Abstract
Arbuscular mycorrhizal fungi (AMF) are the predominant type of mycorrhizal fungi in roots and rhizosphere soil of grass species worldwide. Grasslands are currently experiencing increasing grazing pressure, but it is not yet clear how grazing intensity and host plant grazing preference by large herbivores interact with soil- and root-associated AMF communities. Here, we tested whether the diversity and community composition of AMF in the roots and rhizosphere soil of two dominant perennial grasses, grazed differently by livestock, change in response to grazing intensity. We conducted a study in a long-term field experiment in which seven levels of field-manipulated grazing intensities were maintained for 13 years in a typical steppe grassland in northern China. We extracted DNA from the roots and rhizosphere soil of two dominant grasses, Leymus chinense (Trin.) Tzvel. and Stipa grandis P. Smirn, with contrasting grazing preference by sheep. AMF DNA from root and soil samples was then subjected to molecular analysis. Our results showed that AMF α-diversity (richness) at the virtual taxa (VT) level varied as a function of grazing intensity. Different VT showed completely different responses along the gradient, one increasing, one decreasing, and others showing no response. Glomeraceae was the most abundant AMF family along the grazing gradient, which fits well with the theory of disturbance tolerance of this group. In addition, sheep-grazing preference for host plants did not explain much of the variation in AMF α-diversity. However, the two grass species exhibited different AMF community composition in their roots and rhizosphere soils. Roots exhibited a lower α-diversity and higher β-diversity within the AMF community than soils. Overall, our results suggest that long-term grazing intensity might have changed the abundance of functionally diverse AMF taxa in favor of those with disturbance-tolerant traits. We suggest our results would be useful in informing the choice of mycorrhizal fungi indicator variables when assessing the impacts of grassland management choices on grassland ecosystem functioning.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Arbuscular mycorrhizal fungi (AMF) are a key part of the soil-root biota, playing a crucial role in maintaining grassland productivity and stability [5, 37]. AMF are the predominant type of mycorrhizal fungi in grasslands and form unique communities in the roots and rhizosphere soil of grass species [17, 51]. Grasslands are currently experiencing increasing grazing pressure, but it is not yet clear whether and how grazing intensity by large herbivores drives AMF diversity and community composition. The extent of the grazing impact on AMF function and community structure is primarily determined by intensity of grazing [6, 21, 58] as it has disparate effects on above- and below-ground biodiversity and productivity [57]. However, most studies compare the effects of grazing on AMF communities in grazed and un-grazed plots [24, 38, 52], with very few investigating impacts along a gradient of grazing pressure such as that ranging from light grazing pressure through to overgrazing [6, 20, 36]. This is a major shortcoming as livestock grazing of grasslands is a major agriculture practice worldwide [14, 39]. Besides, identification of AMF communities has mostly been confined to spore isolation approaches, thus potentially losing much of the relative diversity of the active component of the mycorrhizal communities that could be better captured through the use of novel molecular techniques [17, 29].
The effects of grazing on the AMF community can be highly species-specific. Grazing impacts will depend on the host plant identity due to their different palatability/digestibility traits that influence selection by grazing livestock [23, 51]. Although AMF are considered non-specific symbiotic partners at species level [47], non-random patterns in host-fungal interaction suggests some degree of host specificity or preference which is regulated by the functional characteristics of both partners [16, 46, 51]. For example, the use of pyrosequencing analysis of AMF communities colonizing the roots of three common grass species (Poaceae) with different sensitivities to trampling and grazing revealed that host plant identity is critical in shaping the AMF community structure and composition [51]. It is also known that AMF root colonization differs among certain grasses that are selected differentially by grazing livestock [11]. Cavagnaro et al. [11] reported a highly significant decrease in AMF colonization as a result of increased grazing intensity for species preferred by the grazers and less pronounced effects for the less-preferred species. This evidence, therefore, suggests that the effect of grazing on AMF community is also dependent on host plant identity. Given that herbivory-caused defoliation limits below-ground carbon allocation from the plant to their associated fungal partners [58], preferred or not preferred plant species by grazers within a community could be expected to modify the rhizosphere differentially, thus creating distinct associated AMF communities and mycorrhizal functioning.
There is also evidence that the AMF communities differ between the host roots and the rhizosphere soil [26, 32, 46]. It has been suggested that the AMF community structure in these two environments is driven by a number of different factors, with the AMF community composition colonizing the roots mainly determined by the host plant, while the AMF assemblage of the rhizosphere soil is more related to environmental conditions including soil properties [32, 49]. However, most studies have investigated AMF assemblages in either root [51] or soil [52], and few have assessed both simultaneously. Studies based on the identification of root-colonizing and soil-borne AMF assemblages between plants with different grazing preferences by livestock could provide further insights into the impact of grazing on AMF communities and the underlying mechanisms. Changes in AMF community structure could be used as an indicator of plant-soil system health.
The objective of this study, therefore, is to find out whether AMF communities in the roots and rhizosphere soil of two dominant perennial grasses (Leymus chinense (Trin.) Tzvel. and Stipa grandis P. Smirn) change as a function of grazing intensity. L. chinense is a highly mycorrhizal and palatable bunchgrass, while S. grandis has lower mycorrhizal colonization and low palatability [56]. We conducted a study in a long-term experimental site with a gradient of seven grazing intensities maintained for 13 years at a typical steppe grassland in northern China. In our study, grazing intensity is represented in a gradient of seven levels, which may be better than traditional replicated designs that consider only extreme environmental conditions [15, 28], where the influence of intermediate levels of grazing intensity may not be detected.
We extracted and sequenced AMF DNA from the root and rhizosphere soil samples of each species and evaluated (1) how AMF communities associated with both root and rhizosphere soil varied between L. chinense and S. grandis along the grazing gradient and (2) how AMF α-diversity and β-diversity were affected by grazing intensity and host plant grazing preferences by sheep. We hypothesized that the sheep-preferred L. chinense would have a different composition of AMF taxa in its root and rhizosphere soil than the less-preferred S. grandis. In addition, AMF taxa respond differently to grazing intensity because of their different traits. To our knowledge, this study is the first to thoroughly characterize the AMF community using molecular techniques in two common grass species with different palatability and mycorrhizal dependence along a grazing gradient in a typical temperate steppe. Such information will contribute to a deeper understanding of changes within the AMF community caused by livestock grazing and disturbance, provide opportunities for developing AMF ecological indicators, and improve restoration strategies for reestablishment of native vegetation in temperate grasslands.
Materials and Methods
Study Site
This study was set up in a steppe grassland in a semi-arid zone with continental climate located at the Sino-German Inner Mongolia Grassland Ecosystem Research Station (IMGERS) in the Xilin River Basin (116° 42′ E; 43° 38′ N), Inner Mongolia, China. We conducted our experiment in seven plots, each with different levels of grazing intensities (GI); each plot contained a flat area of 2 ha and was subjected to one level of grazing intensity, from 0 to 9 ewes ha−1 with interval increases of 1.5 ewe ha−1 giving a range of grazing intensities: 0 (no grazing), 1.5 (very light), 3 (light), 4.5 (light-moderate), 6 (moderate), 7.5 (heavy), and 9 (overgrazing) (Appendix A-1, Fig S1). The grassland was grazed by young female sheep (ewes) ca. 35 kg live weight. The ewes were put in plots for 90 days throughout the growing season (June–September) every year. The different grazing intensity treatments have been run continuously for 13 years before our study in 2019; hence, the impact of the differing grazing intensities will have stabilized [33, 43]. A detailed description of vegetation cover, climate, and soil characteristics of the experimental site can be found in the supplementary information (Appendix A-1).
Root and Soil Sampling
Root and rhizosphere soil sampling of two dominant grass species, S. grandis and L. chinense, was conducted in the middle of the growing season in July 2019. Three parallel transects, 50 m apart, were set up in each experimental plot (100 m × 200 m). Transect lines were 150 m long, and nine sampling points were placed at regular distance along each transect. Along each transect, nine S. grandis and nine L. chinense were collected with their roots and adhering soil from 0 to 20 cm of depth with a garden trowel. Twenty-seven samples of root and adhering soil from the same plant species were taken in each plot. Collected samples on each transect were then mixed to give a composite soil and plant samples of the same species. A schematic representation of the experimental design can be found in Appendix A-2, Fig S2.
Root samples were rinsed with tap water, and non-lignified, light-fine secondary roots were hand-picked from the main root and cut into 1 cm pieces. 10 g fresh weight of roots was sub-sampled and stored at −80 °C for DNA extraction. Rhizosphere soil samples, i.e., soil tightly adhering to the roots, were sampled, sieved (2-mm sieve), and frozen at −80 °C for DNA extraction and sequencing.
Molecular Analysis
DNA Extraction and PCR
DNA extraction and PCR were conducted by Guangdong Magigene Biotechnology Co. Ltd. (Guangzhou, China). Total genomic DNA was extracted from 84 samples (7 plots × 3 transects × 2 plant species × 2 environments (root vs. soil)), consisting of 42 rhizosphere soil samples and 42 root samples, using a DNA extraction kit (MinkaGene Bacterial DNA Kit). Six of 42 rhizosphere soil samples did not provide usable data and were removed from subsequent analysis (Appendix A-3, Table S1). DNA amplification was then performed using AMF-specific primers by nested polymerase chain reaction (nested PCR). Nested PCR was carried out using BioRad S1000 (Bio-Rad Laboratory, CA) and two sets of primers: AML1 (5′-ATCAACTTTCGATGGTAGGATAGA-3′)/AML2 (5′-GAACCCAAACACTTTGGTTTCC-3′) [30] in first PCR and primers AMV4.5NF (5′-AAGCTCGTAGTTGAATTTCG-3′) and AMDGR (5′-CCCAACTATCCCTATTAATCAT-3′) [45] in the second PCR.
Premix Taq™ DNA polymerase (Takara Bio, USA) was used to provide nearly error-free amplification and high amplicon yield. PCR reactions, containing 25 μl 2 × Premix Taq™ DNA polymerase, 1 μl each primer (10 mM), and 3 μl DNA (20 ng/μl) template in a volume of 50 μl, were amplified. For the nested PCRs, the DNA template was the first PCR product diluted 1:10 in ultrapure H2O. The thermocycler settings were as follows: 94°C for 3 min then 35 cycles at 94°C for 45 s, 51°C for 40 s, 72°C for 60 s, and a final elongation step by 72°C for 10 min for the first PCR followed by 94°C for 3 min, then 35 cycles at 94°C for 40 s, 58°C for 60 s, and 72°C for 60 s and a final elongation step of 72°C for 10 min for the nested PCR.
The DNA fragments resulting from PCR were separated by size via agarose gel electrophoresis within a 1.5% agarose gel in 0.5 × TAE (40 mM Tris-acetate, 1 mM EDTA) as running buffer. After separation, the resulting DNA fragments were visible as clearly defined bands. The AxyPrepDNA gel extraction kit was then used for the rapid purification of DNA fragments from TAE agarose gels (AXYGEN, California, USA). The DNA concentrations of the purified PCR products were quantified on the NanoDrop 1000TM Spectrophotometer (Thermo Fisher Scientific, MA, USA) according to the manufacturer’s instructions. PCR products were mixed in equimolar ratios according to the GeneTools Analysis Software (Version4.03.05.0, SynGene). Then, mixture of PCR products was purified with EZNA Gel Extraction Kit (Omega, USA). Sequencing libraries were generated using NEBNext® Ultra™ DNA Library Prep Kit for Illumina® (New England Biolabs, USA) following manufacturer’s recommendations, and index codes were added. The library quality was assessed on the Qubit@ 2.0 Fluorometer (Thermo Scientific) and Agilent Bioanalyzer 2100 system.
Next-Generation Sequencing and Species Annotation
Illumina next-generation sequencing (NGS) technology was used for paired-end sequencing (2 × 250 bp) on an Illumina NovaSeqTM6000 sequencing platform. To check the quality of raw sequence data and calculation of quality values, FastQC tool (Version 0.11.9) [4] was used. In addition, we used MultiQC [18] to aggregate the reports from FastQC into a single report with interactive plots for multiple bioinformatics analyses (Appendix B).
Following the quality check, CutAdapt (V 3.2) [34] was applied to cut adapters. Forward and reverse reads were then processed using the SEED 2.1.1 pipeline [55],paired-end reads were merged, and reads with an average quality below 30 and a length of less than 200 bp, as well as sequences with ambiguous bases, were discarded. High-quality sequences were clustered into operational taxonomic units (OTUs) with 97% sequence identity using the VSEARCH clustering program [44]. Representative sequences for each OTU was screened for further annotation. We excluded singletons and doubletons as well as OTUs that represented 0.002% of the total number of sequences to reduce the number of spurious OTUs due to sequencing errors, following the recommendation of Bokulich et al. [9]. Representative sequences of the OTUs were BLASTed [1] against the maarjAM database (Opik et al., 2010) to verify the Glomeromycotan origin of the sequences and to categorize the reads to virtual taxa (VT). Raw sequencing data were deposited on the Sequence Read Archive under accession number PRJNA758203.
Statistical Analyses
We used three-way nested data to examine differences in AMF diversity and community composition of the roots and rhizosphere soil from two different plant hosts. We also tested how AMF α- and β-diversity at the virtual taxa (VT) level are affected by grazing intensity, host plant identity, and mycorrhizal environment as well as their interactions. The data are nested in the sense that samples were taken from seven sites (hereafter referred to as plots) with seven rates of grazing intensity. Although replication is a key principle for dealing with random or spatially structured heterogeneity between individuals or experimental units, adding experimental units along the gradient instead of increasing the number of replicates can improve the success of overall prediction in ecological studies [28]. In each plot, composite samples were collected from two host plants (L. chinense and S. grandis) along three transects, and from each host, samples included both roots and rhizosphere soil, where mycorrhizal community composition was measured (see Appendix A-2 (Fig S2) for a schematic illustration of the experimental design).
AMF α-Diversity
Rarefaction curves were plotted to test how well a sample reflects the true diversity of root- and soil-associated AMF and to compare observed OTUs richness among AMF communities in the roots and rhizosphere soil (Fig. 1).
AMF α-diversity was calculated using the richness and Shannon–Wiener diversity index based on virtual taxa (VT). Linear mixed effects models were applied to test the impact of grazing intensity on AMF Shannon diversity in the roots and rhizosphere soil of the two dominant plant species. The response variable was the AMF diversity. Fixed explanatory variables were grazing intensity, plant species, and mycorrhizal environment (root and rhizosphere soil), and the random variable was transect ID (nested by grazing intensity). We first fitted a model with all terms as well as all their interactions. Then, automated model selections using Akaike’s Information Criterion (AIC) [10] were conducted to select the best fitting model [60]. Due to the design of this large-scale, long-term field experiment, grazing intensity was treated as a continuous variable.
AMF β-Diversity
To visualize the differences between AMF community composition in the root and rhizosphere soil of the dominant plant species, non-metric multidimensional scaling (NMDS) was applied based on the Bray-Curtis distance [3].
To test how much of the variation in AMF community composition at VT level (β-diversity) was explained by grazing intensity, mycorrhizal environment, and host plant in the AMF community data of all 78 samples (6 missing samples) at VT level, we performed redundancy analysis (RDA). RDA was conducted on VT with relative abundance of >5% in soil and root. RDA was computed based on Hellinger pre-transformed AMF composition data with three measured variables as explanatory factors [31]. Monte Carlo permutation test with 999 permutations was applied to test the significance level for the variation explained by explanatory variables [59]. We also calculated the significance of each constrained axis independently. RDA ordinations were plotted as a triplot and type II scaling which was considered correlative relationships between variables.
All statistical analyses were carried out using R, version 3.6.3 [42]. Generalized linear mixed effect models were applied using lme function from “nlme” package [41]. Automated model selection was carried out using dredge() function [8] from “MuMIn” package. All models were validated by checking the distribution of residuals and standard model validation graphs to verify homogeneity and normality and to identify influential observations according to Zuur et al. [60]. Visual inspection of the residual plots did not show any obvious deviations from homoscedasticity or normality. Species accumulation curves, NMDS, RDA, and the significance for constrained ordination test (Monte Carlo permutation test) were conducted using functions available in “vegan” package (Oksanen, 2013).
Results
Characterization of AMF Community
A total of 1,515,589 AMF sequences were obtained from 36 soil (6 samples failed to produce useable data) and 42 root samples. 705376 and 810213 sequences appeared in soil and root samples, respectively. In addition, 727530 and 788059 sequences appeared in S. grandis and L. chinense, respectively. AMF sequences were then clustered into 1340 OTUs according to the ≥ 97% similarity threshold. 1340 OTUs belong to 71 virtual taxa annotated in MaarjAM Glomeromycota database. These VT belong to seven families (Fig. 1): 61 Glomeraceae, 4 Claroideoglomeraceae, 2 Archaeosporaceae, 1 Ambisporaceae, 1 Paraglomeraceae, 1 Diversisporaceae, and 1 Gigasporaceae.
The most abundant VT in soil were Glomus_VTX00063 (12.11%), Glomus_VTX00222 (10.47%), Glomus_VTX00167 (9.93%), Glomus_VTX00329 (9.90%), Glomus_VTX00156 (7.33%), Glomus_VTX00387 (5.50%), and Glomus_VTX00304 (4.97%), respectively. The most abundant VTs in roots were Glomus_VTX00387 (25.87%), Glomus_VTX00390 (11.76%), Glomus_VTX00156 (11.27%), Glomus_VTX00167 (9.69%), Glomus_VTX00304 (6.77%), Glomus_VTX00166 (5.83%), and Glomus_VTX00386 (5.43%), respectively (Fig. 1).
AMF α-Diversity
Rarefaction analysis showed that the number of samples was sufficient to identify the major AMF in the root and soil environment (Fig. 2). The rarefaction curves of the AMF communities also showed different OTU richness in the soil and root AMF communities; the AMF community had greater species richness in the soil compared to the roots (Fig. 2). The linear mixed effects models revealed the significant effect of mycorrhizal environment on VT richness and VT diversity in the best fitted models (Table 1). The significant effect of grazing intensity, however, was only observed on VT richness. In addition, VT richness significantly impacted by host plant species, with L. chinense harboring more VT than S. grandis (Appendix A-7, Fig S6).
Species accumulation curves of estimated OTU richness for both root and rhizosphere soil-associated AMF communities collected along a grazing gradient in a steppe grassland in Inner Mongolia. Solid lines and light-colored region refer to the average estimated richness and standard deviation, respectively.
AMF β-Diversity
The NMDS plot of the similarity between AMF communities in the roots and rhizosphere soil of the two dominant plant species showed a clear distinction between the mycorrhizal environment, soil or root (shown with different colors), but the differences in grazing intensity (Appendix A-6, Fig S5) and mycorrhizal communities between the two grass species were more subtle (visualized as different shapes) as two groups of samples are mixed in the plot (Fig. 3) (Appendix A-5, Fig S4).
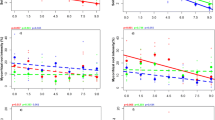
RDA analysis computed the proportion of variance in AMF composition at the VT level explained by grazing intensity, mycorrhizal environment, and host plant. The analysis yielded three canonical axes (RDA1 to RDA3) and three additional unconstrained axes for the residuals. The three explanatory variables together explained 42% of the variance (P=0.001, adjusted R2 = 0.394). The first (RDA1), second (RDA2), and third constrained axis (RDA3) explained 36.87%, 4.3%, and 0.59% of the variance, respectively. Calculating the significance of each constrained axis using Monte Carlo permutation tests showed that the first (P=0.001) and second RDA (P=0.001) axes were significant, but the third RDA was not (P=0.591). Furthermore, the RDA plot clearly visualized the grazing intensity impact on the different AMF VT; e.g., the relative abundance of Glomus_VTX00390 was positively correlated with grazing intensity, and the relative abundance of Glomus_VTX00329 was negatively correlated with grazing intensity (Fig. 4).
RDA triplot of 78 Illumina-sequenced root and soil samples collected along a grazing gradient in a steppe grassland in Inner Mongolia. Objects are ordinated as points, while both response and explanatory variables are plotted as vectors. Explanatory environmental variables including grazing intensity, AMF environment, and host identity are indicated by line arrows in green. The most abundant AMF VT (>5%) in root and soil environments are shown by brown arrows.
Discussion
We determined AMF diversity and community composition in the roots and rhizosphere soil of two dominant grasses with contrasting palatability/digestibility traits that influence different sheep-grazing preferences within a 13-year grazing experiment with seven levels of grazing intensity on a steppe grassland in China. Overall, we found long-term grazing significantly affected AMF α-diversity (VT richness), but Shannon diversity at VT was not significantly affected by grazing intensity. The β-diversity (the relative abundance of AMF taxa) differed as a function of grazing intensity. Grazing preference did not determine the composition of AMF taxa in two perennial grass species. However, mycorrhizal environment, i.e., where sampling occurred, played a determining role in shaping the AMF community composition.
While we acknowledge that true replications of grazing intensity on each plot would have improved the statistical rigor of the work and allowed for the detection of differences among grazing treatments, conducting replications would not have been feasible in a large-scale study of this type, where plots must be relatively large (in this case, 2 ha per plot). In particular, a replicated design would not exploit response patterns along the grazing gradient of interest in our study [28].
AMF α-Diversity and Grazing Intensity
AMF α-diversity (Shannon diversity at the VT level) was not significantly affected by grazing intensity in our study (Table 1), confirming Ambrosino et al. [2], who found no effect of grazing-induced defoliation on diversity of AMF spores in Argentine pastures, and van der Heyde et al. [52], who reported no effects of grazing on AMF communities detected in soil in Canadian grasslands. However, we found a significant effect of grazing intensity on AMF richness (Table 1). There have been conflicting results from studies reporting either a positive or negative response of AMF richness and diversity to grazing intensity [6, 7]. These conflicting results may be caused by multiple approaches used to quantify AMF communities, ranging from spore isolation to molecular techniques [54]. Furthermore, Kusakabe et al. [29] found different responses of AMF richness and diversity to grazing intensity at three different grazing sites in Mongolian grasslands. They found a negative correlation between grazing intensity and AMF diversity in one site and attributed this to decrease in shoot biomass of mycorrhizal plant species and increase in non- or weakly mycorrhizal plants [29]. In contrast, a positive correlation with grazing intensity in another site was associated with higher AMF host biomass at the grazed sites [29]. Overall, the grazing effects on AMF species diversity and underlying mechanisms are still controversial as they depend on grazing-induced changes in mycorrhizal environment which varies along different ecosystems [22].
AMF β-Diversity and Grazing Intensity
It has been argued that traits of AMF communities vary in response to environmental disturbances such as grazing, particularly at the species level [19]. Our results showed various responses among AMF virtual taxa to grazing intensity. We found significantly positive responses of some taxa such as Glomus_VTX00329 and significantly negative responses of some other species such as Glomus_VTX00390 to grazing intensity. Some taxa such as Glomus_VTX00304 fungi showed no remarkable variation in response to grazing intensity. Differential responses among virtual AMF taxa to grazing-caused disturbance can be partly explained by differences in their life-history traits such as growth rate, sporulation, and resource conservation strategies [17, 53]. These traits most likely generate a wide range of mycorrhizal functional groups, from grazing tolerant to non-tolerant.
We found that the most abundant AMF VT in the roots and soil environments belonged to the Glomeraceae, and most of them were positively correlated with grazing intensity (Fig. 4). The Glomeraceae has often been described as the most disturbance-tolerant AMF family [12, 50, 53]. Glomeraceae is dominant in a broad range of ecosystems worldwide, including agricultural systems, owning to their tolerance to environmental disturbances achieved by its ruderal life-history strategy through high carbon use efficiency, fast producing of large spores, and good adaptation to various host plants [2, 12, 49]. There is some evidence that AMF species within the Glomeraceae tolerate consistent low levels of carbon supply from the host plants as a result of herbivory and above-ground tissue loss,hence, they are tolerant to grazing [17, 50]. In addition, the high rate of sporulation [40] and hyphal turnover [48], as well as reproduction from both hyphal fragments and spores probably, makes AMF species of Glomeraceae resistant to hyphal disruption and mycelial loss caused by grazing. Likewise, Glomus species were detected in greater abundance in a long-term, overgrazed steppe compared with naturally restored and non-grazed sites in typical steppes of Inner Mongolia [56]. These findings suggest that functional life-history strategies in the Glomeraceae may have enhanced the capability of the most species in this family to withstand grazing disturbance.
Taken together, our results suggest that long-term grazing intensity may have caused the dominance of grazing-adapted mycorrhizal fungi in our experimental site. Therefore, grazing intensity will have favored members of the AMF community with grazing-tolerant traits.
AMF and Host Plant Grazing Preference
Host plant identity and traits have been identified as an important driver of AMF community structure in some cases [35, 51]. In an effort to investigate the effects of land-use intensity on the AMF community of three common grass species with contrasting mowing, trampling, and grazing tolerance in pastures with either mown or grazed, or both management, Vályi et al. [51] found that host plant-specific traits explained a large proportion of variation in the AMF community in response to the type and degree of disturbance at land-use sites. However, contrary to our prediction, we found no meaningful differences in AMF diversity and community composition between the plant species we studied (L. chinense and S. grandis) with contrasting grazing preferences by sheep. This can be explained by the fact that the two grass species are grazing tolerant and remained the dominant plant species in all grazing treatments despite subtle changes in their relative abundance [33]. Similar to our findings, AMF spore abundance and diversity were not significantly affected by grazing in three native perennial grass species with different livestock grazing preferences in Argentine rangelands [2]. Given that the grasses we studied had different grazing preferences by sheep, the lack of effect of host plant on AMF community composition suggests that there is no strong preference among AMF communities or functional groups to colonize the root and rhizosphere soil of sheep-preferred and non-preferred grass species. In fact, the response of AMF to host plant grazing preferences by sheep appears to be similar within AMF communities. Therefore, the effects of grazing on AMF community composition were independent of the grazing preference of the dominant native grasses in our study site.
AMF Community Structure in Soil and Root
Consistent with previous findings, we found distinct AMF community structures in both the root and rhizosphere soil, regardless of host plant identity [13, 26, 32, 49]. Not surprisingly, we detected lower α-diversity within the AMF community in root compared to soil, suggesting that the AMF root community is less diverse in terms of the number of different OTUs. The disparities between root and soil media are to be expected, as these two mycorrhizal accessible environments represent different parts of the AMF communities in intra-radical and extra-radical structures. Lower α-diversity in roots suggests that some AMF species were dormant and inactive in the root system at the time of sampling [26, 51]. This finding might also reflect the different AMF carbon allocation to internal and external structures. Indeed, AMF invest either in long-lived internal structures or in high-cost short-lived external ones depending on resource accessibility and environmental condition [27]. The internal structures in roots, such as internal hyphae, arbuscules, and vesicules, are those which are involved in transferring nutrients to the plant, whereas external hyphal structure are involved in foraging soil and nutrient acquisition [47]. Given that AMF cannot invest considerably in both internal and external components simultaneously [25], some level of differential allocation to structures in soil and root under grazing stress is highly likely. Furthermore, a greater homogeneity in root samples in terms of AMF community composition can be interpreted as an evidence that AMF species in roots are not random selections of AMF species from the soil. Therefore, AMF taxa may have evolved with properties to colonize different mycorrhizal plants based on their life-history traits and symbiont selection by their plant partners.
In contrast to α-diversity, we found higher β-diversity in the root samples compared to those of soil. Although AMF community composition within the two environments did not vary at family level, the relative abundance of AMF VT in the rhizosphere soils and roots was different (Fig. 1). The relative abundance of some VT is greater in soil samples (e.g., Glomus_VTX00063 and Glomus_VTX00222), whereas the relative abundance of some others is particularly greater in root samples (e.g., Glomus_VTX00387, Glomus_VTX00390, and Glomus_VTX00156) (Fig. 1). There are also some common VT that occur in root and soil with the same proportion, e.g., Glomus_VTX00167 (9.69% in root 9.93% in soil). This finding is expected as AMF isolates at the species level differ considerably in the rate and extent of colonization in soil and root [25]. A greater proportion of some VT in root compared to soil, e.g., Glomus_VTX00387 (25.87% in root and 5.50% in soil) or Glomus_VTX00390 (11.76% in root and in 3.7% soil), indicates a strong plant dependency for these species and potentially a less mutualistic relationship.
In conclusion, by integrating the effects of grazing intensity, host plant, and assessment of mycorrhizal community composition in different components (root vs. soil), we observed various functional traits among AMF taxa at VT level that could explain some of the grazing responses. We observed a significant increase in the relative abundances of most of AMF taxa belonging to the Glomeraceae, such as Glomus_VTX00390. Some AMF taxa in contrast decreased with increasing grazing intensity, such as Glomus_VTX00390. The differences among AMF taxa in their response to grazing stress reflect their various resource requirements and abilities to acquire resources which affect their impacts on communities and ecosystems.
Grass species with contrasting grazing preferences by sheep did not differ in the composition of their associated AMF communities in our study, suggesting that AMF functional groups do not respond differently to host plant grazing preference. The mycorrhizal environment in which the AMF community was measured (soil or root), regardless of host plant identity, determined to a large degree the composition of AMF communities with lower Shannon diversity, higher β-diversity in the root compared to the soil. Overall, our findings suggest that a decade of grazing by large herbivores has resulted in functional changes in AMF communities in response to grazing. Gaining knowledge on functionally diverse traits of AMF taxa may improve our understanding of the role of AMF in rehabilitating and restoring degraded grasslands around the world. This would be beneficial to both the sustainable use of these grasslands for livestock production and other ecosystem services such as climate change mitigation.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Ambrosino ML, Cabello MN, Busso CA, Velázquez MS, Torres YA, Cardillo DS et al (2018) Communities of arbuscular mycorrhizal fungi associated with perennial grasses of different forage quality exposed to defoliation. J Arid Environ 154:61–69
Anderson MJ, Ellingsen KE, McArdle BH (2006) Multivariate dispersion as a measure of beta diversity. Ecol Lett 9:683–693
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. In: Babraham Bioinformatics. Babraham Institute, Cambridge. http://www.bioinformatics.babraham.ac.uk/projects/fastqc
Asmelash F, Bekele T, Birhane E (2016) The potential role of arbuscular mycorrhizal fungi in the restoration of degraded lands. Front Microbiol 7:1095
Ba L, Ning J, Wang D, Facelli E, Facelli JM, Yang Y, Zhang L (2012) The relationship between the diversity of arbuscular mycorrhizal fungi and grazing in a meadow steppe. Plant Soil 352:143–156
Bai G, Bao Y, Du G, Qi Y (2013) Arbuscular mycorrhizal fungi associated with vegetation and soil parameters under rest grazing management in a desert steppe ecosystem. Mycorrhiza 23:289–301
Barton K (2018) MuMIn: multi-model inference. R package version 1.40.4. https://CRAN.R-project.org/package=MuMIn
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon JI, Knight R et al (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10:57–59
Burnham KP, Anderson DR (2004) Multimodel inference: understanding AIC and BIC in model selection. Sociol Methods Res 33(2):261–304
Cavagnaro RA, Pero E, Dudinszky N, Golluscio RA, Grimoldi AA (2019) Under pressure from above: overgrazing decreases mycorrhizal colonization of both preferred and unpreferred grasses in the Patagonian steppe. Fungal Ecol 40:92–97
Chagnon P-L, Bradley RL, Maherali H, Klironomos JN (2013) A trait-based framework to understand life history of mycorrhizal fungi. Trends Plant Sci 18:484–491
Chen Y-L, Zhang X, Ye J-S, Han H-Y, Wan S-Q, Chen B-D (2014) Six-year fertilization modifies the biodiversity of arbuscular mycorrhizal fungi in a temperate steppe in Inner Mongolia. Soil Biol Biochem 69:371–381
Conant RT (2010) Challenges and Opportunities for Carbon Sequestration in Grassland Systems: A Technical Report on Grassland Management and Climate Change Mitigation’, Integrated Crop Management, 9 (Rome: FAO).
Cottingham KL, Lennon JT, Brown BL (2005) Knowing when to draw the line: designing more informative ecological experiments. Front Ecol Environ 3(3):145–152
Davison J, García de León D, Zobel M, Moora M, Bueno CG, Barceló M et al (2020) Plant functional groups associate with distinct arbuscular mycorrhizal fungal communities. New Phytol 226:1117–1128
Dudinszky N, Cabello MN, Grimoldi AA, Schalamuk S, Golluscio RA (2019) Role of grazing intensity on shaping arbuscular mycorrhizal fungi communities in patagonian semiarid steppes. Rangel Ecol Manage 72:692–699
Ewels P, Magnusson M, Lundin S, Käller M (2016) MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32:3047–3048
Faggioli VS, Cabello MN, Grilli G, Vasar M, Covacevich F, Öpik M, Environment, (2019) Root colonizing and soil borne communities of arbuscular mycorrhizal fungi differ among soybean fields with contrasting historical land use. J Agriculture, Ecosystems 269:174–182
Faghihinia M, Zou Y, Bai Y, Marrs R, Staddon PL (2020) Seasonal variation in the response of arbuscular mycorrhizal fungi to grazing intensity. Mycorrhiza 30:635–646
Faghihinia M, Zou Y, Chen Z, Bai Y, Li W, Marrs R, Staddon PL (2020) The response of grassland mycorrhizal fungal abundance to a range of long-term grazing intensities. Rhizosphere 13:100178
Faghihinia M, Zou Y, Chen Z, Bai Y, Li W, Marrs R, Staddon PL (2020) Environmental drivers of grazing effects on arbuscular mycorrhizal fungi in grasslands. Applied Soil Ecology 153:103591
González JB, Petipas RH, Franken O, Kiers ET, Veblen KE, Brody AK (2018) Herbivore removal reduces influence of arbuscular mycorrhizal fungi on plant growth and tolerance in an East African savanna. Oecologia 187:123–133
Guo Y, Du Q, Li G, Ni Y, Zhang Z, Ren W, Hou X (2016) Soil phosphorus fractions and arbuscular mycorrhizal fungi diversity following long-term grazing exclusion on semi-arid steppes in Inner Mongolia. Geoderma 269:79–90
Hart MM, Reader RJ (2002) Taxonomic basis for variation in the colonization strategy of arbuscular mycorrhizal fungi. New Phytol 153:335–344
Hempel S, Renker C, Buscot F (2007) Differences in the species composition of arbuscular mycorrhizal fungi in spore, root and soil communities in a grassland ecosystem. Environ Microbiol 9:1930–1938
Johnson NC, Wilson GWT, Bowker MA, Wilson JA, Miller RM (2010) Resource limitation is a driver of local adaptation in mycorrhizal symbioses. Proc Natl Acad Sci 107:2093–2098
Kreyling J, Schweiger AH, Bahn M, Ineson P, Migliavacca M, Morel‐Journel T, Christiansen JR, Schtickzelle N, Larsen KS (2018) To replicate, or not to replicate–that is the question: how to tackle nonlinear responses in ecological experiments. Ecol Lett 21(11):1629–1638
Kusakabe R, Taniguchi T, Goomaral A, Undarmaa J, Yamanaka N, Yamato M (2018) Arbuscular mycorrhizal fungal communities under gradients of grazing in Mongolian grasslands of different aridity. Mycorrhiza 28:621–634
Lee J, Lee S, Young JPW (2008) Improved PCR primers for the detection and identification of arbuscular mycorrhizal fungi. FEMS Microbiol Ecol 65:339–349
Legendre P, Gallagher ED (2001) Ecologically meaningful transformations for ordination of species data. Oecologia 129:271–280
Li S, Shakoor A, Wubet T, Zhang N, Liang Y, Ma K (2018) Fine-scale variations of fungal community in a heterogeneous grassland in Inner Mongolia: Effects of the plant community and edaphic parameters. Soil Biol Biochem 122:104–110
Li W, Xu F, Zheng S, Taube F, Bai Y (2017) Patterns and thresholds of grazing-induced changes in community structure and ecosystem functioning: species-level responses and the critical role of species traits. J Appl Ecol 54:963–975
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet journal 17:10–12
Martínez-García LB, Richardson SJ, Tylianakis JM, Peltzer DA, Dickie IA (2015) Host identity is a dominant driver of mycorrhizal fungal community composition during ecosystem development. New Phytol 205:1565–1576
Mendoza R, Cabello M, Anchorena J, García I, Marbán L (2011) Soil parameters and host plants associated with arbuscular mycorrhizae in the grazed Magellanic steppe of Tierra del Fuego. Agr Ecosyst Environ 140:411–418
Moora M, Zobel M (2010) Arbuscular mycorrhizae and plant-plant interactions. Positive plant interactions and community dynamics, pp 79–98
Murray TR, Frank DA, Gehring CA (2010) Ungulate and topographic control of arbuscular mycorrhizal fungal spore community composition in a temperate grassland. Ecology 91:815–827
O’Mara FP (2012) The role of grasslands in food security and climate change. Ann Bot 110:1263–1270
Oehl F, Sieverding E, Ineichen K, Maeder P, Wiemken A, Boller T (2009) Distinct sporulation dynamics of arbuscular mycorrhizal fungal communities from different agroecosystems in long-term microcosms. Agr Ecosyst Environ 134:257–268
Pinheiro J, Bates D, DebRoy S, Sarkar D, Team RC (2018) nlme: linear and nonlinear mixed effects models. R package version 3.1-137. https://CRAN.R-project.org/package=nlme
R Core Team (2021) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Ren H, Eviner VT, Gui W, Wilson GW, Cobb AB, Yang G et al (2018) Livestock grazing regulates ecosystem multifunctionality in semi-arid grassland. Funct Ecol 32:2790–2800
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. J PeerJ 4:e2584
Sato K, Suyama Y, Saito M, Sugawara K (2005) A new primer for discrimination of arbuscular mycorrhizal fungi with polymerase chain reaction-denature gradient gel electrophoresis. Grassland Sci 51:179–181
Sepp SK, Davison J, Jairus T, Vasar M, Moora M, Zobel M, Öpik M (2019) Non-random association patterns in a plant–mycorrhizal fungal network reveal host–symbiont specificity. Mol Ecol 28:365–378
Smith SE, Read DJ (2008) Mycorrhizal symbiosis. Academic press, San Diego
Staddon PL, Ramsey CB, Ostle N, Ineson P, Fitter AH (2003) Rapid turnover of hyphae of mycorrhizal fungi determined by AMS microanalysis of 14C. Science 300:1138–1140
Stevens BM, Propster JR, Öpik M, Wilson GW, Alloway SL, Mayemba E, Johnson NC, (2020) Arbuscular mycorrhizal fungi in roots and soil respond differently to biotic and abiotic factors in the Serengeti. Mycorrhiza 30(1):79–95
Stover HJ, Naeth MA, Boldt-Burisch K (2018) Soil disturbance changes arbuscular mycorrhizal fungi richness and composition in a fescue grassland in Alberta Canada. Appl Soil Ecol 131:29–37
Vályi K, Rillig MC, Hempel S (2015) Land-use intensity and host plant identity interactively shape communities of arbuscular mycorrhizal fungi in roots of grassland plants. New Phytol 205:1577–1586
van der Heyde M, Bennett JA, Pither J, Hart M (2017) Longterm effects of grazing on arbuscular mycorrhizal fungi. Agr Ecosyst Environ 243:27–33
van der Heyde M, Ohsowski B, Abbott LK, Hart M (2017) Arbuscular mycorrhizal fungus responses to disturbance are context-dependent. Mycorrhiza 27:431–440
van der Heyde M, Abbott LK, Gehring C, Kokkoris V, Hart MM (2019) Reconciling disparate responses to grazing in the arbuscular mycorrhizal symbiosis. Rhizosphere 11:100167
Větrovský T, Baldrian P (2013) Analysis of soil fungal communities by amplicon pyrosequencing: current approaches to data analysis and the introduction of the pipeline SEED. Biol Fertil Soils 49:1027–1037
Wang Q, Bao Y, Liu X, Du G (2014) Spatio-temporal dynamics of arbuscular mycorrhizal fungi associated with glomalin-related soil protein and soil enzymes in different managed semiarid steppes. Mycorrhiza 24:525–538
Yan L, Zhou G, Zhang F (2013) Effects of different grazing intensities on grassland production in China: a meta-analysis. PLoS One 8:e81466
Yang X, Chen J, Shen Y, Dong F, Chen J (2020) Global negative effects of livestock grazing on arbuscular mycorrhizas: a meta-analysis. Sci Total Environ 708:134553
Zhu L (2005) Monte Carlo Tests. In: Zhu L (ed) Nonparametric Monte Carlo Tests and Their Applications. Springer New York, New York, pp 1–9
Zuur AF, Ieno EN, Walker NJ, Saveliev AA, Smith GM (2009) Mixed effects modelling for nested data. In: Mixed effects models and extensions in ecology with R. Springer, New York, NY, pp 101–142
Acknowledgements
We acknowledge the kind cooperation of the staff in the Inner Mongolia Grassland Ecosystem Research Station (IMGERS) led by Chinese Academy of Sciences to provide equipment and facilities. Particular thanks to our research team Chuanyu Zhou, Zirui Hu, Ye Wang, Hanyu Jian, Shasha Zou, Fanjing Liu and Yuwei He for their collaborative effort during data collection.
Funding
This work has been funded by the Xian-Jiatong-Liverpool University Research Development Fund (RDF-15-02-13).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Faghihinia, M., Zou, Y., Bai, Y. et al. Grazing Intensity Rather than Host Plant’s Palatability Shapes the Community of Arbuscular Mycorrhizal Fungi in a Steppe Grassland. Microb Ecol 84, 1062–1071 (2022). https://doi.org/10.1007/s00248-021-01920-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-021-01920-7