Abstract
Lignin is a biomass-derived aromatic polymer that has been identified as a potential renewable source of aromatic chemicals and other valuable compounds. The valorization of lignin, however, represents a great challenge due to its high inherent functionalization, what compromises the identification of chemical routes for its selective depolymerization. In this work, an in vitro biocatalytic depolymerization process is presented, that was applied to lignin samples obtained from beech wood through OrganoCat pretreatment, resulting in a mixture of lignin-derived aromatic monomers. The reported biocracking route comprises first a laccase-mediator system to specifically oxidize the Cα hydroxyl group in the β-O-4 structure of lignin. Subsequently, selective β-O-4 ether cleavage of the oxidized β-O-4 linkages is achieved with β-etherases and a glutathione lyase. The combined enzymatic approach yielded an oily fraction of low-molecular-mass aromatic compounds, comprising coniferylaldehyde and other guaiacyl and syringyl units, as well as some larger (soluble) fractions. Upon further optimization, the reported biocatalytic route may open a valuable approach for lignin processing and valorization under mild reaction conditions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The valorization of lignin represents a great challenge due to its complex chemical structure and heterogeneity (Rinaldi et al. 2016; Hendriks and Zeeman 2009; Ragauskas et al. 2006). Importantly, delivering added value products out of lignin would significantly enhance the economics of lignocellulose-based biorefineries (Ragauskas et al. 2014; Viell et al. 2013; Zakzeski et al. 2010). The β-O-4-ether linkage is the most prevalent type of intermolecular bond in lignin, typically accounting for 45–60% of the total linkages (Adler 1977), being an essential target for selective lignin depolymerization processes. In that respect, the selective oxidation of the β-O-4 structure combined with a subsequent treatment to cleave the β-ether bond (C-O bond) has recently emerged as a promising route to achieve a selective lignin depolymerization. Relevant examples propose the catalytic oxidation of lignin promoted by the 2,3-dichloro-5,6-dicyano-1,4-benzoquinone/tert-butyl nitrite/O2 (DDQ/tBuONO/O2) system (Lancefield et al. 2015); the oxidation of lignin with 4-acetamido 2,2,6,6-tetramethylpiperidine-1-oxyl (TEMPO) (5 mol%) in acidic media (Rahimi et al. 2013); or the formic acid-mediated depolymerization of oxidized lignin (Rahimi et al. 2014).
Apart from those successful chemical cases, also “White Biotechnology” holds potential for an efficient and selective lignin valorization (e.g., by applying processes at milder conditions to valorize more thermo-labile fractions of lignin). In nature, a β-aryl ether degradation pathway has been elucidated in Sphingobium sp. SYK-6 (Masai et al. 1989, 1993a, 1999, 2007), comprising several enzymes: (i) at least four different (NAD+)-dependent stereospecific alcohol dehydrogenases (LigD, LigL, LigN, and LigO) which catalyze the oxidation of the Cα alcohol to ketone (Masai et al. 1993b; Sato et al. 2009); (ii) stereospecific β-etherases LigE, LigF, and LigP to cleave either β(R)- or β(S)-ether linked substrate enantiomers with glutathione (GSH) consumption (Gall et al. 2014a; Masai et al. 2003; Picart et al. 2014); and (iii) at least one glutathione lyase (LigG) to further convert the formed glutathione adduct (LigG is stereospecific for the GS-β(R)-enantiomer with very low activity on the corresponding (S)-enantiomer) (Picart et al. 2015a). In the latter case, the GS group attached to the aromatic compound is eliminated, resulting in the release of oxidized glutathione (GSSG) (Gall et al. 2014b). A similar pathway was also reported for a Novosphingobium sp. strain MBES04 recently (Ohta et al. 2015). Following this biochemical pathway, Reiter et al. (2013) evaluated the cleavage of β-aryl ether linkages in organosolv- and Kraft-type lignins, reporting traces of monomeric and dimeric aromatic units with these lignin samples. When starting from milled wood lignin, Ohta et al. (2016) achieved a higher proportion of monomeric aromatic species (up to 6.6 wt%), which might be related to the lower degree of polymerization found in milled wood lignin (Crestini et al. 2011). Other enzymes that are applied in nature to degrade lignin include different peroxidases (lignin peroxidase, manganese peroxidase, versatile peroxidase, and dye-decolorizing peroxidases) as well as laccases (Bugg and Rahmanpour 2015; Chen et al. 2012; Pollegioni et al. 2015). Due to their intrinsic radical-based mechanisms, these enzymes catalyze a random lignin depolymerization, and may also cause repolymerization of already released monolignols (Martínez et al. 2005).
In the present work, a novel biocatalytic approach combining a laccase-mediator system (LMS), two β-etherases, and a glutathione lyase to depolymerize biomass-derived lignin samples is presented. The proof of concept of this biocatalytic approach is successfully demonstrated for the cleavage of β-O-4 bonds of lignin samples obtained from an OrganoCat biomass pretreatment (vom Stein et al. 2011), leading to promising yields of low-molecular-mass aromatics.
Materials and methods
Materials
All commercially available compounds were purchased and used as received, unless otherwise stated. The β-O-4 lignin model compounds 1 and 2, as well as the degradation products 3 and 4, were synthesized according to our previously reported procedures (Picart et al. 2014, 2015a). Guaiacol (>99% pure) was purchased from Sigma-Aldrich, Taufkirchen, Germany. β-Etherases LigE from Sphingobium sp. SYK-6 (GenBank: WP_014075192) and LigF-NA from Novosphingobium aromaticivorans (GenBank: WP_041551020), glutathione lyase LigG-TD from Thiobacillus denitrificans (GenBank: AAZ97003), and laccase lcc2 M3 from Trametes versicolor (GenBank: CAA77015) were prepared as described previously (Liu et al. 2013; Picart et al. 2014, 2015a).
Conversion of lignin model compound
Oxidation of β-O-4 lignin model compound 1 with the LMS
A 15 mL tube was loaded with the lignin model compound 1 (adlerol, 33.4 mg), laccase lcc2 M3 (1 U) and violuric acid monohydrate (18 mg) in 3.15 mL sodium acetate buffer (0.1 M, pH 5), and 0.35 mL of the ionic liquid 1-ethyl-3-methylimidazolium ethylsulfate ([EMIM] [EtSO4]) (99%, Iolitec, Heilbronn, Germany) were added. The reaction mixture was incubated at room temperature with shaking (250 rpm) for 5 days. The resulting precipitated white solid (adlerone, 2) was washed with water and then used for enzymatic ether bond cleavage without any further purification. Under these conditions, 92% conversion was achieved as confirmed by high-performance liquid chromatography (HPLC) analysis.
β-Etherase cleavage and deglutathionylation of oxidized β-O-4 lignin model compound 2
Enzymatic cleavage of the oxidized lignin model compound 2 was carried out in an Eppendorf tube containing 50 mM glycine/NaOH buffer, pH 9.5, 0.5 mM 2 dissolved in dimethylsulfoxide (DMSO; final concentration, 25 vol%), 1 mM GSH (reduced), and each 10 μg of purified β-etherases LigE and LigF-NA. The reaction mixture was incubated at 25 °C, and after 1-h incubation, the supernatant was analyzed by HPLC to obtain full conversion of 2 into guaiacol and the GSH-conjugated product 3.
Afterwards, water was added to the reaction mixture to reach a final DMSO concentration of 10% (concentrations higher than that drastically inhibited the activity of LigG-TD), and 10 μg purified glutathione lyase LigG-TD were added. The reaction mixture was incubated for 6 h at room temperature to achieve 88% conversion of the final product VG, as determined by HPLC. As the enzyme LigG-TD exhibits enantiopreference for one of the enantiomers, no full conversion could be achieved within the applied reaction time. The identity of 3 (GS-GVG) as well as 4 (VG) was confirmed by HPLC when compared to the retention time of an authentic standard.
Lignin fraction of OrganoCat
The OrganoCat process is a biphasic lignocellulose fractionation based on the dilute acid approach, using biogenic oxalic acid as catalyst (vom Stein et al. 2011). OrganoCat lignin was extracted from beech wood as follows:
In a glass-made high pressure reactor (Ace pressure tube, 80 mL) 20 mL 2-methyl-tetrahydrofuran (2-MeTHF) and the same volume of 0.1 M aqueous oxalic acid solution (referring to the aqueous phase) were introduced. One hundred grams per liter (referring to the aqueous phase) 10-mm beech wood particles were added, and the reactor was closed tightly. The mixture was magnetically stirred at room temperature for 20 min, before heating to 140 °C for 3 h. After reaching the desired reaction time, the reactor was cooled down to room temperature and opened. The resulting mixture was centrifuged at 4000 rpm for 5 min, and the organic phase containing the lignin was separated by decantation. To obtain the lignin, 2-MeTHF was removed under reduced pressure. The resulting OrganoCat lignin was used in subsequent experiments without further processing.
Conversion of OrganoCat lignin
Lignin oxidation by LMS
Two hundred milligrams of dry OrganoCat lignin, 40 mg of violuric acid, and 2 U of laccase lcc2 M3 were added to a mixture of 4 mL [EMIM] [EtSO4] and 36 mL sodium acetate buffer (0.1 M, pH 5). The reaction mixture was incubated at room temperature for 5 days to ensure complete oxidation of benzylic α-hydroxy groups. Afterwards, the solid (oxidized lignin) in the reaction mixture was collected by centrifugation (16,100×g, 4 °C), washed twice with distilled water, and dried.
The aqueous reaction mixture after lignin separation was extracted three times with ethyl acetate, and the combined organic extracts after solvent evaporation served as blank control in subsequent LC-MS measurements.
Lignin depolymerization
One hundred milligrams of oxidized lignin were dissolved in pure DMSO (1 mL) and added to 3 mL glycine-NaOH buffer, pH 9.5 (50 mM final concentration), containing reduced GSH (final concentration 1 mM) and each 0.1 mg of β-etherases LigE and LigF-NA. Thus, the final DMSO concentration was set to 25 vol% as β-etherases are inhibited at higher DMSO concentrations. The reaction mixture was incubated overnight at 25 °C to enable the β-ether bond cleavage in the lignin polymer. The reaction mixture was then added to 6 mL water (10-mL total volume) to lower the DMSO concentration to 10% as glutathione lyase LigG-TD is inhibited at a higher DMSO concentration. After addition of 0.1 mg LigG-TD, the reaction mixture was incubated overnight at 20 °C to remove the glutathione group attached to the aromatic conjugates. The reaction mixture was then acidified to pH 1 by the addition of 1 M hydrochloric acid (HCl), triggering the flocculation of polymeric lignin. This lignin residue was washed with ethyl acetate, and the aqueous phase was also extracted with ethyl acetate (3 × 6 mL). The organic soluble extracts and washings were combined and concentrated under reduced pressure to obtain a viscous yellowish oil. The organic insoluble material (residual lignin) was washed with water and dried under reduced pressure. The mass of the resulting material was 87.5 mg (corresponding to 87.5 wt% of the initial lignin).
Analytical methods
General
2D HSQC NMR spectra were recorded on a Bruker (Rheinstetten, Germany) AV 600 instrument. HPLC was performed on a Prominence modular HPLC system (Shimadzu, Duisburg, Germany) equipped with a Nucleosil 100-5 C18 column (4.6 × 150 mm; particle size 5 μm, Macherey-Nagel, Düren, Germany) at 25 °C. A mixture of water (49%), acetonitrile (50%), and phosphoric acid (1%) was used as the mobile phase with a flow rate of 1.0 mL min−1. All compounds were detected with a UV detector (SPD-M30A, Shimadzu) at a wavelength of λ = 280 nm.
LC-MS identification of low-molecular-weight aromatics
The organic soluble material (yellowish oil) after lignin biocracking and the organic soluble fraction extracted from only oxidized lignin (blank control) were analyzed by LC-MS. The depolymerization products were separated via liquid chromatography on an 1260 Infinity HPLC system (Agilent, Waldbronn, Germany) using a RP-8 LiChrospher column (125 × 2 mm, 5 μm, CS-Chromatographie Service, Langerwehe Germany) with the following conditions: injection volume 5 μL, column temperature 30 °C, an isocratic elution with methanol (45 vol%), water (54.9 vol%) and formic acid (0.1 vol%), flow rate of 0.5 mL min−1, and an overall measurement time of 30 min. The HPLC system was coupled both to UV detection (Agilent 1260 Infinity DAD, wavelength range 190–640 nm) and to electrospray ionization-quadrupole time-of-flight mass spectrometry (Agilent 6530 accurate mass Q-ToF LC/MS) for the identification of depolymerized products. The LC-ESI-Q-ToF-MS was also employed to separate and identify possible glutathione adducts of lignin-derived compounds in the aqueous DMSO fraction in addition to lignin-derived monomers in the yellowish oil and in the organic soluble fraction extracted from oxidized lignin. Thus, an ESI technique in the positive ionization mode was used for the measurement. The ESI conditions were set as follows: capillary voltage 3500 V, nozzle voltage 1500 V, gas temperature 300 °C, drying gas 8 L min−1, sheath gas temperature 350 °C, sheath gas flow 11 L min−1, fragmentor 100 V, and skimmer 30 V. The software MassHunter was used for measurement and data calculation. Identification of lignin-derived compounds was based on the use of exact mass, relative retention time, and peak shape data in the context of expected depolymerization products.
Results
Enzymatic depolymerization of lignin model compound 1
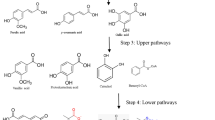
In the natural metabolic pathway for β-aryl ether bond cleavage involving bacterial β-etherases, the Cα hydroxy groups are first oxidized by multiple stereoselective alcohol dehydrogenases (ADHs), due to the fact that β-etherases are unable to cleave the respective non-oxidized ether linkages (Picart et al. 2014). Recently, laccases have also been shown to catalyze this oxidation reaction in lignin model compounds, avoiding the use of expensive cofactors (Majumdar et al. 2014). Hence, we hypothesized that a combination of laccase-mediator system (LMS) and bacterial β-etherases should enable the efficient depolymerization of lignin, while reducing the number of required enzymes for lignin biocracking. To prove this hypothesis, the biocracking route proposed in this work was first evaluated (and validated) on lignin model compound 1 (Fig. 1). Thus, a laccase-mediator system, composed of engineered laccase lcc2 M3 from T. versicolor and violuric acid as a mediator (Liu et al. 2013), was applied to specifically oxidize the Cα alcohol of 1 to the respective ketone. In a subsequent step, the oxidized β-O-4 structure 2 was subjected to two sequential reactions catalyzed by stereospecific β-etherases LigF-NA from N. aromaticivorans and LigE from Sphingobium sp. SYK-6 (Picart et al. 2014), cleaving the β-O-4 ether bond, and by glutathione lyase LigG-TD from T. denitrificans (Picart et al. 2015a), that cleaves the thioether linkage in the formed GSH adduct 3 leading to the release of β-deoxy-α-veratrylglycerone (VG, 4). In the first reaction step, the LMS treatment resulted in the selective oxidation of the benzylic alcohol of the model compound 1 (in the presence of 0.5 eq violuric acid and 10 vol% [EMIM] [EtSO4] as cosolvent). HPLC analysis (Fig. 2a, b) confirmed almost full conversion of 1 to 2a and 2b with complete selectivity for oxidation of the secondary benzylic alcohol over the primary alcohol. Subsequently, selective β-O-4 ether bond cleavage of 2a and 2b by LigE and LigF-NA, respectively, proceeded with complete conversion into GSH-conjugated intermediates 3a and 3b as shown by HPLC analysis (Fig. 1, step II; Fig. 2c). No cleavage products were observed when the non-oxidized model compound 1 was directly subjected to the β-etherase treatment. In the third reaction step (Fig. 1, step III; Fig. 2d), LigG-TD efficiently released the glutathione (GS-) group attached to 3a and 3b to achieve high yields (88% conversion) of the final product 4. As the latter enzyme still exhibits stereopreference for the conversion of 3b, compound 3a was likely not fully converted by LigG-TD within the 6-h reaction time.
HPLC-UV chromatograms of conversions of substrate 1 catalyzed by the multi-enzymatic system studied in this work. a Chromatogram of 1 without enzyme addition. b Oxidation of 1 with the LMS composed of lcc2 M3 and violuric acid. c Conversion of 2 with β-etherases LigE and LigF-NA. d Reaction of 3 with glutathione lyase LigG-TD. All products were identified by comparing their retention times with those of authentic standards
Enzymatic depolymerization of OrganoCat lignin
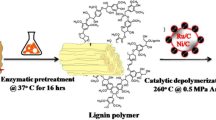
After successful cleavage of the β-O-4 model compound, the enzymatic biocracking system was further tested on authentic lignin polymer. To this end, lignin obtained from beech wood pretreated with the OrganoCat pulping system (vom Stein et al. 2011) was used. An exhaustive analytic characterization of the OrganoCat lignin has been published elsewhere recently (including 2D-NMR, elemental analysis, FTIR, ESI-MS, etc.) (Grande et al. 2015; Wiermans et al. 2015). A schematic representation of our enzymatic depolymerization process for real lignin is depicted in Fig. 3.
In the first step, OrganoCat lignin was successfully oxidized by the LMS system as revealed by NMR analysis of untreated and oxidized lignin samples (Fig. 4). In the presence of DMSO-d6/pyridin-d5 (4:1, v/v), signals of Aα and Aβ (Fig. 4, structure A) quantitatively disappeared and signal Aγ shifted to Aγ’ (Fig. 4, structure A’) due to the oxidation. Afterwards, insoluble oxidized lignin polymer was isolated from the aqueous reaction mixture (containing 10% [EMIM] [EtSO4]) by centrifugation. Sugars, or other low-molecular-mass compounds that might have been present in the original OrganoCat lignin (or were released during LMS treatment), remained soluble at 10% [EMIM] [EtSO4], and therefore were removed during lignin separation. The oxidized lignin polymer was then subjected to depolymerization by the combined use of β-etherases LigE and LigF-NA as well as the glutathione lyase LigG-TD. After 16-h incubation at room temperature, the reaction mixture was acidified with 1 M HCl to precipitate the residual lignin polymer. That insoluble fraction accounted for 87.5 wt% of the initially applied oxidized lignin, indicating a weight reduction of 12.5 wt% due to enzymatic depolymerization. The soluble fraction was extracted three times with ethyl acetate, and the combined organic extracts were concentrated under vacuum to obtain a viscous yellowish oil (Fig. 5), corresponding to a low-molecular-mass aromatics stream (Fig. 3).
Gel permeation chromatography (GPC) analysis of the lignin after oxidation by LMS and after the complete biocracking route (i.e., oxidation plus ether bond cleavage) revealed significant differences among them, as well as when compared to the untreated lignin, confirming that the lignin is indeed modified upon the sequence of different enzymatic treatments (supplementary Fig. S1). GPC analysis of oxidized lignin further suggests significant repolymerization, which was previously reported for laccase-treated lignin (Munk et al. 2015).
Identification of hydrolysis products from OrganoCat lignin
The obtained yellowish oil, containing the lignin-derived low-molecular-mass compounds, was further analyzed by high-resolution mass spectrometry coupled with liquid chromatography (LC-ESI-Q-ToF-MS) to identify low-molecular-mass products of β-etherase- and glutathione lyase-catalyzed OrganoCat lignin depolymerization (Table 1). The extracted soluble fraction of OrganoCat lignin after oxidation by the LMS served as a blank for LC-MS measurements. Thus, molar masses comparable to lignin-derived low-molecular-mass products that were not present in the blank control were found in the yellowish oil (Table 1, Fig. 6). The most abundant m/z value is 179 corresponding to coniferylaldehyde (7), a known degradation product of lignin (Eriksson et al. 1990). Quantification by LC-MS using an authentic standard resulted in about 1 mg of coniferylaldehyde, which corresponds to 8% of the total oily fraction. Additionally, m/z values of 153, 169, and 181 were detected and assigned to guaiacyl and syringyl units derived from lignin as well as a coniferylaldehyde isotope (Table 1) (Banoub and Delmas 2003; Morreel et al. 2010a, b; Saito et al. 2005). Besides these monomeric species, also other, e.g., oligomeric, species might be present in the oily fraction. Due to a lack of commercially available standards, a further clear allocation of these compounds remains speculative. Furthermore, also residual glutathione adducts of low-molecular-mass compounds, that were not converted by LigG-TD within the given reaction time, still seemed to be present in the aqueous DMSO fraction after extraction (supplementary Table S1). Possible explanations for the incomplete glutathione removal by LigG-TD could be the enzyme’s stereoselectivity (as explained previously for the conversion of model compound 3) as well as inhibition or deactivation of the enzyme over time.
Individual time profiles of the most abundant lignin depolymerization products. a Results obtained after enzymatic lignin biocracking (LMS + β-etherase + glutathione lyase). b Results obtained after lignin oxidation by LMS only. The first row corresponds to the total ion chromatogram, whereas the successive rows show the spectra of the major molecular ion peaks identified by LC-ESI-Q-ToF-MS with their m/z value
Discussion
In recent years, the valorization of lignocellulose has drawn significant attention as renewable starting material for the provision of chemicals and biofuels (Amore et al. 2016; Isikgor and Becer 2015). While the carbohydrate fractions have become the main focus for decades, since a few years also lignin valorization is attempted either using chemical or enzymatic catalysts. Whereas numerous chemical approaches for lignin depolymerization have been reported (Rinaldi et al. 2016; Zakzeski et al. 2010), only few of them allow for a selective lignin breakdown. On the other hand, microorganisms produce a range of enzymes to facilitate lignin degradation in nature (de Gonzalo et al. 2016; Palazzolo and Kurina-Sanz 2016; Pollegioni et al. 2015). Of these enzymes, only β-etherases enable a selective lignin depolymerization by cleaving the β-O-4 aryl ether bonds exclusively (Picart et al. 2015b). Though bacterial β-etherases are likely intracellular enzymes, due to the absence of respective secretion signal sequences, they have been shown to act also on polymeric substrates if carbonyl groups in the Cα position of β-O-4 aryl ether linkages are present (Picart et al. 2014). In real lignin, however, Cα hydroxyl groups are present instead, which need to be oxidized first. In the natural ether bond cleavage pathway, this oxidation is catalyzed by at least two stereoselective alcohol dehydrogenases (Sato et al. 2009). Since these alcohol dehydrogenases are cofactor (NAD+) dependent, its regeneration would be required for large-scale applications. Albeit different valuable approaches for NAD+ regeneration have been reported (Weckbecker et al. 2010), the use of cofactor-independent enzymes for lignin oxidation, such as laccases, constitutes an appealing alternative. Hence, we investigated the combination of β-etherases and laccase-mediator system for lignin conversion. To validate the hypothesis, we first applied the system to a model compound, which was successfully cleaved by the combination of enzymes. Results on model compounds, however, cannot be directly extrapolated to lignin polymer, as repolymerization of lignin and other chemical modifications may occur during lignocellulose pretreatment (Tolbert et al. 2014). Thus, experiments with real lignin are always needed to finally proof the strategy. Gratifyingly, our results clearly show that etherase-catalyzed β-O-4 aryl ether bond cleavage of polymeric lignin also works efficiently after prior LMS-catalyzed lignin oxidation. Hence, the combination of LMS with (R)- and (S)-selective β-etherases and glutathione lyase constitutes a promising alternative to the natural β-O-4 aryl ether bond cleavage pathway for application in biocatalytic lignin depolymerization under mild conditions.
The overall yield of low-molecular-weight aromatic compounds from beech wood lignin after etherase and glutathione lyase treatment, achieved under yet non-optimized conditions, was about 12.5 wt%, suggesting already a significant breakdown of the β-O-4 linkages present in the lignin polymer. In recent studies performed with aspen lignin, it was stated that the release of a monomer requires 2 β-O-4 linkages to be located next to each other, resulting in a probability to release a monomer of 0.16 (estimating 40% of the β-O-4 linkages present in aspen lignin), that is, releasable monomers would reach a maximum of 16% (Lancefield et al. 2015). In this respect, the overall yield of low-molecular-mass aromatics obtained in this work using OrganoCat lignin seems to be actually significant. For a reliable yield discussion, however, further characterization of OrganoCat lignin regarding its actual content of β-O-4 linkages would be necessary. Of the 12.5 wt% oily fraction of low-molecular-weight aromatics, 1 wt% (8% of the oily fraction) could be assigned to the formation of coniferylaldehyde, a known degradation product of lignin (Jones et al. 2010). This 1 wt% corresponds to 6% of the maximum obtainable yield of monomeric species if a 40% content of β-O-4 linkages present in OrganoCat lignin is assumed. Though further guaiacyl and syringyl units could be identified by LC–ESI-Q-ToF-MS, their actual yield could not be quantified. Additionally, other mono- and oligomeric species present in the oil could not be assigned due to the lack of commercial standards. Ohta et al. (2016) reported the formation of guaiacyl- and syringylhydroxypropanone as main products after depolymerization of milled wood lignins from Japanese cedar and Eucalyptus globulus wood using a combination of alcohol dehydrogenases, β-etherases, and glutathione lyase. Their yields of phenylpropanone monomers ranged from 2.4 to 4.7 wt% depending on the source of milled wood lignin. Likewise, Bouxin et al. (2015) highlighted the influence of the lignin source as well as the process of lignin preparation on the maximal obtainable yield of aromatic monomers in lignin depolymerization processes.
In future work, optimization of the multi-enzymatic process—with regard to the applied mediator, the use of cosolvents, and reaction conditions as well as by engineering of the employed enzymes—will be required to improve the depolymerization efficiency and to further increase the yield of low-molecular-weight aromatics. In this first proof of concept, lignin oxidation and depolymerization were performed as two separate steps due to non-matching pH and solvent requirements of the employed biocatalysts. Moreover, the applied mediator violuric acid was found to inhibit β-etherase activity. A careful fine-tuning of reaction conditions as well as engineering of the biocatalysts to meet process demands will thus enable the setup of one-pot reactions in the future.
Another aspect to be tackled for β-etherases is the enzymes’ dependence on glutathione as cofactor. Overall, two molecules of reduced glutathione (GSH) are required for the reductive cleavage of one β-O-4 aryl ether bond resulting in the formation of oxidized glutathione dimer (GSSG). The incorporation of a glutathione reductase for GSSG reduction during lignin depolymerization would allow GSH recycling and, hence, the use of only catalytic amounts of this cofactor (Reiter et al. 2013).
In a nutshell, the synergistic combination of three different enzyme activities (laccase + mediator/etherase/glutathione lyase)—catalyzing alcohol oxidation and subsequent ether bond cleavage—enables the selective depolymerization of OrganoCat lignin. A stream of low-molecular-mass lignins is generated under rather mild reaction conditions, leading to the isolation of an oily lignin derivative. Further optimization of the process could enable the production of an array of compounds from lignin, giving strong support to the setup of integrated biorefineries in which lignin can efficiently be valorized.
References
Adler E (1977) Lignin chemistry—past, present and future. Wood Sci Technol 11:169–218
Amore A, Ciesielski PN, Lin C-Y, Salvachúa D, Sànchez I Nogué V (2016) Development of lignocellulosic biorefinery technologies: recent advances and current challenges. Austr J Chem 69:1201–1218. doi:10.1071/CH16022
Banoub JH, Delmas M (2003) Structural elucidation of the wheat straw lignin polymer by atmospheric pressure chemical ionization tandem mass spectrometry and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J Mass Spectrom 38:900–903. doi:10.1002/jms.503
Bouxin FP, McVeigh A, Tran F, Westwood NJ, Jarvis MC, Jackson SD (2015) Catalytic depolymerisation of isolated lignins to fine chemicals using a Pt/alumina catalyst: part 1—impact of the lignin structure. Green Chem 17:1235–1242. doi:10.1039/c4gc01678e
Bugg TDH, Rahmanpour R (2015) Enzymatic conversion of lignin into renewable chemicals. Curr Opin Chem Biol 29:10–17. doi:10.1016/j.cbpa.2015.06.009
Chen YR, Sarkanen S, Wang YY (2012) Lignin-degrading enzyme activities. Methods Mol Biol 908:251–268. doi:10.1007/978-1-61779-956-3_21
Crestini C, Melone F, Sette M, Saladino R (2011) Milled wood lignin: a linear oligomer. Biomacromolecules 12:3928–3935. doi:10.1021/bm200948r
De Gonzalo G, Colpa DI, Habib MHM, Fraaije MW (2016) Bacterial enzymes involved in lignin degradation. J Biotechnol 236:110–119. doi:10.1016/j.jbiotec.2016.08.011
Eriksson K-E, Blanchette RA, Ander P (1990) Microbial and enzymatic degradation of wood and wood components. Springer, Berlin
Gall DL, Ralph J, Donohue TJ, Noguera DR (2014a) A group of sequence-related sphingomonad enzymes catalyzes cleavage of β-aryl ether linkages in lignin β-guaiacyl and β-syringyl ether dimers. Environ Sci Technol 48:12454–12463. doi:10.1021/es503886d
Gall DL, Kim H, Lu F, Donohue TJ, Noguera DR, Ralph J (2014b) Stereochemical features of glutathione-dependent enzymes in the Sphingobium sp. strain SYK-6 β-aryl etherase pathway. J Biol Chem 289:8656–8667. doi:10.1074/jbc.M113.536250
Grande PM, Viell J, Theyssen N, Marquardt W, Domínguez de María P, Leitner W (2015) Fractionation of lignocellulosic biomass using the OrganoCat process. Green Chem 17:3533–3539. doi:10.1039/C4GC02534B
Hendriks AT, Zeeman G (2009) Pretreatments to enhance the digestibility of lignocellulosic biomass. Bioresour Technol 100:10–18. doi:10.1016/j.biortech.2008.05.027
Isikgor FH, Becer R (2015) Lignocellulosic biomass: a sustainable platform for the production of bio-based chemicals and polymers. Polym Chem 6:4497–4559. doi:10.1039/c5py00263j
Jones RW, Reinot T, McClelland JF (2010) Molecular analysis of primary vapor and char products during stepwise pyrolysis of poplar biomass. Energy Fuel 24:5199–5209. doi:10.1021/ef100655n
Lancefield CS, Ojo OS, Tran F, Westwood NJ (2015) Isolation of functionalized phenolic monomers through selective oxidation and C-O bond cleavage of the β-O-4 linkages in lignin. Angew Chem Int Ed Engl 54:258–262. doi:10.1002/ange.201409408
Liu H, Zhu L, Bocola M, Chen N, Spiess AC, Schwaneberg U (2013) Directed laccase evolution for improved ionic liquid resistance. Green Chem 15:1348–1355. doi:10.1039/C3GC36899H
Majumdar S, Lukk T, Solbiati JO, Bauer S, Nair SK, Cronan JE, Gerlt JA (2014) Roles of small laccases from Streptomyces in lignin degradation. Biochemistry 53:4047–4058. doi:10.1021/bi500285t
Martínez AT, Speranza M, Ruiz-Dueñas FJ, Ferreira P, Camarero S, Guillén F, Martínez MJ, Gutiérrez A, del Río JC (2005) Biodegradation of lignocellulosics: microbial, chemical, and enzymatic aspects of the fungal attack of lignin. Int Microbiol 8:195–204
Masai E, Katayama Y, Nishikawa S, Yamasaki M, Morohoshi N, Haraguchi T (1989) Detection and localization of a new enzyme catalyzing the β-aryl ether cleavage in the soil bacterium (Pseudomonas paucimobilis SYK-6). FEBS Lett 249:348–352. doi:10.1016/0014-5793(89)80656-8
Masai E, Katayama Y, Kubota S, Kawai S, Yamasaki M, Morohoshi N (1993a) A bacterial enzyme degrading the model lignin compound β-etherase is a member of the glutathione-S-transferase superfamily. FEBS Lett 323:135–140. doi:10.1016/0014-5793(93)81465-C
Masai E, Kubota S, Katayama Y, Kawai S, Yamasaki M, Morohoshi N (1993b) Characterization of the Cα-dehydrogenase gene involved in the cleavage of β-aryl ether by Pseudomonas paucimobilis. Biotechnol Biochem 57:1655–1659. doi:10.1271/bbb.57.1655
Masai E, Katayama Y, Nishikawa S, Fukuda M (1999) Characterization of Sphingomonas paucimobilis SYK-6 genes involved in degradation of lignin-related compounds. J Ind Microbiol Biotechnol 23:364–373. doi:10.1038/sj.jim.2900747
Masai E, Ichimura A, Sato Y, Miyauchi K, Katayama Y, Fukuda M (2003) Roles of the enantioselective glutathione-S-transferases in cleavage of β-aryl ether. J Bacteriol 185:1768–1775. doi:10.1128/JB.185.6.1768-1775.2003
Masai E, Katayama Y, Fukuda M (2007) Genetic and biochemical investigations on bacterial catabolic pathways for lignin-derived aromatic compounds. Biosci Biotechnol Biochem 71:1–15. doi:10.1271/bbb.60437
Morreel K, Dima O, Kim H, Lu F, Niculaes C, Vanholme R, Dauwe R, Goeminne G, Inzé D, Messens E, Ralph J, Boerjan W (2010a) Mass spectrometry-based sequencing of lignin oligomers. Plant Physiol 153:1464–1478. doi:10.1104/pp.110.156489
Morreel K, Kim H, Lu F, Dima O, Akiyama T, Vanholme R, Niculaes C, Goeminne G, Inzé D, Messens E, Ralph J, Boerjan W (2010b) Mass spectrometry-based fragmentation as an identification tool in lignomics. Anal Chem 82:8095–8105. doi:10.1021/ac100968g
Munk L, Sitarz AK, Kalyani DC, Mikkelsen JD, Meyer AS (2015) Can laccases catalyze bond cleavage in lignin? Biotechnol Adv 33:13–24. doi:10.1016/j.biotechadv.2014.12.008
Ohta Y, Nishi S, Hasegawa R, Hatada Y (2015) Combination of six enzymes of a marine Novosphingobium converts the stereoisomers of β-O-4 lignin model dimers into the respective monomers. Sci Rep 5:15105–15118. doi:10.1038/srep15105
Ohta Y, Hasegawa R, Kurosawa K, Maeda AH, Koizumi T, Nishimura H, Okada H, Qu C, Saito K, Watanabe T, Hatada Y (2016) Enzymatic specific production and chemical functionalization of phenylpropanone platform monomers from lignin. ChemSusChem 10:425–433. doi:10.1002/cssc.201601235
Palazzolo MA, Kurina-Sanz M (2016) Microbial utilization of lignin: available biotechnologies for its degradation and valorization. World J Microbiol Biotechnol 32:173. doi:10.1007/s11274-016-2128-y
Picart P, Müller C, Mottweiler J, Wiermans L, Bolm C, Domínguez de María P, Schallmey A (2014) From gene towards selective biomass valorization: bacterial β-etherases with catalytic activity on lignin-like polymers. ChemSusChem 7:3164–3171. doi:10.1002/cssc.201402465
Picart P, Sevenich M, Domínguez de María P, Schallmey A (2015a) Exploring glutathione lyases as biocatalysts: paving the way for enzymatic lignin depolymerization and future stereoselective applications. Green Chem 17:4931–4940. doi:10.1039/C5GC01078K
Picart P, Domínguez de María P, Schallmey A (2015b) From gene to biorefinery: microbial β-etherases as promising biocatalysts for lignin valorization. Front Microbiol 6:916. doi:10.3389/fmicb.2015.00916
Pollegioni L, Tonin F, Rosini E (2015) Lignin-degrading enzymes. FEBS J 282:1190–1213. doi:10.1111/febs.13224
Ragauskas AJ, Williams CK, Davison BH, Britovsek G, Cairney J, Eckert CA, Frederick WJ Jr, Hallet JP, Leak DJ, Liotta CL, Mielenz JR, Murphy R, Templer R, Tschaplinki T (2006) The path forward for biofuels and biomaterials. Science 311:484–489. doi:10.1126/science.1114736
Ragauskas AJ, Beckham GT, Biddy MT, Chandra R, Chen F, Davis MF, Davison BH, Nixon RA, Glina P, Keller M, Langan P, Naskar AK, Saddler JN, Tschaplinski TJ, Tuskan GA, Wyman CE (2014) Lignin valorization: improving lignin processing in the biorefinery. Science 344:1246843. doi:10.1126/science.1246843
Rahimi A, Azarpira A, Kim H, Ralph J, Stahl SS (2013) Chemoselective metal-free aerobic alcohol oxidation in lignin. J Am Chem Soc 135:6415–6418. doi:10.1021/ja401793n
Rahimi A, Ulbrich A, Coon JJ, Stahl SS (2014) Formic-acid induced depolymerization of oxidized lignin to aromatics. Nature 13:249–252. doi:10.1038/nature13867
Reiter J, Strittmatter H, Wiemann LO, Schieder D, Sieber V (2013) Enzymatic cleavage of lignin β-O-4 aryl ether bonds via net internal hydrogen transfer. Green Chem 15:1373–1381. doi:10.1039/C3GC40295A
Rinaldi R, Jastrzebski R, Clough MT, Ralph J, Kennema M, Bruijnincx PC, Weckhuysen BM (2016) Paving the way for lignin valorisation: recent advances in bioengineering, biorefining and catalysis. Angew Chem Int Ed Engl 55:8164–8215. doi:10.1002/anie.201510351
Saito K, Kato T, Tsuji Y, Fukushima K (2005) Identifying the characteristic secondary ions of lignin polymer using ToF-SIMS. Biomacromolecules 6:678–683. doi:10.1021/bm049521v
Sato Y, Moriuchi H, Hishiyama S, Otsuka Y, Oshima K, Kasai D, Nakamura M, Ohara S, Katayama Y, Fukuda M, Masai E (2009) Identification of three alcohol dehydrogenase genes involved in the stereospecific catabolism of arylglycerol-β-aryl ether by Sphingobium sp. strain SYK-6. Appl Environ Microbiol 75:5195–5201. doi:10.1128/AEM.00880-09
Tolbert H, Akinosho R, Khunsupat AK, Naskar A, Ragauskas J (2014) Characterization and analysis of the molecular weight of lignin for biorefining studies. Biofuels Bioprod Biorefin 8:836–856. doi:10.1002/bbb.1500
Viell J, Harwardt A, Seiler J, Marquardt W (2013) Is biomass fractionation by Organosolv-like processes economically viable? A conceptual design study. Bioresour Technol 150:89–97. doi:10.1016/j.biortech.2013.09.078
vom Stein T, Grande PM, Kayser H, Sibilla F, Leitner W, Domínguez de María P (2011) From biomass to feedstock: one-step fractionation of lignocellulose components by the selective organic acid-catalyzed depolymerization of hemicellulose in a biphasic system. Green Chem 13:1772–1777. doi:10.1039/C1GC00002K
Weckbecker A, Gröger H, Hummel W (2010) Regeneration of nicotinamide coenzymes: principles and applications for the synthesis of chiral compounds. Adv Biochem Engin/Biotechnol 120:195–242. doi:10.1007/10_2009_55
Wiermans L, Schumacher H, Klaassen CM, Domínguez de María P (2015) Unprecedented catalyst-free lignin dearomatization with hydrogen peroxide and dimethyl carbonate. RSC Adv 5:4009–4018. doi:10.1039/C4RA13113D
Zakzeski J, Bruijnincx PCA, Jongerius AL, Weckhuysen BM (2010) The catalytic valorization of lignin for the production of renewable chemicals. Chem Rev 110:3552–3599. doi:10.1021/cr900354u
Acknowledgements
We thank Dr. Jakob Mottweiler and Prof. Carsten Bolm (Institute of Organic Chemistry, RWTH Aachen University) for the generous provision of lignin model substrates 1 + 2. Additionally, we thank Dr. Christoph Räuber (Institute of Organic Chemistry, RWTH Aachen University) for the preparation of 2D NMR figures.
This work was performed as part of the Cluster of Excellence “Tailor-Made Fuels from Biomass” [grant EXC 236], which is funded by the Excellence Initiative of the German Research Foundation to promote science and research at German universities.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 276 kb)
Rights and permissions
About this article
Cite this article
Picart, P., Liu, H., Grande, P.M. et al. Multi-step biocatalytic depolymerization of lignin. Appl Microbiol Biotechnol 101, 6277–6287 (2017). https://doi.org/10.1007/s00253-017-8360-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8360-z