Abstract
The engineered Saccharomyces cerevisiae strain △faa1△faa4 [Acot5s] was demonstrated to accumulate more free fatty acids (FFA) previously. Here, comparative proteomic analysis was performed to get a global overview of metabolic regulation in the strain. Over 500 proteins were identified, and 82 of those proteins were found to change significantly in the engineered strains. Proteins involved in glycolysis, acetate metabolism, fatty acid synthesis, TCA cycle, glyoxylate cycle, the pentose phosphate pathway, respiration, transportation, and stress response were found to be upregulated in △faa1△faa4 [Acot5s] as compared to the wild type. On the other hand, proteins involved in glycerol, ethanol, ergosterol, and cell wall synthesis were downregulated. Taken together with our metabolite analysis, our results showed that the disruption of Faa1 and Faa4 and expression of Acot5s in the engineered strain △faa1△faa4 [Acot5s] not only relieved the feedback inhibition of fatty acyl-CoAs on fatty acid synthesis, but also caused a major metabolic rearrangement. The rearrangement redirected carbon flux toward the pathways which generate the essential substrates and cofactors for fatty acid synthesis, such as acetyl-CoA, ATP, and NADPH. Therefore, our results help shed light on the mechanism for the increased production of fatty acids in the engineered strains, which is useful in providing information for future studies in biofuel production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbial synthesized fatty acids have increasingly attracted attention in recent decades as a potential feedstock for biofuel production. This is because microbially synthesized fatty acids are easily converted into many fuel molecules such as alkanes, alcohols, and biodiesel through biochemical or catalytic chemical methods (Choi and Lee 2013; Howard et al. 2013; Liu et al. 2013; Steen et al. 2010). Recently, there has been great progress in engineering Saccharomyces cerevisiae to produce free fatty acids (FFA). It was shown that by disruption of the two main acyl-CoA synthases, Faa1 and Faa4, the yeast strain secreted 220 μmol/L (∼60 mg/L) of FFA (Scharnewski et al. 2008). Besides FAA1 and FAA4 disruption, overexpression of Escherichia coli acyl-ACP thioesterase in S. cerevisiae yielded 207 mg/L FFA, and combined with overexpression of the fatty acid synthase (FAS) and acetyl-CoA carboxylase, resulted in the production of up to 400 mg/L FFA (Runguphan and Keasling 2014). In another study, it was demonstrated that the disruption of the β-oxidation pathway and the acyl-CoA synthetases, overexpression of different thioesterases, and enhancement of the supply of acetyl-CoA resulted in the production of more than 120 mg/L FFA (Li et al. 2014). In a latest study, it was found that disruption of the acyl-CoA synthetase genes FAA1, FAA4, and FAT1 yielded 490 mg/L FFA, combined with disruption of the β-oxidation pathway and co-expression of lipid body forming enzyme DGA1 and lipase TGL3, 2.2 g/L FFA was produced (Leber et al. 2015).
In a previous study, we successfully increased the accumulation of FFA to ∼500 mg/L in minimal medium by engineering the acyl-CoA metabolism of S. cerevisiae. This was done by double deletion of acyl-CoA synthases Faa1 and Faa4, and cytosolic expression of truncated version of acyl-CoA thioesterase Acot5 (Acot5s) from Mus musculus (Chen et al. 2014). We found a significant increase in extracellular fatty acids in the engineered strains △faa1△faa4 and △faa1△faa4 [Acot5s], as compared to the wild-type (WT) strain. This showed that the enzymes acyl-CoA synthetases Faa1, Faa4, and thioesterase Acot5s had a direct effect on FFA accumulation. In addition, our RT-PCR results showed the genes involved in fatty acid synthesis were de-repressed in the strains △faa1△faa4 and △faa1△faa4 [Acot5s], although it should be taken into account that mRNA levels do not always correlate with protein expression (Griffin et al. 2002; Ideker et al. 2001). Therefore, to understand the mechanism for the increase of FFA in the engineered strains, proteomic analysis is needed.
The regulation of the gene, protein, and metabolic network are closely connected in S. cerevisiae (Smedsgaard and Nielsen 2005). The enzymes involved in fatty acid synthesis and elongation are regulated on both the gene expression and protein level. In addition, compounds needed in the fatty acid synthesis process such as acetyl-CoA, ATP, and NADPH are competed for by many other cellular reactions and metabolic pathways (Tehlivets et al. 2007).
Our previous study on acyl-CoA metabolism in engineered cells has provided important insights into the regulation of fatty acid synthesis. The use of proteomic analysis will further aid the understanding of the effects of gene modification, and also enable a global overview of metabolic regulation. Hence, we conduct comparative proteomic analysis between the WT, △faa1△faa4, and △faa1△faa4 [Acot5s] strains in this study. The changes in proteins involved in different metabolic pathways were identified and analyzed. Furthermore, the global metabolic changes in the engineered strains were mapped.
Materials and methods
Strain growth and cell harvest
All strains used in this study were constructed based on S. cerevisiae WT strain BY4741 (MATa; his3△1; leu2△0; met15△0; ura3△0, EUROSCARF). As previously described (Chen et al. 2014), the control strain WT [pVTU260] was constructed by transforming the WT strain with plasmid pVTU260 (also shown as WT). The double deletion strain △faa4△faa1 was transformed with the plasmid pVTU260 to construct the strain △faa4△faa1 [pVTU260] (also shown as △faa4△faa1). The strain △faa1△faa4 [Acot5s] was constructed by transforming △faa4△faa1 with the plasmid pVTU260-Acot5s. All strains were streaked on YNBD-Ura agar plates (0.67 % yeast nitrogen base (with no amino acids but with ammonium sulfate, Invitrogen), 2 % dextrose, amino acid drop out (without Ura, Clonetech), and 2 % agar), and incubated at 30 °C for 2–3 days. Five milliliters of YNBD-Ura liquid medium were inoculated with a single colony and incubated overnight at 30 °C, 250 rpm. This pre-culture was then used to inoculate 50 mL YNBD-Ura liquid medium with an initial OD600 (optical density at 600 nm) of 0.2. OD600 was measured using Nano Drop 2000 (Thermo Scientific) to monitor growth status. Cell culture samples were collected at different time points. The culture supernatant and cell pellet were collected separately by centrifugation at 10,000g for 10 min at 4 °C.
Protein Extraction, Digestion, and Labeling with iTRAQ Reagents
Cells with an OD600 of 20 from each sample were collected, washed with PBS buffer, and re-suspended in 500 μL NP40 cell lysis buffer (Invitrogen) supplemented with 1 mM PMSF and protease inhibitor cocktail. Then, 300 μL acid-washed glass beads were added and the cells were lysed using a FastPrep®-24 Instrument (6004–500, MP Biomedicals) for 20 s, and chilled on ice for 30 s. This was repeated six times. Next, the cells were centrifuged at 12,000g for 10 min at 4 °C. The clear lysate was then transferred to new sterile microcentrifuge tubes which were pre-chilled on ice.
Protein concentration was determined by 2-D Quant Kit (GE Healthcare). Then, 100 μg of protein from each sample was cleaned up using acetone precipitation. Four volumes of ice cold acetone were added to each sample, and the protein was precipitated at −20 °C for at least 2 h. After centrifugation at 12,000g for 10 min, the acetone was removed. Then, the sample was digested and labeled using the iTRAQ Reagent Multiplex Kit (AB SCIEX) according to manufacturer’s instructions. First, each protein sample was re-dissolved in 20 μL dissolution buffer and 1 μL denaturant, followed by adding 2 μL reducing reagent and incubated at 60 °C for 1 h. Then, 1 μL cysteine blocking reagent was added into each tube and incubated at room temperature for 10 min. Next, 20 μL of 0.25 μg/μL sequence-grade modified trypsin (Promega) was added into each sample. The digestion reaction was carried out at 37 °C for 12–16 h. Samples from three strains were labeled with a different iTRAQ tag, respectively. The labeled peptides were mixed together accordingly and vacuum evaporated to proper concentration.
Online 2-D Nano-LC-MS/MS analysis
The labeled samples were analyzed using an Agilent 1200 series nanoflow liquid chromatography system (Agilent Technologies) interfaced with 6530 Q-TOF mass spectrometer (Agilent Technologies) (Shi et al. 2013). A HPLC-Chip Cube MS Interface (Agilent Technologies) coupled with the columns PolySulfoethyl A SCX column (0.32 × 50 mm, 5 μm) and HPLC-Chip (Zorbax 300SB C18 enrichment column (0.3 mm × 5 mm, 5 μm) and Zorbax 300SB C18 reversed-phase column (75 μm × 50 mm, 3.5 μm) was used for two-dimensional analyses.
In the first dimension, 4 μL of the combined peptides mixture was loaded onto the SCX column and eluted stepwise by 8 μL of ammonium formate solutions in a series of concentrations—20, 40, 60, 80, 100, 300, 500, and 1000 mM. In the second dimension, the peptides were eluted from SCX column and trapped on the Zorbax 300SB C18 enrichment column using buffer A (5 % acetonitrile and 0.1 % formic acid in water) at a flow rate of 4 μL/min. The peptides which were trapped on the enrichment column were then eluted and separated by the Zorbax 300SB C18 reversed-phase column. The elution buffers used were Buffer B (0.1 % formic acid in water) and Buffer C (a nanoflow gradient of 5–80 % acetonitrile + 0.1 % formic acid in water) at a flow rate of 300 nL/min. The total run time was 62 min for each round running. In all, nine runs were carried out for each sample due to the gradient elution. The effluent was analyzed by the MS. Survey scans were obtained from m/z 300 to 2000 at acquisition rate of 4 spectra per second, with up to two precursors selected for MS/MS analysis from m/z 50 to 2000 (Shi et al. 2013).
LC-MS/MS data analysis and interpretation
Protein identification and quantification were performed using Spectrum Mill software (Agilent Technologies). The MS/MS spectra acquired were searched against the database UniProtKB/Swiss-Prot with the following criteria: two maximum missed cleavages, 20 ppm precursor mass tolerance, 50 ppm product mass tolerance, methylmethanethiosulfate-labeled cysteine as the fixed modification, and iTRAQ modification of free amine in the amino terminus. For further statistical analysis, the following criteria were used: two or more identified unique peptides, protein MS/MS search score greater than 11, and peptide score greater than 6. Three independent experiments were carried out to calculate the standard deviation and confidence value. Those with more than 95 % confidence were selected for further analysis (Shi et al. 2013).
HPLC analysis of intracellular and extracellular metabolites
Metabolites in the filtered culture supernatant such as glucose, ethanol, glycerol, and acetate were quantified using an Agilent 1200 HPLC system (Singapore) equipped with a Rezex ROA-organic acid H+ (8 %) column (300 × 7.8 mm, Omega Scientific Phenomenex, Singapore). 5 mM H2SO4 was used as the mobile phase with a flow rate of 0.6 mL/min. The oven was set at 40 °C, and a refractive index detector was used. The samples were quantitated using standards.
ATP and NADPH assay
Intracellular ATP and NADPH were detected using ATP Colorimetric/Fluorometric Assay Kit and NADP/NADPH Quantitation Kit (BioVision, USA). Because they are labile during sampling, we used comparative ratio to indicate the changes between the engineered cells and the WT.
Results
In our previous work, it was shown that FFA production was enhanced in engineered strains △faa1△faa4 and △faa1△faa4 [Acot5s]. By comparing the fatty acids levels in the WT [Acot5s], △faa1, △faa1 [Acot5s], △faa4, △faa4 [Acot5s], △faa1△faa4, △faa1△faa4 [Acot5s], and WT strains, we observed a significant increase in extracellular FFA in the engineered strains △faa1△faa4 and △faa1△faa4 [Acot5s], while no significant differences were observed in the WT [Acot5s], △faa1, △faa1 [Acot5s], △faa4, and △faa4 [Acot5s]. The removal of genes FAA1 and FAA4 which encode for acyl-CoA synthetases led to the accumulation of a more than 4-fold extracellular fatty acid (326.97 μg/mL) as compared to the WT strain (76.5 μg/mL). The expression of cytosolic acyl-CoA thioesterase Acot5s in △faa1△faa4 [Acot5s] enhanced the total FFA production to about 500 μg/mL, which is more than 6-fold as compared to the WT, and 1.5-fold increase as compared to the strain △faa1△faa4. In addition, we also detected a high proportion of unsaturated extracellular fatty acids (UFA: C16:1 and C18:1) in △faa1△faa4 and △faa1△faa4 [Acot5s], whereas these could not be detected in the WT or the single deletion strains (Chen et al. 2014). Acyl-CoAs act as feedback inhibitors on fatty acid biosynthesis. The modification of acyl-CoA metabolism reduced the accumulation of acyl-CoAs, therefore relieving the inhibition and enhanced FFA production (Chen et al. 2014). The ability of S. cerevisiae to secrete FFA into medium also provided a driving force for fatty acid production, which is also seen in Runguphan and Keasling (2014). To obtain a global overview of the in vivo proteome changes in the engineered strains, a comparative proteomic analysis was performed.
iTRAQ analysis and protein identification
iTRAQ labeling was used to compare and analyze the in vivo protein changes between the WT, △faa1△faa4, and △faa1△faa4 [Acot5s] strains. Proteins were extracted from samples harvested at 24 h, and labeled using the iTRAQ Reagent Multiplex Kit (AB SCIEX). Proteomic analysis was performed at least three times for statistical analysis. In total, over 500 proteins were detected. Among them, 82 proteins were found to be significantly altered in the engineered strains as compared to the WT as listed in Table S1 with their relative fold changes. Thirty proteins were found to be significantly altered in △faa1△faa4 [Acot5s] as compared to △faa1△faa4 as listed in Table 1.
Relative changes in protein expression between the WT and engineered strains
The identified proteins which were shown to have significant changes could be classified into different biological process categories according to their functions as annotated in the UniProtKB database (http://www.uniprot.org/) (Fig. 1). These functions included carbon metabolism, amino acid and nucleotide metabolism, protein biosynthesis, respiration, transportation, stress response, antioxidants, and others. The majority of proteins belonged to the category of carbon metabolism, which accounted for 37.80 % of all proteins (Fig. 1), showing that a major metabolic rearrangement occurred in the engineered cells as a result of the genetic modifications.
Functional classification of proteins identified. Proteins were classified into eight groups according to their functions annotated in the UniProtKB database (http://www.uniprot.org/). The percentage of proteins of each function group out of the whole proteins identified was exhibited
Among these proteins involved in carbon metabolism (Table S1), eight proteins (Glk1, Pyk1, Tpi1, Pgi1, Hxk1, Hxk2, Eno2, and Pdb1) which are involved in the glycolysis were upregulated in the strain △faa4△faa1, whereas five proteins (Glk1, Hxk1, Gpm2, Eno2, and Pdb1) were upregulated in the strain △faa4△faa1 [Acot5s]. Enzyme Adh1 which is responsible for ethanol production showed upregulation in the strain △faa4△faa1, whereas it remained unchanged in the strain △faa4△faa1 [Acot5s]. Four enzymes (Ald6, Ald4, Acs2, and Ach1) involved in acetate metabolism were upregulated in both △faa4△faa1 and △faa4△faa1 [Acot5s] strains. Apart from the acetate biosynthetic process (Saint-Prix et al. 2004), Ald6 and Ald4 are also involved in NADPH regeneration (Grabowska and Chelstowska 2003). Acetyl-CoA synthetase 2 (Acs2) is involved in acetyl-CoA synthesis from acetate in the pyruvate-acetaldehyde-acetate pathway (Falcón et al. 2010). Ach1 is involved in shuttling mitochondrial acetyl-CoAs for cytosolic C2 provision (Chen et al. 2015) and acetate detoxification (Fleck and Brock 2009). The enzyme Gpp1 (Påhlman et al. 2001) which takes part in the glycerol biosynthetic process was downregulated in both △faa4△faa1 and △faa4△faa1 [Acot5s] strains, whereas the enzyme Gut2 (Wang et al. 2001) involved in the glycerol catabolic process was upregulated in both △faa4△faa1 and △faa4△faa1 [Acot5s] strains. Moreover, two enzymes (Acc1 and Acb1) which participate in fatty acid metabolism, one enzyme (Icl1) involved in glyoxylate cycle (Schöler and Schüller 1993), and six enzymes (Cit1, Idh1, Idh2, Kgd1, Fum1, and Mdh1) involved in the TCA cycle were upregulated in both △faa4△faa1 and △faa4△faa1 [Acot5s] strains. Four enzymes (Gnd1, Tkl1, Tal1, and Nqm1) involved in the pentose phosphate pathway were upregulated in the strain △faa4△faa1, whereas two enzymes (Gnd1 and Tal1) were upregulated in the strain △faa4△faa1 [Acot5s]. For enzymes involved in carbohydrate biosynthesis, the major phosphoglucomutase isoform Pgm2 (Fu et al. 2000) responsible for hexose metabolism showed upregulation in both △faa4△faa1 and △faa4△faa1 [Acot5s] strains. A previous study showed that protein abundance of Pmg2 also increased in response to various stresses (Hirata et al. 2003). For enzymes involved in the ergosterol biosynthetic process, Erg13 (Barker et al. 2003) was upregulated in △faa4△faa1, whereas it was downregulated in △faa4△faa1 [Acot5s].
Three enzymes (Gas1, Mpg1, and Pmi40) involved in cell wall synthesis were downregulated in both △faa4△faa1 and △faa4△faa1 [Acot5s] strains. Six enzymes (Oye2, Atp1, Atp2, Atp3, Atp5, and Aac2) involved in respiration were upregulated in △faa4△faa1, while five enzymes (Atp1, Atp2, Atp3, Atp5, and Aac2) were detected upregulated in △faa4△faa1 [Acot5s]. For proteins involved in the amino acid biosynthesis pathway, it was detected that eight enzymes (Arg1, Arg4, Sah1, Shm2, Lys1, Lys12, Lys21, and His4) were downregulated and four enzymes (Cys3, Ser3, Gdh1, and Uga1) were upregulated in △faa4△faa1. Ten enzymes (Arg1, Arg4, Sah1, Ser3, Shm2, Lys1, Lys12, Lys21, Gdh1, and His4) were downregulated and one enzyme Cys3 was upregulated in △faa4△faa1 [Acot5s]. For proteins involved in the protein biosynthesis pathway, four enzymes (Pdi1, Ils1, Fpr3, and Stm1) were upregulated in the strain △faa4△faa1, and three enzymes (Pdi1, Fpr3, and Rps2) were upregulated in the strain △faa4△faa1 [Acot5s].
For proteins involved in sugar, protein, and other ion transport process, six proteins (Hxt6, Pma1, Trx2, Por1, Chc1, and Grx1) were upregulated in the strain △faa4△faa1. Among these proteins, all other five proteins except Chc1 showed similar increase in the strain △faa4△faa1 [Acot5s]. Twelve proteins (Gre2, Mcr1, Tps1, Tps2, Dcs1, Ynk1, Prb1, Glr1, Hsp26, Hsp42, Hsp104, and Ara1) involved in stress response showed increase in their expression levels in the strain △faa4△faa1. Comparatively, six proteins (Gre2, Mcr1, Dcs1, Hsp26, Hsp30, and Hsp104) involved in stress response showed increase in the strain △faa4△faa1 [Acot5s].
The level of antioxidant proteins Ahp1 (Lee et al. 1999), Sod1 (Sturtz et al. 2001), and Sod2 (Saffi et al. 2006) were increased in the strain △faa4△faa1, while only Sod1 was increased in the strain △faa4△faa1 [Acot5s]. Moreover, Rib4 involved in cofactor riboflavin biosynthesis (García-Ramírez et al. 1995), inositol-3-phosphate synthase (Ino1) (Shirra et al. 2001) involved in inositol synthesis, and phosphoinositide phosphatase Sac1 (Whitters et al. 1993) involved in the coordination of the activities of the secretory pathway showed upregulation in both strains.
Relative changes in protein expression between △faa4△faa1 [Acot5s] and △faa4△faa1
To investigate the influence of Acot5s expression on proteome changes, a direct comparison of protein changes between △faa4△faa1 [Acot5s] and △faa4△faa1 was conducted. As shown in Table 1, 10 proteins which play a role in carbon metabolism were found to be markedly changed in △faa1△faa4 [Acot5s] as compared to △faa1△faa4. Among these proteins, three enzymes (Pyk1, Pgi1, and Hxk2) involved in glycolysis, one enzyme (Adh1) involved in ethanol production, one enzyme (Gpp1) involved in the glycerol biosynthetic process, and one enzyme (Erg13) involved in the ergosterol biosynthesis pathway were downregulated in the strain △faa4△faa1 [Acot5s]. One enzyme (Ach1) that is involved in acetate metabolism, two enzymes (Kgd1 and Mdh1) involved in the TCA cycle, and one enzyme (Icl1) involved in glyoxylate cycle were upregulated.
Moreover, three proteins (Atp1, Atp3, and Aac2) involved in respiration and one phosphoinositide phosphatase Sac1 were upregulated in △faa4△faa1 [Acot5s]. Six proteins involved in stress response (Tps1, Tps2, Glr1, Hsp42, Hsp104, and Ara1) and two antioxidant proteins (Ahp1 and Sod2) were found downregulated. Two enzymes (Gas1 and Pmi40) involved in cell wall synthesis and one enzyme (Rib4) involved in cofactor biosynthesis were also downregulated in △faa4△faa1 [Acot5s]. Besides, five enzymes involved in amino acid biosynthesis were found differentially expressed.
Physiological and metabolite analysis of the engineered and WT strains
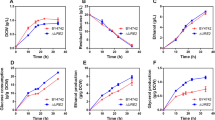
The WT and engineered strains were cultured in shake flask with YNB medium containing 20 g/L glucose as the carbon source. Growth curve and the concentration of several extracellular metabolites were studied. As shown in Fig. 2a, strain △faa1△faa4 had a defective growth and slower exponential growth rate, while strain △faa1△faa4 [Acot5s] displayed a higher growth rate as compared with the WT strain. This was also seen in our previous study (Chen et al. 2014). As showed in Fig. 2b, glucose consumption was delayed in the strain △faa1△faa4 as compared with the WT and △faa1△faa4 [Acot5s], the delayed glucose consumption in strain △faa1△faa4 was consistent with its slow growth rate. As for glycerol accumulation (Fig. 2c), △faa1△faa4 [Acot5s] had the lowest glycerol concentration, whereas the WT strain had the highest glycerol concentration. For acetate accumulation (Fig. 2d), △faa1△faa4 and △faa1△faa4 [Acot5s] had much lower final acetate concentration as compared with the WT strain. Regarding ethanol accumulation (Fig. 2e), the three strains had similar ethanol concentration curve. The strain △faa1△faa4 [Acot5s] gradually accumulated a lower level of ethanol after 24 h than the WT and △faa1△faa4 strains. In addition, intracellular ATP and NADPH levels of three strains were examined. As expected, the strains △faa1△faa4 and △faa1△faa4 [Acot5s] had higher ATP and NADPH levels as compared to the WT strain in both exponential phase and stationary phase (Figs. 3 and 4).
Physiological features of the engineered and WT strains. The WT strain and engineered strains were cultured in YNBD medium. Cell growth (a), extracellular concentration of glucose (b), glycerol (c), acetate (d), and ethanol (e) were measured at different time points as indicated in method. Data are the mean values of three independent experiments. The error bar indicates the S.D.
Discussion
In our previous study, it was shown that FFA production was enhanced by disrupting both acyl-CoA synthetases Faa1 and Faa4, and expressing acyl-CoA thioesterase Acot5s in S. cerevisiae. This modification restrained FFA activation, relieved the feedback inhibition of acyl-CoAs on fatty acid synthesis, and directed carbon flux toward FFA accumulation (Chen et al. 2014). However, how the cells sensed the changes in fatty acid metabolism and responded accordingly was unknown. Therefore, we carried out proteomic analysis in this study, and mapped the changes in proteins in relation to the biochemical pathways to give a global overview of the pathway regulations (Figs. 5 and 6).
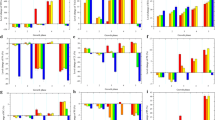
Changes of metabolic pathways in the strain △faa1△faa4 as compared with the WT. The primary pathways are described according to the changes in protein expression as listed in Table S1. Red boxes and red text indicate upregulated pathways and proteins. Blue boxes and blue text indicate downregulated pathways and proteins
Changes of metabolic pathways in the strain △faa1△faa4 [Acot5s] as compared with the WT. The primary pathways are described according to the changes in protein expression as listed in Table S1. Red boxes and red text indicate upregulated pathways and proteins. Blue boxes and blue text indicate downregulated pathways and proteins
Comparative proteome analysis of the engineered strains as compared to the WT
There were many changes in enzymes involved in crucial biological pathways, due to the double deletion of genes FAA1 and FAA4 and expression of Acot5s. As shown in Table S1, several enzymes involved in glycolysis, a fundamental pathway in glucose metabolism to provide energy in yeast (Bruce Alberts and Lewis 2002), were upregulated in both engineered strains. The average increment of these enzymes was 30.75 % in △faa1△faa4 and 30.2 % in △faa1△faa4 [Acot5s]. In the glycerol metabolism pathway, the abundance of the enzyme Gpp1 (Påhlman et al. 2001), which is involved in the glycerol biosynthetic process, was found to decrease to 79 and 53 % in the strains △faa1△faa4 and △faa1△faa4 [Acot5s], respectively, as compared with the WT strain. The relative abundance of the enzyme Gut2 (Wang et al. 2001), which is involved in the glycerol catabolic process, increased more than 30 % in both engineered strains. This was consistent with the data that both engineered strains accumulated less glycerol than the WT strain (Fig. 4c). In addition, decrease in intracellular glycerol levels were also seen in both engineered strains in our previous metabolite profiling data (Chen et al. 2014). The decrease in glycerol synthesis probably provided extra pyruvate for other pathways. In the pyruvate-ethanol pathway, the relative abundance of protein Adh1, which is the main alcohol dehydrogenase that catalyzes the conversion of acetaldehyde to ethanol (Leskovac et al. 2002), increased 2-fold in the strain △faa1△faa4, while it remained unchanged in the strain △faa1△faa4 [Acot5s]. Because several proteins in the pyruvate-acetaldehyde-acetate pathway were upregulated in △faa1△faa4 [Acot5s], it is not a surprise that the strain △faa1△faa4 [Acot5s] had lower concentration of ethanol than the WT and △faa1△faa4 after 24 h (Fig. 2e). However, we did not find significant increase of ethanol concentration in the strain △faa1△faa4. This may be due to it had slower growth rate and lower biomass accumulation than the WT strain, and several proteins in its pyruvate-acetaldehyde-acetate pathway were also upregulated. There were three enzymes (Erg10, Erg13, and Erg20) detected in the ergosterol biosynthesis pathway. Erg13 were upregulated to 112 % in △faa4△faa1 and downregulated to 64 % in △faa4△faa1 [Acot5s] as compared to the WT. There were no significant changes in the level of Erg10 and Erg20 (data not shown). This might indicate that there was decreased carbon flux toward ergosterol production in the strain △faa4△faa1 [Acot5s]. Further experiment to estimate the regulation of the ergosterol pathway in △faa4△faa1 would be required.
Besides the abovementioned carbon metabolic regulation, multiple levels of regulation in fatty acid metabolism of S. cerevisiae exist. Fatty acid synthesis is a highly energy-demanding process, which requires large amounts of acetyl-CoA, ATP, and NADPH (Tehlivets et al. 2007). The supply of these compounds has significant effect on fatty acid synthesis. Acetyl-CoA in S. cerevisiae can be produced from both the mitochondrial pyruvate dehydrogenase pathway and the cytosolic pyruvate-acetaldehyde-acetate pathway (Chen et al. 2012; Pronk et al. 1996). As shown in Table S1, enzyme Pdb1 (Miran et al. 1993), which is part of the pyruvate dehydrogenase complex and involved in catalyzing the overall conversion of pyruvate to acetyl-CoA in the mitochondria, was upregulated in both strains. The levels of several enzymes in the pyruvate-acetaldehyde-acetate pathway were also increased. Cytosolic aldehyde dehydrogenase (Ald6) which plays a major role in acetate formation and a compensative mitochondrial aldehyde dehydrogenase (Ald4) (Saint-Prix et al. 2004) were upregulated in both engineered strains. Acetyl-CoA synthetase 2 (Acs2) (Carman et al. 2008; Van Den Berg and Steensma 1995), which generates acetyl-CoA by direct activation of acetate in the cytoplasm, was also upregulated by about 15 % in both strains. This was consistent with the observation that both engineered strains accumulated less acetate than the WT strain (Fig.2d). In addition, the enzyme Icl1 (Schöler and Schüller 1993), which catalyzes the formation of succinate and glyoxylate from isocitrate, a key step in the glyoxylate cycle, was also upregulated. Glyoxylate cycle is one known route for transporting acetyl-CoA from the peroxisomes/cytosol into the cytosol/mitochondria through converting acetyl-CoA into glyoxylate cycle intermediates (van Roermund et al. 1995). The upregulation of Icl1 in the glyoxylate cycle implied there may be increased need for acetyl-CoA transportation. Furthermore, the upregulation of Ach1 may imply the same increased need, since mitochondrial Ach1 can transport acetyl-CoA produced in the mitochondria to the cytosol via conversion of acetyl-CoA into acetate, which can cross the mitochondrial membrane freely (Chen et al. 2015). Although we did not observe significant increase in cellular acetyl-CoA level in the engineered strains (data not shown), the significant increase of FFA production implied that there should be increased supply of acetyl-CoA.
ATP can be generated through oxidation of glucose in glycolysis, the TCA cycle, and oxidative phosphorylation. The amount of energy released by oxidative phosphorylation is much higher as compared to glycolysis (Rich 2003). In addition to the upregulation of enzymes in glycolysis, six enzymes involved in the TCA cycle, and several subunits of ATP synthase (Reinders et al. 2007) in aerobic respiration were observed to be significantly upregulated in both engineered strains. In the strain △faa1△faa4, enzymes in the TCA cycle were upregulated from 1.13- to 1.59-fold, and subunits of ATP synthase were upregulated from 1.18- to 1.24-fold as compared to the WT strain. In the strain △faa1△faa4 [Acot5s], enzymes in the TCA cycle were upregulated from 1.21- to1.80-fold, and subunits of ATP synthase were upregulated from 1.26- to 1.62-fold as compared to the WT strain. Besides, the major ADP/ATP carrier of the mitochondrial inner membrane Aac2 was also upregulated in both engineered strains. Aac2 imports cytosolic ADP into the mitochondria, where it can be phosphorylated by ATP synthase, and exports ATP into the cytosol (Klingenberg 2008; Ruprecht et al. 2014). So, the upregulation of Aac2 was consistent with the upregulation of ATP synthase. The analysis of the ATP level in three strains demonstrated there were higher levels of ATP in two engineered strains as compared to the WT, which further demonstrated the upregulation of ATP synthesis.
NADPH can be provided by several sources such as malic enzyme, the pentose phosphate pathway, NADPH isocitrate dedydrogenase, NAD kinase (Tehlivets et al. 2007), and the aldehyde dehydrogenase family (Grabowska and Chelstowska 2003; Minard and McAlister-Henn 2005). In the present analysis, we found that several enzymes involved in NADPH generation were upregulated. For example, the level of Gnd1 (6-phosphogluconate dehydrogenase), which catalyzes the NADPH generation step in the pentose phosphate pathway, increased by about 55 % in both strains. Ald6 and Ald4, which utilize NADP+ as the preferred coenzyme when performing the conversion of acetaldehyde to acetate and play important roles in NADPH regeneration (Grabowska and Chelstowska 2003), were upregulated by 25 and 26 % in △faa1△faa4, 15 and 43 % in △faa1△faa4 [Acot5s]. The increased intracellular NADPH level in both engineered strains further proved there was increased NADPH synthesis in both engineered strains.
Furthermore, the enzymes involved in fatty acid synthesis were also altered, although the results were different from the transcriptional level analysis (Chen et al. 2014); that is, the expression of genes related to fatty acid synthesis which are repressed by fatty acyl-CoAs were increased significantly in the engineered strains △faa1△faa4 and △faa1△faa4 [Acot5s], especially OLE1 which encodes a △ − 9 fatty acid desaturase. However, the protein Acc1 was increased by only about 15 %, while the RT-PCR result showed several fold increase. The increase in Acc1 protein level was smaller than its increase in gene expression level. There was no observed significant increase in the level of Fas1 and Fas2. △ − 9 fatty acid desaturase was not detected, which may be due to its location in the endoplasmic reticulum membrane (Kim et al. 2006). Also, regulation of these proteins is complex and occur at multiple levels, including histone acetylation, mRNA stability, protein stability, and coordination with other genes expression (Tehlivets et al. 2007). The increase in mRNA level with no subsequent increase in protein expression level is understandable and also supported by other studies (Griffin et al. 2002; Ideker et al. 2001). Although Acc1 affects fatty acid synthesis directly and overexpression of Acc1 could enhance fatty acid synthesis (Davis et al. 2000; Lennen et al. 2010; Ruenwai et al. 2009), overexpression of Acc1 in eukaryotic organism generally results in small improvement of fatty acid production (Li et al. 2014; Runguphan and Keasling 2014). Further study is needed to explore the role of upregulation of Acc1 in the increased fatty acid production. Moreover, the level of acyl-CoA binding protein Acb1, which facilitates the cellular transportation of acyl-CoA and lessens the inhibition effect of acyl-CoA on fatty acid synthesis (Rasmussen et al. 1993; Vock et al. 2010), was increased 2.45-fold and 2.43-fold, respectively, in △faa1△faa4 and △faa1△faa4 [Acot5s].
Several proteins involved in transportation were found to be upregulated in both engineered strains, especially the high-affinity hexose transporter Hxt6, which plays an important role in glucose uptake and utilization (Reifenberger et al. 1995). It was upregulated more than 3-fold in both strains. This was consistent with the upregulation of glycolysis in both strains. Previous study showed that overproduction of fatty acids in E. coli resulted in the change of membrane properties because of fatty acid intercalation in the membrane, fatty acid accumulation in the periplasm, and alteration of membrane lipids; these changes then led to membrane stress and compromised membrane integrity (Lennen et al. 2011). In our engineered strains, the downregulation of enzymes (Gas1, Mpg1, and Pmi40) involved in cell wall synthesis (Payton et al. 1991; Tomishige et al. 2003; Yoda et al. 2000) and upregulation of enzyme Sac1 (phosphoinositide phosphatase) involved in the secretory pathway (Whitters et al. 1993) should also be the response of fatty acid overproduction, such regulation may facilitate FFA to be secreted outside the cells.
In cells, amino acid biosynthesis pathways are not only an essential process for protein synthesis, they are also critical for yeast cells to achieve coordinated regulation between nitrogen source utilization and nucleotide biosynthetic pathways (Ljungdahl and Daignan-Fornier 2012). In our observation, many enzymes involved in amino acid biosynthesis (Table S1) were differentially expressed in two engineered strains as compared to the WT. Among these enzymes, Shm2 (McNeil et al. 1994) is the major enzyme responsible for converting serine to glycine. Lys1, Lys12, and Lys21 (Mark Zabriskie and Jackson 2000) are responsible for L-lysine biosynthesis. The downregulation of these enzymes was consistent with the observed decrease in L-lysine and glycine levels in previous metabolic profiling result (Fig. 6 in (Chen et al. 2014)). For other enzymes involved in amino acid synthesis, there were no significant changes detected in the relevant amino acids levels in our previous study. Since cells have a natural ability to reduce biosynthetic cost to optimize growth and reproduction, the regulation of amino acid, protein, and nucleotide biosynthesis should be under the need of metabolic cost optimization (Barton et al. 2010).
Also, DNA microarray experiment showed that environmental stresses, including temperature shock and oxidative stress, greatly stimulate upregulation of the transcription of stress-responsive genes in yeast cells (Gasch et al. 2000). In our study, we observed that more enzymes involved in stress response and antioxidants were upregulated in the strain △faa1△faa4 as compared to the strain △faa1△faa4 [Acot5s]. For example, D-arabinose dehydrogenase Ara1 (Hu et al. 2013; van Bergen et al. 2006), which was re-annotated as α, β-dicarbonyl reductase and found involved in reducing a variety of toxic metabolic by-products such as methylglyoxal, diacetyl, and pentanedione produced during stress, was only upregulated in △faa1△faa4. This was consistent with the growth defect of the strain △faa1△faa4 and recovered growth of the strain △faa1△faa4 [Acot5s] (Fig. 2a), which indicated the stresses caused by removing Faa1 and Faa4 were relieved in a certain degree by introducing Acot5s.
The impact of Acot5s expression on proteome changes
As stated above, the growth phenotype of the strain △faa4△faa1 is more severe than the strain △faa4△faa1 [Acot5s]; thus, it is not surprising that there are more changes about proteins involved in central metabolism in △faa4△faa1 as opposed to △faa4△faa1 [Acot5s]. We wonder how the △faa4△faa1 [Acot5s] cells responded to the driving force provided by Acot5s and had higher FFA production and growth rate compared to △faa4△faa1 and WT strains. Through comparatively analyzing protein expression levels between △faa4△faa1 [Acot5s] and △faa4△faa1, the affected proteins that take part in crucial metabolic pathways were mapped in Fig. 7. Among them, the level of alcohol dehydrogenase Adh1, which is involved in ethanol production, decreased to 49 % in △faa4△faa1 [Acot5s] as compared to △faa4△faa1. The level of enzyme Gpp1 involved in the glycerol biosynthetic process was downregulated to 70 % in △faa4△faa1 [Acot5s] as compared to △faa4△faa1. This indicated that the added driving force provided by Acot5s made cells a direct part of their carbon flux from ethanol and glycerol production to acetyl-CoA synthesis. Although we did not observe additional increase in protein level involved in the pyruvate-acetaldehyde-acetate pathway, we did found decrease in ethanol and glycerol production and increase of FFA production in the strain △faa4△faa1 [Acot5s]. In addition, Erg13 involved in the ergosterol biosynthesis pathway were downregulated to 58 % of that in △faa4△faa1, which may imply decreased carbon flux toward ergosterol.
Changes of metabolic pathways in the strain △faa1△faa4 [Acot5s] as compared with △faa1△faa4. The primary pathways are described according to the changes in protein expression as listed in Table 1. Red boxes and red text indicate upregulated pathways and proteins. Blue boxes and blue text indicate downregulated pathways and proteins
Protein Ach1, which is involved in acetyl-CoA transportation and acetate detoxification (Chen et al. 2015), was upregulated 1.9-fold in △faa4△faa1 [Acot5s] as compared to △faa4△faa1. In addition, the reported study (Chen et al. 2015) also showed that overexpression of ACH1 in the △ach1 strain not only rescued growth on glucose but also led to slightly faster growth than the △ach1 strain. The higher expression level of Ach1 may explain the higher growth rate of △faa4△faa1 [Acot5s]. Furthermore, the downregulation of stress response and antioxidants proteins also showed the △faa4△faa1 [Acot5s] cells adapted to the new balanced state built by Acot5s. Moreover, the upregulation of protein Kgd1 and Mdh1 (1.19- and 1.31-fold) involved in the TCA cycle, and Atp1, Atp3, and Aac2 (1.54-, 1.22- and 1.27-fold) involved in respiration were consistent with the higher ATP level.
The direct effect of removing acyl-CoA synthetases Faa1 and Faa4 was restraining FFA activation, and led to FFA secretion into the culture medium as demonstrated before (Scharnewski et al. 2008). Although the mechanism of fatty acid secretion has not been unraveled completely yet, it was reported that the fatty acid transport could be driven by the concentration gradient between extracellular and intracellular fatty acid and need the activities of fatty acyl-CoA synthetase (Faa1 and Faa4) and fatty acid transport protein 1 (Fat1). The deletion of FAA1 and FAA4 increases the concentration of FFA in the cytosol which leads to the secretion of FFA (Scharnewski et al. 2008), it also decreases the concentration of intracellular acyl-CoA which results in changes in intracellular signaling to fatty acid-responsive genes (Black and DiRusso 2007; Færgeman et al. 2001). In our study, the expression of thioesterase reinforced the effect; thus, the modification provided a higher driving force toward higher fatty acid production. Compared with other recent works (Leber et al. 2015; Li et al. 2014; Runguphan and Keasling 2014), besides using different background strains, the main difference between these studies is the driving force, which is provided by different thioesterases (Li et al. 2014; Runguphan and Keasling 2014) or lipid body forming enzymes with lipase (Leber et al. 2015). Our study showed that Acot5s is a pretty efficient enzyme to transform acyl-CoA to FFA. The expression of Acot5s affected the regulation of proteins involved in crucial metabolic pathways, which led to a major metabolic rearrangement while not harming growth.
In summary, to gain insights into the underlying metabolic regulation associated with the genetic modification in the engineered strains, we set out to investigate the proteomic profile changes between WT and engineered strains. It was found that the disruption of genes FAA1 and FAA4 and expression of Acot5s in S. cerevisiae contributed to elevated carbon flux toward FFA accumulation. It was identified that proteins involved in glycolysis, the pentose phosphate pathway, TCA cycle, fatty acid synthesis, and ATP synthesis and transportation were upregulated, while proteins involved in glycerol synthesis, pyruvate-ethanol synthesis, ergosterol synthesis, and cell wall synthesis were downregulated in the strain △faa1△faa4 [Acot5s]. Our detected changes in metabolites supported the finding regarding the changes in protein expression in the different metabolic pathways. Previous study demonstrated that the genetic modification relieved the feedback inhibition of fatty acyl-CoAs (Chen et al. 2014). The global metabolic regulation found in this study indicated that the cells upregulated certain pathways to facilitate fatty acid synthesis and substrates supply, and efficiently utilize the carbon flux by downregulating some pathways and channeling the carbon flux toward FFA accumulation. This information is of great help to unravel the mechanism for the increased production of FFA in the engineered strain. It also shows the great potential of the engineered strain in the metabolic adjustment, and it will certainly aid future work in rational engineering of the yeast system for biofuel production. For example, the flux toward alcohol biosynthesis could be further decreased in the engineered strain, and enzymes in the pyruvate-acetaldehyde-acetate pathway or Ach1p could be overexpressed to reinforce the carbon flux toward cytosolic acetyl-CoA supply. Several genes in β-oxidation could be deleted to diminish cellular fatty acid degradation. More importantly, the driving force could be enhanced to further promote FFA production.
References
Barker KS, Pearson MM, Rogers PD (2003) Identification of genes differentially expressed in association with reduced azole susceptibility in Saccharomyces cerevisiae. J Antimicrob Chemother 51(5):1131–1140. doi:10.1093/jac/dkg217
Barton MD, Delneri D, Oliver SG, Rattray M, Bergman CM (2010) Evolutionary systems biology of amino acid biosynthetic cost in yeast. PLoS One 5(8):e11935. doi:10.1371/journal.pone.0011935
Black PN, DiRusso CC (2007) Yeast acyl-CoA synthetases at the crossroads of fatty acid metabolism and regulation. BBA-Mol Cell Biol L 1771(3):286–298. doi:10.1016/j.bbalip.2006.05.003
Bruce Alberts AJ, Lewis J (2002) Molecular biology of the cell, 4th edn. Garland Science, New York
Carman AJ, Vylkova S, Lorenz MC (2008) Role of acetyl coenzyme A synthesis and breakdown in alternative carbon source utilization in Candida albicans. Eukaryotic Cell 7(10):1733–1741. doi:10.1128/ec.00253-08
Chen Y, Siewers V, Nielsen J (2012) Profiling of cytosolic and peroxisomal acetyl-CoA metabolism in Saccharomyces cerevisiae. PLoS One 7(8):e42475. doi:10.1371/journal.pone.0042475
Chen L, Zhang J, Lee J, Chen W (2014) Enhancement of free fatty acid production in Saccharomyces cerevisiae by control of fatty acyl-CoA metabolism. Appl Microbiol Biotechnol 98(15):6739–6750. doi:10.1007/s00253-014-5758-8
Chen Y, Zhang Y, Siewers V, Nielsen J (2015) Ach1 is involved in shuttling mitochondrial acetyl units for cytosolic C2 provision in Saccharomyces cerevisiae lacking pyruvate decarboxylase. FEMS Yeast Res 15(3) doi:10.1093/femsyr/fov015
Choi YJ, Lee SY (2013) Microbial production of short-chain alkanes. Nature 502(7472):571–574 doi:10.1038/nature12536 http://www.nature.com/nature/journal/v502/n7472/abs/nature12536.html#supplementary-information
Davis MS, Solbiati J, Cronan JE (2000) Overproduction of acetyl-CoA carboxylase activity increases the rate of fatty acid biosynthesis in Escherichia coli. J Biol Chem 275(37):28593–28598. doi:10.1074/jbc.M004756200
Færgeman NJ, Black PN, Zhao XD, Knudsen J, DiRusso CC (2001) The Acyl-CoA synthetases encoded within FAA1 and FAA4 in Saccharomyces cerevisiae function as components of the fatty acid transport system linking import, activation, and intracellular utilization. J Biol Chem 276(40):37051–37059. doi:10.1074/jbc.M100884200
Falcón A, Chen S, Wood M, Aris J (2010) Acetyl-coenzyme A synthetase 2 is a nuclear protein required for replicative longevity in Saccharomyces cerevisiae. Mol Cell Biochem 333(1–2):99–108. doi:10.1007/s11010-009-0209-z
Fleck CB, Brock M (2009) Re-characterisation of Saccharomyces cerevisiae Ach1p: fungal CoA-transferases are involved in acetic acid detoxification. Fungal Genet Biol 46(6–7):473–485. doi:10.1016/j.fgb.2009.03.004
Fu L, Miseta A, Hunton D, Marchase RB, Bedwell DM (2000) Loss of the major isoform of phosphoglucomutase results in altered calcium homeostasis in Saccharomyces cerevisiae. J Biol Chem 275(8):5431–5440. doi:10.1074/jbc.275.8.5431
García-Ramírez JJ, Santos MA, Revuelta JL (1995) The Saccharomyces cerevisiae RIB4 gene codes for 6,7-dimethyl- 8-ribityllumazine synthase involved in riboflavin biosynthesis: molecular characterization of the gene and purification of the encoded protein. J Biol Chem 270(40):23801–23807. doi:10.1074/jbc.270.40.23801
Gasch AP, Spellman PT, Kao CM, Carmel-Harel O, Eisen MB, Storz G, Botstein D, Brown PO (2000) Genomic expression programs in the response of yeast cells to environmental changes. Mol Biol Cell 11(12):4241–4257. doi:10.1091/mbc.11.12.4241
Grabowska D, Chelstowska A (2003) The ALD6 gene product is indispensable for providing NADPH in yeast cells lacking glucose-6-phosphate dehydrogenase activity. J Biol Chem 278(16):13984–13988. doi:10.1074/jbc.M210076200
Griffin TJ, Gygi SP, Ideker T, Rist B, Eng J, Hood L, Aebersold R (2002) Complementary profiling of gene expression at the transcriptome and proteome levels in Saccharomyces cerevisiae. Mol Cell Proteomics 1(4):323–333. doi:10.1074/mcp.M200001-MCP200
Hirata Y, Andoh T, Asahara T, Kikuchi A (2003) Yeast glycogen synthase kinase-3 activates Msn2p-dependent transcription of stress responsive genes. Mol Biol Cell 14(1):302–312. doi:10.1091/mbc.E02-05-0247
Howard TP, Middelhaufe S, Moore K, Edner C, Kolak DM, Taylor GN, Parker DA, Lee R, Smirnoff N, Aves SJ, Love J (2013) Synthesis of customized petroleum-replica fuel molecules by targeted modification of free fatty acid pools in Escherichia coli. Proc Natl Acad Sci U S A. doi:10.1073/pnas.1215966110
Hu X-Q, Guo P-C, Ma J-D, Li W-F (2013) Structures of Saccharomyces cerevisiae d-arabinose dehydrogenase Ara1 and its complex with NADPH: implications for cofactor-assisted substrate recognition. Acta Crystallogr Sect F 69(11):1190–1195. doi:10.1107/S1744309113026857
Ideker T, Thorsson V, Ranish JA, Christmas R, Buhler J, Eng JK, Bumgarner R, Goodlett DR, Aebersold R, Hood L (2001) Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 292(5518):929–934. doi:10.1126/science.292.5518.929
Kim H, Melén K, Österberg M, von Heijne G (2006) A global topology map of the Saccharomyces cerevisiae membrane proteome. Proc Natl Acad Sci U S A 103(30):11142–11147. doi:10.1073/pnas.0604075103
Klingenberg M (2008) The ADP and ATP transport in mitochondria and its carrier. Biochim Biophys Acta Biomembr 1778(10):1978–2021. doi:10.1016/j.bbamem.2008.04.011
Leber C, Polson B, Fernandez-Moya R, Da Silva NA (2015) Overproduction and secretion of free fatty acids through disrupted neutral lipid recycle in Saccharomyces cerevisiae. Metab Eng 28(0):54–62. doi:10.1016/j.ymben.2014.11.006
Lee J, Spector D, Godon C, Labarre J, Toledano MB (1999) A new antioxidant with alkyl hydroperoxide defense properties in yeast. J Biol Chem 274(8):4537–4544. doi:10.1074/jbc.274.8.4537
Lennen RM, Braden DJ, West RM, Dumesic JA, Pfleger BF (2010) A process for microbial hydrocarbon synthesis: overproduction of fatty acids in Escherichia coli and catalytic conversion to alkanes. Biotechnol Bioeng 106(2):193–202. doi:10.1002/bit.22660
Lennen RM, Kruziki MA, Kumar K, Zinkel RA, Burnum KE, Lipton MS, Hoover SW, Ranatunga DR, Wittkopp TM, Marner WD, Pfleger BF (2011) Membrane Stresses Induced by Overproduction of Free Fatty Acids in Escherichia coli. Appl Environ Microbiol 77(22):8114–8128 doi:10.1128/aem.05421-11
Leskovac V, Trivić S, Peričin D (2002) The three zinc-containing alcohol dehydrogenases from baker’s yeast, Saccharomyces cerevisiae. FEMS Yeast Res 2(4):481–494. doi:10.1111/j.1567-1364.2002.tb00116.x
Li X, Guo D, Cheng Y, Zhu F, Deng Z, Liu T (2014) Overproduction of fatty acids in engineered Saccharomyces cerevisiae. Biotechnol Bioeng 111(9):1841–1852. doi:10.1002/bit.25239
Liu J-F, Nie K-L, Fan L-H, Wang F, Tan T-W, Deng L (2013) Increased production of FAEEs for biodiesel with lipase enhanced Saccharomyces cerevisiae. Process Biochem 48(8):1212–1215. doi:10.1016/j.procbio.2013.06.003
Ljungdahl PO, Daignan-Fornier B (2012) Regulation of amino acid, nucleotide, and phosphate metabolism in Saccharomyces cerevisiae. Genetics 190(3):885–929. doi:10.1534/genetics.111.133306
Mark Zabriskie T, Jackson MD (2000) Lysine biosynthesis and metabolism in fungi. Nat Prod Rep 17(1):85–97. doi:10.1039/A801345D
McNeil JB, McIntosh EM, Taylor BV, Zhang FR, Tang S, Bognar AL (1994) Cloning and molecular characterization of three genes, including two genes encoding serine hydroxymethyltransferases, whose inactivation is required to render yeast auxotrophic for glycine. J Biol Chem 269(12):9155–9165
Minard KI, McAlister-Henn L (2005) Sources of NADPH in yeast vary with carbon source. J Biol Chem 280(48):39890–39896. doi:10.1074/jbc.M509461200
Miran SG, Lawson JE, Reed LJ (1993) Characterization of PDHβ1, the structural gene for the pyruvate dehydrogenase β subunit from Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 90(4):1252–1256. doi:10.1073/pnas.90.4.1252
Påhlman A-K, Granath K, Ansell R, Hohmann S, Adler L (2001) The yeast glycerol 3-phosphatases Gpp1p and Gpp2p are required for glycerol biosynthesis and differentially involved in the cellular responses to osmotic, anaerobic, and oxidative stress. J Biol Chem 276(5):3555–3563. doi:10.1074/jbc.M007164200
Payton MA, Rheinnecker M, Klig LS, DeTiani M, Bowden E (1991) A novel Saccharomyces cerevisiae secretory mutant possesses a thermolabile phosphomannose isomerase. J Bacteriol 173(6):2006–2010
Pronk JT, Yde Steensma H, Van Dijken JP (1996) Pyruvate metabolism in Saccharomyces cerevisiae. Yeast 12(16):1607–33. doi:10.1002/(SICI)1097-0061(199612)12:16
Rasmussen JT, Rosendal J, Knudsen J (1993) Interaction of acyl-coA-binding protein (ACBP) on processes for which acyl-coA is a substrate, product or inhibitor. Biochem J 292:907–913
Reifenberger E, Freidel K, Ciriacy M (1995) Identification of novel HXT genes in Saccharomyces cerevisiae reveals the impact of individual hexose transporters on qlycolytic flux. Mol Microbiol 16(1):157–167. doi:10.1111/j.1365-2958.1995.tb02400.x
Reinders J, Wagner K, Zahedi RP, Stojanovski D, Eyrich B, van der Laan M, Rehling P, Sickmann A, Pfanner N, Meisinger C (2007) Profiling phosphoproteins of yeast mitochondria reveals a role of phosphorylation in assembly of the ATP synthase. Mol Cell Proteomics 6(11):1896–1906. doi:10.1074/mcp.M700098-MCP200
Rich PR (2003) The molecular machinery of Keilin’s respiratory chain. Biochem Soc Trans 31(Pt 6):1095–105. doi:10.1042/BST0311095
Ruenwai R, Cheevadhanarak S, Laoteng K (2009) Overexpression of Acetyl-CoA carboxylase gene of Mucor rouxii enhanced fatty acid content in Hansenula polymorpha. Mol Biotechnol 42(3):327–332. doi:10.1007/s12033-009-9155-y
Runguphan W, Keasling JD (2014) Metabolic engineering of Saccharomyces cerevisiae for production of fatty acid-derived biofuels and chemicals. Metab Eng 21(0):103–113. doi:10.1016/j.ymben.2013.07.003
Ruprecht JJ, Hellawell AM, Harding M, Crichton PG, McCoy AJ, Kunji ERS (2014) Structures of yeast mitochondrial ADP/ATP carriers support a domain-based alternating-access transport mechanism. Proc Natl Acad Sci U S A 111(4):E426–E434. doi:10.1073/pnas.1320692111
Saffi J, Sonego L, Varela QD, Salvador M (2006) Antioxidant activity of L-ascorbic acid in wild-type and superoxide dismutase deficient strains of Saccharomyces cerevisiae. Redox Rep 11(4):179–184. doi:10.1179/135100006X116691
Saint-Prix F, Bönquist L, Dequin S (2004) Functional analysis of the ALD gene family of Saccharomyces cerevisiae during anaerobic growth on glucose: the NADP + −dependent Ald6p and Ald5p isoforms play a major role in acetate formation. Microbiology 150(7):2209–2220. doi:10.1099/mic.0.26999-0
Scharnewski M, Pongdontri P, Mora G, Hoppert M, Fulda M (2008) Mutants of Saccharomyces cerevisiae deficient in acyl-CoA synthetases secrete fatty acids due to interrupted fatty acid recycling. FEBS J 275(11):2765–2778. doi:10.1111/j.1742-4658.2008.06417.x
Schöler A, Schüller HJ (1993) Structure and regulation of the isocitrate lyase gene ICL1 from the yeast Saccharomyces cerevisiae. Curr Genet 23(5–6):375–381. doi:10.1007/bf00312621
Shi J, Feng H, Lee J, Ning Chen W (2013) Comparative proteomics profile of lipid-cumulating oleaginous yeast: An iTRAQ-coupled 2-D LC-MS/MS analysis. PLoS One 8(12):e85532. doi:10.1371/journal.pone.0085532
Shirra MK, Patton-Vogt J, Ulrich A, Liuta-Tehlivets O, Kohlwein SD, Henry SA, Arndt KM (2001) Inhibition of acetyl coenzyme A carboxylase activity restores expression of the INO1 gene in a snf1 mutant strain of Saccharomyces cerevisiae. Mol Cell Biol 21(17):5710–5722. doi:10.1128/mcb.21.17.5710-5722.2001
Smedsgaard J, Nielsen J (2005) Metabolite profiling of fungi and yeast: from phenotype to metabolome by MS and informatics. J Exp Biol 56(410):273–286. doi:10.1093/jxb/eri068
Steen EJ, Kang YS, Bokinsky G, Hu ZH, Schirmer A, McClure A, del Cardayre SB, Keasling JD (2010) Microbial production of fatty-acid-derived fuels and chemicals from plant biomass. Nature 463(7280):559–U182. doi:10.1038/nature08721
Sturtz LA, Diekert K, Jensen LT, Lill R, Culotta VC (2001) A fraction of yeast Cu/Zn superoxide dismutase and its metallochaperone, CCS, localize to the intermembrane space of mitochondria: a physiological role for SOD1 in guarding against mitochondrial oxidative damage. J Biol Chem. doi:10.1074/jbc.M105296200
Tehlivets O, Scheuringer K, Kohlwein SD (2007) Fatty acid synthesis and elongation in yeast. BBA-Mol Cell Biol L 1771(3):255–270. doi:10.1016/j.bbalip.2006.07.004
Tomishige N, Noda Y, Adachi H, Shimoi H, Takatsuki A, Yoda K (2003) Mutations that are synthetically lethal with a gas1Δ allele cause defects in the cell wall of Saccharomyces cerevisiae. Mol Gen Genomics 269(4):562–573. doi:10.1007/s00438-003-0864-9
van Bergen B, Strasser R, Cyr N, Sheppard JD, Jardim A (2006) α,β-dicarbonyl reduction by Saccharomyces d-arabinose dehydrogenase. Biochim Biophys Acta Gen Subj 1760(11):1636–1645. doi:10.1016/j.bbagen.2006.08.025
Van Den Berg MA, Steensma HY (1995) ACS2, a Saccharomyces Cerevisiae gene encoding acetyl-coenzyme A synthetase, essential for growth on glucose. Eur J Biochem 231(3):704–713. doi:10.1111/j.1432-1033.1995.0704d.x
van Roermund CW, Elgersma Y, Singh N, Wanders RJ, Tabak HF (1995) The membrane of peroxisomes in Saccharomyces cerevisiae is impermeable to NAD(H) and acetyl-CoA under in vivo conditions. EMBO J 14(14):3480–3486
Vock C, Biedasek K, Boomgaarden I, Heins A, Nitz I, Doring F (2010) ACBP knockdown leads to down-regulation of genes encoding rate-limiting enzymes in cholesterol and fatty acid metabolism. Cell Physiol Biochem 25(6):675–686. doi:10.1159/000315087
Wang Z, Zhuge J, Fang H, Prior BA (2001) Glycerol production by microbial fermentation: a review. Biotechnol Adv 19(3):201–223. doi:10.1016/S0734-9750(01)00060-X
Whitters E, Cleves A, McGee T, Skinner H, Bankaitis V (1993) SAC1p is an integral membrane protein that influences the cellular requirement for phospholipid transfer protein function and inositol in yeast. J Cell Biol 122(1):79–94. doi:10.1083/jcb.122.1.79
Yoda K, Kawada T, Kaibara C, Fujie A, Abe M, Hashimoto H, Shimizu J, Tomishige N, Noda Y, Yamasaki M (2000) Defect in cell wall integrity of the yeast Saccharomyces cerevisiae caused by a mutation of the gdp-mannose pyrophosphorylase gene VIG9. Biosci Biotechnol Biochem 64(9):1937–1941. doi:10.1271/bbb.64.1937
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was funded by a Competitive Research Programme (CRP) grant from the National Research Foundation of Singapore (grant number NRF-CRP5-2009-03).
Conflict of interest
Liwei Chen declares that she has no conflict of interest. Jaslyn Jie Lin Lee declares that she has no conflict of interest. Jianhua Zhang declares that he has no conflict of interest. Wei Ning Chen declares that he has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
(PDF 45 kb)
Rights and permissions
About this article
Cite this article
Chen, L., Lee, J.J.L., Zhang, J. et al. Comparative proteomic analysis of engineered Saccharomyces cerevisiae with enhanced free fatty acid accumulation. Appl Microbiol Biotechnol 100, 1407–1420 (2016). https://doi.org/10.1007/s00253-015-7028-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-7028-9










