Abstract
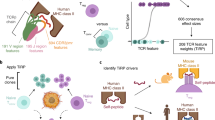
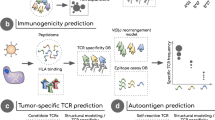
Recognition of cancer epitopes by T cells is fundamental for the activation of targeted antitumor responses. As such, the identification and study of epitope-specific T cells has been instrumental in our understanding of cancer immunology and the development of personalized immunotherapies. To facilitate the study of T-cell epitope specificity, we developed a prediction tool, TCRex, that can identify epitope-specific T-cell receptors (TCRs) directly from TCR repertoire data and perform epitope-specificity enrichment analyses. This chapter details the use of the TCRex web tool.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Roth DB (2014) V(D)J recombination: mechanism, errors, and fidelity. Microbiol Spectr 2:313–324
Ping Y, Liu C, Zhang Y (2018) T-cell receptor-engineered T cells for cancer treatment: current status and future directions. Protein Cell 9:254–266
Bentzen AK, Hadrup SR (2017) Evolution of MHC-based technologies used for detection of antigen-responsive T cells. Cancer Immunol Immunother 66:657–666
Chevalier MF, Bobisse S, Costa-Nunes C et al (2015) High-throughput monitoring of human tumor-specific T-cell responses with large peptide pools. Oncoimmunology 4:e1029702
Lorenz FKM, Ellinger C, Kieback E et al (2017) Unbiased identification of T-cell receptors targeting immunodominant peptide—MHC complexes for T-cell receptor immunotherapy. Hum Gene Ther 28:1158–1168
De Neuter N, Bittremieux W, Beirnaert C et al (2018) On the feasibility of mining CD8+ T cell receptor patterns underlying immunogenic peptide recognition. Immunogenetics 70:159–168
Shugay M, Bagaev DV, Zvyagin IV et al (2018) VDJdb: a curated database of T-cell receptor sequences with known antigen specificity. Nucleic Acids Res 46:D419–D427
Tickotsky N, Sagiv T, Prilusky J et al (2017) McPAS-TCR: a manually curated catalogue of pathology-associated T cell receptor sequences. Bioinformatics 33:2924–2929
Dean J, Emerson RO, Vignali M et al (2015) Annotation of pseudogenic gene segments by massively parallel sequencing of rearranged lymphocyte receptor loci. Genome Med 7:123
Lefranc MP, Giudicelli V, Duroux P et al (2015) IMGT, the international ImMunoGeneTics information system 25 years on. Nucleic Acids Res 43:D413–D422
Adaptive biotechnologies immunoSEQ. https://www.adaptivebiotech.com/products-services/immunoseq/
Bolotin DA, Poslavsky S, Mitrophanov I et al (2015) MiXCR: software for comprehensive adaptive immunity profiling. Nat Methods 12:380–381
Acknowledgments
This work was funded by the University of Antwerp with a University Research Fund (BOF Concerted Research Action) and an Industrial Research Fund (IOF), by the Research Foundation Flanders (FWO) [Personal PhD grants to NDN (1S29816N), SG (1S48819N), and PMo (1141217 N); postdoctoral grant to WB (12W0418N); senior clinical investigator grant to BO (1861219 N); and research project grant (G067118 N)], and by the Belgian American Educational Foundation (BAEF) [postdoctoral grant to WB].
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Gielis, S. et al. (2020). Identification of Epitope-Specific T Cells in T-Cell Receptor Repertoires. In: Boegel, S. (eds) Bioinformatics for Cancer Immunotherapy. Methods in Molecular Biology, vol 2120. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0327-7_13
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0327-7_13
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0326-0
Online ISBN: 978-1-0716-0327-7
eBook Packages: Springer Protocols