Abstract
The first global genomic, proteomic, and secondary metabolomic characterization of the filamentous fungus Aspergillus nidulans following growth onboard the International Space Station (ISS) is reported. The investigation included the A. nidulans wild-type and three mutant strains, two of which were genetically engineered to enhance secondary metabolite production. Whole genome sequencing revealed that ISS conditions altered the A. nidulans genome in specific regions. In strain CW12001, which features overexpression of the secondary metabolite global regulator laeA, ISS conditions induced the loss of the laeA stop codon. Differential expression of proteins involved in stress response, carbohydrate metabolic processes, and secondary metabolite biosynthesis was also observed. ISS conditions significantly decreased prenyl xanthone production in the wild-type strain and increased asperthecin production in LO1362 and CW12001, which are deficient in a major DNA repair mechanism. These data provide valuable insights into the adaptation mechanism of A. nidulans to spacecraft environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
It has been well documented that fungal populations persist in extreme conditions, such as various temperatures and pH (López-Archilla et al. 2001; Amaral Zettler et al. 2002; Onofri et al. 2007; de Crecy et al. 2009; Gunde-Cimerman et al. 2009), desiccation (Barnard et al. 2013), ionizing radiation (Dadachova and Casadevall 2008; Tkavc et al. 2018), Mars-like conditions (de la Torre Noetzel et al. 2018; Onofri et al. 2018a; b), and the spacecraft environment (Onofri et al. 2008; Checinska et al. 2015). Through adaptation, fungi have developed the capacity to sense and respond to external stimuli, enabling their survival in a wide variety of ecological niches (Onofri et al. 2007; Onofri et al. 2008; Dadachova and Casadevall 2008; de Crecy et al. 2009; Checinska et al. 2015). These omnipresent microorganisms can be both beneficial and detrimental to human health, as fungi produce a myriad of secondary metabolites (SMs) in response to environmental stressors with activities ranging from therapeutic to toxic (Segal 2009; Newman and Cragg 2012; Sharpe et al. 2015). SMs have had a tremendous impact on human health, as the majority of small molecule drugs introduced between 1981 and 2010 were either SMs, SM derivatives, SM mimics, or possessed a SM pharmacophore. In fact, approximately 49% of all anticancer drugs are SMs or were inspired by SMs (Newman and Cragg 2012). Fungi are also potent producers of enzymes and therefore have various industrial applications (de Vries and Visser 2001; MacCabe et al. 2002). We are on the cusp of significant advances in human interplanetary space exploration, as the National Aeronautics and Space Administration (NASA) aims to send humans back to the Moon and subsequently to Mars in the 2030s. In entering this new era of human spaceflight, a thorough understanding of how fungi respond and adapt to the various stimuli encountered while in space will play an important role in ensuring a healthy living environment for crew members. Further, such studies enable the evaluation of fungi as drug production hosts during these exploration class missions, as fungi currently play an indispensable role in pharmaceutical biotechnology on Earth.
Fungi residing onboard the International Space Station (ISS) are exposed to microgravity and increased levels of radiation due to being outside of the Earth’s protective atmosphere (Horneck et al. 2010). Microgravity is thought to decrease the transfer of extracellular nutrients and metabolic by-products, which may alter the chemical environment that the cell is exposed to (Horneck et al. 2010; Huang et al. 2018). Radiation alters biological processes by acting as a promoter of mutagenesis, which may result in an increased rate of biological evolution and lead to the development of adaptive responses (Horneck et al. 2006). Although it has been established that fungi are ubiquitous in spacecraft environments (Pierson 2001; Novikova et al. 2006; Van Houdt et al. 2012; Venkateswaran et al. 2014; Checinska et al. 2015), our understanding of how fungi respond and adapt to the various conditions encountered during spaceflight remains in its infancy (Council NR 2011). The objective of this study was to investigate the changes encountered in various aspects of fungal “omics” under ISS conditions using the well-characterized organism Aspergillus nidulans. Of 30 distinct fungal species retrieved from ISS habitat surfaces during one microbial monitoring study, Aspergillus was the dominant genus, featuring a diverse population of 13 species, with A. nidulans being one of four fungal species isolated from both surfaces and air onboard the ISS (Novikova et al. 2006). Of particular interest is how the space environment alters secondary metabolism, as fungal SM production is highly variable and dependent on external stimuli (Haas 2015). A. nidulans is an extensively studied model organism and in the last decade many of the genes and regulatory networks involved in SM formation have been characterized (Yaegashi et al. 2014). This new information enabled a comprehensive investigation into genomic, proteomic, and metabolomic alterations in response to the space environment.
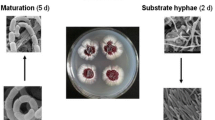
Herein, we report the multi-omic characterization of wild-type (WT) A. nidulans FGSC A4 and three mutant strains, displayed in Supplemental Table S1, following 4 and 7 days of growth onboard the ISS and compared to ground counterparts (Fig. 1). The first mutant strain, LO1362, is the A. nidulans nkuA deletion strain, which is a homolog of the human KU70 gene. These genes are crucial for nonhomologous end joining of DNA double strand break repair, and therefore deletion of nkuA disrupts a major DNA repair mechanism in A. nidulans (Krappmann 2007). The second mutant strain, LO8158, is deficient in mcrA, which is a negative regulator of at least 10 SM gene clusters (Oakley et al. 2017). Deletion of this gene stimulates SM production while impairing fungal growth. The final mutant strain used in this study was an A. nidulans laeA overexpression mutant, CW12001. LaeA is a global positive regulator of secondary metabolism, and therefore overexpression of laeA increases the production levels of a number of SMs (Bok and Keller 2004). ISS- and Earth-grown strains were subjected to genomic, proteomic, and metabolomic analyses with the aim of identifying genetic and molecular alterations that occur in fungi in response to the space environment, as such findings could pave the way to new discoveries that benefit human spaceflight and people on Earth.
Schematic overview of the A. nidulans ISS experiment. The A. nidulans wild-type (FGSC A4) and three mutant strains (LO1362, LO8158, and CW12001) were seeded onto GMM agar OmniTray plates and integrated into PHAB systems. Samples were transported to the ISS at 4 °C, where they remained for approximately 26 or 23 days until being subjected to growth in SABL systems at 37 °C for either 4 or 7 days, respectively. Earth-grown PHABs were simultaneously transferred to on-ground containers mimicking ISS SABL systems. Following growth, all samples were subjected to 4 °C and ISS-grown samples were transported to Earth. All samples were subjected to genomic, proteomic, and metabolomic analyses
Materials and methods
Strains, media, and growth conditions
The WT A. nidulans FGSC A4 strain was obtained from the Fungal Genetics Stock Center (McCluskey et al. 2010). LO1362 and LO8158 were obtained from previous studies (Oakley et al. 2017). The laeA overexpression strain (CW12001) was generated using the constitutive gpdA promoter according to standard protocol (Hunter et al. 1992). Protoplast production, construction of fusion PCR products, and transformation were carried out as described previously (Szewczyk et al. 2006; Nayak et al. 2006). Primers used are listed in Table S2 and correct transformants were verified using diagnostic PCR.
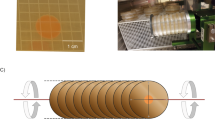
All strains were seeded onto Nunc OmniTray plates (Thermo Fisher Scientific, Waltham, MA, USA) (Supplementary Fig. S1a) containing 46 mL of solid GMM media (6 g/L NaNO3, 0.52 g/L KCl, 0.52 g/L MgSO4⋅7H2O, 1.52 g/L KH2PO4, 10 g/L d-glucose, 15 g/L agar supplemented with 1 mL/L of Hutner’s trace elements). Seeded Nunc OmniTrays were loaded into Plate Habitat (PHAB) systems (BioServe Space Technologies, Boulder, CO, USA) (Supplementary Fig. S1b), with six OmniTray plates in each PHAB, and immediately transferred to 4 °C. The PHAB system is a growth platform for biological materials that allows for gas exchange. Each PHAB was equipped with a temperature logger (HOBO) (Supplementary Fig. S1c) that accurately measured the temperature throughout the duration of the experiment. PHABs containing cultures bound for the ISS were transferred to 4 °C cold bags approximately 28 h prior to launch and transported to the ISS on the SpaceX CRS-8 (Space Exploration Technologies) mission that launched on April 8, 2016. On the ISS, PHABs were loaded into Space Automated Bioproduct Lab (SABL) systems (Supplementary Fig. S1d), where the 7- and 4-day samples remained at 4 °C for approximately 23 and 26 days, respectively. Ground culture PHABs were near-synchronously transferred to on-ground containers mimicking ISS SABL systems. To initiate growth, ISS and ground cultures were subjected to 37 °C, where they remained for either 4 or 7 days, which are optimal growth conditions for SM production in A. nidulans. Following growth, all samples were subjected to 4 °C, where they remained until ISS-grown agar plates were transported back to Earth on May 11, 2016. A more detailed explanation of space flight hardware, science preparation and loading, the flight operation timeline, and ISS environmental parameters can be found in the Supplemental Methods. All subsequent analyses of ISS- and ground-grown fungal samples, including genomic, proteomic, and metabolomic characterization, were conducted on Earth.
Genomic DNA extraction, library preparation, and genome sequencing
Mycelia were collected from Earth-grown (7-day) and space-grown (4- and 7-day) GMM agar OmniTrays for all strains (FGSC A4, LO1362, LO8158, and CW12001), frozen with liquid nitrogen, and ground using a mortar and pestle. DNA was extracted using the Mo Bio PowerMax Soil DNA Isolation Kit (Mo Bio Laboratories, Carlsbad, CA, USA) according to the manufacturer’s protocol. Library preparation and whole genome sequencing were performed at the Duke Center for Genomic and Computational Biology (Duke University Medical Center, Durham, NC, USA). DNA quality and quantity were checked using the Agilent 2100 Bioanalyzer DNA assay (Agilent Technologies, Santa Clara, CA, USA) and Qubit 2.0 Fluorometer (Thermo Fisher Scientific, Waltham, MA, USA). The library was prepared for paired-end sequencing using the TruSeq Nano DNA Library Preparation Kit (Illumina, San Diego, CA, USA). Samples were sequenced using a HiSeq 4000 Illumina Sequencer and 101 base read lengths were generated.
Genetic mutation identification
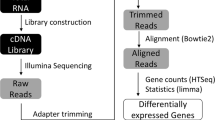
Illumina sequence reads were trimmed using Trimmomatic v 0.36 (Bolger et al. 2014) and quality was checked using FastQC v 0.11.7 (Andrews 2010). The genome and annotation files for A. nidulans FGSC A4 (Galagan et al. 2005) were downloaded from the FungiDB web portal (Stajich et al. 2012). Reads were mapped to the FGSC A4 reference genome using the Burrows-Wheeler Aligner (BWA) software package v 0.7.12 (Li and Durbin 2009) and further processed with SAMtools v 1.6 to generate sorted BAM files (Li et al. 2009). Single nucleotide polymorphisms (SNPs) and insertion/deletion mutations (INDELs) were identified using GATK v 3.7 (DePristo et al. 2011). Duplicates were marked using Picard tools MarkDuplicates (https://broadinstitute.github.io/picard/) to remove PCR artifacts. Sequence reads containing putative INDELs were realigned using GATK’s IndelRealigner to generate an updated BAM file. Variants within each sample were called using GATK’s Haplotype Caller, and the resulting Variant Call Format (VCF) files were combined using GATK’s Genotype GVCFs so that there was one VCF file for each strain (four in total). GATK’s VariantFiltration was used to filter each VCF file based on stringent cutoffs for quality and coverage {SNPs: QD < 2.0, MQ < 40.0, QUAL < 100, FS > 60.0, MQRankSum < − 12.5, SOR > 4.0, ReadPosRankSum < − 8.0; INDELs: QD < 2.0, FS > 200.0, MQRankSum < − 12.5, SOR > 4, InbreedingCoeff < − 0.8, ReadPosRankSum < − 20.0}, so that only high-quality variants remained.
Protein extraction
Mycelia were collected from Earth-grown (7-day) and space-grown (4- and 7-day) GMM agar OmniTrays for all strains (FGSC A4, LO1362, LO8158, and CW12001), frozen with liquid nitrogen, and ground using a mortar and pestle. For protein extraction, the lysis buffer consisted of 100 mM triethylammonium bicarbonate (TEAB) with 1× Halt Protease Inhibitor Cocktail (100×) (Thermo Fisher Scientific, Waltham, MA, USA) and 1 mM phenylmethylsulfonyl fluoride (Sigma-Aldrich, St. Louis, MO, USA). The frozen ground mycelia were transferred and subjected to a Precellys 24 homogenizer (Bertin Instruments, Rockville, MD, USA) in which each sample was processed inside a 2-mL cryotube with 1.0 mm glass beads for three times (at 4 °C, 6500 rpm, 1 min, repeated three times with 15 s pauses in between). The lysed cells were centrifuged at 17,000×g for 15 min. Protein concentrations in the supernatants were measured using the Bradford assay (Bio-Rad Laboratories, Inc., Hercules, CA, USA).
Tandem mass tag labeling
About 200 μg proteins from each sample were precipitated in 20% trichloroacetic acid (TCA) at 4 °C. Protein pellets were obtained by centrifugation (17,000×g), washed with ice-cold acetone, and resuspended in 25 μL TEAB (100 mM) and 25 μL 2,2,2-trifluoroethanol (TFE). Proteins were reduced by adding 1 μL of tris(2-carboxyethyl) phosphine (TCEP, 500 mM) and incubated for 1 h at 37 °C (10 mM final TCEP concentration). Proteins were alkylated in the presence of iodoacetamide (IAA, 30 mM) in the dark for 1 h at room temperature, and 2.5 μg per sample of mass spec grade trypsin/lysC (Promega, Madison, WI, USA) was used to digest the peptides overnight at 37 °C.
The digested peptides were quantified using the Pierce Quantitative Colorimetric Peptide Assay (Thermo Fisher Scientific, Waltham, MA, USA). Forty micrograms of peptides from each sample were labeled with the Thermo Scientific Tandem Mass Tag 6-plex (TMT6) Isobaric Mass Tagging Kit according to the manufacturer’s protocol. The TMT6-130 label was used as either the 4- or 7-day strains’ reference that contained 5 μg of peptides from each of the eight strains. The TMT6-131 label was used as a total mixture reference that contained 2.5 μg of peptides from each of the 16 strains. All six labeled peptide mixtures were combined into a single tube, mixed, and fractionated using the Pierce High pH Reversed-Phase Peptide Fractionation Kit (Thermo Fisher Scientific, Waltham, MA, USA). While this kit usually uses only eight fractions with step elution of up to 50% acetonitrile, we added a ninth fraction eluting at 100% acetonitrile. Nine fractionated samples were dried using a SpeedVac concentrator and resuspended in 1% formic acid prior to LC-MS/MS analysis.
LC-MS/MS analysis
The samples were analyzed on an Orbitrap Fusion Tribrid mass spectrometer with an EASY-nLC 1000 Liquid Chromatograph, a 75-μm × 2-cm Acclaim PepMap100 C18 trapping column, a 75-μm × 25-cm PepMap RSLC C18 analytical column, and an Easy-Spray ion source (Thermo Fisher Scientific, Waltham, MA, USA). The column temperature was maintained at 45 °C and the peptides were eluted at a flow rate of 300 nL/min over a 110-min gradient, from 3 to 30% solvent B (100 min), 30–50% solvent B (3 min), 50–90% solvent B (2 min), and 90% solvent B (2 min). Solvent A was 0.1% formic acid in water and solvent B was 0.1% formic acid in acetonitrile.
The full MS survey scan (m/z 400–1500) was acquired in the Orbitrap at a resolution of 120,000 and with an automatic gain control (AGC) target of 2 × 105. The maximum injection time for MS scans was 50 ms. Monoisotopic precursor ions were selected with charge states 2–7 with a ± 10-ppm mass window using a 70-s dynamic exclusion. The MS2 scan (m/z 400–2000) was performed using the linear ion trap with the CID collision energy set to 35%. The ion trap scan rate was set to “rapid,” with an AGC target of 4 × 103 and a maximum injection time of 150 ms. Ten fragment ions from each MS2 experiment were subsequently simultaneously selected for an MS3 experiment. The MS3 scan (m/z 100–500) was performed to generate the TMT reporter ions in the linear ion trap using HCD at a collision energy setting of 55%, a rapid scan rate and an AGC target of 5 × 103, and a maximum injection time of 250 ms.
Quantitative proteomics analysis
All MS/MS spectra were analyzed using the Proteome Discoverer v 2.2.0.388 (Thermo Fisher Scientific, Waltham, MA, USA) with the Sequest-HT searching engines against an Aspergillus nidulans FGSC A4 database containing 10,525 protein group sequences based on the annotated genome (NCBI, BioProject PRJEA40559, Assembly GCA_000011425.1, Release date 2009 September 24). The search was performed with the following parameters: 2 maximum missed cleavage sites, 6 minimum peptide length, 5 ppm tolerance for precursor ion masses, and 0.6 Da tolerance for fragment ion masses. The static modification settings included carbamidomethyl of cysteine residues and dynamic modifications included oxidation of methionine, TMT6plex modification of lysine ε-amino groups and peptide N-termini, and acetyl modification of protein N-terminus. A false discovery rate (FDR) of 1% for peptides and proteins was obtained using a target-decoy database search. The reporter ion integration tolerance was 0.5 Da, while the co-isolation threshold was 75%. The average signal-to-noise threshold of all reported peaks was greater than 10. The quantitative abundance of each protein was determined from the total intensity of the detected reporter ions. The ratios between the reporter and the reference reporter ion (TMT6-131) were used to estimate the abundance ratio of each protein.
For the statistical analysis, technical triplicate measurements for each protein were averaged. Only proteins that were identified and quantified with at least one peptide detected in all three technical replicates were considered for the analysis. The normalization across two biological sample sets in eight TMT experiments was carried out according to Plubell et al. (2017) with modifications. Briefly, the data from the eight TMT experiments were first corrected for small systematic differences resulting from sample loading variations and labeling efficiency by normalizing the reporter ion totals for each channel. The trimmed mean of M values (TMM) normalization corrected the compositional bias by aligning the median of the distribution of abundance intensities between samples (Robinson and Oshlack 2010). Internal reference scaling was used to adjust eight TMT data sets onto the same intensity scale. The normalized data was then averaged and log2 transformed. One-way ANOVA was preformed to identify proteins that were differentially expressed among strains in either 4 or 7 days, respectively (P value ≤ 0.05). The identified proteins were also evaluated for up- and downregulation by setting a ± 2-fold change cutoff.
Secondary metabolite extraction and analysis
Organic compounds were extracted by taking three plugs of fungal-grown agar and extracting with 3 mL methanol (MeOH), followed by 3 mL 1:1 MeOH–dichloromethane, each with 1 h of sonication and filtration. The extract was evaporated in vacuo using a rotary evaporator, redissolved in 250 μL of 20% dimethyl sulfoxide in MeOH, and a portion (10 μL) was examined by high-performance liquid chromatography-photodiode array detection-mass spectroscopy (HPLC-DAD-MS) analysis. HPLC-DAD-MS was carried out using a ThermoFinnigan LCQ Advantage ion trap mass spectrometer with a reverse-phase C18 column (3 μm; 2.1 × 100 μm; Alltech Prevail) at a flow rate of 125 μL/min. The solvent gradient for HPLC-DAD-MS was 95% MeCN/H2O (solvent B) in 5% MeCN/H2O (solvent A) both containing 0.05% formic acid, as follows: 0% solvent B from 0 to 5 min, 0 to 100% solvent B from 5 to 35 min, 100 to 0% solvent B from 40 to 45 min, and re-equilibration with 0% solvent B from 45 to 50 min. For quantification, positive-ion electrospray ionization (ESI) was used for the detection of austinol, dehydroaustinol, sterigmatocystin, nidulanin A and analogues, emericellamides A and C–F, emericellin, shamixanthone, and epishamixanthone. Negative-ion ESI was used for the detection of asperthecin, terrequinone, and sterigmatocystin intermediate. Relative production levels were quantified by integrating the area under each SM’s ESI trace.
Data availability
Raw WGS data for ISS-grown A. nidulans strains and ground-grown counterparts are available in the NCBI SRA under accession numbers SRR7724113–SRR7724124 and BioProject accession number PRJNA486827. Proteomics data is accessible through the ProteomeXchange Consortium via PRIDE with the dataset identifier PXD010778.
Results
Genome variation among ISS-grown samples
To identify genomic alterations occurring in A. nidulans strains in response to ISS conditions, whole genome paired-end sequencing (WGS) was performed on ISS-grown samples (4- and 7-day) and ground-grown control strains (7-day). Reads were aligned to the FGSC A4 reference genome and any SNP present in ground controls was removed from each strain’s sample set, which resulted in the removal of 208, 194, 231, and 224 SNPs from FGSC A4, LO1362, LO8158, and CW12001, respectively. This revealed 129, 136, 108, and 106 SNPs and 36, 39, 41, and 31 INDELs in ISS-grown FGSC A4, LO1362, LO8158, and CW12001, respectively, that occurred in ISS-grown samples when compared to ground controls, the features of which are displayed in Table 1. All identified SNPs and INDELs are summarized in Supplementary Tables S3–6, presenting FGSC A4, LO1362, LO8158, and CW12001, respectively. The total number of missense mutations observed in ISS-grown strains ranged from 9 to 15 for each strain’s sample set. These mutations were observed within only five protein-coding genes, with the same mutation often present in multiple samples (Table 2). A total of 13 unique missense base mutations were observed within AN5254, which encodes a protein containing domains predicted to be involved in RNA binding and RNA-directed DNA polymerase activity. Two unique mutations that resulted in premature stop codon were also observed within AN5254, one of which occurred in both the 4- and 7-day LO8158 and CW12001 ISS-grown samples. The remaining missense mutations occurred within AN0532, AN0535, AN0537, and AN0538, which are within a few genes away from one another in the genome. AN0532 encodes a predicted DDE1 transposable element gene, while the products of AN05235, AN0537, and AN0538 are uncharacterized. In both CW12001 ISS-grown samples, identical mutations that resulted in the loss of the original stop codon were observed within the laeA (AN0807) gene. The same frameshift stop-gain mutation independently occurred in all space-grown LO1362 and LO8158 samples within the mnpA gene (AN10311) (Supplementary Tables S4–5), which encodes a hyphal cell wall mannoprotein that may influence the surface structure (Jeong et al. 2004).
For all strains, most SNPs (> 77%) and INDELs (> 89%) occurred in intergenic regions. Most intergenic SNPs were clustered nearby several specific genes and did not appear to be strain specific (Supplementary Tables S3–6). Among all strains, many intergenic SNPs occurred near genes involved in transcription and translation, including the putative C6 transcription factor AN4972, the transcription elongation factor AN11131, the U3 small nucleolar ribonucleoprotein AN4298, the S-adenosylmethionine-dependent methyltransferase AN10829 with a predicted role in translational read-through, and AN6968 which is predicted to have RNA-directed DNA polymerase activity. Intergenic SNPs were also clustered nearby the putative alanine-tRNA ligase AN9419, the putative C4 sterol methyl oxidase AN6973 which has a predicted role in sterol metabolism, and AN9410 which has a predicted role in lipid metabolic processes. Intergenic SNPs were also observed near AN0538 and AN0539, which are within the uncharacterized cluster of genes that were reported above to possess high numbers of missense mutations. Most of the remaining intergenic SNPs occurred near AN6972, AN7848, AN10328, and AN11577, the products of which remain uncharacterized.
Proteomic profiling of ISS-grown A. nidulans
To investigate alterations in the proteome of A. nidulans strains following growth on the ISS, total protein was extracted from two biological replicates of each ISS-grown sample and Earth-grown counterpart. All samples were subjected to TMT labeling and LC-MS analysis. The resulting MS data were analyzed using Proteome Discoverer with the Sequest-HT search engine against the A. nidulans FGSC A4 protein database (NCBI). The abundance ratios for all ISS-grown samples were normalized to their Earth-grown counterparts (Supplementary Table S7, 4-day samples; Supplemental Table S8, 7-day samples), which led to the identification of up- and downregulated proteins (fold change (FC) > ∣2∣, P < 0.05) in response to the ISS environment (Fig. 2a). Interestingly, only two proteins, QutC (AN1140), which is involved in quinic acid utilization, and AN2704, a putative aryl-alcohol oxidase-related protein, were upregulated in the A. nidulans WT strain. The number of proteins upregulated and downregulated in the three ISS-grown mutant strains ranged from 4 to 28 and 2 to 77, respectively, of 10,525 predicted proteins in A. nidulans FGSC A4 in total. Distribution of AspGD Gene Ontology (GO) Slim terms (Arnaud et al. 2010) among the differentially expressed proteins for ISS-grown mutant strains is displayed in Fig. 2b–d. The GO Slim categories that possessed the highest number of differentially expressed proteins in LO1362 were stress response and carbohydrate metabolic processes. Similarly, in both LO8158 and CW12001, most differentially expressed proteins were involved in carbohydrate metabolism.
Overview of proteomic analysis. a Number of up- and downregulated proteins in ISS-grown strains (FC > |2|, P < 0.05) compared to ground-grown counterparts. b–d Biological process GO Slim categories of differentially expressed proteins. Differentially expressed proteins in b LO1362, c LO8158, and d CW12001 were mapped to terms representing various biological processes using AspGD Gene Ontology (GO) Slim Mapper
Our study revealed differential abundance of proteins involved in the A. nidulans stress response following growth on the ISS (Table 3). The heat shock protein Hsp20 (AN10507) was highly affected by ISS conditions, displaying a 2- and 4-fold increase in protein abundance in LO1362 and CW12001, respectively, and a 2-fold decrease in protein abundance in LO8158. Induction of Hsp20 has been reported following exposure to osmotic stress in A. nidulans (Wu et al. 2016). Conversely, the osmotic stress response protein CipB (AN7895) was downregulated in LO1362, and the chaperone/heat shock protein Awh11 (AN3725) was downregulated in LO1362 and LO8158. Differential abundance of proteins involved in oxidative stress response was observed among all ISS-grown strains relative to Earth-grown counterparts. The glutathione S-transferase GstB (AN6024), which has been reported to be significantly induced in response to menadione-induced oxidative stress (Pusztahelyi et al. 2011), was upregulated over 2-fold in LO8158 but downregulated 1.7- and 1.8-fold in LO1362 and CW12001, respectively, after 7 days of growth on the ISS. Two catalases, CatA (AN8637) and AN8553, exhibited decreased protein abundance in LO8158 and increased protein abundance in CW12001. Similarly, both the nitrosative stress response protein AN2470 and the menadione stress-induced protein AN5564 were downregulated approximately 3-fold in LO8158 and upregulated approximately 1.7-fold in CW12001. Upregulation was observed for phosphatidylinositol phospholipase C AN3636 whose ortholog plays a major role in responding to nutrient deprivation in Candida albicans (Uhl et al. 2003). Additionally, the GPI-anchored protein EcmA (AN4390), whose ortholog plays a major role in cell wall integrity, morphogenesis, and virulence, was upregulated 2-fold in LO8158 after 7 days of growth on the ISS (Martinez-Lopez et al. 2004).
Among all strains, proteins involved in secondary metabolism were differentially expressed in response to ISS conditions (Table 3). The polyketide synthase AptA (AN6000), which is required for biosynthesis of the spore pigment asperthecin (Szewczyk et al. 2008), was upregulated over 2-fold in CW12001 and nearly 2-fold in FGSC A4 strain after 7 days of growth on the ISS. LO1362 also exhibited upregulation of the monooxygenase MdpD (AN0147) after 7 days of growth. The MdpD protein is the product of a gene in the prenyl xanthone gene cluster, which is responsible for the production of monodictyphenone, emericellin, shamixanthone, and epishamixanthone (Chiang et al. 2010; Sanchez et al. 2011). Interestingly, the opposite was observed in CW12001, as MdpJ (AN10038), which is the product of another gene in the prenyl xanthone gene cluster, exhibited decreased protein abundance in ISS-grown samples after 4 days. A similar trend was observed for proteins involved in the biosynthesis of the potent carcinogenic mycotoxin sterigmatocystin (ST) (Fujii et al. 1976; Brown et al. 1996b). ST gene cluster products AN7817 and StcN (AN7812) exhibited increased protein abundance in LO8158, while StcN was downregulated in CW12001, following growth in ISS conditions.
The proteome of ISS-grown A. nidulans samples also revealed differential levels of proteins involved in carbohydrate metabolism when compared to Earth-grown controls (Table 3). Many glycoside hydrolases involved in carbohydrate degradation processes were upregulated more than 2-fold in ISS-grown LO8158 samples, including beta-1,4-endoglucanase AN8068 and beta-glucosidase BglM (AN7396), both of which are involved in cellulose degradation; alpha-arabinofuranosidase AbfC (AN1277), involved in pectin degradation; endo-1,4-beta-xylanase XlnC (AN1818), involved in xylan degradation; and beta-galactosidase LacA (AN0756), involved in xyloglucan, xylan, pectin, and galactomannan degradation. These glycoside hydrolases also exhibited decreased protein abundance in CW12001 when compared to ground counterparts. A similar trend was observed with alcohol oxidase AN0567, beta-glycosidase AN10124, ketose-1,6-bisphosphate aldolase AN2334, dehydratase AN6035, and rhamnogalacturonan lyase RglA (AN7135), which were upregulated at least 2-fold in ISS-grown LO8158 samples and downregulated at least 1.5-fold in CW12001. Conversely, mannose-6-phosphate isomerase AN1715, isocitrate lyase AcuD (AN5634), phosphoenolpyruvate carboxykinase AcuF (AN1918), and NAD+ dependent glycerol 3-phosphate dehydrogenase GfdB (AN6792) were downregulated at least 2.5-fold in ISS-grown LO8158 samples and upregulated at least 1.8-fold in CW12001 samples.
Secondary metabolome alterations in ISS-grown A. nidulans
Alterations in SM production of A. nidulans in response to ISS conditions were assessed by extracting SMs from three biological replicates of each ISS- and Earth-grown counterpart, which were analyzed using HPLC-DAD-MS. All SMs were identified based on mass, UV absorption, and retention time, which led to the identification of austinol and dehydroaustinol (Lo et al. 2012), terrequinone (Bok et al. 2006), sterigmatocystin and its intermediate (Brown et al. 1996a), nidulanin A and its analogues (Andersen et al. 2013), emericellamides (Chiang et al. 2008), prenyl xanthones (Sanchez et al. 2011), and asperthecin (Sanchez et al. 2012) (Supplementary Fig. S2). Relative differences in SM production levels of ISS-grown samples and Earth-grown counterparts were quantified by integrating the area under each SM’s EIC trace (Supplementary Fig. S3). ISS conditions induced asperthecin production in LO1362 and CW12001 after 7 days of growth, with production levels increased by over 300 and 150%, respectively (Fig. 3). Production levels of prenyl xanthones decreased approximately 5-fold in FGSC A4 ISS-grown samples (Supplementary Fig. S3f). In LO8158 samples, emericellamide and terrequinone production decreased (Supplementary Fig. S3b&e), while nidulanin and sterigmatocystin production increased in ISS-grown samples (Supplementary Fig. S3c&d).
Asperthecin production in ISS-grown LO1362 and CW12001. a LC-MS profiles depicting asperthecin production after 7 days of growth on the ISS, as detected by UV total scan. b Quantification of asperthecin production showing percent change for ISS-grown samples relative to Earth-grown counterparts. Significance was determined using Welch’s t test. c Chemical structure of asperthecin
Discussion
Although the persistence of fungi within space vessels is well documented and unavoidable (Novikova 2004; Novikova et al. 2006; Checinska et al. 2015), little is understood about how fungi respond and adapt to spacecraft conditions, such as microgravity and enhanced radiation. To date, most microbiological studies conducted in such environments have focused on changes occurring within bacteria or the microbiome as a whole (Tixador et al. 1985; Wilson et al. 2008; Checinska et al. 2015; Mora et al. 2016; Huang et al. 2018). Additionally, despite the various therapeutic and industrial applications of SMs, few studies have analyzed the global influence of space conditions on fungal secondary metabolism, as previous investigations have often focused on the production of a single SM (Lam et al. 1998; Lam et al. 2002; Benoit et al. 2006). Therefore, with the duration of space missions expected to increase, a major goal of this study was to investigate space-induced alterations in fungal “omics” to identify specific adaptation biomarkers and acquire insight into potential benefits of fungi that may not be discernable using traditional methodology.
This study revealed that the spacecraft environment alters the A. nidulans genome in specific regions. Five protein-coding genes displayed signatures of positive selection in the form of a high ratio of nonsynonymous to synonymous SNPs across all ISS-grown samples. These data stand in agreement with a study that investigated genomic alterations occurring in the bacterium Staphylococcus aureus during spaceflight, in which missense mutations were clustered and occurred within only nine protein-coding genes (Guo et al. 2015). High numbers of intergenic mutations were clustered near genes encoding transcriptional and translational machinery. One gene with several nonsynonymous and two unique stop-gain mutations encodes a putative retrotransposon, suggesting that its suppression confers selective advantage during growth in spacecraft environments. Other missense and intergenic mutations were clustered within and around a specific region of the genome (AN0532–AN0538), suggesting that it underwent positive selection and therefore plays a role in adapting to the space environment. One of these genes also encoded an uncharacterized transposable element gene, underscoring the significance of alterations in transposable element activity in response to growth in ISS conditions. These findings are consistent with the results from the aforementioned S. aureus study, as variations were also observed within a putative transposase (Guo et al. 2015). Interestingly, transposable element activity has been associated with stress response due to the novel variation it introduces into the genome (Capy et al. 2000). Future work should focus on characterizing the activities of the rest of these genes, as many of their functions remain unknown. Such knowledge may provide information key to elucidating the genetic adaptation mechanisms of fungi residing in spacecraft environments.
Correlations linking genomic and proteomic data were observed with the two strains genetically engineered to increase SM production. Exposure to the space environment appeared to curtail enhanced SM production in the laeA overexpression strain through the introduction of a point mutation that resulted in loss of the laeA stop codon, thereby activating nonstop decay degradation of laeA mRNA (van Hoof et al. 2002; Klauer and van Hoof 2012). The ISS-induced downregulation of laeA was also observed in the proteome of those samples, which exhibited an expression profile opposite that of the mcrA deletion strain for many proteins. Proteins involved in SM biosynthesis pathways regulated by laeA exhibited decreased and increased abundance in ISS-grown CW12001 and LO8158, respectively. The extensively studied protein LaeA forms a nuclear complex with VeA and VelB that coordinates secondary metabolism regulation with fungal development (Bayram et al. 2010). LaeA is often referred to as a global regulator of secondary metabolism (Bok and Keller 2016) and has been reported to also influence proteins involved in carbohydrate metabolism and oxidative stress response (Lv et al. 2018). Accordingly, several proteins involved in carbohydrate and antioxidant metabolic processes exhibited opposite protein expression profiles in LO8158 and CW12001. Regulation by laeA has been reported for some of these proteins in Aspergillus flavus, including endo-1,4-beta-xylanase, beta-glycoside, endo-beta-1,4-glucanase, and oxidative stress response proteins CatA and GstB (Lv et al. 2018). These findings verify the extensive regulatory role of laeA and highlight the complexity involved in identifying the laeA-controlled processes responsible for conferring selective advantage of the observed laeA stop lost mutation. This is compounded by reports that laeA also alters chromatin remodeling, cell growth and metabolism, conidiogenesis, conidial-chain elongation, sporulation, pigmentation, and colony hydrophobicity in various fungal species (Kosalková et al. 2009; Chang et al. 2012; Brakhage 2013; Lv et al. 2018). It is also possible that curtailing laeA overexpression is favored in ISS conditions to reduce energy expenditures in a stressful environment.
ISS conditions significantly increased production of asperthecin, an anthraquinone pigment, in LO1362 and CW12001. Both mutants are deficient in NkuA production, which facilitates nonhomologous end-joining (NHEJ) DNA repair, the favored DNA double strand break repair pathway in filamentous fungi (Krappmann 2007). We had anticipated that NkuA-deficient strains would be particularly interesting in flight studies due to impairment of a preferred DNA repair pathway in a mutagenic, high-radiation environment. Additionally, deletion of the nkuA homolog has been reported to increase sensitivity toward gamma irradiation in other fungal species (Meyer et al. 2007). Although we cannot discriminate the specific ISS condition that induced asperthecin production in LO1362, we hypothesize that it served as an alternative protective mechanism from the high levels of radiation present on the ISS. Our observation stays in agreement with other reports suggesting that pigment production is a key adaptive response of fungi exposed to similar environments (Volz and Dublin 1973; Singaravelan et al. 2008; Dadachova and Casadevall 2008). It is therefore possible that asperthecin can be used to protect other forms of life present on the spacecraft.
Cultivating plants for food in space will be crucial for the success of future space missions. However, space radiation can generate mutations in plant DNA, including base substitutions, deletions, and chromosomal alterations, which can result in genetic changes in seeds or tissue damage (Micco et al. 2011; Arena et al. 2014). The transformation of asperthecin biosynthesis genes into plants may potentially minimize plant DNA damage and optimize plant and astronaut health. Future studies should be conducted both on the ground and on the ISS to verify this hypothesis. Interestingly, space conditions did not increase asperthecin production in LO8158, which also possesses the nkuA- genetic background. Global regulation of SM production was altered to increase SM production in both LO8158 and CW12001, but the genetic alteration was reversed through a stop lost point mutation only in CW12001. It is therefore possible that alternative metabolomic protective mechanisms were sufficient in LO8158, and therefore, asperthecin production was not enhanced in ISS conditions.
Only a small proportion of proteins were differentially expressed in ISS-grown A. nidulans samples, which emphasizes the potential and safety of A. nidulans as a therapeutic production host during outer space missions. This finding may not hold true across the Aspergillus genus, as increased virulence has been reported in Aspergillus fumigatus strains isolated from the ISS (Knox et al. 2016). Currently, if pharmaceutical stocks in space are depleted, a small, unmanned spacecraft is launched to restock crew supplies. In the era of long-term space travel, the duration of future space missions is expected to drastically increase. The inability to deliver required drugs to astronauts in a timely manner could result in serious complications. Through heterologous expression of specific genes, A. nidulans introduces the ability to biosynthesize a wide range of pharmaceutical drugs within a week, which could significantly improve astronauts’ safety during long-term manned space missions.
In summary, this work has revealed the multi-omic response of the well-characterized model filamentous fungus A. nidulans to spacecraft conditions. These findings illustrate the potential of asperthecin to confer radiation resistance and of A. nidulans to be utilized as a small molecule production host in space. Further, specific genetic mutations involved in the adaptive mechanism of fungi in space environments were identified. Such knowledge may be valuable to NASA’s Space Biology Program (https://www.nasa.gov/spacebio) in planning for future outer space explorations.
References
Amaral Zettler LA, Gómez F, Zettler E, Keenan BG, Amils R, Sogin ML (2002) Microbiology: eukaryotic diversity in Spain’s River of Fire. Nature 417:137. https://doi.org/10.1038/417137a
Andersen MR, Nielsen JB, Klitgaard A, Petersen LM, Zachariasen M, Hansen TJ, Blicher LH, Gotfredsen CH, Larsen T, Nielsen KF, Mortensen UH (2013) Accurate prediction of secondary metabolite gene clusters in filamentous fungi. Proc Natl Acad Sci 110:E99–E107. https://doi.org/10.1073/pnas.1205532110
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data
Arena C, De Micco V, Macaeva E, Quintens R (2014) Space radiation effects on plant and mammalian cells. Acta Astronaut 104:419–431. https://doi.org/10.1016/j.actaastro.2014.05.005
Arnaud MB, Chibucos MC, Costanzo MC, Crabtree J, Inglis DO, Lotia A, Orvis J, Shah P, Skrzypek MS, Binkley G, Miyasato SR, Wortman JR, Sherlock G (2010) The Aspergillus Genome Database, a curated comparative genomics resource for gene, protein and sequence information for the Aspergillus research community. Nucleic Acids Res 38:D420–D427. https://doi.org/10.1093/nar/gkp751
Barnard RL, Osborne CA, Firestone MK (2013) Responses of soil bacterial and fungal communities to extreme desiccation and rewetting. ISME J 7:2229–2241. https://doi.org/10.1038/ismej.2013.104
Bayram ÖS, Bayram Ö, Valerius O, Park HS, Irniger S, Gerke J, Ni M, Han K-H, Yu J-H, Braus GH (2010) LaeA control of velvet family regulatory proteins for light-dependent development and fungal cell-type specificity. PLoS Genet 6:e1001226. https://doi.org/10.1371/journal.pgen.1001226
Benoit MR, Li W, Stodieck LS, Lam KS, Winther CL, Roane TM, Klaus DM (2006) Microbial antibiotic production aboard the International Space Station. Appl Microbiol Biotechnol 70:403–411. https://doi.org/10.1007/s00253-005-0098-3
Bok JW, Keller NP (2004) LaeA, a regulator of secondary metabolism in Aspergillus spp. Eukaryot Cell 3:527–535
Bok JW, Keller NP (2016) Insight into fungal secondary metabolism from ten years of LaeA research. In: Biochemistry and molecular biology. Springer, Cham, pp 21–29
Bok JW, Hoffmeister D, Maggio-Hall LA, Murillo R, Glasner JD, Keller NP (2006) Genomic mining for Aspergillus natural products. Chem Biol 13:31–37. https://doi.org/10.1016/j.chembiol.2005.10.008
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Brakhage AA (2013) Regulation of fungal secondary metabolism. Nat Rev Microbiol 11:21–32. https://doi.org/10.1038/nrmicro2916
Brown DW, Adams TH, Keller NP (1996a) Aspergillus has distinct fatty acid synthases for primary and secondary metabolism. Proc Natl Acad Sci 93:14873–14877. https://doi.org/10.1073/pnas.93.25.14873
Brown DW, Yu JH, Kelkar HS, Fernandes M, Nesbitt TC, Keller NP, Adams TH, Leonard TJ (1996b) Twenty-five coregulated transcripts define a sterigmatocystin gene cluster in Aspergillus nidulans. Proc Natl Acad Sci 93:1418–1422. https://doi.org/10.1073/pnas.93.4.1418
Capy P, Gasperi G, Biémont C, Bazin C (2000) Stress and transposable elements: co-evolution or useful parasites? Heredity 85:101–106. https://doi.org/10.1046/j.1365-2540.2000.00751.x
Chang P-K, Scharfenstein LL, Ehrlich KC, Wei Q, Bhatnagar D, Ingber BF (2012) Effects of laeA deletion on Aspergillus flavus conidial development and hydrophobicity may contribute to loss of aflatoxin production. Fungal Biol 116:298–307. https://doi.org/10.1016/j.funbio.2011.12.003
Checinska A, Probst AJ, Vaishampayan P, White JR, Kumar D, Stepanov VG, Fox GE, Nilsson HR, Pierson DL, Perry J, Venkateswaran K (2015) Microbiomes of the dust particles collected from the International Space Station and Spacecraft Assembly Facilities. Microbiome 3:50. https://doi.org/10.1186/s40168-015-0116-3
Chiang Y-M, Szewczyk E, Nayak T, Davidson AD, Sanchez JF, Lo H-C, Ho W-Y, Simityan H, Kuo E, Praseuth A, Watanabe K, Oakley BR, Wang CCC (2008) Molecular genetic mining of the Aspergillus secondary metabolome: discovery of the emericellamide biosynthetic pathway. Chem Biol 15:527–532. https://doi.org/10.1016/j.chembiol.2008.05.010
Chiang Y-M, Szewczyk E, Davidson AD, Entwistle R, Keller NP, Wang CCC, Oakley BR (2010) Characterization of the Aspergillus nidulans monodictyphenone gene cluster. Appl Environ Microbiol 76:2067–2074. https://doi.org/10.1128/AEM.02187-09
Council NR (2011) Recapturing a future for space exploration: life and physical sciences research for a new era
de Crecy E, Jaronski S, Lyons B, Lyons TJ, Keyhani NO (2009) Directed evolution of a filamentous fungus for thermotolerance. BMC Biotechnol 9:74. https://doi.org/10.1186/1472-6750-9-74
Dadachova E, Casadevall A (2008) Ionizing radiation: how fungi cope, adapt, and exploit with the help of melanin. Curr Opin Microbiol 11:525–531. https://doi.org/10.1016/j.mib.2008.09.013
DePristo MA, Banks E, Poplin RE, Garimella KV, Maguire JR, Hartl C, Philippakis AA, del Angel G, Rivas M, Hanna M, McKenna A, Fennell TJ, Kernytsky AM, Sivachenko AY, Cibulskis K, Gabriel SB, Altshuler D, Daly MJ (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43:491–498. https://doi.org/10.1038/ng.806
Fujii K, Kurata H, Odashima S, Hatsuda Y (1976) Tumor induction by a single subcutaneous injection of sterigmatocystin in newborn mice. Cancer Res 36:1615–1618
Galagan JE, Calvo SE, Cuomo C, Ma L-J, Wortman JR, Batzoglou S, Lee S-I, Baştürkmen M, Spevak CC, Clutterbuck J, Kapitonov V, Jurka J, Scazzocchio C, Farman M, Butler J, Purcell S, Harris S, Braus GH, Draht O, Busch S, D’Enfert C, Bouchier C, Goldman GH, Bell-Pedersen D, Griffiths-Jones S, Doonan JH, Yu J, Vienken K, Pain A, Freitag M, Selker EU, Archer DB, Peñalva MA, Oakley BR, Momany M, Tanaka T, Kumagai T, Asai K, Machida M, Nierman WC, Denning DW, Caddick M, Hynes M, Paoletti M, Fischer R, Miller B, Dyer P, Sachs MS, Osmani SA, Birren BW (2005) Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. Nature 438:1105–1115. https://doi.org/10.1038/nature04341
Gunde-Cimerman N, Ramos J, Plemenitas A (2009) Halotolerant and halophilic fungi. Mycol Res 113:1231–1241. https://doi.org/10.1016/j.mycres.2009.09.002
Guo J, Han N, Zhang Y, Wang H, Zhang X, Su L, Liu C, Li J, Chen C, Liu C (2015) Use of genome sequencing to assess nucleotide structure variation of Staphylococcus aureus strains cultured in spaceflight on Shenzhou-X, under simulated microgravity and on the ground. Microbiol Res 170:61–68. https://doi.org/10.1016/j.micres.2014.09.001
Haas H (2015) Microbial ecology: how to trigger a fungal weapon. eLife 4:e10504. https://doi.org/10.7554/eLife.10504
van Hoof A, Frischmeyer PA, Dietz HC, Parker R (2002) Exosome-mediated recognition and degradation of mRNAs lacking a termination codon. Science 295:2262–2264. https://doi.org/10.1126/science.1067272
Horneck G, Baumstark-Khan C, Facius R (2006) Radiation biology. In: Clément G, Slenzka K (eds) Fundamentals of space biology. Springer, New York, pp 291–336
Horneck G, Klaus DM, Mancinelli RL (2010) Space microbiology. Microbiol Mol Biol Rev 74:121–156. https://doi.org/10.1128/MMBR.00016-09
Huang B, Li D-G, Huang Y, Liu C-T (2018) Effects of spaceflight and simulated microgravity on microbial growth and secondary metabolism. Mil Med Res 5:18. https://doi.org/10.1186/s40779-018-0162-9
Hunter GD, Bailey CR, Arst HN (1992) Expression of a bacterial aspartase gene in Aspergillus nidulans: an efficient system for selecting multicopy transformants. Curr Genet 22:377–383. https://doi.org/10.1007/BF00352439
Jeong H-Y, Chae K-S, Whang SS (2004) Presence of a mannoprotein, MnpAp, in the hyphal cell wall of Aspergillus nidulans. Mycologia 96:52–56. https://doi.org/10.1080/15572536.2005.11832996
Klauer AA, van Hoof A (2012) Degradation of mRNAs that lack a stop codon: a decade of nonstop progress. Wiley Interdiscip Rev RNA 3:649–660. https://doi.org/10.1002/wrna.1124
Knox BP, Blachowicz A, Palmer JM, Romsdahl J, Huttenlocher A, Wang CCC, Keller NP, Venkateswaran K (2016) Characterization of Aspergillus fumigatus isolates from air and surfaces of the International Space Station. mSphere 1:e00227–e00216. https://doi.org/10.1128/mSphere.00227-16
Kosalková K, García-Estrada C, Ullán RV, Godio RP, Feltrer R, Teijeira F, Mauriz E, Martín JF (2009) The global regulator LaeA controls penicillin biosynthesis, pigmentation and sporulation, but not roquefortine C synthesis in Penicillium chrysogenum. Biochimie 91:214–225. https://doi.org/10.1016/j.biochi.2008.09.004
Krappmann S (2007) Gene targeting in filamentous fungi: the benefits of impaired repair. Fungal Biol Rev 21:25–29. https://doi.org/10.1016/j.fbr.2007.02.004
de la Torre Noetzel R, Miller AZ, de la Rosa JM, Pacelli C, Onofri S, García Sancho L, Cubero B, Lorek A, Wolter D, de Vera JP (2018) Cellular responses of the lichen Circinaria gyrosa in Mars-like conditions. Front Microbiol 9. https://doi.org/10.3389/fmicb.2018.00308
Lam KS, Mamber SW, Pack EJ, Forenza S, Fernandes PB, Klaus DM (1998) The effects of space flight on the production of monorden by Humicola fuscoatra WC5157 in solid-state fermentation. Appl Microbiol Biotechnol 49:579–583
Lam KS, Gustavson DR, Pirnik DL, Pack E, Bulanhagui C, Mamber SW, Forenza S, Stodieck LS, Klaus DM (2002) The effect of space flight on the production of actinomycin D by Streptomyces plicatus. J Ind Microbiol Biotechnol 29:299–302. https://doi.org/10.1038/sj.jim.7000312
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, 1000 Genome Project Data Processing Subgroup (2009) The Sequence Alignment/Map format and SAMtools. Bioinforma Oxf Engl 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Lo H-C, Entwistle R, Guo C-J, Ahuja M, Szewczyk E, Hung J-H, Chiang Y-M, Oakley BR, Wang CCC (2012) Two separate gene clusters encode the biosynthetic pathway for the meroterpenoids austinol and dehydroaustinol in Aspergillus nidulans. J Am Chem Soc 134:4709–4720. https://doi.org/10.1021/ja209809t
López-Archilla AI, Marin I, Amils R (2001) Microbial community composition and ecology of an acidic aquatic environment: the Tinto River. Spain Microb Ecol 41:20–35. https://doi.org/10.1007/s002480000044
Lv Y, Lv A, Zhai H, Zhang S, Li L, Cai J, Hu Y (2018) Insight into the global regulation of laeA in Aspergillus flavus based on proteomic profiling. Int J Food Microbiol 284:11–21. https://doi.org/10.1016/j.ijfoodmicro.2018.06.024
MacCabe AP, Orejas M, Tamayo EN, Villanueva A, Ramón D (2002) Improving extracellular production of food-use enzymes from Aspergillus nidulans. J Biotechnol 96:43–54
Martinez-Lopez R, Monteoliva L, Diez-Orejas R, Nombela C, Gil C (2004) The GPI-anchored protein CaEcm33p is required for cell wall integrity, morphogenesis and virulence in Candida albicans. Microbiology 150:3341–3354. https://doi.org/10.1099/mic.0.27320-0
McCluskey K, Wiest A, Plamann M (2010) The Fungal Genetics Stock Center: a repository for 50 years of fungal genetics research. J Biosci 35:119–126. https://doi.org/10.1007/s12038-010-0014-6
Meyer V, Arentshorst M, El-Ghezal A, Drews A-C, Kooistra R, van den Hondel CAMJJ, Ram AFJ (2007) Highly efficient gene targeting in the Aspergillus niger kusA mutant. J Biotechnol 128:770–775. https://doi.org/10.1016/j.jbiotec.2006.12.021
Micco VD, Arena C, Pignalosa D, Durante M (2011) Effects of sparsely and densely ionizing radiation on plants. Radiat Environ Biophys 50:1–19. https://doi.org/10.1007/s00411-010-0343-8
Mora M, Perras A, Alekhova TA, Wink L, Krause R, Aleksandrova A, Novozhilova T, Moissl-Eichinger C (2016) Resilient microorganisms in dust samples of the International Space Station—survival of the adaptation specialists. Microbiome 4:65. https://doi.org/10.1186/s40168-016-0217-7
Nayak T, Szewczyk E, Oakley CE, Osmani A, Ukil L, Murray SL, Hynes MJ, Osmani SA, Oakley BR (2006) A versatile and efficient gene-targeting system for Aspergillus nidulans. Genetics 172:1557–1566. https://doi.org/10.1534/genetics.105.052563
Newman DJ, Cragg GM (2012) Natural products as sources of new drugs over the 30 years from 1981 to 2010. J Nat Prod 75:311–335. https://doi.org/10.1021/np200906s
Novikova ND (2004) Review of the knowledge of microbial contamination of the Russian manned spacecraft. Microb Ecol 47:127–132. https://doi.org/10.1007/s00248-003-1055-2
Novikova N, De Boever P, Poddubko S, Deshevaya E, Polikarpov N, Rakova N, Coninx I, Mergeay M (2006) Survey of environmental biocontamination on board the International Space Station. Res Microbiol 157:5–12. https://doi.org/10.1016/j.resmic.2005.07.010
Oakley CE, Ahuja M, Sun W-W, Entwistle R, Akashi T, Yaegashi J, Guo C-J, Cerqueira GC, Russo Wortman J, Wang CCC, Chiang Y-M, Oakley BR (2017) Discovery of McrA, a master regulator of Aspergillus secondary metabolism. Mol Microbiol 103:347–365. https://doi.org/10.1111/mmi.13562
Onofri S, Selbmann L, de Hoog GS, Grube M, Barreca D, Ruisi S, Zucconi L (2007) Evolution and adaptation of fungi at boundaries of life. Adv Space Res 40:1657–1664. https://doi.org/10.1016/j.asr.2007.06.004
Onofri S, Barreca D, Selbmann L, Isola D, Rabbow E, Horneck G, de Vera JPP, Hatton J, Zucconi L (2008) Resistance of Antarctic black fungi and cryptoendolithic communities to simulated space and Martian conditions. Stud Mycol 61:99–109. https://doi.org/10.3114/sim.2008.61.10
Onofri S, Selbmann L, Pacelli C, de Vera JP, Horneck G, Hallsworth JE, Zucconi L (2018a) Integrity of the DNA and cellular ultrastructure of cryptoendolithic fungi in space or Mars conditions: a 1.5-year study at the International Space Station. Life Basel Switz 8:. https://doi.org/10.3390/life8020023
Onofri S, Selbmann L, Pacelli C, Zucconi L, Rabbow E, de Vera J-P (2018b) Survival, DNA, and ultrastructural integrity of a cryptoendolithic Antarctic fungus in Mars and lunar rock analogues exposed outside the International Space Station. Astrobiology. https://doi.org/10.1089/ast.2017.1728
Pierson DL (2001) Microbial contamination of spacecraft. Gravit Space Biol Bull 14:1–6
Plubell DL, Wilmarth PA, Zhao Y, Fenton AM, Minnier J, Reddy AP, Klimek J, Yang X, David LL, Pamir N (2017) Extended multiplexing of tandem mass tags (TMT) labeling reveals age and high fat diet specific proteome changes in mouse epididymal adipose tissue. Mol Cell Proteomics 16:873–890. https://doi.org/10.1074/mcp.M116.065524
Pusztahelyi T, Klement É, Szajli E, Klem J, Miskei M, Karányi Z, Emri T, Kovács S, Orosz G, Kovács KL, Medzihradszky KF, Prade RA, Pócsi I (2011) Comparison of transcriptional and translational changes caused by long-term menadione exposure in Aspergillus nidulans. Fungal Genet Biol 48:92–103. https://doi.org/10.1016/j.fgb.2010.08.006
Robinson MD, Oshlack A (2010) A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol 11:R25. https://doi.org/10.1186/gb-2010-11-3-r25
Sanchez JF, Entwistle R, Hung J-H, Yaegashi J, Jain S, Chiang Y-M, Wang CCC, Oakley BR (2011) Genome-based deletion analysis reveals the prenyl xanthone biosynthesis pathway in Aspergillus nidulans. J Am Chem Soc 133:4010–4017. https://doi.org/10.1021/ja1096682
Sanchez JF, Somoza AD, Keller NP, Wang CCC (2012) Advances in Aspergillus secondary metabolite research in the post-genomic era. Nat Prod Rep 29:351–371. https://doi.org/10.1039/c2np00084a
Segal BH (2009) Aspergillosis. N Engl J Med 360:1870–1884. https://doi.org/10.1056/NEJMra0808853
Sharpe RA, Bearman N, Thornton CR, Husk K, Osborne NJ (2015) Indoor fungal diversity and asthma: a meta-analysis and systematic review of risk factors. J Allergy Clin Immunol 135:110–122. https://doi.org/10.1016/j.jaci.2014.07.002
Singaravelan N, Grishkan I, Beharav A, Wakamatsu K, Ito S, Nevo E (2008) Adaptive melanin response of the soil fungus Aspergillus niger to UV radiation stress at “Evolution Canyon”, Mount Carmel, Israel. PLoS One 3:e2993. https://doi.org/10.1371/journal.pone.0002993
Stajich JE, Harris T, Brunk BP, Brestelli J, Fischer S, Harb OS, Kissinger JC, Li W, Nayak V, Pinney DF, Stoeckert CJ, Roos DS (2012) FungiDB: an integrated functional genomics database for fungi. Nucleic Acids Res 40:D675–D681. https://doi.org/10.1093/nar/gkr918
Szewczyk E, Nayak T, Oakley CE, Edgerton H, Xiong Y, Taheri-Talesh N, Osmani SA, Oakley BR, Oakley B (2006) Fusion PCR and gene targeting in Aspergillus nidulans. Nat Protoc 1:3111–3120. https://doi.org/10.1038/nprot.2006.405
Szewczyk E, Chiang Y-M, Oakley CE, Davidson AD, Wang CCC, Oakley BR (2008) Identification and characterization of the asperthecin gene cluster of Aspergillus nidulans. Appl Environ Microbiol 74:7607–7612. https://doi.org/10.1128/AEM.01743-08
Tixador R, Richoilley G, Gasset G, Templier J, Bes J, Moatti N, Lapchine L (1985) Study of minimal inhibitory concentration of antibiotics on bacteria cultivated in vitro in space (Cytos 2 experiment). Aviat Space Environ Med 56:748–751
Tkavc R, Matrosova VY, Grichenko OE, Gostinčar C, Volpe RP, Klimenkova P, Gaidamakova EK, Zhou CE, Stewart BJ, Lyman MG, Malfatti SA, Rubinfeld B, Courtot M, Singh J, Dalgard CL, Hamilton T, Frey KG, Gunde-Cimerman N, Dugan L, Daly MJ (2018) Prospects for fungal bioremediation of acidic radioactive waste sites: characterization and genome sequence of Rhodotorula taiwanensis MD1149. Front Microbiol 8. https://doi.org/10.3389/fmicb.2017.02528
Uhl MA, Biery M, Craig N, Johnson AD (2003) Haploinsufficiency-based large-scale forward genetic analysis of filamentous growth in the diploid human fungal pathogen C. albicans. EMBO J 22:2668–2678. https://doi.org/10.1093/emboj/cdg256
Van Houdt R, Mijnendonckx K, Leys N (2012) Microbial contamination monitoring and control during human space missions. Planet Space Sci 60:115–120. https://doi.org/10.1016/j.pss.2011.09.001
Venkateswaran K, Vaishampayan P, Cisneros J, Pierson DL, Rogers SO, Perry J (2014) International Space Station environmental microbiome—microbial inventories of ISS filter debris. Appl Microbiol Biotechnol 98:6453–6466. https://doi.org/10.1007/s00253-014-5650-6
Volz PA, Dublin M (1973) Filamentous fungi exposed to spaceflight stresses including known levels of ultraviolet irradiations. Space Life Sci 4:402–414. https://doi.org/10.1007/BF00930352
de Vries RP, Visser J (2001) Aspergillus enzymes involved in degradation of plant cell wall polysaccharides. Microbiol Mol Biol Rev 65:497–522. https://doi.org/10.1128/MMBR.65.4.497-522.2001
Wilson JW, Ott CM, Quick L, Davis R, zu Bentrup KH, Crabbé A, Richter E, Sarker S, Barrila J, Porwollik S, Cheng P, McClelland M, Tsaprailis G, Radabaugh T, Hunt A, Shah M, Nelman-Gonzalez M, Hing S, Parra M, Dumars P, Norwood K, Bober R, Devich J, Ruggles A, CdeBaca A, Narayan S, Benjamin J, Goulart C, Rupert M, Catella L, Schurr MJ, Buchanan K, Morici L, McCracken J, Porter MD, Pierson DL, Smith SM, Mergeay M, Leys N, Stefanyshyn-Piper HM, Gorie D, Nickerson CA (2008) Media ion composition controls regulatory and virulence response of Salmonella in spaceflight. PLoS One 3:e3923. https://doi.org/10.1371/journal.pone.0003923
Wu J, Wang M, Zhou L, Yu D (2016) Small heat shock proteins, phylogeny in filamentous fungi and expression analyses in Aspergillus nidulans. Gene 575:675–679. https://doi.org/10.1016/j.gene.2015.09.044
Yaegashi J, Oakley BR, Wang CCC (2014) Recent advances in genome mining of secondary metabolite biosynthetic gene clusters and the development of heterologous expression systems in Aspergillus nidulans. J Ind Microbiol Biotechnol 41:433–442. https://doi.org/10.1007/s10295-013-1386-z
Acknowledgements
Part of the research described in this publication was carried out at the Jet Propulsion Laboratory, California Institute of Technology, under a contract with NASA. We would like to thank astronauts Tim Peake, Tim Kopra, and Jeff Williams for handling the samples aboard the ISS, the Implementation Team at NASA Ames Research Center, and BioServe Space Technologies for coordinating this effort. © 2018 California Institute of Technology. Government sponsorship acknowledged.
Reference herein to any specific commercial product, process, or service by trade name, trademark, manufacturer, or otherwise does not constitute or imply its endorsement by the U.S. Government or the Jet Propulsion Laboratory, California Institute of Technology.
Funding
This research was funded by a 2012 Space Biology NNH12ZTT001N grant nos. 19-12829-26 under Task Order NNN13D111T awarded to CW and KV, which also funded JR and AB.
Author information
Authors and Affiliations
Contributions
JR drafted the manuscript, contributed to sample processing, and was responsible for data analysis and interpretation. AB contributed to sample processing and data interpretation. AC and MK conducted protein sample processing, LC/MS analyses, and proteome data processing. YC contributed to secondary metabolic analysis and interpretation. SM contributed to variant analysis. JY generated the CW12001 strain. SC was responsible for sample integration into flight hardware. FK was responsible for project implementation and generating metadata from the ISS. JS contributed to genome data processing and variant analysis. KV and CW designed the study, interpreted the data, and drafted the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 36056 kb)
Rights and permissions
About this article
Cite this article
Romsdahl, J., Blachowicz, A., Chiang, A.J. et al. International Space Station conditions alter genomics, proteomics, and metabolomics in Aspergillus nidulans. Appl Microbiol Biotechnol 103, 1363–1377 (2019). https://doi.org/10.1007/s00253-018-9525-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9525-0