Abstract
Recent advances in molecular and bioinformatic methods have greatly improved our ability to study the formation of an adaptive immune response towards foreign pathogens, self-antigens, and cancer neoantigens. T cell receptors (TCR) are the key players in this process that recognize peptides presented by major histocompatibility complex (MHC). Owing to the huge diversity of both TCR sequence variants and peptides they recognize, accumulation and complex analysis of large amounts of TCR-antigen specificity data is required for understanding the structure and features of adaptive immune responses towards pathogens, vaccines, cancer, as well as autoimmune responses. In the present review, we summarize recent efforts on gathering and interpreting TCR-antigen specificity data and outline the critical role of tighter integration with other immunoinformatics data sources that include epitope MHC restriction, TCR repertoire structure models, and TCR/peptide/MHC structural data. We suggest that such integration can lead to the ability to accurately annotate individual TCR repertoires, efficiently estimate epitope and neoantigen immunogenicity, and ultimately, in silico identify TCRs specific to yet unstudied antigens and predict self-peptides related to autoimmunity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The advent of high-throughput TCR sequencing techniques has changed the way T cell responses are studied (Benichou et al. 2012). As deep sequencing is required to study the highly diverse set of TCR sequences present in each individual, such techniques soon became one of the methods of choice for assaying the TCR repertoire (Rubelt et al. 2017). TCR repertoire sequencing can generate huge amounts of data and can be successfully applied to compare repertoires between individuals and tissues, as well as to track and monitor selected TCR sequences. However, it soon became obvious that the exact functional role is known for only a minor fraction of TCR sequences, which is one of the major bottlenecks in extracting all the available information from TCR repertoire sequencing data (Shugay et al. 2015).
A large number of published assays that involve enriching a T cell sample for cells specific to an antigen of interest and sequencing their TCRs predate high-throughput sequencing era. Such data, while being sparsely organized until recent, carries an important source of information that can be applied to annotate TCR sequences and provide additional information for high-throughput TCR repertoire profiling studies. Recent efforts in summarizing such data gave rise to databases listing TCRs with known antigen specificity: McPAS-TCR (Tickotsky et al. 2017) and VDJdb (Shugay et al. 2018; Bagaev et al. 2019) databases contain ~ 20 k and ~ 55 k specific TCR sequences as of October 2019. Since 2018, immune epitope database (IEDB) also provides TCR and antibody sequences linked with certain antigens having ~ 30 k TCR and ~ 2 k antibody sequences as of October 2019 (Mahajan et al. 2018).
These databases complement the existing set of related immunoinformatics resources that include epitope immunogenicity databases such as IEDB (Vita et al. 2015), MHC-binding analysis tools such as NetMHCpan (Jurtz et al. 2017), and various other T cell epitope prediction tools as those reviewed in Kar et al.( 2018). Both McPAS-TCR and VDJdb are directly applicable to the analysis of large-scale TCR repertoire studies; however, as suggested below, any comprehensive adaptive immunity study requires integrated usage of all of these data sources to delineate interactions between TCRs, antigens, and MHCs involved in the immune response.
Accumulating TCR antigen specificity data
There are several types of T cell assays that can serve as a source of data for the TCR antigen specificity database. The most basic approach is to stimulate a culture of T cells with an antigen of interest in order to select T cells that have undergone an antigen-driven expansion. The stimulation is basically carried out together with a series of limiting dilutions leading to what is known as a limiting dilution assay (Sharrock et al. 1990). Alternatively, primary T cells can be stimulated and sorted on the basis of expression of surface molecules such as CD69, CD154, CD137 (Bacher and Scheffold 2013) or production of effector molecules such as IFNg and perforin. Resulting T cell cultures or cloned TCR sequences can be validated by either antigen re-stimulation and monitoring for intra- and extracellular molecules (cytokine secretion, granzyme B, perforin, etc), or by direct lysis of antigen-expressing target cells (such as 51Cr release assay) (Saade et al. 2012).
Culture-based assays can be carried out using either a specific epitope, a set of peptides, or an entire protein. In the case of protein and peptide mixes, the information about the actual cognate epitope and presenting MHC allele may be lost. It is necessary to note that the VDJdb database includes only those TCR specificity records that list both the exact cognate epitope sequence and the HLA allele presenting it. McPAS-TCR and IEDB, on the other hand, also report TCR specificities resolved up to polypeptide, protein, pathogen, or even pathology level.
One of the main limitations of conventional antigen specificity assays is that they are quite laborious and have a relatively small yield in terms of TCR sequences. The development of peptide-MHC multimer-based techniques allowed accurate identification of specific T cells in both primary samples and expanded cultures (Altman et al. 1996; Dolton et al. 2015). While originally followed by conventional cloning and Sanger sequencing of sorted T cells, currently such approaches are followed by high-throughput sequencing yielding several thousands TCR sequences (Rius et al. 2018). This assay became the main source of specificity records for corresponding databases, as it can yield several-fold more sequences compared to experiments that use Sanger sequencing. As this technique yields many TCR sequences of frequency lower than 1%, it can be biased by sorting contaminations and other artefacts. However, as it was recently shown, antigen specificity can be validated even for low-abundance TCR variants coming from such experiments (Rius et al. 2018).
A combinatorial approach based on parallel sorting of donor cells for several combinations of peptide-MHC multimers followed by TCR repertoire sequencing of positive and negative fractions can be applied to survey a list of epitopes in a single assay (Klinger et al. 2015; Napolitani et al. 2018). The number of false-positives in this case can be minimized by combining the set of positive fractions sorted for each particular epitope. Further developments in the field of TCR specificity assays are related to the application of DNA-barcoded MHC-multimers and single-cell sequencing allowing to survey multiple antigen specificities for an individual T cell in a high-throughput manner (Bentzen et al. 2016). Recent improvements in methods for tagged MHC-multimer library production and specific T cell isolation promise to greatly increase the yield of T cell specificity assays in the near future (Zhang et al. 2018; Moritz et al. 2019; Saini et al. 2019; Ng et al. 2019).
Peptide-MHC libraries in yeast is a powerful technology for high-throughput screening of dozens peptides against target TCR technology (Birnbaum et al. 2014). Except studying of TCR cross-reactivity, the technology is able to provide data for identification of TCR targets without prior knowledge of antigen for the TCR of interest (Gee et al. 2018). Two recently developed promising methods take advantages of “natural” antigen-presentation process in specific cell line for large-scale screening of potential epitopes against T cells (Kula et al. 2019; Sharma et al. 2019). Combination of both library of antigen-presenting cells with library of cells expressing TCRs and co-stimulatory molecules can lead to the technology for large-scale identification of TCR-epitope pairs (Siewert et al. 2011).
Such studies can greatly increase the size of available TCR specificity knowledgebase in the near future; however, a thorough assessment of the amount of non-specific TCR binding should be performed in order to guarantee that TCRs specific to multiple antigens do represent real T cell cross-reactivity events.
Studying TCR repertoire structure
Several large datasets of human TCR repertoires were generated recently, most notable the collection of nearly 786 samples from Emerson et al. (Emerson et al. 2017) and 79 samples from Britanova et al. aging study (Britanova et al. 2016) that were used as a reference set for a number of recent bioinformatic studies (DeWitt et al. 2018; Pogorelyy et al. 2018). Such a large collection of donor TCR repertoires is useful in defining public and rare TCR sequences that can be met frequently across the population or are private to specific donor (Venturi et al. 2008; Shugay et al. 2013; Bagaev et al. 2016).
Moreover, these datasets allow to quantify the incidence specific T cells across the general population and can serve as a reliable baseline for specific TCR frequency estimates. As was demonstrated previously, the precursor frequency of T cells specific to a selected epitope varies greatly and may be as small as 10−6 (Alanio et al. 2010; Neller et al. 2015), reaching the detection limit of a regular high-throughput sequencing study (Britanova et al. 2016). On the other hand, there should be a direct link between epitope immunogenicity, i.e., its ability to elicit an immune response in a given individual, and the incidence rate of specific T cells: given an estimate of ~ 3 × 1011 T cells in a human body and a typical naive T cell clone size of ~ 5 cells (Mora and Walczak 2016), encountering an antigen by an individual specific T cell is an extremely rare event that is limited by specific T cell abundance. Some recent results do indeed confirm this link suggesting that the overall frequency of specific T cells is important for forming an immune response (Pogorelyy et al. 2018).
The structure of unperturbed T cell repertoire can be also summarized using a model originally developed by Murugan et al. (2012). This elegant model involves a relatively small set of parameters including specific variable, diversity, and joining segment choices, their 5′ and 3′ sequence trimmings and features of randomly added N-bases, and can be trained using non-functional TCR sequences that do not undergo thymic selection. Interestingly, for TCRs with known antigen specificity, TCR population frequencies predicted using this model are in a good agreement with those observed in the Emerson et al. dataset (Pogorelyy et al. 2018). Another advantage of using such model is the ability to subtract the background TCR repertoire structure produced by intrinsic biases of the VDJ rearrangement process. For example, one of the recent studies combined a large TCR repertoire sequencing dataset and a VDJ rearrangement model in order to identify a set of rare TCR sequences associated with an autoimmune disease (Komech et al. 2018).
Applying MHC-restriction rules
MHC presentation of foreign peptides is a limiting factor for TCR-mediated recognition, while thymic selection and TCR-MHC interactions at the periphery shape the T cell repertoire of an individual (Zvyagin et al. 2014; Sharon et al. 2016; Chen et al. 2017). A huge body of knowledge on which epitopes are actually presented by MHC molecules and recognized by TCRs is summarized in the IEDB database (Vita et al. 2015). Peptidomes of various MHC molecules can be utilized to build highly accurate predictors of MHC binding, such as the one implemented in netMHCpan software (Jurtz et al. 2017). Searching for TCRs that recognize epitopes that are not presented by donor MHCs even in case they are related to pathogens of interest has little sense as it is unlikely that they can ever elicit a strong immune response. One of the recent studies probing the Emerson et al. dataset shows strong patterns of MHC-associated clusters of TCR sequences that are related to common infections (DeWitt et al. 2018). A ubiquitous infection such as EBV that affects nearly 95% percent of the adult population can leave a distinctive mark on the repertoire that can, in theory, be used for MHC typing in case actual donor MHCs are unknown (Pogorelyy et al. 2018). Thus, pathology-associated TCR sequences can be identified based on their HLA linkage in case of pathologies with strong HLA association. Overall, this suggests that donor MHC haplotype is critical for any study that aims at donor TCR repertoire annotation and should be utilized for increasing the precision of this procedure (Pogorelyy and Shugay 2019).
Studying crystal structures of TCR/peptide/MHC complex
TCR antigen specificity is determined by a set of complex interaction between complementarity determining region (CDR) loops and a peptide presented by the MHC complex (Rossjohn et al. 2015). The ability to model and predict these interactions will ultimately lead to the ability to predict potential binding for TCRs and epitopes that were not studied previously. As current knowledge of TCR specificity is limited to a relatively small set of peptides, studying and modeling interactions in the TCR/peptide/MHC complex is a key for resolving unknown epitope specificity for autoimmunity-linked TCRs and predicting optimal TCRs targeting cancer neoantigens.
The number of available TCR/peptide/MHC structures resolved so far is relatively small, less than 200 structures (Leem et al. 2018). Several databases and web resources dedicated to TCR/peptide/MHC complexes were recently developed, such as the TCRmodel web server (Gowthaman and Pierce 2018). There are, however, several lessons that can be learned from structural data analysis: for example, only the central part of hypervariable CDR3 region is in direct contact with the antigen (Rossjohn et al. 2015) and that CDR3 regions of alpha and beta chains contact the central part of the epitope, while CDR1/2 regions contact epitope termini (Egorov et al. 2018).
Integrated analysis of TCR repertoire sequencing data using immunoinformatics resources
A large set of immunoinformatics resources related to various aspects of TCR repertoire and immunopeptidome analysis outlined in previous paragraphs is summarized in Table 1. Integrating various resources is necessary for proper analysis of TCR repertoire sequencing data since straightforward application of a TCR specificity database for TCR repertoire annotation may give rise to many false-positive calls. Moreover, certain tasks, especially those related to tumor immunotherapy and autoimmunity studies, may require complex analysis of both TCR repertoire and peptidome.
For example, in case one would like to identify the imprint of past and ongoing infections (Fig. 1a), donor HLA haplotype information should be taken into account, as it allows to filter a majority of TCRs that most probably would not recognize a pathogen in a given context (Pogorelyy and Shugay 2019). This strategy can be also applied when dealing with an immune response directed towards rare epitopes: controlling for the baseline structure of the TCR repertoire allows inferring TCR motifs that are present in the data, while common pathogen-specific responses that are always present in donor repertoires can be filtered using the TCR specificity database.
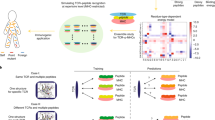
Answering some of the key questions of adaptive immunity studies by integrating TCR repertoire sequencing, HLA typing, epitope discovery, structural modeling, and TCR specificity predictors. a Exploring the imprint of common pathogen exposure on donor TCR repertoire. Pathogen-reactive TCRs can be inferred by analyzing hyperexpanded TCR variants and groups of homologous TCRs that are unlikely to appear simultaneously in an individual repertoire by chance. Resulting TCR set can be queried against a database of TCR sequences of known antigen specificity to infer the pathogen exposure landscape. HLA-restriction of pathogen epitopes can be exploited to filter out irrelevant TCR matches and decrease false discovery rate. b Estimating immunogenicity of pathogenic and autologous epitopes. Viral and self-peptides that are presented by HLA molecules and are homologous to known epitopes from TCR specificity database are rated based on the frequency of TCRs that can potentially recognize them. Specific TCR frequency can be estimated using the V(D)J rearrangement model and correlated with epitope features. Links between specific TCR frequency and epitope features can be further validated using sets of immunogenic and non-immunogenic epitopes from a database containing T cell assay results for various epitopes. c, d Unraveling T cell reactivity to tumor neoantigens and self-peptides related to autoimmunity. c Candidate TCRs are screened for reactivity towards highly immunogenic tumor neoantigens presented by donor HLAs. A set of potential neoantigens is selected based on their presentation on patient HLAs, distance from self, and overall immunogenicity. Candidate TCRs can be either selected using TCR specificity database in case the tumor expresses known neoantigens, selected from the set of TCR sequences of tumor-infiltrating lymphocytes, or predicted using structural modeling or machine learning methods otherwise. d Potential immunogenic self-peptides presented by donor HLAs are screened based on their affinity towards TCRs observed exclusively in patients with autoimmune disease. Screening process can be guided by HLA association of the pathology in question and by considering molecular mimicry and immunogenicity of self-antigens. Self-antigen ranking can again be performed using molecular modeling of corresponding TCR/peptide/MHC complexes or machine learning algorithms
Integrating TCR specificity and epitope assay databases can be applied to quantify peptide immunogenicity for both pathogenic and self-peptides (Fig. 1b). Population frequencies of T cells recognizing a given epitope can be estimated using both existing theoretical models and available TCR repertoire sequencing data. As one would expect, specific TCR frequencies are a good predictor of antigen immunogenicity (Pogorelyy et al. 2018) and can be correlated to certain antigen features allowing immunogenicity scoring for novel epitopes.
Prediction of potential targets for cancer immunotherapy also relies on selection of a set of neoantigens that are both immunogenic and can be presented by donor HLAs (Fig. 1c). A number of recent studies (Bjerregaard et al. 2017; Thorsson et al. 2018; Rubinsteyn et al. 2018; Schenck et al. 2019) aim at identifying prospective candidates among mutated self-peptides. Methods outlined in the previous paragraph can be also considered to rank neoantigens, while TCR specificity prediction algorithms and TCR/peptide/MHC structural modeling can be utilized to select candidate TCRs for tumor immunotherapy.
In case when only a weak immune response is expected, such as responses towards non-mutated tumor-associated antigens or self-peptides in autoimmunity, structural modeling may aid in identifying cognate epitopes for corresponding TCRs (Fig. 1d). Up to this date, despite the emergence of several successful database-driven TCR specificity prediction algorithms (Table 1, “TCR specificity” section), no study reported a pipeline for TCR/peptide/MHC structural modeling that can successfully predict cognate TCRs and epitopes using their primary sequences alone, which remains a major challenge. However, a number of TCR modeling software were published recently (Table 1, “Structural modeling” section) and we hope that TCR specificity databases such as VDJdb can aid in calibrating such tools in order to allow de novo prediction of T cell antigen specificity.
References
Alanio C, Lemaitre F, Law HK, Hasan M, Albert ML (2010) Enumeration of human antigen-specific naive CD8+ T cells reveals conserved precursor frequencies. Blood. 115:3718–3725. https://doi.org/10.1182/blood-2009-10-251124
Altman JD, Moss PA, Goulder PJ, Barouch DH, McHeyzer-Williams M, Bell JI, McMichael A, Davis MM (1996) Phenotypic analysis of antigen-specific T lymphocytes. Science 274:94–96. https://doi.org/10.1126/science.274.5284.94
Bacher P, Scheffold A (2013) Flow-cytometric analysis of rare antigen-specific T cells. Cytometry A 83A:692–701. https://doi.org/10.1002/cyto.a.22317
Bagaev DV, Zvyagin IV, Putintseva EV, Izraelson M, Britanova OV, Chudakov DM, Shugay M (2016) VDJviz: a versatile browser for immunogenomics data. BMC Genomics 17(1):453. https://doi.org/10.1186/s12864-016-2799-7
Bagaev DV, Vroomans RMA, Samir J, Stervbo U, Rius C, Dolton G, Greenshields-Watson A, Attaf M, Egorov ES, Zvyagin IV, Babel N, Cole DK, Godkin AJ, Sewell AK, Kesmir C, Chudakov DM, Luciani F, Shugay M (2019) VDJdb in 2019: database extension, new analysis infrastructure and a T-cell receptor motif compendium. Nucleic Acids Res. https://doi.org/10.1093/nar/gkz874
Benichou J, Ben-Hamo R, Louzoun Y, Efroni S (2012) Rep-Seq: uncovering the immunological repertoire through next-generation sequencing. Immunology 135(3):183–191. https://doi.org/10.1111/j.1365-2567.2011.03527.x
Bentzen AK, Marquard AM, Lyngaa R, Saini SK, Ramskov S, Donia M, Such L, Furness AJ, McGranahan N, Rosenthal R, Straten PT, Szallasi Z, Svane IM, Swanton C, Quezada SA, Jakobsen SN, Eklund AC, Hadrup SR (2016) Large-scale detection of antigen-specific T cells using peptide-MHC-I multimers labeled with DNA barcodes. Nat Biotechnol 34:1037–1045. https://doi.org/10.1038/nbt.3662
Birnbaum ME, Mendoza JL, Sethi DK, Dong S, Glanville J, Dobbins J, Ozkan E, Davis MM, Wucherpfennig KW, Garcia KC (2014) Deconstructing the peptide-MHC specificity of T cell recognition. Cell 157:1073–1087. https://doi.org/10.1016/j.cell.2014.03.047
Bjerregaard AM, Nielsen M, Hadrup SR, Szallasi Z, Eklund AC (2017) MuPeXI: prediction of neo-epitopes from tumor sequencing data. Cancer Immunol Immunother 66(9):1123–1130. https://doi.org/10.1007/s00262-017-2001-3
Borrman T, Cimons J, Cosiano M, Purcaro M, Pierce BG, Baker BM, Weng Z (2017) ATLAS: A database linking binding affinities with structures for wild-type and mutant TCR-pMHC complexes. Proteins 85:908–916. https://doi.org/10.1002/prot.25260
Britanova OV, Shugay M, Merzlyak EM et al (2016) Dynamics of Individual T Cell Repertoires: From Cord Blood to Centenarians. J Immunol 196:5005–5013. https://doi.org/10.4049/jimmunol.1600005
Calis JJA, Maybeno M, Greenbaum JA et al (2013) Properties of MHC Class I Presented Peptides That Enhance Immunogenicity. PLOS Comput Biol 9:e1003266. https://doi.org/10.1371/journal.pcbi.1003266
Chen X, Poncette L, Blankenstein T (2017) Human TCR-MHC coevolution after divergence from mice includes increased nontemplate-encoded CDR3 diversity. J Exp Med 214:3417–3433. https://doi.org/10.1084/jem.20161784
Chowell D, Krishna S, Becker PD, Cocita C, Shu J, Tan X, Greenberg PD, Klavinskis LS, Blattman JN, Anderson KS (2015) TCR contact residue hydrophobicity is a hallmark of immunogenic CD8+ T cell epitopes. Proc Natl Acad Sci 112:E1754–E1762. https://doi.org/10.1073/pnas.1500973112
Dash P, Fiore-Gartland AJ, Hertz T et al (2017) Quantifiable predictive features define epitope specific T cell receptor repertoires. Nature 547:89–93. https://doi.org/10.1038/nature22383
DeWitt WS, Smith A, Schoch G et al (2018) Human T cell receptor occurrence patterns encode immune history, genetic background, and receptor specificity. eLife 7. https://doi.org/10.7554/eLife.38358
Dolton G, Tungatt K, Lloyd A et al (2015) More tricks with tetramers: a practical guide to staining T cells with peptide–MHC multimers. Immunology 146:11–22. https://doi.org/10.1111/imm.12499
Egorov ES, Kasatskaya SA, Zubov VN et al (2018) The Changing Landscape of Naive T Cell Receptor Repertoire With Human Aging. Front Immunol 9. https://doi.org/10.3389/fimmu.2018.01618
Emerson RO, DeWitt WS, Vignali M, Gravley J, Hu JK, Osborne EJ, Desmarais C, Klinger M, Carlson CS, Hansen JA, Rieder M, Robins HS (2017) Immunosequencing identifies signatures of cytomegalovirus exposure history and HLA-mediated effects on the T cell repertoire. Nat Genet 49:659–665. https://doi.org/10.1038/ng.3822
Gee MH, Han A, Lofgren SM et al (2018) Antigen identification for orphan T cell receptors expressed on tumor-infiltrating lymphocytes. Cell 172:549–563. https://doi.org/10.1016/j.cell.2017.11.043
Gielis S, Moris P, Neuter ND et al (2018) TCRex: a webtool for the prediction of T-cell receptor sequence epitope specificity. bioRxiv:373472. https://doi.org/10.1101/373472
Glanville J, Huang H, Nau A, Hatton O, Wagar LE, Rubelt F, Ji X, Han A, Krams SM, Pettus C, Haas N, Arlehamn CSL, Sette A, Boyd SD, Scriba TJ, Martinez OM, Davis MM (2017) Identifying specificity groups in the T cell receptor repertoire. Nature 547:94–98. https://doi.org/10.1038/nature22976
Gowthaman R, Pierce BG (2018) TCRmodel: high resolution modeling of T cell receptors from sequence. Nucleic Acids Res 46:W396–W401. https://doi.org/10.1093/nar/gky432
Gowthaman R, Pierce BG (2019) TCR3d: The T cell receptor structural repertoire database. Bioinforma Oxf Engl. https://doi.org/10.1093/bioinformatics/btz517
Hoffmann T, Marion A, Antes I (2017) DynaDom: structure-based prediction of T cell receptor inter-domain and T cell receptor-peptide-MHC (class I) association angles. BMC Struct Biol 17:2. https://doi.org/10.1186/s12900-016-0071-7
Jokinen E, Huuhtanen J, Mustjoki S et al (2019) Determining epitope specificity of T cell receptors with TCRGP. bioRxiv:542332. https://doi.org/10.1101/542332
Jurtz V, Paul S, Andreatta M, Marcatili P, Peters B, Nielsen M (2017) NetMHCpan-4.0: Improved Peptide-MHC Class I Interaction Predictions Integrating Eluted Ligand and Peptide Binding Affinity Data. J Immunol 199:3360–3368. https://doi.org/10.4049/jimmunol.1700893
Jurtz VI, Jessen LE, Bentzen AK et al (2018) NetTCR: sequence-based prediction of TCR binding to peptide-MHC complexes using convolutional neural networks. bioRxiv:433706. https://doi.org/10.1101/433706
Kar P, Ruiz-Perez L, Arooj M, Mancera RL (2018) Current methods for the prediction of T-cell epitopes. Pept Sci 110:e24046. https://doi.org/10.1002/pep2.24046
Klinger M, Pepin F, Wilkins J, Asbury T, Wittkop T, Zheng J, Moorhead M, Faham M (2015) Multiplex Identification of Antigen-Specific T Cell Receptors Using a Combination of Immune Assays and Immune Receptor Sequencing. PLoS One. 10:e0141561. https://doi.org/10.1371/journal.pone.0141561 eCollection 2015
Komech EA, Pogorelyy MV, Egorov ES et al (2018) CD8+ T cells with characteristic T cell receptor beta motif are detected in blood and expanded in synovial fluid of ankylosing spondylitis patients. Rheumatology 57:1097–1104. https://doi.org/10.1093/rheumatology/kex517
Konstantinou GN (2017) T-Cell Epitope Prediction. Methods Mol Biol Clifton NJ 1592:211–222. https://doi.org/10.1007/978-1-4939-6925-8_17
Kula T, Dezfulian MH, Wang CI et al (2019) T-Scan: a genome-wide method for the systematic discovery of T cell epitopes. Cell 178:1016–1028. https://doi.org/10.1016/j.cell.2019.07.009
Leem J, de Oliveira SHP, Krawczyk K, Deane CM (2018) STCRDab: the structural T-cell receptor database. Nucleic Acids Res 46:D406–D412. https://doi.org/10.1093/nar/gkx971
Mahajan S, Vita R, Shackelford D et al (2018) Epitope Specific Antibodies and T Cell Receptors in the Immune Epitope Database. Front Immunol 9. https://doi.org/10.3389/fimmu.2018.02688
Marcou Q, Mora T, Walczak AM (2018) High-throughput immune repertoire analysis with IGoR. Nat Commun 9:561. https://doi.org/10.1038/s41467-018-02832-w
Mora T, Walczak AM (2016) Quantifying lymphocyte receptor diversity. bioRxiv:046870. https://doi.org/10.1101/046870
Moritz A, Anjanappa R, Wagner C et al (2019) High-throughput peptide-MHC complex generation and kinetic screenings of TCRs with peptide-receptive HLA-A*02:01 molecules. Sci Immunol 4:eaav0860. https://doi.org/10.1126/sciimmunol.aav0860
Murugan A, Mora T, Walczak AM, Callan CG (2012) Statistical inference of the generation probability of T-cell receptors from sequence repertoires. Proc Natl Acad Sci 109:16161–16166. https://doi.org/10.1073/pnas.1212755109
Napolitani G, Kurupati P, Teng KWW, Gibani MM, Rei M, Aulicino A, Preciado-Llanes L, Wong MT, Becht E, Howson L, de Haas P, Salio M, Blohmke CJ, Olsen LR, Pinto DMS, Scifo L, Jones C, Dobinson H, Campbell D, Juel HB, Thomaides-Brears H, Pickard D, Bumann D, Baker S, Dougan G, Simmons A, Gordon MA, Newell EW, Pollard AJ, Cerundolo V (2018) Clonal analysis of Salmonella-specific effector T cells reveals serovar-specific and cross-reactive T cell responses. Nat Immunol 19(7):742–754. https://doi.org/10.1038/s41590-018-0133-z
Neller MA, Ladell K, McLaren JE, Matthews KK, Gostick E, Pentier JM, Dolton G, Schauenburg AJ, Koning D, Fontaine Costa AI, Watkins TS, Venturi V, Smith C, Khanna R, Miners K, Clement M, Wooldridge L, Cole DK, van Baarle D, Sewell AK, Burrows SR, Price DA, Miles JJ (2015) Naive CD8+ T-cell precursors display structured TCR repertoires and composite antigen-driven selection dynamics. Immunol Cell Biol 93:625–633. https://doi.org/10.1038/icb.2015.17
Ng AHC, Peng S, Xu AM, Noh WJ, Guo K, Bethune MT, Chour W, Choi J, Yang S, Baltimore D, Heath JR (2019) MATE-Seq: microfluidic antigen-TCR engagement sequencing. Lab Chip. 19:3011–3021. https://doi.org/10.1039/c9lc00538b
Pogorelyy MV, Shugay M (2019) A framework for annotation of antigen specificities in high-throughput T-cell repertoire sequencing studies. bioRxiv:676239. https://doi.org/10.1101/676239
Pogorelyy MV, Fedorova AD, McLaren JE, Ladell K, Bagaev DV, Eliseev AV, Mikelov AI, Koneva AE, Zvyagin IV, Price DA, Chudakov DM, Shugay M (2018) Exploring the pre-immune landscape of antigen-specific T cells. Genome Med 10:68. https://doi.org/10.1186/s13073-018-0577-7
Pogorelyy MV, Minervina AA, Shugay M, Chudakov DM, Lebedev YB, Mora T, Walczak AM (2019) Detecting T cell receptors involved in immune responses from single repertoire snapshots. PLOS Biol 17:e3000314. https://doi.org/10.1371/journal.pbio.3000314
Ritvo P-G, Saadawi A, Barennes P, Quiniou V, Chaara W, el Soufi K, Bonnet B, Six A, Shugay M, Mariotti-Ferrandiz E, Klatzmann D (2018) High-resolution repertoire analysis reveals a major bystander activation of Tfh and Tfr cells. Proc Natl Acad Sci 115:9604–9609. https://doi.org/10.1073/pnas.1808594115
Rius C, Attaf M, Tungatt K et al (2018) Peptide–MHC Class I Tetramers Can Fail To Detect Relevant Functional T Cell Clonotypes and Underestimate Antigen-Reactive T Cell Populations. J Immunol. https://doi.org/10.4049/jimmunol.1700242
Rossjohn J, Gras S, Miles JJ et al (2015) T Cell Antigen Receptor Recognition of Antigen-Presenting Molecules. Annu Rev Immunol 33:169–200. https://doi.org/10.1146/annurev-immunol-032414-112334
Rubelt F, Busse CE, Bukhari SAC et al (2017) Adaptive Immune Receptor Repertoire Community recommendations for sharing immune-repertoire sequencing data. Nat Immunol 18(12):1274–1278. https://doi.org/10.1038/ni.3873
Rubinsteyn A, Hodes I, Kodysh J et al (2018) Vaxrank: A computational tool for designing personalized cancer vaccines. bioRxiv:142919. https://doi.org/10.1101/142919
Saade F, Gorski SA, Petrovsky N (2012) Pushing the frontiers of T-cell vaccines: accurate measurement of human T-cell responses. Expert Rev Vaccines 11:1459–1470. https://doi.org/10.1586/erv.12.125
Saini SK, Tamhane T, Anjanappa R et al (2019) Empty peptide-receptive MHC class I molecules for efficient detection of antigen-specific T cells. Sci Immunol 4:eaau9039. https://doi.org/10.1126/sciimmunol.aau9039
Schenck RO, Lakatos E, Gatenbee C, Graham TA, Anderson ARA (2019) NeoPredPipe: high-throughput neoantigen prediction and recognition potential pipeline. BMC Bioinformatics 20:264. https://doi.org/10.1186/s12859-019-2876-4
Schritt D, Li S, Rozewicki J et al (2019) Repertoire Builder: high-throughput structural modeling of B and T cell receptors. Mol Syst Des Eng 4:761–768. https://doi.org/10.1039/C9ME00020H
Sethna Z, Elhanati Y, Callan CG et al (2019) OLGA: fast computation of generation probabilities of B- and T-cell receptor amino acid sequences and motifs. Bioinforma Oxf Engl. https://doi.org/10.1093/bioinformatics/btz035
Sharma G, Rive CM, Holt RA (2019) Rapid selection and identification of functional CD8+ T cell epitopes from large peptide-coding libraries. Nat Commun 10:4553. https://doi.org/10.1038/s41467-019-12444-7
Sharon E, Sibener LV, Battle A, Fraser HB, Garcia KC, Pritchard JK (2016) Genetic variation in MHC proteins is associated with T cell receptor expression biases. Nat Genet 48:995–1002. https://doi.org/10.1038/ng.3625
Sharrock CE, Kaminski E, Man S (1990) Limiting dilution analysis of human T cells: a useful clinical tool. Immunol Today 11:281–286
Shugay M, Bolotin DA, Putintseva EV, Pogorelyy MV, Mamedov IZ, Chudakov DM (2013) Huge overlap of individual TCR beta repertoires. Front Immunol 4:466. https://doi.org/10.3389/fimmu.2013.00466
Shugay M, Bagaev DV, Turchaninova MA, Bolotin DA, Britanova OV, Putintseva EV, Pogorelyy MV, Nazarov VI, Zvyagin IV, Kirgizova VI, Kirgizov KI, Skorobogatova EV, Chudakov DM (2015) VDJtools: Unifying Post-analysis of T Cell Receptor Repertoires. PLOS Comput Biol 11:e1004503. https://doi.org/10.1371/journal.pcbi.1004503
Shugay M, Bagaev DV, Zvyagin IV, Vroomans RM, Crawford JC, Dolton G, Komech EA, Sycheva AL, Koneva AE, Egorov ES, Eliseev AV, van Dyk E, Dash P, Attaf M, Rius C, Ladell K, McLaren J, Matthews KK, Clemens EB, Douek DC, Luciani F, van Baarle D, Kedzierska K, Kesmir C, Thomas PG, Price DA, Sewell AK, Chudakov DM (2018) VDJdb: a curated database of T-cell receptor sequences with known antigen specificity. Nucleic Acids Res 46:D419–D427. https://doi.org/10.1093/nar/gkx760
Siewert K, Malotka J, Kawakami N, Wekerle H, Hohlfeld R, Dornmair K (2011) Unbiased identification of target antigens of CD8+ T cells with combinatorial libraries coding for short peptides. Nat Med 18:824–828. https://doi.org/10.1038/nm.2720
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, Porta-Pardo E, Gao GF, Plaisier CL, Eddy JA, Ziv E, Culhane AC, Paull EO, Sivakumar IKA, Gentles AJ, Malhotra R, Farshidfar F, Colaprico A, Parker JS, Mose LE, Vo NS, Liu J, Liu Y, Rader J, Dhankani V, Reynolds SM, Bowlby R, Califano A, Cherniack AD, Anastassiou D, Bedognetti D, Mokrab Y, Newman AM, Rao A, Chen K, Krasnitz A, Hu H, Malta TM, Noushmehr H, Pedamallu CS, Bullman S, Ojesina AI, Lamb A, Zhou W, Shen H, Choueiri TK, Weinstein JN, Guinney J, Saltz J, Holt RA, Rabkin CS, Cancer Genome Atlas Research Network, Lazar AJ, Serody JS, Demicco EG, Disis ML, Vincent BG, Shmulevich I (2018) The Immune Landscape of Cancer. Immunity 48:812–830. https://doi.org/10.1016/j.immuni.2018.03.023
Tickotsky N, Sagiv T, Prilusky J et al (2017) McPAS-TCR: a manually curated catalogue of pathology-associated T cell receptor sequences. Bioinforma Oxf Engl 33:2924–2929. https://doi.org/10.1093/bioinformatics/btx286
Venturi V, Price DA, Douek DC, Davenport MP (2008) The molecular basis for public T-cell responses? Nat Rev Immunol 8:231–238. https://doi.org/10.1038/nri2260
Vita R, Overton JA, Greenbaum JA, Ponomarenko J, Clark JD, Cantrell JR, Wheeler DK, Gabbard JL, Hix D, Sette A, Peters B (2015) The immune epitope database (IEDB) 3.0. Nucleic Acids Res 43:D405–D412. https://doi.org/10.1093/nar/gku938
Zhang SQ, Ma KY, Schonnesen AA et al (2018) High-throughput determination of the antigen specificities of T cell receptors in single cells. Nat Biotechnol 36:1156–1159. https://doi.org/10.1038/nbt.4282
Zvyagin IV, Pogorelyy MV, Ivanova ME et al (2014) Distinctive properties of identical twins' TCR repertoires revealed by high-throughput sequencing. Proc Natl Acad Sci U S A 111(16):5980–5985. https://doi.org/10.1073/pnas.1319389111
Acknowledgments
We would like to thank anonymous reviewers for their valuable suggestions and comments.
Funding
This work was supported by RFBR grant No 19-34-70011.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on “Nomenclature, databases and bioinformatics in Immunogenetics”
Rights and permissions
About this article
Cite this article
Zvyagin, I.V., Tsvetkov, V.O., Chudakov, D.M. et al. An overview of immunoinformatics approaches and databases linking T cell receptor repertoires to their antigen specificity. Immunogenetics 72, 77–84 (2020). https://doi.org/10.1007/s00251-019-01139-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-019-01139-4