Abstract
Listeria monocytogenes is a pathogen of significant concern in many ready to eat foods due to its ability to survive and multiply even under significant environmental stresses. Listeriosis in humans is a concern, especially to high-risk populations such as those who are immunocompromised or pregnant, due to the high rates of morbidity and mortality. Whole genome sequencing has become a routine part of assessing L. monocytogenes isolated from patients, and the frequency of different genetic subtypes associated with listeriosis is now being reported. The recent abundance of genome sequences for L. monocytogenes has provided a wealth of information regarding the variation in core and accessory genomic elements. Newly described accessory genomic regions have been linked to greater virulence capabilities as well as greater resistance to environmental stressors such as sanitizers commonly used in food processing facilities. This review will provide a summary of our current understanding of stress response and virulence phenotypes of L. monocytogenes, within the context of the genetic diversity of the pathogen.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Listeria monocytogenes is a Gram positive, non-spore-forming bacterium that can be isolated from a number of environments, including soil, water, and vegetation. It is considered to be an environmental saprotroph (Swaminathan et al. 2007). This bacterium is also an opportunistic pathogen of humans and animals that is acquired by consumption of contaminated food. Compared to other foodborne pathogens, L. monocytogenes can thrive in numerous environments outside the host, such as soil and water, which are entry points for contamination of the food supply (Freitag et al. 2009). The robust stress adaptive capabilities of L. monocytogenes contribute to its ability to survive and grow in diverse, non-host environments. For example, L. monocytogenes can grow with salt concentrations as high as 1.9 M NaCl (Ribeiro et al. 2014), at pH 4.5 (Vermeulen et al. 2007), and is a psychrotroph, able to grow at temperatures as low as 2 °C (Swaminathan et al. 2007).
The population structure of L. monocytogenes consists of four lineages, each of which have multiple clonal complexes (Haase et al. 2014). Analyses of multi-locus sequence data indicate that L. monocytogenes has a low rate of homologous recombination (Ragon et al. 2008), though the rates of recombination differ between lineages 1 and 2, with recombination more prevalent in lineage 2 (den Bakker et al. 2008). Comparative genomics analyses identified that ~ 43% (2360) of the L. monocytogenes genome is composed of core genes, with the remaining ~ 57% (3109) composed of accessory genes (Tan et al. 2015). An assessment of the accessory genome of L. monocytogenes found a number of lineage-specific accessory genes (den Bakker et al. 2013). Variation in stress resistance (Adriao et al. 2008; Bergholz et al. 2010; Lianou et al. 2006) as well as virulence (Gray et al. 2004; Rakic Martinez et al. 2017) phenotypes has been observed among genotypes of L. monocytogenes, though direct associations between specific genetic elements and phenotype are not always clear. While in the USA and many other countries, L. monocytogenes regulation is based on finding any L. monocytogenes in foods and/or the food processing environment, it has become increasingly clear that all L. monocytogenes strains are not equally represented among clinical and food isolates, whether these isolates are from outbreaks or sporadic cases (Norton and Braden 2007; Ward et al. 2008). There is an overwhelming preponderance of serotypes 4b, 1/2a, and 1/2b among clinical and food isolates, clearly pointing to differences in ability to survive in foods and/or cause disease.
Recent studies capitalizing on the ease and rapidity of whole genome sequencing (WGS) have uncovered to a greater extent the genetic variation among clonal complexes of L. monocytogenes (Moura et al. 2016). The scope of this mini-review includes the changing landscapes of different clonal complexes around the globe, and new insights arising from this extensive WGS data combined with phenotypic assessments, establishing connections with variation in genetic information and phenotypes that influence survival of L. monocytogenes in extra-host and in host environments.
Newly emerging clones of L. monocytogenes
L. monocytogenes phylogenetic studies have made use of various typing methods, including multi-locus sequence typing (MLST) and multi-locus virulence genes sequence typing (MLVST) to evaluate the relationships between strains and to identify lineages and clonal groups that correlate with infection and/or source. Both these schemes use about 3000 nucleotides from six different genes for phylogenetic analyses. In 2008, Ragon et al. used MLST sequence types (ST) to group strains by clonal complex (CC), which are comprised of ST groups that had a recent common ancestor (Ragon et al. 2008). Subsequent studies have led to the identification of additional CCs, though for some lineages, such as lineages 1 and 2, few new CCs were found, suggesting that most of the clinically relevant CCs have been identified (Haase et al. 2014). Many of these CCs have been linked to epidemic clones (Cantinelli et al. 2013), responsible for multiple outbreaks of listeriosis, suggesting these strains may represent more fit pathogens, though whether that is due to virulence attributes, higher incidence in food processing environments or increased stress resistance properties is unclear. As such, determination of dominant CCs over time may provide clues about fitness as well as indicate alterations in selective pressures that may lead to changes in the CCs that are predominant.
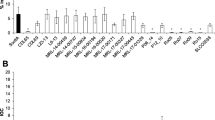
Several studies have focused on the distribution of CCs and STs during different time periods and in different geographic regions. An evaluation of these studies individually, as well as holistically, provides clues to changes in the environment L. monocytogenes resides within. Data collected from ten studies, with redundant strains removed when identified, were evaluated and found that certain groups represent potentially emerging CCs (Fig. 1). It is apparent that historically dominant CCs, such as CC1 and CC2 are represented less frequently in recent years. Instead, previously uncommon CCs are rising in frequency, particularly CC5, CC6, CC9, and CC121.
Relative frequency of clonal complexes before and since 2000. Isolates from ten studies (Bertrand et al. 2016; Chenal-Francisque et al. 2011; Haase et al. 2014; Huang et al. 2015; Jennison et al. 2017; Jensen et al. 2016; Kwong et al. 2016; Maury et al. 2016; Ragon et al. 2008; Wang et al. 2012) were used to evaluate CC frequency before (black) and since (gray) 2000. Data has been ranked based on frequency prior to 2000
Within these datasets are observations of other possible CCs that represent an emerging population. Haase et al. noted that CC101 was common in the 1950s but became less frequent until more recently when it expanded notably relative to other CCs (Haase et al. 2014). Additionally, Haase et al. noted that these isolates are typically associated with human clinical sources, a bias suggesting a higher pathogenic potential. However, an examination of CC101 from Australia (Jennison et al. 2017) and Europe (Bertrand et al. 2016), as well as the aggregated data presented here (Fig. 1), does not show the same expansion of this CC as seen by Haase et al. Similarly, the apparent expansion of CC121 may represent the impact of certain geographic regions. Evaluation of data from Australia suggests that CC1 and CC3 show no sign of decline and evidence of any emergent clones is less clear, though CC121 appears to be expanding in Australia as well as Europe. Studies in Asia are more limited but suggest that other CCs play a larger role in listeriosis in that region. Similarly, while not included in the aggregated data, an evaluation of clinical isolates from Canada showed a heavy predominance of CC8 (Knabel et al. 2012). While the aggregated dataset showed an increase in the merged CC (CC8 and CC16) since 2000, the percentage of CC8-CC16 (8%) observed is still lower than incidence of CC8 alone in the Canadian dataset (58%). The presence of differing dominant CCs in different geographic regions is expected as dietary choices and environmental factors are likely to impact the required survival adaptations necessary for L. monocytogenes transmission within different food matrices.
Besides phylogenetically defined groups, other subtypes may represent an emerging trend in listeriosis with their own risk factors to consider. Among the 13 serotypes of L. monocytogenes, over 95% of listeriosis cases are associated with serotype 1/2a, 1/2b, and 4b, with serotype 4b causing over half of these cases. This suggests that serotypes, which are broadly distinct phylogenetic groups, have differences in fitness depending on their evolutionary niche. Within these serotypes there may be subgroups representing emerging trends in listeriosis. One example is the recent series of outbreaks that have been linked to a variant of serotype 4b, termed 4bV or IVb-v1 (Burall et al. 2017a). This clade of L. monocytogenes was linked to a few atypical foods, specifically fresh produce, and presented atypical cases. The 4bV subtype of L. monocytogenes has been observed previously, with isolates dating back to 1959; however, these isolates historically had no epidemiological links and cases prior to 2014 were considered sporadic (Leclercq et al. 2011). Evaluation of some of these isolates by WGS (Burall et al. 2017b; Laksanalamai et al. 2013) further supports the possibility of these isolates being part of unrecognized outbreaks, though due to the lack of epidemiological linkage, this possibility cannot be proven (Burall et al. 2017b; Lee et al. 2014). Prior to 2013, only about 40 4bV isolates had been reported (Lee et al. 2014) while nearly 300 have been collected subsequently. This higher number may be due to the large number of isolates associated with a few outbreaks, which further supports an emerging trend associated with these strains. Additionally, evaluation of the geographic sources of these 4bV strains found that the clade associated with these outbreaks has been linked to a specific region in California, while the other large clade of 4bV strains has been predominantly linked to isolates from the eastern half of the USA, again underscoring the possibility of geographic differences influencing strain association.
Critically lacking in these studies is evaluation of these trends from both a temporal and geographic perspective. While there is clear evidence for CC6 as an emerging group within Europe, its presence wasn’t mentioned in either Australian study and was low in the Asian surveillance efforts. However, due to smaller sample sizes compared to European efforts, this may not be reflective of the overall population. Similarly, the 4bV expansion may represent the effects of a localized microcosm altered by local environmental factors, such as agricultural practices, within the overall global population dynamics. Whether these expansions represent persistent emerging global and/or regional trends or simply reflect a brief increase remains to be seen and underscores the need for on-going evaluation of CCs, 4bV, and other L. monocytogenes subgroups of relevance as they are identified for shifts in frequency globally and regionally, especially when considering the changing trends in listeriosis related to patient demographics (more elderly individuals compared to pregnant women), food, and serotypes (Cartwright et al. 2013; Goulet et al. 2008; Mammina et al. 2013; Munoz et al. 2012; Pontello et al. 2012).
L. monocytogenes clones and hyper virulence
The disease-causing potential of different L. monocytogenes strains has been difficult to measure because of the numerous variables associated with disease manifestation. These variables include, but are not limited to, small genetic differences between strains, sources of these strains which could affect epigenetic make-up, physiology, and genetics of the host population, the amount of L. monocytogenes ingested, and consumers’ health, as well as the treatment regimen they undergo. Nevertheless, the severity of different outbreaks and sporadic cases in terms of morbidity and mortality as well as the number of people affected and attack rate (number of people infected/number of people exposed) often point to some intrinsic difference in genomic composition of the strains associated with these illnesses. Recent efforts with WGS data have led to the identification of STs and CCs associated with outbreaks of invasive disease and sporadic cases (Hain et al. 2007; Orsi et al. 2011; Piffaretti et al. 1989). For example, CC1 (also known as Epidemic Clone 1) strains were associated with several major outbreaks in the USA from 1985 to 2000 (Kathariou 2002). Although the vast majority of listeriosis cases are restricted to the elderly, immunocompromised, and neonates, a few cases have been recorded where the infected individuals did not fall in these categories (Painter and Slutsker 2007). For example, in a recent outbreak involving caramel apples, three cases were found to be outside the “susceptible group” (Centers for Disease Control 2015). These cases and a few other recent outbreaks were caused by a clade of 4bV L. monocytogenes strains, indicating definite genetic differences from other 4b and 4bV strains (Burall et al. 2017a). However, absence of clinical data associated with the listeriosis cases as well as a reliable and sensitive ex vivo or animal model for virulence assessment also adds to the complexity of assessing the virulence potential of L. monocytogenes strains. Many animal models require a substantial number of laboratory grown bacterial cells (105–108 CFU) for any measurable end-point, which is not a true reflection of the number of cells that would be consumed. Maury et al. recently published a retrospective study in which they identified a few CCs correlated with cases of meningitis while other CCs were associated with cases of septicemia (Maury et al. 2016). They identified that the CCs involved in the meningitis cases had a more virulent phenotype than the strains associated with the septicemia cases. Interestingly, these meningitis-associated CC4 strains were also found to be more virulent in tissue culture and humanized mouse model of infection. In addtion to other genetic differences the so called “hyper virulent” CC4 strains contain an ~ 8 kb DNA fragment, termed LIPI-4 (Listeria pathogenicity island 4). At this point, the role of LIPI-4 genes, which are involved in carbohydrate metabolism, is debatable as this island was also found in L. innocua strain (Lecuit personal communication). It is still possible that LIPI-4, in addition to other virulence factors which are not present in L. innocua, could make 4b strains more virulent. Although this study was the first to identify the existence of such a pathogenicity island, its association with other listeriosis cases, whether they are meningitis, septicemia, or just self-limiting gastroenteritis requires further study.
Variation in virulence phenotypes
Given that certain clones of L. monocytogenes differ in their severity of disease based on clinical data, assessing variation in virulence can lead to identifying genetic elements that may be associated with virulence phenotypes. In addition to identifying LIPI-4 and hyper virulence of CC1, CC4, and CC6, Maury et al. demonstrated that strains of CCs more commonly isolated from foods, such as CC9 and CC121, were not as virulent in mice. This was represented by significantly higher loss of body weight and concentration of cells isolated from the spleen, liver, and mesenteric lymph nodes of mice infected with CC1, CC4, and CC6 strains compared to CC9 and CC121 strains (Maury et al. 2016). This work is one of the only studies assessing virulence phenotypes by clonal complex; other studies have measured variation in virulence by serotypes (Kuenne et al. 2013) or source of isolation (Rakic Martinez et al. 2017). While these studies have identified that serotype 4b strains or clinical isolates typically exhibit greater virulence, they have not identified any underlying genetic features that are associated with these differences (Kuenne et al. 2013; Rakic Martinez et al. 2017). Taken together, listeriosis severity based on outbreak data and in vitro and ex vivo virulence phenotypes provide evidence for variability in virulence properties of L. monocytogenes, likely linked to genetic differences among strains. Assessing virulence phenotypes among CCs or other WGS based subtypes will provide more insights into these genetic associations.
Variation in stress resistance phenotypes—food processing environment
L. monocytogenes is exposed to a number of environmental stressors both in foods and in the food processing environment. The ability of L. monocytogenes to adapt to these stresses facilitates its survival and transmission in the food supply. In food processing environments, persistence of L. monocytogenes has been associated with numerous outbreaks, as the processing environment was the most likely source of contamination for different foods (Angelo et al. 2017; McCollum et al. 2013). The ability of L. monocytogenes to survive and persist after cleaning and disinfection in food processing environments have been attributed to numerous factors, including colonization of niches and reservoirs out of disinfectant reach, biofilm formation, and activation of resistance mechanisms (Martinez-Suarez et al. 2016). Following prolonged exposure to sub-lethal concentrations of disinfectants, bacteria may develop increased resistance to the biocide in use (To et al., 2002). Increased resistance of bacteria in biofilms is a result of adaptation of bacteria to the environment within the biofilm, in addition to the intrinsic factors which may vary between the strains (Bridier et al. 2011; Kastbjerg and Gram 2009).
Environmental adaptations of L. monocytogenes are also associated with resistance to heavy metals, most commonly cadmium (Cd) and arsenic (As) (McLauchlin et al. 2004). Recent reports indicated that resistance to cadmium may correlate with resistance to quaternary ammonium compounds (QAC) (Katharios-Lanwermeyer et al. 2012). QACs are water soluble, broad spectrum antibacterial agents most commonly used in the food processing industry. Benzalkonium chloride (BC) is the most frequently used QAC in food processing plants (McDonnell and Russell 1999). Reports of tolerance to BC among L. monocytogenes isolates from food processing facilities vary between 10 and 46% (Aase et al. 2000; Mereghetti et al. 2000; Mullapudi et al. 2008; Soumet et al. 2005). So far, only a few molecular mechanisms for tolerance to BC have been elucidated. The genetic elements responsible for BC tolerance include bcrABC, a resistance cassette on a putative transposon harbored by pLM80-like plasmids (Elhanafi et al. 2010), a novel transposon Tn6188 (Muller et al. 2013), and small multidrug-resistant (SMR) efflux pump encoded by emrE carried by a novel genomic island LGI1 (Kovacevic et al. 2015).
The distribution of genetic elements associated with BC tolerance varies. Some elements, such as bcrABC and Tn6188, are widely distributed among strains as they are carried on mobile elements. The bcrABC resistance cassette has been identified in a strain related to the 1998/1999 hot dog-associated listeriosis outbreak in the USA (Elhanafi et al. 2010). The cassette is comprised of a transcriptional regulator bcrA and a multidrug resistance protein transporter bcrBC. This cassette is found to be widely disseminated among L. monocytogenes regardless of the serotype (Elhanafi et al. 2010). Its presence in nonpathogenic Listeria spp. indicates the possibility of these strains as reservoirs of BC and other resistance determinants for L. monocytogenes through the possibility of conjugative transfer (Katharios-Lanwermeyer et al. 2012). A novel transposon Tn6188, related to Tn554 from Staphylococcus aureus, can contribute to L. monocytogenes resistance to BC. When present in L. monocytogenes strains, this transposon along with other Tn554-like transposons have been shown to contribute to higher minimum inhibitory concentration of BC and significantly higher expression of qacH (coding a protein responsible for export of BC) in the presence of BC (Muller et al. 2013).
During the deadliest outbreak of listeriosis in Canada in 2008, a novel genomic island LGI1, specific to L. monocytogenes CC8 strains was discovered (Gilmour et al. 2010). Strains harboring this island belong to CC8, found to be predominant in listeriosis outbreaks and sporadic cases over two decades in Canada (Knabel et al. 2012; Kovacevic et al. 2015). LGI1 carries a small multidrug-resistant efflux pump encoded by emrE believed to be responsible for the increased tolerance to QACs. Expression of emrE gene in the presence of BC is upregulated raising the concern of possible adaptation and persistence of Listeria strains harboring this gene in the food processing environment (Kovacevic et al. 2015).
Resistance to heavy metals can correlate with resistance to QACs. Resistance to cadmium is frequent in Listeria, with several major outbreaks of listeriosis involving cadmium resistant L. monocytogenes isolates (Elhanafi et al. 2010). The same plasmid that carries bcrABC also harbors the cadmium efflux determinant cadA2 and trans-conjugants acquire both determinants (Katharios-Lanwermeyer et al. 2012). In addition to cadA2, the resistance to Cd is also conferred by plasmid-borne cadA1 and chromosomal cadA3—members of the cadAC efflux system (Lee et al. 2013). Recently, chromosomal cadA4 has been identified in strains belonging to CC1, CC2, and CC6 (Lee et al. 2013). Although the presence of cadA4 repressed virulence of L. monocytogenes in the Galleria mellonella insect model (Parsons et al. 2017), its effect on human listeriosis is not known.
Resistance to arsenic is chromosomally encoded with specific genes described recently (Harter et al. 2017; Lee et al. 2013). Harter et al. demonstrated the role of stress survival islet 2 (SSI-2) specific to the L. monocytogenes strains of sequence type (ST) 121 in pathogen survival under alkaline and oxidative stress caused by exposure to some commonly used disinfectants in the food processing environment (Harter et al. 2017). L. monocytogenes strains of serotype 4b harbor a 35-kb chromosomal Listeria genomic island 2 (LGI2) associated with arsenic resistance genes arsA1 and arsA2 (Lee et al. 2013). Lee et al. described LGI2 as a “floating island “with capacity to mobilize heavy metal resistance genes into different location in L. monocytogenes. They also speculated that due to the wide presence of this island among serotype 4b strains, especially CC1, CC2 and CC4, this island could have a role in L. monocytogenes virulence (Lee et al. 2017).
Variation in resistance to food-related stresses
Survival and growth capabilities of L. monocytogenes in foods, particularly in ready to eat (RTE) foods, are important from a food safety perspective. As L. monocytogenes is capable of growing at refrigerated temperature and has a high tolerance to salt, which are two main hurdles used to preserve foods, it is important to understand how L. monocytogenes cope with these stresses. Besides high salt and cold, foods containing other standard, generally recognized as safe (GRAS) preservatives have also been associated with human listeriosis. For this review, we will limit ourselves to L. monocytogenes response to low temperature, high salt, low pH, and presence of a few GRAS compounds including nisin. For a more comprehensive review on microbial stress response and adaptations, please refer to recent reviews on the topic (Milillo et al. 2012; NicAogain and O’Byrne 2016).
Various salts including NaCl are commonly used to control bacterial growth in foods. This is most relevant for RTE foods with a long shelf life including cheese, a food often associated with listeriosis. Detailed studies involving various strains of L. monocytogenes in laboratory media as well as isolation of L. monocytogenes from various foods showed that L. monocytogenes can tolerate as high as 10% NaCl although the growth rate decreases as function of salt concentration (Swaminathan et al. 2007). L. monocytogenes salt tolerance varies considerably from strain to strain. A recent study by Hingston et al. showed a wide range of variability among strains representing various CCs, with CC2 and CC11 exhibiting greater salt tolerance (Hingston et al. 2017b). While the study by Hingston et al. is one of the few to assess salt stress phenotypes by CC, older studies have examined variation in salt stress phenotypes by genetic lineage or serotype (Ribeiro and Destro 2014). Growth under salt stress varies by lineage and is also dependent on growth temperature. Lineage 1 strains grew at a significantly faster rate under 6% NaCl compared to lineage 2 strains, but this difference was only evident at 37 °C, not at 7 °C (Bergholz et al. 2010).
The use of osmoprotectants or compatible solutes by L. monocytogenes to manage osmotic stress is well-described (Burgess et al. 2016). These compounds include glycine betaine, carnitine, proline, glycerol, and trehalose, which can be found in foods at varying levels. The ability of L. monocytogenes to utilize compatible solutes or osmoprotectants to manage salt stress in foods would not only depend upon the genetic composition of the strain but also on the presence of osmoprotectants in that food. For example, foods such as alfalfa sprouts, beetroot, spinach, clams, and chicken contain elevated levels of glycine betaine, proline betaine, or trigonelline, and it would be expected that L. monocytogenes would be able to tolerate higher levels of salt in these foods compared to others with lower levels, e.g., cream cheese, celery, or butter (de Zwart et al. 2003). In addition to mechanisms to utilize osmoprotectants, L. monocytogenes utilize other mechanisms to alleviate osmotic stress which are not as well characterized (Bergholz et al. 2012). The increase in transcription of these genes in high-salt-containing media indicates that these genes play important roles in L. monocytogenes salt tolerance. Random mutagenesis experiments have identified several genes which encode functions involving salt transport as important for survival under salt stress, though the function of many of these genes is poorly understood (Burall et al. 2012; Burall et al. 2015).
L. monocytogenes has been isolated from several low acid foods indicating its ability to tolerate low pH (Koutsoumanis et al. 2003). The ability to tolerate acid varies from strain to strain, and can be related to growth phase and prior exposure to low pH (Datta and Benjamin 1997; Davis et al. 1996). In general, L. monocytogenes growth has been found to be completely inhibited at pH 4.5 and below, although the recent findings following the L. monocytogenes outbreak linked to caramel apples and several in vitro studies indicated that L. monocytogenes survival and growth in apples may be a function of other factors including titratable acidity and temperature of storage (Glass et al. 2015; Salazar et al. 2016). One of the most well-studied acid tolerance systems in L. monocytogenes is the glutamate decarboxylase system. Strain to strain differences in acid resistance were linked to differences in accumulation of the decarboxylated glutamate (γ-amino butyric acid, GABA) intracellulary, as well as differences in regulation of the genes encoding the decarboxylases and transporters involved in this system (Feehily et al. 2014; Karatzas et al. 2012).
Many GRAS substances are commonly used to suppress L. monocytogenes growth in RTE foods. Nisin, a well-characterized GRAS bacteriocin, has been studied in detail for its role in the control of L. monocytogenes in food (Datta and Benjamin 1997). The effectiveness of nisin depends on pH, salt concentration, and storage temperature of the food. Variation in sensitivity among strains has been reported (Begley et al. 2010). Strains belonging to CC6 and CC7 were found to have significantly higher innate resistance to nisin compared to strains belonging to CC2, CC3, CC5, CC9, and CC11 (Malekmohammadi et al. 2017). Adaptation of L. monocytogenes to either low pH (van Schaik et al. 1999) or osmotic stress (Bergholz et al. 2013) can significantly increase nisin resistance, with the potential for limiting the efficacy of nisin in some foods (Bonnet and Montville 2005).
L. monocytogenes is one of the few foodborne pathogens which grows relatively well at cold temperature (4–10 °C) provided that other conditions, e.g., pH, nutrients, and water activity are favorable. Several mechanisms have been implicated in the L. monocytogenes response to cold, and several genes have been identified whose functions are associated with optimum cold tolerance (Tasara and Stephan 2006). Strains can vary in cold tolerance, and greater levels of cold tolerance have been linked to increased transcript levels of known cold tolerance genes, including cspA and pgpH, demonstrating an association between genetic factors and cold tolerance (Arguedas-Villa et al. 2010; Lianou et al. 2006). Hingston et al. compared cold tolerance capabilities across different CCs of L. monocytogenes, and while they did not find any significant differences among CCs, they did find that strains of serotype 1/2c had lower cold tolerance compared to other serotypes (Hingston et al. 2017b). As low temperature is a very important tool for food safety, a thorough understanding of cold tolerance in L. monocytogenes will be extremely useful to develop new intervention strategies benefiting both public health and food industry.
L. monocytogenes can overcome many of the standard hurdles used to control microbes in the food industry. Development of increased tolerance to one hurdle may provide cross protection to another. As shown by genetic and phenotypic studies, genes which are involved in stress tolerance often have multiple overlapping functions thereby achieving a tolerance response to multiple stresses. Some of the genes contributing to the different stress responses have been identified using random or site-directed mutagenesis in commonly used reference strains (Begley et al. 2002; Burall et al. 2012; Hingston et al. 2015). The exact mechanisms by which these genes influence stress tolerance is yet to be determined.
Links between stress resistance and human listeriosis
Adaptation of L. monocytogenes to environmental stresses present in foods can influence their ability to survive subsequent stresses that may be encountered in the gastrointestinal tract of a potential host. For example, L. monocytogenes adapted to the pH and osmotic stresses present in different types of cheeses had significantly greater survival during subsequent challenge in a simulated gastric environment (Kapetanakou et al. 2017). Growth of L. monocytogenes in an oxygen-limited environment, similar to vacuum packaging of a food, significantly increased subsequent survival in a simulated gastric system (Sewell et al. 2015). It would be useful to assess a wider range of strains to determine if these phenomena are universally applicable.
As noted in the “Newly emerging clones of L. monocytogenes” section, certain subtypes of L. monocytogenes have been increasing in frequency, in particular CC6 in the Netherlands (Koopmans et al. 2017). Isolates subtyped as ST6 (belonging to CC6) were significantly associated with cases of listerial meningitis which had unfavorable outcomes (Koopmans et al. 2017). A genome-wide association analysis was conducted to identify genetic elements from these CC6 isolates that were associated with unfavorable outcomes, including mortality (Kremer et al. 2017). A novel phage, phiLMST6, and a novel plasmid, pLMST6, were both identified as strongly associated with ST6 isolates that were associated with more severe disease. A gene encoding a QAC efflux pump, emrC, was present on pLMST6. It is interest to note that additional isolates were surveyed for the presence of pLMST6, and it was found predominantly in CC6 isolates, though also occurred in one isolate each from CC9, CC8, and CC101 (Kremer et al. 2017). Isolates carrying pLMST6 had greater resistance to BC, as well as to gentamycin and amoxicillin (Kremer et al. 2017). This is the first study that provided links between an increased frequency of a given L. monocytogenes subtype and specific genetic elements that may contribute to its ability to survive in the environment and resist treatments in the host.
Future perspectives
With the significant increase in WGS data for L. monocytogenes, there is the opportunity to identify additional genetic elements, i.e., accessory genes that are associated with virulence and/or stress resistance phenotypes (Table 1). In addition to assessing these factors among CCs recently reported to be associated with more severe disease (including CC1, CC4, and CC6 (Kremer et al. 2017; Maury et al. 2016)), investigation of potential factors associated with the emergence of ST382 in the USA is also needed. Our understanding of the genes and regulatory elements involved in different stress responses has slowly expanded beyond that from the reference strains LO28, EGD, and EDGe, to include relevant outbreak- and food processing environment-associated strains (Burall et al. 2012; Hingston et al. 2017a; Liu et al. 2017; Stasiewicz et al. 2011). Genome-wide association studies have the potential to significantly transform our knowledge of the underlying genetic mechanisms leading to variation in stress response and virulence by utilizing the range of genetic diversity among L. monocytogenes strains.
References
Aase B, Sundheim G, Langsrud S, Rorvik LM (2000) Occurrence of and a possible mechanism for resistance to a quaternary ammonium compound in Listeria monocytogenes. Int J Food Microbiol 62(1–2):57–63
Adriao A, Vieira M, Fernandes I, Barbosa M, Sol M, Tenreiro RP, Chambel L, Barata B, Zilhao I, Shama G, Perni S, Jordan SJ, Andrew PW, Faleiro ML (2008) Marked intra-strain variation in response of Listeria monocytogenes dairy isolates to acid or salt stress and the effect of acid or salt adaptation on adherence to abiotic surfaces. Int J Food Microbiol 123(1–2):142–150
Angelo KM, Conrad AR, Saupe A, Dragoo H, West N, Sorenson A, Barnes A, Doyle M, Beal J, Jackson KA, Stroika S, Tarr C, Kucerova Z, Lance S, Gould LH, Wise M, Jackson BR (2017) Multistate outbreak of Listeria monocytogenes infections linked to whole apples used in commercially produced, prepackaged caramel apples: United States, 2014-2015. Epidemiol Infect 145(5):848–856. https://doi.org/10.1017/S0950268816003083
Arguedas-Villa C, Stephan R, Tasara T (2010) Evaluation of cold growth and related gene transcription responses associated with Listeria monocytogenes strains of different origins. Food Microbiol 27(5):653–660. https://doi.org/10.1016/j.fm.2010.02.009
Begley M, Cotter PD, Hill C, Ross RP (2010) Glutamate decarboxylase-mediated nisin resistance in Listeria monocytogenes. Appl Environ Microbiol 76(19):6541–6546
Begley M, Gahan CG, Hill C (2002) Bile stress response in Listeria monocytogenes LO28: adaptation, cross-protection, and identification of genetic loci involved in bile resistance. Appl Environ Microbiol 68(12):6005–12
Bergholz TM, den Bakker HC, Fortes ED, Boor KJ, Wiedmann M (2010) Salt stress phenotypes in Listeria monocytogenes vary by genetic lineage and temperature. Foodborne Pathog Dis 7(12):1537–1549
Bergholz TM, Bowen B, Wiedmann M, Boor KJ (2012) Listeria monocytogenes shows temperature-dependent and -independent responses to salt stress, including responses that induce cross-protection against other stresses. Appl Environ Microbiol 78(8):2602–2612. https://doi.org/10.1128/AEM.07658-11
Bergholz TM, Tang S, Wiedmann M, Boor KJ (2013) Nisin resistance of Listeria monocytogenes is increased by exposure to salt stress and is mediated via LiaR. Appl Environ Microbiol 79(18):5682–5688. https://doi.org/10.1128/AEM.01797-13
Bertrand S, Ceyssens PJ, Yde M, Dierick K, Boyen F, Vanderpas J, Vanhoof R, Mattheus W (2016) Diversity of Listeria monocytogenes strains of clinical and food chain origins in Belgium between 1985 and 2014. PLoS One 11(10):e0164283. https://doi.org/10.1371/journal.pone.0164283
Bonnet M, Montville TJ (2005) Acid-tolerant Listeria monocytogenes persist in a model food system fermented with nisin-producing bacteria. Lett Appl Microbiol 40(4):237–242. https://doi.org/10.1111/j.1472-765X.2005.01661.x
Bridier A, Briandet R, Thomas V, Dubois-Brissonnet F (2011) Resistance of bacterial biofilms to disinfectants: a review. Biofouling 27(9):1017–1032. https://doi.org/10.1080/08927014.2011.626899
Burall LS, Laksanalamai P, Datta AR (2012) Listeria monocytogenes mutants with altered growth phenotypes at refrigeration temperature and high salt concentrations. Appl Environ Microbiol 78(4):1265–1272. https://doi.org/10.1128/AEM.06576-11
Burall LS, Simpson AC, Chou L, Laksanalamai P, Datta AR (2015) A novel gene, lstC, of Listeria monocytogenes is implicated in high salt tolerance. Food Microbiol 48:72–82. https://doi.org/10.1016/j.fm.2014.12.008
Burall LS, Grim CJ, Datta AR (2017a) A clade of Listeria monocytogenes serotype 4b variant strains linked to recent listeriosis outbreaks associated with produce from a defined geographic region in the US. PLoS One 12(5):e0176912. https://doi.org/10.1371/journal.pone.0176912
Burall LS, Grim CJ, Mammel MK, Datta AR (2017b) A comprehensive evaluation of the genetic relatedness of Listeria monocytogenes serotype 4b variant strains. Front Public Health 5:241. https://doi.org/10.3389/fpubh.2017.00241
Burgess CM, Gianotti A, Gruzdev N, Holah J, Knochel S, Lehner A, Margas E, Esser SS, Sela Saldinger S, Tresse O (2016) The response of foodborne pathogens to osmotic and desiccation stresses in the food chain. Int J Food Microbiol 221:37–53. https://doi.org/10.1016/j.ijfoodmicro.2015.12.014
Cantinelli T, Chenal-Francisque V, Diancourt L, Frezal L, Leclercq A, Wirth T, Lecuit M, Brisse S (2013) Epidemic clones of Listeria monocytogenes are widespread and ancient clonal groups. J Clin Microbiol 51:3770–3779. https://doi.org/10.1128/JCM.01874-13
Cartwright EJ, Jackson KA, Johnson SD, Graves LM, Silk BJ, Mahon BE (2013) Listeriosis outbreaks and associated food vehicles, United States, 1998-2008. Emerg Infect Dis 19(1):1–9; quiz 184. https://doi.org/10.3201/eid1901.120393
Centers for Disease Control (2015) Multistate outbreak of Listeriosis linked to commercially produced, prepackaged caramel apples made from Bidart bros. Apples (Final Update). PUblisher. https://www.cdc.gov/listeria/outbreaks/caramel-apples-12-14/index.html
Chenal-Francisque V, Lopez J, Cantinelli T, Caro V, Tran C, Leclercq A, Lecuit M, Brisse S (2011) Worldwide distribution of major clones of Listeria monocytogenes. Emerg Infect Dis 17(6):1110–1112. https://doi.org/10.3201/eid/1706.101778
Datta AR, Benjamin MM (1997) Factors controlling acid tolerance of Listeria monocytogenes: effects of nisin and other ionophores. Appl Environ Microbiol 63(10):4123–4126
Davis MJ, Coote PJ, O’Byrne CP (1996) Acid tolerance in Listeria monocytogenes: the adaptive acid tolerance response (ATR) and growth-phase-dependent acid resistance. Microbiology 142(Pt 10):2975–2982
de Zwart FJ, Slow S, Payne RJ, Lever M, George PM, Gerrard JA, Chambers ST (2003) Glycine betaine and glycine betaine analogues in common foods. Food Chem 83(2):197–204. https://doi.org/10.1016/S0308-8146(03)00063-3
den Bakker HC, Didelot X, Fortes ED, Nightingale KK, Wiedmann M (2008) Lineage specific recombination rates and microevolution in Listeria monocytogenes. BMC Evol Biol 8:277
den Bakker HC, Desjardins CA, Griggs AD, Peters JE, Zeng Q, Young SK, Kodira CD, Yandava C, Hepburn TA, Haas BJ, Birren BW, Wiedmann M (2013) Evolutionary dynamics of the accessory genome of Listeria monocytogenes. PLoS One 8(6):e67511. https://doi.org/10.1371/journal.pone.0067511
Elhanafi D, Dutta V, Kathariou S (2010) Genetic characterization of plasmid-associated benzalkonium chloride resistance determinants in a Listeria monocytogenes strain from the 1998-1999 outbreak. Appl Environ Microbiol 76(24):8231–8238. https://doi.org/10.1128/AEM.02056-10
Feehily C, Finnerty A, Casey PG, Hill C, Gahan CG, O’Byrne CP, Karatzas KA (2014) Divergent evolution of the activity and regulation of the glutamate decarboxylase systems in Listeria monocytogenes EGD-e and 10403S: roles in virulence and acid tolerance. PLoS One 9(11):e112649. https://doi.org/10.1371/journal.pone.0112649
Freitag NE, Port GC, Miner MD (2009) Listeria monocytogenes—from saprophyte to intracellular pathogen. Nat Rev Microbiol 7(9):623–628
Gilmour MW, Graham M, Van Domselaar G, Tyler S, Kent H, Trout-Yakel KM, Larios O, Allen V, Lee B, Nadon C (2010) High-throughput genome sequencing of two Listeria monocytogenes clinical isolates during a large foodborne outbreak. BMC Genomics 11:120. https://doi.org/10.1186/1471-2164-11-120
Glass KA, Golden MC, Wanless BJ, Bedale W, Czuprynski C (2015) Growth of Listeria monocytogenes within a caramel-coated apple microenvironment. mBio 6(5):e01232-15. https://doi.org/10.1128/mBio.01232-15
Goulet V, Hedberg C, Le Monnier A, de Valk H (2008) Increasing incidence of listeriosis in France and other European countries. Emerg Infect Dis 14(5):734–740. https://doi.org/10.3201/eid1405.071395
Gray MJ, Zadoks RN, Fortes ED, Dogan B, Cai S, Chen Y, Scott VN, Gombas DE, Boor KJ, Wiedmann M (2004) Listeria monocytogenes isolates from foods and humans form distinct but overlapping populations. Appl Environ Microbiol 70(10):5833–5841
Haase JK, Didelot X, Lecuit M, Korkeala H, The Listeria monocytogenes MLST Study Group, Achtman M (2014) The ubiquitous nature of Listeria monocytogenes clones: a large-scale multilocus sequence typing study. Environ Microbiol 16(2):405–416. https://doi.org/10.1111/1462-2920.12342
Hain T, Chatterjee SS, Ghai R, Kuenne CT, Billion A, Steinweg C, Domann E, Karst U, Jansch L, Wehland J, Eisenreich W, Bacher A, Joseph B, Schar J, Kreft J, Klumpp J, Loessner MJ, Dorscht J, Neuhaus K, Fuchs TM, Scherer S, Doumith M, Jacquet C, Martin P, Cossart P, Rusniock C, Glaser P, Buchrieser C, Goebel W, Chakraborty T (2007) Pathogenomics of Listeria spp. Int J Med Microbiol 297(7–8):541–557. https://doi.org/10.1016/j.ijmm.2007.03.016
Harter E, Wagner EM, Zaiser A, Halecker S, Wagner M, Rychli K (2017) Stress survival islet 2, predominantly present in Listeria monocytogenes strains of sequence type 121, is involved in the alkaline and oxidative stress responses. Appl Environ Microbiol 83(16):e00827–e00817. https://doi.org/10.1128/AEM.00827-17
Hingston P, Chen J, Allen K, Truelstrup Hansen L, Wang S (2017a) Strand specific RNA-sequencing and membrane lipid profiling reveals growth phase-dependent cold stress response mechanisms in Listeria monocytogenes. PLoS One 12(6):e0180123. https://doi.org/10.1371/journal.pone.0180123
Hingston P, Chen J, Dhillon BK, Laing C, Bertelli C, Gannon V, Tasara T, Allen K, Brinkman FS, Truelstrup Hansen L, Wang S (2017b) Genotypes associated with Listeria monocytogenes isolates displaying impaired or enhanced tolerances to cold, salt, acid, or desiccation stress. Front Microbiol 8:369. https://doi.org/10.3389/fmicb.2017.00369
Hingston PA, Piercey MJ, Truelstrup Hansen L (2015) Genes associated with desiccation and osmotic stress in Listeria monocytogenes as revealed by insertional mutagenesis. Appl Environ Microbiol 81(16):5350–5362. https://doi.org/10.1128/AEM.01134-15
Huang YT, Ko WC, Chan YJ, Lu JJ, Tsai HY, Liao CH, Sheng WH, Teng LJ, Hsueh PR (2015) Disease burden of invasive listeriosis and molecular characterization of clinical isolates in Taiwan, 2000-2013. PLoS One 10(11):e0141241. https://doi.org/10.1371/journal.pone.0141241
Jennison AV, Masson JJ, Fang NX, Graham RM, Bradbury MI, Fegan N, Gobius KS, Graham TM, Guglielmino CJ, Brown JL, Fox EM (2017) Analysis of the Listeria monocytogenes population structure among isolates from 1931 to 2015 in Australia. Front Microbiol 8:603. https://doi.org/10.3389/fmicb.2017.00603
Jensen AK, Bjorkman JT, Ethelberg S, Kiil K, Kemp M, Nielsen EM (2016) Molecular typing and epidemiology of human listeriosis cases, Denmark, 2002–2012. Emerg Infect Dis 22(4):625–633. https://doi.org/10.3201/eid2204.150998
Kapetanakou AE, Gkerekou MA, Vitzilaiou ES, Skandamis PN (2017) Assessing the capacity of growth, survival, and acid adaptive response of Listeria monocytogenes during storage of various cheeses and subsequent simulated gastric digestion. Int J Food Microbiol 246:50–63. https://doi.org/10.1016/j.ijfoodmicro.2017.01.015
Karatzas KA, Suur L, O’Byrne CP (2012) Characterization of the intracellular glutamate decarboxylase system: analysis of its function, transcription, and role in the acid resistance of various strains of Listeria monocytogenes. Appl Environ Microbiol 78(10):3571–3579. https://doi.org/10.1128/AEM.00227-12
Kastbjerg VG, Gram L (2009) Model systems allowing quantification of sensitivity to disinfectants and comparison of disinfectant susceptibility of persistent and presumed nonpersistent Listeria monocytogenes. J Appl Microbiol 106(5):1667–1681. https://doi.org/10.1111/j.1365-2672.2008.04134.x
Katharios-Lanwermeyer S, Rakic-Martinez M, Elhanafi D, Ratani S, Tiedje JM, Kathariou S (2012) Coselection of cadmium and benzalkonium chloride resistance in conjugative transfers from nonpathogenic Listeria spp. to other Listeriae. Appl Environ Microbiol 78(21):7549–7556. https://doi.org/10.1128/AEM.02245-12
Kathariou S (2002) Listeria monocytogenes virulence and pathogenicity, a food safety perspective. J Food Prot 65(11):1811–1829
Knabel SJ, Reimer A, Verghese B, Lok M, Ziegler J, Farber J, Pagotto F, Graham M, Nadon CA, Canadian Public Health Laboratory Network, Gilmour MW (2012) Sequence typing confirms that a predominant Listeria monocytogenes clone caused human listeriosis cases and outbreaks in Canada from 1988 to 2010. J Clin Microbiol 50(5):1748–1751. https://doi.org/10.1128/JCM.06185-11
Koopmans MM, Bijlsma MW, Brouwer MC, van de Beek D, van der Ende A (2017) Listeria monocytogenes meningitis in the Netherlands, 1985–2014: a nationwide surveillance study. The J Infection 75(1):12–19. https://doi.org/10.1016/j.jinf.2017.04.004
Koutsoumanis KP, Kendall PA, Sofos JN (2003) Effect of food processing-related stresses on acid tolerance of Listeria monocytogenes. Appl Environ Microbiol 69(12):7514–7516
Kovacevic J, Ziegler J, Walecka-Zacharska E, Reimer A, Kitts DD, Gilmour MW (2015) Tolerance of Listeria monocytogenes to quaternary ammonium sanitizers is mediated by a novel efflux pump encoded by emrE. Appl Environ Microbiol 82(3):939–953. https://doi.org/10.1128/AEM.03741-15
Kremer PH, Lees JA, Koopmans MM, Ferwerda B, Arends AW, Feller MM, Schipper K, Valls Seron M, van der Ende A, Brouwer MC, van de Beek D, Bentley SD (2017) Benzalkonium tolerance genes and outcome in Listeria monocytogenes meningitis. Clin Microbiol Infect 23(4):265 e1–265 e7. https://doi.org/10.1016/j.cmi.2016.12.008
Kuenne C, Billion A, Mraheil MA, Strittmatter A, Daniel R, Goesmann A, Barbuddhe S, Hain T, Chakraborty T (2013) Reassessment of the Listeria monocytogenes pan-genome reveals dynamic integration hotspots and mobile genetic elements as major components of the accessory genome. BMC Genomics 14:47. https://doi.org/10.1186/1471-2164-14-47
Kwong JC, Mercoulia K, Tomita T, Easton M, Li HY, Bulach DM, Stinear TP, Seemann T, Howden BP (2016) Prospective whole-genome sequencing enhances National Surveillance of Listeria monocytogenes. J Clin Microbiol 54(2):333–342. https://doi.org/10.1128/JCM.02344-15
Laksanalamai P, Steyert SR, Burall LS, Datta AR (2013) Genome sequences of Listeria monocytogenes serotype 4b variant strains isolated from clinical and environmental sources. Genome Announcements 1(5). https://doi.org/10.1128/genomeA.00771-13
Leclercq A, Chenal-Francisque V, Dieye H, Cantinelli T, Drali R, Brisse S, Lecuit M (2011) Characterization of the novel Listeria monocytogenes PCR serogrouping profile IVb-v1. Int J Food Microbiol 147(1):74–77. https://doi.org/10.1016/j.ijfoodmicro.2011.03.010
Lee S, Rakic-Martinez M, Graves LM, Ward TJ, Siletzky RM, Kathariou S (2013) Genetic determinants for cadmium and arsenic resistance among Listeria monocytogenes serotype 4b isolates from sporadic human listeriosis patients. Appl Environ Microbiol 79(7):2471–2476. https://doi.org/10.1128/AEM.03551-12
Lee S, Ward TJ, Graves LM, Tarr CL, Siletzky RM, Kathariou S (2014) Population structure of Listeria monocytogenes serotype 4b isolates from sporadic human listeriosis cases in the United States from 2003 to 2008. Appl Environ Microbiol 80(12):3632–3644. https://doi.org/10.1128/AEM.00454-14
Lee S, Ward TJ, Jima DD, Parsons C, Kathariou S (2017) The arsenic resistance-associated Listeria genomic island LGI2 exhibits sequence and integration site diversity and a propensity for three Listeria monocytogenes clones with enhanced virulence. Appl Environ Microbiol 83(21):e01189–e01117. https://doi.org/10.1128/AEM.01189-17
Lianou A, Stopforth JD, Yoon Y, Wiedmann M, Sofos JN (2006) Growth and stress resistance variation in culture broth among Listeria monocytogenes strains of various serotypes and origins. J Food Prot 69(11):2640–2647
Liu X, Basu U, Miller P, McMullen LM (2017) Differential gene expression and filamentation of Listeria monocytogenes 08-5923 exposed to sodium lactate and sodium diacetate. Food Microbiol 63:153–158. https://doi.org/10.1016/j.fm.2016.11.009
Malekmohammadi S, Kodjovi KK, Sherwood J, Bergholz TM (2017) Genetic and environmental factors influence Listeria monocytogenes nisin resistance. J Appl Microbiol 123:262–270. https://doi.org/10.1111/jam.13479
Mammina C, Parisi A, Guaita A, Aleo A, Bonura C, Nastasi A, Pontello M (2013) Enhanced surveillance of invasive listeriosis in the Lombardy region, Italy, in the years 2006-2010 reveals major clones and an increase in serotype 1/2a. BMC Infect Dis 13:152. https://doi.org/10.1186/1471-2334-13-152
Martinez-Suarez JV, Ortiz S, Lopez-Alonso V (2016) Potential impact of the resistance to quaternary ammonium disinfectants on the persistence of Listeria monocytogenes in food processing environments. Front Microbiol 7:638. https://doi.org/10.3389/fmicb.2016.00638
Maury MM, Tsai YH, Charlier C, Touchon M, Chenal-Francisque V, Leclercq A, Criscuolo A, Gaultier C, Roussel S, Brisabois A, Disson O, Rocha EP, Brisse S, Lecuit M (2016) Uncovering Listeria monocytogenes hypervirulence by harnessing its biodiversity. Nat Genet 48:308–313. https://doi.org/10.1038/ng.3501
McCollum JT, Cronquist AB, Silk BJ, Jackson KA, O'Connor KA, Cosgrove S, Gossack JP, Parachini SS, Jain NS, Ettestad P, Ibraheem M, Cantu V, Joshi M, DuVernoy T, Fogg NW Jr, Gorny JR, Mogen KM, Spires C, Teitell P, Joseph LA, Tarr CL, Imanishi M, Neil KP, Tauxe RV, Mahon BE (2013) Multistate outbreak of listeriosis associated with cantaloupe. N Engl J Med 369(10):944–953. https://doi.org/10.1056/NEJMoa1215837
McDonnell G, Russell AD (1999) Antiseptics and disinfectants: activity, action, and resistance. Clin Microbiol Rev 12(1):147–179
McLauchlin J, Mitchell RT, Smerdon WJ, Jewell K (2004) Listeria monocytogenes and listeriosis: a review of hazard characterisation for use in microbiological risk assessment of foods. Int J Food Microbiol 92(1):15–33. https://doi.org/10.1016/S0168-1605(03)00326-X
Mereghetti L, Quentin R, Marquet-Van Der Mee N, Audurier A (2000) Low sensitivity of Listeria monocytogenes to quaternary ammonium compounds. Appl Environ Microbiol 66(11):5083–5086
Milillo SR, Friedly EC, Saldivar JC, Muthaiyan A, O’Bryan C, Crandall PG, Johnson MG, Ricke SC (2012) A review of the ecology, genomics, and stress response of Listeria innocua and Listeria monocytogenes. Crit Rev Food Sci Nutr 52(8):712–725. https://doi.org/10.1080/10408398.2010.507909
Moura A, Criscuolo A, Pouseele H, Maury MM, Leclercq A, Tarr C, Bjorkman JT, Dallman T, Reimer A, Enouf V, Larsonneur E, Carleton H, Bracq-Dieye H, Katz LS, Jones L, Touchon M, Tourdjman M, Walker M, Stroika S, Cantinelli T, Chenal-Francisque V, Kucerova Z, Rocha EP, Nadon C, Grant K, Nielsen EM, Pot B, Gerner-Smidt P, Lecuit M, Brisse S (2016) Whole genome-based population biology and epidemiological surveillance of Listeria monocytogenes. Nature microbiology 2:16185. https://doi.org/10.1038/nmicrobiol.2016.185
Mullapudi S, Siletzky RM, Kathariou S (2008) Heavy-metal and benzalkonium chloride resistance of Listeria monocytogenes isolates from the environment of turkey-processing plants. Appl Environ Microbiol 74(5):1464–1468. https://doi.org/10.1128/AEM.02426-07
Muller A, Rychli K, Muhterem-Uyar M, Zaiser A, Stessl B, Guinane CM, Cotter PD, Wagner M, Schmitz-Esser S (2013) Tn6188—a novel transposon in Listeria monocytogenes responsible for tolerance to benzalkonium chloride. PLoS One 8(10):e76835. https://doi.org/10.1371/journal.pone.0076835
Munoz P, Rojas L, Bunsow E, Saez E, Sanchez-Cambronero L, Alcala L, Rodriguez-Creixems M, Bouza E (2012) Listeriosis: an emerging public health problem especially among the elderly. The Journal of infection 64(1):19–33. https://doi.org/10.1016/j.jinf.2011.10.006
NicAogain K, O’Byrne CP (2016) The role of stress and stress adaptations in determining the fate of the bacterial pathogen Listeria monocytogenes in the food chain. Front Microbiol 7:1865. https://doi.org/10.3389/fmicb.2016.01865
Norton DM, Braden C (2007) Foodborne Listeriosis. In: Ryser ET (ed) Listeria, Listeriosis, and food safety. 3rd edn. CRC Press, Boca Raton, pp 305–356
Orsi RH, den Bakker HC, Wiedmann M (2011) Listeria monocytogenes lineages: genomics, evolution, ecology, and phenotypic characteristics. Int J Med Microbiol 301(2):79–96. https://doi.org/10.1016/j.ijmm.2010.05.002
Painter J, Slutsker L (2007) Listeriosis in humans. In: Ryser ET, Marth EH (eds) Listeria, Listeriosis, and food safety. 3rd edn. CRC press, Boca Raton, pp 85–110
Parsons C, Lee S, Jayeola V, Kathariou S (2017) Novel cadmium resistance determinant in Listeria monocytogenes. Appl Environ Microbiol 83(5):e02580–e02516. https://doi.org/10.1128/AEM.02580-16
Piffaretti JC, Kressebuch H, Aeschbacher M, Bille J, Bannerman E, Musser JM, Selander RK, Rocourt J (1989) Genetic characterization of clones of the bacterium Listeria monocytogenes causing epidemic disease. Proc Natl Acad Sci U S A 86(10):3818–3822
Pontello M, Guaita A, Sala G, Cipolla M, Gattuso A, Sonnessa M, Gianfranceschi MV (2012) Listeria monocytogenes serotypes in human infections (Italy, 2000-2010). Annali dell'Istituto Superiore di Sanita 48(2):146–150. https://doi.org/10.4415/ANN_12_02_07
Quereda JJ, Meza-Torres J, Cossart P, Pizarro-Cerda J (2017) Listeriolysin S: a bacteriocin from epidemic Listeria monocytogenes strains that targets the gut microbiota. Gut Microbes 8(4):384–391. https://doi.org/10.1080/19490976.2017.1290759
Ragon M, Wirth T, Hollandt F, Lavenir R, Lecuit M, Le Monnier A, Brisse S (2008) A new perspective on Listeria monocytogenes evolution. PLoS Pathog 4(9):e1000146. https://doi.org/10.1371/journal.ppat.1000146
Rakic Martinez M, Wiedmann M, Ferguson M, Datta AR (2017) Assessment of Listeria monocytogenes virulence in the Galleria mellonella insect larvae model. PLoS One 12(9):e0184557. https://doi.org/10.1371/journal.pone.0184557
Ribeiro VB, Destro MT (2014) Listeria monocytogenes serotype 1/2b and 4b isolates from human clinical cases and foods show differences in tolerance to refrigeration and salt stress. J Food Prot 77(9):1519–1526. https://doi.org/10.4315/0362-028X.JFP-13-548
Ribeiro VB, Mujahid S, Orsi RH, Bergholz TM, Wiedmann M, Boor KJ, Destro MT (2014) Contributions of sigma(B) and PrfA to Listeria monocytogenes salt stress under food relevant conditions. Int J Food Microbiol 177:98–108. https://doi.org/10.1016/j.ijfoodmicro.2014.02.018
Salazar JK, Carstens CK, Bathija VM, Narula SS, Parish M, Tortorello ML (2016) Fate of Listeria monocytogenes in fresh apples and caramel apples. J Food Prot 79(5):696–702. https://doi.org/10.4315/0362-028X.JFP-15-442
Sewell D, Allen S, Phillips CA (2015) Oxygen limitation induces acid tolerance and impacts simulated gastro-intestinal transit in Listeria monocytogenes J0161. Gut pathogens 7:11. https://doi.org/10.1186/s13099-015-0058-0
Soumet C, Ragimbeau C, Maris P (2005) Screening of benzalkonium chloride resistance in Listeria monocytogenes strains isolated during cold smoked fish production. Lett Appl Microbiol 41(3):291–296. https://doi.org/10.1111/j.1472-765X.2005.01763.x
Stasiewicz MJ, Wiedmann M, Bergholz TM (2011) The transcriptional response of Listeria monocytogenes during adaptation to growth on lactate and diacetate includes synergistic changes that increase fermentative acetoin production. Appl Environ Microbiol 77(15):5294–5306
Swaminathan B, Cabanes D, Zhang W, Cossart P (2007) Listeria monocytogenes. In: Doyle MP, Beuchat LR (eds) Food microbiology: fundamentals and frontiers. 3rd edn. ASM press, Washington DC
Tan MF, Siow CC, Dutta A, Mutha NV, Wee WY, Heydari H, Tan SY, Ang MY, Wong GJ, Choo SW (2015) Development of ListeriaBase and comparative analysis of Listeria monocytogenes. BMC Genomics 16(1):755. https://doi.org/10.1186/s12864-015-1959-5
Tasara T, Stephan R (2006) Cold stress tolerance of Listeria monocytogenes: a review of molecular adaptive mechanisms and food safety implications. J Food Prot 69(6):1473–1484
To MS, Favrin S, Romanova N, Griffiths MW (2002) Postadaptational resistance to benzalkonium chloride and subsequent physicochemical modifications of Listeria monocytogenes. Appl Environ Microbiol 68:5258–5264
van Schaik W, Gahan CG, Hill C (1999) Acid-adapted Listeria monocytogenes displays enhanced tolerance against the lantibiotics nisin and lacticin 3147. J Food Prot 62(5):536–539
Vermeulen A, Gysemans KP, Bernaerts K, Geeraerd AH, Van Impe JF, Debevere J, Devlieghere F (2007) Influence of pH, water activity and acetic acid concentration on Listeria monocytogenes at 7 degrees C: data collection for the development of a growth/no growth model. Int J Food Microbiol 114(3):332–341. https://doi.org/10.1016/j.ijfoodmicro.2006.09.023
Wang Y, Zhao A, Zhu R, Lan R, Jin D, Cui Z, Wang Y, Li Z, Wang Y, Xu J, Ye C (2012) Genetic diversity and molecular typing of Listeria monocytogenes in China. BMC Microbiol 12:119. https://doi.org/10.1186/1471-2180-12-119
Ward TJ, Ducey TF, Usgaard T, Dunn KA, Bielawski JP (2008) Multilocus genotyping assays for single nucleotide polymorphism-based subtyping of Listeria monocytogenes isolates. Appl Environ Microbiol 74(24):7629–7642
Acknowledgements
Teresa M. Bergholz and the work in the Bergholz lab is partially supported by the National Institute of Food and Agriculture, U.S. Department of Agriculture, under project no. ND02426. Mira Rakic-Martinez was supported by the Oak Ridge Institute for Science and Education Research Participation Program to the FDA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any experiments with human participants or animals performed by the authors.
Rights and permissions
About this article
Cite this article
Bergholz, T.M., Shah, M.K., Burall, L.S. et al. Genomic and phenotypic diversity of Listeria monocytogenes clonal complexes associated with human listeriosis. Appl Microbiol Biotechnol 102, 3475–3485 (2018). https://doi.org/10.1007/s00253-018-8852-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-8852-5