Abstract
Bacteriocins are gene-encoded antimicrobial peptides produced by bacteria. These peptides are heterogeneous in terms of structure, antimicrobial activities, biosynthetic clusters, and regulatory mechanisms. Bacteriocins are widespread in nature and may contribute to microbial diversity due to their capacity to target specific bacteria. Primarily studied as food preservatives and therapeutic agents, their function in natural settings is however less known. This review emphasizes the ecological significance of bacteriocins as multifunctional peptides by exploring bacteriocin distribution, mobility, and their impact on bacterial population dynamics and biofilms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the environment, many microbes live in polymicrobial communities occupying ecological niches which may be limited in space and resources [1, 2]. To circumvent these conditions, microbes have developed competition and communication strategies that allow them to survive and coexist with other strains and the surrounding environment [3]. Competition may occur by exploitation when one strain restricts competitors’ access to resources [3], or by interference when an individual causes direct harm to their competitors [3, 4]. Microbe-microbe interactions by interference competition can occur by contact-dependent mechanisms such as secretion systems [5, 6] or by the release of antimicrobial compounds such as bacteriocins that target the opponent cell [2].
Bacteriocins constitute a diverse group of ribosomally synthesized antimicrobial peptides or proteins, that have been described across all bacterial phyla. While bacteriocins have been extensively studied for their potential as food preservatives or antimicrobials [7], the actual role of bacteriocins in bacterial communities is less clear. The conventional hypothesis is that bacteria use bacteriocins to kill neighboring cells in the fight for common resources, but their exact role in different competitive settings in nature remains poorly understood. Bacteriocins may also have additional ecological functions, such as mediating cell lysis to release DNA for natural transformation or interact with regulatory networks in the bacterial cells [8, 9]. Here, we first present an overview of the broad range of structures, biosynthesis pathways, and mechanism of action among bacteriocins, and then we discuss different aspects of bacteriocins in relation to their distribution and impact on the surrounding niche as well as their potential antimicrobial and non-antimicrobial roles.
The Diverse Nature of Bacteriocins
Bacteriocins were first described in Escherichia coli in the early twentieth century [10] and the term quickly embraced the antimicrobial peptides produced by gram-positive bacteria [11,12,13]. The extended bacteriocin family exhibits remarkable diversity in terms of size, genetic organization, structures, antimicrobial spectrum, and mode of action as outlined in recent comprehensive reviews [14, 15]. In the following paragraphs, we offer a brief glimpse into the diverse nature of bacteriocins to emphasize the breadth of this diversity. Classification of bacteriocins is continuously evolving [16,17,18,19]. Here, bacteriocins of gram-negative and gram-positive bacteria are presented separately, but it should be noted that there are examples of similar bacteriocins which can be produced by bacteria across the phylogenetic tree.
Among gram-negative bacteria, colicins (30–80 kDa) and microcins (1–10 kDa) from E. coli have been studied most extensively, although other colicin-like bacteriocins, such as the S-type pyocins produced by Pseudomonas aeruginosa and Klebsiella spp. [14, 20] have been described. Both colicins and microcins generally have relatively narrow antimicrobial spectra, targeting phylogenetically related bacteria [14, 20]. Colicins exert their antimicrobial function either by pore formation, acting as nucleases, or interfering with the peptidoglycan metabolism. Colicins are encoded on small (group A) or larger (group B) plasmids with a variable genetic organization which usually includes a structural bacteriocin gene, an immunity gene encoding a protein for self-protection, and a gene encoding a lytic protein involved in the release of colicin [21]. On the other hand, microcins can be plasmid or chromosome encoded and are commonly divided into class I (small peptides < 5 kDa), which undergo posttranslational modifications, e.g. microcin J25, microcin B17 or microcin E492 [22,23,24]; and class II (larger peptides, 5–10 kDa), which include modified and unmodified peptides such as microcin V and microcin H47 [25, 26]. Their biosynthetic gene clusters include a variable number of genes involved in the production, immunity, and export of the peptide [27] (see Fig. 1 for examples of bacteriocin cluster organization). Due to the different modifications, there is a large structural and functional diversity among microcins. Among the most prominent examples are microcin J25, a cyclic peptide containing a C-terminal segment that is threaded through the cycle (thus referred to as a lasso-peptide) which acts as an RNA-polymerase inhibitor; and microcins E492 and H47 which carry a siderophore-like structure on their C-termini needed for the peptides to cross the outer membrane of target cells via recognition by bacterial siderophore receptors to cause membrane depolarization [28]. As exemplified here, the mechanisms of action of microcins and colicins are diverse, and can also include DNase or RNase activity and inhibitors of protein or cell wall synthesis [21], in addition to membrane disruption and transcription inhibition mentioned above.
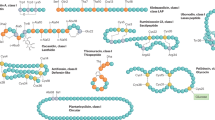
Schematic representation of bacteriocin gene clusters showing the diversity of gene arrangement for bacteriocin synthesis in different bacteriocin groups. Genes are colored according to their function: structural (purple), modification (light green), transport (orange), immunity (dark blue), cleavage (dark green), regulation (grey), and lysis (light blue). References: microcin V [25], colicin E2 [29], nisin A [30], pediocin PA1 [31, 32], lactococcin A [33, 34], garvicin ML [35, 36], garvicin KS [37], halocin C8 [38]. Note that the gene involved in the circularization (modification) of garvicin ML has not been definitely established but likely involves either garX or the garBCDE operon [36]
The study of gram-positive bacteriocins has primarily focused on those produced by lactic acid bacteria. The first classification by Klaenhammer in 1993 [16] was expanded by Nes et al. (1996), then Cotter et al. (2005), and more recently updated by Alvarez-Sieiro et al. (2016) [17,18,19]. In the latter, three classes are suggested: class I are heat-stable peptides of less than 10 kDa which undergo modifications during their biosynthesis (includes the subgroups lantibiotics, circular bacteriocins, sactibiotics, lasso peptides, and glycosylated bacteriocins); class II are also small heat-stable peptides of less than 10 kDa, but these peptides are not modified post-translationally (includes subgroups like the pediocin-like bacteriocins, the two peptide bacteriocins, leaderless bacteriocins, and other non-modified peptides); lastly class III, are larger (> 10 kDa) heat-labile bacteriocins which include both bacteriolysins and proteins with non-lytic modes of action. Despite most bacteriocins showing a narrow antimicrobial spectrum targeting species related to the producer, some gram-positive bacteriocins such as the canonic lantibiotic nisin show a broader antimicrobial spectrum [39,40,41].
Like bacteriocins from gram-negative bacteria, bacteriocin class I and II biosynthetic gene clusters include a variable number of genes and operons on the chromosome or on plasmids. These include at least one structural gene, encoding the antimicrobial precursor peptide, and an immunity gene [15]. Most bacteriocins also have a dedicated transporter encoded in the same locus. Intriguingly, the bacteriocin immunity mechanism of some class I and II bacteriocins involves both the immunity protein and a dedicated ABC transporter system, highlighting a complex regulation for self-protection. This is exemplified by the class I lantibiotic nisin [42,43,44], the circular bacteriocins AS-48 [45,46,47], or the class II leaderless bacteriocin aureocin A53 [48]. In the case of class I bacteriocins, genes involved in modifying the peptides post-translationally are also present. Finally, regulatory genes for control of production are also often present in these loci, making it possible for the bacteria to adapt the energy costly bacteriocin production to the social environmental conditions [49] (see Fig. 1 one for examples of bacteriocin cluster organization).
Gram-positive bacteriocins of class I and II commonly exert their killing action through cell permeabilization and pore formation that causes the cell to leak essential components and eventually lead to the cell death [50]. However, the mode of action varies between different classes of bacteriocins, and exact mechanisms are mostly uncharacterized. Docking molecules such as lipid II [51,52,53,54], the mannose phosphotransferase system (man-PTS) [55, 56] or the intramembrane site-2 protease RseP [57,58,59,60] have been shown to be necessary for the function of some gram-positive bacteriocins; however, it is also possible that some peptides can target bacterial membranes without the need for a specific receptor [61].
In addition to the bacteriocin groups mentioned above, archaeal antimicrobial peptides and proteins, known as archaeocins, have been described, including halocins produced by halophilic Archaea [62, 63]. Furthermore, there are also other peptides and proteins often referred to as bacteriocins, such as the so-called phage tail-like bacteriocins which have been found across gram-positive (e.g., Clostridium spp.) and gram-negative bacteria (e.g., type R- and F-pyocins from P. aeruginosa, or carotovoricin from (Erwinia carotovora) [64,65,66,67].
Bacteriocin Producers are Widespread in Natural Environments
It has been stated that every bacteria is able to produce at least one bacteriocin [16]. Multi-bacteriocinogenic strains have also been frequently described [68,69,70,71]. A vast number of studies over the years have explored different environments for the presence of bacteriocin producer strains, including but not limited to recent studies of fermented foods and dairy products [72,73,74,75], the human and animal gastrointestinal tracts [76,77,78,79,80,81], clinical samples [79, 82, 83], medical devices [84], and environmental samples such as the rhizosphere [85] as well as hypersaline [63] or high-temperature habitats [86]. Moreover, the rise of whole genome and metagenomic sequencing in combination with in silico bioinformatic tools for bacteriocin mining such as Bagel4 [87], antiSMASH [88], Bactibase [89], BADASS [90], and BaPreS, [91] are giving a great impulse to the discovery of bacteriocins and bacteriocin-related genes in sequencing data. Bacteriocin-related genes are indeed found in the vast majority of surveyed microbial communities [73, 81, 92,93,94,95]. The widespread occurrence of bacteriocins and their associated genes, either complete or incomplete gene clusters, could be the result of evolutionary conservation and habitat adaptation driven by the roles they play in resource competition. However, it may also point to a versatile nature of these peptides in fulfilling multiple functions.
Nevertheless, accurately quantifying the prevalence of bacteriocins and bacteriocin-associated genes in different populations remains challenging. Notably, current wet-lab studies are skewed towards finding pathogen-killing molecules, often overlooking bacteriocins with activity against bacteria less relevant from a clinical point of view. Furthermore, determining the bacteriocin prevalence is also impeded by the diverse range of identification methods employed and the difficulty in mimicking the optimal conditions for bacteriocin production in laboratory settings. Indeed, most of the in silico detected bacteriocin gene clusters are “silent,” indicating low or no expression under laboratory conditions [93, 96,97,98]. To investigate the potential antimicrobial function of such silent bacteriocin clusters, in silico-identified bacteriocin genes can be revived by heterologous expression, chemical synthesis, or cell-free peptide synthesis systems [96, 99,100,101,102]. Nevertheless, the roles of these silent gene clusters in nature often remain obscure. Techniques that can be employed to decipher the expression of silent secondary metabolites under different conditions include using high-throughput elicitor screening with reporter genes to evaluate expression levels and identify inducers [103, 104]. Such approaches, combined with large-scale proteomics from diverse environments, will be critical in the future to get a more holistic view of the prevalence of bacteriocins in natural settings.
Regulation of Bacteriocin Production
Bacteria have often evolved to express the bacteriocins only when they are needed and give benefits for the producer. Understanding the regulation of bacteriocins will thus also give important clues about their functional roles. Different regulatory mechanisms have been described and bacterial cells are able to regulate bacteriocin production in response to the growth phase, the physical state of the cell, and environmental factors such as nutrient scarcity, iron availability or osmolarity. The most well-known regulatory systems are quorum-sensing mechanisms governed by dedicated two- or three-component regulatory systems, allowing activation of bacteriocin production by autoinducers as a response to high cell densities and/or other environmental factors, relevant for instance in biofilm settings [105,106,107] (Fig. 2A). Such systems are discussed in detail below. In addition, bacteriocin production has been shown to be regulated by dedicated regulators [108], conserved sigma factors [109], or extracytoplasmic sigma factors [110]. A comprehensive overview of regulatory mechanisms controlling the synthesis of modified peptides was published recently [111].
Regulation of bacteriocin production via three-component systems (TCS). Bacteriocin production can be regulated through the bacteriocin biosynthetic machinery by autoinduction or as part of natural competence systems. A In a typical three-component system a dedicated inducer peptide is produced in a cell density-dependent manner in response to environmental signals. When it reaches a certain threshold, the transmembrane histidine kinase sensor is autophosphorylated and transfers the phosphate group to the response regulator. The phosphorylated response regulator binds to target DNA elements activating the transcription and production of bacteriocin genes. B Examples of bacteriocin production via TCS: autoinduction of nisin production in Lactococcus lactis using nisin as autoinducer [112], autoinduction of plantaricin production in Lactobacillus plantarum NC8 through the pheromone PLNC8IF [49], and CibAB and Blp bacteriocin production in Streptococcus pneumoniae via the ComCDE natural competence system [113,114,115]
In two- or three-component regulatory systems (Fig. 2A), the autoinducer may be the bacteriocins themselves (Fig. 2B). This is the case of the lantibiotics nisin, subtilin, or mersacidin, or the class II bacteriocin plantaricin A [112, 116,117,118]. In other cases, dedicated bacteriocin-like autoinducers without antimicrobial activity function to control bacteriocin production. Examples of the latter include the BlpC peptide of the streptococcal Blp-system, the PLNC8IF in Lactiplantibacillus plantarum [49] or SppIP, in Latilactobacillus sakei, respectively [119]. Interestingly, plantaricin A shows a bacteriocin or pheromone activity depending on its concentration adopting different modes of action [118]. On another aspect, environmental external stimuli can also affect bacteriocin expression. Low salt concentration and short times of exposure to gastric acid, simulating gastrointestinal conditions, stimulated bactofencin A promoter activity [120], antibiotics triggered Blp production in Streptococcus pneumoniae [113] and acetic acid stress increased bacteriocin production in Lacticasibacillus paracasei [121].
An intriguing regulatory and potentially functional link between bacteriocins and horizontal gene transfer is known from streptococci. In several streptococcal species, including S. pneumoniae, Streptococcus mutans, and Streptococcus salivarius, intertwined regulation between expression of bacteriocins and competence for natural transformation has been documented [107, 114, 122] (Fig. 2B). In these species, competent cells are able to take up naked DNA from their surroundings, and the co-regulation with bacteriocins suggests that bacteriocin-mediated killing and lysis of cells may contribute to rendering DNA from neighboring cells accessible for uptake [9]. Regulation of bacteriocin expression is intertwined with competence regulation in different ways [107, 122]. In S. pneumoniae, competence is regulated by quorum-sensing via the two-component system ComCDE, where a competence-stimulating peptide (CSP) encoded by ComC is produced and exported to bind to the histidine kinase ComD, which in turn phosphorylates and leads to transcriptional activation of the competent state via the response regulator ComE (see comprehensive reviews [123, 124]). The most straightforward example of co-regulation is that the phosphorylated response regulator ComE directly activates the transcription of the bacteriocin genes cibABC, a two-peptide bacteriocin found in S. pneumoniae [115]. A more intricate example involves crosstalk between the ComCDE system and a second related two-component system, BlpCRH, known to regulate the expression of bacteriocin-like peptides (Blp) [113, 114]. Here, CSP induces the production of Blp bacteriocins [113, 125] and the corresponding Blp-inducing peptide BlpC is dependent on the competence-expressed transporter for export [113].
Similar to the example above, the Streptococcus gordonii ComCDE competence system has also been shown to be linked to bacteriocin production [126]. Further examples of intertwined regulation are found in S. salivarius [127] where the ComRS regulatory system, another type of two-component system regulating competence in this species, also controls the production of bacteriocins [128]. Most likely, the same is true for bacteriocin production also in Streptococcus thermophilus [128]. Intriguingly, some species such as the dental caries associated S. mutans have both the ComCDE and ComRS competence regulatory systems [129]. In this species, the ComRS system is the main pathway for natural competence regulation while the ComCDE system is involved together with the VicRK signal transduction system in the regulation of the bacteriocins known as mutacins IV, V and VI [130].
The crosstalk and overlaps described here point to a common evolutionary origin of the regulation of bacteriocins and competence in these species [131], highlighting the potential relevance of bacteriocins in the evolution of natural competence. These intricate regulatory links may be a way for bacteriocins to facilitate evolution; bacteriocin-mediated lysis of cells releases DNA to the environment which may be taken up by natural transformation allowing the bacteriocin-producing cells to acquire new features. It remains to be determined how functionally relevant this feature is in nature and whether the coupling of competence and bacteriocin production observed in streptococci is a typical tactic to assist competent, bacteriocin-producing bacteria in other phylogenetic groups to acquire new traits.
Broad and Narrow Spectrum Antimicrobials: Do They Have Different Roles?
Bacteriocins and antibiotics are both part of the bacterial antimicrobial arsenal. Although exceptions exist, bacteriocins generally show a narrower antimicrobial spectrum, most often targeting only bacteria closely related to the bacteriocin producer. Canonical antibiotics, on the other hand, have a wider spectrum of activity and even if their spectrum is restricted, they do not show preferential effect for strains related to their producer. As a result, it has been suggested that bacteriocins have a more limited impact on the surrounding microbiota, causing less disruption to the overall microbial community [132, 133]. Palmer and Foster developed an evolutionary model to explain why bacteria produce both broad-spectrum antibiotics and narrower-spectrum bacteriocins. These models suggest that narrow-spectrum bacteriocins are produced to eliminate relevant competitive threats efficiently while conserving resources, while broader-spectrum antimicrobials are produced when bacterial populations are abundant, aiming to establish dominance by displacing the wider community [134]. Thus, bacteriocins might target direct competitors while protecting others in their cooperative network. One could hypothesize that the production of narrow-spectrum bacteriocins represents a strategy to increase or maintain a high population diversity while effectively eliminating the primary competitors.
This is indeed in line with some studies of the distribution of bacteriocins in different environments. Garcia-Garcera and Rocha studied the production of extracellular proteins by bacteria, including bacteriocins in relation to the habitat spatial structure and the community diversity [135]. Their findings revealed a higher frequency of genes encoding bacteriocins in bacteria from diverse communities, and spatially structured habitats were also found to promote bacteriocin production. Similarly, bacteriocins have been suggested to play a vital role in promoting strain diversity during the establishment of microbiota in the infant human gut [136]. In this metagenome-based study, larger strain diversity was associated with the presence of bacteriocin genes in infant gut samples. Future experimental studies should aim at deciphering how and to what extent bacteriocins affect strain diversity in different environments.
Biofilms and the Dynamics of Bacteriocin-Producing Populations
The surroundings and physical architecture of the microbial habitats are likely to have a great influence on the role bacteriocins play in nature. Some bacteriocins may, for instance, be prone to degradation by proteases and/or physically interact with surfaces, thereby limiting their stability and diffusion in different environments [137,138,139,140]. Therefore, bacteriocin activity is also dependent on the physical distance from bacteriocin producers to the susceptible target cells.
Biofilms, which are organized communities of microorganisms encased in extracellular substances, are the most common form of microbial life in nature [141]. One of the hallmarks of biofilms is the high density of cells, which may facilitate efficient antimicrobial activity of the bacteriocins due to the proximity of cells, but also promote activation of bacteriocin production through quorum-sensing regulation [49] (see section on “Regulation of bacteriocin production”).
The role of bacteriocins in biofilms has been studied to some extent. Bacteriocins have been widely tested as anti-biofilm agents inhibiting pathogenic biofilms [142] and shown to reduce cell surface adhesion inhibiting the early stages of biofilm formation and even disrupting established biofilms at subinhibitory concentrations [143,144,145]. This highlights the potential of bacteriocins as biofilm modulators also in natural settings. Indeed, when a bacteriocin-producing and a bacteriocin-sensitive strain are grown together in liquid culture, the bacteriocin producer outcompetes the sensitive one [146]. However, when the same strains are used in biofilm-experiments, they may be able to coexist [146]. This observation is attributable to the spatial localization of the cells within biofilms and highlights the impact of physical architecture on bacterial interactions [147, 148]. Biofilms may have variable degree of spatial segregation, ranging from well-mixed to complete spatial segregation of cellular subpopulations. It has been argued that bacteriocin production will be most favorable in intermediately mixed biofilms with local competition between strains. This may confer an optimal density of producer strains to initiate an effective attack and target susceptible cells in the vicinity [149, 150]. Too large degree of mixing may preclude the bacteriocin-producing bacteria to initiate a potent attack on target cells, while too large degree of segregation would limit the physical interaction between bacteriocins and susceptible cells. In the latter situation, bacteriocin-sensitive cells may indeed coexist with bacteriocin-producing cells.
With respect to bacteriocin production, bacteria can, in a simplified manner, be classified as bacteriocin-producers, bacteriocin-sensitive or bacteriocin-resistant. The outcome of the different interactions between these groups of organisms and the influence of bacteriocins in these settings is dependent on the physical architecture of the microbial habitat. Colicin-producing E. coli have long served as a model to study complex bacteriocinogenic interactions in in vitro and in in vivo mouse studies [151,152,153]. In environments with spatial segregation, the coexistence of the three phenotypes (bacteriocin-producer, bacteriocin-sensitive, or bacteriocin-resistant) has been shown to follow a non-transitive competition model in which there is no dominant competitor but a balance in which each of the phenotypes can outcompete one of the competitors [151]. The bacteriocin-producer displaces bacteriocin-sensitive cells, the producers are outcompeted by the resistant cells, which do not have a metabolic burden to produce the toxin, and the resistant cells can also be outcompeted by sensitive cells, since the latter would grow faster. This model, referred to as the rock-paper-scissors cyclic model, enables coexistence and favors microbial diversity on a short time scale when there is spatial separation [154]. In well-mixed systems, such as culture flasks, this microbial balance seems to be lost after prolonged incubation, and the resistant strain displaces the bacteriocin-producing and the sensitive ones [151, 152]. Resistant cells avoid the metabolic cost of bacteriocin production while still benefitting from the producers killing sensitive cells.
However, stable cyclic interactions, such as in the rock-paper-scissors model, seem to be unlikely in the long term [155], and different authors have suggested that interactions within the range from purely transitive to purely non-transitive competition are more likely to occur [156, 157]. Interestingly, in a study where engineered E. coli strains producing colicins of different antimicrobial potencies and mechanisms of action (colicin E3, E7, and V) were cultured together, the strain producing the less potent bacteriocin of the three, colicin V in this setup, outcompeted the others [155]. Epidemiological models for bacteriocinogenic S. pneumoniae have shown that diverse bacteriocin profiles from different bacterial populations can also be maintained either by the rock-paper-scissors model or by a “competition-colonization trade-off” mechanism [158]. In the latter, the bacteriocin producer would outcompete its non-producer counterpart within the same bacterial population, but pay a fitness cost in a context of other bacteria that are not sensitive [158]. Pyocins from P. aeruginosa are also important factors in driving competitive interactions in biofilms of these species. While different R-pyocin producers were able to compete each other, R-pyocins deletion mutants were outcompeted by wild-type R-pyocins producers, and combinations of R mutans were able to coexist spatially separated [159].
In a study with pyocin-producing P. aeruginosa isolated from different house locations, the isolates were more likely to outcompete each other when they were at an intermediate genetic distance [160]. The authors suggest that isolates from close communities are more similar to each other as a result of co-evolution and exclusion of sensitive phenotypes, while more separated isolates are more diverse with different pyocin profiles [160]. On a similar setup with Pseudomonas fluorescens isolated from soil at different locations in a park, no relation was found between inhibition and genetic similarity but inhibition among isolates was more frequent within strains from overlapping areas pointing to an important role of bacteriocin in resource utilization and niche competition [161].
A growing body of research highlights the links between bacteriocin producers and biofilm dynamics. However, the extensive diversity of bacteriocins presents a challenge in generalizing models for these complex interactions. Further studies are thus necessary to attain a comprehensive understanding of the intricate bacteriocin-based interactions, encompassing the various types of bacteriocins and experimental setups. Additional aspects, such as phenotypic heterogeneity, which could play a crucial role in sustaining resource-intensive traits like bacteriocin production [162] should also be explored in these investigations. Addressing these factors will contribute to a more complete understanding of the complexities underlying bacteriocin dynamics and their impact on microbial communities.
Mobilization of Bacteriocin Genes by Horizontal Gene Transfer
Horizontal gene transfer (HGT), including transformation via natural competence, conjugation, and transduction, can have significant ecological and evolutionary implications in bacterial communities by transferring genes that modify the competitiveness of the strains. HGT is indeed important for the spread of bacteriocin genes between bacteria. Bacteriocin genes are often encoded on mobile genetic elements such as plasmids, and integrative and conjugative elements which facilitate their transfer between bacterial species or strains by HGT. Such phenomena have been observed for a large number of species, including the gram-positive S. mutans [8], S. salivarius [163], Enterococcus faecalis [164], Staphylococcus aureus [165] or the gram-negatives E. coli [166], and Salmonella [167]. Likewise, bacteriophage-associated bacteriocins have been detected in Bacillus subtilis [168], Klebsiella [169], and E. coli [170].
Strains acquiring complete loci gain the ability to produce bacteriocins, which may be advantageous for competition in natural settings. However, such uptake may also incur a fitness cost [171]. Krauss et al. showed that S. aureus acquiring a plasmid encoding micrococcin P1, a bacteriocin targeting protein synthesis, resulted in immediate production of the bacteriocin [165]. At the same time, this caused a major metabolic burden on the cell, which, interestingly, could be alleviated by increased expression of the central metabolic enzyme, citrate synthase [165]. Whether similar phenomena are true for bacteriocins targeting the cell membranes remains to be determined, but such phenotypes may represent hurdles for the spread of bacteriocin loci by HGT in nature.
In the context of HGT, it is also interesting to note that numerous studies have identified bacteriocin gene clusters that lack one or several essential components of a complete biosynthetic cluster. Some examples of incomplete bacteriocin clusters include Pseudomonas strains that carry the immunity genes but not the corresponding M-type bacteriocin structural genes [172], cyanobacteria harboring separated modification or transportation genes [95], and lactobacilli with partial operons that only contain structural and/or immunity genes [93]. Furthermore, several incomplete bacteriocin clusters have been identified when searching for circular bacteriocins in public databases [173] and so-called lasso peptides in ruminal bacteria [174]. Such incomplete loci may be the result of gene rearrangements, gene loss, or that these partial loci have been acquired by HGT [95, 175]. One potential result of the presence of such loci is the emergence of bacteriocin cheater phenotypes that exploit bacteriocin production by other members in the community without paying the cost of producing the bacteriocin. Isolated bacteriocin immunity genes can be utilized as a protective mechanism conferring cross-immunity against different bacteriocins [96, 176], and such cheaters may thus have important consequences for the dynamics of bacterial populations.
It is also possible that these incomplete clusters are in fact involved in functional bacteriocin production processes. For example, a phenomenon known as coexpression networks has been observed for fungal secondary metabolites in Aspergillus niger [177] and for the production of prochlorosins by cyanobacteria [178]. In these networks, genes involved in the production of specific metabolites are found at different locations in the genome. In bacteriocins from gram-positive bacteria, it is known that translocation and processing can occur via the general sec-dependent pathway, when no dedicated ATP-binding cassette transporters are found in the bacteriocin biosynthetic cluster [69, 179,180,181,182]. Experimentally, it is rare that all the genes required for bacteriocin production or genes in the bacteriocin structural gene vicinity are verified for their function, and therefore it is possible that bacteriocin production involves different pathways and circuits that might be located elsewhere in the bacterial genome.
Non-antimicrobial Effects of Bacteriocins on Target Cells
Bacteriocins may have dual actions. In addition to their antimicrobial effect, they may also act as regulatory molecules at sub-lethal concentrations, similarl to what has been reported for antibiotics and other antimicrobial peptides [183, 184]. In addition to bacteriocin production itself being regulated by quorum sensing systems, sub-inhibitory concentrations of bacteriocins may induce a variety of cellular responses. Among bacteriocins produced by gram-negative bacteria, colicin M sub-inhibitory concentrations induced transcriptional changes in E. coli strains affecting genes involved in the cell envelope, osmotic stress, exopolysaccharide motility, and biofilm-associated genes [185]. Other examples include changes in the architecture S. aureus and Staphylococcus epidermidis biofilms after nisin application [186], and reduced levels of auto inducer 2 with low concentrations of subtilosin, a lantibiotic produced by B. subtilis [187].
In addition, it is known that proteins involved in regulatory pathways are directly influenced by bacteriocins. For instance, the leaderless LsbB family of bacteriocins targets the membrane-bound protease RseP [59] which serves as a key regulator for the stress response through the regulatory intramembrane proteolysis cascade across bacteria [57, 58]. Upon bacteriocin exposure, resistant enterococci acquire mutations in the site-2 protease rseP rendering them with an altered stress response capability [59]. Another example is streptomonomicin, a lasso peptide produced by Streptomonospora alba which influences regulation by WalR, a response regulator linked to cell wall metabolism [188]. Bacillus anthracis acquire streptomonomicin resistance by mutating walR rendering phenotypic defects including clumping of the cells, resistance to forming pellets, and long chains of cells. Moreover, the streptomonomicin-resistant strains showed upregulation of liaI and liaH, and downregulation of ftsE and lytE involved in the stress regulon and cell elongation, respectively [188]. Mutations in such regulatory systems typically render the cells more susceptible to various stressors, and the resistance to bacteriocins thus comes with a fitness cost.
To which extent the bacteriocins’ interactions within these systems can generate additional regulatory responses remains to be determined, but it could be speculated that bacteriocins (such as the leaderless bacteriocins targeting RseP [59]) act as communication modules at subinhibitory concentrations to target and modify diverse regulatory pathways in the population.
Interkingdom interactions are another aspect of the multifaceted nature of bacteriocins. In humans, bacteriocins have been found to interact at different levels with our immune system. For instance, plantaricin A is known to stimulate the synthesis of human-beta defensin 2 in keratinocytes [189], acidocin A and avicin A modulate cytokines and growth factors in human monocytes [190] and both nisin A and Z have immunomodulatory effects as reviewed elsewhere [191, 192]. In plants, bacteriocins from Bacilli spp. isolated from soils, for example, thuricin H7 [193, 194] and bacteriocin IH7 [195] can act as growth promotors.
Conclusion and Outlook
Bacteriocins are ubiquitous among bacteria and extensive research has highlighted their versatility as competition molecules, population modulators, mobile elements, or as part of the cell communication pathways. However, studies on bacteriocins are scattered across the very different and diverse bacteriocin groups. Unraveling the full ecological significance of bacteriocins in different bacteriocin groups and their role in the microbial dialogue poses a challenge. Further multidisciplinary research is essential to gain a comprehensive understanding of bacteriocin circuits.
Data Availability
No datasets were generated or analysed during the current study.
References
Gonzalez JM, Aranda B (2023) Microbial growth under limiting conditions-future perspectives. Microorganisms 11:1641. https://doi.org/10.3390/microorganisms11071641
Hibbing ME, Fuqua C, Parsek MR, Peterson SB (2010) Bacterial competition: surviving and thriving in the microbial jungle. Nat Rev Microbiol 8:15–25. https://doi.org/10.1038/nrmicro2259
Ghoul M, Mitri S (2016) The ecology and evolution of microbial competition. Trends Microbiol 24:833–845. https://doi.org/10.1016/j.tim.2016.06.011
Cornforth DM, Foster KR (2013) Competition sensing: the social side of bacterial stress responses. Nat Rev Microbiol 11:285–293. https://doi.org/10.1038/nrmicro2977
Hayes CS, Aoki SK, Low DA (2010) Bacterial contact-dependent delivery systems. Annu Rev Genet 44:71–90. https://doi.org/10.1146/annurev.genet.42.110807.091449
Ikryannikova LN, Kurbatov LK, Gorokhovets NV, Zamyatnin AA (2020) Contact-dependent growth inhibition in bacteria: do not get too close! Int J Mol Sci 21:7990. https://doi.org/10.3390/ijms21217990
Mathur H, Field D, Rea MC et al (2017) Bacteriocin-antimicrobial synergy: a medical and food perspective. Front Microbiol 8:1205. https://doi.org/10.3389/fmicb.2017.01205
Kreth J, Merritt J, Shi W, Qi F (2005) Coordinated bacteriocin production and competence development: a possible mechanism for taking up DNA from neighbouring species. Mol Microbiol 57:392–404. https://doi.org/10.1111/j.1365-2958.2005.04695.x
Veening J-W, Blokesch M (2017) Interbacterial predation as a strategy for DNA acquisition in naturally competent bacteria. Nat Rev Microbiol 15:629. https://doi.org/10.1038/nrmicro.2017.89
Gratia JP (2000) André Gratia: a forerunner in microbial and viral genetics. Genetics 156:471–476. https://doi.org/10.1093/genetics/156.2.471
Jack RW, Tagg JR, Ray B (1995) Bacteriocins of gram-positive bacteria. Microbiol Rev 59:171–200. https://doi.org/10.1128/mr.59.2.171-200.1995
Tagg JR, Dajani AS, Wannamaker LW (1976) Bacteriocins of gram-positive bacteria. Bacteriol Rev 40:722–756
Drider D, Fimland G, Héchard Y et al (2006) The continuing story of class IIa bacteriocins. Microbiol Mol Biol Rev 70:564–582. https://doi.org/10.1128/MMBR.00016-05
Marković KG, Grujović MŽ, Koraćević MG et al (2022) Colicins and microcins produced by Enterobacteriaceae: characterization, mode of action, and putative applications. Int J Environ Res Public Health 19:11825. https://doi.org/10.3390/ijerph191811825
Simons A, Alhanout K, Duval RE (2020) Bacteriocins, antimicrobial peptides from bacterial origin: overview of their biology and their impact against multidrug-resistant bacteria. Microorganisms 8:639. https://doi.org/10.3390/microorganisms8050639
Klaenhammer TR (1993) Genetics of bacteriocins produced by lactic acid bacteria. FEMS Microbiol Rev 12:39–85. https://doi.org/10.1111/j.1574-6976.1993.tb00012.x
Nes IF, Diep DB, Håvarstein LS et al (1996) Biosynthesis of bacteriocins in lactic acid bacteria. Antonie Van Leeuwenhoek 70:113–128. https://doi.org/10.1007/BF00395929
Cotter PD, Hill C, Ross RP (2005) Bacteriocins: developing innate immunity for food. Nat Rev Microbiol 3:777–788. https://doi.org/10.1038/nrmicro1273
Alvarez-Sieiro P, Montalbán-López M, Mu D, Kuipers OP (2016) Bacteriocins of lactic acid bacteria: extending the family. Appl Microbiol Biotechnol 100:2939–2951. https://doi.org/10.1007/s00253-016-7343-9
Rebuffat S (2011) Bacteriocins from gram-negative bacteria: a classification? In: Drider D, Rebuffat S (eds) Prokaryotic antimicrobial peptides: from genes to applications. Springer, New York, NY, pp 55–72
Cascales E, Buchanan SK, Duché D et al (2007) Colicin biology. Microbiol Mol Biol Rev 71:158–229. https://doi.org/10.1128/MMBR.00036-06
Salomón RA, Farías RN (1992) Microcin 25, a novel antimicrobial peptide produced by Escherichia coli. J Bacteriol 174:7428–7435. https://doi.org/10.1128/jb.174.22.7428-7435.1992
Liu J (1994) Microcin B17: posttranslational modifications and their biological implications. Proc Natl Acad Sci U S A 91:4618–4620
Nolan EM, Fischbach MA, Koglin A, Walsh CT (2007) Biosynthetic tailoring of microcin E492m: post-translational modification affords an antibacterial siderophore-peptide conjugate. J Am Chem Soc 129:14336. https://doi.org/10.1021/ja074650f
Gilson L, Mahanty HK, Kolter R (1987) Four plasmid genes are required for colicin V synthesis, export, and immunity. J Bacteriol 169:2466–2470. https://doi.org/10.1128/jb.169.6.2466-2470.1987
Laviña M, Gaggero C, Moreno F (1990) Microcin H47, a chromosome-encoded microcin antibiotic of Escherichia coli. J Bacteriol 172:6585–6588. https://doi.org/10.1128/jb.172.11.6585-6588.1990
Parker JK, Davies BW (2022) Microcins reveal natural mechanisms of bacterial manipulation to inform therapeutic development. Microbiology 168:001175. https://doi.org/10.1099/mic.0.001175
Massip C, Oswald E (2020) Siderophore-microcins in Escherichia coli: Determinants of digestive colonization, the first step toward virulence. Front Cell Infect Microbiol 10:381. https://doi.org/10.3389/fcimb.2020.00381
Cole ST, Saint-Joanis B, Pugsley AP (1985) Molecular characterisation of the colicin E2 operon and identification of its products. Mol Gen Genet MGG 198:465–472. https://doi.org/10.1007/BF00332940
Kuipers OP, Beerthuyzen MM, Siezen RJ, De Vos WM (1993) Characterization of the nisin gene cluster nisABTCIPR of Lactococcus lactis. requirement of expression of the nisA and nisI genes for development of immunity. Eur J Biochem 216:281–291. https://doi.org/10.1111/j.1432-1033.1993.tb18143.x
Gonzalez CF, Kunka BS (1987) Plasmid-associated bacteriocin production and sucrose fermentation in Pediococcus acidilactici. Appl Environ Microbiol 53:2534–2538. https://doi.org/10.1128/aem.53.10.2534-2538.1987
Marugg JD, Gonzalez CF, Kunka BS et al (1992) Cloning, expression, and nucleotide sequence of genes involved in production of pediocin PA-1, and bacteriocin from Pediococcus acidilactici PAC1.0. Appl Environ Microbiol 58:2360–2367. https://doi.org/10.1128/aem.58.8.2360-2367.1992
Holo H, Nilssen O, Nes IF (1991) Lactococcin A, a new bacteriocin from Lactococcus lactis subsp. cremoris: isolation and characterization of the protein and its gene. J Bacteriol 173:3879–3887. https://doi.org/10.1128/jb.173.12.3879-3887.1991
Varcamonti M, Nicastro G, Venema G, Kok J (2001) Proteins of the lactococcin A secretion system: lcnD encodes two in-frame proteins. FEMS Microbiol Lett 204:259–263. https://doi.org/10.1111/j.1574-6968.2001.tb10894.x
Borrero J, Brede DA, Skaugen M et al (2011) Characterization of garvicin ML, a novel circular bacteriocin produced by Lactococcus garvieae DCC43, isolated from mallard ducks (Anas platyrhynchos). Appl Environ Microbiol 77:369–373. https://doi.org/10.1128/AEM.01173-10
Gabrielsen C, Brede DA, Salehian Z et al (2014) Functional genetic analysis of the GarML gene cluster in Lactococcus garvieae DCC43 gives new insights into circular bacteriocin biosynthesis. J Bacteriol 196:911–919. https://doi.org/10.1128/JB.01115-13
Ovchinnikov KV, Chi H, Mehmeti I et al (2016) Novel group of leaderless multipeptide bacteriocins from gram-positive bacteria. Appl Environ Microbiol 82:5216–5224. https://doi.org/10.1128/AEM.01094-16
Besse A, Vandervennet M, Goulard C et al (2017) Halocin C8: an antimicrobial peptide distributed among four halophilic archaeal genera: Natrinema, Haloterrigena, Haloferax, and Halobacterium. Extrem Life Extreme Cond 21:623–638. https://doi.org/10.1007/s00792-017-0931-5
Okuda K, Zendo T, Sugimoto S et al (2013) Effects of bacteriocins on methicillin-resistant Staphylococcus aureus biofilm. Antimicrob Agents Chemother 57:5572–5579. https://doi.org/10.1128/AAC.00888-13
Tong Z, Ni L, Ling J (2014) Antibacterial peptide nisin: a potential role in the inhibition of oral pathogenic bacteria. Peptides 60:32–40. https://doi.org/10.1016/j.peptides.2014.07.020
Udompijitkul P, Paredes-Sabja D, Sarker MR (2012) Inhibitory effects of nisin against Clostridium perfringens food poisoning and nonfood-borne isolates. J Food Sci 77:M51-56. https://doi.org/10.1111/j.1750-3841.2011.02475.x
Stein T, Heinzmann S, Solovieva I, Entian K-D (2003) Function of Lactococcus lactis nisin immunity genes nisI and nisFEG after coordinated expression in the surrogate host Bacillus subtilis. J Biol Chem 278:89–94. https://doi.org/10.1074/jbc.M207237200
Ra R, Beerthuyzen MM, de Vos WM et al (1999) Effects of gene disruptions in the nisin gene cluster of Lactococcus lactis on nisin production and producer immunity. Microbiol Read Engl 145(Pt 5):1227–1233. https://doi.org/10.1099/13500872-145-5-1227
Siegers K, Entian KD (1995) Genes involved in immunity to the lantibiotic nisin produced by Lactococcus lactis 6F3. Appl Environ Microbiol 61:1082–1089
Diaz M, Valdivia E, Martínez-Bueno M et al (2003) Characterization of a new operon, as-48EFGH, from the as-48 gene cluster involved in immunity to enterocin AS-48. Appl Environ Microbiol 69:1229–1236. https://doi.org/10.1128/AEM.69.2.1229-1236.2003
Cebrián R, Rodríguez-Ruano S, Martínez-Bueno M et al (2014) Analysis of the promoters involved in enterocin AS-48 expression. PLoS ONE 9:e90603. https://doi.org/10.1371/journal.pone.0090603
Martínez-Bueno M, Valdivia E, Gálvez A et al (1998) Analysis of the gene cluster involved in production and immunity of the peptide antibiotic AS-48 in Enterococcus faecalis. Mol Microbiol 27:347–358. https://doi.org/10.1046/j.1365-2958.1998.00682.x
dos Santos Nascimento J, Coelho MLV, Ceotto H et al (2012) Genes involved in immunity to and secretion of aureocin A53, an atypical class II bacteriocin produced by Staphylococcus aureus A53. J Bacteriol 194:875–883. https://doi.org/10.1128/JB.06203-11
Maldonado-Barragán A, West SA (2020) The cost and benefit of quorum sensing-controlled bacteriocin production in Lactobacillus plantarum. J Evol Biol 33:101–111. https://doi.org/10.1111/jeb.13551
Pérez-Ramos A, Madi-Moussa D, Coucheney F, Drider D (2021) Current knowledge of the mode of action and immunity mechanisms of LAB-Bacteriocins. Microorganisms 9:2107. https://doi.org/10.3390/microorganisms9102107
Breukink E, Wiedemann I, van Kraaij C et al (1999) Use of the cell wall precursor lipid II by a pore-forming peptide antibiotic. Science 286:2361–2364. https://doi.org/10.1126/science.286.5448.2361
Hasper HE, Kramer NE, Smith JL et al (2006) An alternative bactericidal mechanism of action for lantibiotic peptides that target lipid II. Science 313:1636–1637. https://doi.org/10.1126/science.1129818
Grein F, Schneider T, Sahl H-G (2019) Docking on lipid II-A widespread mechanism for potent bactericidal activities of antibiotic peptides. J Mol Biol 431:3520–3530. https://doi.org/10.1016/j.jmb.2019.05.014
Martin NI, Breukink E (2007) Expanding role of lipid II as a target for lantibiotics. Future Microbiol 2:513–525. https://doi.org/10.2217/17460913.2.5.513
Diep DB, Skaugen M, Salehian Z et al (2007) Common mechanisms of target cell recognition and immunity for class II bacteriocins. Proc Natl Acad Sci 104:2384–2389. https://doi.org/10.1073/pnas.0608775104
Kjos M, Salehian Z, Nes IF, Diep DB (2010) An extracellular loop of the mannose phosphotransferase system component IIC is responsible for specific targeting by class IIa bacteriocins. J Bacteriol 192:5906–5913. https://doi.org/10.1128/JB.00777-10
Kristensen SS, Oftedal TF, Røhr ÅK et al (2022) The extracellular domain of site-2-metalloprotease RseP is important for sensitivity to bacteriocin EntK1. J Biol Chem 298:102593. https://doi.org/10.1016/j.jbc.2022.102593
Kristensen SS, Diep DB, Kjos M, Mathiesen G (2023) The role of site-2-proteases in bacteria: a review on physiology, virulence, and therapeutic potential. microlife 4:uqad025. https://doi.org/10.1093/femsml/uqad025
Ovchinnikov KV, Kristiansen PE, Straume D et al (2017) The leaderless bacteriocin enterocin K1 is highly potent against Enterococcus faecium: a study on structure, target spectrum and receptor. Front Microbiol 8:774. https://doi.org/10.3389/fmicb.2017.00774
Uzelac G, Kojic M, Lozo J et al (2013) A Zn-dependent metallopeptidase is responsible for sensitivity to LsbB, a class II leaderless bacteriocin of Lactococcus lactis subsp. lactis BGMN1-5. J Bacteriol 195:5614–5621. https://doi.org/10.1128/JB.00859-13
Perez RH, Zendo T, Sonomoto K (2018) Circular and leaderless bacteriocins: biosynthesis, mode of action, applications, and prospects. Front Microbiol 9:2085. https://doi.org/10.3389/fmicb.2018.02085
Kumar V, Singh B, van Belkum MJ et al (2021) Halocins, natural antimicrobials of Archaea: exotic or special or both? Biotechnol Adv 53:107834. https://doi.org/10.1016/j.biotechadv.2021.107834
Najjari A, Mejri H, Jabbar M et al (2020) Halocins, bacteriocin-like antimicrobials produced by the Archaeal domain: occurrence and phylogenetic diversity in Halobacteriales. In: Najjari A, Cherif A, Sghaier H, Ouzari HI (eds) Extremophilic Microbes and Metabolites. IntechOpen. https://doi.org/10.5772/intechopen.94765
Gebhart D, Williams SR, Bishop-Lilly KA et al (2012) Novel high-molecular-weight, R-Type bacteriocins of Clostridium difficile. J Bacteriol 194:6240–6247. https://doi.org/10.1128/JB.01272-12
Nakayama K, Takashima K, Ishihara H et al (2000) The R-type pyocin of Pseudomonas aeruginosa is related to P2 phage, and the F-type is related to lambda phage. Mol Microbiol 38:213–231. https://doi.org/10.1046/j.1365-2958.2000.02135.x
Scholl D (2017) Phage tail-like bacteriocins. Annu Rev Virol 4:453–467. https://doi.org/10.1146/annurev-virology-101416-041632
Yamada K, Hirota M, Niimi Y et al (2006) Nucleotide sequences and organization of the genes for carotovoricin (Ctv) from Erwinia carotovora indicate that Ctv evolved from the same ancestor as Salmonella typhi prophage. Biosci Biotechnol Biochem 70:2236–2247. https://doi.org/10.1271/bbb.60177
Campanero C, Muñoz-Atienza E, Diep DB et al (2020) Biochemical, genetic and transcriptional characterization of multibacteriocin production by the anti-pneumococcal dairy strain Streptococcus infantarius LP90. PLoS ONE 15:e0229417. https://doi.org/10.1371/journal.pone.0229417
Cintas LM, Casaus P, Herranz C et al (2000) Biochemical and genetic evidence that Enterococcus faecium L50 produces enterocins L50A and L50B, the sec-dependent enterocin P, and a novel bacteriocin secreted without an N-terminal extension termed enterocin Q. J Bacteriol 182:6806–6814. https://doi.org/10.1128/JB.182.23.6806-6814.2000
Gordon DM, O’Brien CL (2006) Bacteriocin diversity and the frequency of multiple bacteriocin production in Escherichia coli. Microbiol Read Engl 152:3239–3244. https://doi.org/10.1099/mic.0.28690-0
Nes IF, Diep DB, Holo H (2007) Bacteriocin diversity in Streptococcus and Enterococcus. J Bacteriol 189:1189–1198. https://doi.org/10.1128/JB.01254-06
Gontijo MTP, de S Silva J, Vidigal PMP, Martin JGP (2020) Phylogenetic distribution of the bacteriocin repertoire of lactic acid bacteria species associated with artisanal cheese. Food Res Int 128:108783. https://doi.org/10.1016/j.foodres.2019.108783
Leech J, Cabrera-Rubio R, Walsh AM et al (2020) Fermented-food metagenomics reveals substrate-associated differences in taxonomy and health-associated and antibiotic resistance Determinants. mSystems 5:e00522-20. https://doi.org/10.1128/mSystems.00522-20
Simova ED, Beshkova DB, Dimitrov ZhP (2009) Characterization and antimicrobial spectrum of bacteriocins produced by lactic acid bacteria isolated from traditional Bulgarian dairy products. J Appl Microbiol 106:692–701. https://doi.org/10.1111/j.1365-2672.2008.04052.x
Yi L, Qi T, Hong Y et al (2020) Screening of bacteriocin-producing lactic acid bacteria in Chinese homemade pickle and dry-cured meat, and bacteriocin identification by genome sequencing. LWT 125:109177. https://doi.org/10.1016/j.lwt.2020.109177
Arbulu S, Jiménez JJ, Gútiez L et al (2016) Evaluation of bacteriocinogenic activity, safety traits and biotechnological potential of fecal lactic acid bacteria (LAB), isolated from Griffon vultures (Gyps fulvus subsp. fulvus). BMC Microbiol 16:228. https://doi.org/10.1186/s12866-016-0840-2
Garcia-Gutierrez E, Mayer MJ, Cotter PD, Narbad A (2018) Gut microbiota as a source of novel antimicrobials. Gut Microbes 10:1–21. https://doi.org/10.1080/19490976.2018.1455790
Kuznetsova MV, Mihailovskaya VS, Remezovskaya NB, Starčič Erjavec M (2022) Bacteriocin-producing Escherichia coli isolated from the gastrointestinal tract of farm animals: prevalence, molecular characterization and potential for application. Microorganisms 10:1558. https://doi.org/10.3390/microorganisms10081558
Miri ST, Sotoodehnejadnematalahi F, Amiri MM et al (2022) Comparison of the prevalence of bacteriocin encoding genes in Lactobacillus spp. isolated from fecal samples of healthy volunteers, IBD-patient and IBD-recovered. Iran J Microbiol 14:219–226. https://doi.org/10.18502/ijm.v14i2.9191
Sánchez J, Basanta A, Gómez-Sala B et al (2007) Antimicrobial and safety aspects, and biotechnological potential of bacteriocinogenic enterococci isolated from mallard ducks (Anas platyrhynchos). Int J Food Microbiol 117:295–305. https://doi.org/10.1016/j.ijfoodmicro.2007.04.012
Wosinska L, Walsh CJ, O’Connor PM et al (2022) In vitro and in silico based approaches to identify potential novel bacteriocins from the athlete gut microbiome of an elite athlete cohort. Microorganisms 10:701. https://doi.org/10.3390/microorganisms10040701
Angelopoulou A, Warda AK, O’Connor PM et al (2020) Diverse bacteriocins produced by strains from the human milk microbiota. Front Microbiol 11:788. https://doi.org/10.3389/fmicb.2020.00788
Mazurek-Popczyk J, Pisarska J, Bok E, Baldy-Chudzik K (2020) Antibacterial activity of bacteriocinogenic commensal Escherichia coli against zoonotic strains resistant and sensitive to antibiotics. Antibiot Basel Switz 9:411. https://doi.org/10.3390/antibiotics9070411
Snopkova K, Dufkova K, Klimesova P et al (2020) Prevalence of bacteriocins and their co-association with virulence factors within Pseudomonas aeruginosa catheter isolates. Int J Med Microbiol 310:151454. https://doi.org/10.1016/j.ijmm.2020.151454
Subramanian S, Smith DL (2015) Bacteriocins from the rhizosphere microbiome - from an agriculture perspective. Front Plant Sci 6:909. https://doi.org/10.3389/fpls.2015.00909
Egan K, Field D, Ross RP et al (2018) In silico prediction and exploration of potential bacteriocin gene clusters within the bacterial genus Geobacillus. Front Microbiol 9:2116. https://doi.org/10.3389/fmicb.2018.02116
van Heel AJ, de Jong A, Song C et al (2018) BAGEL4: a user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res 46:W278–W281. https://doi.org/10.1093/nar/gky383
Blin K, Shaw S, Kloosterman AM et al (2021) antiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res 49:W29–W35. https://doi.org/10.1093/nar/gkab335
Hammami R, Zouhir A, Le Lay C et al (2010) BACTIBASE second release: a database and tool platform for bacteriocin characterization. BMC Microbiol 10:22. https://doi.org/10.1186/1471-2180-10-22
Costa SS, da Silva MG, Silva A et al (2023) BADASS: BActeriocin-Diversity ASsessment Software. BMC Bioinformatics 24:24. https://doi.org/10.1186/s12859-022-05106-x
Akhter S, Miller JH (2023) BaPreS: a software tool for predicting bacteriocins using an optimal set of features. BMC Bioinformatics 24:313. https://doi.org/10.1186/s12859-023-05330-z
Azevedo AC, Bento CBP, Ruiz JC et al (2015) Distribution and genetic diversity of bacteriocin gene clusters in rumen microbial genomes. Appl Environ Microbiol 81:7290–7304. https://doi.org/10.1128/AEM.01223-15
Collins FWJ, O’Connor PM, O’Sullivan O et al (2017) Bacteriocin gene-trait matching across the complete Lactobacillus pan-genome. Sci Rep 7:3481. https://doi.org/10.1038/s41598-017-03339-y
Walsh CJ, Guinane CM, Hill C et al (2015) In silico identification of bacteriocin gene clusters in the gastrointestinal tract, based on the Human Microbiome Project’s reference genome database. BMC Microbiol 15:183. https://doi.org/10.1186/s12866-015-0515-4
Wang H, Fewer DP, Sivonen K (2011) Genome mining demonstrates the widespread occurrence of gene clusters encoding bacteriocins in cyanobacteria. PLoS ONE 6:e22384. https://doi.org/10.1371/journal.pone.0022384
Collins FWJ, Mesa-Pereira B, O’Connor PM et al (2018) Reincarnation of bacteriocins from the Lactobacillus pangenomic graveyard. Front Microbiol 9:1298. https://doi.org/10.3389/fmicb.2018.01298
Li H, Guo L, Zhang X et al (2022) Whole-genome sequencing combined with mass spectrometry to identify bacteriocin and mine silent genes. LWT 169:113975. https://doi.org/10.1016/j.lwt.2022.113975
Amos GCA, Awakawa T, Tuttle RN et al (2017) Comparative transcriptomics as a guide to natural product discovery and biosynthetic gene cluster functionality. Proc Natl Acad Sci 114:E11121–E11130. https://doi.org/10.1073/pnas.1714381115
Peña N, Bland MJ, Sevillano E et al (2022) In vitro and in vivo production and split-intein mediated ligation (SIML) of circular bacteriocins. Front Microbiol 13:1052686. https://doi.org/10.3389/fmicb.2022.1052686
Gabant P, Borrero J (2019) PARAGEN 1.0: a standardized synthetic gene library for fast cell-free bacteriocin synthesis. Front Bioeng Biotechnol 7:213. https://doi.org/10.3389/fbioe.2019.00213
van Heel AJ, Mu D, Montalbán-López M et al (2013) Designing and producing modified, new-to-nature peptides with antimicrobial activity by use of a combination of various lantibiotic modification enzymes. ACS Synth Biol 2:397–404. https://doi.org/10.1021/sb3001084
Knerr PJ, van der Donk WA (2012) Discovery, biosynthesis, and engineering of lantipeptides. Annu Rev Biochem 81:479–505. https://doi.org/10.1146/annurev-biochem-060110-113521
Han EJ, Lee SR, Hoshino S, Seyedsayamdost MR (2022) Targeted discovery of cryptic metabolites with antiproliferative activity. ACS Chem Biol 17:3121–3130. https://doi.org/10.1021/acschembio.2c00588
Seyedsayamdost MR (2014) High-throughput platform for the discovery of elicitors of silent bacterial gene clusters. Proc Natl Acad Sci U S A 111:7266–7271. https://doi.org/10.1073/pnas.1400019111
Kreth J, Merritt J, Zhu L et al (2006) Cell density- and ComE-dependent expression of a group of mutacin and mutacin-like genes in Streptococcus mutans. FEMS Microbiol Lett 265:11–17. https://doi.org/10.1111/j.1574-6968.2006.00459.x
Quadri LEN (2002) Regulation of antimicrobial peptide production by autoinducer-mediated quorum sensing in lactic acid bacteria. Antonie Van Leeuwenhoek 82:133–145
Shanker E, Federle MJ (2017) Quorum sensing regulation of competence and bacteriocins in Streptococcus pneumoniae and mutans. Genes 8:E15. https://doi.org/10.3390/genes8010015
Rawlinson ELA, Nes IF, Skaugen M (2005) Identification of the DNA-binding site of the Rgg-like regulator LasX within the lactocin S promoter region. Microbiol Read Engl 151:813–823. https://doi.org/10.1099/mic.0.27364-0
Fomenko D, Veselovskii A, Khmel I (2001) Regulation of microcin C51 operon expression: the role of global regulators of transcription. Res Microbiol 152:469–479. https://doi.org/10.1016/s0923-2508(01)01220-7
Foulston L, Bibb M (2011) Feed-forward regulation of microbisporicin biosynthesis in Microbispora corallina. J Bacteriol 193:3064–3071. https://doi.org/10.1128/JB.00250-11
Bartholomae M, Buivydas A, Viel JH et al (2017) Major gene-regulatory mechanisms operating in ribosomally synthesized and post-translationally modified peptide (RiPP) biosynthesis. Mol Microbiol 106:186–206. https://doi.org/10.1111/mmi.13764
Kuipers OP, Beerthuyzen MM, de Ruyter PGGA et al (1995) Autoregulation of nisin biosynthesis in Lactococcus lactis by signal transduction. J Biol Chem 270:27299–27304. https://doi.org/10.1074/jbc.270.45.27299
Kjos M, Miller E, Slager J et al (2016) Expression of Streptococcus pneumoniae bacteriocins is induced by antibiotics via regulatory interplay with the competence system. PLoS Pathog 12:e1005422. https://doi.org/10.1371/journal.ppat.1005422
Miller EL, Kjos M, Abrudan MI et al (2018) Eavesdropping and crosstalk between secreted quorum sensing peptide signals that regulate bacteriocin production in Streptococcus pneumoniae. ISME J 12:2363–2375. https://doi.org/10.1038/s41396-018-0178-x
Guiral S, Mitchell TJ, Martin B, Claverys J-P (2005) Competence-programmed predation of noncompetent cells in the human pathogen Streptococcus pneumoniae : Genetic requirements. Proc Natl Acad Sci 102:8710–8715. https://doi.org/10.1073/pnas.0500879102
Stein T, Borchert S, Kiesau P et al (2002) Dual control of subtilin biosynthesis and immunity in Bacillus subtilis. Mol Microbiol 44:403–416. https://doi.org/10.1046/j.1365-2958.2002.02869.x
Schmitz S, Hoffmann A, Szekat C et al (2006) The lantibiotic mersacidin is an autoinducing peptide. Appl Environ Microbiol 72:7270–7277. https://doi.org/10.1128/AEM.00723-06
Hauge HH, Mantzilas D, Moll GN et al (1998) Plantaricin A is an amphiphilic alpha-helical bacteriocin-like pheromone which exerts antimicrobial and pheromone activities through different mechanisms. Biochemistry 37:16026–16032. https://doi.org/10.1021/bi981532j
Brurberg MB, Nes IF, Eijsink VG (1997) Pheromone-induced production of antimicrobial peptides in Lactobacillus. Mol Microbiol 26:347–360. https://doi.org/10.1046/j.1365-2958.1997.5821951.x
Guinane CM, Piper C, Draper LA et al (2015) Impact of environmental factors on bacteriocin promoter activity in gut-derived Lactobacillus salivarius. Appl Environ Microbiol 81:7851–7859. https://doi.org/10.1128/AEM.02339-15
Kang J, Zhou X, Zhang W et al (2022) Transcriptomic analysis of bacteriocin synthesis and stress response in Lactobacillus paracasei HD1.7 under acetic acid stress. LWT 154:112897. https://doi.org/10.1016/j.lwt.2021.112897
Reck M, Tomasch J, Wagner-Döbler I (2015) The alternative sigma factor sigX controls bacteriocin synthesis and competence, the two quorum sensing regulated traits in Streptococcus mutans. PLoS Genet 11:e1005353. https://doi.org/10.1371/journal.pgen.1005353
Fontaine L, Wahl A, Fléchard M et al (2015) Regulation of competence for natural transformation in streptococci. Infect Genet Evol 33:343–360. https://doi.org/10.1016/j.meegid.2014.09.010
Straume D, Stamsås GA, Håvarstein LS (2015) Natural transformation and genome evolution in Streptococcus pneumoniae. Infect Genet Evol J Mol Epidemiol Evol Genet Infect Dis 33:371–380. https://doi.org/10.1016/j.meegid.2014.10.020
Wholey W-Y, Kochan TJ, Storck DN, Dawid S (2016) Coordinated bacteriocin expression and competence in Streptococcus pneumoniae contributes to genetic adaptation through neighbor predation. PLOS Pathog 12:e1005413. https://doi.org/10.1371/journal.ppat.1005413
Vickerman MM, Iobst S, Jesionowski AM, Gill SR (2007) Genome-wide transcriptional changes in Streptococcus gordonii in response to competence signaling peptide. J Bacteriol 189:7799–7807. https://doi.org/10.1128/JB.01023-07
Wang CY, Dawid S (2018) Mobilization of bacteriocins during competence in streptococci. Trends Microbiol 26:389–391. https://doi.org/10.1016/j.tim.2018.03.002
Mignolet J, Fontaine L, Sass A et al (2018) Circuitry rewiring directly couples competence to predation in the gut dweller Streptococcus salivarius. Cell Rep 22:1627–1638. https://doi.org/10.1016/j.celrep.2018.01.055
Håvarstein LS (2010) Increasing competence in the genus Streptococcus. Mol Microbiol 78:541–544. https://doi.org/10.1111/j.1365-2958.2010.07380.x
Senadheera DB, Cordova M, Ayala EA et al (2012) Regulation of bacteriocin production and cell death by the VicRK signaling system in Streptococcus mutans. J Bacteriol 194:1307–1316. https://doi.org/10.1128/JB.06071-11
Martin B, Quentin Y, Fichant G, Claverys J-P (2006) Independent evolution of competence regulatory cascades in streptococci? Trends Microbiol 14:339–345. https://doi.org/10.1016/j.tim.2006.06.007
Drissi F, Buffet S, Raoult D, Merhej V (2015) Common occurrence of antibacterial agents in human intestinal microbiota. Front Microbiol 6:441. https://doi.org/10.3389/fmicb.2015.00441
Umu ÖCO, Bäuerl C, Oostindjer M et al (2016) The potential of class II bacteriocins to modify gut microbiota to improve host health. PLoS ONE 11:e0164036. https://doi.org/10.1371/journal.pone.0164036
Palmer JD, Foster KR (2022) The evolution of spectrum in antibiotics and bacteriocins. Proc Natl Acad Sci 119:e2205407119. https://doi.org/10.1073/pnas.2205407119
Garcia-Garcera M, Rocha EPC (2020) Community diversity and habitat structure shape the repertoire of extracellular proteins in bacteria. Nat Commun 11:758. https://doi.org/10.1038/s41467-020-14572-x
Ormaasen I, Rudi K, Diep DB, Snipen L (2023) Metagenome-mining indicates an association between bacteriocin presence and strain diversity in the infant gut. BMC Genomics 24:295. https://doi.org/10.1186/s12864-023-09388-0
Melia S, Juliyarsi I, Kurnia YF et al (2021) Characteristics of antibacterial activity stability of crude bacteriocin Pediococcus acidilactici BK01. IOP Conf Ser Earth Environ Sci 782:032074. https://doi.org/10.1088/1755-1315/782/3/032074
Soltani S, Zirah S, Rebuffat S et al (2021) Gastrointestinal stability and cytotoxicity of bacteriocins from Gram-positive and Gram-negative bacteria: a comparative in vitro study. Front Microbiol 12:780355. https://doi.org/10.3389/fmicb.2021.780355
Blom H, Katla T, Hagen BF, Axelsson L (1997) A model assay to demonstrate how intrinsic factors affect diffusion of bacteriocins. Int J Food Microbiol 38:103–109. https://doi.org/10.1016/S0168-1605(97)00098-6
Azevedo PO de S de, Molinari F, Oliveira RP de S (2018) Importance of the agar-media in the evaluation of bacteriocin activity against the same test-microorganisms. Braz J Pharm Sci 54:e17533https://doi.org/10.1590/s2175-97902018000117533
Flemming H-C, Wuertz S (2019) Bacteria and Archaea on Earth and their abundance in biofilms. Nat Rev Microbiol 17:247–260. https://doi.org/10.1038/s41579-019-0158-9
Mathur H, Field D, Rea MC et al (2018) Fighting biofilms with lantibiotics and other groups of bacteriocins. Npj Biofilms Microbiomes 4:1–13. https://doi.org/10.1038/s41522-018-0053-6
Kim N-N, Kim WJ, Kang S-S (2019) Anti-biofilm effect of crude bacteriocin derived from Lactobacillus brevis DF01 on Escherichia coli and Salmonella Typhimurium. Food Control 98:274–280. https://doi.org/10.1016/j.foodcont.2018.11.004
de Jesus Pimentel-Filho N, de Freitas Martins MC, Nogueira GB et al (2014) Bovicin HC5 and nisin reduce Staphylococcus aureus adhesion to polystyrene and change the hydrophobicity profile and Gibbs free energy of adhesion. Int J Food Microbiol 190:1–8. https://doi.org/10.1016/j.ijfoodmicro.2014.08.004
Shayesteh F, Ahmad A, Usup G (2020) In vitro anti-biofilm activity of bacteriocin from a marine Bacillus sp. strain Sh10 against Proteus mirabilis. Iran J Microbiol 12:52–61
Tait K, Sutherland IW (2002) Antagonistic interactions amongst bacteriocin-producing enteric bacteria in dual species biofilms. J Appl Microbiol 93:345–352. https://doi.org/10.1046/j.1365-2672.2002.01692.x
van Gestel J, Weissing FJ, Kuipers OP, Kovács ÁT (2014) Density of founder cells affects spatial pattern formation and cooperation in Bacillus subtilis biofilms. ISME J 8:2069–2079. https://doi.org/10.1038/ismej.2014.52
Van Dyken JD, Müller MJI, Mack KML, Desai MM (2013) Spatial population expansion promotes the evolution of cooperation in an experimental prisoner’s dilemma. Curr Biol 23:919–923. https://doi.org/10.1016/j.cub.2013.04.026
Bucci V, Nadell CD, Xavier JB (2011) The evolution of bacteriocin production in bacterial biofilms. Am Nat 178:E162-173. https://doi.org/10.1086/662668
Nadell CD, Drescher K, Foster KR (2016) Spatial structure, cooperation and competition in biofilms. Nat Rev Microbiol 14:589–600. https://doi.org/10.1038/nrmicro.2016.84
Kerr B, Riley MA, Feldman MW, Bohannan BJM (2002) Local dispersal promotes biodiversity in a real-life game of rock–paper–scissors. Nature 418:171–174. https://doi.org/10.1038/nature00823
Kirkup BC, Riley MA (2004) Antibiotic-mediated antagonism leads to a bacterial game of rock–paper–scissors in vivo. Nature 428:412–414. https://doi.org/10.1038/nature02429
Majeed H, Gillor O, Kerr B, Riley MA (2011) Competitive interactions in Escherichia coli populations: the role of bacteriocins. ISME J 5:71–81. https://doi.org/10.1038/ismej.2010.90
Soliveres S, Allan E (2018) Everything you always wanted to know about intransitive competition but were afraid to ask. J Ecol 106:807–814. https://doi.org/10.1111/1365-2745.12972
Liao MJ, Miano A, Nguyen CB et al (2020) Survival of the weakest in non-transitive asymmetric interactions among strains of E. coli. Nat Commun 11:6055. https://doi.org/10.1038/s41467-020-19963-8
Gallien L, Zimmermann NE, Levine JM, Adler PB (2017) The effects of intransitive competition on coexistence. Ecol Lett 20:791–800. https://doi.org/10.1111/ele.12775
Laird RA, Schamp BS (2006) Competitive intransitivity promotes species coexistence. Am Nat 168:182–193. https://doi.org/10.1086/506259
Lehtinen S, Croucher NJ, Blanquart F, Fraser C (2022) Epidemiological dynamics of bacteriocin competition and antibiotic resistance. Proc R Soc B Biol Sci 289:20221197. https://doi.org/10.1098/rspb.2022.1197
Oluyombo O, Penfold CN, Diggle SP (2019) Competition in biofilms between cystic fibrosis isolates of Pseudomonas aeruginosa is shaped by R-Pyocins. mBio 10:e01828-18. https://doi.org/10.1128/mBio.01828-18
Mojesky AA, Remold SK (2020) Spatial structure maintains diversity of pyocin inhibition in household Pseudomonas aeruginosa. Proc R Soc B Biol Sci 287:20201706. https://doi.org/10.1098/rspb.2020.1706
Bruce JB, West SA, Griffin AS (2017) Bacteriocins and the assembly of natural Pseudomonas fluorescens populations. J Evol Biol 30:352–360. https://doi.org/10.1111/jeb.13010
Cooper GA, Liu M, Peña J, West SA (2022) The evolution of mechanisms to produce phenotypic heterogeneity in microorganisms. Nat Commun 13:195. https://doi.org/10.1038/s41467-021-27902-4
Dahmane N, Libante V, Charron-Bourgoin F et al (2017) Diversity of integrative and conjugative elements of Streptococcus salivarius and their intra- and interspecies transfer. Appl Environ Microbiol 83:e00337-e417. https://doi.org/10.1128/AEM.00337-17
Devi SM, Halami PM (2013) Conjugal transfer of bacteriocin plasmids from different genera of lactic acid bacteria into Enterococcus faecalis JH2-2. Ann Microbiol 63:1611–1617. https://doi.org/10.1007/s13213-013-0624-y
Krauss S, Harbig TA, Rapp J et al (2023) Horizontal transfer of bacteriocin biosynthesis genes requires metabolic adaptation to improve compound production and cellular fitness. Microbiol Spectr 11:e0317622. https://doi.org/10.1128/spectrum.03176-22
Micenková L, Bosák J, Kucera J et al (2019) Colicin Z, a structurally and functionally novel colicin type that selectively kills enteroinvasive Escherichia coli and Shigella strains. Sci Rep 9:11127. https://doi.org/10.1038/s41598-019-47488-8
Johnson TJ, Thorsness JL, Anderson CP et al (2010) Horizontal gene transfer of a ColV plasmid has resulted in a dominant avian clonal type of Salmonella enterica serovar Kentucky. PLoS ONE 5:e15524. https://doi.org/10.1371/journal.pone.0015524
Dragoš A, Andersen AJC, Lozano-Andrade CN et al (2021) Phages carry interbacterial weapons encoded by biosynthetic gene clusters. Curr Biol 31:3479-3489.e5. https://doi.org/10.1016/j.cub.2021.05.046
Chavan M, Rafi H, Wertz J et al (2005) Phage associated bacteriocins reveal a novel mechanism for bacteriocin diversification in Klebsiella. J Mol Evol 60:546–556. https://doi.org/10.1007/s00239-004-0263-9
Spriewald S, Stadler E, Hense BA et al (2020) Evolutionary stabilization of cooperative toxin production through a bacterium-plasmid-phage interplay. mBio 11:e00912-20. https://doi.org/10.1128/mBio.00912-20
Heilbronner S, Krismer B, Brötz-Oesterhelt H, Peschel A (2021) The microbiome-shaping roles of bacteriocins. Nat Rev Microbiol 19:726–739. https://doi.org/10.1038/s41579-021-00569-w
Ghequire MGK, Kemland L, De Mot R (2017) Novel immunity proteins associated with colicin M-like bacteriocins exhibit promiscuous protection in Pseudomonas. Front Microbiol 8:93. https://doi.org/10.3389/fmicb.2017.00093
Vezina B, Rehm BHA, Smith AT (2020) Bioinformatic prospecting and phylogenetic analysis reveals 94 undescribed circular bacteriocins and key motifs. BMC Microbiol 20:77. https://doi.org/10.1186/s12866-020-01772-0
Sabino YNV, de Araújo KC, do Val de Assis FG et al (2020) In silico screening unveil the great potential of ruminal bacteria synthesizing lasso peptides. Front Microbiol 11:576738. https://doi.org/10.3389/fmicb.2020.576738
Popin RV, Alvarenga DO, Castelo-Branco R et al (2021) Mining of cyanobacterial genomes indicates natural product biosynthetic gene clusters located in conjugative plasmids. Front Microbiol 12:684565. https://doi.org/10.3389/fmicb.2021.684565
Draper LA, Grainger K, Deegan LH et al (2009) Cross-immunity and immune mimicry as mechanisms of resistance to the lantibiotic lacticin 3147. Mol Microbiol 71:1043–1054. https://doi.org/10.1111/j.1365-2958.2008.06590.x
Kwon MJ, Steiniger C, Cairns TC et al (2021) Beyond the biosynthetic gene cluster paradigm: genome-wide coexpression networks connect clustered and unclustered transcription factors to secondary metabolic pathways. Microbiol Spectr 9:e0089821. https://doi.org/10.1128/Spectrum.00898-21
Li B, Sher D, Kelly L et al (2010) Catalytic promiscuity in the biosynthesis of cyclic peptide secondary metabolites in planktonic marine cyanobacteria. Proc Natl Acad Sci 107:10430–10435. https://doi.org/10.1073/pnas.0913677107
Zheng S, Sonomoto K (2018) Diversified transporters and pathways for bacteriocin secretion in gram-positive bacteria. Appl Microbiol Biotechnol 102:4243–4253. https://doi.org/10.1007/s00253-018-8917-5
Herranz C, Driessen AJM (2005) Sec-mediated secretion of bacteriocin enterocin P by Lactococcus lactis. Appl Environ Microbiol 71:1959–1963. https://doi.org/10.1128/AEM.71.4.1959-1963.2005
Liu G, Wang Y, Li X et al (2019) Genetic and biochemical evidence that Enterococcus faecalis Gr17 produces a novel and sec-dependent bacteriocin, enterocin Gr17. Front Microbiol 10:1806. https://doi.org/10.3389/fmicb.2019.01806
Sánchez J, Diep DB, Herranz C et al (2007) Amino acid and nucleotide sequence, adjacent genes, and heterologous expression of hiracin JM79, a sec-dependent bacteriocin produced by Enterococcus hirae DCH5, isolated from Mallard ducks (Anas platyrhynchos). FEMS Microbiol Lett 270:227–236. https://doi.org/10.1111/j.1574-6968.2007.00673.x
Romero D, Traxler MF, López D, Kolter R (2011) Antibiotics as signal molecules. Chem Rev 111:5492–5505. https://doi.org/10.1021/cr2000509
Vasilchenko AS, Rogozhin EA (2019) Sub-inhibitory effects of antimicrobial peptides. Front Microbiol 10:1160. https://doi.org/10.3389/fmicb.2019.01160
Kamenšek S, Žgur-Bertok D (2013) Global transcriptional responses to the bacteriocin colicin M in Escherichia coli. BMC Microbiol 13:42. https://doi.org/10.1186/1471-2180-13-42
Andre C, de Jesus P-F, de Almeida Costa PM, Vanetti MCD (2019) Changes in the composition and architecture of staphylococcal biofilm by nisin. Braz J Microbiol 50:1083–1090. https://doi.org/10.1007/s42770-019-00135-w
Algburi A, Zehm S, Netrebov V et al (2017) Subtilosin prevents biofilm formation by inhibiting bacterial quorum sensing. Probiotics Antimicrob Proteins 9:81–90. https://doi.org/10.1007/s12602-016-9242-x
Metelev M, Tietz JI, Melby JO et al (2015) Structure, bioactivity, and resistance mechanism of streptomonomicin, an unusual lasso peptide from an understudied halophilic actinomycete. Chem Biol 22:241–250. https://doi.org/10.1016/j.chembiol.2014.11.017
Marzani B, Pinto D, Minervini F et al (2012) The antimicrobial peptide pheromone plantaricin A increases antioxidant defenses of human keratinocytes and modulates the expression of filaggrin, involucrin, β-defensin 2 and tumor necrosis factor-α genes. Exp Dermatol 21:665–671. https://doi.org/10.1111/j.1600-0625.2012.01538.x
Antoshina DV, Balandin SV, Bogdanov IV et al (2022) Antimicrobial activity and immunomodulatory properties of acidocin A, the pediocin-like bacteriocin with the non-canonical structure. Membranes 12:1253. https://doi.org/10.3390/membranes12121253
Guryanova SV (2023) Immunomodulation, bioavailability and safety of bacteriocins. Life 13:1521. https://doi.org/10.3390/life13071521
Małaczewska J, Kaczorek-Łukowska E (2021) Nisin-A lantibiotic with immunomodulatory properties: A review. Peptides 137:170479. https://doi.org/10.1016/j.peptides.2020.170479
Gray EJ, Lee KD, Souleimanov AM et al (2006) A novel bacteriocin, thuricin 17, produced by plant growth promoting rhizobacteria strain Bacillus thuringiensis NEB17: isolation and classification. J Appl Microbiol 100:545–554. https://doi.org/10.1111/j.1365-2672.2006.02822.x
Lee KD, Gray EJ, Mabood F et al (2009) The class IId bacteriocin thuricin-17 increases plant growth. Planta 229:747–755. https://doi.org/10.1007/s00425-008-0870-6
Hammami I, Triki MA, Rebai A (2011) Purification and characterization of the novel bacteriocin Bac IH7 with antifungal and antibacterial properties. J Plant Pathol 93:443–454. https://doi.org/10.4454/JPP.V93I2.1200
Funding
Open access funding provided by Norwegian University of Life Sciences. This work was supported by Marie Skłodowska-Curie Actions H2020-MSCA-EF-ST-2020 grant #101029099 and Research Council of Norway, Joint Programming Initiative on Antimicrobial Resistance grant #296906.
Author information
Authors and Affiliations
Contributions
SA and MK designed the article, SA performed literature search, SA and MK wrote the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors have no relevant financial or non-financial interests to disclose.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arbulu, S., Kjos, M. Revisiting the Multifaceted Roles of Bacteriocins. Microb Ecol 87, 41 (2024). https://doi.org/10.1007/s00248-024-02357-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-024-02357-4