Abstract
To all organisms, sulfur is an essential and important element. The assimilation of inorganic sulfur molecules such as sulfate and thiosulfate into organic sulfur compounds such as l-cysteine and l-methionine (essential amino acid for human) is largely contributed by microorganisms. Of these, special attention is given to thiosulfate (S2O32−) assimilation, because thiosulfate relative to often utilized sulfate (SO42−) as a sulfur source is proposed to be more advantageous in microbial growth and biotechnological applications like l-cysteine fermentative overproduction toward industrial manufacturing. In Escherichia coli as well as other many bacteria, the thiosulfate assimilation pathway is known to depend on O-acetyl-l-serine sulfhydrylase B. Recently, another yet-unidentified CysM-independent thiosulfate pathway was found in E. coli. This pathway is expected to consist of the initial part of the thiosulfate to sulfite (SO32−) conversion, and the latter part might be shared with the final part of the known sulfate assimilation pathway [sulfite → sulfide (S2−) → l-cysteine]. The catalysis of thiosulfate to sulfite is at least partly mediated by thiosulfate sulfurtransferase (GlpE). In this mini-review, we introduce updated comprehensive information about sulfur assimilation in microorganisms, including this topic. Also, we introduce recent advances of the application study about l-cysteine overproduction, including the GlpE overexpression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
To all organisms, sulfur is an essential element as well as carbon and nitrogen, although its elemental composition in living organisms is generally much low (Ingenbleek and Kimura 2013). As well-known and important sulfur-containing biomolecules, there are two proteinogenic amino acids, l-cysteine (Cys) and l-methionine (Met). For mammals including human, Met is considered to be an essential amino acid, because they cannot biosynthesize it de novo, but can do Cys from Met. Thus, mammals must obtain Met from diet intake, subsequently metabolizing it into Cys and a wide range of other functional sulfur compounds, such as l-glutathione (GSH) and coenzyme A. They finally catabolize such organic sulfur compounds into inorganic sulfur molecules. In contrast, most microorganisms and plants can basically biosynthesize Cys and Met from environmental inorganic sulfur sources such as sulfate (SO42−), thiosulfate (S2O32−), sulfite (SO32−), and sulfide (S2−). In this sense, biological sulfur cycle on the Earth is supported by the ability of the sulfur assimilation, namely, the Cys biosynthesis dwelling in microorganisms and plants. So, understanding of Cys biosynthetic metabolism in microorganisms is important study for us.
Considering functional difference between Cys and Met, it is mainly attributed to structural difference of its sulfur form—Cys harbors sulfur atom as its thiol group [HS-CH2CH(NH2)COOH] responsible for high redox reactivity, but Met as its thioether group [CH3-S-(CH2)2CH(NH2)COOH] with basically low redox reactivity. This is why Cys is involved in many redox reactions to exert critical biological functions as catalytic residue of various enzymes, as donor of sulfur atom required for assembly of Fe/S cluster, and as a component of GSH (5-l-glutamyl-l-cysteinylglycine). GSH play significant roles in various cellular functions and is required for glutaredoxin, glutathione peroxidase, dehydroascorbate reductase, glutathione S-transferase, etc. In addition, we recently reported a novel cellular antioxidative mechanism mediated by free-form Cys itself in E. coli, designated as “cysteine/cystine shuttle system” (Ohtsu et al. 2015; Ohtsu et al. 2010): Cys is exported from the cytoplasm to the periplasm, then detoxifies H2O2 by converting it to H2O as reducing equivalents, and alternatively generated oxidized product l-cystine (a Cys dimer via disulfide bond) is imported back to the cytoplasm to regenerate Cys for its recycling.
Such redox property of Cys makes it valuable and significant as a functional material in the applicational industries such as food (dough conditioner, flavor enhancer, and dietary supplement), pharmacy (expectorant agent and freckles preventive medicine), and cosmetic (perm assistant and whitening agent). So, its need is becoming higher, and global Cys market is growing (ca. 5000 tons per year) (Takagi and Ohtsu 2017). Currently, commercial Cys is mainly supplied by the following production processes: (1) extraction from keratin (human hair and animal feather) hydrolysis product, (2) enzymatic production of biochemical approach (Ryu et al. 1997; Tamura et al. 1998), and (3) fermentative production by genetically engineered microorganisms. Historically, almost all Cys has been manufactured by the extraction method. However, this extraction method has some unfavorable factors, i.e., generation of unpleasant smell, abundant utilization of environmentally unfriendly concentrated hydrochloric acid (Hunt 1985), and utilization of animal-derived source, which is unsuitable for people with certain religious beliefs and also due to health concern about bovine spongiform encephalopathy (BSE) (Wada and Takagi 2006). On the other hand, the fermentative production method does not basically accompany such disadvantages. Moreover, the recent technological progress based on understanding of sulfur metabolism is making it possible to dramatically improve the productivity, thereby realizing higher cost-effectiveness and resulting price competitiveness. In this way, the market share of the extraction method and also the enzymatic production method is being replaced by that of the microorganism fermentation method, although it is now still at the beginning on the way to go.
This mini-review highlights Cys mainly in terms of microorganisms, introduces its general biosynthetic metabolism and its recent progresses of scientific understanding, and provides recent advances of its fermentative production. Besides, based on this survey, we discuss about the perspective with respect to upcoming sulfur biology and required leading-edge analytical chemistry of sulfur metabolites as its driving force. Furthermore, we also discuss next-generation targets of unique and valuable sulfur compounds for commercial manufacturing by means of the establishing first-generation technic, i.e., Cys fermentative production.
l-Cysteine biosynthetic metabolism in microorganism
For sulfur assimilation to biosynthesize Cys in microorganisms, various forms of inorganic sulfur sources exist in natural environments. For example, elemental sulfur is one of them, and its assimilation is summarized elsewhere (Le Faou et al. 1990). Of such sulfur sources, we here focus on sulfate (SO42−) and thiosulfate (S2O32−), which are often utilized in the fermentative production by microorganisms.
Sulfate assimilation pathway
Sulfate is oxidized and stable state of inorganic sulfur molecule, ubiquitously exists in various environments such as soil and sea. Most microorganisms can utilize sulfate as a sulfur source. Also, for many scientists, sulfate is the most popular and familiar sulfur source in their cultivation of microorganisms, e.g., ammonium sulfate, sodium sulfate, and magnesium sulfate. For de novo Cys biosynthesis, firstly, sulfate is reduced to sulfide (S2−) in most microorganisms in common, and it can be then incorporated into two types of carbon skeleton molecules such as O-acyl-l-serine and O-acyl-l-homoserine to generate organic sulfur molecules, Cys and l-homocysteine (HmCys), respectively. Subsequently, interconversion of Cys to HmCys or HmCys to Cys can be promoted by transsulfuration pathway or reverse transsulfuration pathway, respectively. Met is biosynthesized from HmCys by its S-methylation. For de novo biosynthesis of Cys and Met, each microorganism divergently equips each combination of pathways above. For example, Escherichia coli equips Cys biosynthesis derived from O-acetyl-l-serine (OAS) and transsulfuration pathway, and Saccharomyces cerevisiae does HmCys biosynthesis derived from O-acetyl-l-homoserine (OAH) and reverse transsulfuration pathway. We here introduce these representative two types of Cys biosynthetic pathways from sulfate in these model microorganisms. Detailed description of the kinds of combinations and catalytic reactions (e.g., acyl-groups of substrates, its responsible genes for catalyzes, and its genetic distribution across biological kingdoms) is excellently reviewed elsewhere (Ferla and Patrick 2014; Ruckert 2016).
E. coli
E. coli, model prokaryotic microorganism, is one of the most utilized bacterial strain for genetic engineering field. E. coli has been well studied with regard to sulfate assimilation pathway (Fig. 1, sulfate pathway). At first, sulfate is taken up into cells from ambient environment via an ATP-dependent transporter complex CysUWA (Aguilar-Barajas et al. 2011). CysU and CysW are subunits to form the channel in the inner membrane, and CysA is the predicted ATPase subunit for driving sulfate translocation. Sbp is the periplasmic sulfate-binding protein to deliver sulfate to the transporter for high affinity uptake (Sirko et al. 1995). After the import, sulfate is assimilated into Cys by five-step sequential enzymatic reactions. (1) Sulfate is activated by CysDN complex (sulfate adenylyltransferase) utilizing ATP to generate adenosine 5′-phosphosulfate (APS) (Leyh et al. 1988). (2) APS is phosphorylated by CysC (adenylylsulfate kinase) utilizing another ATP to generate 3′-phosphoadenosine 5′-phosphosulfate (PAPS) (Satishchandran and Markham 1989; Satishchandran and Markham 2000). (3) PAPS is converted into sulfite (SO32−) and adenosine 3′,5′-bisphosphate (PAP) in the coupled catalytic reactions by CysH (3′-phospho-adenylylsulfate reductase), thioredoxin, and thioredoxin reductase using a NADPH (Berendt et al. 1995; Lillig et al. 1999). (4) Sulfite is reduced by CysIJ complex (sulfite reductase) to generate sulfide (S2−) utilizing 3 NADPH (Siegel and Davis 1974). Siroheme, which is a prosthetic group in CysI subunit of the sulfite reductase, is biosynthesized from uroporphyrinogen-III by CysG (siroheme synthase) (Spencer et al. 1993). (5) Sulfide and OAS are converted into Cys simultaneously releasing acetate by CysK (O-acetyl-l-serine sulfhydrylase A) (Boronat et al. 1984; Fimmel and Loughlin 1977; Kredich and Tomkins 1966). OAS is generated by CysE (l-serine O-acetyltransferase) from l-serine and acetyl-CoA (Denk and Bock 1987; Kredich and Tomkins 1966). Altogether, sulfate pathway needs two ATPs and four NADPHs as the expense of cellular energy to biosynthesize 1 Cys. Another inevitable sulfur-containing amino acid, Met, is biosynthesized from Cys and O-succinyl-l-homoserine (OSH) via l-cystathionine and HmCys by transsulfuration pathway [MetB (cystathionine γ-synthase), MetC (cystathionine β-lyase), and MetE (cobalamin-independent homocysteine transmethylase) or MetH (cobalamin-dependent methionine synthase)] (Fig. 1). Other than E. coli, many microorganisms such as Salmonella typhimurium (Kredich 1992) and Schizosaccharomyces pombe (fission yeast) (Fujita and Takegawa 2004) possess similar fashion of Cys biosynthesis.
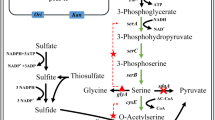
Schematic metabolic pathway of the inorganic sulfur assimilation/l-cysteine biosynthesis. For the comparison, that of E. coli is in the left and that of S. cerevisiae is in the right. Newly identified GlpE-dependent CysM-independent thiosulfate pathway in E. coli is represented as yellow arrows: thiosulfate conversion into sulfite coupled with the persulfuration of the thiol group of GlpE is emphasized. Enzymes catalyzing each step are underlined. Basically, sulfur metabolic flow from sulfate is represented as red arrows and that from thiosulfate is blue arrows, and carbon metabolic flow for l-cysteine biosynthesis is represented as green arrows
S. cerevisiae
S. cerevisiae (budding yeast) is a valuable organism utilized for fermentation industry such as alcoholic beverage, food flavorings, bread, and bioethanol. S. cerevisiae, a model eukaryotic microorganism, is also shown to assimilate sulfate (Marzluf 1997). In S. cerevisiae, sulfate is assimilated to sulfide (S2−) in the basically same steps as E. coli (Fig. 1). However, subsequent step to accept the sulfide is different from E. coli: HmCys is synthesized from the sulfide and OAH by l-homoserine O-acetyltransferase (Met17) (Cherest et al. 1969; Kerjan et al. 1986; Yamagata et al. 1974). Met is biosynthesized from HmCys by Met6 (Cobalamin-independent methionine synthase). Also, HmCys is utilized for Cys biosynthesis by reverse transsulfuration pathway via l-cystathionine as follows: (1) HmCys and l-serine are converted into l-cystathionine by Cys4 (l-cystathionine β-synthase) (Thomas and Surdin-Kerjan 1997), (2) the l-cystathionine is converted into Cys by Cys3 (l-cystathionine γ-lyase) (Cherest et al. 1993; Thomas and Surdin-Kerjan 1997). OAH to accept sulfide is generated by Met2 (l-homoserine O-acetyltransferase) from l-homoserine (HmSer) and acetyl-CoA (Thomas and Surdin-Kerjan 1997; Yamagata 1987).
Thiosulfate assimilation pathway
Thiosulfate is an inorganic sulfur molecule including sulfane sulfur (S=SO32−), which is more reduced state of sulfur atom than that of sulfate. It ubiquitously exists in the various environment such as soil and sea like sulfate, but relatively in anoxic conditions. For many scientists, thiosulfate might be unfamiliar sulfur source in their cultivation of microorganisms. Some microorganisms are known to be capable of utilizing thiosulfate as a sulfur source, although such reports are currently much less than that of sulfate assimilation. We here summarize conventional thiosulfate pathway, whose sulfur metabolic route is basically independent from sulfate pathway. Further, recently found new thiosulfate pathway in E. coli, which bypasses to sulfate assimilation pathway, is also shown.
Conventional thiosulfate pathway
E. coli
At first, thiosulfate is taken up from ambient environment via an ATP-dependent transporter complex CysUWA (Aguilar-Barajas et al. 2011). Namely, CysUWA is a common transporter for uptake of both sulfate and thiosulfate. However, unlike in the case of sulfate uptake, CysP, but not Sbp, is the periplasmic thiosulfate-binding protein to deliver thiosulfate to the transporter for high affinity uptake (Sirko et al. 1995). After the import, thiosulfate is assimilated into Cys by two-step sequential enzymatic reactions (Fig. 1). (1) Thiosulfate is incorporated into OAS to generate S-sulfo-l-cysteine (SSC) releasing acetate by CysM (O-acetyl-l-serine sulfhydrylase B) (Maier 2003). So, carbon skeleton molecule is the common as that in the sulfate pathway. (2) SSC is subsequently metabolized into Cys by the reductive cleavage reaction of its disulfide bond by NrdH (glutaredoxin-like protein) or Grx1 (glutaredoxin) (Nakatani et al. 2012). For continuity of these reductive reactions, NrdH itself must be reduced again after the reaction by thioredoxin reductase utilizing NADPH, and similarly Grx1 by reduced form of GSH, which is reduced again in parallel by glutathione reductase utilizing NADPH. Altogether, thiosulfate pathway needs at least only one NADPH as the expense of cellular energy to biosynthesize one Cys, although the simultaneously released sulfite (SO32−) is found to be also assimilated into another Cys via the sulfate pathway, which needs only three NADPHs (Nakatani et al. 2012).
Remarkably, the thiosulfate pathway is comparatively more efficient than sulfate pathway to biosynthesize Cys in terms of cellular bioenergetics. This is consistent with the physicochemical property that thiosulfate originally contains more reduced state of sulfur atom (sulfane sulfur) than that in sulfate (fully-oxidized state) (Toohey and Cooper 2014). Indeed, compatible with such principle, E. coli cells cultured with thiosulfate as the sole sulfur source can grow slightly faster than the cells cultured with sulfate as the sole sulfur source, especially in anaerobic conditions (personal communication). This phenomenon is probably due to the effect for saving the consumption of cellular fundamental energy metabolites (ATP and NADPH) in Cys biosynthesis by utilization of thiosulfate pathway. The saved energy equivalents can be alternatively allocated to other cellular processes limiting the growth.
S. cerevisiae
Like E. coli, S. cerevisiae is also known to assimilate thiosulfate as a sole sulfur source (Funahashi et al. 2015; Kankipati et al. 2015; Thomas et al. 1992). At first, thiosulfate is taken up from ambient environment via an transporters Sul1, Sul2, and Soa1 (Holt et al. 2017). After the import, thiosulfate is eventually assimilated into Cys and Met, but the metabolic route is still elusive. However, previous genetic study partly showed its outline that one sulfur atom of thiosulfate passes though sulfite-to-sulfide part and both two sulfur atoms of thiosulfate do sulfide-to-HmCys part of sulfate assimilation pathway (Thomas et al. 1992) (Fig. 1). In other words, for thiosulfate assimilation, it is converted into sulfite and sulfide and the both enter into sulfate assimilation pathway. Although thiosulfate can be generally cleaved into sulfite and sulfide by thiosulfate reductase, the corresponding gene functional for thiosulfate assimilation has been not found to date.
From the aspect of physiological significance, thiosulfate-grown S. cerevisiae cells exhibit slightly faster growth and ethanol production than the sulfate-grown cells (Funahashi et al. 2015). As is the case for E. coli, these behaviors of S. cerevisiae should also arise from thiosulfate-utilization-derived cellular energy-saving effect in Cys biosynthesis. In the thiosulfate-grown cells, intracellular NADPH content is shown to be indeed higher during logarithmic growth phase. This suggests that S. cerevisiae cells actually save cellular NADPH consumption more in assimilation of thiosulfate than that of sulfate. Also, the thiosulfate-grown cells consistently exhibited low carbon metabolic flux in the pentose phosphate pathway, whose important function is de novo production of cellular NADPH. In this way, advantage of thiosulfate compared to sulfate as sulfur source act as a common principle beyond biological species.
Recently identified new thiosulfate pathway
E. coli
As mentioned above as conventional thiosulfate pathway, thiosulfate assimilation in E. coli have been generally believed to be carried out by CysM-dependent thiosulfate pathway (CysM TSAP). However, in our recent study, hitherto-unknown thiosulfate assimilation pathway, namely CysM-independent thiosulfate pathway (non-CysM TSAP) was discovered (Fig. 1) (Kawano et al. 2017). The beginning of the study is the fact that even ΔcysM cells can very slowly and slightly but surely grow in the conditions of thiosulfate as a sole sulfur source. This means existence of non-CysM TSAP in E. coli. The sulfur metabolic flow of non-CysM TSAP toward Cys is suggested to be quite slow compared to that of CysM-dependent TSAP. In fact, thiosulfate-grown ΔcysM relative to WT cells can accumulate equivalent level of Cys, but cannot accumulate GSH at all, which located at downstream of Cys. The non-CysM TSAP is indicated to consist of the initial part of the thiosulfate-to-sulfite conversion (bypass from thiosulfate to sulfite) and share the latter part with the sulfite-to-sulfide-to-Cys conversion of the sulfate pathway. This is because thiosulfate-grown ΔcysM cells can (i) accumulate sulfite and sulfide equivalently to WT cells and (ii) completely lose growth capability by additional disruption of cysK gene, whose product catalyzes the final step (sulfide-to-Cys) in the sulfate pathway. The bypass from thiosulfate to sulfite in non-CysM TSAP is at least in part mediated by GlpE [cytoplasmic thiosulfate sulfurtransferase harboring a single “rhodanese-like domain” (Ray et al. 2000)]. GlpE is known to catalyze thiosulfate disproportionation, making the thiol group (-SH) of the catalytic cysteine residue persulfurated (-SSH) by the sulfane sulfur of thiosulfate (S=SO32−), simultaneously releasing sulfite (SO32−) (Cheng et al. 2008). The reason for the GlpE function in thiosulfate assimilation is based on the results that its overexpression can (i) enhance in vitro cellular thiosulfate sulfurtransferase activity, (ii) restore the GSH pool abolished in thiosulfate-grown ΔcysM cells up to the WT level in vivo, and (iii) completely complement the crucial phenotype of restricted and slow growth in thiosulfate-grown ΔcysM cells. (iv) Additionally, in its gene disruption, thiosulfate-grown ΔcysMΔglpE cells consistently exhibited more defective growth than ΔcysM cells, suggesting that GlpE intrinsically plays the role in non-CysM TSAP. In summary, GlpE catalyzes thiosulfate to sulfite in a novel non-CysM TSAP, which forms a bypass of sulfur metabolic flow from thiosulfate to a sulfate pathway to biosynthesize Cys. To the best of our knowledge, this is the first comprehensive report at the genetic and molecular level demonstrating that a thiosulfate sulfurtransferase can function in thiosulfate assimilation.
S. cerevisiae
Based on current knowledge, in S. cerevisiae, thiosulfate assimilation pathway is believed to pass the sulfite and sulfide in sulfate assimilation pathway. However, the responsible gene(s) for the conversion is unidentified to date. Considering newly identified GlpE-dependent non-CysM TSAP in E. coli above, thiosulfate sulfurtransferase family gene with “rhodanese-like domain” might be a part of the missing link. In S. cerevisiae, there are five such genes (TUM1, RDL1, RDL2, UBA4, and YCH1). These genes are known to individually have individual functions respectively, e.g., sulfur relay regarding 2-thiouridine biogenesis at tRNA wobble positions (Noma et al. 2009) or hydrogen sulfide metabolism (Melideo et al. 2014). For example, Rdl1 is known to actually catalyze the reaction making its thiol group (-SH) of the catalytic cysteine residue persulfurated (-SSH) by the sulfane sulfur of thiosulfate, simultaneously releasing sulfite (Melideo et al. 2014). Of these genes, some might have the function responsible for thiosulfate assimilation.
Recent progress in l-cysteine fermentative production
As mentioned, GlpE overexpression was found to be effective for Cys biosynthesis via non-CysM TSAP. This led us to apply this mechanism in fermentative Cys overproduction. In this regard, we have already reported marked production of extracellular Cys from glucose in E. coli by genetic engineering involving sulfur metabolism, using thiosulfate as the sulfur source (Kawano et al. 2015a; Nakatani et al. 2012; Nonaka et al. 2012; Wiriyathanawudhiwong et al. 2009). As the result, GlpE overexpression is found to be effective for enhancement of Cys production from thiosulfate regardless of the presence or absence of CysM function (Kawano et al. 2017), and achieved 1.5 g/L Cys production. For other thiosulfate sulfurtransferases, it is known that PspE in the periplasm is highly active in one E. coli: most cellular thiosulfate sulfurtransferase activity is derived from PspE and GlpE (Cheng et al. 2008). Thus, like GlpE, PspE might be involved in non-CysM TSAP, and thus it might be another candidate for the genetic engineering.
Other than our study above, Cys overproduction technic is currently progressing by recent studies. For example, Liu et al. achieved 5.1 g/L Cys production in E. coli, applying combined rational metabolic engineering and modular strategy (enhancing biosynthesis and weakening degradation) (Liu et al. 2018). Takumi et al. achieved 2.2 g/L Cys production in Pantoea ananatis by applying genetic modification of metabolic engineering approach (enhancing/optimizing biosynthesis, weakening degradation, and enhancing efflux) (Takumi et al. 2017). Joo et al. achieved 60 mg/L Cys production in Corynebacterium glutamicum by combinatorial overexpression of genes involved in Cys production (enhancing biosynthesis) (Joo et al. 2017). At present, a few companies have recently and already established industrial fermentative Cys overproduction using thiosulfate, and such Cys is being marketed. However, the production cost remains higher than that of other amino acids. Thus, further improvement of the productive efficiency is still required from now on.
Perspective and conclusion
Understanding of sulfur metabolism in microorganism and also other biological species is recently progressing rapidly. One major contributing reason is development and improvement of quantitative analysis method to monitor the dynamics of sulfur metabolites, whose biological contents are quite low and thus difficult to quantify, relative to abundantly existing carbon or nitrogen metabolites. As current mainstream for sulfur metabolomic analysis, the method combining (i) chemical derivatization of target thiol molecules by monobromobimane (mBBr) and (ii) its detection and quantitation by the LC-MS/MS system (Kawano et al. 2015b) is prevailing because of preceding cutting-edge studies (Ida et al. 2014; Nakano et al. 2015; Ohmura et al. 2015). In this method, critical point is that the chemical derivatization of thiols enables the method to avoid chemical redox and autoxidation reactions of thiol molecules in the sample during metabolite extraction and analytic sample preparation generally performed in open air existing oxygen. Also, such analysis has been already available commercially (http://www.euglena.jp/sulfurindex/).
Such methodological advance for sulfur metabolites allowed detailed analysis of sulfane sulfur: sulfane sulfur is basically in the form of persulfide (R-SS-H) and polysulfide (R-S-Sn-H) among many thiol-containing proteins, small organic thiol compounds (GSH, Cys, HmCys, etc.), and inorganic sulfur such as thiosulfate. Recently, such polysulfides are shown to be ubiquitously exists in a wide variety of organisms in multiple forms (Ida et al. 2014). The fashion of dynamics is considered to be complex and is unique to sulfur. The polysulfide can function as new redox player, can also be ready-to-use-form of stock of sulfur and reducing equivalent, and can function as a new form of signal depending on its polysulfide state, leading to the establishment of a new biological concept, i.e., polysulfidomics (Ida et al. 2014). Elucidation of this concept is challenging but will bring about new, real, and comprehensive cellular biology. In this context, as for GlpE-associated thiosulfate assimilation, it is still questions regarding (i) the mechanism and relevance of GlpE persulfide (or polysulfide) and (ii) the fate of sulfur. This theme can provide valuable clues to understand the largely obscure subject of polysulfidomics.
Nowadays, technic of Cys fermentative production has reached the level of practical application, although further improvement is strongly needed for cost-effectiveness. So, in the near future, it is becoming significant to study and develop the sophisticated technology for fermentative production of other valuable and functional sulfur compounds on the fundamental technic for Cys fermentative production. The accomplishment should be carried out by genetic engineering and synthetic biology approach applying diverse and extensive genetic resource in microorganisms. As one practical such compound, much attention is now paid to ergothioneine (ERG). ERG is naturally occurring sulfur-containing amino acid derivative with unique structure, a histidine betaine derivative with a thione group attached to the C-2 atom of the imidazole ring. ERG is functional antioxidant and biosynthesized by actinobacteria, cyanobacteria, and a fission yeast (Genghof 1970; Pfeiffer et al. 2011; Pluskal et al. 2014), etc. Humans obtain ERG from intake of diet, such as mushrooms and red beans, because they cannot biosynthesize it. ERG is valuable for humans, because it functions as unique antioxidant especially effective for scavenging hydroxyl radical (Franzoni et al. 2006), highly toxic reactive oxygen species giving crucial damage to cellular system. On the contrary, ERG hold only a disadvantage with regard to its availability as material source, namely, its too high cost in the current market. Thus, we are now trying to establish the efficient fermentative production by application of genetic engineering and synthetic biology approach (Osawa et al. 2018) and have developed the production to the level of gram-scale at present (personal communication). Toward enjoyment of the merit of ERG for human health care, it is another rational and potential approach to develop ERG content in already-marketed health food derived from microorganisms such as Euglena (Suzuki 2017), Chlorella, etc., through exhaustive survey of ERG presence, its amount, and its potential changeability in the culture conditions (e.g., a kind of sulfur source). Finally, we propose again that utilization of thiosulfate as sulfur source rather than sulfate is promising for the improvement of biological production of various target molecules, even non-sulfur compounds. Authors hope that this mini-review can provide some help and encouragement for many readers involving applied microbiology and biotechnology.
References
Aguilar-Barajas E, Diaz-Perez C, Ramirez-Diaz MI, Riveros-Rosas H, Cervantes C (2011) Bacterial transport of sulfate, molybdate, and related oxyanions. Biometals 24(4):687–707. https://doi.org/10.1007/s10534-011-9421-x
Berendt U, Haverkamp T, Prior A, Schwenn JD (1995) Reaction mechanism of thioredoxin: 3′-phospho-adenylylsulfate reductase investigated by site-directed mutagenesis. Eur J Biochem 233(1):347–356. https://doi.org/10.1111/j.1432-1033.1995.347_1.x
Boronat A, Britton P, Jones-Mortimer MC, Kornberg HL, Lee LG, Murfitt D, Parra F (1984) Location on the Escherichia coli genome of a gene specifying O-acetylserine (thiol)-lyase. J Gen Microbiol 130(3):673–685. https://doi.org/10.1099/00221287-130-3-673
Cheng H, Donahue JL, Battle SE, Ray WK, Larson TJ (2008) Biochemical and genetic characterization of PspE and GlpE, two single-domain Sulfurtransferases of Escherichia coli. Open Microbiol J 2:18–28. https://doi.org/10.2174/1874285800802010018
Cherest H, Eichler F, Robichon-Szulmajster H (1969) Genetic and regulatory aspects of methionine biosynthesis in Saccharomyces cerevisiae. J Bacteriol 97(1):328–336
Cherest H, Thomas D, Surdin-Kerjan Y (1993) Cysteine biosynthesis in Saccharomyces cerevisiae occurs through the transsulfuration pathway which has been built up by enzyme recruitment. J Bacteriol 175(17):5366–5374. https://doi.org/10.1128/jb.175.17.5366-5374.1993
Denk D, Bock A (1987) L-cysteine biosynthesis in Escherichia coli: nucleotide sequence and expression of the serine acetyltransferase (cysE) gene from the wild-type and a cysteine-excreting mutant. J Gen Microbiol 133(3):515–525. https://doi.org/10.1099/00221287-133-3-515
Ferla MP, Patrick WM (2014) Bacterial methionine biosynthesis. Microbiology 160(Pt 8):1571–1584. https://doi.org/10.1099/mic.0.077826-0
Fimmel AL, Loughlin RE (1977) Isolation and characterization of cysK mutants of Escherichia coli K12. J Gen Microbiol 103(1):37–43. https://doi.org/10.1099/00221287-103-1-37
Franzoni F, Colognato R, Galetta F, Laurenza I, Barsotti M, Di Stefano R, Bocchetti R, Regoli F, Carpi A, Balbarini A, Migliore L, Santoro G (2006) An in vitro study on the free radical scavenging capacity of ergothioneine: comparison with reduced glutathione, uric acid and trolox. Biomed Pharmacother 60(8):453–457. https://doi.org/10.1016/j.biopha.2006.07.015
Fujita Y, Takegawa K (2004) Characterization of two genes encoding putative cysteine synthase required for cysteine biosynthesis in Schizosaccharomyces pombe. Biosci Biotechnol Biochem 68(2):306–311. https://doi.org/10.1271/bbb.68.306
Funahashi E, Saiki K, Honda K, Sugiura Y, Kawano Y, Ohtsu I, Watanabe D, Wakabayashi Y, Abe T, Nakanishi T, Suematsu M, Takagi H (2015) Finding of thiosulfate pathway for synthesis of organic sulfur compounds in Saccharomyces cerevisiae and improvement of ethanol production. J Biosci Bioeng 120:666–669. https://doi.org/10.1016/j.jbiosc.2015.04.011
Genghof DS (1970) Biosynthesis of ergothioneine and hercynine by fungi and Actinomycetales. J Bacteriol 103(2):475–478
Holt S, Kankipati H, De Graeve S, Van Zeebroeck G, Foulquie-Moreno MR, Lindgreen S, Thevelein JM (2017) Major sulfonate transporter Soa1 in Saccharomyces cerevisiae and considerable substrate diversity in its fungal family. Nat Commun 8:14247. https://doi.org/10.1038/ncomms14247
Hunt S (1985) Degradation of amino acids accompanying in vitro protein hydrolysis. In: Barrett GC (ed) Chemistry and biochemistry of the amino acids. Springer Netherlands, Dordrecht, pp 376–398
Ida T, Sawa T, Ihara H, Tsuchiya Y, Watanabe Y, Kumagai Y, Suematsu M, Motohashi H, Fujii S, Matsunaga T, Yamamoto M, Ono K, Devarie-Baez NO, Xian M, Fukuto JM, Akaike T (2014) Reactive cysteine persulfides and S-polythiolation regulate oxidative stress and redox signaling. Proc Natl Acad Sci U S A 111(21):7606–7611. https://doi.org/10.1073/pnas.1321232111
Ingenbleek Y, Kimura H (2013) Nutritional essentiality of sulfur in health and disease. Nutr Rev 71(7):413–432. https://doi.org/10.1111/nure.12050
Joo YC, Hyeon JE, Han SO (2017) Metabolic design of Corynebacterium glutamicum for production of l-cysteine with consideration of sulfur-supplemented animal feed. J Agric Food Chem 65(23):4698–4707. https://doi.org/10.1021/acs.jafc.7b01061
Kankipati HN, Rubio-Texeira M, Castermans D, Diallinas G, Thevelein JM (2015) Sul1 and Sul2 sulfate transceptors signal to protein kinase a upon exit of sulfur starvation. J Biol Chem 290(16):10430–10446. https://doi.org/10.1074/jbc.M114.629022
Kawano Y, Ohtsu I, Takumi K, Tamakoshi A, Nonaka G, Funahashi E, Ihara M, Takagi H (2015a) Enhancement of l-cysteine production by disruption of yciW in Escherichia coli. J Biosci Bioeng 119(2):176–179. https://doi.org/10.1016/j.jbiosc.2014.07.006
Kawano Y, Ohtsu I, Tamakoshi A, Shiroyama M, Tsuruoka A, Saiki K, Takumi K, Nonaka G, Nakanishi T, Hishiki T, Suematsu M, Takagi H (2015b) Involvement of the yciW gene in l-cysteine and l-methionine metabolism in Escherichia coli. J Biosci Bioeng 119(3):310–313. https://doi.org/10.1016/j.jbiosc.2014.08.012
Kawano Y, Onishi F, Shiroyama M, Miura M, Tanaka N, Oshiro S, Nonaka G, Nakanishi T, Ohtsu I (2017) Improved fermentative L-cysteine overproduction by enhancing a newly identified thiosulfate assimilation pathway in Escherichia coli. Appl Microbiol Biotechnol 101(18):6879–6889. https://doi.org/10.1007/s00253-017-8420-4
Kerjan P, Cherest H, Surdin-Kerjan Y (1986) Nucleotide sequence of the Saccharomyces cerevisiae MET25 gene. Nucleic Acids Res 14(20):7861–7871. https://doi.org/10.1093/nar/14.20.7861
Kredich NM (1992) The molecular basis for positive regulation of cys promoters in Salmonella typhimurium and Escherichia coli. Mol Microbiol 6(19):2747–2753. https://doi.org/10.1111/j.1365-2958.1992.tb01453.x
Kredich NM, Tomkins GM (1966) The enzymic synthesis of L-cysteine in Escherichia coli and Salmonella typhimurium. J Biol Chem 241(21):4955–4965
Le Faou A, Rajagopal BS, Daniels L, Fauque G (1990) Thiosulfate, polythionates and elemental sulfur assimilation and reduction in the bacterial world. FEMS Microbiol Rev 6(4):351–381. https://doi.org/10.1016/0378-1097(90)90688-M
Leyh TS, Taylor JC, Markham GD (1988) The sulfate activation locus of Escherichia coli K12: cloning, genetic, and enzymatic characterization. J Biol Chem 263(5):2409–2416
Lillig CH, Prior A, Schwenn JD, Aslund F, Ritz D, Vlamis-Gardikas A, Holmgren A (1999) New thioredoxins and glutaredoxins as electron donors of 3′-phosphoadenylylsulfate reductase. J Biol Chem 274(12):7695–7698. https://doi.org/10.1074/jbc.274.12.7695
Liu H, Fang G, Wu H, Li Z, Ye Q (2018) L-Cysteine production in Escherichia coli based on rational metabolic engineering and modular strategy. Biotechnol J. https://doi.org/10.1002/biot.201700695
Maier TH (2003) Semisynthetic production of unnatural L-alpha-amino acids by metabolic engineering of the cysteine-biosynthetic pathway. Nat Biotechnol 21(4):422–427. https://doi.org/10.1038/nbt807
Marzluf GA (1997) Molecular genetics of sulfur assimilation in filamentous fungi and yeast. Annu Rev Microbiol 51:73–96. https://doi.org/10.1146/annurev.micro.51.1.73
Melideo SL, Jackson MR, Jorns MS (2014) Biosynthesis of a central intermediate in hydrogen sulfide metabolism by a novel human sulfurtransferase and its yeast ortholog. Biochemistry 53(28):4739–4753. https://doi.org/10.1021/bi500650h
Nakano S, Ishii I, Shinmura K, Tamaki K, Hishiki T, Akahoshi N, Ida T, Nakanishi T, Kamata S, Kumagai Y, Akaike T, Fukuda K, Sano M, Suematsu M (2015) Hyperhomocysteinemia abrogates fasting-induced cardioprotection against ischemia/reperfusion by limiting bioavailability of hydrogen sulfide anions. J Mol Med (Berl) 93:879–889. https://doi.org/10.1007/s00109-015-1271-5
Nakatani T, Ohtsu I, Nonaka G, Wiriyathanawudhiwong N, Morigasaki S, Takagi H (2012) Enhancement of thioredoxin/glutaredoxin-mediated L-cysteine synthesis from S-sulfocysteine increases L-cysteine production in Escherichia coli. Microb Cell Factories 11:62. https://doi.org/10.1186/1475-2859-11-62
Noma A, Sakaguchi Y, Suzuki T (2009) Mechanistic characterization of the sulfur-relay system for eukaryotic 2-thiouridine biogenesis at tRNA wobble positions. Nucleic Acids Res 37(4):1335–1352. https://doi.org/10.1093/nar/gkn1023
Nonaka G, Yamazaki S, Takumi K (2012) Method for producing l-cysteine. WO patent WO 2012/137689 A1
Ohmura M, Hishiki T, Yamamoto T, Nakanishi T, Kubo A, Tsuchihashi K, Tamada M, Toue S, Kabe Y, Saya H, Suematsu M (2015) Impacts of CD44 knockdown in cancer cells on tumor and host metabolic systems revealed by quantitative imaging mass spectrometry. Nitric Oxide 46:102–113. https://doi.org/10.1016/j.niox.2014.11.005
Ohtsu I, Kawano Y, Suzuki M, Morigasaki S, Saiki K, Yamazaki S, Nonaka G, Takagi H (2015) Uptake of L-cystine via an ABC transporter contributes defense of oxidative stress in the L-cystine export-dependent manner in Escherichia coli. PLoS One 10(3):e0120619. https://doi.org/10.1371/journal.pone.0120619
Ohtsu I, Wiriyathanawudhiwong N, Morigasaki S, Nakatani T, Kadokura H, Takagi H (2010) The L-cysteine/L-cystine shuttle system provides reducing equivalents to the periplasm in Escherichia coli. J Biol Chem 285(23):17479–17487. https://doi.org/10.1074/jbc.M109.081356
Osawa R, Kamide T, Satoh Y, Kawano Y, Ohtsu I, Dairi T (2018) Heterologous and high production of ergothioneine in Escherichia coli. J Agric Food Chem 66(5):1191–1196. https://doi.org/10.1021/acs.jafc.7b04924
Pfeiffer C, Bauer T, Surek B, Schomig E, Grundemann D (2011) Cyanobacteria produce high levels of ergothioneine. Food Chem 129(4):1766–1769. https://doi.org/10.1016/j.foodchem.2011.06.047
Pluskal T, Ueno M, Yanagida M (2014) Genetic and metabolomic dissection of the ergothioneine and selenoneine biosynthetic pathway in the fission yeast, S. pombe, and construction of an overproduction system. PLoS One 9(5):e97774. https://doi.org/10.1371/journal.pone.0097774
Ray WK, Zeng G, Potters MB, Mansuri AM, Larson TJ (2000) Characterization of a 12-kilodalton rhodanese encoded by glpE of Escherichia coli and its interaction with thioredoxin. J Bacteriol 182(8):2277–2284. https://doi.org/10.1128/JB.182.8.2277-2284.2000
Ruckert C (2016) Sulfate reduction in microorganisms-recent advances and biotechnological applications. Curr Opin Microbiol 33:140–146. https://doi.org/10.1016/j.mib.2016.07.007
Ryu OH, Ju JY, Shin CS (1997) Continuous L-cysteine production using immobilized cell reactors and product extractors. Process Biochem 32(3):201–209. https://doi.org/10.1016/S0032-9592(96)00061-1
Satishchandran C, Markham GD (1989) Adenosine-5′-phosphosulfate kinase from Escherichia coli K12. Purification, characterization, and identification of a phosphorylated enzyme intermediate. J Biol Chem 264(25):15012–15021
Satishchandran C, Markham GD (2000) Mechanistic studies of Escherichia coli adenosine-5′-phosphosulfate kinase. Arch Biochem Biophys 378(2):210–215. https://doi.org/10.1006/abbi.2000.1841
Siegel LM, Davis PS (1974) Reduced nicotinamide adenine dinucleotide phosphate-sulfite reductase of enterobacteria. IV The Escherichia coli hemoflavoprotein: subunit structure and dissociation into hemoprotein and flavoprotein components. J Biol Chem 249(5):1587–1598
Sirko A, Zatyka M, Sadowy E, Hulanicka D (1995) Sulfate and thiosulfate transport in Escherichia coli K-12: evidence for a functional overlapping of sulfate- and thiosulfate-binding proteins. J Bacteriol 177(14):4134–4136. https://doi.org/10.1128/jb.177.14.4134-4136.1995
Spencer JB, Stolowich NJ, Roessner CA, Scott AI (1993) The Escherichia coli cysG gene encodes the multifunctional protein, siroheme synthase. FEBS Lett 335(1):57–60. https://doi.org/10.1016/0014-5793(93)80438-Z
Suzuki K (2017) Large-scale cultivation of Euglena. Adv Exp Med Biol 979:285–293. https://doi.org/10.1007/978-3-319-54910-1_14
Takagi H, Ohtsu I (2017) L-cysteine metabolism and fermentation in microorganisms. Adv Biochem Eng Biotechnol 159:129–151. https://doi.org/10.1007/10_2016_29
Takumi K, Ziyatdinov MK, Samsonov V, Nonaka G (2017) Fermentative production of cysteine by Pantoea ananatis. Appl Environ Microbiol 83(5):e02502–e02516. https://doi.org/10.1128/AEM.02502-16
Tamura Y, Nishino M, Ohmachi T, Asada Y (1998) N-Carbamoyl-L-Cysteine as an Intermediate in the bioconversion from D,L-2-Amino-Delta (2)-Thiazoline-4-Carboxylic Acid to L-Cysteine by Pseudomonas sp. ON-4a. Biosci Biotechnol Biochem 62(11):2226–2229. https://doi.org/10.1271/bbb.62.2226
Thomas D, Barbey R, Henry D, Surdin-Kerjan Y (1992) Physiological analysis of mutants of Saccharomyces cerevisiae impaired in sulphate assimilation. J Gen Microbiol 138(10):2021–2028. https://doi.org/10.1099/00221287-138-10-2021
Thomas D, Surdin-Kerjan Y (1997) Metabolism of sulfur amino acids in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 61(4):503–532
Toohey JI, Cooper AJ (2014) Thiosulfoxide (sulfane) sulfur: new chemistry and new regulatory roles in biology. Molecules 19(8):12789–12813. https://doi.org/10.3390/molecules190812789
Wada M, Takagi H (2006) Metabolic pathways and biotechnological production of L-cysteine. Appl Microbiol Biotechnol 73(1):48–54. https://doi.org/10.1007/s00253-006-0587-z
Wiriyathanawudhiwong N, Ohtsu I, Li ZD, Mori H, Takagi H (2009) The outer membrane TolC is involved in cysteine tolerance and overproduction in Escherichia coli. Appl Microbiol Biotechnol 81(5):903–913. https://doi.org/10.1007/s00253-008-1686-9
Yamagata S (1987) Partial purification and some properties of homoserine O-acetyltransferase of a methionine auxotroph of Saccharomyces cerevisiae. J Bacteriol 169(8):3458–3463. https://doi.org/10.1128/jb.169.8.3458-3463.1987
Yamagata S, Takeshima K, Naiki N (1974) Evidence for the identity of O-acetylserine sulfhydrylase with O-acetylhomoserine sulfhydrylase in yeast. J Biochem 75(6):1221–1229. https://doi.org/10.1093/oxfordjournals.jbchem.a130505
Acknowledgements
We would like to thank Taka-Aki Sato (Ph.D. Program in Human Biology, School of Integrative and Global Majors, University of Tsukuba, Tsukuba, Japan; Shimadzu Co., Kyoto, Japan) for excellent discussion.
Funding
This review was supported in part by JSPS KAKENHI Grant Numbers JP26450091 and JP15KT0028, by Science and Technology Research Promotion Program for Agriculture, Forestry, Fisheries and Food Industry (26027AB) from MAFF, Japan, and by the grant from The SKYLARK Food Science Institute, Japan, to I.O. This work was also supported in part by JSPS KAKENHI Grant Numbers JP16K18675 and JP15KT0028 to Y.K. The funders had no role in manuscript design or the decision to submit the work for publication.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Kawano, Y., Suzuki, K. & Ohtsu, I. Current understanding of sulfur assimilation metabolism to biosynthesize l-cysteine and recent progress of its fermentative overproduction in microorganisms. Appl Microbiol Biotechnol 102, 8203–8211 (2018). https://doi.org/10.1007/s00253-018-9246-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9246-4