Abstract
Type A chitinases (EC 3.2.1.14), GH family 18, attack chitin ((1 → 4)-2-acetamido-2-deoxy-β-d-glucan) and chito-oligosaccharides from the reducing end to catalyze release of chitobiose (N,N′-diacetylchitobiose) via hydrolytic cleavage of N-acetyl-β-d-glucosaminide (1 → 4)-β-linkages and are thus “exo-chitobiose hydrolases.” In this study, the chitinase type A from Serratia marcescens (SmaChiA) was used as a template for identifying two novel exo-chitobiose hydrolase type A enzymes, FbalChi18A and MvarChi18A, originating from the marine organisms Ferrimonas balearica and Microbulbifer variabilis, respectively. Both FbalChi18A and MvarChi18A were recombinantly expressed in Escherichia coli and were confirmed to exert exo-chitobiose hydrolase activity on chito-oligosaccharides, but differed in temperature and pH activity response profiles. Amino acid sequence comparison of the catalytic β/α barrel domain of each of the new enzymes showed individual differences, but ~69% identity of each to that of SmaChiA and highly conserved active site residues. Superposition of a model substrate on 3D structural models of the catalytic domain of the enzymes corroborated exo-chitobiose hydrolase type A activity for FbalChi18A and MvarChi18A, i.e., substrate attack from the reducing end. A main feature of both of the new enzymes was the presence of C-terminal 5/12 type carbohydrate-binding modules (SmaChiA has no C-terminal carbohydrate binding module). These new enzymes may be useful tools for utilization of chitin as an N-acetylglucosamine donor substrate via chitobiose.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chitin, a β-(1,4)-linked polymer of N-acetylglucosamine moieties (GlcNAc), is the second most abundant natural polysaccharide in the world besides cellulose. Chitin serves as the structural component in the exoskeleton of the arthropods (including crustaceans such as crabs and shrimp) and is also present in the cell walls of fungi and yeast (Synowiecki and Al-Khateeb 2003; Rinaudo 2006). Partial deacetylation of chitin results in chitosan; in contrast to chitin, chitosan is soluble in acidic aqueous solution. Both chitin and chitosan are of commercial interest due to their functional properties as well as their presumed biocompatibility and have recently been reported to have potential in applications ranging from biomaterials to drug delivery (Anitha et al. 2014). Since chitin is abundantly available, chitinases are interesting as processing aids for production of chito-oligosaccharides which present several possible medical applications, including inhibition of tumor growth (Shen et al. 2009) and of TH2-induced inflammation in asthma (Elias et al. 2005). In addition, chitobiose was recently shown to act as an N-acetylglucosamine donor substrate for enzymatic trans-glycosylation reactions to produce human milk oligosaccharide mimics (Nyffenegger et al. 2015). Chitinases (EC 3.2.1.14) are glycoside hydrolases that catalyze hydrolytic depolymerization of chitin into oligosaccharides by catalyzing the cleavage of the β-(1,4) linkages in the polymer backbone (Cohen-Kupiec and Chet 1998). Based on the similarity of amino acid sequences and their catalytic mechanism, chitinases can be divided into two glycosyl hydrolase (GH) families: GH family 18 and 19 (Lombard et al. 2014). Chitinases of family 18 are distributed among microbes, animals, and other organisms while GH 19 chitinases were first found in higher-order plants, and somewhat later, the first bacterial GH 19 chitinase was identified in Streptomyces griseus (Ohno et al. 1996). Up to now (January 2017), according to the CAZy database, chitinases of family 19 have been found in bacteria, eukaryotes, and viruses (Cantarel et al. 2009). Most of the available knowledge of microbial GH 18 chitinases is based on detailed work done on the three Serratia marcescens chitinases, ChiA (SmaChiA), ChiB (SmaChiB) (Fuchs et al. 1986; Van Aalten et al. 2000; Horn et al. 2006a, b), and ChiC (SmaChiC) (Suzuki et al. 1999; Synstad et al. 2008). In addition to the GH18 SmaChiA, SmaChiB, and SmaChiC, S. marcescens produces a lytic polysaccharide monooxygenase, CBP21, active on chitin (Suzuki et al. 1998; Vaaje-Kolstad et al. 2005). SmaChiA and SmaChiB are described as processive exolytic enzymes that catalyze the release of chitobiose from crystalline chitin acting from the reducing and non-reducing ends, respectively (Hult et al. 2005), whereas SmaChiC is an endo-acting enzyme attacking randomly in amorphous regions of the chitin backbone (Horn et al. 2006a, b; Sikorski et al. 2006). SmaChiA and SmaChiB are known as multidomain enzymes both having a central catalytic domain including a (β/α)8 triosephosphate isomerase (TIM) barrel fold structure containing the key catalytic motif DXXDXDXE (Van Aalten et al. 2001) and minimum one chitin-binding domain: SmaChiA has an N-terminal chitin-binding module, whereas SmaChiB has a C-terminal carbohydrate-binding module (CBM) 5/12 type chitin-binding module (and thus SmaChiA does not have a C-terminal CBM module and SmaChiB does not have an N-terminal chitin-binding module) (Vaaje-Kolstad et al. 2013).
The objective of the present work was to identify new chitinases able to catalyze the release of chitobiose from chitin as part of our quest to produce designed human milk oligosaccharides via enzymatic reactions on natural substrates, i.e., including using chitin derivatives as a source of GlcNAc moieties (Nyffenegger et al. 2015). In addition, the objective of the work was to improve the understanding of the action of chitinases on chito-oligosaccharides, and not least to investigate the chitin-binding domains (or modules) of GH18 chitinases. The SmaChiA enzyme was used as a sequence template in a genomic database mining approach to identify putative chitinases. On this basis, we report here the identification and characterization of two novel bacterial chitinases from marine microbes. Both enzymes are able to cleave the chito-oligosaccharides to produce chitobiose, and both enzymes contain the classic GH18 chitinase catalytic domain in addition to a C-terminal CBM with only low homology to SmaChiB.
Materials and methods
Chemicals
Chitin (from shrimp shells), isopropyl β-d-1-thiogalactopyranoside (IPTG), and imidazole were purchased from Sigma-Aldrich (Steinheim, Germany). 4-Nitrophenyl-N-acetyl-β-d-glucosamine (pNP-GlcNAc), 4-nitrophenyl N,N′-diacetyl-β-d-chitobioside (pNP-GlcNAc2), and 4-nitrophenyl N,N′,N″-triacetyl-β-d-chitotriose (pNP-GlcNAc3) were purchased from Carbosynth (Compton, UK). N,N′-Diacetyl chitobiose, N,N′,N″-triacetyl chitotriose, N,N′,N″,N′″-tetraacetyl chitotetraose, N,N′,N″,N′″,N′″′-pentaacetyl chitopentaose, and N,N′,N″,N′″,N′″′,N′″″-hexaactyl chitohexaose were purchased from Megazyme (Wicklow, Ireland). Restriction enzymes were purchased from Thermo Fischer Scientific (MA, USA).
Strains and genes
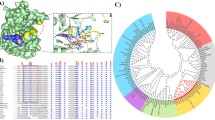
The SmaChiA amino acid sequence (GenBank accession no. AAA26551.1) was used as a sequence template for alignment analysis (blast) to select putative GH18 chitinases or GH18 hypothetical proteins of bacteria in the GenBank database. Selection of putative chitinases was focused on candidates that had 40–70% sequence similarity with SmaChiA and which were from non-pathogenic sources. Clustal Omega (Sievers et al. 2011) was used to construct the multiple alignment for the selected putative chitinases (20 different sequences) and further used to generate the neighbor-joining phylogenetic tree using SeaView (Gouy et al. 2010). Two bacterial chitinases from Ferrimonas balearica (original GenBank accession no. ADN76700.1 and new GenBank accession no. KY131984 (codon-optimized sequence for Escherichia coli expression)) and Microbulbifer variabilis (original GenBank accession no. WP_051089467.1 and new GenBank accession no. KY100262), respectively, were selected based on their close distance in the phylogenetic tree (Fig. S1) to the benchmark enzyme SmaChiA from Serratia marcescens (GenBank accession no. AAA26551.1) (Brurberg et al. 1994). These two new chitinase enzymes are referred to as FbalChi18A and MvarChi18A, respectively.
The putative chitinase-encoding genes for FbalChi18A and MvarChi18A were synthesized by DNA 2.0© (Newark, CA, USA) with a C-terminal His6 tag and codon-optimized for E. coli expression. The expression vector pJ414 was used as provided by DNA 2.0©. The E. coli strain DH5α (Invitrogen® Life Technologies, Thermo Fisher Scientific, MA, USA), was used as cloning host and E. coli BL21 (DE3), C41 (DE3), C43 (DE3), TOP10 (DE3), and TUNER (DE3) (Novagen and Lucigen, USA) were used as transformation and expression hosts of the putative chitinases.
Expression and purification of novel chitinases
Enzyme expression was done principally as described previously (Nyffenegger et al. 2015). The E. coli transformants (BL21 (DE3) turned out to be best) harboring the individual recombinant chitinase genes (FbalChi18A and MvarChi18A, respectively) were induced for overexpression with 1 mM IPTG as the cell culture reached an OD600 = 0.6 and were grown further overnight at 30 °C (for FbalChi18A) and 25 °C (for MvarChi18A) in LB medium with 100 μg ml−1 ampicillin. Cells were harvested by centrifugation, and the pellets were resuspended in binding buffer (20 mM sodium phosphate buffer, 500 mM NaCl, 20 mM imidazole, pH 7.8) before being disrupted by sonication and further centrifugation (20,000×g, 20 min at 4 °C) for removal of cell debris. The supernatant obtained by centrifugation was then filtered through a 0.22-μm filter and applied to a 5-ml Ni2+ Sepharose HisTrap HP column (GE Healthcare, Uppsala, Sweden) which was equilibrated with binding buffer using an Äkta purifier (GE Healthcare, Uppsala, Sweden). Proteins were eluted by a linear gradient of elution buffer (20 mM sodium phosphate buffer, 500 mM NaCl, 500 mM imidazole, pH 7.8). The eluted fractions were analyzed by SDS-PAGE to assess the purity, and the homogenous fractions were pooled.
Protein determination
Protein concentration was determined according to the bicinchoninic acid assay using bovine serum albumin as the standard (Smith et al. 1985).
Enzyme activity assays
Chitinase activity was assayed using 50 mM phosphate citrate buffer at pH 6.0 at 25 °C with 0.15 mM pNP-GlcNAc2 as substrate. The reaction was terminated by adding Na2CO3 (to 0.4 M) after 30 min of reaction, and the absorbance was then read at 410 nm in an Infinite 200 microplate reader (Tecan, Grödig, Austria). One unit of enzyme activity was defined as released 1 μmol of pNP per minute per milliliter at 25 °C, pH 6, in the assay. The optimal temperature for the enzymes was determined by assaying in 50 mM phosphate citrate buffer (pH 6) at different temperatures from 20 to 70 °C; analogously, the optimal pH for each enzyme was determined using 50 mM phosphate citrate buffer at different pH values ranging from 2.2 to 9 at 25 °C, in both cases using 0.15 mM pNP-GlcNAc2 as substrate, and the assay was run according to the method mentioned above. Thermal stability was determined by pre-incubation of the individual enzymes at different temperatures ranging from 30 to 55 °C for 5, 10, 15, and 20 min. The residual activity was then measured as the initial rate using 0.15 mM pNP-GlcNAc2 in 50 mM phosphate citrate buffer (pH 6) at 25 °C. Reactions were terminated by adding Na2CO3 (to 0.4 M), and the absorbance was read at 410 nm as described above. All reactions were run as true replicate runs (n = 2).
Substrate degradation
Colloidal chitin was used as the insoluble chitin substrate and prepared according to Rojas Avelizapa et al. (1999), briefly as follows: 10 g chitin was dissolved in 100 ml 85% (v/v) phosphoric acid, and the homogenized mixture was incubated overnight at 4 °C, then Milli-Q water was added in excess until visible chitin precipitation occurred; the mixture was then filtered through a cheesecloth, and the chitin precipitate was washed repeatedly (minimum 4 times) with Milli-Q water until acid removal was complete (neutral pH). The enzymatic hydrolysis of the colloidal chitin (1% (w/v)) was then performed in 50 mM sodium acetate buffer, pH 6, at 25 °C with 0.6 μM FbalChi18A or 0.9 μM MvarChi18A for 18 h. The reaction was stopped by heat inactivation, and the reaction mixture was applied to a Vivaspin 500 centrifugal concentrator with 5000 MWCO (Sartorius, Goettingen, Germany) for enzyme removal. The permeate was analyzed by mass spectrometry (see below).
One millimole of each chito-oligomer (chitobiose, chitotriose, chitotetraose, chitopentaose, and chitohexaose) was dissolved individually in 50 mM sodium acetate buffer, pH 6 at 25 °C, and reactions were initiated by addition of 0.02 μM FbalChi18A or MvarChi18A; during reaction, aliquots of 100 μl were taken at 5, 10, 15, 30, and 45 min and immediately heat inactivated for 10 min in 100 μl of Milli-Q water at 100 °C. After inactivation, the enzyme was removed by filtration using a Vivaspin 500 centrifugal concentrator with 5000 MWCO (Sartorius, Goettingen, Germany). The permeate was then further analyzed by mass spectrometry (see below).
Mass spectrometry analysis
Identification and quantification of chito-oligosaccharides and their degradation products were performed by liquid chromatography electrospray ionization mass spectrometry (LC-ESI-MS) on an amaZon SL ion trap (Bruker Daltonics, Bremen, Germany) coupled to an UltiMate 3000 UHPLC from Dionex (Sunnyvale, CA USA). Five-microliter samples were injected on a porous graphitized carbon column (Hypercarb PGC, 150 mm × 2.1 mm; 3 μm, Thermo Fisher Scientific, Waltham, MA, USA). The chromatography was performed at 0.4 ml min−1 at 70 °C on a two-eluent system with eluent A (0.1% formic acid in water) and eluent B (acetonitrile). The elution profile was as follows (time indicated in min): 0–1, 0% B; 1–28, linear gradient to 27.5% B; 28–31, linear gradient to 60% B; 31–35, isocratic 60% B; 35–40, isocratic 0% B. The electrospray was operated in positive mode with ultrascan mode and a scan range from 100 to 1500 m/z, smart parameter setting of 500 m/z, capillary voltage at 4.5 kV, end plate offset 0.5 kV, nebulizer pressure at 3.0 bar, dry gas flow at 12.0 l min−1, and dry gas temperature at 280 °C. Chitinase insertion domain fragmentation was performed using SmartFrag enhanced amplitude ramping from 80 to 120% and fragmentation time 20 ms. Quantification was performed based on manual MS/MS fragmentation of selected masses corresponding to chito-oligosaccharides DP 2-6 using the Compass QuantAnalysis 2.2 Program (Bruker Daltonics, Bremen, Germany).
3D structure modeling and substrate superposition
A 3D structure model for both identified enzymes was generated by the I-TASSER modeling tool (Yang et al. 2015) through homology modeling based on the 3D structure of SmaChiA (PDB code 1X6L) (Aronson et al. 2006). Structure comparisons of the active site amino acids were done by using PyMOL (PyMOL Molecular Graphics System, Delano Scientific, Portland, USA). Substrate superposition was based on 3D structure models for the catalytic domain for each of the identified enzymes generated by the ModWeb modeling web server (Pieper et al. 2011) through homology modeling based on the 3D structure of SmaChiA (PDB code 1EDQ; the 1X6L chitinase A is the W167A mutant of the (wild-type) 1EDQ chitinase A) (Papanikolau et al. 2003). For substrate superposition, the crystal structure (i.e., PDB code 1NH6; SmaChiA active site mutant E315L co-crystallized with chitohexaose) was aligned with the homology model of the catalytic domains of either FbalChi18A or MvarChi18A using YASARA 16.8.19 (YASARA Biosciences GmbH, Vienna, Austria). The model substrate was superposed by deleting the SmaChiA backbone structure and water molecules and joining the chitohexaose and the respective homology model into one object.
Results
Sequence analysis
Both FbalChi18A and MvarChi18A had a sequence similarity of 67 and 63%, respectively, with SmaChiA (Uniprot: P07254) (Perrakis et al. 1994). The neighbor-joining phylogenetic tree showed that FbalChi18A and MvarChi18A were close in distance to SmaChiA, the benchmark enzyme (indicated by black arrows, Fig. S1). SmaChiA was chosen as the benchmark enzyme as it has been well studied and characterized by others and is known to catalyze release of chitobiose from chitin (from the reducing end of chitin) (Horn et al. 2006a, b; Sikorski et al. 2006). Based on the similarity of amino acid sequences and their catalytic mechanism, both FbalChi18A and MvarChi18A are classified into family 18 of glycosyl hydrolases (GH18), the GH family harboring most chitinases identified so far (Lombard et al. 2014) (the more detailed sequence alignment analysis and structural prediction will be discussed further below).
Enzyme expression and purification
The two recombinant enzymes, FbalChi18A and MvarChi18A, were successfully expressed in E. coli strain BL21 (DE3) and purified (Fig. S2) (the enzymes were also expressed successfully in the TUNER (DE3) strain, data not shown). The yields of the purified FbalChi18A and MvarChi18A were approximately 275 and 860 mg l−1 of E. coli cell culture, respectively.
The two chitinases, FbalChi18A and MvarChi18A, are encoded by genes with length of 2610 and 2565 bp which correspond to 870 and 855 amino acids, respectively. The predicted molecular weight from the deduced amino acid sequence plus the His-tag of FbalChi18A and MvarChi18A were 93.8 and 92.5 kDa, respectively. The purified protein FbalChi18A and MvarChi18A correspondingly had a molecular weight of approx. 95 and 100 kDa using SDS-PAGE as shown in Fig. S2 (interpretation of the minor discrepancies between the predicted molecular weight and the molecular weight read off from the SDS-PAGE gels may require further analysis by amino acid sequencing).
Catalytic characterization of the two novel chitinases
Both FbalChi18A and MvarChi18A were found to have optimum temperature at 30 °C and optimum pH at 6 (Fig. 1). FbalChi18A was active in a broader temperature range (20–40 °C) and a broader pH range (pH 5–8) than MvarChi18A (20–30 °C; pH 6–7.5). Regarding thermal stability, FbalChi18A was found to be stable from room temperature up to 45 °C with a t1/2 of 75 min at 45 °C, but unstable at 55 °C with a t1/2 of 9 min (Fig. 2). MvarChi18A was stable from room temperature up to 35 °C with a t1/2 of 52 min, but unstable at 45 °C and above with a t1/2 of 1.5 min at 45 °C (Fig. 2). The specific activities for the purified FbalChi18A and MvarChi18A on the assay substrate pNP-GlcNAc2 were 0.128 and 0.056 U·(μmol enzyme−1), respectively (none of the enzymes had activity on pNP-GlcNAc, data not shown).
Thermal inactivation for both chitinases under different temperature ranging from 30 to 55 °C. Both enzymes, FbalChi18A (a) and MvarChi18A (b), were incubated for 5, 10, 15, and 20 min before the residual activity was determined. kd is the inactivation constant at the specific temperature as indicated
Substrate degradation of the two novel chitinases
The end products from the enzymatic hydrolysis of the chito-oligosaccharides by the two new chitinases were determined by mass spectrometry. From the trimer (GlcNAc3), tetramer (GlcNAc4), pentamer (GlcNAc5), and hexamer (GlcNAc6), the enzymes both catalyzed release of the dimer (GlcNAc2 = chitobiose) (Fig. 3a–d), but none of the two chitinases were able to cleave the GlcNAc2 dimer (data not shown). Analysis of colloidal chitin hydrolysis by both enzymes confirmed catalytic production of the dimer (GlcNAc2), besides indicating some trimer releasing activity, but at a 15- and 6-fold lower rate of the dimer release for FbalChi18A and MvarChi18A, respectively (Table 1). No products of higher chain length were detected. The final dimer/trimer ratios are different between the two enzymes as it is affected by the specific binding preferences of the individual enzymes (Horn et al. 2012).
Functional domains of the two novel chitinases
Most enzymes in the GH18 family exhibit a multidomain architecture with different distribution or arrangement of the domains. Based on sequence homology and tertiary structure prediction (via the web-based analysis tool InterPro program (http://www.ebi.ac.uk/interpro/) (Mitchell et al. 2015)), FbalChi18A and MvarChi18A were both predicted to each have four major domains: (1) the chitinase A N-terminal domain, (2) the catalytic β/α barrel domain (classified as glycoside hydrolase family 18; GH18), (3) a polycystic kidney disease (PKD) domain, and (4) a C-terminal chitin-binding domain (Table 2). For FbalChi18A, the sequence identity of the chitinase A N-terminal domain (amino acids 24–128) to that of SmaChiA was 62%, whereas it was only 50% for MvarChi18A (amino acids 19–153) (Table 2). The similarity to SmaChiA of the catalytic β/α barrel structure, indicated as β1–β8 in Fig. 4, was 69% for both enzymes (Table 2) (discussed further below). The PKD domain in each enzyme was found to consist of tandem repeats of about 85–90 amino acid residues with the two individual “repeat” stretches having 38% intra-identity for FbalChi18A and 49% identity for MvarChi18A (residues 569–659, 668–752 in FbalChi18A and residues 566–655, 662–747 in MvarChi18A, Table 2). Similarly, the C-terminal chitin-binding domain was also found to consist of tandem repeats, each repeat being ~44 residues, and having 43 and 44% identity among them for FbalChi18A and MvarChi18A, respectively.
Sequence alignment of GH18 chitinases from Ferrimonas balearica (FbalChi18A; ADN76700.1), Microbulbifer variabilis (MvarChi18A; WP_051089467.1), and Serratia marcescens (SmaChiA; AAA26551.1). The secondary structure of all the chitinases is indicated at the bottom of the sequence alignment by e (β-strand) and h (α-helix). Red boxes indicate the most conserved motifs, the SXGG and DXDXE motifs, which are the fully conserved motifs in the GH18 family of chitinases, while the asterisk symbol indicates the ten conserved residues in GH18 chitinases (color figure online)
Domain analysis
Sequence alignment comparison with SmaChiA with recognition of the α-helix and β-strand structures indicated that in both of the enzymes, FbalChi18A and MvarChi18A, the GH 18 catalytic β/α barrel domain consisted of a (β/α)8 TIM barrel fold with a small α+β domain insertion between β7 and β8 (Fig. 4). The catalytic domain of family 18 can be classified into three subfamilies A, B, and C based on the amino acid sequence similarity (Watanabe et al. 1993, 1994; Suzuki et al. 1999). The main structural differences between these three subfamilies is the existence of a small α+β domain insertion between β7 and β8 for subfamily A, as this insertion domain is absent in subfamilies B and C (Suzuki et al. 1999). This kind of insertion, which is also known as chitinase insertion domain, is usually composed of five or six anti-parallel β-strands and one α-helix (Li and Greene 2010). The occurrence of a chitinase insertion domain in the catalytic (β/α)8 barrel domain of chitinases has been proposed to increase the depth of the substrate-binding cleft (Perrakis et al. 1994; Suzuki et al. 1999). Hence, the chitinase insertion domain appears to play a role in substrate specificity by promoting the orientation and binding of the enzyme to longer substrates (Li and Greene 2010). It is proposed that both of the newly identified chitinases, FbalChi18A and MvarChi18A, are classified into subfamily A.
The two short conserved motifs of family 18 chitinases, SXGG and DXDXE, which are located at the third and fourth β-strands were observed in the amino acid sequence of both FbalChi18A and MvarChi18A (Fig. 4). The conserved glutamate residue, E313 in FbalChi18A and E312 in MvarChi18A (E315 in SmaChiA), in the DXDXE motif has been predicted to act as a catalytic residue where it facilitates the cleavage of the glycosidic bond via the protonation of the glucosidic oxygen (Vaaje-Kolstad et al. 2013).
The PKD domain which was observed in both FbalChi18A and MvarChi18A but not present in SmaChiA was first identified in human polycystin-1. The PKD domain is known to be common in hydrolases from marine bacteria and has been reported found in proteases, cellulases, and other chitinases in, e.g., Alteromonas sp. O-7 (Miyamoto et al. 2002), Stenotrophomonas maltophilia (Kobayashi et al. 2002), and Vibrio proteolyticus (Itoi et al. 2007). This PKD domain consists of a β-sandwich fold which is similar to Ig-like folds found in proteins and the fibronectin III (FnIII) superfamily (Bycroft et al. 1999). The PKD domain has been suggested to facilitate the binding of chitinase towards chitin (Orikoshi et al. 2005) besides may support the protein-protein interactions (Frederiksen et al. 2013). However, the importance of such functionality for bacterial chitinases has yet to be investigated.
The tandem “repeats” (Table 2) in the C-terminal regions of both chitinases were predicted as two putative chitin-binding domains encoding a CBM (See also Fig. 5). Further verification using the dbCAN annotation server (http://csbl.bmb.uga.edu/dbCAN/) (Yin et al. 2012) also confirmed the existence of two putative CBM5/12 in the C-terminal region of both chitinases. Alignment of these predicted C-terminal chitin-binding domains of FbalChi18A and MvarChi18A with those of other bacterial chitinases being in close proximity in the phylogenetic tree, and with the C-terminal chitin-binding domain sequence of SmaChiB, showed some homology, but also some key differences: (1) Whereas the sequences grouped together in the phylogenetic tree search (based on SmaChiA) contained some hallmark repeats of the signature sequence AKWWTQ or AKYWTQ (Fig. 6), it was evident that FbalChi18A was slightly different in this otherwise conserved region, but also that this segment differed from that of SmaChiB. Nevertheless, based on this analysis, we propose that the two newly identified chitinases FbalChi18A and MvarChi18A do contain a CBM 5/12 type chitin-binding domain in the C-terminus.
Comparison of predicted C-terminal CBM5/12 of two identified chitinases, FbalChi18A and MvarChi18A, with other closely related putative GH18 chitinases from the phylogenetic tree which have a repetitive CBM 5/12. The red box indicates the two aromatic residues (tyrosine and tryptophan) that are responsible for the interaction of the binding domain and chitin/carbohydrate molecules (color figure online)
It can be added that part of the CBM 5/12 sequence, i.e., the second part of the C-terminal repeated sequence of FbalChi18A (amino acids 824–864), showed sequence similarities to those of chitinases from Aeromonas sp. No. 10S-24 (41% identity) (Shiro et al. 1996), Clostridium paraputrificum (38% identity) (Morimoto et al. 1997), and Alteromonas sp. O-7 (33% identity) (Hiroshi et al. 1998) (data not shown). While for MvarChi18A, a similar analysis for the CBM 5/12-type chitin-binding domain (amino acids 809–849) showed sequence similarities of 51, 50, and 43% identity with Aeromonas sp. No. 10S-24, Cl. Paraputrificum, and Alteromonas sp. O-7, respectively (data not shown).
The highly conserved stWWst motif in the CBM 5/12-type domain is presumed to play a specific role in substrate binding. Binding of substrate to the enzyme via the CBM 5/12-type domain is thus believed to occur through hydrophobic interactions between the two exposed aromatic residues (WW) and the chitin substrate moieties (Brun et al. 1997; Suzuki et al. 1999). Chitinases from Aeromonas hydrophila, Bacillus circulans, and Pyrococcus kodakaraensis lacking a CBM have been shown to lose their binding capacity and activity towards insoluble chitin (Wu et al. 2001; Hashimoto et al. 2000; Tanaka et al. 1999). The CBM-type 5/12 is also considered as contributing to maintain the bound enzyme to the substrate during progressive reaction (Aronson et al. 2003).
Modeling of 3D structure
The 3D models for the catalytic domains of both FbalChi18A and MvarChi18A (Fig. 7a, b) appeared as being accurate as evaluated by Qualitative Model Energy ANalysis (QMEAN) (Benkert et al. 2011), Z-score analysis, and PROCHECK (Laskowski et al. 1993). The visual impression of the catalytic domain showed a characteristic barrel-type structure with a hollow center surrounded by β-sheets and with eight α-helices forming the barrel (Fig. 7a, b). The QMEAN Z-score for both FbalChi18A and MvarChi18A were −0.86 and −0.36, respectively (Fig. S3). The Ramachandran plot generated through the PROCHEK program which evaluated the stereochemical quality of the protein structures showed that for FbalChi18A, 92.3% of the residues were in the most favored region while 7.7% of the residues were in allowed regions (Fig. S4). While for MvarChi18A, 92.5% of the residues were in most favored regions while 7.5% of the residues in allowed regions (Fig. S5). Both of the evaluation, QMEAN Z-score and PROCHECK, indicated the favorability of these two models. Hence, this analysis showed that the active site cores of both enzymes agreed highly with the active site structure of SmaChiA and also that the active site regions of the two enzymes (as defined by the selected 10 amino acids) were conserved among the two new chitinases in terms of overlay (Fig. 7c, d).
Theoretical model of a 3D structure for the catalytic domain of both identified enzymes, a FbalChi18A b and MvarChi18A, which contain the TIM barrel (α/β)8 structure. An alpha-helix is indicated by red, beta-sheet by yellow, and loop by green color. An overlay of ten conserved active site residues for the c FbalChi18A d and MvarChi18A homology models (cyan sticks) with 1X6L chitinase from Serratia marcescens (green sticks) (color figure online)
Modeling of substrate binding
The possible binding mode for the hexamer substrate (GlcNAc6) to each of the enzymes FbalChi18A and MvarChi18A was predicted via superimposing the model of the catalytic domain of each enzyme onto the 3D crystal structure model of the catalytic domain of SmaChiA with its chitohexaose substrate (PDB code 1NH6). This superimposition modeling suggested that both FbalChi18A and MvarChi18A had a narrow substrate cleft that spanned across the whole side of the catalytic domain and that the cleft, as expected, was aligned with aromatic residues (Fig. 8). This type of substrate binding cleft is one of the characteristics for subfamily A in the catalytic domain of GH18 (Suzuki et al. 1999). Such substrate-binding clefts, fitting the substrate, are also observed for both SmaChiA and SmaChiB and are due to the presence of small so-called α+β chitinase insertion domains in the catalytic domain—appearing in FbalChi18A from residues 444–516 and in MvarChi18A from residues 442–514. The modeling of FbalChi18A with its substrate was more mischievous than the modeling of MvarChi18A, indicating that despite the 69% identity of both domains to the SmaChiA domain, the FbalChi18A and MvarChi18A sequences were not 100% identical (Fig. 4). The 3D visualization nevertheless corroborated that both enzymes indeed appeared to be exo-chitobiose hydrolases since the structure did indicate fitting of the straight substrate backbone into a “pocket” in the right hand side of the tunnel (as opposed to being a widely open groove in the structure) (Fig. 8b, d). A closer look at the superimposition models supported that the substrate was positioned by the aromatic tryptophan residues aligning the cleft, and this positioning corresponded to subsites −4 to +2 in both enzymes. The enlargement of the substrate interaction with the amino acids aligning the active site-binding cleft showed how the target bond in the substrate was positioned near the catalytic glutamate (E313 in FbalChi18A and E312 in MvarChi18A) to release the dimeric chitobiose product from the reducing end (subsite +1 and +2) of the hexamer (Fig. 8a, c). Thus, this analysis supported the interpretation that both enzymes indeed seem to be type A exo-chitobiose hydrolases, i.e., catalyzing release of chitobiose from the reducing end of chitin and chito-oligosaccharides.
Discussion
Enzyme characterization
By use of the genome database mining approach using the GH18 chitinase A from S. marcescens as template, we managed to identify two novel chitinases, FbalChi18A and MvarChi18A, which we propose to be type A exo-chitobiose hydrolases. The deduced amino acid sequence of FbalChi18A has about 62% homology with chitinase from Aeromonas molluscorum (Uniprot: R1GWE5), 54% with chitinase from Photobacterium angustum (Uniprot: A0A0D8QYT5), and 53% with chitinase from Enterobacter cloacae (Uniprot; G8LGW3). While for MvarChi18A, its amino acid sequence has about 58% homology with chitinase from Shewanella piezotolerans (Uniprot: B8CII9), 56% with chitinase from Aeromonas salmonicida (Uniprot: T0PPB8), and 51% with chitinase from E. cloacae (Uniprot: G8LGW3).
Both FbalChi18A and MvarChi18A had an optimum of activity at pH 6 and an optimum activity at 30 °C and were stable for up to 75 min at 45 °C and 52 min at 35 °C, respectively. The sequence and structural domain comparisons between the two enzymes did not provide any obvious clues pointing towards any structural reasons for their slight temperature optimum activity divergence. The thermal optimum characteristics make FbalChi18A and MvarChi18A mesophilic enzymes compared to the benchmark enzyme SmaChiA, which is more thermally robust and has an optimum activity between 50 and 60 °C and is stable for more than 400 h at 37 °C (Brurberg et al. 1996). Through domain predictions, FbalChi18A and MvarChi18A were both predicted to have four major domains each including the repetitive C-terminal CBM-type 5/12, and an N-terminal chitin-binding domain. The presence of a C-terminal CBM type 5/12 sets the two enzymes apart from SmaChiA which does not have a C-terminal CBM. To our knowledge, C-terminal CBM type 5/12 are specific for chitinases. However, several chitinases have been reported to have more than one chitin-binding domain in their structure, e.g., Chi92 from A. hydrophila (Wu et al. 2001), ChiC from Salinivibrio costicola (Aunpad and Panbangred 2003), and ChiA from P. kodakaraensis (Tanaka et al. 1999). Chi92 has three chitin-binding domains. In addition to two C-terminal chitin-binding domains, Chi92 also has an N-terminal chitin-binding domain (Chi92-N) that is similar to the ChiA N-terminal-binding domain of SmaChiA. Wu et al. (2001) reported that the truncated Chi92 which only contains a catalytic domain and the N-terminal Chi92 still display insoluble chitin-binding and hydrolytic activities. It was also described that the two C-terminal chitin-binding domains were functioning independently of each other (Wu et al. 2001). ChiA from P. kodakaraensis was also reported to have three chitin-binding domains with one residing at the N-terminus while the other two are residing at the C-terminus. Mutants with deletion of either N-terminal or C-terminal chitin-binding domain showed that each of these chitin-binding domains was independently functional in binding insoluble chitin (Tanaka et al. 1999). Both chitinases, FbalChi18A and MvarChi18A, contained the two short conserved motifs of family 18 chitinases: SXGG and DXDXE. A mutation study of SmaChiA by Papanikolau et al. (2001) has shown that residues Asp313 and Tyr390 together with Glu315 play a vital role in the enzyme catalysis. It was proposed that Asp313 which interacts with Asp311 will flip to an alternative position after protonation of the substrate glycosidic bond by Glu315. Since Asp313 also interacts with Glu315, it thus will force the distortion of the substrate acetamido group of the monomeric moiety in the −1 position (Papanikolau et al. 2001). As a consequence of these structural changes, the water molecule that is hydrogen-bonded to both the hydroxy group of Tyr390 and the NH of the acetamido group is displaced to a position that permits the completion of hydrolysis. Brameld and Goddard (1998) also reported that the hydrogen bonding of water molecules to Tyr390 helps to position the N-acetyl group prior to formation of an oxazoline ion. In contrast, the catalytic mechanism of SmaChiB (of Serratia marcescens) depends on the combination action of at least seven conserved active site residues: Tyr10, Ser93, Asp140, Asp 142, Glu144, Tyr 214, and Asp 215 and that Tyr10 and Ser93 are important for stabilization of the charge on Asp140 while Asp142 points towards Glu144 (Synstad et al. 2004). The partly buried Asp140 residue is essential for keeping the Asp142 protonated. Asp142 plays an interconnected role, and its rotation towards Glu144 contributes to a crucial distortion of the N-acetyl group of the −1 GlcNAc moiety. The phenolic hydroxyl of Tyr214 contributes to the positioning of the N-acetyl group of the −1 GlcNAc moiety. The Asp215 residue plays a role in the catalysis by being involved in the binding of the −1 GlcNAc in its distorted conformation. A direct comparison of the active site residue positions in SmaChiA versus SmaChiB (Fig. 4) shows some obvious similarities.
Substrate degradation
The enzymatic hydrolysis of the chito-oligosaccharides and colloidal chitin by each of the enzymes released mainly the dimeric product. The processivity of the family 18 chitinase may be assessed by studying the monomer/dimer ratio in product mixtures (Horn et al. 2006a, b). The product profiles thus indicated that both of them are exo-chitinases or exo-chitobiose hydrolases. The exo-chitinase-catalyzed hydrolysis of substrate may occur either at the reducing end or non-reducing end of the substrate. The C-terminal CBM 5/12 type in ChiB from Serratia marcescens helps locating the substrate-binding cleft so that the dimeric chitobiose products are released from subsites −1 and −2, being the non-reducing end of the substrate. In contrast, the N-terminus in SmaChiA (containing the FnIII-like structure that is likely involved in interactions with the chitin chain during catalysis) provides an active site topology that locates the substrate-binding cleft at the non-reducing end of the substrate, and hence, the dimeric products are thought to be released from subsites +1 and +2 (Perrakis et al. 1994; van Aalten et al. 2000; Vaaje-Kolstad et al. 2013).
Structure function analysis
The aromatic residues (notably tryptophan) in the active site, which were previously shown to be important for substrate binding in chitinases, are well conserved, and they interact with the GlcNAc units of the bound oligomer through stacking interactions (Uchiyama et al. 2001). Three tryptophan residues, Trp167, Trp275, and Trp539 (referred to SmaChiA), in the substrate-binding cleft were identified as the aromatic residues that most likely interact with GlcNAc units of the hexamer substrate. Trp273 (FbalChi18A) and Trp272 (MvarChi18A) (corresponding to Trp275 for SmaChiA) apparently interact with the substrate at subsite +1 while Trp537 (Fbalchi18A) and Trp537 (MvarChi18A) (corresponding to Trp539 for SmaChiA) interact with substrate at subsite −1. Trp165 (FbalChi18A) and Trp165 (MvarChi18A) (corresponding to Trp167 for SmaChiA) interact with the substrate at subsite −3. The significant roles of these aromatic residues towards the processive action of the enzymes have been reported by Aronson et al. (2003) and Zakariassen et al. (2009), who suggested that these aromatic residues are presumed to facilitate processivity by functioning as a flexible and hydrophobic patch, which the substrate chain can slide during the processive action. In accord with the high specific activity on the GlcNAc4, the products obtained from hydrolysis of GlcNAc5 and GlcNAc6 as substrates and the substrate binding analysis of the two new enzymes corroborated that FbalChi18A and MvarChi18A are most likely exo-chitobiose hydrolases that catalyze release of chitobiose from the reducing end of chitin and chito-oligosaccharides. The two new enzymes may find use in the production of chitobiose from chitin or merely for chitin degradation. Their use for production of chitobiose is highly relevant in an enzymatic cascade production of human milk oligosaccharides as the chitobiose product can act as a substrate for subsequent enzymatic synthesis of human milk oligosaccharide backbone structures with N-acetylglucosamine (Nyffenegger et al. 2015).
References
Anitha A, Sowmya S, Kumar PTS, Deepthi S, Chennazhi KP, Ehrlich H, Tsurkan M, Jayakumar R (2014) Chitin and chitosan in selected biomedical applications. Prog Polym Sci 39:1644–1667
Aronson NN, Halloran BA, Alexyev MF, Amable L, Madura JD, Pasupulati L, Worth C, Van Roey P (2003) Family 18 chitinase-oligosaccharide substrate interaction: subsite preference and anomer selectivity of Serratia marcescens chitinase A. Biochem J 376:87–95
Aronson NN, Halloran BA, Alexeyev MF, Zhou XE, Wang Y, Meehan EJ, Chen L (2006) Mutation of a conserved tryptophan in the chitin-binding cleft of Serratia marcescens chitinase A enhances transglycosylation. Biosci Biotechnol Biochem 70:243–251
Aunpad R, Panbangred W (2003) Cloning and characterization of the constitutively expressed chitinase C gene from a marine bacterium, Salinivibrio costicola strain 5SM-1. J Biosci Bioeng 96:529–536
Benkert P, Biasini M, Schwede T (2011) Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 27:343–350
Brameld KA, Goddard WA (1998) Substrate distortion to a boat conformation at subsite - 1 is critical in the mechanism of family 18 chitinases. J Am Chem Soc 120:3571–3580
Brun E, Moriaud F, Gans P, Blackledge MJ, Barras F, Marion D (1997) Solution structure of the cellulose-binding domain of the endoglucanase Z secreted by Erwinia chrysanthemi. Biochemistry 36:16074–16086
Brurberg MB, Eijsink VGH, Nes IF (1994) Characterization of a chitinase gene (chiA) from Serratia marcescens BJL200 and one-step purification of the gene product. FEMS Microbiol Lett 124:399–404
Brurberg MB, Nes IF, Eijsink VG (1996) Comparative studies of chitinases A and B from Serratia marcescens. Microbiology 142(7):1581–1589
Bycroft M, Bateman A, Clarke J, Hamill SJ, Sandford R, Thomas LR, Chothia C (1999) The structure of a PKD domain from polycystin-1: implications for polycystic kidney disease. EMBO J 18:297–305
Cantarel BI, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) The carbohydrate-active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res 37:233–238
Cohen-Kupiec R, Chet I (1998) The molecular biology of chitin digestion. Curr Opin Biotechnol 9:270–277
Elias JA, Homer RJ, Hamid Q, Chun GL (2005) Chitinases and chitinase-like proteins in TH2 inflammation and asthma. J Allergy Clin Immunol 116:497–500
Frederiksen RF, Paspaliari DK, Larsen T, Storgaard BG, Larsen MH, Ingmer H, Palcic MM, Leisner JJ (2013) Bacterial chitinases and chitin-binding proteins as virulence factors. Microbiol (United Kingdom) 159:833–847
Fuchs RL, McPherson SA, Drahos DJ (1986) Cloning of a Serratia marcescens gene encoding chitinase. Appl Environ Microbiol 51:504–509
Gouy M, Guindon S, Gascuel O (2010) SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27:221–224
Hashimoto M, Ikegami T, Seino S, Ohuchi N, Fukada H, Sugiyama J, Shirakawa M, Watanabe T (2000) Expression and characterization of the chitin-binding domain of chitinase A1 from Bacillus circulans WL-12. J Bacteriol 182:3045–3054
Hiroshi T, Hideyuki O, Kayoko S, Miyuki H, Junko U, Katsushiro M, Chiaki I, Yoshiro O, Yoshihiko I (1998) Characterization of chitinase C from a marine bacterium, Alteromonas sp. strain O-7, and its corresponding gene and domain structure. Appl Environ Microbiol 64:472–478
Horn SJ, Sorbotten A, Synstad B, Sikorski P, Sorlie M, Varum KM, Eijsink VGH (2006a) Endo/exo mechanism and processivity of family 18 chitinases produced by Serratia marcescens. FEBS J 273:491–503
Horn SJ, Sørlie M, Vaaje-Kolstad G, Norberg AL, Synstad B, Vårum KM, Eijsink VGH (2006b) Comparative studies of chitinases A, B and C from Serratia marcescens. Biocatal Biotransformation 24:39–53
Horn SJ, Sorlie M, Vårum KM, Väljamäe P, Eijsink VGH (2012) Measuring processivity. Methods Enzymol 510:69–95
Hult E-L, Katouno F, Uchiyama T, Watanabe T, Sugiyama J (2005) Molecular directionality in crystalline beta-chitin: hydrolysis by chitinases A and B from Serratia marcescens 2170. Biochem J 388:851–856
Itoi S, Kanomata Y, Koyama Y, Kadokura K, Uchida S, Nishio T, Oku T, Sugita H (2007) Identification of a novel endochitinase from a marine bacterium Vibrio proteolyticus strain no. 442. Biochim Biophys Acta - Proteins Proteomics 1774:1099–1107
Kobayashi DY, Reedy RM, Bick J, Oudemans PV (2002) Characterization of a chitinase gene from Stenotrophomonas maltophilia strain 34S1 and its involvement in biological control. Appl Environ Microbiol 68:1047–1054
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291
Li H, Greene LH (2010) Sequence and structural analysis of the chitinase insertion domain reveals two conserved motifs involved in chitin-binding. PLoS One 5:e8654
Lombard V, Golaconda Ramulu H, Drula E, Coutinho PM, Henrissat B (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42:490–495
Mitchell A, Chang HY, Daugherty L, Fraser M, Hunter S, Lopez R, McAnulla C, McMenamin C, Nuka G, Pesseat S, Sangrador-Vegas A, Scheremetjew M, Rato C, Yong SY, Bateman A, Punta M, Attwood TK, Sigrist CJA, Redaschi N, Rivoire C, Xenarios I, Kahn D, Guyot D, Bork P, Letunic I, Gough J, Oates M, Haft D, Huang H, Natale DA, Wu CH, Orengo C, Sillitoe I, Mi H, Thomas PD, Finn RD (2015) The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res 43:D213–D221
Miyamoto K, Nukui E, Itoh H, Sato T, Kobayashi T, Imada C, Watanabe E, Inamori Y, Tsujibo H (2002) Molecular analysis of the gene encoding a novel chitin-binding protease from Alteromonas sp. strain O-7 and its role in the chitinolytic system. J Bacteriol 184:1865–1872
Morimoto K, Karita S, Kimura T, Sakka K, Ohmiya K (1997) Cloning, sequencing, and expression of the gene encoding Clostridium paraputrificum chitinase ChiB and analysis of the functions of novel cadherin-like domains and a chitin-binding domain. J Bacteriol 179:7306–7314
Nyffenegger C, Nordvang RT, Zeuner B, Łężyk M, Difilippo E, Logtenberg MJ, Schols H A, Meyer AS, Mikkelsen JD (2015) Backbone structures in human milk oligosaccharides: trans-glycosylation by metagenomic β-N-acetylhexosaminidases. Appl Microbiol Biotechnol 19:7997–8009
Ohno T, Armand S, Hata T, Nikaidou N, Henrissat B, Mitsutomi M, Watanabe T (1996) A modular family 19 chitinase found in the prokaryotic organism Streptomyces griseus HUT 6037. J Bacteriol 178:5065–5070
Orikoshi H, Nakayama S, Hanato C, Miyamoto K, Tsujibo H (2005) Role of the N-terminal polycystic kidney disease domain in chitin degradation by chitinase A from a marine bacterium, Alteromonas sp. strain O-7. J Appl Microbiol 99:551–557
Papanikolau Y, Prag G, Tavlas G, Vorgias CE, Oppenheim AB, Petratos K (2001) High resolution structural analyses of mutant chitinase A complexes with substrates provide new insight into the mechanism of catalysis. Biochemistry 40:11338–11343
Papanikolau Y, Tavlas G, Vorgias CE, Petratos K (2003) De novo purification scheme and crystallization conditions yield high-resolution structures of chitinase A and its complex with the inhibitor allosamidin. Acta Crystallogr - Sect D Biol Crystallogr 59:400–403
Perrakis A, Tews I, Dauter Z, Oppenheim AB, Chet I, Wilson KS, Vorgias CE (1994) Crystal structure of a bacterial chitinase at 2.3 A resolution. Structure 2:1169–1180
Pieper U, Webb BM, Barkan DT, Schneidman-Duhovny D, Schlessinger A, Braberg H, Yang Z, Meng EC, Pettersen EF, Huang CC, Datta RS, Sampathkumar P, Madhusudhan MS, Sjölander K, Ferrin TE, Burley SK, Sali A (2011) ModBase,a database of annotated comparative protein structure models,and associated resources. Nucleic Acids Res 39:465–474
Rinaudo M (2006) Chitin and chitosan: properties and applications. Prog Polym Sci 31:603–632
Rojas Avelizapa L, Cruz-Camarillo R, Guerrero M, Rodríguez R, Ibarra JE (1999) Selection and characterization of a proteo- chitinolytic strain of Bacillus thuringiensis, able to grow in shrimp waste media. World J Microbiol Biotechnol 15:299–308
Shen K-T, Chen M-H, Chan H-Y, Jeng J-H, Wang Y-J (2009) Inhibitory effects of chitooligosaccharides on tumor growth and metastasis. Food Chem Toxicol 47:1864–1871
Shiro M, Ueda M, Kawaguchi T, Arai M (1996) Cloning of a cluster of chitinase genes from Aeromonas sp. no. 10S-24. Biochim Biophys Acta - Gene Struct Expr 1305:44–48. doi:10.1016/0167-4781(95)00213-8
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Sikorski P, Sørbotten A, Horn SJ, Eijsink VGH, Vårum KM (2006) Serratia marcescens chitinases with tunnel-shaped substrate-binding grooves show endo activity and different degrees of processivity during enzymatic hydrolysis of chitosan. Biochemistry 45:9566–9574
Smith PK, Krohn RI, Hermanson GT, Mallia AK, Gartner FH, Provenzano MD, Fujimoto EK, Goeke NM, Olson BJ, Klenk DC (1985) Measurement of protein using bicinchoninic acid. Anal Biochem 150:76–85
Suzuki K, Suzuki M, Taiyoji M, Nikaidou N, Watanabe T (1998) Chitin binding protein (CBP21) in the culture supernatant of Serratia marcescens 2170. Biosci Biotechnol Biochem 62:128–135
Suzuki K, Taiyoji M, Sugawara N, Nikaidou N, Henrissat B, Watanabe T (1999) The third chitinase gene (chiC) of Serratia marcescens 2170 and the relationship of its product to other bacterial chitinases. Biochem J 343:587–596
Synowiecki J, Al-Khateeb NA (2003) Production, properties, and some new applications of chitin and its derivatives. Crit Rev Food Sci Nutr 43:145–171
Synstad B, Gaseidnes S, van Aalten DMF, Vriend G, Nielsen JE, Eijsink VGH (2004) Mutational and computational analysis of the role of conserved residues in the active site of a family 18 chitinase. Eur J Biochem 271:253–262
Synstad B, Vaaje-Kolstad G, Cederkvist FH, Saua SF, Horn SJ, Eijsink VGH, Sørlie M (2008) Expression and characterization of endochitinase C from Serratia marcescens BJL200 and its purification by a one-step general chitinase purification method. Biosci Biotechnol Biochem 72:715–723
Tanaka T, Fujiwara S, Nishikori S, Fukui T, Takagi M, Imanaka T (1999) A unique chitinase with dual active sites and triple substrate binding sites from the hyperthermophilic archaeon Pyrococcus kodakaraensis KOD1. Appl Environ Microbiol 65:5338–5344
Uchiyama T, Katouno F, Nikaidou N, Nonaka T, Sugiyama J, Watanabe T (2001) Roles of the exposed aromatic residues in crystalline chitin hydrolysis by chitinase a from Serratia marcescens 2170. J Biol Chem 276:41343–41349
Vaaje-Kolstad G, Horn SJ, van Aalten DMF, Synstad B, Eijsink VGH (2005) The non-catalytic chitin-binding protein CBP21 from Serratia marcescens is essential for chitin degradation. J Biol Chem 280:28492–28497
Vaaje-Kolstad G, Horn SJ, Sørlie M, Eijsink VGH (2013) The chitinolytic machinery of Serratia marcescens—a model system for enzymatic degradation of recalcitrant polysaccharides. FEBS J 280:3028–3049
van Aalten DM, Synstad B, Brurberg MB, Hough E, Riise BW, Eijsink VG, Wierenga RK (2000) Structure of a two-domain chitotriosidase from Serratia marcescens at 1.9-A resolution. Proc Natl Acad Sci U S A 97:5842–5847
van Aalten DM, Komander D, Synstad B, Gåseidnes S, Peter MG, Eijsink VG (2001) Structural insights into the catalytic mechanism of a family 18 exo-chitinase. Proc Natl Acad Sci U S A 98:8979–8984
Watanabe T, Kobori K, Miyashita K, Fujii T, Sakai H, Uchida M, Tanaka H (1993) Identification of glutamic acid 204 and aspartic acid 200 in chitinase A1 of Bacillus circulans WL-12 as essential residues for chitinase activity. J Biol Chem 268:18567–18572
Watanabe T, Ito Y, Yamada T, Hashimoto M, Sekine S, Tanaka H (1994) The roles of the C-terminal domain and type III domains of chitinase A1 from Bacillus circulans WL-12 in chitin degradation. J Bacteriol 176:4465–4472
Wu ML, Chuang YC, Chen JP, Chen CS, Chang MC (2001) Identification and characterization of the three chitin-binding domains within the multidomain chitinase Chi92 from Aeromonas hydrophila JP101. Appl Environ Microbiol 67:5100–5106
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER suite: protein structure and function prediction. Nat Meth 12:7–8
Yin Y, Mao X, Yang J, Chen X, Mao F, Xu Y (2012) DbCAN: a web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 40:445–451. doi:10.1093/nar/gks479
Zakariassen H, Aam BB, Hom SJ, Vårum KM, Sørlie M, Eijsink VGH (2009) Aromatic residues in the catalytic center of chitinase a from Serratia marcescens affect processivity, enzyme activity, and biomass converting efficiency. J Biol Chem 284:10610–10617
Acknowledgements
This study was funded by the Malaysian Ministry of Education (MoE) and University Malaysia Pahang, Malaysia, and the Center for BioProcess Engineering, Department of Chemical and Biochemical Engineering, Technical University of Denmark, Denmark.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(PDF 677 kb)
Rights and permissions
About this article
Cite this article
Jamek, S.B., Nyffenegger, C., Muschiol, J. et al. Characterization of two novel bacterial type A exo-chitobiose hydrolases having C-terminal 5/12-type carbohydrate-binding modules. Appl Microbiol Biotechnol 101, 4533–4546 (2017). https://doi.org/10.1007/s00253-017-8198-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8198-4