFW, serum levers of TC, TG, HDL-C, LDL-C, ALT, INS, and FBG were significantly increased in HFD-fed mice (P < 0.05). In addition, adipocyte hypertrophy, hepatocellular microvesicular steatosis, inflammatory cell infiltration, and proliferation of intestinal mucosal epithelium cells were observed after HFD supplementation. It has been proven that diet-induced obesity, adipose inflammation, and liver steatosis were associated with bile acid profiles and farnesoid X receptor (FXR) depending on intestinal microbes, which contributed to impaired host metabolism (Dahiya et al. 2017; Janssen et al. 2017; Parseus et al. 2017). We found that feeding mice a HFD was associated with striking changes in phylogenetic composition of gut microbiota, as shown by decreased abundances of Bacteroidetes and Chloroflexi and increased abundances of Firmicutes, Proteobacteria, Verrucomicrobia, Deferribacteres, Cyanobacteria, and Saccharibacteria. These changes in intestinal microbial composition were consistent with previous reports, in which was demonstrated that HFD-induced gut dysbiosis with high ratio of F/B may contribute to obesity (Anhe et al. 2015; Evans et al. 2014; Wang et al. 2014; Zhang et al. 2018a). In this study, we also found increased diversity and richness after feeding mice a HFD; however, this was inconsistent with the data presented in other reports (Reijnders et al. 2016; Suarez-Zamorano et al. 2015) and may be due to the combined protocol when harvested mouse fecal samples in this study.
It has previously been shown that antibiotics may cause variations in specific microorganism populations, which then affect obesity-related metabolic phenotypes (Cox et al. 2014; Reijnders et al. 2016; Suarez-Zamorano et al. 2015; Zarrinpar et al. 2018). Low-dose antibiotics exposure during maturation may alter host metabolism and promote adiposity in mice given the consistent effects of disturbed gut microbiota on the host (Cox et al. 2014). Early-life subtherapeutic antibiotic treatment has been shown to produce significant growth-promoting effects, including increased fat mass, altered metabolic hormones, enhanced hepatic metabolism, and regulated microbiota, all of which have been applied to growth promotion in agriculture (Cox et al. 2014). Similar to animals, antibiotics administration to children can significantly modify the microbiota composition by reducing phylogenetic diversity and microbial load and increase obesity at maturity (Principi and Esposito 2016; Stark et al. 2019). However, high doses of antibiotics may play a role in resisting the onset of obesity (Suarez-Zamorano et al. 2015). Numerous studies have shown that high-dose antibiotics-caused obesity resistance in mice was related to gut microbial perturbation and their mediation on the profiles of bile acids, SCFA, LPS, Angiopoietin-related protein 4 (ANGPTL4), Fasting-induced adipose factor (FIAF), eosinophil, type 2 cytokines, and M2 macrophages in the host (Ge et al. 2017; Leong et al. 2018; Rastelli et al. 2018; Suarez-Zamorano et al. 2015; Zarrinpar et al. 2018).This suggested that high doses of broad-spectrum antibiotics may inhibit HFD-induced obesity while perturbing gut microbiota. In this study, HFD-induced metabolic features, including BWG, LW, EFW, TC, LDL-C, HDL-C, ALT, INS, and FBG, were reduced by antibiotics treatment to different degrees. These phenotypic changes in the treatment groups observed in the current study were similar to those previously reported in GF mice that were fed a HFD (Kusumoto et al. 2017; Lee et al. 2012; Shin et al. 2014; Suarez-Zamorano et al. 2015; Woting et al. 2015; Zhang et al. 2015, 2018a, b, c) Hansen and Suarez-Zamorano indicated that high doses of antibiotics caused a striking decrease in intestinal microbes in mice. The disappearance of most of the flora caused them to physiologically resist diet-induced obesity as in GF mice (Hansen et al. 2015; Suarez-Zamorano et al. 2015). Ellekilde et al. (2014) transplanted intestinal microbes from obese individuals to antibiotic-treated mice. Like GF mice, the obese phenotype was transferred into antibiotic-treated mice. This indicated that the antibiotic-induced depletion and perturbation of gut microbiota might play a key role in metabolic changes in mice. Zarrinpar et al. (2018) stated that microbiome depletion by oral gavage of antibiotics changed the metabolism of mice and altered glucose homeostasis by potentially shifting colonocyte energy utilization from SCFAs to glucose. However, it has also been shown that not all effects of high dose of antibiotics on intestinal microbes were manifested as obesity resistance. The authors found that 2 weeks of antibiotics treatment caused microbiome depletion and altered metabolic homeostasis especially hypoglycemic state without altering adiposity when mice were fed a normal chow diet. Hu et al. (2015) also proved that the antibiotics induced perturbation of gut microbiota aggravated cholesterol accumulation and liver injury in rats fed a high-cholesterol diet during 4 weeks of intervention. The differences observed between studies may depend on the antimicrobial spectrum (Modi et al. 2014), dosage of antibiotics (Cox et al. 2014; Hu et al. 2015), diet used, administration time, and strains of mice (Cox et al. 2014; Hansen et al. 2015).
In this study, the gut microbial diversity, demonstrated as OTU, Chao and Shannon indexes, and phylogenetic composition as shown in PCA and UPGAM were strikingly changed by antibiotics treatment. It should be noted that the increase in Bacteroidetes and the decrease in Firmicutes associated with antibiotic-treatment were opposite of the changes caused by HFD. Our results confirmed previous studies that showed that antibiotics treatment induced F/B ratio reduction in the gut of mice and were considered to be protective against obesity (Leong et al. 2018). Reduced levels of Bacteroidetes, Firmicutes, and F/B were observed when mice were treated with antibiotics and received an HFD for 8 weeks (Carvalho et al. 2012). These changes in intestinal microbes were also found in high-cholesterol diet-fed rats after antibiotics treatment (Hu et al. 2015).
The major decrease in the F/B ratio can affect the production of a host’s key metabolites, including SCFA, bile acids, and lipopolysaccharide (LPS) and ultimately prevent the development of obesity through a series of steps in nutrient metabolism, hormonal regulation, and neuromodulation (Rastelli et al. 2018; Zarrinpar et al. 2018; Zhang et al. 2018a). The reduction in F/B ratio also associated with upregulation of regenerating islet-derived protein 3 gamma (Reg3γ) expression in intestinal epithelial cells and effectively inhibited lipogenesis in mature adipocytes (Cox et al. 2014). An HFD may cause an increase in circulating LPS levels, thereby activating the immune signaling pathway that may ultimately lead to obesity (Turnbaugh et al. 2006). Intestinal microorganisms are reservoirs of LPS, and its composition, especially the F/B ratio, may affect the concentration of bacterial LPS in the circulation (Yadav et al. 2016). LPS mediates toll-like receptor 4 (TLR4), cell surface molecule (CD14), and nuclear factor-κB (NF-κB), which can regulate obesity and other diseases (Carmody et al. 2015; Ussar et al. 2015). Moreover, Bacteroides can change the capacity and composition of bile acid stores to regulate the FXR, nuclear receptor subfamily 1, group H, member 4 (NR1H4), and G protein-coupled receptors (GPCRs), thereby regulating body fat and glucose metabolism (Tuomi et al. 2014). In addition, the decrease in Bacteroides positively correlated with the expression of FIAF, an important lipid regulating gene, in mouse intestinal epithelial cells (Kusumoto et al. 2017)
It is interesting to note that in this study the critical F/B ratio in gut microbiota was regulated by antibiotics in the direction of obesity resistance. Therefore, we could not rule out the possibility that regulation of the F/B ratio by antibiotics may affect the metabolism of HFD-fed mice in many ways and play a role in reduction of obesity.
It has previously been shown that SCFA, including butyrate, acetate, and propionate stimulate the secretion of gut peptides, such as glucagon-like peptide 1 (GLP-1) and cheese casein peptide (PYY), which can reduce the level of ghrelin and brake caloric intake (Coyte et al. 2015). SCFA can also participate in the intestinal gluconeogenesis (GNG) pathway and nerve conduction (Backhed et al. 2015), thereby resulting in increased insulin sensitivity and appetite reduction (Sivan et al. 2015). These SCFAs are produced by specific gut microbes, especially, acetate is produced by Bifidobacterium adolescentis, Bacteroides thetaiotaomicron and Ruminococcus bromii, butyrate by Eubacterium rectale and Faecalibacterium prausnitzii, and propionate by Bacteroides thetaiotaomicron (Shoaie et al. 2015).
In this study, Lachnospiraceae, Ruminiclostridium, and Helicobacter, which were identified as major biomarkers of gut microbiota in mice before antibiotic treatment, were strikingly inhibited by antibiotic treatment. They were also significantly associated with phenotypes of obesity. Therefore, the effect of antibiotic treatment on HFD-induced obesity may be associated with decreased SCFA producing bacteria such as Lachnospiraceae, Ruminiclostridium, and Helicobacter.
In conclusion, HFD-induced metabolic features, including BWG, LW, EFW, epididymal adipocyte size, and serum levels of TC, LDL-C, HDL-C, ALT, INS, and FBG were significantly reduced by oral administration of antibiotics (P < 0.05). The antibiotics-induced depletion of Lachnospiraceae, Ruminiclostridium, and Helicobacter, and the decrease in F/B ratio in the gut microbiota of mice were causal antecedents to obesity resistance. Given that these ameliorative effects of antibiotics were similar to those found in GF mice (Kusumoto et al. 2017; Lee et al. 2012; Shin et al. 2014; Suarez-Zamorano et al. 2015; Woting et al. 2015; Zhang et al. 2015, 2018a, b, c),we demonstrated that it is feasible to produce pseudo-germ-free mice by antibiotics treatment for studying the effect of intestinal microorganisms on obesity. Certainly, additional metagenomics approaches are warranted to verify metabolic pathways altered by antibiotics and to evaluate the contribution of gut microbiota on HFD-induced obesity; however, the current study is an important step in that direction.
References
Al-Assal K, Martinez AC, Torrinhas RS, Cardinelli C, Waitzberg D (2018) Gut microbiota and obesity. Clin Nutr Exp 20:60–64. https://doi.org/10.1016/j.yclnex.2018.03.001
Anhe FF, Roy D, Pilon G, Dudonne S, Matamoros S, Varin TV, Garofalo C, Moine Q, Desjardins Y, Levy E, Marette A (2015) A polyphenol-rich cranberry extract protects from diet-induced obesity, insulin resistance and intestinal inflammation in association with increased Akkermansia spp. population in the gut microbiota of mice. Gut 64(6):872–883. https://doi.org/10.1136/gutjnl-2014-307142
Backhed F, Roswall J, Peng Y, Feng Q, Jia H, Kovatcheva-Datchary P, Li Y, Xia Y, Xie H, Zhong H, Khan MT, Zhang J, Li J, Xiao L, Al-Aama J, Zhang D, Lee YS, Kotowska D, Colding C, Tremaroli V, Yin Y, Bergman S, Xu X, Madsen L, Kristiansen K, Dahlgren J, Wang J (2015) Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17(5):690–703. https://doi.org/10.1016/j.chom.2015.04.004
Bianchi F, Larsen N, Tieghi TD, Adorno MAT, Kot W, Saad SMI, Jespersen L, Sivieri K (2018) Modulation of gut microbiota from obese individuals by in vitro fermentation of citrus pectin in combination with Bifidobacterium longum BB-46. Appl Microbiol Biotechnol 102(20):8827–8840. https://doi.org/10.1007/s00253-018-9234-8
Bluemel S, Williams B, Knight R, Schnabl B (2016) Precision medicine in alcoholic and nonalcoholic fatty liver disease via modulating the gut microbiota. Am J Physiol-Gastr L 311(6):G1018–G1036. https://doi.org/10.1152/ajpgi.00245.2016
Carmody RN, Gerber GK, Luevano JM Jr, Gatti DM, Somes L, Svenson KL, Turnbaugh PJ (2015) Diet dominates host genotype in shaping the murine gut microbiota. Cell Host Microbe 17(1):72–84. https://doi.org/10.1016/j.chom.2014.11.010
Carvalho BM, Guadagnini D, Tsukumo DML, Schenka AA, Latuf-Filho P, Vassallo J, Dias JC, Kubota LT, Carvalheira JBC, Saad MJA (2012) Modulation of gut microbiota by antibiotics improves insulin signalling in high-fat fed mice. Diabetologia 55(10):2823–2834. https://doi.org/10.1007/s00125-012-2648-4
Chen G, Xie M, Wan P, Chen D, Dai Z, Ye H, Hu B, Zeng X, Liu Z (2018) Fuzhuan brick tea polysaccharides attenuate metabolic syndrome in high-fat diet induced mice in association with modulation in the gut microbiota. J Agric Food Chem 66(11):2783–2795. https://doi.org/10.1021/acs.jafc.8b00296
Chen GY, Shaw MH, Redondo G, Nunez G (2008) The innate immune receptor Nod1 protects the intestine from inflammation-induced tumorigenesis. Cancer Res 68(24):10060–10067. https://doi.org/10.1158/0008-5472.CAN-08-2061
Ciciliot S, Albiero M, Campanaro S, Poncina N, Tedesco S, Scattolini V, Dalla Costa F, Cignarella A, Vettore M, Di Gangi IM, Bogialli S, Avogaro A, Fadini GP (2018) Interplay between gut microbiota and p66Shc affects obesity-associated insulin resistance. FASEB J 32(7):4004–4015. https://doi.org/10.1096/fj.201701409R
Cox LM, Yamanishi S, Sohn J, Alekseyenko AV, Leung JM, Cho I, Kim SG, Li H, Gao Z, Mahana D, Rodriguez JGZ, Rogers AB, Robine N, Loke P, Blaser MJ (2014) Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 158(4):705–721. https://doi.org/10.1016/j.cell.2014.05.052
Coyte KZ, Schluter J, Foster KR (2015) The ecology of the microbiome: networks, competition, and stability. Science 350(6261):663–669. https://doi.org/10.1126/science.aad2602
Dahiya DK, Renuka PM, Shandilya UK, Dhewa T, Kumar N, Kumar S, Puniya AK, Shukla P (2017) Gut microbiota modulation and its relationship with obesity using prebiotic fibers and probiotics: a review. Front Microbiol 8:563–580. https://doi.org/10.3389/fmicb.2017.00563
Damms-Machado A, Mitra S, Schollenberger AE, Kramer KM, Meile T, Königsrainer A, Huson DH, Bischoff SC (2015) Effects of surgical and dietary weight loss therapy for obesity on gut microbiota composition and nutrient absorption. Biomed Res Int 2015:806248–806260. https://doi.org/10.1155/2015/806248
De Vadder F, Kovatcheva-Datchary P, Goncalves D, Vinera J, Zitoun C, Duchampt A, Backhed F, Mithieux G (2014) Microbiota-generated metabolites promote metabolic benefits via gut-brain neural circuits. Cell 156(1–2):84–96. https://doi.org/10.1016/j.cell.2013.12.016
Donohoe CL, Lysaght J, O'Sullivan J, Reynolds JV (2017) Emerging concepts linking obesity with the hallmarks of cancer. TEM 28(1):46–62. https://doi.org/10.1016/j.tem.2016.08.004
Duraffourd C, De Vadder F, Goncalves D, Delaere F, Penhoat A, Brusset B, Rajas F, Chassard D, Duchampt A, Stefanutti A, Gautier-Stein A, Mithieux G (2012) Mu-opioid receptors and dietary protein stimulate a gut-brain neural circuitry limiting food intake. Cell 150(2):377–388. https://doi.org/10.1016/j.cell.2012.05.039
Ellekilde M, Selfjord E, Larsen CS, Jakesevic M, Rune I, Tranberg B, Vogensen FK, Nielsen DS, Bahl MI, Licht TR, Hansen AK, Hansen CH (2014) Transfer of gut microbiota from lean and obese mice to antibiotic-treated mice. Sci Rep 4:5922–5930. https://doi.org/10.1038/srep05922
Evans CC, LePard KJ, Kwak JW, Stancukas MC, Laskowski S, Dougherty J, Moulton L, Glawe A, Wang Y, Leone V, Antonopoulos DA, Smith D, Chang EB, Ciancio MJ (2014) Exercise prevents weight gain and alters the gut microbiota in a mouse model of high fat diet-induced obesity. PLoS One 9(3):e92193–e92207. https://doi.org/10.1371/journal.pone.0092193
Ge X, Ding C, Zhao W, Xu L, Tian H, Gong J, Zhu M, Li J, Li N (2017) Antibiotics-induced depletion of mice microbiota induces changes in host serotonin biosynthesis and intestinal motility. J Transl Med 15(1):13–22. https://doi.org/10.1186/s12967-016-1105-4
Gerard P (2016) Gut microbiota and obesity. Cell Mol Life Sci 73(1):147–162. https://doi.org/10.1007/s00018-015-2061-5
Grivennikov SI, Wang K, Mucida D, Stewart CA, Schnabl B, Jauch D, Taniguchi K, Yu GY, Osterreicher CH, Hung KE, Datz C, Feng Y, Fearon ER, Oukka M, Tessarollo L, Coppola V, Yarovinsky F, Cheroutre H, Eckmann L, Trinchieri G, Karin M (2012) Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature 491(7423):254–258. https://doi.org/10.1038/nature11465
Grover M, Kashyap PC (2014) Germ-free mice as a model to study effect of gut microbiota on host physiology. Neurogastroenterol Motil 26(6):745–748. https://doi.org/10.1111/nmo.12366
Hansen AK, Krych L, Nielsen DS, Hansen CHF (2015) A review of applied aspects of dealing with gut microbiota impact on rodent models. ILAR J 56(2):250–264. https://doi.org/10.1093/ilar/ilv010
Hu X, Wang T, Liang S, Li W, Wu X, Jin F (2015) Antibiotic-induced imbalances in gut microbiota aggravates cholesterol accumulation and liver injuries in rats fed a high-cholesterol diet. Appl Microbiol Biotechnol 99(21):9111–9122. https://doi.org/10.1007/s00253-015-6753-4
Hu Y, Wong FS, Wen L (2017) Antibiotics, gut microbiota, environment in early life and type 1 diabetes. Pharmacol Res 119:219–226. https://doi.org/10.1016/j.phrs.2017.01.034
Janssen AWF, Dijk W, Boekhorst J, Kuipers F, Groen AK, Lukovac S, Hooiveld GJEJ, Kersten S (2017) ANGPTL4 promotes bile acid absorption during taurocholic acid supplementation via a mechanism dependent on the gut microbiota. BBA-Mol Cell Biol L 1862(10):1056–1067. https://doi.org/10.1016/j.bbalip.2017.07.005
Josefsdottir KS, Baldridge MT, Kadmon CS, King KY (2017) Antibiotics impair murine hematopoiesis by depleting the intestinal microbiota. Blood 129(6):729–739. https://doi.org/10.1182/blood-2016-03-708594
Just S (2017) Impact of the interplay between bile acids, lipids, intestinal coriobacteriaceae and diet on host metabolism. München
Kusumoto Y, Irie J, Iwabu K, Tagawa H, Itoh A, Kato M, Kobayashi N, Tanaka K, Kikuchi R, Fujita M, Nakajima Y, Morimoto K, Sugizaki T, Yamada S, Kawai T, Watanabe M, Oike Y, Itoh H (2017) Bile acid binding resin prevents fat accumulation through intestinal microbiota in high-fat diet-induced obesity in mice. Metabolism 71:1–6. https://doi.org/10.1016/j.metabol.2017.02.011
Le Roy CI, Woodward MJ, Ellis RJ, La Ragione RM, Claus SP (2019) Antibiotic treatment triggers gut dysbiosis and modulates metabolism in a chicken model of gastro-intestinal infection. BMC Vet Res 15(1):37–50. https://doi.org/10.1186/s12917-018-1761-0
Lee SH, An JH, Lee HJ, Jung BH (2012) Evaluation of pharmacokinetic differences of acetaminophen in pseudo germ-free rats. Biopharm Drug Dispos 33(6):292–303. https://doi.org/10.1002/bdd.1799
Leong KSW, Derraik JGB, Hofman PL, Cutfield WS (2018) Antibiotics, gut microbiome and obesity. Clin Endocrinol 88(2):185–200. https://doi.org/10.1111/cen.13495
Lomasney KW, Houston A, Shanahan F, Dinan TG, Cryan JF, Hyland NP (2014) Selective influence of host microbiota on cAMP-mediated ion transport in mouse colon. Neurogastroenterol Motil 26(6):887–890. https://doi.org/10.1111/nmo.12328
Luczynski P, Neufeld KAM, Oriach CS, Clarke G, Dinan TG, Cryan JF (2016) Growing up in a bubble: using germ-free animals to assess the influence of the gut microbiota on brain and behavior. Int J Neuropsychopharmacol 19(8):1–17. https://doi.org/10.1093/ijnp/pyw020
Luo J, Jin F (2014) Recent advances in understanding the impact of intestinal microbiota on host behavior. Chin Sci Bull 59(22):2169–2190doi. https://doi.org/10.1360/n972014-00120
Marginean CO, Marginean C, Melit LE (2018) New insights regarding genetic aspects of childhood obesity: a minireview. Front Pediatr 6(271):1–8. https://doi.org/10.3389/fped.2018.00271
Meijnikman AS, Gerdes VE, Nieuwdorp M, Herrema H (2018) Evaluating causality of gut microbiota in obesity and diabetes in humans. Endocr Rev 39(2):133–153. https://doi.org/10.1210/er.2017-00192
Membrez M, Blancher F, Jaquet M, Bibiloni R, Cani PD, Burcelin RG, Corthesy I, Mace K, Chou CJ (2008) Gut microbiota modulation with norfloxacin and ampicillin enhances glucose tolerance in mice. FASEB J 22(7):2416–2426. https://doi.org/10.1096/fj.07-102723
Modi SR, Collins JJ, Relman DA (2014) Antibiotics and the gut microbiota. J Clin Invest 124(10):4212–4218. https://doi.org/10.1172/JCI72333
Mokkala K, Roytio H, Munukka E, Pietila S, Ekblad U, Ronnemaa T, Eerola E, Laiho A, Laitinen K (2016) Gut microbiota richness and composition and dietary intake of overweight pregnant women are related to serum zonulin concentration, a marker for intestinal permeability. J Nutr 146(9):1694–1700. https://doi.org/10.3945/jn.116.235358
Parseus A, Sommer N, Sommer F, Caesar R, Molinaro A, Stahlman M, Greiner TU, Perkins R, Backhed F (2017) Microbiota-induced obesity requires farnesoid X receptor. Gut 66(3):429–437. https://doi.org/10.1136/gutjnl-2015-310283
Pearl RL, Wadden TA, Allison KC, Chao AM, Alamuddin N, Berkowitz RI, Walsh O, Tronieri JS (2018) Causal attributions for obesity among patients seeking surgical versus behavioral/pharmacological weight loss treatment. Obes Surg 28(11):3724–3728. https://doi.org/10.1007/s11695-018-3490-7
Portune KJ, Benitez-Paez A, Del Pulgar EM, Cerrudo V, Sanz Y (2017) Gut microbiota, diet, and obesity-related disorders-the good, the bad, and the future challenges. Mol Nutr Food Res 61(1):1–38. https://doi.org/10.1002/mnfr.201600252
Principi N, Esposito S (2016) Antibiotic administration and the development of obesity in children. Int J Antimicrob Ag 47(3):171–177. https://doi.org/10.1016/j.ijantimicag.2015.12.017
Rakoff-Nahoum S, Paglino J, Eslami-Varzaneh F, Edberg S, Medzhitov R (2004) Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell 118(2):229–241. https://doi.org/10.1016/j.cell.2004.07.002
Rastelli M, Knauf C, Cani PD (2018) Gut microbes and health: a focus on the mechanisms linking microbes, obesity, and related disorders. Obesity 26(5):792–800. https://doi.org/10.1002/oby.22175
Reijnders D, Goossens GH, Hermes GD, Neis EP, van der Beek CM, Most J, Holst JJ, Lenaerts K, Kootte RS, Nieuwdorp M, Groen AK, Olde Damink SW, Boekschoten MV, Smidt H, Zoetendal EG, Dejong CH, Blaak EE (2016) Effects of gut microbiota manipulation by antibiotics on host metabolism in obese humans: a randomized double-blind placebo-controlled trial. Cell Metab 24(1):63–74. https://doi.org/10.1016/j.cmet.2016.06.016
Reikvam DH, Erofeev A, Sandvik A, Grcic V, Jahnsen FL, Gaustad P, McCoy KD, Macpherson AJ, Meza-Zepeda LA, Johansen FE (2011) Depletion of murine intestinal microbiota: effects on gut mucosa and epithelial gene expression. PLoS One 6(3):e17996–e18009. https://doi.org/10.1371/journal.pone.0017996
Ridaura VK, Faith JJ, Rey FE, Cheng J, Duncan AE, Kau AL, Griffin NW, Lombard V, Henrissat B, Bain JR, Muehlbauer MJ, Ilkayeva O, Semenkovich CF, Funai K, Hayashi DK, Lyle BJ, Martini MC, Ursell LK, Clemente JC, Treuren WV, Walters WA, Knight R, Newgard CB, Heath AC, Gordon JI (2013) Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 341(6150):1241214–1241226. https://doi.org/10.1126/science.1241214
Roopchand DE, Carmody RN, Kuhn P, Moskal K, Rojas-Silva P, Turnbaugh PJ, Raskin I (2015) Dietary polyphenols promote growth of the gut bacterium Akkermansia muciniphila and attenuate high fat diet-induced metabolic syndrome. Diabetes 64(8):2847–2858. https://doi.org/10.2337/db14-1916
Sara-Assar MMFTMA (2017) Evidence-based psychotherapeutic interventions and mhealth for weight management in overweight: a biopsychosocial framework. Dissertation, Alliant International University
Scheithauer TP, Dallinga-Thie GM, de Vos WM, Nieuwdorp M, van Raalte DH (2016) Causality of small and large intestinal microbiota in weight regulation and insulin resistance. Mol Metab 5(9):759–770. https://doi.org/10.1016/j.molmet.2016.06.002
Schweiger M, Romauch M, Schreiber R, Grabner GF, Hütter S, Kotzbeck P, Benedikt P, Eichmann TO, Yamada S, Knittelfelder O, Diwoky C, Doler C, Mayer N, De Cecco W, Breinbauer R, Zimmermann R, Zechner R (2017) Pharmacological inhibition of adipose triglyceride lipase corrects high-fat diet-induced insulin resistance and hepatosteatosis in mice. Nat Commun Nat Commun 8:14859–14873. https://doi.org/10.1038/ncomms14859
Seki E, De Minicis S, Österreicher CH, Kluwe J, Osawa Y, Brenner DA, Schwabe RF (2007) TLR4 enhances TGF-beta signaling and hepatic fibrosis. Nat Med 13(11):1324–1332. https://doi.org/10.1038/nm1663
Shin NR, Lee JC, Lee HY, Kim MS, Whon TW, Lee MS, Bae JW (2014) An increase in the Akkermansia spp. population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut 63(5):727–735. https://doi.org/10.1136/gutjnl-2012-303839
Shoaie S, Ghaffari P, Kovatcheva-Datchary P, Mardinoglu A, Sen P, Pujos-Guillot E, de Wouters T, Juste C, Rizkalla S, Chilloux J, Hoyles L, Nicholson JK, Consortium MI-O, Dore J, Dumas ME, Clement K, Backhed F, Nielsen J (2015) Quantifying diet-induced metabolic changes of the human gut microbiome. Cell Metab 22(2):320–331. https://doi.org/10.1016/j.cmet.2015.07.001
Sivan A, Corrales L, Hubert N, Williams JB, Aquino-Michaels K, Earley ZM, Benyamin FW, Lei YM, Jabri B, Alegre ML, Chang EB, Gajewski TF (2015) Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-pd-l1 efficacy. Science 350(6264):1084–1089. https://doi.org/10.1126/science.aac4255
Stark CM, Susi A, Emerick J, Nylund CM (2019) Antibiotic and acid-suppression medications during early childhood are associated with obesity. Gut 68(1):62–69. https://doi.org/10.1136/gutjnl-2017-314971
Suarez-Zamorano N, Fabbiano S, Chevalier C, Stojanovic O, Colin DJ, Stevanovic A, Veyrat-Durebex C, Tarallo V, Rigo D, Germain S, Ilievska M, Montet X, Seimbille Y, Hapfelmeier S, Trajkovski M (2015) Microbiota depletion promotes browning of white adipose tissue and reduces obesity. Nat Med 21(12):1497–1501. https://doi.org/10.1038/nm.3994
Sun L, Ma L, Ma Y, Zhang F, Zhao C, Nie Y (2018) Insights into the role of gut microbiota in obesity: pathogenesis, mechanisms, and therapeutic perspectives. Protein Cell 9(5):397–403. https://doi.org/10.1007/s13238-018-0546-3
Tannock GW (2005) New perceptions of the gut microbiota: implications for future research. Gastroenterol Clin N Am 34(3):361–382, vii. https://doi.org/10.1016/j.gtc.2005.05.006
Tuomi T, Santoro N, Caprio S, Cai M, Weng J, Groop L (2014) The many faces of diabetes: a disease with increasing heterogeneity. Lancet 383(9922):1084–1094. https://doi.org/10.1016/s0140-6736(13)62219-9
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI (2006) An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444(7122):1027–1031. https://doi.org/10.1038/nature05414
Ussar S, Griffin NW, Bezy O, Fujisaka S, Vienberg S, Softic S, Deng L, Bry L, Gordon JI, Kahn CR (2015) Interactions between gut microbiota, host genetics and diet modulate the predisposition to obesity and metabolic syndrome. Cell Metab 22(3):516–530. https://doi.org/10.1016/j.cmet.2015.07.007
Velikonja A, Lipoglavsek L, Zorec M, Orel R, Avgustin G (2018) Alterations in gut microbiota composition and metabolic parameters after dietary intervention with barley beta glucans in patients with high risk for metabolic syndrome development. Anaerobe 55:67–77. https://doi.org/10.1016/j.anaerobe.2018.11.002
Wang J, Tang H, Zhang C, Zhao Y, Derrien M, Rocher E, van-Hylckama Vlieg JET, Strissel K, Zhao L, Obin M, Shen J (2014) Modulation of gut microbiota during probiotic-mediated attenuation of metabolic syndrome in high fat diet-fed mice. The ISME J 9(1):1–15. https://doi.org/10.1038/ismej.2014.99
Winer DA, Luck H, Tsai S, Winer S (2016) The intestinal immune system in obesity and insulin resistance. Cell Metab 23(3):413–426. https://doi.org/10.1016/j.cmet.2016.01.003
Woting A, Pfeiffer N, Hanske L, Loh G, Klaus S, Blaut M (2015) Alleviation of high fat diet-induced obesity by oligofructose in gnotobiotic mice is independent of presence of Bifidobacterium longum. Mol Nutr Food Res 59(11):2267–2278. https://doi.org/10.1002/mnfr.201500249
Yadav R, Kumar V, Baweja M, Shukla P (2018a) Gene editing and genetic engineering approaches for advanced probiotics a review. Crit Rev Food Sci 58(10):1735–1746. https://doi.org/10.1080/10408398.2016.1274877
Yadav R, Shukla P (2017) An overview of advanced technologies for selection of probiotics and their expediency: a review. Crit Rev Food Sci 57(15):3233–3242. https://doi.org/10.1080/10408398.2015.1108957
Yadav R, Singh PK, Puniya AK, Shukla P (2016) Catalytic interactions and molecular docking of bile salt hydrolase (bsh) from L. plantarum RYPR1 and its prebiotic utilization. Front Microbiol 7:2116–2123. https://doi.org/10.3389/fmicb.2016.02116
Yadav R, Singh PK, Shukla P (2018b) Metabolic engineering for probiotics and their genome-wide expression profiling. Curr Protein Pept Sci 19(1):68–74. https://doi.org/10.2174/1389203718666161111130157
Zarrinpar A, Chaix A, Xu ZJZ, Chang MW, Marotz CA, Saghatelian A, Knight R, Panda S (2018) Antibiotic-induced microbiome depletion alters metabolic homeostasis by affecting gut signaling and colonic metabolism. Nat Commun 9(1):2872–2885. https://doi.org/10.1038/S41467-018-05336-9
Zhang X, Zhao Y, Xu J, Xue Z, Zhang M, Pang X, Zhang X, Zhao L (2015) Modulation of gut microbiota by berberine and metformin during the treatment of high-fat diet-induced obesity in rats. Sci Rep 5(1):14405–14415. https://doi.org/10.1038/srep14405
Zhang X, Chen Y, Zhu J, Zhang M, Ho CT, Huang Q, Cao J (2018a) Metagenomics analysis of gut microbiota in a high fat diet-induced obesity mouse model fed with (−)-epigallocatechin 3-O-(3-O-methyl) gallate (egcg3”me). Mol Nutr Food Res 62(13):e1800274-e1800309 doi:https://doi.org/10.1002/mnfr.201800274
Zhang X, Zhang M, Ho CT, Guo X, Wu Z, Weng P, Yan M, Cao J (2018b) Metagenomics analysis of gut microbiota modulatory effect of green tea polyphenols by high fat diet-induced obesity mice model. J Funct Foods 46:268–277. https://doi.org/10.1016/j.jff.2018.05.003
Zhang X, Zhu J, Zhang X, Cheng M, Zhang Z, Cao J (2018c) The modulatory effect of nanocomplexes loaded with EGCG3″me on intestinal microbiota of high fat diet-induced obesity mice model. J Food Biochem 42(3):e12501–e12509. https://doi.org/10.1111/jfbc.12501
Funding
The authors wish to thank the Key Laboratory of Ministry of Education for Tea Science in China and the National Research Center of Engineering Technology for Utilization of Botanical Functional Ingredients in China for their financial support. This study was supported by the National Major R & D Project in China (2017YFD0400803) and China Tea Research System Project (CARS19-09B).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The animal protocol used in this study was approved by the Animal Care Committee at Hunan Agricultural University and performed in accordance with the Guide for the Care and Use of Laboratory Animals published by the Ministry of Health, People’s Republic of China.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, D., Wen, B., Zhu, K. et al. Antibiotics-induced perturbations in gut microbial diversity influence metabolic phenotypes in a murine model of high-fat diet-induced obesity. Appl Microbiol Biotechnol 103, 5269–5283 (2019). https://doi.org/10.1007/s00253-019-09764-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09764-5