Abstract
Pig manures are frequently used as fertilizer or co-substrate in biogas plants (BGPs) and typically contain antibiotic residues (ARs), as well as bacteria carrying resistance genes (RGs) and mobile genetic elements (MGEs). A survey of manures from eight pig fattening and six pig breeding farms and digestates from eight BGPs in Lower Saxony, Germany was conducted to evaluate the link between antibiotic usage and ARs to RGs and MGEs present in organic fertilizers. In total, 11 different antibiotics belonging to six substance classes were applied in the farms investigated. Residue analysis revealed concentrations of tetracycline up to 300 mg kg−1 dry weight (DW) in manures and of doxycycline up to 10.1 mg kg−1 DW in digestates indicating incomplete removal during anaerobic digestion. RGs (sul1, sul2, tet(A), tet(M), tet(X), qacE∆1) were detected in total community DNA of all samples by PCR-Southern blot hybridization. Broad-host range plasmids (IncP-1, IncQ, IncN, and IncW) and integron integrase genes (intI1, intI2) were found in most manure samples with IncN and IncW plasmids being more abundant in manure from pig breeding compared to pig fattening farms. IntI1, IncQ, and IncW plasmids were also detected in all digestates, while IncP-1, IncN, and LowGC plasmids were detected only sporadically. Our findings strongly reinforce the need for further research to identify mitigation strategies to reduce the level of contamination of organic fertilizers with ARs and transferable RGs that are applied to soil and that might influence the mobile resistome of the plant microbiome.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Conventional livestock husbandry is due to increasing numbers of animals kept in a constantly decreasing number of farms (Udo et al. 2011) inevitably linked to the application of antibiotics (Boxall et al. 2004; Sarmah et al. 2006; Widyasari-Mehta et al. 2016a). An estimated total of 1619 tons of antibiotics was used in Germany in 2012 (Bundesamt für Verbraucherschutz und Lebensmittelsicherheit (BVL) Germany 2015). Among the most frequently applied antibiotic classes are the first generation antibiotics of tetracyclines and sulfonamides. Depending on the target animals, antibiotic application patterns and frequencies typically differ. The fate of the antibiotics, however, mainly depends on their physicochemical characteristics. Thus, sulfonamides and tetracyclines are partially excreted as parent compounds and/or metabolites (up to 96 and 90 %, respectively) (Hirsch et al. 1999; Sarmah et al. 2006; Lamshöft et al. 2007; Heuer et al. 2008). Recent studies showed that antibiotic residues (ARs) in pig manures reached up to 770 mg oxytetracycline kg−1 dry weight (DW) concentrations (Martínez-Carballo et al. 2007; Gans et al. 2010; Ratsak et al. 2013; Spielmeyer et al. 2014, 2015; Widyasari-Mehta et al. 2016a). Furthermore, it was revealed by Heuer et al. (2008) that, due to deacetylation of one major metabolite, the concentration of 14C-labeled sulfadiazine even increased during the storage of manure from 14C-labeled sulfadiazine treated pigs over time. Besides the direct application of manure as fertilizer on agricultural soil, its usage as co-substrate in biogas plants (BGPs) is of increasing importance. Digestates resulting from the biogas production are also applied as organic fertilizers. ARs were recently reported from digestates as well, e.g. sulfadimidine up to 76.2 mg kg−1 DW (Ratsak et al. 2013; Spielmeyer et al. 2014, 2015; Widyasari-Mehta et al. 2016a), suggesting that their application to soil may also contribute to selection of resistant bacteria in digestate treated soils.
Livestock manure does not only frequently contain ARs but it also represents a reservoir of diverse resistance genes (RGs), integrons, and mobile genetic elements (MGEs) such as transferable plasmids (Binh et al. 2008; Heuer et al. 2012; Zhu et al. 2013; Marti et al. 2013; Kyselková et al. 2015). Also, several RGs and integrons were detected in samples from biogas digesters (Diehl and LaPara 2010; Wolters et al. 2016). When manures or digestates are applied as fertilizers to field soil, these RGs and MGEs might be transferred to the indigenous bacterial soil communities (Jechalke et al. 2014). For instance, the repeated application of pig manure spiked with different concentrations of sulfadiazine was shown under microcosm conditions to enhance the abundance of sulfonamide RGs (sul genes) up to 2 months after their entry into soils (Heuer et al. 2011). Several transferable plasmids conferring resistances toward antibiotics that are typically associated with livestock manure have a broad host range (BHR) (Götz et al. 1996; Binh et al. 2008; Heuer et al. 2012) and were shown to transfer to bacteria indigenous to the rhizosphere of plants (Musovic et al. 2006) or to Pseudomonas putida when introduced with manure into field soil (Götz and Smalla 1997). Recently, BHR plasmids belonging to the IncP-1ε plasmid subgroup were captured (based on sulfadiazine or tetracycline resistances conferred) from BGP digestate bacteria into Escherichia coli and P. putida recipients (Wolters et al. 2015). Thus, following the application of manure or digestate as fertilizer, these plasmids may potentially also be taken up by bacteria associated with crops and may subsequently be transferred to consumers. As the intestine is considered to be a hot spot for horizontal gene transfer (HGT) (Kurokowa et al. 2007; Baquero 2012), in a worst case scenario, manure fertilization might contribute via this route to the increasing hazard posed by multiple antibiotic resistant bacteria in particular pathogens of clinical relevance (Forsberg et al. 2012). The combined introduction of RGs, MGEs, and ARs into agricultural soil via manure spread might in this context be of particular relevance, as antibiotics present in soil at subinhibitory concentrations might foster HGT processes and the proliferation of antibiotic-resistant bacteria (Gullberg et al. 2011, 2014; Jechalke et al. 2014).
Therefore, the aims of the current study were to link data on the application of antibiotics in 19 pig fattening and breeding farms without/with farm-owned BGPs in Lower Saxony, Germany to ARs found via liquid chromatography tandem mass spectrometry (LC-MS/MS) analysis and to the presence of RGs, integrons, and MGEs in manures and digestates of BGPs. In the present study, we have used PCR-Southern blot hybridization for highly sensitive and specific detection of RGs and MGEs in total community (TC-) DNA. Additionally, we evaluated whether RGs, integrons, and MGEs present in manures were also detectable in digestates of BGPs fed with manures as co-substrate.
Materials and methods
Sampling, farm operating data, and antibiotic applications
In order to investigate pig manures and digestates from farms located in Lower Saxony, Germany, in autumn 2012, samples from eight pig fattening and six breeding farms, as well as digestates from eight BGPs (five pig breeding farms with farm-owned BGPs and three corporate BGPs without the corresponding antibiotic application data) fed with manure as co-substrate, were collected. The details of samplings were described by Widyasari-Mehta et al. (2016a). Manures were sampled from cellars, silos, or lagoons using a probe sampler, a bypass sampler or in backflush mode from the vacuum tanker. Digestate samples were taken from closed or open silos via the outlet valves. At each farm, four 8-L samples were taken. Sample aliquots of 300 mL were transferred into polyethylene bottles and kept in cooling boxes during transport to the laboratory. In addition, farm operating data and information on antibiotic applications from spring 2012 until autumn 2012 (Table 1) were gathered. A complete data set of antibiotic applications and antibiotic residues for samples from spring 2012 to 2013 is detailed in Widyasari-Mehta et al. (2016a).
Residue analysis
The manure and digestate samples were analyzed for 16 antibiotics and three metabolites, namely, oxytetracycline, tetracycline, chlortetracycline, doxycycline, sulfadiazine, acetyl-sulfadiazine, sulfadimidine, acetyl-sulfadimidine, sulfamerazine, sulfadoxine, sulfadimethoxine, sulfamethoxypyridazine, trimethoprim, enrofloxacin, ciprofloxacin, danofloxacin, marbofloxacin, tiamulin, and tylosin. The analytical procedure is detailed by Widyasari-Mehta et al. (2016a, b). All samples were characterized for pH and DW content. Thereafter, aliquots of 25-g fresh samples were treated with ethylenediaminetetraacetic acid solution, citrate buffer, and hydrochloric acid to adjust pH 3.0. After lyophilization, ARs in the solids were extracted using methanol/ethyl acetate mixture. For clean-up, liquid-liquid partition with n-hexane/water followed by solid phase extraction was conducted. Target compound analysis was finally performed using LC-MS/MS with electrospray ionization in multiple reaction monitoring mode. Single-point standard addition technique was applied for quantification purpose. Method quantitation limits (MQLs) of all target compounds were 0.2 mg kg−1 DW in manure and digestate samples.
Bioanalytics
Extraction of TC-DNA
TC-DNA was extracted from manure samples and BGP digestates using the spin kit for soil (MP Biomedicals, Heidelberg, Germany) according to the manufacturer’s recommendation. Freshly homogenized samples (14 mL) were centrifuged in 15-mL Falcon tubes (Sarstedt, Nümbrecht, Germany) at 3100×g for 10 min; the supernatants were discarded and pellets were homogenized with a sterile spatula. Homogenized sample material (0.1 g) was transferred into lysis tubes and used for extraction of TC-DNA. Prior to PCRs, TC-DNA was diluted 1:5 with 1× TE buffer (pH 8.0).
Detection of sequences specific for antibiotic resistance genes, quaternary ammonium compound resistance genes, integron integrase genes, and plasmids
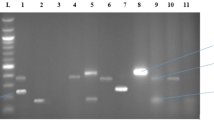
PCR assays used for the detection of RGs, integrons, and plasmid-specific sequences of IncN, IncP-1, IncQ, IncW, and LowGC plasmids were previously described and are listed in Table 2. To increase the sensitivity and specificity of the detection method, PCR products were Southern blotted and hybridized with the corresponding digoxigenin-labeled probes. The generation of the probes and the Southern blot hybridization were done as described in detail by Götz et al. (1996) and Binh et al. (2008). PCR amplicons obtained from reference plasmids (Table 2) were labeled with digoxigenin using the DIG-labeling Kit (Roche Diagnostic, Mannheim, Germany).
Results
Antibiotic application patterns and antibiotic residues in manures and digestates
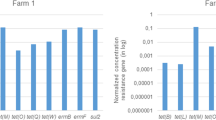
Antibiotics from tetracycline, sulfonamide, diaminopyrimidine, pleuromutiline, and macrolide classes were used on 19 pig husbandry farms with conventional manure management (Table 1). Seven antibiotics from these classes were applied at the eight pig fattening farms F1 to F8. The highest usage rates were found for tetracycline at farm F1 keeping 1000 fattening pigs, where 140–208 pigs were treated eight times. Here, tetracycline was consistently found in manure at 300 mg tetracycline kg−1 DW. Tetracyclines were also used to treat up to 385 pigs at five other pig fattening farms (except for F2 and F8). At these farms, tetracycline was detected in manures, ranging from 5.5 to 287 mg kg−1 DW. Doxycycline and chlortetracycline were found only in single manure samples from farms F2 and F7, respectively. These findings correlated with the data on the rate of antibiotic usage. A single application of the sulfonamide sulfadiazine in combination with trimethoprim was only recorded for farms F6 and F7. At farm F6 where 220 pigs were treated with sulfadiazine/trimethoprim, 0.7 mg sulfadiazine residues were detected per kilogram DW of manure. In addition, acetyl-sulfadiazine was simultaneously detected at 11.5 mg kg−1 DW. Even though tylosin was frequently applied to 20–355 pigs at farm F1, it could not be detected in the corresponding manure samples.
Broader application patterns were observed at six pig breeding farms (B1 to B6), where up to 11 antibiotics were administered. On those farms, sulfonamides in combination with trimethoprim were most frequently applied, nine times to 180 pigs, while tetracycline was applied only once to 400 pigs at B5. Alternatively, doxycycline was delivered to four pig breeding farms, while chlortetracycline and oxytetracycline were less frequently administered. In manure from pig breeding farms, doxycycline was most frequently detected at concentrations from 9.4 to 101 mg kg−1 DW of manure. The detection of doxycycline up to 19.0 mg kg−1 DW in both cellar and silo manures of farm B2, however, did not reflect doxycycline usage. The highest concentration of oxytetracycline with 211 mg kg−1 DW manure was detected at farm B3. Moreover, the application of sulfadimidine to a larger animal group, up to 180 pigs, resulted in sulfadimidine residues of 23.0 mg kg−1 DW manure of B6. Here, the corresponding acetyl-sulfadimidine was not detected above MQL. Enrofloxacin was applied at four pig breeding farms but was detected only in manure from farm B5 which reported the highest application frequency (4 times/4–50 pigs) at 1.3 mg kg−1 DW. In contrast, 1.4 mg tiamulin kg−1 DW was found in the manure from farm B5 after the single application to 50 pigs.
On pig breeding farms with farm-owned BGPs (BGP1 to BGP5), frequent applications of doxycycline (21 times) were recorded to treat up to 400 pigs at BGP4. On those farms, residues of doxycycline up to 10.1 mg kg−1 DW digestate were detected. More doxycycline was applied to larger animal groups at BGP2, where up to 2000 pigs were treated and 7.4 mg doxycycline kg−1 DW of digestate could be detected. Particularly for enrofloxacin, up to seven applications were administered at BGP2 and BGP4; however, there was only one positive digestate sample (BGP4 at 0.2 mg kg−1 DW digestate). In addition, digestate samples from BGP6 to BGP8, fed with input materials from different farms, were also analyzed. At BGP6, tetracycline and doxycycline were detected in the digestate at 6.4 and 2.2 mg kg−1 DW, respectively. At BGP7, 0.4 mg tetracycline kg−1 DW digestate was found. The digestate from BGP8 was free of residues of all 19 antibiotics analyzed.
Antibiotic RGs and disinfectant RGs in manures derived from different pig producing systems and digestates of BGPs
A very similar occurrence of RGs was detected via PCR/Southern blot hybridization for manures originating from different pig production systems and for digestates of BGPs fed with manures as co-substrate (Tables 3 and 4). Except for the sulfonamide resistance gene sul3, all antibiotic RGs investigated in this study were detected in all samples analyzed. Thus, genes conferring all three known mechanisms conferring resistance toward tetracycline (efflux, ribosomal protection, enzymatic modification) were detected in manures and in BGP digestates. In most cases, signal intensities were similar for manures from pig breeding and fattening farms. However, more intense signals for tet(M) were found in manures from pig fattening farms than in manures from breeding farms. While a high abundance of the sulfonamide resistance genes sul1 and sul2 was indicated by strong hybridization signal intensities for all samples, sul3 was exclusively detected (with considerably less intense signals) in samples from BGP3.
The complete disinfectant RG qacE was not detected in any of the manure samples derived from pig fattening or in digestates, but it was found in one manure sample from the pig breeding B2 (Table 3). In contrast, the functional fragment of this gene, qacE∆1, was present in all manure samples and digestates, and strong hybridization signals of Southern blotted amplicons indicated a high abundance of qacE∆1.
Detection of sequences specific for transferable plasmids and integron integrases in manures derived from different pig production systems and BGP digestates
Manure
A higher diversity of transferable plasmids was revealed in TC-DNA of manures from pig breeding facilities than from pig fattening farms (Table 3). All manure samples contained both IncP-1 and IncQ plasmids, and hybridization signal intensities were comparable except for considerably weaker hybridization signals obtained from manure samples derived from pig fattening farms F5 and F7. Furthermore, plasmids of the incompatibility group IncW were detected in TC-DNA from eight of nine manures from pig breeding farms and three of eight manures from pig fattening farms. IncN plasmids were detected as well but exclusively in manures derived from pig breeding farms (Tables 3 and 4), whereas LowGC plasmids were detected only in one manure sample (B1) originating from a pig breeding facility. Class 1 and 2 integron integrase genes were detected in TC-DNA from all manure samples investigated with very strong hybridization signals indicating a high abundance.
BGP digestates
PCR-Southern blot hybridization indicated that IncW and IncQ plasmids were abundant in all digestate samples taken from BGPs. The trfA-based detection system for IncP-1 plasmids revealed that plasmids belonging to the IncP-1 group were detectable in TC-DNA from digestates of all BGPs except BGP7. A more diverse range of plasmids was observed in biogas digestate samples compared to farm manures due to the frequent detection of LowGC plasmids BGP digestates (Table 4). Plasmids belonging to the incompatibility group IncN were only detected in one BGP digestate with very weak signal intensity. Furthermore, integrase genes of class 1 and 2 integrons were also detected in TC-DNA from BGPs samples. The hybridization signal intensity indicated a similar abundance in the TC-DNA from different BGPs.
Discussion
The current study aimed to link data on antibiotic usage from a survey conducted at 19 pig farms without/with farm-owned BGPs in Lower Saxony, Germany to the occurrence of ARs, RGs, and plasmid- and integron-specific sequences in liquid manures and digestates of BGPs feeding manure as co-substrate. The survey provides unique data on the extent of antibiotic usage in conventional pig farms and reveals that a more diverse range of antibiotics is applied in pig breeding farms and in farms with BGPs.
The AR concentrations measured were higher in manures compared to digestates. The results often showed a clear dependency of the AR detected on antibiotic application data. The frequent detection of different tetracycline residues at rather high concentrations may be explained by the higher frequency of tetracycline application at the different farms and the concomitantly high excretion rates of those production animals (up to 90 %) (Table 1). However, the detected residues did not always reflect the application rates. For example, low concentrations of tetracyclines were detected in manure samples from farm B2 where no tetracyclines were used in the time period surveyed, and thus might indicate background contaminations with tetracyclines from previous applications. Although sulfonamide antibiotics were applied at 7 out of 19 farms, they were less often detected in the respective manure samples. In addition to the rather lower recovery rates of 46 ± 10 % in manures indicating a rapid formation of non-extractable residues (Kreuzig and Höltge 2005), sampling time and the application frequency might have contributed to the detection of sulfonamide residues in manures. Furthermore, sulfadiazine may be transformed into the phase II metabolite acetyl-sulfadiazine in the digestive systems of animals (Lamshöft et al. 2007) and then may be excreted with the remaining parent compound. High recovery rates of 90 ± 10 % for acetyl-metabolites in manures may also be the reason why acetyl-sulfadiazine was found at higher concentration, which is 11.5 mg kg−1 DW of manure. Although trimethoprim was applied as a synergist of sulfonamides, it was not detected in any sample, which might be explained by rapid biodegradation as already reported by Haller et al. (2002), Mohring et al. (2009), and Spielmeyer et al. (2015). Similar to the sulfonamides, enrofloxacin residues were detected only once. Single animal treatments and high dilution factors may be one reason contributing to the rare detection of enrofloxacin. Another factor might be the strong sorption of enrofloxacin to the biosolids of manure (Tolls 2001; Uslu et al. 2008). Tiamulin, however, was reported to remain unchanged during a 180-day degradation experiment (Schlüsener et al. 2006). This might explain the finding that tiamulin was detected in manure of B5, even though tiamulin was applied only once to a group of 50 pigs. Moreover, due to rapid degradation and strong sorption to biosolids (Kolz et al. 2005), tylosin residues could not be detected in any samples even though it was frequently applied.
In comparison to manure, the concentrations of ARs detected in digestates from the mesophilic BGPs under study were clearly lower, ranging from 0.2 to 10.1 mg kg−1 DW (Table 1). The current results supported the survey studies conducted in the BGPs as well as laboratory testing which showed that ARs are only partially removed during anaerobic digestion (Mohring et al. 2009; Ratsak et al. 2013; Spielmeyer et al. 2014, 2015). Corresponding to the frequent applications of enrofloxacin in the farm with the BGP4, enrofloxacin was also detected in one digestate sample from BGP4. Tetracycline residues were also detected at low mg kg−1 DW concentrations in digestates. The highest concentrations of doxycycline were observed in BGP4, where 21 applications of doxycycline to bigger groups of pigs were recorded. At BGP7 fed with cattle manures, only a trace of tetracycline antibiotic was detected. This is consistent to the regular practice at the dairy farm where limited amounts of antibiotics are applied (Spielmeyer et al. 2015). Various elimination rates of tetracyclines in the BGP feeding manures as co-substrate were reported ranging from 14 to 89 % (Arikan et al. 2006; Arikan 2008; Spielmeyer et al. 2015). Besides biodegradation, strong sorption to biosolids and dilution factors from other co-substrates of the BGP, such as maize silage up to 65 %, might have contributed to the lower concentrations of ARs detected in digestates.
The detection of sequences specific for antibiotic RGs, qac genes, integron integrase genes, and for a range of transferable plasmids confirmed previous data by Binh et al. (2008) for farm-scale manures taken from farms in different regions in Germany, supporting their conclusion that manure from conventional farms is a reservoir of transferable antibiotic RGs. While Binh et al. (2008) determined the occurrence of sul1, sul2, and TEM-1 genes, in the present study, we focused on the detection of genes conferring resistances toward tetracycline antibiotics. We selected three tet genes (tet(A), tet(M), and tet(X)) that were previously reported to occur in manure (Lanz et al. 2003; Thames et al. 2012; Chee-Sanford et al. 2001; Kyselkova et al. 2013; Zhang et al. 2013) and that represent the three major resistance mechanisms toward tetracyclines (efflux, ribosomal protection protein, and enzymatic degradation, respectively). Although the Southern blot hybridization provided only semi-quantitative results, hybridization signal intensities indicated that all tet genes with a few exceptions were highly abundant in manure bacteria. The high abundances of tet genes and the detection of tetracycline residues in the same manures suggest that selective pressure increased the abundance of tet genes carrying bacteria. Likely, vertical and horizontal gene transfer processes might have played an important role in the dissemination of tetracycline RGs. In addition, these genes might have been subject to co-selection, as for instance, tet(A) and tet(X) were previously reported as linked with various RGs on transferable plasmids (Heuer et al. 2009; Wolters et al. 2015), while tet(M) was shown to co-occur with a diversity of other RGs in Staphylococcus aureus ST398 isolates from livestock related sources (Argudín et al. 2011). The sulfonamide RGs sul1 and sul2 seemed to be even more abundant than the tet genes, although sulfonamide residues were only detected in manure from F6 and B6. Explanations for this strikingly high abundance of sul genes and the absence of detected sulfonamide residues could be either their co-selection or their analytical problems due to the tight binding of sulfonamides to organic matter (Heuer et al. 2008). Also, in all digestates investigated in the present study, sul1, sul2, and tet genes were highly abundant; although in none of the samples, sulfonamide residues were detected, while in contrast, tetracycline residues were found in digestates from all BGPs except BGP8, but at approximately tenfold lower concentrations compared to manures. This high abundance of sul1, sul2, and tet(M) in the digestate samples was recently confirmed by quantitative real-time PCR with relative abundances in the range of Log −2.16 to Log −3.03, Log −1.80 to Log −2.75 and Log −1.94 to Log −2.52, respectively (Wolters et al. 2016). In the present study, we showed that qacE∆1 genes (conferring resistance toward quaternary ammonium compounds that are frequently used as disinfectants in animal husbandries) were detected in high abundance in all manure samples. Recently, Heuer et al. (2012) and Jechalke et al. (2014) reported on IncP-1ε plasmids that were exogenously isolated from manure and manure treated soils. The molecular analysis of a collection of 50 IncP-1ε plasmids revealed the presence of tet(A) and class 1 integrons with the presence of qacE∆1 and sul1 genes at the 3′end of the gene cassette region. Thus, a co-selection not only of sul genes but also of qacE∆1 can be assumed. In addition, the exogenous isolation of IncP-1ε plasmids from the digestates investigated in the presented study was recently reported by Wolters et al. (2015). All IncP-1ε plasmids captured carried tet(A) and class 1 integrons with sul1 and qacE∆1 at the 3′end of the gene cassette region, providing further support for the assumption of co-selection. The high proportion of IncP-1ε among the collection of exogenously captured plasmids that were obtained based on the sulfonamide or tetracycline resistance conferred was rather surprising as they were also obtained from BGPs with no IncP-1 plasmids detected in the present study (BGP7; Table 4). This finding also shows that plasmids with low abundance can be significant in the dissemination of RGs. The ability of IncP-1ε plasmids to efficiently transfer in soil and their BHR (Heuer et al. 2012; Musovic et al. 2014) makes them likely important vectors contributing to the dissemination of RGs in the agro-ecosystem. In digestates, the abundance of intI1, sul1, and qacE∆1 was high, while IncP-1 plasmids seemed much less abundant (Table 4). These findings were recently confirmed by qPCR (Wolters et al. 2016). Interestingly, the relative abundances of IncP-1ε plasmid-specific trfA gene was low (Log −4.62 to Log −6.67 that is close to or below the detection limit) compared to the relative abundances of intI1 (Log −3.12 to Log −3.50), sul1 (Log −2.16 to Log −3.03), or qacE∆1 (Log −2.26 to Log −2.88). Integrons can, however, also be localized chromosomally or on plasmids belonging to other incompatibility groups such as IncW plasmids (Revilla et al. 2008) that were detected in the digestates from all BGPs but not in all manures. In contrast to the study by Binh et al. (2008), IncN plasmids seemed to be less abundant in the manures from pig fattening farms and the digestates. Another plasmid type that seemed to be highly abundant in manure and in digestates were IncQ plasmids. Like the BHR plasmids, IncP-1, IncN, and IncW, plasmids belonging to the IncQ group have a BHR but they can only be transferred by mobilizing plasmids. IncQ plasmids typically carry sul2 genes. Smalla et al. (2000) showed not only the presence of mobilizing plasmids in pig manure but also the isolated IncQ plasmids conferring resistances toward tetracycline, streptomycin, sulfonamide, gentamycin, and streptothricin antibiotics. In contrast to recent reports on the frequent occurrence of LowGC plasmids (Binh et al. 2008; Heuer et al. 2009; Kopmann et al. 2012), these plasmids seemed to be less abundant in the present study and were only detected in a few manure and digestate samples, e.g., from BGP2.
The broad-host range plasmid groups detected in our study (IncP-1, IncN, IncW, and IncQ) in manures or digestates were previously also reported from clinical isolates. Indeed, many of these groups were originally discovered in clinical isolates (Datta et al. 1971; Loftie-Eaton and Rawlings 2012). Furthermore, they were frequently detected among Gram-negative bacteria isolated from wounds, urine, or sputum of hospital patients in a recent study by Zhang et al. (2014).
Comparing the antibiotic application patterns, detected ARs in the samples and corresponding detected RGs and MGEs, it was not only found that pig breeding facilities had the broadest overall antibiotic application patterns (Table 1) and that the spectrum of ARs detected in corresponding manures was larger than in pig fattening farms (Table 1) but also that the diversity of MGEs detected was higher (Table 3). As antibiotic treatment of livestock is inevitably impacting the composition of bacterial communities resident in the animals intestine and hence also in respective excrements (Heuer et al. 2011; Reichel et al. 2013), antibiotic application patterns may also influence the occurrence of RGs and MGEs in manure not necessarily accompanied by subsequent detection of ARs.
Although the detection of RG- and MGE-specific sequences in TC-DNA by means of Southern blot hybridization provides only semi-quantitative results, it was observed that the signals detected for tet(M) were often stronger in manures derived from pig fattening compared to pig breeding (Table 3). At the same time, the highest concentrations of tetracyclines were detected in pig fattening manures. Among the more than 50 known tetracycline RGs, of which only a small subset was monitored within the present study, tet(M) appears to have the broadest host range (http://faculty.washington.edu/marilynr/tetweb2.pdf&/tetweb3.pdf). In contrast to the studies of Kopmann et al. (2012), Heuer et al. (2012), and Jechalke et al. (2013), who reported on direct correlations of antibiotic treatment and antibiotic RGs (sul1 and sul2) and MGEs (IncP-1 and LowGC plasmids) in manures and manure treated soil, no differences related to concentrations of the antibiotics detected were found for RGs monitored in the present study, except for tet(M) as already mentioned.
The present study clearly showed that antibiotics such as tetracyclines that are frequently applied in conventional pig husbandry systems were detectable in liquid pig manures and digestates of BGPs, where they provide a selective advantage for bacterial populations carrying RGs and likely foster horizontal gene transfer. The fate and effects of veterinary antibiotics entering soils might strongly depend on the physicochemical characteristics of the antibiotics. Therefore, when being spread onto fields, manures and digestates might pose a potential risk of RG transfer via plasmids to bacteria associated with soil and plant environments.
The detection of antibiotics, RGs, and MGEs in digestates showed that the mesophilic fermentation process in BGPs is not an effective mitigation strategy. In order to reduce contamination of farm fertilizers with ARs and antibiotic resistant bacteria, the frequency of antibiotic usage needs to be reduced by optimizing the conditions of conventional pig husbandry systems. Furthermore, additional factors co-selecting antibiotic resistance such as disinfectants or metal compounds used as feed additives need to be better understood. It will be important to gain better understanding of the uptake of ARs by plants and to determine whether root exudates in the presence of ARs facilitates the horizontal spread of multiple antibiotic resistances. Raw leafy greens are not only transiently but also endophytically colonized by a subset of the soil microbiome, and thus, their mobile resistome might provide a link to the human gut microbiome.
References
Argudín MA, Tenhagen B-A, Fetsch A, Sachsenröder J, Käsbohrer A, Schroeter A, Hammerl JA, Hertwig S, Helmuth R, Bräunig J, Mendoza MC, Appel B, Rodicio MR, Guerra B (2011) Virulence and resistance determinants of German Staphylococcus aureus ST398 isolates from nonhuman sources. Appl Environ Microbiol 77(9):3052–3060
Arikan OA, Sikora LJ, Mulbry W, Khan SU, Rice C, Foster GD (2006) The fate and effect of oxytetracycline during the anaerobic digestion of manure from therapeutically treated calves. Process Biochem 41:1637–1643
Arikan OA (2008) Degradation and metabolization of chlortetracycline during the anaerobic digestion of manure from medicated calves. J Hazard Mater 158:485–490
Bahl MI, Burmølle M, Meisner A, Hansen LH, Sørensen SJ (2009) All IncP-1 plasmid subgroups, including the novel ε subgroup, are prevalent in the influent of a Danish wastewater treatment plant. Plasmid 62:134–139
Baquero F (2012) Metagenomic epidemiology: a public health need for the control of antimicrobial resistance. Clin Microbiol Infect 18(Suppl. 4):67–73
Bartha NA, Sóki J, Urbán E, Nagy E (2011) Investigation of the prevalence of tetQ, tetX and tetX1 genes in Bacteroides strains with elevated tigecycline minimum inhibitory concentrations. Int J Antimicrob Ag 38:522–525
Binh CTT, Heuer H, Kaupenjohann M, Smalla K (2008) Piggery manure used for fertilization is a reservoir for transferable antibiotic resistance plasmids. FEMS Microbiol Ecol 66:25–37
Boxall ABA, Fogg LA, Blackwell PA, Kay P, Pemberton EJ, Croxford A (2004) Veterinary medicines in the environment. Rev Environ Contam Toxicol 180: 1–91
Bundesamt für Verbraucherschutz und Lebensmittelsicherheit (BVL) (Federal Office of Consumer Protection and Food Safety), Germany (2015): http://www.bvl.bund.de
Chee-Sanford JC, Aminov RI, Krapac IJ, Garrigues-Jeanjean N, Mackie RI (2001) Occurrence and diversity of tetracycline resistance genes in lagoons and groundwater underlying two swine production facilities. Appl Environ Microbiol 67(4):1494–1502
Datta N, Hedges RW, Shaw EJ, Sykes RB, Richmond MH (1971) Properties of an R factor from Pseudomonas aeruginosa. J Bacteriol 108(3):1244–1249
Diehl DL, LaPara TM (2010) Effect of temperature on the fate of genes encoding tetracycline resistance and the integrase of class 1 integrons within anaerobic and aerobic digesters treating wastewater solids. Environ Sci Technol 44:9128–9133
Forsberg KJ, Reyes A, Wang B, Selleck EM, Sommer MOA, Dantas G (2012) The shared antibiotic resistome of soil bacteria and human pathogens. Science 337: 1107–1111.
Gans O, Pfundtner E, Winckler C, Bauer A (2010) Antibiotika in Biogasanlagen. Umweltbundesamt Wien, Austria, Report REP-0287, ISBN 978–3–99004-088-1, 1–48.
Goldstein C, Lee MD, Sanchez S, Hudson C, Phillips B, Register B, Grady M, Liebert C, Summers AO, White DG, Maurer JJ (2001) Incidence of class 1 and 2 integrases in clinical and commensal bacteria from livestock, companion animals, and exotics. Antimicrob Ag Chemother 45(3):723–726
Götz A, Pukall R, Smit E, Tietze E, Prager R, Tschäpe H, van Elsas JD, Smalla K (1996) Detection and characterization of broad-host-range plasmids in environmental bacteria by PCR. Appl Environ Microbiol 62(7):2621–2628
Götz A, Smalla K (1997) Manure enhances plasmid mobilization and survival of Pseudomonas putida introduced into field soil. Appl Environ Microbiol 63(5):1980–1986
Gullberg E, Cao S, Berg OG, Ilbäck C, Sandegren L, Hughes D, Andersson DI (2011) Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog 7(7):e1002158. doi:10.1371/journal.ppat.1002158
Gullberg E, Albrecht LM, Karlsson C, Sandegren L, Andersson DI (2014) Selection of a multidrug resistance plasmid by sublethal levels of antibiotics and heavy metals. mBIO 5(5):e01918–e01914. doi:10.1128/mBio.01918-14
Haller MY, Müller SR, McArdell CS, Alder AC, Suter MJF (2002) Quantification of veterinary antibiotics (sulfonamides and trimethoprim) in animal manure by liquid chromatography–mass spectrometry. J Chromatogr A 952:111–120
Heuer H, Focks A, Lamshöft M, Smalla K, Matthies M, Spiteller M (2008) Fate of sulfadiazine administered to pigs and its quantitative effect on the dynamics of bacterial resistance genes in manure and manured soil. Soil Biol Biochem 40:1892–1900
Heuer H, Kopmann C, Binh CTT, Top E, Smalla K (2009) Spreading antibiotic resistance through spread manure: characteristics of a novel plasmid type with low % G + C content. Environ Microbiol 11(4):937–949
Heuer H, Solehati Q, Zimmerling U, Kleineidam K, Schloter M, Müller T, Focks A, Thiele-Brun S, Smalla K (2011) Accumulation of sulfonamide resistance genes in arable soils due to repeated application of manure containing sulfadiazine. Appl Environ Microbiol 77(7):2527–2530
Heuer H, Binh CTT, Jechalke S, Kopmann C, Zimmerling U, Krögerrecklenfort E, Ledger T, González B, Top E, Smalla K (2012) IncP-1ε plasmids are important vectors of antibiotic resistance genes in agricultural systems: diversification driven by class 1 integron gene cassettes. Front Microbiol 3:Article 2. doi:10.3389/fmicb.2012.00002
Hirsch R, Ternes T, Haberer K, Kratz K-L (1999) Occurrence of antibiotics in the aquatic environment. Sci Total Environ 225:109–118
Jechalke S, Kopmann C, Rosendahl I, Groeneweg J, Weichelt V, Krögerrecklenfort E, Brandes N, Nordwig M, Ding G-C, Siemens J, Heuer H, Smalla K (2013) Increased abundance and transferability of resistance genes after field application of manure from sulfadiazine-treated pigs. Appl Environ Microbiol 79(5):1704–1711
Jechalke S, Schreiter S, Wolters B, Dealtry S, Heuer H, Smalla K (2014) Widespread dissemination of class 1 integron components in soils and related ecosystems as revealed by cultivation-independent analysis. Front Microbiol 4:article 420. doi:10.3389/fmicb.2013.00420
Kerrn MB, Klemmensen T, Frimodt-Möller N, Espersen F (2002) Susceptibility of Danish Escherichia coli strains isolated from urinary tract infections and bacteraemia, and distribution of sul genes conferring sulphonamide resistance. J Antimicrob Chemother 50:513–516
Kolz AC, Moorman TB, Ong SK, Scoggin KD, Douglass EA (2005) Degradation and metabolite production of tylosin in anaerobic and aerobic swine-manure lagoons. Water Environ Res 77:49–56
Kopmann C, Jechalke S, Rosendahl I, Groeneweg J, Krögerrecklenfort E, Zimmerling U, Weichelt V, Siemens J, Amelung W, Heuer H, Smalla K (2012) Abundance and transferability of antibiotic resistance as related to the fate of sulfadiazine in maize rhizosphere and bulk soil. FEMS Microbiol Ecol
Kraft CA, Timbury MC, Platt DJ (1986) Distribution and genetic location of Tn7 in trimethoprim-resistant Escherichia coli. J Med Microbiol 22:125–131
Kreuzig R, Höltge S (2005) Investigations on the fate of sulfadiazine in manured soil: laboratory experiments and test plot studies. Environ Toxicol Chem 24:771–776
Kurokawa K, Itoh T, Kuwahara T, Oshima K, Toh H, Toyoda A, Takami H, Morita H, Sharma VK, Srivastava TP, Taylor TD, Noguchi H, Mori H, Ogura Y, Ehrlich DS, Itoh K, Takagi T, Sakaki Y, Hayashi T, Hattori M (2007) Comparative metagenomics revealed commonly enriched gene sets in human gut microbiomes. DNA Res 14:169–181
Kyselková M, Jirout J, Chronáková A, Vrchotová N, Bradley R, Schmitt H, Elhottová D (2013) Cow excrements enhance the occurrence of tetracycline resistance genes in soil regardless of their oxytetracycline content. Chemosphere 93:2413–2418
Kyselková M, Jirout J, Vrchotová N, Schmitt H, Elhottová D (2015) Spread of tetracycline resistance genes at a conventional dairy farm. Front Microbiol 6:article 536. doi:10.3389/fmicb.2015.00536
Lamshöft M, Sukul P, Zühlke S, Spiteller M (2007) Metabolism of 14C-labelled and non-labelled sulfadiazine after administration to pigs. Anal Bioanal Chem 388:1733–1745
Lanz R, Kuhnert P, Boerlin P (2003) Antimicrobial resistance and resistance gene determinants in clinical Escherichia coli from different animal species in Switzerland. Vet Microbiol 91:73–84
Loftie-Eaton W, Rawlings DE (2012) Diversity, biology and evolution of IncQ-family plasmids. Plasmid 67:15–34
Marti R, Scott A, Tien YC, Murray R, Salbourin L, Zhang Y, Topp E (2013) Impact of manure fertilization on the abundance of antibiotic-resistant bacteria and frequency of detection of antibiotic resistance genes in soil and on vegetables at harvest. Appl Environ Microbiol 79(18):5701–5709
Martínez-Carballo E, González-Barreiro C, Scharf S, Gans O (2007) Environmental monitoring study of selected veterinary antibiotics in animal manure and soils in Austria. Environ Pollut 148:570–579
Mohring SA, Strzysch I, Fernandes MR, Kiffmeyer TK, Tuerk J, Hamscher G (2009) Degradation and elimination of various sulfonamides during anaerobic fermentation: a promising step on the way to sustainable pharmacy? Environ Sci Technol 43:2569–2574
Musovic S, Oregaard G, Krøer N, Sørensen SJ (2006) Cultivation-independent examination of horizontal transfer and host range of an IncP-1 plasmid among gram-positive and gram-negative bacteria indigenous to the barley rhizosphere. Appl Environ Microbiol 72(10):6687–6692
Musovic S, Klümper U, Dechesne A, Magid J, Smets BF (2014) Long-term manure exposure increases soil bacterial community potential for plasmid uptake. Environ Microbiol Rep 6(2):125–130
Ng L-K, Martin I, Alfa M, Mulvey M (2001) Multiplex PCR for the detection of tetracycline resistant genes. Mol Cell Probes 15:209–215
Ratsak C, Guhl B, Zühlke S, Delschen T (2013) Veterinärantibiotika-Rückstände in Gülle und Gärresten aus Nordrhein-Westfalen. Environ Sci Europe 25:1–11
Reichel R, Rosendahl I, Peeters ETHM, Focks A, Groeneweg J, Bierl R, Schlichting A, Amelung W, Thiele-Bruhn S (2013) Effects of slurry from sulfadiazine- (SDZ) and difloxacin- (DIF) medicated pigs on the structural diversity of microorganisms in bulk and rhizosphere soil. Soil Biol Biochem 62:82--91
Revilla C, Garcillán-Barcia MP, Fernández-Lopez R, Thomson NR, Sanders M, Cheung M, Thomas CM, de la Cruz F (2008) Different pathways to acquiring resistance genes illustrated by the recent evolution of IncW plasmids. Antimicrob Agents Chemother 52(4):1472–1480
Sandvang D, Aarestrup FM, Jensen LB (1997) Characterisation of integrons and antibiotic resistance genes in Danish multiresistant Salmonella enterica Typhimurium DT104. FEMS Microbiol Lett 157:177–181
Sarmah AK, Meyer MT, Boxall ABA (2006) A global perspective on the use, sales, exposure pathways, occurrence, fate and effects of veterinary antibiotics (VAs) in the environment. Chemosphere 65:725–759
Schlüsener MP, Von Arb MA, Bester K (2006) Elimination of macrolides, tiamulin, and salinomycin during manure storage. Arch Environ Contam Toxicol 51:21–28
Smalla K, Heuer H, Götz A, Niemeyer D, Krögerrecklenfort E, Tietze E (2000) Exogenous isolation of antibiotic resistance plasmids from piggery manure slurries reveals a high prevalence and diversity of IncQ-like plasmids. Appl Environ Microbiol 66(11):4854–4862
Spielmeyer A, Ahlborn J, Hamscher G (2014) Simultaneous determination of 14 sulfonamides and tetracyclines in biogas plants by liquid-liquid-extraction and liquid chromatography tandem mass spectrometry. Anal Bioanal Chem 406:2513–2524
Spielmeyer A, Breier B, Groißmeier K, Hamscher G (2015) Elimination patterns of worldwide used sulfonamides and tetracyclines during anaerobic fermentation. Bioresour Technol 193:307–314
Thames CH, Pruden A, James RE, Ray PP, Knowlton KF (2012) Excretion of antibiotic resistance genes by dairy calves fed milk replacers with varying doses of antibiotics. Front Microbiol 3:article 139. doi:10.3389/fmicb.2012.00139
Tolls J (2001) Sorption of veterinary pharmaceuticals in soils: a review. Environ Sci Technol 35:3397–3406
Udo HMJ, Aklilu HA, Phong LT, Bosma RH, Budisatria IGS, Patil BR, Samdup T, Bebe BO (2011) Impact of intensification of different types of livestock production in smallholder crop-livestock systems. Livest Sci 139:22–29
Uslu MÖ, Yediler A, Balcıoğlu IA, Schulte-Hostede S (2008) Analysis and sorption behavior of fluoroquinolones in solid matrices. Water Air Soil Pollut 190:55–63
Widyasari-Mehta A, Hartung S, Kreuzig R (2016a) From the application of antibiotics to antibiotic residues in liquid manures and digestates: a screening study in one European center of conventional pig husbandry. J Environ Manage 177:129–137
Widyasari-Mehta A, Suwito HRKA, Kreuzig R (2016b) Laboratory testing on the removal of the veterinary antibiotic doxycycline during long-term liquid pig manure and digestate storage. Chemosphere 149:154–160
Wolters B, Kyselková M, Krögerrecklenfort E, Kreuzig R, Smalla K (2015) Transferable antibiotic resistance plasmids from biogas plant digestates often belong to the IncP-1ε subgroup. Front Microbiol 5:article 765. doi:10.3389/fmicb.2014.00765
Wolters B, Ding G-C, Kreuzig R, Smalla K (2016) Full-scale mesophilic biogas plants using manure as C-source: bacterial community shifts along the process cause changes in the abundance of resistance genes and mobile genetic elements. FEMS Microbiol Ecol 92:fiv163. doi:10.1093/femsec/fiv163
Wu S, Dalsgaard A, Hammerum AM, Porsbo LJ, Jensen LB (2010) Prevalence and characterization of plasmids carrying sulfonamide resistance genes among Escherichia coli from pigs, pig carcasses and human. Acta Vet Scand 52:article 47
Zhang Y, Zhang C, Parker DB, Snow DD, Zhou Z, Li X (2013) Occurrence of antimicrobials and antimicrobial resistance genes in beef cattle storage ponds and swine treatment lagoons. Sci Total Environ 463-464:631–638
Zhang G-Q, Yao Y-H, X-I Y, Niu J-J (2014) A survey of five broad-host-range plasmids in gram-negative bacilli isolated from patients. Plasmid 74:9–14
Zhu Y-G, Johnson TA, J-Q S, Qiao M, Gou G-X, Stedtfeld RD, Hashsham SA, Tiedje JM (2013) Diverse and abundant antibiotic resistance genes in Chinese swine farms. PNAS 110(9):3435–3440
Acknowledgments
The authors acknowledge the financial support from the German Federal Ministry of Food, Agriculture, and Consumer Protection through the Federal Office for Agriculture and Food, Bonn, Germany (grant number 2810HS032). We thank Dr. G. Steffens, T. Eilers, Dr. A. Freitag, and Dr. K. Lacü, Chamber of Agriculture, Oldenburg and Braunschweig, Germany, as well as the farmers in Lower Saxony, Germany, who substantially support these research activities. Additionally, we thank Ilse-Marie Jungkurth (Julius Kühn-Institut, Braunschweig) and Kim Cook (US Department of Agriculture, Athens, GA, USA) for proofreading of the manuscript and Ina-Kristin Behrens for her contribution to this work in the frame of a bachelor’s thesis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was funded by the German Federal Ministry of Food, Agriculture, and Consumer Protection through the Federal Office for Agriculture and Food, Bonn, Germany (grant number 2810HS032).
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Birgit Wolters and Arum Widyasari-Mehta equally contributed to this work.
Rights and permissions
About this article
Cite this article
Wolters, B., Widyasari-Mehta, A., Kreuzig, R. et al. Contaminations of organic fertilizers with antibiotic residues, resistance genes, and mobile genetic elements mirroring antibiotic use in livestock?. Appl Microbiol Biotechnol 100, 9343–9353 (2016). https://doi.org/10.1007/s00253-016-7742-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7742-y