Abstract
Acetic acid is present in cellulosic hydrolysate as a potent inhibitor, and the superior acetic acid tolerance of Saccharomyces cerevisiae ensures good cell viability and efficient ethanol production when cellulosic raw materials are used as substrates. In this study, a mutant strain of S. cerevisiae ATCC4126 (Sc4126-M01) with improved acetic acid tolerance was obtained through screening strains transformed with an artificial zinc finger protein transcription factor (ZFP-TF) library. Further analysis indicated that improved acetic acid tolerance was associated with improved catalase (CAT) activity. The ZFP coding sequence associated with the improved phenotype was identified, and real-time RT-PCR analysis revealed that three of the possible genes involved in the enhanced acetic acid tolerance regulated by this ZFP-TF, namely YFL040W, QDR3, and IKS1, showed decreased transcription levels in Sc4126-M01 in the presence of acetic acid, compared to those in the control strain. Sc4126-M01 mutants having QDR3 and IKS1 deletion (ΔQDR3 and ΔIKS1) exhibited higher acetic acid tolerance than the wild-type strain under acetic acid treatment. Glucose consumption rate and ethanol productivity in the presence of 5 g/L acetic acid were improved in the ΔQDR3 mutant compared to the wild-type strain. Our studies demonstrated that the synthetic ZFP-TF library can be used to improve acetic acid tolerance of S. cerevisiae and that the employment of an artificial transcription factor can facilitate the exploration of novel functional genes involved in stress tolerance of S. cerevisiae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Production of biofuels using lignocellulosic feedstocks has been extensively studied in recent years and is becoming an attractive alternative to fossil fuels. However, the economics and production efficiency of biofuels derived from cellulosic materials are still not satisfactory. One of the main factors that negatively affect the fermentation capability of Saccharomyces cerevisiae is the low cell viability in the presence of various toxic inhibitors in the cellulosic hydrolysate (Almeida et al. 2011; Hasunuma and Kondo 2012; Demeke et al. 2013; Lv et al. 2014).
Acetic acid is an important inhibitory agent in cellulosic hydrolysate (Almeida et al. 2011; Hasunuma and Kondo 2012; Jönsson et al. 2013), and it can negatively affect cell growth and ethanol production of S. cerevisiae (Ullah et al. 2012; Woo et al. 2014). Acetic acid inhibits the uptake of nutrients by the cells, leading to energy depletion and decreased activities of metabolic enzymes (Kitanovic et al. 2012; Ding et al. 2013). At the same time, the emergence of acetic acid-resistant spoilage yeasts is also a serious problem in food preservation (Stratford et al. 2013). Therefore, understanding the mechanisms of acetic acid tolerance in yeast and being able to improve such property is of great importance to various biological applications, especially the production of ethanol from cellulosic materials.
It is well known that multiple genes are involved in the stress tolerance of microbes (reviewed by Zhao and Bai 2009; Nicolaou et al. 2010), and it is therefore a very challenging task to improve this property by genetic engineering. Among the various methods for developing robust strains with improved stress tolerance, global transcriptional machinery engineering (gTME) has been proven to be an effective method (Alper et al. 2006; Yang et al. 2011), which achieves global transcription network regulation by manipulating the TATA-binding protein encoding gene SPT15. It is conceivable that other transcription factors can also be applied in gTME studies concerned with the metabolic engineering of microbial strains.
Zinc finger proteins (ZFPs) belong to an important family of transcription factors and play vital roles in various aspects of cellular metabolism (Klug 2010). The modular structure of ZFPs consists of a zinc finger DNA-binding domain responsible for DNA binding and an effector domain that acts a transcriptional activator or repressor (Klug 2010). To obtain optimal metabolic properties (including stress tolerance), plasmid libraries containing genes encoding artificial transcription factor (ATF) based on ZFP (designated as ZFP-TF in the following text) were constructed and employed as an attractive tool for gTME studies. Each ZFP-TF consists of a randomly shuffled zinc finger DNA-binding domain fused to a regulatory domain (Park et al. 2003, 2005; Lee et al. 2008, 2011, 2013). The zinc finger DNA-binding domain is usually selected on the basis of its ability to recognize various sequences, while the regulatory domain such as Gal4, Ume6, or CRP (cyclic AMP receptor protein) acts as an effector domain. Transformation of Escherichia coli or S. cerevisiae with these ZFP-TF libraries followed by screening would yield stress-tolerant cells from which the ZFP-TF sequences on the expression plasmids could be subsequently identified (Park et al. 2003, 2005; Lee et al. 2008, 2011, 2013). However, it remains a mystery as how a ZFP-TF interacts with the metabolic regulation network of the host cells, especially in the case of S. cerevisiae (Park et al. 2003; Lee et al. 2013). On the other hand, although ZFP-TF libraries have been used to improve heat shock tolerance and drug resistance in S. cerevisiae (Park et al. 2003; Lee et al. 2013), there has been no study looking into the application of ZFP-TF for breeding acid-tolerant yeast strains.
In this study, we obtained acetic-acid-tolerant yeast mutants using a ZFP-TF library based on zinc finger proteins. We also showed that two genes, QDR3 and IKS, which contain the predicted binding sites for a specific ZFP-TF with a repression domain, were indeed related to acetic acid tolerance and deletion of QDR3 improved the ability of S. cerevisiae to produce ethanol in the presence of acetic acid. These studies showed that a synthetic ZFP-TF library can be used to improve the acetic acid tolerance of S. cerevisiae, demonstrating the usefulness of an artificial transcription factor for exploring novel functional genes involved in stress tolerance in S. cerevisiae.
Materials and methods
Strains and culture media
All yeast strains used in this study are listed in Table 1. E. coli DH5α was incubated in Luria-Bertani (LB) medium (10 g/L tryptone, 5 g/L yeast extract, and 10 g/L NaCl) at 37 °C with shaking at 200 rpm. The culture was supplemented with ampicillin (100 μg/mL) when necessary. Yeast strains were maintained on YPD agar medium (10 g/L yeast exact, 20 g/L peptone, and 20 g/L glucose). Solid YPD plates were prepared by adding Bacto agar to a final concentration of 20 g/L. The fermentation medium for Sc4126-M00 and Sc4126-M01 was YPDA, which consisted of 100 g/L glucose, 4 g/L yeast exact, 3 g/L peptone, and 5 g/L acetic acid. Yeast transformants were selected on YPD agar plates containing 200 μg/mL of G418.
DNA manipulations and genetic transformation
DNA isolation, manipulations, and transformation of E. coli DH5α were carried out following standard methods (Green and Sambrook 2012). Plasmid purification and the extraction and purification of DNA from the gel were performed using commercial kits. All polymerase chain reaction (PCR) primers used in this study are listed in Supplementary Table S1.
Construction of yeast strains transformed with a ZFP-TF library and selection of stress-tolerant mutants
The ZFP-TF library used in this study was kindly provided by Professor Jin-Soo Kim in ToolGen Inc., Daejeon, South Korea. The preparation of the ZFP-TF library has been described previously (Park et al. 2003). Yeast cells (strain S288c) were transformed with the ZFP-TF library by electroporation (Amberg et al. 2005), and the transformants were subsequently screened by growth on G418-containing YPD plates. A total of about 10,000 transformants were obtained, which were cultured on YPD plates containing 5 g/L acetic acid. Colonies with larger sizes were picked and again grown on acetic acid containing plates, and those that exhibited the best growth were chosen for further studies.
Tolerance assay with the inhibitor
Analysis of acetic acid tolerance of S. cerevisiae was performed using a plate spot assay. The cells were inoculated in YPD liquid medium and grown at 30 °C for 24 h, and 2 μl of a tenfold diluted culture was spotted onto YPD plates containing 5 g/L acetic acid. Cell growth was examined after 18-h incubation or 2 to 7 days on the YPD plates containing acetic acid.
Determination of CAT, SODs activities, and reduced GSH/GSSG ratios
Catalase (CAT) and superoxide dismutases (SOD) activities as well as glutathione/oxidized glutathione (GSH/GSSH) ratios were measured using the crude extracts prepared from S. cerevisae cells that had been subjected to long-term acetic acid exposure and short-term intensive acetic acid exposure. For long-term exposure, yeast cells were cultured in the presence of 5 g/L acetic acid until they reached the logarithmic growth phase (OD620 0.9), whereas for short-term intense exposure, the cells were collected from a non-acetic acid containing culture at the logarithmic growth phase (OD620 0.9) and then treated with 30 g/L acetic acid for 30 min. Yeast cells grown in YPD liquid medium were used as the control. Yeast cells were harvested by centrifugation 6000×g for 10 min at 4 °C, and cell extract was prepared by violent vortexing with glass beads (MiniBeadbeater-16, BioSpec Products Inc., Bartlesville, OK, USA) in 0.9 % NaCl solution for 60 s, with cooling in an ice bath for 120 s between vortexing (in total six cycles). The extract was centrifuged at 6000×g for 15 min at 4 °C, and the supernatant was collected and subjected to protein assay, CAT and SOD activity assays, and GSH/GSSH ratio measurement. Protein assay was determined with a BCA Protein Assay Kit (Pierce, Rockford, IL, USA). CAT activity was determined with a catalase assay kit according to the manufacturer’s instructions (Nanjing Jiancheng Bioengineering Institute, Nanjing, China) and calculated as previously described (Zheng et al. 2013). The activities of SODs and GSH/GSSG ratios were determined using an SOD assay kit (WST-1 method) and a total glutathione/oxidized glutathione assay kit, respectively, according to the manufacturer’s instructions (Nanjing Jiancheng Bioengineering Institute, Nanjing, China) and calculated as previously described (Zheng et al. 2013).
Prediction of the possible target genes regulated by the ZFP-TF in Sc4126-M01
The conserved binding sequences of the ZFP-TF in strain Sc4126-M01 were revealed to be 5′-GCWAATGAWGTT-3′ (W represents A or T) according to the reference described previously (Park et al. 2003). The possible target genes regulated by the ZFP-TF were identified by BLAST analysis (http://blast.ncbi.nlm.nih.gov/Blast.cgi) using the abovementioned binding sequences.
Real-time quantitative PCR analysis
S. cerevisiae cells grown in YPDA medium at 30 °C and 180 rpm were harvested during the logarithmic phase. Total RNA was isolated from the yeast cells using a TransZol Plant Kit (TransGen Biotech, Beijing, China) and then reversely transcribed into cDNA using a PrimeScript RT Reagent Kit (TaKaRa, Dalian, China) as described by the manufacturer. TransReal-time quantitative PCR (RT-qPCR) used to verify the expression of mRNA was performed according to the method described by Zheng et al. (2013), with ACT1 taken as a reference gene. The primers used are listed in Supplementary Table S1.
Construction of yeast deletion mutants
Based on the results of real-time quantitative PCR analysis, mutants of S. cerevisiae S288c with deletion of YFL040W, QDR3, or IKS1 were constructed by a kanMX4 gene disruption cassette following the method described elsewhere (Baudin et al. 1993). The in-frame deletion regions for YFL040W, QDR3, and IKS1 covered nucleotides +39 to +1584, +39 to +2031, and +39 to +1965, respectively. Yeast transformation was performed by electroporation using the purified PCR products (Amberg et al. 2005). The deletion mutants were verified by diagnostic PCR. The primers used are listed in Supplementary Table S1.
Statistical analysis
All experiments were independently repeated three times, and reproducible results were presented. The results of real-time quantitative PCR, enzyme activities, and fermentation test were expressed as mean and standard deviation (SD). Statistical analysis was performed using Student’s t test, *p < 0.05; **p < 0.01.
Accession number of genes
The GenBank accession number of the coding sequence for the ZFP-TF in this study is JX982113.
Results
Development of acetic acid-tolerant S. cerevisiae strains and analysis of the zinc finger sequence in the tolerant strain
S288c-M01 was one of the acetic-acid-tolerant colonies selected through screening a population of yeast cells transformed with a ZFP-TF library (Fig. S1a in the Supplementary Material). When the expression plasmid (designated as pRS316-M01) containing a ZFP-TF sequence was isolated from S288c-M01 and introduced into S. cerevisiae ATCC4126 (Sc4126), the resulting transformants also exhibited acetic acid tolerance (Fig. S1b in the Supplementary Material). Compared to the control strain (Sc4126-M00), which carried the empty plasmid pRS316-M00, Sc4126-M01 showed remarkably more cell growth on acetic acid-containing medium (Fig. S1b). This indicated that the acetic acid tolerance phenotype could be genetically transferred to another host strain.
To further study the possible targets of ZFP-TF, we analyzed the sequence encoding this protein. As shown in Supplementary Fig. S1c, M01-ZFP is composed of four zinc fingers as follows: NH2-VSTR-VSNV-ISNR-HSSR-COOH. The four groups of letters, VSTR, VSNV, ISNR, or HSSR with capital form, are amino acid residues respectively positioned at the −1, −2, −3, and −6 site of the alpha helix of zinc finger domains (Park et al. 2005). Furthermore, we also identified the putative binding site for this ZFP as 5′-GCWAATGAWGTT-3′ (Park et al. 2003).
Response to acetic acid stress in Sc4126-M01 and Sc4126-M00
To test whether the improved acetic acid tolerance benefits ethanol fermentation under stress conditions, ethanol fermentation by Sc4126-M01 Sc4126-M00 was tested in the presence of acetic acid. We chose to study ethanol production ability of Sc4126-M01 and Sc4126-M00 based on our previous study that industrial yeast strains have better stress tolerance than that the laboratory strain S288c (data not shown). Before starting the fermentation experiments, we first confirmed that the ZFP encoding gene was indeed transcribed. As shown in Fig. S2 in the Supplementary material, transcription of the ZFP-TF encoding gene was evident in Sc4126-M01 grown in the presence of acetic acid due to the presence of the expected transcript, whereas no band was observed in the control strain Sc4126-M00 under the same growth conditions, indicating that no ZFP-TF was expressed. These results demonstrated that the artificial ZFP encoding gene was indeed expressed in Sc4126-M01.
Ethanol fermentation by Sc4126-M01 and Sc4126-M00 was further investigated. Both Sc4126-M01 and Sc4126-M00 showed similar cell density (OD620 is about 3.0) when they were cultured in YPD medium without acetic acid for 24 h (data not shown). When the medium was supplemented with 5 g/L acetic acid, Sc4126-M01 showed a shorter lag phase and higher fermentation rate than Sc4126-M00 (Fig. 1a). Although both Sc4126-M01 and Sc4126-M00 yielded similar final ethanol concentrations when YPDA medium was used, Sc4126-M01 consumed glucose much faster than Sc4126-M00 (Fig. 1b). Interestingly, rapid utilization of acetic acid was observed for Sc4126-M01 after 48 h, which coincided with the end of fermentation and slight increases in growth beyond 48 h (Fig. 1a, b).
Comparison of the fermentation performance of Sc4126-M00 (square) and Sc4126-M01 (diamond) in the presence of 5 g/L acetic acid. Fermentation was performed under anaerobic condition at 30 °C at 150 rpm in a 250-mL Erlenmeyer flask with 100-mL culture medium. a Cell growth and residual acetic acid concentration in the culture supernatant. b Consumption of glucose and changes of ethanol concentration in Sc4126-M00 and Sc4126-M01
It has been reported that when yeast cells are exposed to acetic acid, the activity levels of their antioxidant enzymes would increase (Guaragnella et al. 2008). We thus further investigated the activities of antioxidant enzymes in Sc4126-M01 and Sc4126-M00. As shown in Fig. 2, in the absence of acetic acid, Sc4126-M01 showed similar CAT activity to the control strain Sc4126-M00. However, there were obvious differences in CAT activity between the two strains after long-term treatments with 5 g/L acetic acid or short-term treatments with 30 g/L acetic acid. The CAT activity of Sc4126-M01 was much higher than that of Sc4126-M00, and this would be an advantage for the elimination of reactive oxygen species (ROS) from the cells. In contrast, lower SOD activity was observed for Sc4126-M01 than for the control strain Sc4126-M00 (Fig. S3a in the Supplementary Material), indicating that the activities of SOD and CAT were differently regulated, consistent with the work reported by Semchyshyn et al. (2011). A lower GSH/GSSG ratio was observed for Sc4126-M01 compared to Sc4126-M00 (Fig. S3b in the Supplementary Material), indicating the influence of ZFP-TF on glutathione homeostasis. The results indicated that ZFP-TF might regulate the activities of these antioxidant enzymes, as well as the cellular thiol redox state.
Comparison of catalase (CAT) activity between Sc4126-M00 and Sc4126-M01 in the presence of acetic acid. CAT activity of yeast cells cultivated in the presence of 4.5 g/L acetic acid was measured and compared with that of yeast cells treated with 30 g/L acetic acid for 30 min. Yeast cells grown in YPD medium without addition of acid were used as the control
Prediction of the regulated genes for the synthetic ZFP-TF
The specific DNA-binding sites of the DNA-binding domain of ZFP-TF were predicted based on the ZFP-TF sequence encoded in the plasmid of pRS316-M01 (Park et al. 2003). A total of 12 genes were found to have specific binding sites in their open reading frames (ORFs) or upstream of their ORFs. The proposed functions of these genes are listed in Supplementary Table S2. We focused on determining the transcription levels of three particular genes, namely IKS1, YFL040W, and QDR3, because the functions of these genes in relation to acetic acid tolerance were still unclear. According to the functions annotated for these genes in the Saccharomyces Genome Database (SGD, www.yeastgenome.org), IKS1 encodes a protein kinase of unknown cellular role and it is located downstream of ZAP1, a regulatory gene that responses to the zinc status in the cell and controls the expression of various other genes, including the genes TSA1 and CTT1 that are involved in protecting the cells against oxidative stress (Eide 2009). The putative binding site for ZFP-TF is near the 3′ end of IKS1, inside the ORF of ZAP1 (from +2068 to +2079). YFL040W encodes a putative transporter of the sugar porter family (Palma et al. 2007), whereas the protein product of QDR3 is a multidrug transporter of the major facilitator family (Tenreiro et al. 2005). The putative binding sites of ZFP-TF in these two genes were found inside their ORFs.
Detection of transcription levels of key genes in response to acetic acid
To determine the connection between the possible functional genes and acetic acid tolerance in yeast, the level of IKS1, YFL040W, and QDR3 RNA transcripts in response to acetic acid exposure was measured. In addition, the transcription of ENP2 and HAA1 was also examined. ENP2 encodes the 90S subunit of pre-ribosome and contains a putative binding site for ZFP-TF, and HAA1 is the key regulatory gene for acetic acid tolerance (Fernandes et al. 2005). The results showed that the transcript levels of IKS1, YFL040W, and QDR3 of Sc4126-M01 were decreased in response to acetic acid exposure compared to their expression levels in the control strain Sc4126-M00 (Table 2). In contrast, no detectable change in HAA1 and ENP2 transcripts in the two strains was observed following exposure to acetic acid (data not shown).
Construction of deletion mutants and investigation of their acetic acid tolerance
To further investigate the roles of YFL040W, QDR3, and IKS1 in acetic acid tolerance, we constructed mutants of S288c with deletion of these three genes. The deletion of each gene was confirmed by PCR that used primers specific for the G418-resistant cassette (data not shown). We then examined the cell growth of these mutants grown on agar plates containing 4.5 g/L acetic acid. ΔQDR3 showed increased acetic acid resistance, while ΔYFL040W and ΔIKS1 did not show increased resistance, compared to the wild-type strain S288c (Fig. 3). Considering that acetic acid can induce oxidative stress (Semchyshyn et al. 2011), we also checked the growth of the yeast cells that were cultured in the presence of H2O2. Improved cell growth was observed for Δ QDR3 and Δ IKS1 in the presence of acetic acid and H2O2, respectively (Fig. 3). To check whether the resistant phenotype was due to the different detection methods, we also tried acetic acid shock treatments. After treatment with 30 g/L acetic for 30 min, Δ IKSI showed increased viability and growth recovery compared to the wild-type strain (Fig. 3). These results have proven for the first time that QDR3 and IKS1 were involved in acetic acid tolerance. This study is, therefore, the first report to show that possible target genes in S. cerevisiae that could improve certain phenotype could be identified through the prediction of ZFP-TF-binding sites on these genes.
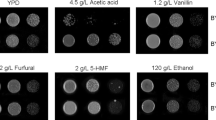
Comparison of acetic acid tolerance and oxidative stress tolerance of the various deletion mutants (S288c-YFL040WΔ, S288c-QDR3Δ, and S288c-IKSIΔ) with that of S288c. Yeast cells were grown on YPD plates supplemented with 4.5 g/L acetic acid or pretreated with 30 g/L acetic acid for 30 min and then cultured on YPD plates without acetic acid supplement. Acetic acid tolerances were tested by comparing their growth rates. In addition, the growth rates of both groups of cells on YPD agar plates containing 6 mM H2O2 were also compared to determine their oxidative stress tolerance. Typical data from one of three independent experiments are shown
Ethanol fermentation by the deletion mutants was also performed in the presence of 3 g/L acetic acid. ΔQDR3 showed higher biomass accumulation and faster sugar utilization than the wild type (Fig. 4a), which is consistent with the result obtained with YPD solid medium supplemented with acetic acid. As for ΔIKSI, lower glucose consumption rate was observed, compared to the wild-type strain (Fig. 4a). Improved ethanol production rate was observed in ΔQDR3 mutant, whereas IKSI deletion resulted in decreased ethanol production rate (Fig. 4b).
Cell growth, glucose consumption, and ethanol production of S288c (circle), S288c-QDR3Δ (square), and S288c-IKSIΔ (triangle) in the presence of 3 g/L acetic acid. The yeast strains were cultivated in YPD liquid medium containing 3 g/L acetic acid, and samples were collected at various time points to determine the biomass accumulation, residual glucose concentration, and ethanol production. Results were obtained from two independent experiments, and the average values are presented. Upper panel shows glucose consumption (closed symbol) and cell growth determined by OD620 values (open symbol). Lower panel shows acetic acid consumption and ethanol production. Ethanol production (closed symbol) and acetic acid concentration (open symbol)
Discussion
ZFP-TF has been used in the metabolic engineering of S. cerevisiae to improve resistance to heat shock and antifungal agents (Park et al. 2003; Lee et al. 2013). Our present work provides the first example of improvement of acetic acid tolerance using a ZFP-TF. More importantly, this is also the first study to demonstrate that deletion of QDR3, IKS1, and YFL040W genes led to higher cell viability in the presence of acetic acid and H2O2. Although in the previous study, genes with the predicted binding sites for ZFP-TF were identified in S. cerevisiae, and none of the predicted target genes shows any relation to the varied phenotypes (Lee et al. 2013). In this study, we showed that the three genes with the predicted binding sites for ZFP-TF were indeed involved in increased stress tolerance, indicating that the function of ZFP-TF could at least be partially attributed to the function of these three genes. These results provided a basis for further exploration of ZFP-TF that may reveal novel functions of genes.
QDR3 encodes a plasma membrane protein (Qdr3p) that has been proposed to act as a multidrug transporter, and its involvement in the efflux of the antimalarial drug quinidine in yeast has been experimentally demonstrated (Tenreiro et al. 2005). Qdr3p also serves as a drug: H+ antiporter that is involved in polyamine homeostasis (Teixeira et al. 2011). However, the transcription level of QDR3 does not change in the presence of the drugs, indicating that its product might have other specific physiological substrates, and the drugs might be transported opportunistically (Tenreiro et al. 2005). The disruption of the FPS1 aquaglyceroporin gene was previously reported to improve acetic acid tolerance and ethanol fermentation performance of a S. cerevisiae strain (Zhang et al. 2011). Fps1p facilitates the diffusion of acetic acid into the cell (Mollapour and Piper 2007), which leads to intracellular acidification and toxicity. It would be interesting to check whether QDR3 also functions in the uptake of acetic acid. Although we could not prove the direct binding of ZFP-TF to the putative binding region of QDR3 using chromatin immunoprecipitation (ChIP), we could still conclude that ZFP-TF may interact with QDR3 directly or indirectly to decrease its transcription, which would then result in improved cell growth in the presence of acetic acid. The putative binding site of ZFP-TF on QDR3 was found within its ORF, which is a very rare event for S. cerevisiae. Nevertheless, a similar finding has previously been reported by Nishizawa et al. (2008), who showed that the binding of Pho4p to the ORF of KCS1 provokes the translation of a truncated Kcs1p, which functions in the phosphate signaling pathway. It could also be possible that QDR3 expression was indirectly regulated by ZFP-TF. Further studies are needed to explore how ZFP-TF regulates QDR3 expression.
IKS1 encodes a protein kinase of unknown cellular role, the expression of which is induced by mild heat stress (Sakaki et al. 2003). Deletion of IKS1 in yeast has led to hypersensitivity to copper sulfate (Rieger et al. 1999) and a sorbate-resistant phenotype (Mollapour et al. 2004). We deduced that IKS1 may be involved in the regulation of key pathways in acute stress defense. Although deletion of IKS1 led to improved cell viability in response to the shock treatment with high concentration of acetic acid, the deletion mutant displayed reduced ability to grow and ferment glucose in liquid culture, implying that IKS1 and QDR3 may play different roles in the defense against acetic acid stress. It will be interesting to further investigate the mechanisms of QDR3 and IKS1 deletion in yeast stress tolerance.
The consumption of acetic acid by Sc4126-M01 was rather interesting. The same consumption profile was not observed for the QDR3-deletion mutant, indicating that the ZFP-TF, but not Qdr3p, may have multiple functions in the regulation of cell metabolism. The decreased acetic acid concentration in Sc4126-M01 coincided with the improved cell viability with slight increases in growth beyond 48 h. Acetic acid can be converted to acetyl-coA by acetyl-coA synthetases, ACS1, and ACS2 (van den Berg et al. 1996). It is still not clear whether ZFP-TF functions in elevating the activity of acetyl-coA synthetase(s). Engineering of global transcriptional machinery using ZFP-TF would have a promising advantage in that a broader phenotypic space can be achieved using the heterologous transcription factor.
ROS can be produced by acetic acid treatment, which may result from intracellular acidification (Guaragnella et al. 2011). We also observed ROS production when a high concentration of acetic acid (10 g/L) was added to the culture medium (data not shown). Cells have developed various strategies for defense against ROS, and SODs and catalases are well known for their functions in the enzymatic detoxification of ROS (Herrero et al. 2008). SODs convert superoxide anion to hydrogen peroxide, whereas peroxisomal catalase detoxifies hydrogen peroxide by converting it to water and oxygen (Herrero et al. 2008). Previous studies by other investigators only focused on the perturbation of global gene expression by ZFP-TF (Park et al. 2003, 2005; Lee et al. 2008, 2011), but regulation of cellular metabolism can also be achieved at the levels of post-transcription and/or translation. We found that the activities of antioxidant enzymes (SOD and CAT) as well as the glutathione redox status also changed by the overexpression of ZFP-TF, indicating the direct or indirect effect of the artificial ZFP on cellular oxidative stress tolerance. Further studies are necessary to provide a clearer picture on the function of ZFP-TF in the remodeling of the cellular metabolic network. The results present in this study would provide a basis for further development of robust yeast strains for ethanol fuel production using gTME methods that involve ZFP-TF.
References
Almeida JR, Runquist D, Sànchez i Nogué V, Lidén G, Gorwa-Grauslund MF (2011) Stress-related challenges in pentose fermentation to ethanol by the yeast Saccharomyces cerevisiae. Biotechnol J 6:286–299
Alper H, Moxley J, Nevoigt E, Fink GR, Stephanopoulos G (2006) Engineering yeast transcription machinery for improved ethanol tolerance and production. Science 314:1565–1568
Amberg DC, Burke DJ, Strathern JN (2005) Methods in yeast genetics: a Cold Spring Harbor Laboratory course material. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Baudin A, Ozier-Kalogeropoulos O, Denouel A, Lacroute F, Cullin C (1993) A simple and efficient method for direct gene deletion in Saccharomyces cerevisiae. Nucleic Acids Res 21:3329
Demeke MM, Dumortier F, Li Y, Broeckx T, Foulquié-Moreno MR, Thevelein JM (2013) Combining inhibitor tolerance and D-xylose fermentation in industrial Saccharomyces cerevisiae for efficient lignocellulose-based bioethanol production. Biotechnol Biofuels 6:120
Ding J, Bierma J, Smith MR, Poliner E, Wolfe C, Hadduck AN, Bakalinsky AT (2013) Acetic acid inhibits nutrient uptake in Saccharomyces cerevisiae: auxotrophy confounds the use of yeast deletion libraries for strain improvement. Appl Microbiol Biotechnol 97:7405–7416
Eide DJ (2009) Homeostatic and adaptive responses to zinc deficiency in Saccharomyces cerevisiae. J Biol Chem 284:18565–18569
Fernandes AR, Mira NP, Vargas RC, Canelhas I, Sá-Correia I (2005) Saccharomyces cerevisiae adaptation to weak acids involves the transcription factor Haa1p and Haa1p-regulated genes. Biochem Biophys Res Commun 337:95–103
Green MR, Sambrook J (2012) Molecular cloning: a laboratory manual. Harbor Laboratory Press, Cold Spring Harbor
Guaragnella N, Antonacci L, Giannattasio S, Marra E, Passarella S (2008) Catalase T and Cu, Zn-superoxide dismutase in the aceticacid-induced programmed cell death in Saccharomyces cerevisiae. FEBS Lett 582:210–214
Guaragnella N, Antonacci L, Passarella S, Marra E, Giannattasio S (2011) Achievements and perspectives in yeast acetic acid-induced programmed cell death pathways. Biochem Soc Trans 39(5):1538–1543
Hasunuma T, Kondo A (2012) Development of yeast cell factories for consolidated bioprocessing of lignocellulose to bioethanol through cell surface engineering. Biotechnol Adv 30:1207–1218
Herrero E, Ros J, Bellí G, Cabiscol E (2008) Redox control and oxidative stress in yeast cells. Biochim Biophys Acta 1780(11):1217–1235
Jönsson LJ, Alriksson B, Nilvebrant N (2013) Bioconversion of lignocellulose: inhibitors and detoxification. Biotechnol Biofuels 6:16
Kitanovic A, Bonowski F, Heigwer F, Ruoff P, Kitanovic I, Ungewiss C, Wölfl S (2012) Acetic acid treatment in S. cerevisiae creates significant energy deficiency and nutrient starvation that is dependent on the activity of the mitochondrial transcriptional complex Hap2-3-4-5. Front Oncol 2:118
Klug A (2010) The discovery of zinc fingers and their development for practical applications in gene regulation and genome manipulation. Q Rev Biophys 43:1–21
Lee JY, Sung BH, Yu BJ, Lee JH, Lee SH, Kim MS, Koob MD, Kim MC (2008) Phenotypic engineering by reprogramming gene transcription using novel artificial transcription factors in Escherichia coli. Nucleic Acids Res 36:1–10
Lee JY, Yang KS, Jang SA, Sung BH, Kim SC (2011) Engineering butanol-tolerance in Escherichia coli with artificial transcription factor libraries. Biotechnol Bioeng 108:742–749
Lee SW, Kim E, Kim JS, Oh MK (2013) Artificial transcription regulator as a tool for improvement of cellular property in Saccharomyces cerevisiae. Chem Eng Sci 103:42–49
Lv YJ, Wang X, Ma Q, Bai X, Li BZ, Zhang W, Yuan YJ (2014) Proteomic analysis reveals complex metabolic regulation in Saccharomyces cerevisiae cells against multiple inhibitors stress. Appl Microbiol Biotechnol 98:2207–2221
Mollapour M, Piper PW (2007) Hog1 mitogen-activated protein kinase phosphorylation targets the yeast Fps1 aquaglyceroporin for endocytosis, thereby rendering cells resistant to acetic acid. Mol Cell Biol 27:6446–6456
Mollapour M, Fong D, Balakrishnan K, Harris N, Thompson S, Schüller C, Kuchler K, Piper PW (2004) Screening the yeast deletant mutant collection for hypersensitivity and hyper-resistance to sorbate, a weak organic acid food preservative. Yeast 21(11):927–946
Nicolaou SA, Gaida SM, Papoutsakis ET (2010) A comparative view of metabolite and substrate stress and tolerance in microbial bioprocessing: from biofuels and chemicals, to biocatalysis and bioremediation. Metab Eng 12:307–331
Nishizawa M, Komai T, Katou Y, Shirahige K, Ito T, Toh-E A (2008) Nutrient-regulated antisense and intragenic RNAs modulate a signal transduction pathway in yeast. PLoS Biol 6:2817–2830
Palma M, Goffeau A, Spencer-Martins I, Baret PV (2007) A phylogenetic analysis of the sugar porters in hemiascomycetous yeasts. J Mol Microbiol Biotechnol 12:241–248
Park KS, Lee DK, Lee H, Lee Y, Jang YS, Kim YH, Kim JS (2003) Phenotypic alteration of eukaryotic cells using randomized libraries of artificial transcription factors. Nat Biotechnol 21:1208–1214
Park KS, Jang YS, Le H, Kim JS (2005) Phenotypic alteration and target gene identification using combinatorial libraries of zinc finger proteins in prokaryotic cells. J Bacteriol 187:5496–5499
Rieger KJ, El-Alama M, Stein G, Bradshaw C, Slonimski PP, Maundrell K (1999) Chemotyping of yeast mutants using robotics. Yeast 15(10B):973–986
Sakaki K, Tashiro K, Kuhara S, Mihara K (2003) Response of genes associated with mitochondrial function to mild heat stress in yeast Saccharomyces cerevisiae. J Biochem 134(3):373–384
Semchyshyn HM, Abrat OB, Miedzobrodzki J, Inoue Y, Lushchak VI (2011) Acetate but not propionate induces oxidative stress in bakers’ yeast Saccharomyces cerevisiae. Redox Rep 16:15–23
Stratford M, Steels H, Nebe-von-Caron G, Novodvorska M, Hayer K, Archer DB (2013) Extreme resistance to weak-acid preservatives in the spoilage yeast Zygosaccharomyces bailii. Int J Food Microbiol 166:126–134
Teixeira MC, Cabrito TR, Hanif ZM, Vargas RC, Tenreiro S, Sá-Correia I (2011) Yeast response and tolerance to polyamine toxicity involving the drug: H+ antiporter Qdr3 and the transcription factors Yap1 and Gcn4. Microbiology 157:945–995
Tenreiro S, Vargas RC, Teixeira MC, Magnani V, Sá-Correia I (2005) The yeast multidrug transporter Qdr3 (Ybr043c): localization and role as a determinant of resistance to quinidine, barban, cisplatin, and bleomycin. Biochem Biophys Res Commun 327:952–959
Ullah A, Orij R, Brul S, Smits GJ (2012) Quantitative analysis of the modes of growth inhibition by weak organic acids in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 78:8377–8387
van den Berg MA, de Jong-Gubbels P, Kortland CJ, van Dijken JP, Pronk JT, Steensma HY (1996) The two acetyl-coenzyme A synthetases of Saccharomyces cerevisiae differ with respect to kinetic properties and transcriptional regulation. J Bio Chem 271(46):28953–28959
Woo JM, Yang KM, Kim SU, Blank LM, Park JB (2014) High temperature stimulates acetic acid accumulation and enhances the growth inhibition and ethanol production by Saccharomyces cerevisiae under fermenting conditions. Appl Microbiol Biotechnol 98:1–10
Yang J, Bae JY, Lee YM, Kwon H, Moon HY, Kang HA, Yee SB, Kim W, Choi W (2011) Construction of Saccharomyces cerevisiae strains with enhanced ethanol tolerance by mutagenesis of the TATA-binding protein gene and identification of novel genes associated with ethanol tolerance. Biotechnol Bioeng 108:1776–1787
Zhang JG, Liu XY, He XP, Guo XN, Lu Y, Zhang BR (2011) Improvement of acetic acid tolerance and fermentation performance of Saccharomyces cerevisiae by disruption of the FPS1 aquaglyceroporin gene. Biotechnol Lett 33:277–284
Zhao XQ, Bai FW (2009) Mechanisms of yeast stress tolerance and its manipulation for efficient fuel ethanol production. J Biotechnol 144:23–30
Zheng DQ, Liu TZ, Chen J, Zhang K, Li O, Zhu L, Zhao YH, Wu XC, Wang PM (2013) Comparative functional genomics to reveal the molecular basis of phenotypic diversities and guide the genetic breeding of industrial yeast strains. Appl Microbiol Biotechnol 97:2067–2076
Acknowledgments
This work was supported by financial support from the National Science Foundation of China (No. 21376043), National High Technology Research and Development Program of China (863 Program, No. 2012AA101805, 2012AA021205), and Program for New Century Excellent Talents, Ministry of Education, China (No. NCET-11-0057). We appreciate the kind help of Dr. Jin-Soo Kim in ToolGen, Inc., South Korea for donating the artificial zinc finger protein (ZFP) library. We also thank Dr. Alan K Chang for improving the language of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1154 kb)
Rights and permissions
About this article
Cite this article
Ma, C., Wei, X., Sun, C. et al. Improvement of acetic acid tolerance of Saccharomyces cerevisiae using a zinc-finger-based artificial transcription factor and identification of novel genes involved in acetic acid tolerance. Appl Microbiol Biotechnol 99, 2441–2449 (2015). https://doi.org/10.1007/s00253-014-6343-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-6343-x