Abstract
Gastric cancer (GC) is the third leading cause of global cancer morbidity and mortality. One of the significant challenges in GC treatment is that most GC patients are diagnosed with advanced-stage disease due to the lack of suitable biomarkers. Recent studies have shown that microRNAs (miRNAs) can acts as a potential biomarker in GC diagnosis and prognosis. I performed a systematic review of published miRNA studies in GC, which includes the miRNA expression profiles between GC tissues and normal tissues and also miRNA studies to evaluate their potential value in the diagnosis and prognosis of GC. Among the studies, upregulation of miR-21, miR-106b, miR-25, miR-214, miR-18a, miR-191, and miR-93 and downregulation of miR-375, miR-148a, miR-92, miR-155, and miR-564 were observed in GC tissues. In evaluating of diagnosis value of miRNAs, the study was performed on a combined miRNA include miR-21, miR-93, miR-106a, and miR-106b indicated the panel of these miRNAs have the highest AUC 0.887 to discriminate GC patients from healthy. Also, miR-940 with a sensitivity of 81.25% and specificity of 98.57% may be used for diagnostic biomarkers for GC. Finally, the pooled prognostic result of miR-21 for hazard ratios (HR) was 1.260 (95% CI 0.370–4.330, P < 0.001), showing that miR-21 could predict poor survival in GC patients. This systematic review can confirm that we need to find a miRNA or a panel of miRNAs with high sensitivity and specificity for further exploration to investigate a better diagnostic or therapeutic tool for personalized management of GC patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer (GC) remains one of the most lethal digestive malignancies and is the fourth most common cancer, with an estimated 989,600 new cases and 738,000 deaths globally in 2008 (Jemal et al. 2011; Torre et al. 2015). GC has no remarkable symptoms, and it is usually found in the advanced stage; therefore, only a few patients were cured. Due to the lack of sensitivity and specificity, the classical biomarkers such as carcinoembryonic antigen (CEA) and carbohydrate antigen 19.9 (CA19.9) cannot be used as a potential biomarker to diagnose or prognosis of GC (Zhang et al. 2012; Cui et al. 2013; Jiexia et al. 2013; Yu et al. 2013; Shao et al. 2016).
In recent years, an increasing number of non-coding RNAs (ncRNAs) containing microRNAs (miRNAs) and long non-coding RNAs (lncRNAs) have been proven to have great potential clinical value in GC. Recent studies have indicated the pivotal role of miRNAs in regulating biological processes such as proliferation, cellular differentiation, apoptosis, and gene regulation (Alvarez-Garcia and Miska 2005; Esquela-Kerscher and Slack 2006). Moreover, miRNAs are dysregulated in many cancers, and miRNA expression profiling has shown that specific miRNAs are associated with cancer development and progression.
Several studies indicated the several aspects of miRNAs interacting with multiple target genes and pathways, making them a potential biomarker for clinical diagnostics. Moreover, miRNAs could maintain its stability in plasma, urine, and saliva (Mitchell et al. 2008). Also, the classifications of miRNAs can investigate tissue source in the cancers of uncertain primaries. Additionally, dysregulation of miRNAs could be observed during the sequential cancer pattern, including the early stage, during progression, and after metastasis. Thus, miRNAs may act as favorable clinical biomarkers for distinguishing tumors and selection of therapeutic approach.
Several studies have been conducted to find a biomarker by identifying the differential expression of miRNAs between GC and normal samples (Guo et al. 2009; Tchernitsa et al. 2010; Ueda et al. 2010; Oh et al. 2011; Saito et al. 2013). Therefore, they are good candidates for diagnostic, prognostic, and predictive biomarkers (Iorio and Croce 2012).
In this study, I conducted a systematic review to identify the differential expression of miRNAs consistently reported in GC to find the miRNAs that could act as a potential biomarker in GC diagnosis and prognosis.
Materials and methods
Search strategy
I searched in the PubMed online database for published articles up to September 1, 2020, with the following keywords: microRNA,” “miRNA,” “gastric cancer,” “expression profiling,” “prognosis,” “diagnosis,” “biomarker,” and “survival.” The data were then extracted from the selected studies and input into tables that contain different characteristics of interest.
Selection criteria
The inclusion criteria were as follows: (1) studies had to be miRNA profiling studies in GC patients and reported on dysregulated miRNAs; (2) studies that investigated the diagnosis value of miRNAs to discriminate GC patients from normal; (3) studies that investigated the association between miRNAs expression and survival outcome and that provided a hazard ratio (HR) and a 95% confidence interval (CI), while the exclusion criteria were as follows: (1) non-English and non-human subject studies; (2) abstracts, reviews, comments, and letters; (3) studies with incomplete or insufficient data.
Data extraction
All the articles were filtered three times, and then the suitable studies were extracted. According to the inclusion and exclusion criteria, the data about GC patients, total samples used for the study on diagnosis value of miRNAs in serum or plasma and platform used in these studies, fold change of statistically differentially expressed miRNAs in GC tissues have been provided. The list of upregulated and downregulated miRNAs was summarized in (Tables 1 and 2).
Ranking of miRNA
In this study, I used the vote-counting strategy-based method developed by Griffith and Chan (Griffith et al. 2006; Chan et al. 2008) to rank miRNAs as potential molecular markers. MiRNAs were ranked based on criteria as (1) the number of studies in agreement, reporting miRNA deregulation with statistical significance, as well as the direction of deregulation (upregulated or downregulated), (2) the miRNA frequency was reported in the studies, and (3) the mean fold change (FC) of each miRNA reported by the studies in agreement.
Results
A total of 320 studies were identified from PubMed using my search strategy. A total of 183 studies were retrieved after screening the titles and abstracts, including irrelevant studies, reviews, and studies that are not a human or English study. Finally, I found that 105 articles lacked sufficient data or were not relevant to diagnoses or prognoses. Therefore, 32 studies, including 14 miRNAs studies for analysis of differential expression of miRNAs in GC tissues, 10 for evaluating the diagnostic value of miRNA expression in plasma or serum to discriminate GC patients from healthy, and 8 for predictive analysis of miRNAs, were included in the analysis. A flow diagram of the study selection process is presented in Fig. 1.
Dysregulation of miRNAs in GC tissues
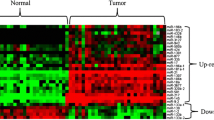
Among differentially expressed miRNAs, the upregulation of miR-21 was reported in four studies, followed by upregulation of miR-106b, miR-25, and miR-214 in six, six, and two studies, respectively. Moreover, the downregulation of miR-375 and miR-148a and upregulation of miR-18a and miR-93 were reported in three studies, followed by two downregulated miRNAs including miR-155 and miR-564 in two and one studies, respectively. I have also summarized miRNAs that were upregulated and downregulated (Table 1).
Diagnostic value of miRNAs in GC
A total of 8 miRNA studies have indicated their potential value in the diagnosis of GC. Its sensitivity ranges from 65.40 to 90%, and specificity ranges from 58 to 100.00%. Among them, nine studies were performed on a single miRNA, and one was performed on a combined miRNA include miR-21, miR-93, miR-106a, and miR-106b; the list of these studies is summarized in Table 2. The sensitivity of one miRNA was higher than 90%, and the specificity of the three miRNAs was higher than 90%. The highest sensitivity was 90% for miR-421 (Wu et al. 2015a), while the highest specificity was 100% for miR-200c (Valladares-Ayerbes et al. 2012). In the study of Liu et al. (2016), the expression level of miR-940 was reduced using a sample of 5 patients, and subsequently, a controlled study using 80 patients confirmed that the AUC reached 0.966, which is the highest in the current study. In the research that was conducted by Zhao et al. (2018), they determined the diagnostic value of a panel of miRNAs include miR-21, miR-93, miR-106a, and miR-106b, and they verified the sensitivity and specificity of the combination were improved compared with the four when independent, with an AUC of 0.887 when combined.
MiRNAs act as a prognosis biomarker in GC
In this study, I used the hazard ratios (HRs) or relative risks (RRs) for each miRNA. The pooled results indicated that miR-21 were significant prognostic biomarkers for GC patients (HR = 1.260, 95% CI 0.370–4.330, P < 0.001). Also, the HR of miR-148 was statistically significant, with a value of 3.076 (95% CI 1.216–8.776; P < 0.017). The significant HR value of two miRNAs includes miR-200c and miR-204, which were found with the HR value of 2.240 (95% CI 1.091–4.614; P = 0.028) and HR value of 3.900 (95% CI 1.300–11.800; P = 0.017), respectively. Additionally, in the list of studies summarized in Table 3, the association between miRNA expression and RR was observed. The highest RR value was for miR-15a with an RR of 1.950, these studies followed by the RR value for miR-125a-3p, miR-106b, and miR-20a, which includes RR = 1.800, 1.600, and 1.110, respectively.
Discussion
In the translational aspect of using miRNAs in cancer, there has been an evolution from profiling altered miRNA expressions in cancers to clinical trials with miRNAs as therapy in the last decade. Recent studies have shown that its promise with miRNA replacement therapy in pre-clinical and clinical pilot studies and its potential use in personalized medicine. These studies have also explored a range of applications from diagnostics, prognostics, disease surveillance to primary therapy, or a tool to sensitize patients to treatment modalities such as chemotherapy and radiotherapy. Advancements in delivering miRNAs, from viral vectors and liposomal delivery to nanoparticle-based, have led to several pre-clinical and clinical applications for miRNA cancer therapeutics (Kwok et al. 2017).
In this systematic review, I conducted a systematic search to retrieve data from studies that explored aberrantly expressed miRNAs as candidate biomarkers for GC diagnosis and prognosis, using either tissue samples or blood samples. With regard to tissue-diagnostic miRNAs, I identified eight consistently upregulated miRNAs (miR-21, miR-223, miR-18a, miR-214, miR-93, miR-191, miR-25, miR-106b) and five miRNAs consistently downregulated (miR-375, miR-564, miR-155, miR-148a, miR-92) (Table 1). MiR-21 was most consistently upregulated, with a differential expression reported among four studies and a median fold change of 4.05. However, miR-21 is vastly cited across the literature for this upregulation in GC. I found only these four studies matching the inclusion criteria that had reported significant upregulation of miR-21.
Furthermore, the upregulation of miR-21 significantly promoted cell proliferation. It revealed a higher proportion of cells at the S phase, and the knockdown of miR-21 expression resulted in a cell-cycle arrest at the G2/M phase and inhibited cell proliferation (Zhong et al. 2012). Additionally, Zhang et al. (2012), De Val and Black (2009), and Yamanaka et al. (2012) demonstrated that miR-21, as an oncomir, contributes to GC progression by inhibiting apoptosis and elevating cell proliferation by targeting the tumor suppressor gene RECK. Moreover, the highest fold change for downregulated miRNAs was found for miR-92 with a median fold change of 2.80. Additionally, in this systematic review, ten diagnostic and eight prognostic studies were included to study whether miRNAs are useful biomarkers for GC. The data showed that specific miRNAs could be used (with moderate sensitivity and specificity) in the diagnosis of GC. The study’s result related to miR-940 showed that the sensitivity and specificity were 81.25% and 98.57%, respectively (Liu et al. 2016), which indicate the potential act of this miRNA as a biomarker in GC diagnosis. Also, a recent study has shown the dysregulation of miR-940 in stage IV of GC patients, and CD276 as a target gene of this miRNA played a significant role in promoting migration and invasion of GC cells (Liu et al. 2016). Also, miR-940 has been proven to promote tumor cell invasion and metastasis with the aid of interacting with ZNF24 in GC (Liu et al. 2015), and plasma miR-940 decreased during gastric carcinogenesis. This miRNA might be applied as a novel biomarker for the diagnosis of GC.
Moreover, the combined diagnostic AUC, sensitivity, and specificity of miR-21, miR-93, miR-106a, and miR-106b in plasma were 0.887, 84.80%, and 79.20%, respectively, for discriminating GC cases. With this combination, the diagnostic efficiency for the early-stage gastric non-cardia adenocarcinoma (GNCA) increased significantly (Zhu et al. 2014). As supporting evidence, a recent study that analyzed the expression level of miR-21, miR-93, miR-106a, and miR-106b in GC samples using ddPCR, the results have shown the association between the increased levels of these miRNAs with advanced TNM stage (Tchernitsa et al. 2010; Shiotani et al. 2013). Due to the overexpression of miR-25, miR-93, and miR-106b in GC stem cells, the functional analysis of these miRNAs might also be essential for GCs diagnosis (Yu et al. 2014). A previous study verified that upregulation of miR-25 could induce cell apoptosis via targeting gene BIM. MiR-106b and miR-93 abrogate TGFβ prompted apoptosis in GC cells by targeting the expression of BIM, encoding the pro-apoptotic protein BCL2-like 11, thereby preventing apoptosis and leads to tumor progression. Also, recent studies indicated that the differential expression of miR-335 had been used as a signature for GC diagnosis, and the expression level of this miRNA is correlated significantly with LNM, distant metastasis, and TNM stage (Li et al. 2011; Ahadi and Safavi 2019). In the next step, I listed the studies on miRNAs potential to act as a prognosis biomarker using HRs and RRs to determine each miRNA overall prognostic performance. The analysis indicated a closer relationship between miR-204 expression and poor survival in GC patients (HR = 3.900, 95% CI 1.300–11.800), and this miRNA can be applied to monitor the therapeutic effects. The upregulation of miR-204 is related to GC cell invasion and epithelial-mesenchymal transition (EMT) by targeting SIRT-1 at the post-transcriptional level. Thus, miR-204 is believed to play a crucial role in regulating the metastasis of GC. Therefore, miR-204 can increase GC cells responsiveness to 5-fluorouracil and oxaliplatin treatment by targeting Bcl-2, indicating that miR-204 might be a therapeutic target for improving GC prognosis (Canu 2012). The results also revealed that miR-15a yielded worse overall survival in GC (RR = 1.950, 95% CI 0.470–9.130). The downregulation of miR-15a induces cell proliferation, EMT, migration, and invasion by Twist1 and YAP1 as the target genes (Wang et al. 2017).
In conclusion, from this systematic review study, I identified that miR-940 and the combination of miR-21, miR-93, miR-106a, and miR-106b with the highest AUC in discriminating of GC patient could act as a diagnostic biomarker, and miR-204 and miR-15a with the highest HR and RR are closely associated to poor survival in GC which can be determined as a prognosis biomarker for further assessment. The study found several promising miRNAs that had been consistently reported. However, more investigations are needed for the clinical studies focusing on these miRNAs to understand these miRNAs potential roles in GC. One of the significant factors limiting the use of miRNAs as a diagnostic tool in clinical settings is associated with the fact that frequently reported miRNA biomarkers are detected in patients with different tumor types. Moreover, other limitations of using miRNAs as a diagnostic biomarker include the range of concentrations in body fluids and modulation depending on various parameters (age, gender, health/disease) that are not yet clearly established (Gillespie et al. 2019). In the future, a miRNA or a miRNA signature should be a better diagnostic or therapeutic tool than a single gene. Ultimately, the personalized management, diagnosis, and prognosis of the disorder can be finished using a panel of miRNAs (Ahadi 2020).
References
Ahadi A (2020) Dysregulation of miRNAs as a signature for diagnosis and prognosis of gastric cancer and their involvement in the mechanism underlying gastric carcinogenesis and progression. IUBMB Life 72(5):884–898
Ahadi A, Safavi MS (2019) miR-335-5p has an important role in the progression of gastric cancer by down-regulation of CEACAM5. Meta Gene 19:65–68
Alvarez-Garcia I, Miska EA (2005) MicroRNA functions in animal development and human disease. Development 132(21):4653–4662
Canu V, Mori F, Di Benedetto A, Lorenzon L, Ercolani C, Di Agostino S, Cambria AM, Germoni S, SABF (2012) miR-204 targets Bcl-2 expression and enhances responsiveness of gastric cancer. Cell Death and Disease 3:e423
Chan SK, Griffith OL, Tai IT, Jones SJ (2008) Meta-analysis of colorectal cancer gene expression profiling studies identifies consistently reported candidate biomarkers. Cancer Epidemiol Prev Biomarkers 17(3):543–552
Chang L, Guo F, Huo B, Lv Y, Wang Y, Liu W (2015) Expression and clinical significance of the microRNA-200 family in gastric cancer. Oncology letters 9(5):2317–2324
Cui L, Lou Y, Zhang X, Zhou H, Deng H, Song H, Yu X, Xiao B, Wang W, Guo J (2011) Detection of circulating tumor cells in peripheral blood from patients with gastric cancer using piRNAs as markers. ClinBiochem 44(13):1050–1057
Cui L, Zhang X, Ye G, Zheng T, Song H, Deng H, Xiao B, Xia T, Yu X, Le Y (2013) Gastric juice MicroRNAs as potential biomarkers for the screening of gastric cancer. Cancer 119(9):1618–1626
De Val S, Black BL (2009) Transcriptional control of endothelial cell development. Dev Cell 16(2):180–195
Ding L, Xu Y, Zhang W, Deng Y, Si M, Du Y, Yao H, Liu X, Ke Y, Si J (2010) MiR-375 frequently downregulated in gastric cancer inhibits cell proliferation by targeting JAK2. Cell Res 20(7):784–793
Esquela-Kerscher A, Slack FJ (2006) Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 6(4):259–269
Gillespie P, Ladame S, O’Hare D (2019) Molecular methods in electrochemical microRNA detection. Analyst 144(1):114–129
Griffith OL, Melck A, Jones SJ, Wiseman SM (2006) Meta-analysis and meta-review of thyroid cancer gene expression profiling studies identifies important diagnostic biomarkers. J ClinOncol 24(31):5043–5051
Guan H, Li W, Li Y, Wang J, Li Y, Tang Y, Lu S (2017) MicroRNA-93 promotes proliferation and metastasis of gastric cancer via targeting TIMP2. PLoS ONE 12(12):e0189490
Guo J, Miao Y, Xiao B, Huan R, Jiang Z, Meng D, Wang Y (2009) Differential expression of microRNA species in human gastric cancer versus non-tumorous tissues. J GastroenterolHepatol 24(4):652–657
Iorio MV, Croce CM (2012) MicroRNA dysregulation in cancer: diagnostics, monitoring and therapeutics. A comprehensive review. EMBO molecular medicine 4(3):143–159
Jemal, A., F. Bray, M. M. Center, J. Ferlay, E. Ward and D. Forman (2011). Global cancer statistics. CA: a cancer journal for clinicians 61(2): 69–90.
Jiexian J, Xiaoqin X, Lili D, Baoguo T, Ting S, Xianwen Z, Cunzhi H (2013) Clinical assessment and prognostic evaluation of tumor markers in patients with gastric cancer. Int J Biol Marker 28(2):192–200
Kim CH, Kim HK, Rettig RL, Kim J, Lee ET, Aprelikova O, Choi IJ, Munroe DJ, Green JE (2011) miRNA signature associated with outcome of gastric cancer patients following chemotherapy. BMC Med Genomics 4(1):79
Kong Y, Ning L, Qiu F, Yu Q, Cao B (2019) Clinical significance of serum miR-25 as a diagnostic and prognostic biomarker in human gastric cancer. Cancer Biomark 24(4):477–483
Kwok GT, Zhao JT, Weiss J, Mugridge N, Brahmbhatt H, MacDiarmid JA, Robinson BG, Sidhu SB (2017) Translational applications of microRNAs in cancer, and therapeutic implications. Non-coding RNA research 2(3–4):143–150
Li B-S, Zhao Y-L, Guo G, Li W, Zhu E-D, Luo X, Mao X-H, Zou Q-M, Yu P-W, Zuo Q-F (2012) Plasma microRNAs, miR-223, miR-21 and miR-218, as novel potential biomarkers for gastric cancer detection. PLoS ONE 7(7):e41629
Li X, Luo F, Li Q, Xu M, Feng D, Zhang G, Wu W (2011) Identification of new aberrantly expressed miRNAs in intestinal-type gastric cancer and its clinical significance. Oncol Rep 26(6):1431–1439
Liu X, Ge X, Zhang Z, Zhang X, Chang J, Wu Z, Tang W, Gan L, Sun M, Li J (2015) MicroRNA-940 promotes tumor cell invasion and metastasis by downregulating ZNF24 in gastric cancer. Oncotarget 6(28):25418
Liu X, Kwong A, Sihoe A, Chu K-M (2016) Plasma miR-940 may serve as a novel biomarker for gastric cancer. Tumor Biology 37(3):3589–3597
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O’Briant KC, Allen A (2008) Circulating microRNAs as stable blood-based markers for cancer detection. ProcNatlAcadSci 105(30):10513–10518
Naito Y, Yasuno K, Tagawa H, Sakamoto N, Oue N, Yashiro M, Sentani K, Goto K, Shinmei S, Oo HZ (2014) MicroRNA-145 is a potential prognostic factor of scirrhous type gastric cancer. Oncol Rep 32(4):1720–1726
Oh H-K, Tan AL-K, Das K, Ooi C-H, Deng N-T, Tan IB, Beillard E, Lee J, Ramnarayanan K, Rha S-Y (2011) Genomic loss of miR-486 regulates tumor progression and the OLFM4 antiapoptotic factor in gastric cancer. Clin Cancer Res 17(9):2657–2667
Saito Y, Suzuki H, Imaeda H, Matsuzaki J, Hirata K, Tsugawa H, Hibino S, Kanai Y, Saito H, Hibi T (2013) The tumor suppressor microRNA-29c is downregulated and restored by celecoxib in human gastric cancer cells. Int J Cancer 132(8):1751–1760
Shao J, Fang P-H, He B, Guo L-L, Shi M-Y, Zhu Y, Bo P (2016) Downregulated microRNA-133a in gastric juice as a clinicopathological biomarker for gastric cancer screening. Asian Pac J Cancer Prev 17(5):2719–2722
Shin J-Y, Kim Y-I, Cho S-J, Lee MK, Kook M-C, Lee JH, Lee SS, Ashktorab H, Smoot DT, Ryu KW (2014) MicroRNA 135a suppresses lymph node metastasis through down-regulation of ROCK1 in early gastric cancer. PLoS ONE 9(1):e85205
Shiotani A, Murao T, Kimura Y, Matsumoto H, Kamada T, Kusunoki H, Inoue K, Uedo N, Iishi H, Haruma K (2013) Identification of serum miRNAs as novel non-invasive biomarkers for detection of high risk for early gastric cancer. Br J Cancer 109(9):2323–2330
Tchernitsa O, Kasajima A, Schäfer R, Kuban RJ, Ungethüm U, Györffy B, Neumann U, Simon E, Weichert W, Ebert MP (2010) Systematic evaluation of the miRNA-ome and its downstream effects on mRNA expression identifies gastric cancer progression. J Pathol 222(3):310–319
Torre, L. A., F. Bray, R. L. Siegel, J. Ferlay, J. Lortet‐Tieulent and A. Jemal (2015). Global cancer statistics, 2012. CA: a cancer journal for clinicians 65(2): 87–108.
Tsujiura M, Ichikawa D, Konishi H, Komatsu S, Shiozaki A, Otsuji E (2014) Liquid biopsy of gastric cancer patients: circulating tumor cells and cell-free nucleic acids. World journal of gastroenterology: WJG 20(12):3265
Tsujiura M, Komatsu S, Ichikawa D, Shiozaki A, Konishi H, Takeshita H, Moriumura R, Nagata H, Kawaguchi T, Hirajima S (2015) Circulating miR-18a in plasma contributes to cancer detection and monitoring in patients with gastric cancer. Gastric Cancer 18(2):271–279
Tsukamoto Y, Nakada C, Noguchi T, Tanigawa M, Nguyen LT, Uchida T, Hijiya N, Matsuura K, Fujioka T, Seto M (2010) MicroRNA-375 is downregulated in gastric carcinomas and regulates cell survival by targeting PDK1 and 14-3-3ζ. Can Res 70(6):2339–2349
Ueda T, Volinia S, Okumura H, Shimizu M, Taccioli C, Rossi S, Alder H, Liu C-G, Oue N, Yasui W (2010) Relation between microRNA expression and progression and prognosis of gastric cancer: a microRNA expression analysis. Lancet Oncol 11(2):136–146
Valladares-Ayerbes M, Reboredo M, Medina-Villaamil V, Iglesias-Díaz P, Lorenzo-Patiño MJ, Haz M, Santamarina I, Blanco M, Fernández-Tajes J, Quindós M (2012) Circulating miR-200c as a diagnostic and prognostic biomarker for gastric cancer. J Transl Med 10(1):186
Volinia S, Calin GA, Liu C-G, Ambs S, Cimmino A, Petrocca F, Visone R, Iorio M, Roldo C, Ferracin M (2006) A microRNA expression signature of human solid tumors defines cancer gene targets. ProcNatlAcadSci 103(7):2257–2261
Wang H, Wang L, Wu Z, Sun R, Jin H, Ma J, Liu L, Ling R, Yi J, Wang L (2014) Three dysregulated microRNAs in serum as novel biomarkers for gastric cancer screening. Med Oncol 31(12):298
Wang M, Gu H, Wang S, Qian H, Zhu W, Zhang L, Zhao C, Tao Y, Xu W (2012) Circulating miR-17-5p and miR-20a: molecular markers for gastric cancer. Mol Med Rep 5(6):1514–1520
Wang T, Hou J, Li Z, Zheng Z, Wei J, Song D, Hu T, Wu Q, Yang JY, Cai J-C (2017) miR-15a-3p and miR-16-1-3p negatively regulate Twist1 to repress gastric cancer cell invasion and metastasis. Int J Biol Sci 13(1):122
Wu, J., G. Li, Z. Wang, Y. Yao, R. Chen, X. Pu and J. Wang (2015). Circulating microRNA-21 is a potential diagnostic biomarker in gastric cancer. Disease markers 2015.
Wu J, Li G, Yao Y, Wang Z, Sun W, Wang J (2015) MicroRNA-421 is a new potential diagnosis biomarker with higher sensitivity and specificity than carcinoembryonic antigen and cancer antigen 125 in gastric cancer. Biomarkers 20(1):58–63
Yamanaka S, Olaru AV, An F, Luvsanjav D, Jin Z, Agarwal R, Tomuleasa C, Popescu I, Alexandrescu S, Dima S (2012) MicroRNA-21 inhibits Serpini1, a gene with novel tumour suppressive effects in gastric cancer. Digestive and Liver Disease 44(7):589–596
Yang Q, Jie Z, Ye S, Li Z, Han Z, Wu J, Yang C, Jiang Y (2014) Genetic variations in miR-27a gene decrease mature miR-27a level and reduce gastric cancer susceptibility. Oncogene 33(2):193–202
Yao Y, Suo A-L, Li Z-F, Liu L-Y, Tian T, Ni L, Zhang W-G, Nan K-J, Song T-S, Huang C (2009) MicroRNA profiling of human gastric cancer. Mol Med Rep 2(6):963–970
Yu D, Shin H-S, Lee YS, Lee YC (2014) miR-106b modulates cancer stem cell characteristics through TGF-β/Smad signaling in CD44-positive gastric cancer cells. Lab Invest 94(12):1370–1381
Yu X, Luo L, Wu Y, Yu X, Liu Y, Yu X, Zhao X, Zhang X, Cui L, Ye G (2013) Gastric juice miR-129 as a potential biomarker for screening gastric cancer. Med Oncol 30(1):365
Zhang BG, Li JF, Yu BQ, Zhu ZG, Liu BY, Yan M (2012) microRNA-21 promotes tumor proliferation and invasion in gastric cancer by targeting PTEN. Oncol Rep 27(4):1019–1026
Zhang X, Cui L, Ye G, Zheng T, Song H, Xia T, Yu X, Xiao B, Le Y, Guo J (2012) Gastric juice microRNA-421 is a new biomarker for screening gastric cancer. Tumor Biology 33(6):2349–2355
Zhao G, Jiang T, Liu Y, Huai G, Lan C, Li G, Jia G, Wang K, Yang M (2018) Droplet digital PCR-based circulating microRNA detection serve as a promising diagnostic method for gastric cancer. Bmc Cancer 18(1):676
Zheng Y, Cui L, Sun W, Zhou H, Yuan X, Huo M, Chen J, Lou Y, Guo J (2012) MicroRNA-21 is a new marker of circulating tumor cells in gastric cancer patients. Cancer biomarkers 10(2):71–77
Zhong Z, Dong Z, Yang L, Gong Z (2012) miR-21 induces cell cycle at S phase and modulates cell proliferation by down-regulating hMSH2 in lung cancer. J Cancer Res ClinOncol 138(10):1781–1788
Zhu C, Ren C, Han J, Ding Y, Du J, Dai N, Dai J, Ma H, Hu Z, Shen H (2014) A five-microRNA panel in plasma was identified as potential biomarker for early detection of gastric cancer. Br J Cancer 110(9):2291–2299
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares that there is no conflict of interest.
Rights and permissions
About this article
Cite this article
Ahadi, A. A systematic review of microRNAs as potential biomarkers for diagnosis and prognosis of gastric cancer. Immunogenetics 73, 155–161 (2021). https://doi.org/10.1007/s00251-020-01201-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-020-01201-6