Abstract
It has been documented that the purification of inclusion bodies from Escherichia coli by size exclusion chromatography (SEC) may benefit subsequent refolding and recovery of recombinant proteins. However, loading volume and the high cost of the column limits its application in large-scale manufacturing of biopharmaceutical proteins. We report a novel process using polyethylene glycol (PEG) precipitation under denaturing conditions to replace SEC for rapid purification of inclusion bodies containing recombinant therapeutic proteins. Using recombinant human interleukin 15 (rhIL-15) as an example, inclusion bodies of rhIL-15 were solubilized in 7 M guanidine hydrochloride, and rhIL-15 was precipitated by the addition of PEG 6000. A final concentration of 5% (w/v) PEG 6000 was found to be optimal to precipitate target proteins and enhance recovery and purity. Compared to the previously reported S-200 size exclusion purification method, PEG precipitation was easier to scale up and achieved the same protein yields and quality of the product. PEG precipitation also reduced manufacturing time by about 50 and 95% of material costs. After refolding and further purification, the rhIL-15 product was highly pure and demonstrated a comparable bioactivity with a rhIL-15 reference standard. Our studies demonstrated that PEG precipitation of inclusion bodies under denaturing conditions holds significant potential as a manufacturing process for biopharmaceuticals from E. coli protein expression systems.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biopharmaceutical drug development has expanded rapidly as recombinant proteins, and monoclonal antibodies are validated as clinical treatments (Zhu 2012). Protein expression systems involving Escherichia coli are frequently used in biopharmaceutical production because of cost-effectiveness, rapid production, and convenient handling and disposal. Heterogeneous expression of biopharmaceuticals in E. coli can result in the formation of insoluble inclusion bodies (IBs) (Jiang et al. 2013; Ouellette et al. 2003; Qi et al. 2015; Rahmen et al. 2015), which require additional protein denaturation and refolding steps to retrieve the biologically active target proteins. Since IBs are often contaminated with bacterial polysaccharides and outer membrane proteins, purification of IBs before denaturation/refolding can considerably improve target protein recovery yield. Size exclusion chromatography (SEC) has been reported as a useful method to purify IBs (Palmer and Wingfield 2012), but low loading volumes and high costs have significantly limited its industrial application. Development of a more efficient, scalable, and cost-effective process to purify IBs is of critical importance for biopharmaceutical production using E. coli protein expression systems.
Recombinant human interleukin 15 (rhIL-15) promotes immune cell proliferation and is relatively non-toxic (Jakobisiak et al. 2011). The cytokine uniquely activates the development and activity of both NK cells and CD8+ T cells (Nabekura and Lanier 2016; Van den Bergh et al. 2017), as well as promoting a persistent immune response through its action on memory T cells (Richer et al. 2015; Schenkel et al. 2016; Wang et al. 2016). In 2008, rhIL-15 was at the top of the National Cancer Institute’s list of potential biopharmaceuticals for tumor immunotherapy (Cheever 2008). Although IL-15 has been studied in multicenter clinical trials since 2012 (Croce et al. 2012), the cytokine is difficult to express in either yeast or mammalian protein expression systems (Huang et al. 2006; Sun et al. 2016). The rhIL-15 currently used in clinical development is manufactured by an E. coli protein expression system (Nellis et al. 2012; Vyas et al. 2012). Because the current manufacturing process produces a large quantity of host cell contaminants along with the raw rhIL-15 IBs, S-200 SEC chromatography was performed under denaturing conditions to remove these high molecular weight contaminants. Several problems were observed during the SEC purification of IBs, including a limited sample loading volume (usually 2–5% of column volume), extended packing height and slow loading speed, which led to a significant bottleneck within the manufacturing process. The development of a process that replaces SEC and separates IBs from other contaminants would be beneficial to biopharmaceutical industry.
Polyethylene glycol (PEG) precipitation has been widely used in purifying biological molecules including recombinant proteins and antibodies (Gagnon et al. 2014; Li et al. 2011; Oelmeier et al. 2013). PEG precipitation provides excellent recovery and purification of macromolecules, and the organic compound can be removed from the purified target protein during downstream ion-exchange or hydrophobic interaction columns that can accommodate large loading volumes (Giese et al. 2013; Li et al. 2011). Because of these advantages, the PEG precipitation method may be a suitable alternative to SEC during the purification of rhIL-15 IBs from E. coli protein expression systems.
In this report, we describe a novel production process using PEG precipitation, which successfully produced rhIL-15 with high purity and biological activity. Comparing to the previous reported method (Vyas et al. 2012), the PEG precipitation reduced the purification time by half and cost significantly less. More importantly, PEG precipitation is scalable to meet industrial production demands. All these advantages make PEG precipitation an ideal approach for protein manufacturing and isolation from E. coli protein expression systems.
Materials and methods
Reagents
E. coli BL21 (DE3) and DH5α were purchased from Microgene Laboratories (Shanghai, China). The anti-IL-15 monoclonal antibody was purchased from R&D Systems (Minneapolis, MN, USA). The rhIL-15 reference standard produced by an E. coli protein expression system was a gift from the Biological Resource Branch of the National Cancer Institute (Frederick, MD, USA). The rhIL-15 reference standard for biological activity was purchased from the National Institute of Biological Standards and Control (NIBSC; code 95/554; UK).
Bacteria fermentation and cytoplasmic protein expression
The coding sequences for human IL-15 were synthesized and cloned into vector pET28b to construct the plasmid pET28b/IL-15, which was transformed into E. coli BL21 (DE3) for rhIL-15 expression. Colonies were grown in tryptone Luria-Bertani (LB) broth (10 g/L tryptone, 10 g/L yeast extract, 10 g/L sodium chloride, supplemented with 100 μg/mL kanamycin). The 0.5 mL aliquot of the overnight culture was used to inoculate a 250-mL flask containing 20 mL soytone LB (10 g/L soytone-replaced tryptone). When the cell culture reached OD600 = 0.6, 1.0 mM IPTG was added to the culture to induce protein expression for 3 h.
For large-scale production, a 30 L fermenter (BIOTECH-30JSA, Shanghai Baoxing Bioengineering Equipment Co. Ltd., Shanghai, China) was prepared with 15 L of production medium (12 g/L soytone, 24 g/L yeast extract, 12 g/L glycerol, 12.5 g/L potassium phosphate dibasic, 3.8 g/L potassium phosphate monobasic, and 0.1 mL/L polypropylene glycol P2000 antifoam). The media was supplemented with 0.4 g/L magnesium sulfate heptahydrate and 0.05 g/L kanamycin. The culture parameters were set as follows: 37 °C, pH 7.0, aeration 0.5–1.0 VVM, and dissolved oxygen higher than 30% of saturation. A seed culture was grown in a flask overnight and inoculated into the fermenter at a final concentration of 2% (w/v). Once the cell density reached OD600 = 4–6, 1 mM IPTG was added to induce protein expression. The culture was harvested by centrifugation at 3 h post induction. The fermentation yielded approximately 450 g of cell paste from 15 L of culture. The cell paste was resuspended at 30% w/v in TES buffer (50 mM Tris-HCl, 20 mM EDTA, 100 mM NaCl, pH 7.4) and homogenized six times at 700–900 bar (ATS 1500, Shanghai, China). The IBs were collected by centrifugation at 23,000g, 4 °C for 30 min, washed three times with identical volumes of TES with 5% Triton X-100, followed by three washes with TES alone, and then stored at −70 to −90 °C.
PEG precipitation
Washed IBs were solubilized for 2 h in denaturing buffer I (50 mM Tris-HCl, 5 mM EDTA, 7 M guanidine hydrochloride [GdnHCl], pH 8.5, and 100 mM dithiothreitol) at 10 mL/g IBs. The supernatant was collected after centrifugation at 23,000g, 4 °C for 30 min. Several 2× PEG 6000 stock solutions (2–40% w/v) were prepared and added to the solubilized inclusion bodies. The mixtures were incubated at 4 °C for 2 h to precipitate rhIL-15. After centrifugation at 23,000g, 4 °C for 30 min, the PEG-precipitated proteins were collected and dissolved in denaturing buffer II (50 mM Tris-HCl, 5 mM EDTA, 7 M GdnHCl, pH 8.0, and 6 mM dithioerythritol) at the same volume as denaturing buffer I and gently solubilized for another 2 h. After centrifugation at 23,000g, 4 °C for 30 min, the supernatant was collected.
SEC purification of denatured IBs
The washed IBs were dissolved directly in denaturing buffer II at 10 mL/g IBs and gently solubilized for another 2 h before centrifugation to collect the supernatant. An S-200 column with 120 mL column volume (CV) was equilibrated with denaturing buffer II until the UV280 reached baseline. For each run, about 5 mL (4% of CV) of sample would be loaded on the column. Peaks were collected based on the absorbance (UV280), and the fractions were analyzed by SDS-PAGE and HPLC. To concentrate rhIL-15 and minimize the subsequent refolding volume, centrifugal ultrafiltration was performed using a Centriprep YM-10 (Millipore, Billerica, MA, USA).
Refolding
RhIL-15 refolding was conducted by drop-wise dilution of proteins (solubilized in denaturing buffer II) into chilled refolding buffer (100 mM Tris, 2 mM EDTA, 500 mM L-arginine, pH 9.5, with 1 mM oxidized glutathione and 1 mM reduced glutathione). Proteins were diluted 1:50 and stirred vigorously at 4 °C for 2–5 h. The pH was adjusted to 7.4 with 6 M HCl, and the conductivity was adjusted to 120 mS/cm with adjustment buffer (100 mM Tris, 2 mM EDTA, 500 mM L-arginine, 4 M NaCl, pH 7.4).
Chromatography purification of rhIL-15
Refolded rhIL-15 was purified by hydrophobic interaction (HIC), ion exchange (IEX), and size-exclusion chromatography. HIC resin (Butyl Sepharose HP, XK16/20, 1.6 × 6-cm bed) was equilibrated with five CVs of buffer (100 mM Tris, 2 mM EDTA, 500 mM L-arginine, 1.2 M NaCl, pH 7.4). Refolded rhIL-15 was loaded onto the column at 60 cm/h, followed by a wash with five CVs of HIC buffer A (100 mM Tris, 2 mM EDTA, 500 mM L-arginine, 1.2 M NaCl, pH 7.4) and three CVs of HIC buffer B (20 mM Tris, 1.2 M NaCl, pH 7.4). The rhIL-15 was eluted using a 10 CV linear gradient to 100% of HIC buffer C (20 mM Tris, 1.2 M NaCl, pH 7.4 to 20 mM Tris, 100 mM GdnHCl, pH 7.4) at a flow rate of 30 cm/h. A sample of each main peak fraction was analyzed by SDS-PAGE and ExRP-HPLC to determine purity and the level of deamidation.
IEX resin (Source 15Q, GE Healthcare, Tricon 5/200, 0.5 × 20-cm bed) was equilibrated with five CVs of IEX buffer A (50 mM Tris, 1 mM EDTA, 100 mM NaCl, pH 7.4). The HIC pool was adjusted to 11 ± 1 mS/cm with 50 mM Tris, 1 mM EDTA, pH 7.4, and loaded onto the IEX column at 600 cm/h, followed by a 10 CV wash. A 40 CV elution gradient was run at 300 cm/h up to 45% IEX buffer B (50 mM Tris, 1 mM EDTA, 1 M NaCl, pH 7.4). Main fractions were analyzed by SDS-PAGE. The eluted fractions were analyzed by ExRP-HPLC, and the non-deamidated proteins were selected and diluted with adjustment buffer (50 mM Tris, 1 mM EDTA, pH 7.4) to achieve a conductivity of 10–14 mS/cm before being loaded onto a pre-packed Q Sepharose XL (QXL) column (CV = 1 mL). The column was pre-equilibrated with IEX buffer C (50 mM Tris, 1 mM EDTA, 100 mM NaCl, and pH 7.4) and eluted directly with IEX buffer D (50 mM Tris, 1 mM EDTA, 1 M NaCl, pH 7.4). The fractions were collected. The main peak fractions were loaded onto a pre-packed Superdex™ 75-pg column (HiLoad™ 16/600, 120 mL, GE) using 25 mM sodium phosphate, 500 mM NaCl, pH 7.4. Collected fractions were sampled for quality analysis.
SDS-PAGE and Western blot
Samples taken at various stages of purification were analyzed by SDS-PAGE using 4–12% Bis–Tris polyacrylamide gels (Invitrogen, Carlsbad, CA, USA) to assess and identify the approximate molecular weight (MW) of rhIL-15. For guanidine-containing samples, cold ethanol precipitation was performed to extract proteins before loading (Sun et al. 2016). For Western blot of rhIL-15, electrophoresed samples were transferred to a PVDF membrane at 200 mA for 60 min. The membrane was incubated with mouse anti-human IL-15 (1:2000; R&D Systems Inc., Minneapolis, MN, USA) and goat anti-mouse IgG-HRP (1:10,000; Kirkegaard and Perry Laboratories Inc., Gaithersburg, MD, USA) for immunocharacterizing. Enhanced chemiluminescence (ECL) Western blot detection reagents (Merck Millipore, Billerica, MA, USA) were used to visualize proteins following the manufacturer’s instructions.
ExRP-HPLC
An expanded resolution reverse-phase HPLC method (ExRP-HPLC) was used to define the deamidation proportion of rhIL-15. An Agilent HPLC 1260 system (Agilent, Santa Clara, CA, USA) was fitted with two tandem-plumbed C18 columns (X-Bridge BEH300, C18, 3.5 μm, 250 mm, Waters, Milford, MA, USA). Mobile phases were 0.08% TFA and 0.02% formic acid in HPLC-grade water or acetonitrile. Elution was performed at 0.14-mL/min flow rate and monitored at 210 nm (min-%B 0–10, 12–0, 16–47, 116–53, 126–100, 128–100, 130–0, 142–10). Chromatograms were integrated and the area under the rhIL-15 peak was reported as a percentage of the total area detected.
SEC-HPLC
SEC-HPLC was used to verify the purity of rhIL-15 final product. Mobile phase buffers were prepared in 1× PBS (0.24 g/L KH2PO4, 1.44 g/L Na2HPO4, 8 g/L NaCl, 0.2 g/L KCl, pH 7.4). Samples were chilled prior to injection (25 μL) onto the column (G2000SWXL; 5 μm, 0.78 × 30 cm, Tosoh Biosciences, King of Prussia, PA), at a flow rate of 0.75 mL/min. Peaks were detected at 280 nm. Gel filtration standards (Bio-Rad Laboratories, Hercules, CA, USA) were run as controls. The purity of rhIL-15 was calculated as a percentage of the total peak area detected.
RPLC-MS
To confirm rhIL-15 identities, a Waters VION IMS quadrupole time-of-flight (QTOF) mass spectrometer interfaced with a Waters Acquity ultrahigh-performance liquid chromatography I class system was used for intact mass measurement. Mobile phases A and B (A 0.1% aqueous formic acid; B 0.1% formic acid in acetonitrile) were used with an Acquity UPLC BEH300 C18 (2.1 × 150 mm, 1.7 μm) column. The column was developed at 0.3 mL/min (min-%B 0–5, 5–5, 6–40, 15–60, 16–90, 20–5) and heated to 60 °C. Mass spectrometer conditions are as follows: capillary was 2.0 kV, sampling cone was 60 V, source temperature was 115 °C, desolvation temperature was 500 °C, and desolvation gas flow was 900 L/min.
In vitro CTLL-2 cell proliferation assay
The potency of rhIL-15 was determined using a colorimetric cell proliferation assay, which quantifies the IL-15-dependent CTLL-2 cell proliferation activity (Soman et al. 2009). Briefly, CTLL-2 cells were cultured in RPMI 1640 supplemented with 10% fetal bovine serum (FBS) and 200 U/mL rhIL-2. For the proliferation assay, cells were washed with rhIL-2 free medium and suspended to a density of 2 × 105 cells/mL. Recombinant human rhIL-15 samples were diluted to an initial concentration of 2.595 ng/mL (20 IU/mL for NIBSC standard) and serially diluted down to 0.045 ng/mL in the assay medium. The diluted samples were to a total of 6 individual wells in the 96-well plate at 50 μL per well. The prepared cell suspensions were seeded in the wells of the 96-well plate that containing rhIL-15 at different concentrations to yield a final cell density of 1 × 104 cells/100 μL/well. After 48 h of incubation at 37 °C in 5% CO2/95% humidity, cell viability was measured using Cell Counting Kit-8 (CCK-8; Donjindo, Japan). The plate was read at 450 nm, with a reference wavelength of 600 nm. The background readings in the wells with blank medium were subtracted from the results from the sample wells. The data was then analyzed using GraphPad Prism software and the four-parameter fit logistic equation.
In vivo stimulating effect of rIL-15 on memory T cells
Animal experiments were approved by the Animal Care and Use Committee of Shanghai Jiao Tong University. The specific pathogen-free, sex-matched, 8- to 10-week old C57BL/6 mice (SLACCAS, Shanghai, China) were held in air-filtered units at 23 ± 5 °C and 50 ± 15% relative humidity throughout the experimental period. RhIL-15 was diluted in 0.2% HSA to serial concentrations of 0, 0.1, 0.5, and 2.5 μg in a volume of 15 μL. Diluted rhIL-15 was injected subcutaneously into mice at days 1, 2, and 3. Mice were sacrificed on day 5, and the splenocytes were isolated and collected as previously described (Qian et al. 2012). Cells were then washed with PBS containing 0.2% BSA and incubated with anti-mouse CD3-PE-Cy5, anti-mouse CD8-APC, and anti-mouse CD44-PE for 30 min in the dark. After washes, the cells were fixed and permeabilized, followed by an incubation with an anti-Ki67 antibody for an additional 30 min. All the fluorescence-conjugated antibodies and the permeabilization solution were purchased from BD Pharmingen (Franklin Lakes, NJ, USA). The stained cells were washed twice and acquired using the CytoFlex flow cytometer (Beckman Coulter, China). Frequency of CD3+CD8+CD44+Ki67+ was analyzed as previously reported (Lu et al. 2012).
Results
RhIL-15 expression and isolation of inclusion bodies
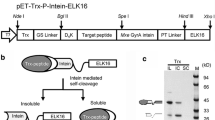
The coding sequence of human interleukin 15 (GeneBank nos. NM_172174 and NM_000585) was synthesized and inserted into the protein expression vector pET28b to obtain pET28b/IL-15, and the optimized stop codon TAATAATGA was placed at the end of the IL-15 coding sequence (Vyas et al. 2012). As shown in Fig. 1, rhIL-15 IBs were prepared by either PEG precipitation or SEC. BL21 (DE3) cells were transformed with pET28b/IL-15 and cultured as described in the “Materials and methods” section (Fig. S1). After cell lysis and recovery, approximately 35 g of rhIL-15 IBs was recovered from 450 g of cell paste (15 L culture).
PEG precipitation
The rhIL-15 IBs were first denatured by 7 M GdnHCl, and the collected supernatant was analyzed by SDS-PAGE and Western blot (Fig. 2a, lane 0). Large amounts of contaminating bacterial proteins and polysaccharides were detected, and it was necessary to remove these contaminants before the refolding step. To investigate the effect of PEG precipitation on contaminant removal, several 2× PEG 6000 solutions (from 1 to 20% w/v) were added to the denatured supernatant and tested for protein recovery (Fig. 2a). We found that a 5% (w/v) PEG 6000 solution could precipitate rhIL-15 with the best recovery and remove the majority of the impurities from the denatured protein solution.
RhIL-15 IB purification by PEG precipitation or SEC. a PEG concentrations used to precipitate rhIL-15. Lane M molecular weight standards, lane R BDP rhIL-15 reference standard (2 μg), lane 0 dissolved IBs in denaturing buffer I, lane S PEG supernatant, lane P PEG pellet. b The 5% (w/v) PEG precipitation data. Lane M molecular weight standards, lane R BDP rhIL-15 reference standard (2 μg), lane 1 dissolved IBs in denaturing buffer I, lane 2 precipitate from the IB dissolution, lane 3 PEG supernatants, lane 4 PEG pellet, lane 5 GdnHCl dissolved PEG pellet, lane 6 precipitate from the PEG pellet dissolution. c SEC elution profile. Peak fraction b was collected for further processing. d SDS-PAGE and Western blot analyses of SEC purification. Lane M molecular weight standards, lane R BDP rhIL-15 reference standard (2 μg), lane 1 dissolved IBs in denaturing II. lanes a–d peaks eluted in SEC
PEG precipitation and the SEC method were compared for manufacturing costs and process times by running the methods in parallel using the same batch of IBs. For the SEC method, an S-200 column was used to purify about 5 mL of samples for each run as described in the “Materials and methods” section. As shown in Fig. 2c, d, peak b contained most of the rhIL-15 and the least concentration of high molecular weight contaminants. For PEG precipitation, 10% PEG 6000 (w/v) was added to the denatured supernatant at a ratio of 1:1. The solution was placed at 4 °C for 2 h for precipitation of protein. After centrifugation at 4 °C, 23,000g for 30 min, the pellet was recovered and redissolved in denaturing buffer II and incubated for another 2 h before refolding (Fig. 2b). Both PEG precipitation and S-200 removed most of the contaminants (Fig. 2b, d). PEG precipitation saved about 50% of the time consumed in processing and 95% of reagent costs compared to the SEC method (Table 1). In addition, the procedure of PEG precipitation was relatively convenient and easy to scale up.
Refolding and purification of PEG precipitated rhIL-15
To validate that PEG precipitation is compatible with conventional chromatography well and facilitate target protein purification, we performed the downstream purification of rhIL-15 with steps of chromatography. The PEG precipitated protein pellet was refolded as described in the “Materials and methods” section and purified by Butyl Sepharose HP chromatography (Fig. 3a). SDS-PAGE analysis revealed that HIC can effectively remove the misfolded rhIL-15 (Fig. 3b, left panel, lane P, upper band), from the correctly folded, active rhIL-15. ExRP-HPLC analysis revealed that there are very low levels of deamidation in all fractions, consisting mainly of the isoaspartate form (Fig. 3c). Therefore, specific HIC fractions (a to g) were selected for further purification by Source 15Q chromatography.
Purification of rhIL-15 by Butyl Sepharose HP chromatography. a Chromatogram of rhIL-15 purified on a Butyl Sepharose HP column. Absorbance of fractions was monitored at 280 nm. Peak fractions (a–g) were pooled for further processing. b Coomassie blue-stained SDS-PAGE analysis of fractions a–g eluted from Butyl Sepharose HP column. left panel Non-reducing samples and right panel reducing samples. Lane M molecular weight markers, lane R BDP rhIL-15 reference of 2 μg, lane P HIC pre-column, lanes a–g HIC elution fractions. c HIC elution fractions “a” through “g” were analyzed by ExRP-HPLC, with reference standards of non-deamidated (“BDP ref”) or partially deamidated BDP rhIL-15 (“deamidated BDP ref”). Peaks are labeled as follows: “N” indicates non-deamidated rhIL-15, “B” indicates the isoaspartate form of deamidated rhIL-15, and “D” indicates the aspartate form of deamidated rhIL-15
From the Source 15Q column, rhIL-15 eluted as a major sharp peak (Fig. 4). According to ExRP-HPLC analysis, there were increasing proportions of deamidated rhIL-15 in the trailing peak (Fig. 4e, fractions (d–f)). The extra negative charge contributed by the deamidation likely made the molecules bind more tightly to the resin. The non-deamidated fractions were pooled (Fig. 4a, fractions (a–c)) and concentrated by QXL chromatography, with further purification by Superdex-75 chromatography (Fig. 4c). The purified protein was eluted in 25 mM sodium phosphate and 500 mM sodium chloride, pH 7.4, and evaluated for bioactivity.
Purification of rhIL-15 by Source 15Q and Superdex 75-pg chromatography. a Source 15Q elution profile. b Coomassie blue-stained SDS-PAGE analysis of reducing samples. Lane M molecular weight markers, lane R BDP rhIL-15 reference standard (2 μg), lanes a–f respective IEX column fractions. c Superdex 75-pg chromatography elution profile. d Coomassie blue-stained SDS-PAGE analysis of reducing samples. Lane M molecular weight markers, lane R BDP rhIL-15 reference standard (2 μg), lane P Superdex 75 pg pre-column samples collected from QXL elution, lanes 1–5 fractions 1–5. e ExRP-HPLC analysis of eluted fractions “a” through “f”. Peaks and reference standards are labeled as in Fig. 3
Chromatography of rhIL-15 on an S-200 column was also performed in parallel. The collected fractions containing rhIL-15 were refolded and purified by sequential Butyl Sepharose HP, Source 15Q, QXL, and Superdex 75 pg as described above (Fig. 5a). The final product from the Superdex 75-pg column was further analyzed by SDS-PAGE, ExRP-HPLC, and SEC-HPLC (Fig. 5b–d). Table 2 summarizes the yield, recovery rate, and deamidation proportion of each chromatography step. Using the two strategies, purified rhIL-15 can be produced at comparable yields and qualities, and PEG precipitation can be successfully combined with downstream chromatography steps to produce a highly purified product.
Purification and purity analysis of rhIL-15 from SEC purified IBs. a Chromatography profiles of (1) Butyl Sepharose HP, (2) Source 15Q, and (3) Superdex 75 pg (3). b SDS-PAGE analysis of reduced samples from Superdex 75 pg. Lane M molecular weight markers, lane R rhIL-15 reference standard (2 μg), lane P Superdex 75 pg pre-column samples, lanes 1–4 fractions 1–4. c ExRP-HPLC analysis of the deamidation proportion in the rhIL-15 final product. d SEC-HPLC analysis of purity of the rhIL-15 final product
ExRP-HPLC, SEC-HPLC, and RPLC-MS analyses of rhIL-15 product from PEG-precipitated IBs
To confirm the quality of the product, purified rhIL-15 was characterized by a panel of assays, including ExRP-HPLC, SEC-HPLC, and mass spectrometry. ExRP-HPLC was used to detect deamidation of the pool collected from each chromatography step, with BDP rhIL-15 as reference. Most of deamidated rhIL-15 was removed by Source 15Q, and <5% of deamidated rhIL-15 were left in the final product (Table 2 and Fig. 6a). SEC-HPLC analysis demonstrated that the purity of the product was at the same level with the BDP reference (Fig. 6b). RPLC-MS analysis was performed and confirmed that the molecular weight of the PEG process-purified rhIL-15 was 12,900.6 Da. This result matched the expected molecular weight of 12,900.5 Da. When the sample was partly deamidated, there was another major species with a molecular weight of 12,901.2 Da (Fig. 6d), consistent with an amide (−CONH2) deamidation and formation of a carboxyl group (−COOH) (Nellis et al. 2012).
ExRP-HPLC, SEC-HPLC, and mass spectroscopy analyses of purified rhIL-15. a Samples from each purification step were analyzed by rapid RP-HPLC. Final product eluted from Superdex 75 pg was analyzed by SEC-HPLC for purity (b) and RPLC-MS for confirmation of non-deamidated (c) and deamidated (d) fractions
The in vitro and in vivo activities of purified rhIL-15
The in vitro activity of the purified rhIL-15 was measured using a cell proliferation assay and a mouse T cell line (CTLL-2). RhIL-15 products purified by the parallel PEG precipitation and SEC procedures were evaluated, and the rhIL-15 NIBSC standard was used as control. There was no significant difference between the bio-activities of rhIL-15 obtained by either purification method, with EC50 of 1.07 × 107 IU/mg for PEG precipitation and 1.13 × 107 IU/mg for SEC process (Fig. 7).
The in vitro evaluation of rhIL-15 activation of CTLL-2 proliferation. Indicated amounts of rhIL-15 products purified by PEG precipitation (a) or SEC (b) processes, and the NIBSC standard (c), were added to the 96-well plates containing CTLL-2 cells at 1 × 104/well. Following 48-h incubation, cell proliferation was measured by CCK-8 assay. The data represents one of six independent experiments. d EC50 values of rhIL-15 products purified by PEG precipitation or SEC processes
We further validated the in vivo activity of purified rhIL-15 in mice by subcutaneously administering serial dosages (0, 0.1, 0.5, 2.5 μg per mouse) of the cytokine, once a day for 3 days. Two days after the last administration, the splenocytes were isolated to determine the frequency of Ki67-positive, proliferating memory T cells in the CD3+CD8+CD44+ subpopulation. As rhIL-15 dosage increased, the expression of Ki67 on memory T cells also increased (Fig. 8). The in vivo stimulation of memory T cells demonstrated that the PEG precipitation-purified rhIL-15 was biologically active.
RhIL-15 stimulates mouse memory T lymphocyte proliferation in vivo. Recombinant rhIL-15 proteins purified by PEG precipitation were administered subcutaneously to mice at serial dosages of 0, 0.1, 0.5, and 2.5 μg per mouse, once a day for 3 days. a The representative FACS profile of memory T cells from each treatment group. b Statistical analysis of the frequency of CD44+Ki67+ proliferating memory T lymphocytes in the CD3+CD8+ subpopulation. Higher dosages of rhIL-15 caused a statistically greater proliferative effect on memory T lymphocytes
Discussion
It has been reported that over 70% of eukaryotic proteins would aggregate to form inclusion bodies when expressed heterologously in E. coli (Yang et al. 2011). IBs must be solubilized and refolded into an active conformation to recover the target proteins (Tsumoto et al. 2003). Since the efficiency of refolding is highly dependent on the purity of the IB, it is important to remove contaminants before or during denaturation/resolubilization (Kumada et al. 2015; Ouellette et al. 2003).
When rhIL-15 was expressed in E. coli, there were many contaminants that co-aggregated with the target protein to form IBs. In a previously reported strategy, rhIL-15 IBs were denatured in 7 M GdnHCl and purified by S-200 SEC before refolding (Nellis et al. 2012; Vyas et al. 2012). Contaminants with high molecular weights were successfully removed, and contaminants with low molecular weights were removed in the downstream chromatography steps. However, due to its very limited loading capacity (2–5% of CV) and low flow rate, SEC is difficult to scale up and is rarely used as the first step in large-scale production. In addition, large amounts of GdnHCl are required for the preparation of SEC buffers, which increases cost, buffer viscosity, and difficulties in handling the buffer. Therefore, the SEC step was the bottleneck of the previous strategy. Development of a simple and cost-effective method for the purification of rhIL-15 inclusion bodies required substitution of a different procedure in place of SEC.
In this report, we studied the purification of rhIL-15 IBs with various concentrations of PEG 6000 (1–20% w/v). Based on SDS-PAGE and Western blot analyses of purified rhIL-15 (Fig. 2a), PEG 6000 removed most of the contaminants in IBs at all concentrations tested. Because of the chemical properties and physical volume of PEG, higher concentrations of PEG could precipitate more proteins. However, lower concentrations (2.5 and 5% w/v PEG) were able to precipitate the majority of rhIL-15, and this data was much lower than the reported 10–20% observed in other studies (Knevelman et al. 2010; Li et al. 2011). This observation could be due to the existence of high concentration of GdnHCl.
There are several advantages for using PEG precipitation to purify IB under denaturing conditions. First, PEG precipitation is a non-specific method to separate proteins by altering solubility of interested protein relative to those of many other proteins and macromolecules in a cell extract system (Sim et al. 2012). Compared to other precipitants such as ammonium sulfate ((NH4)2SO4) and polyethyleneimine (PEI), PEG is easy to remove and compatible with multiple types of chromatography, including HIC, CIEX, and AIEX (Zhao et al. 2012). Additionally, PEG precipitation works on highly concentrated protein solutions, so the working volume and protein mass can be controlled to ensure high scalability. Several reports have confirmed that PEG precipitation can be used to perform a rapid, scalable, and cost-effective purification of target proteins after cell lysis (Branston et al. 2015; Sim et al. 2012; Zhao et al. 2012). Furthermore, it has been recently reported that PEG with long chains prevented denatured protein from aggregation, which can benefit refolding processes in biopharmaceutical manufacturing (Yamamoto et al. 2017).
To evaluate the purification efficiency of PEG precipitation, we also performed the SEC purification of rhIL-15 IBs in our lab according to a previously reported methodology (Chertova et al. 2013). We used a pre-packed S-200 column with a CV of 120 mL, which could handle only about 5 mL of denatured IB for each run. This method required almost 1 day to complete the production steps, including buffer preparation, column manipulation, and subsequent ultrafiltration to further concentrate the protein (Table 1). Compared to the SEC method, PEG precipitation takes about half of the time and <5% of the cost of materials. As a primary capture method, PEG precipitation does not use expensive columns or the sophisticated AKTA system, which further reduces production costs. To evaluate the product quality of the two strategies, we established a series of analysis methods including ExRP-HPLC, SEC-HPLC, RPLC-MS, and the CTLL-2 cell-based proliferation assay. Our assay data demonstrated that the two processes resulted in similar product yield, purity, and bioactivity (Table 2). All the above results confirmed that PEG precipitation could purify high-quality target proteins with improved efficiency compared to SEC.
In conclusion, the PEG precipitation-based IB purification method has been developed and applied to the production of recombinant protein rhIL-15 under denaturing conditions. For the purification of rhIL-15 IBs, PEG precipitation can replace the previously reported S-200 chromatography with improved efficiency, reduced cost, and ease of operation. Our work demonstrated that PEG precipitation, in combination with denaturants, provides a new strategy for the purification of inclusion bodies. This application could potentially be applied in large-scale industrial production of biotechnological drugs from E. coli protein expression systems.
References
Branston SD, Wright J, Keshavarz-Moore E (2015) A non-chromatographic method for the removal of endotoxins from bacteriophages. Biotechnol Bioeng 112(8):1714–1719. doi:10.1002/bit.25571
Cheever MA (2008) Twelve immunotherapy drugs that could cure cancers. Immunol Rev 222:357–368. doi:10.1111/j.1600-065X.2008.00604.x
Chertova E, Bergamaschi C, Chertov O, Sowder R, Bear J, Roser JD, Beach RK, Lifson JD, Felber BK, Pavlakis GN (2013) Characterization and favorable in vivo properties of heterodimeric soluble IL-15.IL-15Ralpha cytokine compared to IL-15 monomer. J Biol Chem 288(25):18093–18103. doi:10.1074/jbc.M113.461756
Croce M, Orengo AM, Azzarone B, Ferrini S (2012) Immunotherapeutic applications of IL-15. Immunotherapy 4(9):957–969. doi:10.2217/imt.12.92
Gagnon P, Toh P, Lee J (2014) High productivity purification of immunoglobulin G monoclonal antibodies on starch-coated magnetic nanoparticles by steric exclusion of polyethylene glycol. J Chromatogr A 1324:171–180. doi:10.1016/j.chroma.2013.11.039
Giese G, Myrold A, Gorrell J, Persson J (2013) Purification of antibodies by precipitating impurities using polyethylene glycol to enable a two chromatography step process. J Chromatogr B Anal Technol Biomed Life Sci 938:14–21. doi:10.1016/j.jchromb.2013.08.029
Huang XQ, Hamilton MJ, Li CL, Schmidt C, Ellem KA (2006) An extraordinarily high level of IL-15 expression by a cell line transduced with a modified BMGneo vector displays hypoxic upregulation. Mol Biotechnol 33(1):49–56. doi:10.1385/MB:33:1:49
Jakobisiak M, Golab J, Lasek W (2011) Interleukin 15 as a promising candidate for tumor immunotherapy. Cytokine Growth Factor Rev 22(2):99–108. doi:10.1016/j.cytogfr.2011.04.001
Jiang H, Xie Y, Burnette A, Roach J, Giardina SL, Hecht TT, Creekmore SP, Mitra G, Zhu J (2013) Purification of clinical-grade disulfide stabilized antibody fragment variable—Pseudomonas exotoxin conjugate (dsFv-PE38) expressed in Escherichia coli. Appl Microbiol Biotechnol 97(2):621–632. doi:10.1007/s00253-012-4319-2
Knevelman C, Davies J, Allen L, Titchener-Hooker NJ (2010) High-throughput screening techniques for rapid PEG-based precipitation of IgG4 mAb from clarified cell culture supernatant. Biotechnol Prog 26(3):697–705. doi:10.1002/btpr.357
Kumada Y, Kang B, Yamakawa K, Kishimoto M, Horiuchi J (2015) Efficient preparation and site-directed immobilization of VHH antibodies by genetic fusion of poly(methylmethacrylate)-binding peptide (PMMA-tag). Biotechnol Prog 31(6):1563–1570. doi:10.1002/btpr.2169
Li H, Wang Y, Xu A, Li S, Jin S, Wu D (2011) Large-scale production, purification and bioactivity assay of recombinant human interleukin-6 in the methylotrophic yeast Pichia pastoris. FEMS Yeast Res 11(2):160–167. doi:10.1111/j.1567-1364.2010.00701.x
Lu H, Zhu S, Qian L, Xiang D, Zhang W, Nie A, Gao J, Wu M, Lu B, Yu Y, Han W, Moldenhauer A (2012) Activated expression of the chemokine Mig after chemotherapy contributes to chemotherapy-induced bone marrow suppression and lethal toxicity. Blood 119(21):4868–4877. doi:10.1182/blood-2011-07-367581
Nabekura T, Lanier LL (2016) Tracking the fate of antigen-specific versus cytokine-activated natural killer cells after cytomegalovirus infection. J Exp Med 213(12):2745–2758. doi:10.1084/jem.20160726
Nellis DF, Michiel DF, Jiang MS, Esposito D, Davis R, Jiang H, Korrell A, Knapp GC, Lucernoni LE, Nelson RE, Pritt EM, Procter LV, Rogers M, Sumpter TL, Vyas VV, Waybright TJ, Yang X, Zheng AM, Yovandich JL, Gilly JA, Mitra G, Zhu J (2012) Characterization of recombinant human IL-15 deamidation and its practical elimination through substitution of asparagine 77. Pharm Res 29(3):722–738. doi:10.1007/s11095-011-0597-0
Oelmeier SA, Ladd-Effio C, Hubbuch J (2013) Alternative separation steps for monoclonal antibody purification: combination of centrifugal partitioning chromatography and precipitation. J Chromatogr A 1319:118–126. doi:10.1016/j.chroma.2013.10.043
Ouellette T, Destrau S, Zhu J, Roach JM, Coffman JD, Hecht T, Lynch JE, Giardina SL (2003) Production and purification of refolded recombinant human IL-7 from inclusion bodies. Protein Expr Purif 30(2):156–166
Palmer I, Wingfield PT (2012) Preparation and extraction of insoluble (inclusion-body) proteins from Escherichia coli. Current protocols in protein science Chapter 6:Unit6 3. doi:10.1002/0471140864.ps0603s70
Qi X, Sun Y, Xiong S (2015) A single freeze-thawing cycle for highly efficient solubilization of inclusion body proteins and its refolding into bioactive form. Microb Cell Factories 14:24. doi:10.1186/s12934-015-0208-6
Qian L, Zhu S, Shen J, Han X, Gao J, Wu M, Yu Y, Lu H, Han W (2012) Expression and purification of recombinant human Mig in Escherichia coli and its comparison with murine Mig. Protein Expr Purif 82(1):205–211. doi:10.1016/j.pep.2011.12.009
Rahmen N, Fulton A, Ihling N, Magni M, Jaeger KE, Buchs J (2015) Exchange of single amino acids at different positions of a recombinant protein affects metabolic burden in Escherichia coli. Microb Cell Factories 14:10. doi:10.1186/s12934-015-0191-y
Richer MJ, Pewe LL, Hancox LS, Hartwig SM, Varga SM, Harty JT (2015) Inflammatory IL-15 is required for optimal memory T cell responses. J Clin Invest 125(9):3477–3490. doi:10.1172/JCI81261
Schenkel JM, Fraser KA, Casey KA, Beura LK, Pauken KE, Vezys V, Masopust D (2016) IL-15-independent maintenance of tissue-resident and boosted effector memory CD8 T cells. J Immunol 196(9):3920–3926. doi:10.4049/jimmunol.1502337
Sim SL, He T, Tscheliessnig A, Mueller M, Tan RB, Jungbauer A (2012) Protein precipitation by polyethylene glycol: a generalized model based on hydrodynamic radius. J Biotechnol 157(2):315–319. doi:10.1016/j.jbiotec.2011.09.028
Soman G, Yang X, Jiang H, Giardina S, Vyas V, Mitra G, Yovandich J, Creekmore SP, Waldmann TA, Quinones O, Alvord WG (2009) MTS dye based colorimetric CTLL-2 cell proliferation assay for product release and stability monitoring of interleukin-15: assay qualification, standardization and statistical analysis. J Immunol Methods 348(1–2):83–94. doi:10.1016/j.jim.2009.07.010
Sun W, Lai Y, Li H, Nie T, Kuang Y, Tang X, Li K, Dunbar PR, Xu A, Li P, Wu D (2016) High level expression and purification of active recombinant human interleukin-15 in Pichia pastoris. J Immunol Methods 428:50–57. doi:10.1016/j.jim.2015.12.002
Tsumoto K, Ejima D, Kumagai I, Arakawa T (2003) Practical considerations in refolding proteins from inclusion bodies. Protein Expr Purif 28(1):1–8. doi:10.1016/s1046-5928(02)00641-1
Van den Bergh JM, Lion E, Van Tendeloo VF, Smits EL (2017) IL-15 receptor alpha as the magic wand to boost the success of IL-15 antitumor therapies: the upswing of IL-15 transpresentation. Pharmacol Ther 170:73–79. doi:10.1016/j.pharmthera.2016.10.012
Vyas VV, Esposito D, Sumpter TL, Broadt TL, Hartley J, Knapp GC, Cheng W, Jiang MS, Roach JM, Yang X, Giardina SL, Mitra G, Yovandich JL, Creekmore SP, Waldmann TA, Zhu J (2012) Clinical manufacturing of recombinant human interleukin 15. I. Production cell line development and protein expression in E. coli with stop codon optimization. Biotechnol Prog 28(2):497–507. doi:10.1002/btpr.746
Wang Y, Rahman D, Mistry M, Lehner T (2016) The effect of cellular stress on T and B cell memory pathways in immunized and unimmunized BALB/c mice. J Biol Chem 291(39):20707–20717. doi:10.1074/jbc.M116.746057
Yamamoto E, Yamaguchi S, Nagamune T (2017) Protein refolding is improved by adding nonionic polyethylene glycol monooleyl ethers with various polyethylene glycol lengths. Biotechnol J. doi:10.1002/biot.201600689
Yang Z, Zhang L, Zhang Y, Zhang T, Feng Y, Lu X, Lan W, Wang J, Wu H, Cao C, Wang X (2011) Highly efficient production of soluble proteins from insoluble inclusion bodies by a two-step-denaturing and refolding method. PLoS One 6(7):e22981. doi:10.1371/journal.pone.0022981
Zhao Y, Kang L, Gao S, Gao X, Xin W, Wang J (2012) PEG precipitation coupled with chromatography is a new and sufficient method for the purification of botulinum neurotoxin type B [corrected]. PLoS One 7(6):e39670. doi:10.1371/journal.pone.0039670
Zhu J (2012) Mammalian cell protein expression for biopharmaceutical production. Biotechnol Adv 30(5):1158–1170. doi:10.1016/j.biotechadv.2011.08.022
Acknowledgements
This work was supported in part by the National Natural Science Foundation of China (No. 81273576 to Lu H., 81473127 to Zhu J.), the Science and Technology Commission of Shanghai Municipality (No. 15431907000 and 15DZ0503700 to Zhu J.), and the Medical and Engineering Cross Research Foundation of Shanghai Jiao Tong University (No. YM2013MS54 and YG2016QN27 to Lu H.). We would also like to thank the Biological Resources Branch (BRB) Preclinical Repository at the National Cancer Institute (Frederick, MD, USA), for supplying rhIL-15 reference.
Author information
Authors and Affiliations
Contributions
Huanhuan Chen, Ninghuan Li, Siwei Shi, Chencen Zhu, Han Luo, Lei Zhang, Junsheng Chen, Menglin Zhao, Lei Feng, and Huili Lu designed and performed the experiments. Yueqing Xie, Hua Jiang, Xiaoyi Yang, Cedric Cagliero, Huili Lu, and Jianwei Zhu analyzed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants. All animal studies were evaluated and approved by the Animal Care and Use Committee of Shanghai Jiao Tong University, prior to the commencement of any experiments.
Electronic supplementary material
ESM 1
(PDF 150 kb)
Rights and permissions
About this article
Cite this article
Chen, H., Li, N., Xie, Y. et al. Purification of inclusion bodies using PEG precipitation under denaturing conditions to produce recombinant therapeutic proteins from Escherichia coli . Appl Microbiol Biotechnol 101, 5267–5278 (2017). https://doi.org/10.1007/s00253-017-8265-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8265-x