Abstract
Bacteriocins are ribosomally synthesised small antimicrobial peptides produced from a wide range of bacteria, and also rich sources for potential alternatives to traditional antibiotics. Many bacteriocins have highly specific antibacterial activity against target pathogens, even including drug-resistant bacteria such as methicillin-resistant Staphylococcus aureus and vancomycin-resistant Enterococcus. As the final and essential step during biosynthesis, the leader sequence removal and exportation of matured bacteriocin are lacking of research and therefore the last to be understood. In respect of production, bacteriocin precursor peptides are processed and exported by a group of membrane proteins from the ATP-binding cassette transporter family. The main aims of this article are to summarise knowledge till now on the leader signal and correlated transporters for bacteriocin secretion in gram-positive bacteria in a review for the first time, to introduce different strategies for higher production, and to offer new insights into many essential but still unanswered questions above for the purpose of more efficient bacteriocin utilisation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The discovery of penicillin by Alexander Fleming in 1928 opened new avenues for the human race to fight against a wide range of bacterial infections and diseases. Within a century of its discovery, a number of other naturally occurring and artificially synthesised antibiotics have been discovered. The abuse and careless utilisation of antibiotics, however, has led to the emergence of drug resistance in bacteria. This poses an alarming situation, and currently, there are limited solutions available to tackle these uprising threats (Cotter et al. 2005). Therefore, there is an urgent need for the development of alternative antibiotic drugs.
Bacteriocins comprise a group of structurally diverse small antimicrobial peptides, which are ribosomally synthesised by a wide range of bacterial strains in the environment (Cotter et al. 2005). Based on their structure, bacteriocins have been classified into two major groups: class I in which peptides undergo post-translational modifications, and class II peptides which remain unmodified (Cotter et al. 2013). In addition to this primary classification system based on secondary modifications, class I bacteriocins have been further subdivided into lantibiotics (containing lanthionine rings), glycocins (containing S-linked glycopeptides), bottromycins (containing macrocyclic amidine), thiopeptides (containing thiazole and other dehydro amino acids with a pyridine/piperidine core), sactibiotics (containing sulphur-α-carbon linkages), and linaridins (linear dehydrated peptides) in gram-positive bacteria (Cotter et al. 2013). Similarly, the class II bacteriocins are subdivided into four groups. The class IIa peptides are pediocin PA-1-like bacteriocins with a conserved YGNGV motif. Bacteriocins from class IIb are composed of two unmodified peptides, both of which should be active. Class IIc consists of circular peptides. All other unmodified, linear, non-pediocin-like single peptides belong to the class IId subgroup (Cotter et al. 2013).
Many bacteriocins have a highly specific mode of action against their bacterial targets, making them potential alternatives to current clinical drugs. In respect of target specificity, a number of bacteriocins demonstrate broad-spectrum antimicrobial activity, albeit narrow-spectrum bacteriocins have been more favoured for the control of targeted pathogens since they do not disrupt the general bacterial communities (Boakes et al. 2012; Rea et al. 2010). In general, bacteriocins from lactic acid bacteria (LAB) are mostly active against other closely related gram-positive bacteria. Although their secretion pathways are not accommodated in this review, it is notable that some other bacteriocins produced by gram-negative bacteria (or termed microcins) show a potential activity against gram-negative bacteria.
Despite an increasing emergence of novel bacteriocins, the knowledge on their transport and processing is lacking. The aim of this review is to summarise all the information till date about the transporters and secretion pathways involved in bacteriocin production from gram-positive bacteria.
Classification of bacteriocin transporters
The genes involved in bacteriocin biosynthesis are commonly assembled and encoded either on plasmids or on the chromosome (Klaenhammer 1993). The biosynthesis cluster typically comprises structural genes, including genes encoding precursor peptide, enzymes involved in post-translational modifications (in class I), a series of transporters with separate or combined protease activity, and enzymes involved in self-immunity. The increasing knowledge on the production of various kinds of bacteriocins has favoured biosynthesis studies by querying related genes (Letzel et al. 2014; Velasquez and van der Donk 2011).
The precursor peptide of bacteriocins contains an N-terminal leader peptide and a C-terminal core peptide (Oman and van der Donk 2010). There are a few exceptions to this rule though, such as bottromycin, which contains a C-terminal extension for removal (Gomez-Escribano et al. 2012) and the leaderless multipeptide bacteriocins (Ovchinnikov et al. 2016). The leader sequence plays an important role in guiding the precursor from beginning of the process till final maturation, by interaction and recognition by correlated enzymes and transporters. In addition, the leader sequence also protects the precursor peptide from degradation and maintains it in an inactive form during the process (Oman and van der Donk 2010; Patton et al. 2008; van Belkum et al. 1997; van der Meer et al. 1994).
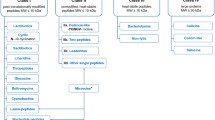
The export of precursor peptides for maturation generally relies on a group of diversified transporters belonging to a large family of ATP-binding cassette (ABC) transporters, which share a common structure and functions involving transportation and ATP hydrolysis (Fath and Kolter 1993; Havarstein et al. 1995). Based on their structure and the signal sequence of their cognitive substrates, bacteriocin secretion transporters in gram-positive bacteria are divided into four different groups (summarised in Table 1 and Fig. 1; for bacteriocin structures and leader, see Fig. S1): NisT-type transporters (Gebhard 2012), SunT-type transporters (Gebhard 2012), sec-dependent pathway (Nes et al. 1996), and other uncharacterised transporters.
Models of different types of transport pathways in bacteriocin production. Main components of three identified major pathways in bacteriocin production are displayed. a NisT type transporter and peptidase. b SunT type transporter. c sec-dependent pathway. Transporters and related components are coloured as blue, peptidases or peptidase region in SunTs are coloured as red, and precursor/matured peptides are coloured as green. Out, extracellular side. In, inside of cells. SP, signal peptidase in sec-dependent pathway
The NisT-type transporters are termed after the transporter involved in nisin production (Kuipers et al. 2004; Qiao and Saris 1996; van den Berg van Saparoea et al. 2008). This group consists of transporters that are approximately 600 amino acids long and are responsible for exporting the precursor peptides to the extracellular region in some lantibiotics (Klein et al. 1992; Meyer et al. 1995; Peschel et al. 1997; Qi et al. 2000; Qiao and Saris 1996). For cleavage of the leader peptide, NisT-type transporters generally require additional enzymes, such as NisP, involved in nisin production because they lack an external peptidase region (van der Meer et al. 1993).
The SunT-type transporters belong to a protein subfamily termed as ABC transporter maturation and secretion (AMS) proteins (Havarstein et al. 1995). Transporters belonging to this class have an additional N-terminal peptidase region and are around 700 amino acids long, due to which they are responsible for both transport and leader peptide removal by class I and II bacteriocins (Birri et al. 2010; Franke et al. 1999; Paik et al. 1998; Xie et al. 2004).
In addition to the above transporters, a number of class II bacteriocins are secreted through a general sec-dependent pathway (Nes et al. 1996; Nes et al. 2001). Recently, it was reported that some novel lantibiotics are likely to rely on this secretion route, in addition to the common NisT or SunT-type transporters (Repka et al. 2017; Zhang et al. 2015).
The lack of detailed information on the secretion/maturation of bacteriocins by their producer strains has limited their industrial applications. These bacteriocins with unclassified transporters are discussed and summarised in the last section.
Roles of NisT-type transporters in lantibiotics
NisT-type transporters are broadly encoded by the structural genes for a number of lantibiotics (Gebhard 2012). The transporters responsible for exporting the precursor peptide are also typically termed as LanT, which are followed by leader signal processing through an additional protease LanP (Fath and Kolter 1993; Hollenstein et al. 2007; McAuliffe et al. 2001). The use of the term LanT, however, can be occasionally confusing, since LanT represents all the transporters related to lantibiotic secretion/maturation, including bifunctional SunT-type transporters and others. This section focuses only on the LanTs related to NisT-type transporters in lantibiotic production.
The NisT-type transporter is a homodimer, with each polypeptide consisting of a hydrophobic transmembrane domain (TMD) and a nucleotide-binding domain (NBD). NisT was the first transporter to be identified in the nisin biosynthesis cluster, and it exhibited sequence similarity with other membrane transporters (Engelke et al. 1992). This was followed by the identification of several other homologous transporters encoded by gene clusters of other lantibiotics (Hillman et al. 1998; Klein et al. 1992; Meyer et al. 1995; Peschel et al. 1997; Qi et al. 2000; van de Kamp et al. 1995).
To understand the process of lantibiotic biosynthesis, several efforts have been made in terms of gene disruption in NisT-type transporters. It has been reported that the inactivation of NisT resulted in the loss of nisin secretion by the cells, leading to detectable nisin activity from cell lysates (Qiao and Saris 1996). In a similar experiment, deletion of the spaT gene from the subtilin biosynthesis cluster led to cell agglutination during the mid-logarithmic growth phase and loss of cell viability, presumably due to intracellular subtilin accumulation (Klein et al. 1992). In-frame deletion of the spaT gene also resulted in a severely abnormal morphology during cell growth (Izaguirre and Hansen 1997). However, lantibiotic production was unaffected in lanT-deleted variant strains of mutacin 1140 (Escano et al. 2015) and epidermin (Schnell et al. 1992). Similarly, a decreased yet consistent production (10%) of Pep5 has been reported in the absence of its transporter PepT, due to other transporters present in the host (Meyer et al. 1995). The fact that these compounds are secreted even in the absence of their respective transporters requires further investigation.
NisT-type transporters have been found to display a broad tolerance towards a number of substrates. They utilise the leader sequence of the precursor peptide for the recognition of these substrates. A number of different peptides (not belonging to the lantibiotic cluster) fused after the cognitive leader signals were successfully secreted outside the cells (Izaguirre and Hansen 1997; Kuipers et al. 2004).
Another intriguing question is whether the NisT-type transporters act alone or they require accessory proteins for lantibiotic secretion. Introduction of a homologous transporter encoding gene gdmT along with a suggested accessory protein gdmH (identified to be an immunity-related protein later (Hille et al. 2001)) from gallidermin clusters in an epiT-deleted variant resulted in an increased epidermin production of up to 7–10 folds (Peschel et al. 1997), possibly due to an increased efficiency of epidermin secretion. However, studies with NisT-type transporters did not reveal any such pattern. Therefore, further in-depth analysis of the genes involved in secretion is required to understand the transport mechanism.
Leader peptide removal after NisTs
The enzyme leading to cleavage of leader peptide in the NisT group is a subtilisin-like serine protease LanP. The LanP proteases differ widely in length from 266 amino acids (LasP) (Skaugen et al. 1997) to 682 amino acids (NisP) (van der Meer et al. 1993).
As a representative LanP enzyme, NisP was first identified by sequence analysis to be composed of four major regions: an N-terminal signal peptide, a prodomain, a serine protease catalytic domain, and a C-terminal domain (~ 110 amino acids) which contains LPXTG to provide anchorage towards the cell wall of the producer strain (Mazmanian et al. 2001). The C-terminal anchor region of NisP is eliminated through auto-cleavage, possibly to free the remaining protease region extracellularly (Xu et al. 2014). A short peptide fragment (from position 635 to 647) was found to be responsible for auto-proteolysis. The crystal structure of the protease region of NisP demonstrated a conformation similar to other S8 clades of subtilisin-like proteases (Kuhn et al. 2014). However, unlike other subtilases that can bind to varying numbers of Ca atoms, the predicted calcium-chelating residues of NisP were found to prevent the binding of calcium ion to NisP in that structure (Xu et al. 2014). High-affinity calcium binding is a common feature of such proteases and is believed to stabilise and activate subtilisin secretion (Bryan et al. 1992).
Some conserved motifs of the leader peptide (such as FNLD motif) are essential and have been demonstrated to contribute to leader peptide removal by NisP for nisin maturation (van der Meer et al. 1994). It also suggests that leader peptide cleavage by NisP occurs as the last step during maturation post secretion. Apart from the leader region of the substrate, the modified core region of NisA was found to play a key role during nisin maturation by NisP (Kuipers et al. 2004). It was found that NisP requires lanthionine formation on the substrate and is unable to cleave off the leader sequence from unmodified NisA lacking a lanthionine ring. This substrate specificity during leader peptide removal is also seen in other LanPs in lantibiotic maturation, such as mutacin 1140 (Escano et al. 2015).
In another study, a second type of LanP, ElxP, which is distinct from NisP, was found to be involved in the maturation of epilancin 15X with low substrate specificity (Ortega et al. 2014; Velasquez et al. 2011). The amino acid sequence of ElxP is similar to the protease EciP, which is involved in the biosynthesis of epicidin 280 (Heidrich et al. 1998). There are two major differences in the structures of ElxP from NisP. First, unlike the auto-cleaved extracellular enzyme NisP, ElxP lacks an N-terminal secretion signal and a C-terminal cell wall anchor sequence (LPXTG). This implies that ElxP is more likely to be functional inside the cytoplasm, where it cleaves off the leader peptide prior to export by ElxT. The second difference lies in the conserved sequences near the cleavage site between leader and core peptide of substrates (Ortega et al. 2014). Such differences among LanPs are yet to be investigated.
Bifunctional transporters in various bacteriocins
In the SunT group, proteins contain an additional N-terminal protease to form an integrated bifunctional transporter, termed as AMS protein (Havarstein et al. 1995). Based on structural analysis, SunT transporters have been demonstrated to consist of three major regions: an N-terminal peptidase domain (PEP) for leader peptide cleavage, a C-terminal NBD for ATP hydrolysis, and a TMD inserted across cell membranes leading to export of the substrate. The SunTs are responsible for both the secretion and maturation in various kinds of bacteriocins, such as NukT for lantibiotic nukacin ISK-1 (Kimura et al. 1997), SunT for glycocin sublancin 168 (Paik et al. 1998), PedD for class IIa bacteriocin pediocin PA-1 (Oppegard et al. 2015), LagD for class IIb lactococcin G (Nissen-Meyer et al. 1992), and LcnC for class IId lactococcin A (Stoddard et al. 1992). Apart from bacteriocin secretion, the bifunctional transporters are also involved in the transport of other peptide products across bacterial species, such as bacteria-communication signal peptide ComC (Hui et al. 1995).
To investigate how secretion and maturation of substrates with widely diversified structures occurs, a number of approaches have been adopted to analyse the different domains of SunTs. In studies with the PEP region, LagD was first characterised to be capable of in vitro cleavage leading to lactococcin G production (Havarstein et al. 1995). Following that, more isolated PEPs of SunTs were investigated to offer insights into the production of bacteriocins and other proteins, such as quorum sensing signal transporter ComA (Ishii et al. 2010; Ishii et al. 2006), transporter CvaB for colicin V secretion (Wu and Tai 2004), and lacticin 481 transporter LctT (Furgerson Ihnken et al. 2008). Crystal structure analysis of PEP-ComA has revealed an overall structure similar to the papain-like protease family (Ishii et al. 2010). As for the catalytic mechanism, Cys17 and His96 form as a thiolate-imidazolium ion pair, and Asp112 functions to stabilise His96 during processing. Further, the conserved residues in the leader sequence of substrate ComC are presumed to form an essential α-helical structure and interact with a predicted hydrophobic surface proximal to the active site of ComA (Ishii et al. 2010). ATP binds to the NBD region, followed by its hydrolysis to provide energy for constructional shifting. This structure is highly conserved among ABC transporters. Fragment analysis experiment of ComA revealed that in SunT-type transporters, a highly conserved glycine residue is critical for functioning of the NBD region (Ishii et al. 2013). At last, research on the TMD region is comparably falling behind, perhaps caused by the lack of identities in this region among SunTs. Structural analysis of LcnC indicated that TMD comprises six transmembrane segments and is mainly responsible for exporting the matured substrate (Franke et al. 1999).
SunTs have been found to be present in conjunction with an accessory protein during the secretion of products, such as LcnD for LcnC in lactococcin A secretion (Stoddard et al. 1992), PedC for PedD in pediocin PA-1 (Oppegard et al. 2015), and ComB for ComA in quorum sensing signal peptide ComC (Hui et al. 1995). However, heterologous expression of pediocin PA-1 revealed that structural genes, in addition to the accessory protein, are sufficient for pediocin secretion, and that the accessory protein had no influence on increasing the production (Mesa-Pereira et al. 2017). Structural gene analysis and review of the production experiments have suggested that many members of SunT-type transporters lack accessory proteins and can still conduct secretion, such as NukT, SunT, and LctT in class I bacteriocin, and StxT in class IIa and IIb bacteriocin (Gebhard 2012). Although a precise mechanism of action of the accessory protein remains unclear, it seems that the majority of SunT-type transporters are capable of bacteriocin secretion and maturation independently.
Leader removal and transport mechanisms in SunT
Similar to the NisT group, the leader sequence also plays an important role in protection and directing the precursor molecule for secretion and maturation in SunT-type transporters (Xie et al. 2004). The leader sequence of the substrate is rich in aspartate and glutamate residues, and has a conserved ELXXBX (B=V, L or I) motif, with a double glycine motif ahead of the cleavage site (Oman and van der Donk 2010). Notably, this double glycine motif has been reported to be critical for leader peptide processing by transporters (Chen et al. 2001; Furgerson Ihnken et al. 2008), but not for incorporating the lanthionine ring by modification enzymes in lantibiotics (Patton et al. 2008). In addition, the double glycine box is found conserved across class II bacteriocins, which do not undergo post-modifications (Ennahar et al. 2000; Oppegard et al. 2015; Vaughan et al. 2003).
In studies demonstrating heterologous expression of SunT transporters, the leader sequence has been shown to be involved in the production of original bacteriocin peptides as well as fused peptides containing other kinds of bacteriocins from different secretion pathways (Allison et al. 1995; Rodriguez et al. 2003; van Belkum et al. 1997). Likewise, it has also been demonstrated that the leader sequence may be essential in protecting the precursor peptide from degrading and guiding the products through secretion by SunTs (Sprules et al. 2004). However, it became controversial in the production analysis concerning substrate specificity, which displayed broad differences among bacteriocins. For example, the lacticin 481 transporter LctT was involved in the production of nukacin ISK-1 instead of NukT, but not vice versa (Nagao et al. 2007). Likewise, LcnC could not be replaced by LctT in lantibiotic processing and production (Uguen and Uguen 2002). In terms of future applications, potential solutions may rely on more dedicated focus on understanding the transport mechanisms of SunTs, which are barely known so far.
As compared to the extensive research in leader peptides, studies on SunTs have had to confront much more practical problems, especially in terms of experimental difficulties to investigate membrane proteins for their structure and function. In a recent study, a putative AMS protein has been reported with its full-length protein structure, which is likely to offer new insights into domain-to-domain cooperation (Lin et al. 2015). This AMS-like protein is discovered from the gram-positive bacterium Clostridium thermocellum and displays two crystal structures in the presence and absence of ATP binding. In another report, the transporter NukT for lantibiotic nukacin ISK-1 production was characterised to recognise lanthionine rings of its substrate. The study also revealed that ATP is necessary for cleavage of the leader peptide (Nishie et al. 2011; Nishie et al. 2009). According to a recent study, a potential mutual regulation of PEP and NBD has been suggested through the analysis of full-length NukT protein (Zheng et al. 2018). Based on the above findings, it can be concluded that these bifunctional transporters utilise potentially dynamic transport machinery involving domain-to-domain interaction for improving the production efficiency.
Sec-dependent pathway utilised by bacteriocins
Apart from SunTs, a number of class II bacteriocins are secreted across cell membranes by a conserved and essential sec-dependent pathway. Some examples of such bacteriocins include the class IIa bacteriocin 31 (Tomita et al. 1996), enterocin P (Cintas et al. 1997), bacteriocin T8 (De Kwaadsteniet et al. 2006), and class IId divergicin A (Worobo et al. 1995), and lactococcin 972 (Martinez et al. 1999). Recently, some novel lantibiotics, such as duramycin and cinnamycin, were also reported to possibly utilise the sec-dependent pathway in addition to the regular NisT or SunT transporter systems (Huo et al. 2017; Widdick et al. 2003).
The sec-dependent pathway was first characterised in gram-negative bacteria (Duong et al. 1997; Pugsley 1993). In the most studied gram-negative bacterium, E. coli, a sec transport complex is composed of SecYEG, supplemented with membrane protein SecDF and an ATPase SecA (Duong et al. 1997). After secretion, the precursor peptide is processed by a protease to remove the leader sequence (Dalbey and Wickner 1985). Using this information, homologous and other related genes of the sec-dependent pathway were studied in gram-positive bacteria (Caspers and Freudl 2008; van Roosmalen et al. 2004). It was found that while the genes encoding SecA and SecY are well conserved, the genes for SecE and SecG are shortened in gram-positive bacteria compared to E. coli homologues (Lee and Schneewind 2001).
The bacteriocin precursor peptides in the sec-dependent pathway possess a diverse N-terminal signal, with varying lengths and amino acid sequences. Although the signal sequence of the precursor peptide is not conserved, leader removal is processed by signal peptidase I (SPI) or signal peptidase II (SPII) depending on its cleavage site (Schneewind and Missiakas 2014). For example, SpsB was identified and characterised to be a SPI protease for leader processing in Staphylococcus aureus (Cregg et al. 1996).
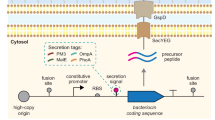
The sec-dependent pathway has also been studied and utilised for heterologous expression in various bacteriocins (Herranz and Driessen 2005; McCormick et al. 1996; Worobo et al. 1995). In addition, abundant efforts have been made to examine the best and most efficient bacteriocin production by analysing different transporter systems, leader peptides, or a combination of the two. For example, the SunT transporter combined with double glycine leader sequence exhibited increased production levels than the general secretion pathway for mesentericin Y105 (Biet et al. 1998). On the other hand, a sec-dependent signal can replace the original double glycine type leader and result in sufficient bacteriocin production (Martin et al. 2007; McCormick et al. 1996). With the increased understanding of varied transport systems, different strategies can be designed for efficient industrial bacteriocin production.
Other unclassified transporters in bacteriocins
In many cases, bacteriocins have been reported to possess potent antimicrobial activity against target strains. However, further application of such bacteriocins is limited due to the lack of information about their biosynthesis mechanisms and detailed mode of action. The information on bacteriocins transported by unidentified transporters or uncharacterised mechanism is discussed in this section.
The bottromycin family of bacteriocins was first identified to be produced from Streptomyces bottropensis (Waisvisz et al. 1957). After decades of research, bottromycin is finally characterised to belong to a group of ribosomally synthesised antimicrobial peptides containing macrocyclic amidine and unusual β-methylated amino acid residues (Shimamura et al. 2009). Bottromycin A2 is the most characterised bacteriocin from this class, with potential antimicrobial activities against many drug-resistant pathogens such as methicillin-resistant S. aureus (MRSA) and vancomycin-resistant Enterococcus (VRE) (Shimamura et al. 2009). Genome sequencing was utilised to identify and understand the biosynthesis mechanism of bottromycin A2 and its derivatives (Hou et al. 2012; Zhang et al. 2013). However, the information on transport and secretion of bottromycin is still lacking. A gene botT has been proposed to encode a multidrug transporter but contains no ATP-binding domain. As a result of heterologous expression, BotT was presumed to play a key role in self-resistance and was speculated to influence greater production (Huo et al. 2012). In addition, the precursor peptide of bottromycin has a unique C-terminal extension, which serves as the signal peptide throughout the biosynthesis process, instead of the typical N-terminal leader in other bacteriocins. The C-“follower” peptide is cleaved off to liberate the N-terminal core peptide before macrocyclisation by the protease BotP (Mann et al. 2016).
The thiopeptides (or thiazolyl peptides) are a group of small heavily modified peptides (12 to 17 amino acid residues), containing sulphur-containing heterocyclic rings, dehydrated residues, and macrocyclic structure (Arnison et al. 2013). Since the first report on thiopeptide micrococcin (Su 1948), a large number of novel variants have been discovered for this family (Arnison et al. 2013). Thiopeptides have been well known for their unique inhibition activity against protein synthesis in many bacterial strains, including the multidrug-resistant pathogens MRSA and VRE (Bagley et al. 2005). Although the essential structural genes for their biosynthesis have been studied (Bowers et al. 2010; Hayashi et al. 2014; Young and Walsh 2011), the precise transport mechanism of thiopeptides has not been understood as yet. Among the structural genes, two potential proteins involved in efflux of products have been discovered (Hayashi et al. 2014). These proteins exhibit similarity to other known transporters, such as the putative ABC transporter tclW and efflux pump tclX in thiocillin (Bowers et al. 2010; Wieland Brown et al. 2009), and ABC transporter-like getB/C in GE37468 (Young and Walsh 2011). Interestingly, some thiopeptides have a double glycine-like motif ahead of the cleavage site within leader sequence (Oman and van der Donk 2010). For removal of the leader sequence, the N-terminal leader signal is cleaved off by an α/β-hydrolase fold protein, TsrB, in order to form a macrocycle structure during thiostrepton biosynthesis (Zheng et al. 2016).
Sactibiotics (or sactipeptides) are a small group of post-translationally modified peptides that contain a sulphur-to-α-carbon link (Arnison et al. 2013). Subtilosin A is the best-studied sactibiotic and is identified to be a negatively charged circular peptide produced by Bacillus subtilis (Babasaki et al. 1985). Bacillus thuringiensis produces a two-peptide sactibiotic thuricin CD, having a narrow spectrum of activity against Clostridium difficile (Rea et al. 2010). Thuricin H is also produced by B. thuringiensis as a single-peptide sactibiotic (Lee et al. 2009). Subtilosin A was first thought to belong to class IIc (circular) bacteriocins. However, after further characterisation, it was classified under a new subgroup (sactibiotics) along with other bacteriocins because of macrocyclisation and thioether installation (Cotter et al. 2013; Rea et al. 2011). The study of subtilosin A biosynthesis (Fluhe et al. 2012; Zheng et al. 2000) revealed that the leader peptide of precursor peptide is cleaved off by a putative protease AlbE or AlbF between installation of the thioether bond by a radical S-adenosylmethionine (SAM) enzyme AlbA and final macrocyclisation. Post macrocyclisation, a putative ABC transporter AlbC exports the mature subtilosin A.
Linaridin is a small group of recently discovered post-translationally modified peptides having a linear structure with dehydrated amino acids, which are similar to the lantibiotics but follow a different biosynthesis pathway (Arnison et al. 2013). Without detection of typical modification enzymes in lantibiotics, the structural gene cluster for a linaridin cypemycin (21 amino acid residues) was identified (Claesen and Bibb 2010). Genome sequencing and mutagenesis experiments have brought in new insights to the biosynthesis studies of cypemycin (Mo et al. 2017). Although the precise transporter and protease remain unknown, it was hypothesised that a gene cypT encoding an ATP-binding subunit of ABC transporter leads to the export of cypemycin through the cell membrane (Claesen and Bibb 2010).
Although most of the class II bacteriocins rely on bifunctional ABC transporters or sec-dependent pathways, the export and maturation of many others still remains unknown. For instance, the dedicated transport and leader cleavage mechanism of class IIc bacteriocins has not been understood as yet (Gabrielsen et al. 2014). By means of heterologous expression studies of enterocin AS-48, a series of genes including as-48BCC 1 D have been found to be essential for production and secretion (Martinez-Bueno et al. 1998). Following this, many studies focused on other circular bacteriocins, such as circularin A (Kemperman et al. 2003), carnocyclin A (van Belkum et al. 2010), and enterocin NKR-5-3B (Perez et al. 2015). It can, therefore, be hypothesised that a complex of several proteins, including membrane proteins and ATPases, may be involved in circularisation and transportation in class IIc bacteriocins.
With recent improvements and updates on novel bacteriocins, a group of leaderless multipeptide bacteriocins do not fit into the classification described above (Ovchinnikov et al. 2016). These bacteriocins share similar biosynthesis-related structural genes and do not contain an N-terminal leader sequence that is commonly found in other bacteriocins. One such bacteriocin, the four-peptide aureocin A70, requires an ABC transporter AurT for its production by S. aureus A70 (Netz et al. 2001). Interestingly, AurT also exhibits phylogenetic similarity with the NisT-type transporters on both NBD and TMD regions (Gebhard 2012).
Concluding remarks
Bacteriocins have emerged as a potent and attractive alternative to antibiotics owing to their numerous desirable characteristics. Current research primarily focuses on their post-modification, mode of action, and immunity mechanism. However, the secretion and maturation of bacteriocins with diverse structures and functions by the numerous transporters and secretion pathways have not been characterised well. In this review, we summarised different types of transporters (ABC transporters especially) to explain the processing and export of various bacteriocins, which may prove to be fundamental in understanding their secretion and maturation and also contribute to their increased production by the strains.
References
Allison GE, Worobo RW, Stiles ME, Klaenhammer TR (1995) Heterologous expression of the lactacin F peptides by Carnobacterium piscicola LV17. Appl Environ Microbiol 61(4):1371–1377
Arnison PG, Bibb MJ, Bierbaum G, Bowers AA, Bugni TS, Bulaj G, Camarero JA, Campopiano DJ, Challis GL, Clardy J, Cotter PD, Craik DJ, Dawson M, Dittmann E, Donadio S, Dorrestein PC, Entian KD, Fischbach MA, Garavelli JS, Goransson U, Gruber CW, Haft DH, Hemscheidt TK, Hertweck C, Hill C, Horswill AR, Jaspars M, Kelly WL, Klinman JP, Kuipers OP, Link AJ, Liu W, Marahiel MA, Mitchell DA, Moll GN, Moore BS, Muller R, Nair SK, Nes IF, Norris GE, Olivera BM, Onaka H, Patchett ML, Piel J, Reaney MJ, Rebuffat S, Ross RP, Sahl HG, Schmidt EW, Selsted ME, Severinov K, Shen B, Sivonen K, Smith L, Stein T, Sussmuth RD, Tagg JR, Tang GL, Truman AW, Vederas JC, Walsh CT, Walton JD, Wenzel SC, Willey JM, van der Donk WA (2013) Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat Prod Rep 30(1):108–160. https://doi.org/10.1039/c2np20085f
Babasaki K, Takao T, Shimonishi Y, Kurahashi K (1985) Subtilosin A, a new antibiotic peptide produced by Bacillus subtilis 168: isolation, structural analysis, and biogenesis. J Biochem 98(3):585–603
Bagley MC, Dale JW, Merritt EA, Xiong X (2005) Thiopeptide antibiotics. Chem Rev 105(2):685–714. https://doi.org/10.1021/cr0300441
Biet F, Berjeaud JM, Worobo RW, Cenatiempo Y, Fremaux C (1998) Heterologous expression of the bacteriocin mesentericin Y105 using the dedicated transport system and the general secretion pathway. Microbiology 144(Pt 10):2845–2854. https://doi.org/10.1099/00221287-144-10-2845
Birri DJ, Brede DA, Forberg T, Holo H, Nes IF (2010) Molecular and genetic characterization of a novel bacteriocin locus in Enterococcus avium isolates from infants. Appl Environ Microbiol 76(2):483–492. https://doi.org/10.1128/AEM.01597-09
Boakes S, Ayala T, Herman M, Appleyard AN, Dawson MJ, Cortes J (2012) Generation of an actagardine A variant library through saturation mutagenesis. Appl Microbiol Biotechnol 95(6):1509–1517. https://doi.org/10.1007/s00253-012-4041-0
Bowers AA, Acker MG, Koglin A, Walsh CT (2010) Manipulation of thiocillin variants by prepeptide gene replacement: structure, conformation, and activity of heterocycle substitution mutants. J Am Chem Soc 132(21):7519–7527. https://doi.org/10.1021/ja102339q
Bryan P, Alexander P, Strausberg S, Schwarz F, Lan W, Gilliland G, Gallagher DT (1992) Energetics of folding subtilisin BPN'. Biochemistry 31(21):4937–4945
Caspers M, Freudl R (2008) Corynebacterium glutamicum possesses two secA homologous genes that are essential for viability. Arch Microbiol 189(6):605–610. https://doi.org/10.1007/s00203-008-0351-0
Chen P, Qi FX, Novak J, Krull RE, Caufield PW (2001) Effect of amino acid substitutions in conserved residues in the leader peptide on biosynthesis of the lantibiotic mutacin II. FEMS Microbiol Lett 195(2):139–144
Cintas LM, Casaus P, Havarstein LS, Hernandez PE, Nes IF (1997) Biochemical and genetic characterization of enterocin P, a novel sec-dependent bacteriocin from Enterococcus faecium P13 with a broad antimicrobial spectrum. Appl Environ Microbiol 63(11):4321–4330
Claesen J, Bibb M (2010) Genome mining and genetic analysis of cypemycin biosynthesis reveal an unusual class of posttranslationally modified peptides. Proc Natl Acad Sci U S A 107(37):16297–16302. https://doi.org/10.1073/pnas.1008608107
Cotter PD, Hill C, Ross RP (2005) Bacteriocins: developing innate immunity for food. Nat Rev Microbiol 3(10):777–788. https://doi.org/10.1038/nrmicro1273
Cotter PD, Ross RP, Hill C (2013) Bacteriocins—a viable alternative to antibiotics? Nat Rev Microbiol 11(2):95–105. https://doi.org/10.1038/nrmicro2937
Cregg KM, Wilding I, Black MT (1996) Molecular cloning and expression of the spsB gene encoding an essential type I signal peptidase from Staphylococcus aureus. J Bacteriol 178(19):5712–5718
Dalbey RE, Wickner W (1985) Leader peptidase catalyzes the release of exported proteins from the outer surface of the Escherichia coli plasma membrane. J Biol Chem 260(29):15925–15931
De Kwaadsteniet M, Fraser T, Van Reenen CA, Dicks LM (2006) Bacteriocin T8, a novel class IIa sec-dependent bacteriocin produced by Enterococcus faecium T8, isolated from vaginal secretions of children infected with human immunodeficiency virus. Appl Environ Microbiol 72(7):4761–4766. https://doi.org/10.1128/AEM.00436-06
Duong F, Eichler J, Price A, Leonard MR, Wickner W (1997) Biogenesis of the gram-negative bacterial envelope. Cell 91(5):567–573
Engelke G, Gutowski-Eckel Z, Hammelmann M, Entian KD (1992) Biosynthesis of the lantibiotic nisin: genomic organization and membrane localization of the NisB protein. Appl Environ Microbiol 58(11):3730–3743
Ennahar S, Sashihara T, Sonomoto K, Ishizaki A (2000) Class IIa bacteriocins: biosynthesis, structure and activity. FEMS Microbiol Rev 24(1):85–106
Escano J, Stauffer B, Brennan J, Bullock M, Smith L (2015) Biosynthesis and transport of the lantibiotic mutacin 1140 produced by Streptococcus mutans. J Bacteriol 197(7):1173–1184. https://doi.org/10.1128/JB.02531-14
Fath MJ, Kolter R (1993) ABC transporters: bacterial exporters. Microbiol Rev 57(4):995–1017
Fluhe L, Knappe TA, Gattner MJ, Schafer A, Burghaus O, Linne U, Marahiel MA (2012) The radical SAM enzyme AlbA catalyzes thioether bond formation in subtilosin A. Nat Chem Biol 8(4):350–357. https://doi.org/10.1038/nchembio.798
Franke CM, Tiemersma J, Venema G, Kok J (1999) Membrane topology of the lactococcal bacteriocin ATP-binding cassette transporter protein LcnC. Involvement of LcnC in lactococcin a maturation. J Biol Chem 274(13):8484–8490
Furgerson Ihnken LA, Chatterjee C, van der Donk WA (2008) In vitro reconstitution and substrate specificity of a lantibiotic protease. Biochemistry 47(28):7352–7363. https://doi.org/10.1021/bi800278n
Gabrielsen C, Brede DA, Nes IF, Diep DB (2014) Circular bacteriocins: biosynthesis and mode of action. Appl Environ Microbiol 80(22):6854–6862. https://doi.org/10.1128/AEM.02284-14
Gebhard S (2012) ABC transporters of antimicrobial peptides in Firmicutes bacteria—phylogeny, function and regulation. Mol Microbiol 86(6):1295–1317. https://doi.org/10.1111/mmi.12078
Gomez-Escribano JP, Song LJ, Bibb MJ, Challis GL (2012) Posttranslational β-methylation and macrolactamidination in the biosynthesis of the bottromycin complex of ribosomal peptide antibiotics. Chem Sci 3(12):3522–3525. https://doi.org/10.1039/c2sc21183a
Havarstein LS, Diep DB, Nes IF (1995) A family of bacteriocin ABC transporters carry out proteolytic processing of their substrates concomitant with export. Mol Microbiol 16(2):229–240
Hayashi S, Ozaki T, Asamizu S, Ikeda H, Omura S, Oku N, Igarashi Y, Tomoda H, Onaka H (2014) Genome mining reveals a minimum gene set for the biosynthesis of 32-membered macrocyclic thiopeptides lactazoles. Chem Biol 21(5):679–688. https://doi.org/10.1016/j.chembiol.2014.03.008
Heidrich C, Pag U, Josten M, Metzger J, Jack RW, Bierbaum G, Jung G, Sahl HG (1998) Isolation, characterization, and heterologous expression of the novel lantibiotic epicidin 280 and analysis of its biosynthetic gene cluster. Appl Environ Microbiol 64(9):3140–3146
Herranz C, Driessen AJ (2005) Sec-mediated secretion of bacteriocin enterocin P by Lactococcus lactis. Appl Environ Microbiol 71(4):1959–1963. https://doi.org/10.1128/AEM.71.4.1959-1963.2005
Hille M, Kies S, Gotz F, Peschel A (2001) Dual role of GdmH in producer immunity and secretion of the staphylococcal lantibiotics gallidermin and epidermin. Appl Environ Microbiol 67(3):1380–1383. https://doi.org/10.1128/AEM.67.3.1380-1383.2001
Hillman JD, Novak J, Sagura E, Gutierrez JA, Brooks TA, Crowley PJ, Hess M, Azizi A, Leung K, Cvitkovitch D, Bleiweis AS (1998) Genetic and biochemical analysis of mutacin 1140, a lantibiotic from Streptococcus mutans. Infect Immun 66(6):2743–2749
Hollenstein K, Dawson RJ, Locher KP (2007) Structure and mechanism of ABC transporter proteins. Curr Opin Struct Biol 17(4):412–418. https://doi.org/10.1016/j.sbi.2007.07.003
Hou Y, Tianero MD, Kwan JC, Wyche TP, Michel CR, Ellis GA, Vazquez-Rivera E, Braun DR, Rose WE, Schmidt EW, Bugni TS (2012) Structure and biosynthesis of the antibiotic bottromycin D. Org Lett 14(19):5050–5053. https://doi.org/10.1021/ol3022758
Hui FM, Zhou L, Morrison DA (1995) Competence for genetic transformation in Streptococcus pneumoniae: organization of a regulatory locus with homology to two lactococcin A secretion genes. Gene 153(1):25–31
Huo L, Rachid S, Stadler M, Wenzel SC, Muller R (2012) Synthetic biotechnology to study and engineer ribosomal bottromycin biosynthesis. Chem Biol 19(10):1278–1287. https://doi.org/10.1016/j.chembiol.2012.08.013
Huo L, Okesli A, Zhao M, van der Donk WA (2017) Insights into the biosynthesis of duramycin. Appl Environ Microbiol 83(3):e02698–e02616. https://doi.org/10.1128/AEM.02698-16
Ishii S, Yano T, Hayashi H (2006) Expression and characterization of the peptidase domain of Streptococcus pneumoniae ComA, a bifunctional ATP-binding cassette transporter involved in quorum sensing pathway. J Biol Chem 281(8):4726–4731. https://doi.org/10.1074/jbc.M512516200
Ishii S, Yano T, Ebihara A, Okamoto A, Manzoku M, Hayashi H (2010) Crystal structure of the peptidase domain of Streptococcus ComA, a bifunctional ATP-binding cassette transporter involved in the quorum-sensing pathway. J Biol Chem 285(14):10777–10785. https://doi.org/10.1074/jbc.M109.093781
Ishii S, Yano T, Okamoto A, Murakawa T, Hayashi H (2013) Boundary of the nucleotide-binding domain of Streptococcus ComA based on functional and structural analysis. Biochemistry 52(15):2545–2555. https://doi.org/10.1021/bi3017069
Izaguirre G, Hansen JN (1997) Use of alkaline phosphatase as a reporter polypeptide to study the role of the subtilin leader segment and the SpaT transporter in the posttranslational modifications and secretion of subtilin in Bacillus subtilis 168. Appl Environ Microbiol 63(10):3965–3971
Kemperman R, Jonker M, Nauta A, Kuipers OP, Kok J (2003) Functional analysis of the gene cluster involved in production of the bacteriocin circularin A by Clostridium beijerinckii ATCC 25752. Appl Environ Microbiol 69(10):5839–5848
Kimura H, Nagano R, Matsusaki H, Sonomoto K, Ishizaki A (1997) A bacteriocin of strain Pediococcus sp. ISK-1 isolated from Nukadoko, bed of fermented rice bran. Biosci Biotechnol Biochem 61(6):1049–1051
Klaenhammer TR (1993) Genetics of bacteriocins produced by lactic acid bacteria. FEMS Microbiol Rev 12(1–3):39–85
Klein C, Kaletta C, Schnell N, Entian KD (1992) Analysis of genes involved in biosynthesis of the lantibiotic subtilin. Appl Environ Microbiol 58(1):132–142
Kuhn ML, Prachi P, Minasov G, Shuvalova L, Ruan J, Dubrovska I, Winsor J, Giraldi M, Biagini M, Liberatori S, Savino S, Bagnoli F, Anderson WF, Grandi G (2014) Structure and protective efficacy of the Staphylococcus aureus autocleaving protease EpiP. FASEB J 28(4):1780–1793. https://doi.org/10.1096/fj.13-241737
Kuipers A, de Boef E, Rink R, Fekken S, Kluskens LD, Driessen AJM, Leenhouts K, Kuipers OP, Moll GN (2004) NisT, the transporter of the lantibiotic nisin, can transport fully modified, dehydrated, and unmodified prenisin and fusions of the leader peptide with non-lantibiotic peptides. J Biol Chem 279(21):22176–22182. https://doi.org/10.1074/jbc.M312789200
Lee VT, Schneewind O (2001) Protein secretion and the pathogenesis of bacterial infections. Genes Dev 15(14):1725–1752. https://doi.org/10.1101/gad.896801
Lee H, Churey JJ, Worobo RW (2009) Biosynthesis and transcriptional analysis of thurincin H, a tandem repeated bacteriocin genetic locus, produced by Bacillus thuringiensis SF361. FEMS Microbiol Lett 299(2):205–213. https://doi.org/10.1111/j.1574-6968.2009.01749.x
Letzel AC, Pidot SJ, Hertweck C (2014) Genome mining for ribosomally synthesized and post-translationally modified peptides (RiPPs) in anaerobic bacteria. BMC Genomics 15:983. https://doi.org/10.1186/1471-2164-15-983
Lin DY, Huang S, Chen J (2015) Crystal structures of a polypeptide processing and secretion transporter. Nature 523(7561):425–430. https://doi.org/10.1038/nature14623
Mann G, Huo L, Adam S, Nardone B, Vendome J, Westwood NJ, Muller R, Koehnke J (2016) Structure and substrate recognition of the bottromycin maturation enzyme BotP. Chembiochem 17(23):2286–2292. https://doi.org/10.1002/cbic.201600406
Martin M, Gutierrez J, Criado R, Herranz C, Cintas LM, Hernandez PE (2007) Chimeras of mature pediocin PA-1 fused to the signal peptide of enterocin P permits the cloning, production, and expression of pediocin PA-1 in Lactococcus lactis. J Food Prot 70(12):2792–2798
Martinez B, Fernandez M, Suarez JE, Rodriguez A (1999) Synthesis of lactococcin 972, a bacteriocin produced by Lactococcus lactis IPLA 972, depends on the expression of a plasmid-encoded bicistronic operon. Microbiology 145(Pt 11):3155–3161. https://doi.org/10.1099/00221287-145-11-3155
Martinez-Bueno M, Valdivia E, Galvez A, Coyette J, Maqueda M (1998) Analysis of the gene cluster involved in production and immunity of the peptide antibiotic AS-48 in Enterococcus faecalis. Mol Microbiol 27(2):347–358
Mazmanian SK, Ton-That H, Schneewind O (2001) Sortase-catalysed anchoring of surface proteins to the cell wall of Staphylococcus aureus. Mol Microbiol 40(5):1049–1057
McAuliffe O, Ross RP, Hill C (2001) Lantibiotics: structure, biosynthesis and mode of action. FEMS Microbiol Rev 25(3):285–308
McCormick JK, Worobo RW, Stiles ME (1996) Expression of the antimicrobial peptide carnobacteriocin B2 by a signal peptide-dependent general secretory pathway. Appl Environ Microbiol 62(11):4095–4099
Mesa-Pereira B, O'Connor PM, Rea MC, Cotter PD, Hill C, Ross RP (2017) Controlled functional expression of the bacteriocins pediocin PA-1 and bactofencin A in Escherichia coli. Sci Rep 7(1):3069. https://doi.org/10.1038/s41598-017-02868-w
Meyer C, Bierbaum G, Heidrich C, Reis M, Suling J, Iglesias-Wind MI, Kempter C, Molitor E, Sahl HG (1995) Nucleotide sequence of the lantibiotic Pep5 biosynthetic gene cluster and functional analysis of PepP and PepC. Evidence for a role of PepC in thioether formation. Eur J Biochem 232(2):478–489
Mo TL, Liu WQ, Ji WJ, Zhao JF, Chen T, Ding W, Yu SN, Zhang Q (2017) Biosynthetic insights into linaridin natural products from genome mining and precursor peptide mutagenesis. ACS Chem Biol 12(6):1484–1488. https://doi.org/10.1021/acschembio.7b00262
Nagao J, Aso Y, Shioya K, Nakayama J, Sonomoto K (2007) Lantibiotic engineering: molecular characterization and exploitation of lantibiotic-synthesizing enzymes for peptide engineering. J Mol Microbiol Biotechnol 13(4):235–242. https://doi.org/10.1159/000104749
Nes IF, Diep DB, Havarstein LS, Brurberg MB, Eijsink V, Holo H (1996) Biosynthesis of bacteriocins in lactic acid bacteria. Anton Leeuw 70(2–4):113–128
Nes IF, Holo H, Fimland G, Hauge HH, Nissen-Meyer J (2001) Unmodified peptide-bacteriocins (class II) produced by lactic acid bacteria. In: Dutton CJ, Haxell MA, McArthur HAI, Wax RG (eds) Peptide antibiotics: discovery, modes of action and applications. Marcel Dekker, Inc, New York. pp 81–115
Netz DJ, Sahl HG, Marcelino R, dos Santos NJ, de Oliveira SS, Soares MB, do Carmo de Freire Bastos M (2001) Molecular characterisation of aureocin A70, a multi-peptide bacteriocin isolated from Staphylococcus aureus. J Mol Biol 311(5):939–949. https://doi.org/10.1006/jmbi.2001.4885
Nishie M, Shioya K, Nagao J, Jikuya H, Sonomoto K (2009) ATP-dependent leader peptide cleavage by NukT, a bifunctional ABC transporter, during lantibiotic biosynthesis. J Biosci Bioeng 108(6):460–464. https://doi.org/10.1016/j.jbiosc.2009.06.002
Nishie M, Sasaki M, Nagao J, Zendo T, Nakayama J, Sonomoto K (2011) Lantibiotic transporter requires cooperative functioning of the peptidase domain and the ATP binding domain. J Biol Chem 286(13):11163–11169. https://doi.org/10.1074/jbc.M110.212704
Nissen-Meyer J, Holo H, Havarstein LS, Sletten K, Nes IF (1992) A novel lactococcal bacteriocin whose activity depends on the complementary action of two peptides. J Bacteriol 174(17):5686–5692
Oman TJ, van der Donk WA (2010) Follow the leader: the use of leader peptides to guide natural product biosynthesis. Nat Chem Biol 6(1):9–18. https://doi.org/10.1038/nchembio.286
Oppegard C, Fimland G, Anonsen JH, Nissen-Meyer J (2015) The pediocin PA-1 accessory protein ensures correct disulfide bond formation in the antimicrobial peptide pediocin PA-1. Biochemistry 54(19):2967–2974. https://doi.org/10.1021/acs.biochem.5b00164
Ortega MA, Velasquez JE, Garg N, Zhang Q, Joyce RE, Nair SK, van der Donk WA (2014) Substrate specificity of the lanthipeptide peptidase ElxP and the oxidoreductase ElxO. ACS Chem Biol 9(8):1718–1725. https://doi.org/10.1021/cb5002526
Ovchinnikov KV, Chi H, Mehmeti I, Holo H, Nes IF, Diep DB (2016) Novel group of leaderless multipeptide bacteriocins from gram-positive bacteria. Appl Environ Microbiol 82(17):5216–5224. https://doi.org/10.1128/AEM.01094-16
Paik SH, Chakicherla A, Hansen JN (1998) Identification and characterization of the structural and transporter genes for, and the chemical and biological properties of, sublancin 168, a novel lantibiotic produced by Bacillus subtilis 168. J Biol Chem 273(36):23134–23142
Patton GC, Paul M, Cooper LE, Chatterjee C, van der Donk WA (2008) The importance of the leader sequence for directing lanthionine formation in lacticin 481. Biochemistry 47(28):7342–7351. https://doi.org/10.1021/bi800277d
Perez RH, Ishibashi N, Inoue T, Himeno K, Masuda Y, Sawa N, Zendo T, Wilaipun P, Leelawatcharamas V, Nakayama J, Sonomoto K (2015) Functional analysis of genes involved in the biosynthesis of enterocin NKR-5-3B, a novel circular bacteriocin. J Bacteriol 198(2):291–300. https://doi.org/10.1128/JB.00692-15
Peschel A, Schnell N, Hille M, Entian KD, Gotz F (1997) Secretion of the lantibiotics epidermin and gallidermin: sequence analysis of the genes gdmT and gdmH, their influence on epidermin production and their regulation by EpiQ. Mol Gen Genet 254(3):312–318
Pugsley AP (1993) The complete general secretory pathway in gram-negative bacteria. Microbiol Rev 57(1):50–108
Qi F, Chen P, Caufield PW (2000) Purification and biochemical characterization of mutacin I from the group I strain of Streptococcus mutans, CH43, and genetic analysis of mutacin I biosynthesis genes. Appl Environ Microbiol 66(8):3221–3229
Qiao M, Saris PE (1996) Evidence for a role of NisT in transport of the lantibiotic nisin produced by Lactococcus lactis N8. FEMS Microbiol Lett 144(1):89–93
Rea MC, Sit CS, Clayton E, O'Connor PM, Whittal RM, Zheng J, Vederas JC, Ross RP, Hill C (2010) Thuricin CD, a posttranslationally modified bacteriocin with a narrow spectrum of activity against Clostridium difficile. Proc Natl Acad Sci U S A 107(20):9352–9357. https://doi.org/10.1073/pnas.0913554107
Rea MC, Ross RP, Cotter PD, Hill C (2011) Classification of bacteriocins from gram-positive bacteria. Springer, New York
Repka LM, Chekan JR, Nair SK, van der Donk WA (2017) Mechanistic understanding of lanthipeptide biosynthetic enzymes. Chem Rev 117(8):5457–5520. https://doi.org/10.1021/acs.chemrev.6b00591
Rodriguez JM, Martinez MI, Horn N, Dodd HM (2003) Heterologous production of bacteriocins by lactic acid bacteria. Int J Food Microbiol 80(2):101–116
Schneewind O, Missiakas D (2014) Sec-secretion and sortase-mediated anchoring of proteins in gram-positive bacteria. Biochim Biophys Acta 1843(8):1687–1697. https://doi.org/10.1016/j.bbamcr.2013.11.009
Schnell N, Engelke G, Augustin J, Rosenstein R, Ungermann V, Gotz F, Entian KD (1992) Analysis of genes involved in the biosynthesis of lantibiotic epidermin. Eur J Biochem 204(1):57–68
Shimamura H, Gouda H, Nagai K, Hirose T, Ichioka M, Furuya Y, Kobayashi Y, Hirono S, Sunazuka T, Omura S (2009) Structure determination and total synthesis of bottromycin A2: a potent antibiotic against MRSA and VRE. Angew Chem Int Ed Engl 48(5):914–917. https://doi.org/10.1002/anie.200804138
Skaugen M, Abildgaard CI, Nes IF (1997) Organization and expression of a gene cluster involved in the biosynthesis of the lantibiotic lactocin S. Mol Gen Genet 253(6):674–686
Sprules T, Kawulka KE, Gibbs AC, Wishart DS, Vederas JC (2004) NMR solution structure of the precursor for carnobacteriocin B2, an antimicrobial peptide from Carnobacterium piscicola. Eur J Biochem 271(9):1748–1756. https://doi.org/10.1111/j.1432-1033.2004.04085.x
Stoddard GW, Petzel JP, van Belkum MJ, Kok J, McKay LL (1992) Molecular analyses of the lactococcin A gene cluster from Lactococcus lactis subsp. lactis biovar diacetylactis WM4. Appl Environ Microbiol 58(6):1952–1961
Su TL (1948) Micrococcin, an antibacterial substance formed by a strain of Micrococcus. Br J Exp Pathol 29(5):473–481
Tomita H, Fujimoto S, Tanimoto K, Ike Y (1996) Cloning and genetic organization of the bacteriocin 31 determinant encoded on the Enterococcus faecalis pheromone-responsive conjugative plasmid pYI17. J Bacteriol 178(12):3585–3593
Uguen M, Uguen P (2002) The LcnC homologue cannot replace LctT in lacticin 481 export. FEMS Microbiol Lett 208(1):99–103
van Belkum MJ, Worobo RW, Stiles ME (1997) Double-glycine-type leader peptides direct secretion of bacteriocins by ABC transporters: colicin V secretion in Lactococcus lactis. Mol Microbiol 23(6):1293–1301
van Belkum MJ, Martin-Visscher LA, Vederas JC (2010) Cloning and characterization of the gene cluster involved in the production of the circular bacteriocin carnocyclin a. Probiotics Antimicrob Proteins 2(4):218–225. https://doi.org/10.1007/s12602-010-9056-1
van de Kamp M, van den Hooven HW, Konings RN, Bierbaum G, Sahl HG, Kuipers OP, Siezen RJ, de Vos WM, Hilbers CW, van de Ven FJ (1995) Elucidation of the primary structure of the lantibiotic epilancin K7 from Staphylococcus epidermidis K7. Cloning and characterisation of the epilancin-K7-encoding gene and NMR analysis of mature epilancin K7. Eur J Biochem 230(2):587–600
van den Berg van Saparoea HB, Bakkes PJ, Moll GN, Driessen AJ (2008) Distinct contributions of the nisin biosynthesis enzymes NisB and NisC and transporter NisT to prenisin production by Lactococcus lactis. Appl Environ Microbiol 74(17):5541–5548. https://doi.org/10.1128/AEM.00342-08
van der Meer JR, Polman J, Beerthuyzen MM, Siezen RJ, Kuipers OP, De Vos WM (1993) Characterization of the Lactococcus lactis nisin A operon genes nisP, encoding a subtilisin-like serine protease involved in precursor processing, and nisR, encoding a regulatory protein involved in nisin biosynthesis. J Bacteriol 175(9):2578–2588
van der Meer JR, Rollema HS, Siezen RJ, Beerthuyzen MM, Kuipers OP, de Vos WM (1994) Influence of amino acid substitutions in the nisin leader peptide on biosynthesis and secretion of nisin by Lactococcus lactis. J Biol Chem 269(5):3555–3562
van Roosmalen ML, Geukens N, Jongbloed JD, Tjalsma H, Dubois JY, Bron S, van Dijl JM, Anne J (2004) Type I signal peptidases of gram-positive bacteria. Biochim Biophys Acta 1694(1–3):279–297. https://doi.org/10.1016/j.bbamcr.2004.05.006
Vaughan A, Eijsink VG, van Sinderen D (2003) Functional characterization of a composite bacteriocin locus from malt isolate Lactobacillus sakei 5. Appl Environ Microbiol 69(12):7194–7203
Velasquez JE, van der Donk WA (2011) Genome mining for ribosomally synthesized natural products. Curr Opin Chem Biol 15(1):11–21. https://doi.org/10.1016/j.cbpa.2010.10.027
Velasquez JE, Zhang X, van der Donk WA (2011) Biosynthesis of the antimicrobial peptide epilancin 15X and its N-terminal lactate. Chem Biol 18(7):857–867. https://doi.org/10.1016/j.chembiol.2011.05.007
Waisvisz JM, Vanderhoeven MG, Vanpeppen J, Zwennis WCM (1957) Bottromycin. I. A new sulfur-containing antibiotic. J Am Chem Soc 79(16):4520–4521. https://doi.org/10.1021/Ja01573a072
Widdick DA, Dodd HM, Barraille P, White J, Stein TH, Chater KF, Gasson MJ, Bibb MJ (2003) Cloning and engineering of the cinnamycin biosynthetic gene cluster from Streptomyces cinnamoneus cinnamoneus DSM 40005. Proc Natl Acad Sci U S A 100(7):4316–4321. https://doi.org/10.1073/pnas.0230516100
Wieland Brown LC, Acker MG, Clardy J, Walsh CT, Fischbach MA (2009) Thirteen posttranslational modifications convert a 14-residue peptide into the antibiotic thiocillin. Proc Natl Acad Sci U S A 106(8):2549–2553. https://doi.org/10.1073/pnas.0900008106
Worobo RW, van Belkum MJ, Sailer M, Roy KL, Vederas JC, Stiles ME (1995) A signal peptide secretion-dependent bacteriocin from Carnobacterium divergens. J Bacteriol 177(11):3143–3149
Wu KH, Tai PC (2004) Cys32 and His105 are the critical residues for the calcium-dependent cysteine proteolytic activity of CvaB, an ATP-binding cassette transporter. J Biol Chem 279(2):901–909. https://doi.org/10.1074/jbc.M308296200
Xie L, Miller LM, Chatterjee C, Averin O, Kelleher NL, van der Donk WA (2004) Lacticin 481: In vitro reconstitution of lantibiotic synthetase activity. Science 303(5658):679–681. https://doi.org/10.1126/science.1092600
Xu Y, Li X, Li R, Li S, Ni H, Wang H, Xu H, Zhou W, Saris PE, Yang W, Qiao M, Rao Z (2014) Structure of the nisin leader peptidase NisP revealing a C-terminal autocleavage activity. Acta Crystallogr D Biol Crystallogr 70(Pt 6):1499–1505. https://doi.org/10.1107/S1399004714004234
Young TS, Walsh CT (2011) Identification of the thiazolyl peptide GE37468 gene cluster from Streptomyces ATCC 55365 and heterologous expression in Streptomyces lividans. Proc Natl Acad Sci U S A 108(32):13053–13058. https://doi.org/10.1073/pnas.1110435108
Zhang H, Zhou W, Zhuang Y, Liang X, Liu T (2013) Draft genome sequence of Streptomyces bottropensis ATCC 25435, a bottromycin-producing actinomycete. Genome Announc 1(2):e0001913. https://doi.org/10.1128/genomeA.00019-13
Zhang Q, Doroghazi JR, Zhao X, Walker MC, van der Donk WA (2015) Expanded natural product diversity revealed by analysis of lanthipeptide-like gene clusters in actinobacteria. Appl Environ Microbiol 81(13):4339–4350. https://doi.org/10.1128/AEM.00635-15
Zheng G, Hehn R, Zuber P (2000) Mutational analysis of the sbo-alb locus of Bacillus subtilis: identification of genes required for subtilosin production and immunity. J Bacteriol 182(11):3266–3273
Zheng Q, Wang S, Duan P, Liao R, Chen D, Liu W (2016) An α/β-hydrolase fold protein in the biosynthesis of thiostrepton exhibits a dual activity for endopeptidyl hydrolysis and epoxide ring opening/macrocyclization. Proc Natl Acad Sci U S A 113(50):14318–14323. https://doi.org/10.1073/pnas.1612607113
Zheng S, Nagao J, Nishie M, Zendo T, Sonomoto K (2018) ATPase activity regulation by leader peptide processing of ABC transporter maturation and secretion protein, NukT, for lantibiotic nukacin ISK-1. Appl Microbiol Biotechnol 102(2):763–772. https://doi.org/10.1007/s00253-017-8645-2
Acknowledgements
Sen Zheng acknowledges the Ajinomoto Scholarship Foundation, Japan fellowship, for the financial aid through this study.
Funding
This work was partially supported by Grants-in-Aid for Scientific Research (KAKENHI) from the Japan Society for the Promotion of Science (JSPS), grant number JP26292040.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 308 kb)
Rights and permissions
About this article
Cite this article
Zheng, S., Sonomoto, K. Diversified transporters and pathways for bacteriocin secretion in gram-positive bacteria. Appl Microbiol Biotechnol 102, 4243–4253 (2018). https://doi.org/10.1007/s00253-018-8917-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-8917-5