Abstract
Host-microbiome dynamics occurring in the yellow fever mosquito (Aedes aegypti) contribute to host life history traits, and particular bacterial taxa are proposed to comprise a “core” microbiota that influences host physiology. Laboratory-based studies are frequently performed to investigate these processes; however, experimental results are often presumed to be generalizable across laboratories, and few efforts have been made to independently reproduce and replicate significant findings. A recent study by Muturi et al. (FEMS Microbiol Ecol 95 (1):213, 2019) demonstrated the food source imbibed by laboratory-reared adult female mosquitoes significantly impacted the host-associated microbiota—a foundational finding in the field of mosquito biology worthy of independent evaluation. Here, we coalesce these data with two additional mosquito-derived 16S rRNA gene sequence data sets using a unifying bioinformatics pipeline to reproduce the characterization of these microbiota, test for a significant food source effect when independent samples were added to the analyses, assess whether similarly fed mosquito microbiomes were comparable across laboratories, and identify conserved bacterial taxa. Our pipeline characterized similar microbiome composition and structure from the data published previously, and a significant food source effect was detected with the addition of independent samples, increasing the robustness of this previously discovered component of mosquito biology. However, distinct microbial communities were identified from similarly fed but independently reared mosquitoes, and surveys across all samples did not identify conserved bacterial taxa. These findings demonstrated that while the main effect of the food source was supported, laboratory-specific conditions may produce inherently differential microbiomes across independent laboratory environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbial communities are an important component of mosquito biology, although the true nature of these dynamic relationships is not yet fully understood. Gut bacteria contribute to the fundamental life history traits of blood digestion and egg production [1], while both gut and whole-body bacteria contribute to development [2,3,4] of the yellow fever mosquito (Aedes aegypti) and other mosquito species. Specifically, the reduction of the gut microbial community negatively impacted digestive processes and reduced egg production [1]. Furthermore, the presence or absence of particular bacterial taxa differentially impacted host development [2, 3], and larval development was significantly stymied when hosts were deprived of microbial communities [4]. However, the relative significance of these mosquito-microbe interactions remains unclear [5]. Certain bacterial taxa were detected across multiple mosquito studies in natural and laboratory environments and have been proposed to constitute a “core” microbiota that plays significant roles in mosquito physiology [6, 7], although no consensus has been reached regarding the validity of this assertion [8].
Variable environmental microbiota contributes to disparate host-associated microbiomes [9, 10], making the identification of significant bacterial taxa difficult when analyzing mosquito-associated microbial communities originating from natural environments. Laboratory-based experiments that rear mosquitoes under controlled conditions are often performed to mitigate these impacts, and thus investigate directly specific experimental treatments’ effects on mosquito-microbiome dynamics. For example, a recent study discovered that the food source fed to laboratory-reared adult female mosquitoes (newly emerged adults that had not consumed a meal [hereafter referred to as Unfed] of sugar, human blood [3- and 7-day digestion], chicken blood, or rabbit blood) impacted various characteristics of the mosquito-associated gut microbiota [11]. These results complement a prior study that revealed influences of host diet on the microbiota associated with the mosquito Anopheles gambiae [12]. Significantly, gut microbial communities can influence host-pathogen interactions and the mosquito’s vector competence (capacity to contract and transmit a variety of pathogens) [13,14,15,16,17,18]. Studying factors that cause shifts in mosquito-associated microbial communities could reveal important characteristics of mosquito biology that drive the global spread of pathogens and impact host life history traits.
Although these findings and those from other laboratory-based studies have proven invaluable to our understanding of how biotic and abiotic factors influence microbial communities, it is often presumed that laboratory dynamics are generalizable, and efforts to reproduce and replicate significant results are rarely performed. Studies characterizing similarly reared mosquito microbiomes across laboratories are essential to validate this widespread assumption and promote the accurate identification of significant mosquito-microbe interactions. Significantly, these studies investigate microbial communities isolated from mosquito guts, other anatomic regions, or the entire animal body, and inter-study comparisons of experimental findings must take these differences into account when interpreting results.
Bacterial taxa in the genus Clostridium identified from whole body microbial communities [4], and the genera Asaia, Enterobacter, Pseudomonas, Elizabethkingia, and Serratia from gut microbiota [19,20,21] have been detected across independent mosquito samples [4, 19, 20] collected from laboratory environments. Although this could indicate that certain significant microbes are conserved across laboratory settings, there is also evidence that the gut microbiome of mosquitoes is determined entirely by the insectary environment in which the animals are reared [22]. Additional research demonstrated that the laboratory environment and other study-specific conditions, such as the sequential cohort of the experimental animals and the microbiota found in the larval rearing water, impact host-associated microbiomes in the mosquito species A. gambiae and Aedes albopictus [23]. These findings suggest results produced from laboratory-based studies may not be reproducible across different laboratory environments, bringing in to question the validity of generalizing conserved taxa-specific host-microbe dynamics without independent reproduction and replication of experimental results. The identification and confirmation of core laboratory microbes could serve as an important first step in the unification of laboratory-based mosquito microbiome studies. Reanalyzing data collected from independent laboratory-based mosquito projects may help to address these questions. We are not aware of any studies that have attempted to compare independently produced laboratory-reared A. aegypti 16S rRNA gene sequence data sets.
Disparate methods used to process 16S rRNA sequencing data introduce challenges for comparative analyses of independently collected data. For example, amplicon sequencing data are prepared with different processing methods, such as differential sequence error-correction strategies, that can affect downstream analyses [24]. Disparate, pipeline-specific sequencing read quality filtering methods may also influence results. Furthermore, different sequence clustering strategies produce operational taxonomic units (OTUs) or amplicon sequence variants (ASVs) that vary in quality and utility [25]. Amplicon sequence variants are now widely considered superior to OTUs; however, past studies that performed OTU clustering have not been re-evaluated with ASV-based methods. In addition, distinct 16S sequence databases are used to assign taxonomy to sequences, possibly influencing taxonomic identifications [26]. Finally, contaminant sequences introduced through DNA extraction kits and other sources [27] are not always accounted for due to the absence of negative control sample sequencing during experimental workflows. Taken together, these factors associated with microbiome research can impede the replication and reproduction of previously published results [28].
Normalizing inter-experimental data through the implementation of a conserved bioinformatics pipeline using the same sequence data processing, clustering, taxonomic assignment, and decontamination strategies could mitigate variation resulting from disparate data handling protocols and promote a more accurate coalescence and analysis of mosquito microbiome data sets. Once normalized, laboratory-specific effects could be incorporated and accounted for when performing standard microbiome comparative analyses to provide a framework to test the reproducibility, replicability, and robustness of previous findings and provide novel opportunities to investigate core laboratory-associated bacterial taxa and the impact of experimental treatments on microbial communities.
To advance the field’s understanding of the impact of food source and differential mosquito-associated microbiota across laboratory environments, we coalesced and re-analyzed the sequencing data collected from gut microbiomes in Muturi et al. 2019 (hereafter, Muturietal_2019), and from two additional laboratory-produced 16S data sets derived from whole-body microbiomes for mosquitoes fed sugar (29 [hereafter, Hegdeetal_2018]) as well as newly emerged adults that had not imbibed a meal (30 [hereafter, FrankelBrickeretal_2020]), by implementing a unifying bioinformatics pipeline to normalize these data for downstream analyses. The pipeline implemented here (DADA2; 31) processed sequencing reads similarly using conserved quality filtering and sequence-error correction methods, merged reads into ASVs, and assigned taxonomy using the same 16S rRNA gene sequence reference database (SILVA v132; 32,33). Furthermore, reads for each sample were rarefied to account for variable sequencing coverage across samples.
First, we evaluated whether our pipeline identified the same prominent bacteria classified from the previous studies to assess whether our pipeline detected microbes similarly to those found previously. Then, we tested whether the significant effect of food source on previously investigated and additional diversity metrics was reproducible and replicable when additional samples from other studies were added to the analysis. We subsequently assessed whether microbial communities harbored in mosquitoes fed the same food source were comparable across laboratories. Finally, we surveyed all experimental samples for the presence of previously proposed core laboratory microbes to evaluate the prevalence of these taxa across food sources and laboratories.
We predicted that our pipeline would successfully identify prominent bacterial taxa from each of the studies analyzed and reproduce the major finding of a significant effect of food source on laboratory mosquito-associated microbiota, however, that distinct microbiome characteristics (the overall diversity and relative abundance distribution of microbial community members) would be detected for microbiota harbored in mosquitoes fed the same food source but originating from different laboratories. We also predicted that core bacterial taxa would be observed due to the significant roles that microbes play in mosquito physiology. Significantly, the studies analyzed here provided data derived from the gut- (Muturietal_2019) and whole body- (Hegdeetal_2018, FrankelBrickeretal_2020) associated microbiota, which required us to tailor our interpretations of results to account for potentially disparate characterizations of microbes found within, and external to, the mosquito gut. We proceeded under the assumption that data from the whole body constituted a characterization of the complete mosquito-associated microbiome (including the gut), whereas gut microbiome samples represented a subset of the whole-body microbiota.
Results
Alpha Diversity Analyses
Differences across food sources were detected for both Simpson (F(5, 201) = 11.134, P < 0.0001; Fig. 1a, Table 1) and Shannon (F(5, 202) = 9.992, P < 0.0001; Fig. 1b, Table 1) diversity indices, showing a significant effect of food source on the overall diversity and evenness of microbial communities. Post-hoc pairwise comparisons revealed significant differences between particular food sources after Bonferroni correction for both metrics, suggesting certain food sources had more pronounced influences on measures of alpha diversity (Table S1).
Box plots of measures of Simpson (a, c), and Shannon (b, d) alpha diversity indices and significant results from pairwise comparisons for samples across all food groups (a, b, respectively; Unfed [n = 49], Sugar [n = 60], Human_3 [n = 27], Human_7 [n = 25], Chicken [n = 36], Rabbit [n = 12]) and studies within shared food groups (c, d, respectively; Unfed [Muturietal_2019 (n = 30), FrankelBrickeretal_2020 (n = 19)], Sugar [Muturietal_2019 (n = 35), Hegdeetal_2018 (n = 25)]). Upper and lower limits of boxes represent quartiles around the mean; horizontal lines within boxes represent median values within each group compared. Significant pairwise comparisons of mean alpha diversity values across all food source groups were calculated with post-hoc simultaneous tests for general linear hypotheses with P-values adjusted with the Bonferroni method (*P < 0.05, **P < 0.01, ***P < 0.001) and Unfed and Sugar samples across studies with Wilcoxon rank sum tests (***P < 0.001)
A significant study effect was detected for the Unfed and Sugar groups for both metrics ([Simpson: W = 442, P < 0.001; W = 777, P < 0.0001, respectively; Fig. 1c, Table 1], [Shannon: W = 457, P < 0.001; W = 750, P < 0.0001, respectively; Fig. 1d, Table 1]), revealing differential microbial community diversity and evenness across studies within shared food source groups (Table 1).
Beta Diversity Analyses
After total read processing, 743 ASVs were identified across 209 samples. A significant food source effect for all beta diversity metrics tested was detected (Bray-Curtis dissimilarity [R = 0.314, P < 0.0001; Fig. 2a], unweighted UniFrac distance [R = 0.417, P < 0.0001; Fig. 2d], weighted UniFrac distance [R = 0.377, P < 0.0001; Fig. 2g]; Table 1), demonstrating an influence of food source on microbiome composition and structure. PCoA plots were constructed to visualize relative clustering of samples for each metric and captured higher relative variation along the x-axis (Bray-Curtis dissimilarity: 23%, unweighted UniFrac distance: 26%, weighted UniFrac distance: 35.9%). Relatively high R values generated from pairwise ANOSIM, as well as distinct clustering of samples from certain food groups, provided high confidence in the significant influence of food source on the metrics tested. In particular, the Unfed and Sugar groups significantly differed for many of the comparisons with the other groups for Bray-Curtis dissimilarity, indicated by low P-values and relatively high R values (Table S1). Furthermore, differences were found between the Human_3 and Human_7 blood groups as well as several other comparisons. Please see Table S1 for a detailed summary of all results from pairwise ANOSIM. Only three significant pairwise comparisons with the Unfed group were identified for unweighted UniFrac distance, whereas Human_3 and Human_7 were the only other significant comparison for this metric. Significant pairwise differences in group dispersions were detected between certain food sources for all metrics and across studies (Table 1; Table S1), revealing differential within-group variation possibly leading to false positives produced by ANOSIM and PERMANOVA. In particular, dispersions of Bray-Curtis dissimilarity and unweighted UniFrac distance in the Unfed group significantly differed with many of the other groups (Table S1). However, PCoA plots show distinct centroids of Unfed clusters relative to many of the other groups (Figs. 2a, d, g), increasing confidence in these results.
Principal coordinates analysis plots of beta diversity measures calculated across all food groups (column 1; Unfed [n = 49], Sugar [n = 60], Human_3 [n = 27], Human_7 [n = 25], Chicken [n = 36], Rabbit [n = 12]), across studies within the Unfed group (column 2; Muturietal_2019 (n = 30), FrankelBrickeretal_2020 (n = 19)), and across studies within the Sugar group (column 3; Muturietal_2019 (n = 35), Hegdeetal_2018 (n = 25)) for Bray-Curtis dissimilarity (a, b, c), unweighted UniFrac distance (d, e, f), and weighted UniFrac distance (g, h, i), respectively. Ninety-five percent confidence ellipses are provided to aid with the visualization of relative clustering
A significant study effect was found for both Unfed and Sugar groups for Bray-Curtis dissimilarity (F (1, 47) = 9.577, R2 = 0.169, P < 0.0001, Fig. 2b; F (1, 58) = 114.13, R2 = 0.663, P < 0.0001, Fig. 2c, respectively), unweighted UniFrac distance (F (1, 47) = 12.75, R2 = 0.213, P < 0.0001, Fig. 2e; F (1, 58) = 29.229, R2 = 0.335, P < 0.0001, Fig. 2f, respectively), and weighted UniFrac distance (F (1, 47) = 16.038, R2 = 0.254, P < 0.0001, Fig. 2h; F (1, 58) = 186.43, R2 = 0.763, P < 0.0001, Fig. 2i, respectively) (Table 1), demonstrating that microbial community composition and structure associated with mosquitoes fed the same food source differed across studies (Table 1). In addition, differential intra-study beta diversity variation was found for comparisons of unweighted UniFrac distance across studies within the shared food groups (Table 1); however, distinct clustering was observed for these comparisons (Figs. 2e, f).
Bacterial Taxa Relative Abundance
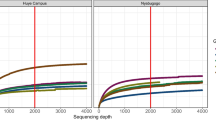
Relative abundances of prominent families across all studies and food groups were calculated (Fig. S1, Table S2). Families identified from the Muturietal_2019 samples reproduced generally the identification and quantification of major taxa published previously. Specifically, Enterobacteriaceae (Proteobacteria) and Weeksellaceae (Bacteroidetes) were present at relatively high abundance across all blood groups (Enterobacteriaceae [Human_3: 70.89%, Human_7: 34.85%, Chicken: 22.7%, Rabbit: 36.48%]; Weeksellaceae [Human_3: 21.93%, Human_7: 32.31%, Chicken: 61.37%, Rabbit: 47.26%]). In contrast, Nocardioidaceae (Actinobacteria) was found at intermediate abundance in Human_7 (22.53%) and Chicken (9.65%) groups, but at very low levels in Human_3 (3.25%) and Rabbit (0.04%) groups. Furthermore, this family was most abundant in the Unfed and Sugar groups (40%, 65.68%, respectively), followed by Microbacteriaceae (Actinobacteria, 26.50%) in the Unfed group and Acetobacteraceae (Proteobacteria, 22.84%) in the Sugar group. In addition, the quantification of major families from FrankelBrickeretal_2020 samples reproduced largely the previously published findings. Burkholderiaceae (Proteobacteria) was identified as the dominant family (40.62%), followed by Pseudomonadaceae (Proteobacteria, 7.93%), Staphylococcaceae (Firmicutes, 7.23%), and Enterobacteriaceae (Proteobacteria, 5.64%). Family-resolution characterization of the Hegdeetal_2018 samples were reproduced largely as well. These communities were dominated by taxa within the family Enterobacteriaceae (Proteobacteria, 89.60%) and lower levels of Acetobacteraceae (Proteobacteria, 7.89%).
Calculations of relative abundance for the most abundant phyla for the shared food groups revealed Proteobacteria, Actinobacteria, Bacteroidetes, and Firmicutes were conserved as the most abundant taxa in the Unfed group and Proteobacteria and Actinobacteria in the Sugar group (Table 2, Fig. 3). However, the relative abundances of Proteobacteria, Actinobacteria, and Firmicutes were not conserved across studies in the Unfed (W = 500, P < 0.0001; W = 54, P < 0.0001; W = 529, P < 0.0001, respectively) and Sugar (W = 870, P < 0.0001; W = 0, P < 0.0001; W = 519, P = 0.04, respectively) groups. In the Unfed group, Actinobacteria was significantly more abundant in Muturietal_2019 (66.96%) relative to FrankelBrickeretal_2020 (10.39%), whereas Proteobacteria and Firmicutes were higher in FrankelBrickeretal_2020 (64.53%, 14.51%) relative to Muturietal_2019 (20.4%, 2.13%), respectively. In the Sugar group, Actinobacteria dominated Muturietal_2019 (65.76%) but was nearly indetectable in Hegdeetal_2018 samples (0.13%). In contrast, Proteobacteria was the prominent phylum in Hegdeetal_2018 (98.42%) and found in intermediate abundance in Muturietal_2019 (33.71%). Inter-sample visualization with heatmaps of relative abundances of families within these phyla show patterns of intra-study variation in both abundances and families present for both shared groups (Fig. S2).
Core Laboratory Microbes
Presence-absence surveys for genera within the Class Clostridia (Firmicutes) for studies and food sources within studies revealed differential prevalence of all genera in samples across studies (Table S3). Furthermore, while certain genera were detected in relatively high numbers of samples within each study, no single genus was universally found across all samples within a study or food group. The genus found in the greatest number of samples for each study and food source was: Muturietal_2019 (Unfed: Ethanoligenens (Ruminococcaceae), 3.33% of samples; Sugar: no genera from Clostridia present; Human_3 blood: No genera from Clostridia present; Human_7 blood: Romboutsia (Peptostreptococcaceae), 4% of samples; Chicken blood: Christensenellaceae_R-7_group (Christensenellaceae), Romboutsia (Peptostreptococcaceae), Roseburia (Lachnospiraceae), Blautia (Lachnospiraceae), 2.78% of samples; Rabbit blood: Ruminococcaceae_NK4A214_group (Ruminococcaceae), Romboutsia (Peptostreptococcaceae), Blautia (Lachnospiraceae), 8.33% of samples); FrankelBrickeretal_2020 (Unfed: Finegoldia (Family_XI), Anaerococcus (Family_XI), 42.11% of samples); Hegdeetal_2018 (Sugar: Paraclostridium (Peptostreptococcaceae), 24% of samples).
Additional presence-absence surveys were conducted for the genera Asaia (Acetobacteraceae), Enterobacter (Enterobacteriaceae), Pseudomonas (Pseudomonadaceae), Elizabethkingia (Weeksellaceae), and Serratia (Enterobacteriaceae) (Table 3) across all samples. Asaia was found in the majority of samples from the Sugar groups from Muturietal_2019 (88.57%) and Hegdeetal_2018 (100%), respectively. However, this genus was not detected in any samples from the other groups. Enterobacter was detected in the majority of samples for the Rabbit (58.33%) and Chicken (52.78%) blood groups from Muturietal_2019, although this genus was found at lower levels or entirely absent across samples from the other groups. Pseudomonas was present in the majority of samples for FrankelBrickeretal_2020 (78.95%) and the Sugar samples from Muturietal_2019 (62.86%) and Hegdeetal_2018 (60%), but at lower levels in the other groups. Elizabethkingia followed similar trends of variable prevalence and was detected across the majority of samples for the blood groups of Muturietal_2019 (Human_3: 62.96%; Human_7: 56%; Chicken: 61.11%; Rabbit: 50%) but in lower numbers or completely absent from the other groups. Finally, Serratia was detected in all samples from the two Human blood groups from Muturietal_2019, but in fewer samples or entirely absent from the other groups.
Discussion
The analyses presented herein are offered as a comprehensive evaluation of the literature on the effect of food source on the adult female A. aegypti-associated microbiota, assessment of laboratory-specific microbiomes, and an investigation of the prevalence of putatively significant core laboratory microbes. Our data workflow implemented a normalized bioinformatics pipeline to coalesce raw 16S sequencing data collected from independent studies to mitigate variation incurred by data processing methods and promote comparisons of host-associated microbiota from laboratory-reared mosquitoes across laboratories.
By processing raw 16S sequencing data collected from different studies, we attempted to minimize differences found across data sets incurred by variable read-processing strategies. By implementing the DADA2 pipeline [31] for all sequencing data to produce ASVs, we minimized the likelihood of differential outputs due to variable sequence-error correction strategies [24]. Furthermore, assigning taxonomy to all ASVs using the SILVA v132 database [32, 33] as a reference ensured consistent taxonomic designations across data sets [26]. Contaminant sequences are often introduced to samples through DNA extraction kits, PCR reagents, and other external sources and can impact downstream microbiome characterization [27]. Only one of the original data sets we analyzed provided negative control sequences (DNA extraction kit reagents) and each study implemented a different DNA extraction protocol. Eleven putative contaminant sequences from the negative controls were identified from FrankelBrickeretal_2020 and four of the eleven from Hegdeetal_2018 samples and were removed prior to analysis. Because no contaminant sequences were subtracted from Muturietal_2019, we cannot be sure whether our decontamination-based strategy served as an effective method across samples from different studies. We acknowledge that this likely influenced the representation of certain ASVs for the data sets that did not provide negative control data; however, we prioritized removing known contaminant ASVs identified by our pipeline. It is possible that the four contaminant ASVs shared between Hegdeetal_2018 and FrankelBrickeretal_2020 represented laboratory independent contaminants, whereas the seven sequences unique to FrankelBrickeretal_2020 may have originated from laboratory dependent sources. It is also possible that the different DNA extraction protocols and mosquito storage conditions could have produced divergent contaminant ASVs. However, the absence of negative controls from two of the three studies prevents further assessment of these concepts. We strongly encourage future microbiome studies to include a comprehensive suite of negative controls and provide all relevant information pertaining to the DNA extraction protocols conducted to promote more accurate removal of putative contaminant sequences.
To assess the capacity of our pipeline to accurately analyze data previously processed by different methods, we investigated whether the prominent bacterial taxa reported in the previous studies could be detected. Our pipeline identified and quantified the same major families for the blood (Enterobacteriaceae, Weeksellaceae, Nocardioidaceae, Burkholderiaceae), Sugar (Nocardioidaceae, Acetobacteraceae), and Unfed (Nocardioidaceae, Microbacteriaceae, Burkholderiaceae) groups originally characterized in Muturietal_2019 (Fig. S1, Table S2), demonstrating taxonomic assignments for these data were conserved despite disparate read processing methods. Similar accuracy was found for our analyses of the data provided in FrankelBrickeretal_2020, which identified Burkholderiaceae, Pseudomonadaceae, and Staphylococcaceae as the prominent families (Fig. S1, Table S2). Family-level characterization was also conserved for Hegdeetal_2018, where Enterobacteriaceae from the Phylum Proteobacteria dominated microbiomes. These general results indicate our pipeline was effective at reproducing prominent taxa identified previously and suggest the characterization and quantification of major microbial community members may not be substantially impacted by differential bioinformatics pipelines implemented during data processing.
In addition, we investigated whether the significant effect of adult food source on the host-associated microbiome could be reproduced for alpha and beta diversity metrics previously analyzed by Muturietal_2019. The significant impact of food source was reproduced for measures of Shannon diversity (Fig. 1b, Table 1) and Bray-Curtis dissimilarity (Fig. 2a, Table 1). The addition of samples to the Unfed and Sugar groups did not impact this overall significant effect, increasing the robustness of the previously published findings. These results suggest that while our pipeline did not replicate all significant pairwise comparisons previously found in Muturietal_2019, the conserved detection of the major impact of food source and the identification of prominent taxa at the family level provide support that our pipeline was suitable for inter-study comparative analyses.
Additional metrics were calculated for alpha (Simpson [Table 1; Table S4, Fig. 1a]) and beta (unweighted and weighted UniFrac distances [Table 1, Figs. 2d, g, respectively]) diversity indices, providing added information to the findings published previously in Muturietal_2019. A strong food source effect for Simpson diversity complements the previous (and now reproduced) finding of distinct Shannon diversity measures across certain food groups, revealing that the microbiota had differential levels of evenness across food sources. Although the main food source treatment effect was reproduced, differences in the specific pairwise comparisons underlying these trends were identified (Table S1). Specifically, our results show microbiota harbored in newly emerged adults were either less diverse or not significantly different from the other groups, contrary to the higher diversity previously reported for these mosquitoes. We also identified previously undetected differences between blood groups for the Shannon diversity index. These divergent results may derive from specific differences in the way data were processed between studies. Our pipeline organized sequences into ASVs rather than OTUs, possibly resulting in differential calculations of these metrics. Furthermore, our implementation of divergent read processing and filtering methods relative to those performed previously may have also contributed to these differences. These findings indicate that the implementation of a different bioinformatics pipeline yielded similar conclusions regarding the main treatment effect but altered the characterization of specific relationships underpinning these results. Analyses of unweighted and weighted UniFrac distances uncovered significant differences for certain pairwise comparisons between the Unfed and the other food groups (Table S1). Significant differences between the human blood groups were identified as well. Including these metrics, along with the previously published finding of a significant food source effect on Bray-Curtis dissimilarity values, indicates that while microbiota structures differ across groups, taxonomic compositions may be relatively similar. Although these trends may indicate truly differential microbiome compositions and relative abundances of microbial community members, they may also reflect skewed measures due to the addition of data derived from whole-body microbiomes to the Unfed and Sugar groups. We also performed essential tests to assess the homogeneity of within-group beta diversity variation [34, 35], which provided important contextual information regarding the potential biases of our statistical tests. Numerous significant differences in the within-group dispersions for the Unfed and Sugar groups relative to other groups may reflect similarly the influence of including samples derived from whole-body microbiomes for these measures.
Our analyses discovered study-specific alpha and beta diversity measures (Table 1) for microbiomes associated with mosquitoes fed with sugar or newly emerged adults, revealing distinct community diversities and disparate compositions and structures (Figs. 1c, d). Genetically and geographically diverse mosquitoes were previously shown to harbor similar microbiomes when fed sugar and reared in a controlled laboratory environment, indicating laboratory-specific conditions influence the results of laboratory-based microbiome studies [22]. Our findings support their conclusions by revealing disparate, laboratory-specific microbiota. Significantly, these results were likely influenced by characterizations of overlapping yet differential microbial communities (gut and whole body). Higher alpha diversity values for whole body samples (FrankelBrickeretal_2020, Hegdeetal_2018) relative to gut samples (Muturietal_2019) for both the Unfed and Sugar groups (Figs. 1c, d, Table 1) suggested that data from whole-body microbiomes were more diverse due to the identification of microbes including and external to the gut microecosystem. These findings do indicate that diverse and distinct communities of microbes inhabit regions outside of the gut, which has been shown in previous studies [36,37,38]. Similarly, strong significant differences and disparate clustering were detected across studies for both shared food groups (Fig. 2b, c, e, f, h, i, Table 1). Although likely impacted by the nature of the microbiomes characterized and compared, the clear and distinct clustering and significant results from statistical tests may reflect truly divergent microbiomes harbored in mosquitoes reared in independent laboratories. Future studies could collect and compare microbiomes derived from the same source to resolve these potential issues in our analyses.
Sugar is a common food source for laboratory-reared mosquitoes [39], yet large-scale microbiome differences at the Phylum level from different laboratories demonstrated that the ingestion of sugar may not have resulted in a conserved host-associated microbiota (Fig. 3b, Table 2). In addition, variation in the relative abundances for families within these phyla indicated laboratory-specific effects could impact the abundances of certain taxa across all taxonomic levels (Fig. S2). These results may have significant implications for the generalizability of laboratory-based mosquito microbiome research. For example, microbiomes harbored in samples collected from Hegdeetal_2018 were dominated by taxa from the phylum Proteobacteria (98.42%), whereas this phylum was detected at intermediate abundance in Muturietal_2019 (33.71%). Conversely, Muturietal_2019 samples contained high levels of Actinobacteria (65.76%), whereas the phylum was nearly indetectable in samples from Hegdeetal_2018 (0.13%). Although these differences may be influenced by disparities between the gut and whole-body microbiomes characterized, the strong significant differences for all metrics tested and taxa characterized between the studies may reflect genuinely distinct microbial communities. The mosquitoes from the two studies could have been reared under disparate environmental conditions, such as temperature and relative humidity, and fed different sugar food sources, contributing to the taxa-specific differences we detected. These results could indicate that certain laboratories rear mosquitoes harboring microbial communities dominated by a single phylum (Hegdeetal_2018, one phylum > 98%), whereas others may rear mosquitoes with more balanced phyla structures (Muturietal_2019, two phyla > 33%). If this is the case, identical experiments conducted in independent laboratories utilizing sugar-fed mosquitoes could yield disparate results when investigating various experimental effects on specific bacterial taxa. Taxa-specific results from laboratory studies may not be applicable or reproducible in other laboratories even if significant experimental treatment effects are conserved. These concepts were demonstrated in another arthropod system, where a treatment effect of temperature on host-associated microbiota was universally detected, but disparate shifts in distinct bacterial taxa were observed [40, 41]. Future research investigating novel experimental questions could also perform comparisons with previously published 16S data sets to gage whether laboratory-specific effects should be accounted for in analyses and interpretations of results.
We similarly identified disparate microbial communities harbored in newly emerged mosquitoes across studies. Specifically, large differences in the relative abundances of Proteobacteria and Actinobacteria suggest that, as with sugar-fed mosquitoes, laboratory-specific conditions may have influenced the relative abundances of prominent taxa at high taxonomic levels (Fig. 3a, Table 2). Newly emerged adults had not consumed a meal; therefore, the host-associated microbiota was not fully established nor influenced by food source. However, it is possible that adults imbibed environmental rearing water that harbored disparate microbes across these studies. As discussed previously, differences across the studies compared may have been affected by the characterizations of gut and whole-body microbiomes, though the strong phylum-level differences we detected do indicate laboratory-specific microbiomes were present. These findings could reveal a potential factor driving the inter-laboratory variation observed in the sugar-fed mosquitoes. Newly emerged adults inherit a subset of microbes from larvae through transstadial transmission (bacteria transferred from larvae through pupae to adult; 4, 12, 42, 43), potentially indicating that the disparate microbiomes we identified were the result of differential transstadial transmission processes of certain bacterial taxa. Larval rearing conditions may have contributed to these dynamics. Significantly, distinct food sources were given to larvae in Muturieetal_2019 (fish food, rabbit food) and FrankelBrickeretal_2020 (fish food), and both larvae and adults may have been reared at different environmental temperatures. Taken together, these and other laboratory- and study-specific conditions, such as host genotype, filial generation, geographic location of laboratories, rearing water conditions, and countless others may inherently contribute to the distinct inter-laboratory microbiota we characterized. Future studies could include larval data along with the adult data to provide additional information on the temporal dynamics of the microbiota across developmental stages for different food sources. Furthermore, larval data could be compared with previous studies to assess whether laboratory-specific effects may influence results.
To investigate whether previously proposed core laboratory microbes were present in our study samples, we performed basic presence-absence surveys to assess the prevalence of these bacteria across studies (Table 3; Table S3). The genus Clostridium in the class Clostridia was suggested as a conserved obligate anaerobe in laboratory environments [4, 7] identified from whole-body microbiomes. We found no evidence for this or any other genus in the class Clostridia being conserved across studies, regardless of food source (Table S3). Though it could be inferred that Clostridium may not inhabit the gut microecosystem, since it was identified from a study that investigated whole-body microbiomes, low detection in the whole body samples from FrankelBrickeretal_2020 and Hegdeetal_2018 suggests the genus and other members of the class Clostridiales do not play universally significant roles in the animals we studied in either gut nor whole-body microbial communities. Further investigations of the prevalence of the genera Asaia, Enterobacter, Pseudomonas, Elizabethkingia, and Serratia across samples within studies (Table 3) challenge the proposed conserved abundance of these genera in laboratory-reared mosquito microbiomes [19,20,21]. Since these genera were identified from studies investigating gut microbiomes, presence of these microbes should have been detected across all of the samples analyzed in our study. However, no single genus was consistently found across samples or studies for any food source. Significantly, particular genera were detected across all or the majority of samples for certain studies and food groups. For example, Asaia was detected in all samples from Hegdeetal_2018 and in 88.57% of the Sugar group samples from Muturietal_2019. Further, Serratia was found in all samples from both human blood groups from Muturietal_2019. These results may suggest that specific microbes are highly prevalent in mosquitoes fed certain food sources; however, we could not detect any universally conserved microbes. Alternatively, particular host genotypes may select for specific core microbes. Each study analyzed herein utilized a different strain of A. aegypti (Rockefeller, Galveston, Gainesville), and coadaptation processes may have driven differential establishment of core microbial communities. In addition, it is possible that a core functional microbiome is conserved across laboratory-reared mosquitoes rather than particular bacterial taxa. Future experiments could implement shotgun sequencing strategies in tandem with 16S rRNA amplicon sequencing to holistically assess mosquito-microbe relationships across different laboratory environments and investigate the functional role of the laboratory-reared mosquito microbiome. Furthermore, non-bacterial microbiome components, such as viruses, eukaryotes, archaea, and fungi, may also be impacted by the dynamics described herein. Future studies could account for these understudied components of mosquito-associated microbiomes to promote a more holistic assessment of biotic factors impacting mosquito biology.
Conclusions
Mosquitoes play important roles in global human health, making studies of the mosquito holobiont scientifically and socially significant. Laboratory-based studies are performed to study various aspects of these microbes and host-microbiome dynamics under controlled laboratory conditions; however, our results suggest that laboratories may inherently rear mosquitoes harboring divergent microbiota potentially influenced by a multitude of laboratory-specific factors not investigated here. If true, the results of these studies may not be generalizable across laboratories. Due to complex and incompletely understood host-microbe interactions, we advise that future laboratory-based studies take into consideration the different laboratory-specific conditions that could impact the host-associated microbiota. Furthermore, the validity and robustness of these studies cannot be fully evaluated until results are replicated under independent laboratory environments.
Materials and Methods
Overview of 16S Mosquito Data Sets
Data sets from previously published laboratory-based studies of adult female A. aegypti microbiomes were selected for analysis based on the following criteria: (1) Raw 16S sequencing reads were publicly available in an accessible and globally recognized data repository, (2) Sequencing data were produced from a modern next-generation sequencing platform, (3) Sequencing data were derived from microbial DNA extractions of individual, rather than pooled, mosquito samples to minimize limitations of assessments of variation in diversity measurements [44], and (4) PCR amplification was performed targeting the V3-V4 hypervariable region of the 16S rRNA gene [45, 46]. Based on a comprehensive literature search, 3 data sets were selected for our study (Table 4).
Muturi et al. 2019 (Muturietal_2019), investigated how different food sources impacted the gut microbiota of age-matched laboratory-reared adult female A. aegypti (Rockefeller strain). The authors found multi-faceted influences of various food sources (Unfed [newly emerged adults], Sugar [10% sucrose solution], human blood 3 days after blood meal [Human_3], human blood 7 days after blood meal [Human_7], chicken blood [Chicken], and rabbit blood [Rabbit]) on the composition and structure of the mosquito-associated microbiota with 2 replicate batches of mosquitoes for each food group. These data are publicly available in the NCBI SRA Bioproject PRJNA494958. All samples were downloaded and processed in our study.
Hegde et al. 2018 (Hegdeetal_2018), analyzed laboratory and wild mosquito-associated whole-body microbiota for multiple species (including A. aegypti; Galveston strain). Laboratory-reared A. aegypti were fed sugar for 5–7 days prior to microbial DNA extraction. Sequencing data are publicly available in the NCBI SRA Bioproject PRJNA422599. The dataset was subset for laboratory-reared A. aegypti samples for our study.
Frankel-Bricker et al. 2020 (FrankelBrickeretal_2020), investigated how an obligate gut fungal symbiont (Zancudomyces culisetae) impacted larval and adult A. aegypti-associated whole-body microbiota. Adult mosquitoes (Gainesville strain) were collected, and microbial DNA extracted immediately after emergence from pupae. These data are publicly available in the NCBI SRA Bioproject PRJNA541017 with negative control sequences provided. The data set was subset for non-fungal adult female samples for our study.
Normalization of 16S rRNA Gene Sequence Data
All data processing and downstream analyses (with the exception of ANOSIM) were performed using the R programming language version 4.0.2 (47; File S1). Sequencing reads from each data set were initially processed using the DADA2 pipeline [31]. For all data sets, reads were trimmed at the location of the first occurrence of a base call with a Phred score less than or equal to 15, reads with any number of N base calls or containing six or more estimated errors were discarded, and forward and reverse reads were merged with a minimum overlap of 12 bases. In addition, 21 base pairs were trimmed from the reverse reads provided by Muturietal_2019 to account for the use of a primer pair (341F/806R; 45) that targeted an overlapping, but larger portion of the 16S V3-V4 hypervariable region targeted by the other two studies (341F/785R; 46). The independently processed sequencing data were combined after chimeric sequences were discarded and merged reads dereplicated. Amplicon sequence variants were produced by the pipeline, and taxonomy was assigned to ASVs using the SILVA v132 database [32, 33].
Phylogenetic Tree and Phyloseq Object Construction
A neighbor-joining tree was inferred using the phangorn package version 2.5.5 in R [48] and a generalized time-reversible with gamma rate variation maximum likelihood tree was fit. The phylogenetic tree was combined with the ASVs, read count data, and sample-associated metadata into a single data object using the Phyloseq package version 1.32 in R [49].
Removal of Contaminant ASV Sequences
The initial Phyloseq object contained data for 267 samples (Table 4). Experimental reagents and other laboratory sources add contaminant sequences to experimental samples [27], yet only one of the three data sets provided sequencing data for negative controls (DNA extraction kit reagents; FrankelBrickeretal_2020, Table 4). Reads from these sources were pooled and putative contaminant ASVs were identified and removed using the decontam package version 1.10.0 in R [50] with the “prevalence” method and the threshold set at 0.5. Eleven putative contaminant sequences were identified and removed from samples in FrankelBrickeretal_2020 and four of the eleven from Hegdeetal_2018. Sequencing data were analyzed for the combined data set of all studies and also subset separately for the 2 shared food sources (Unfed [FrankelBrickeretal_2020; Muturietal_2019], Sugar [Hegdeetal_2018; Muturietal_2019]).
Calculation of Diversity Metrics
Reads were rarefied to 1000 reads per sample to account for the variable read coverage across studies while minimizing the number of samples removed. In total, 209 samples remained for analyses after rarefaction (Table 4). The alpha diversity metrics Simpson and Shannon indices were calculated in Phyloseq (Table S4), and box plots were produced using the ggplot2 package version 3.3.2 in R [51]. Singletons and ASVs not represented by at least five reads or greater in at least one sample were discarded. The beta diversity metrics Bray-Curtis dissimilarity, unweighted UniFrac distance, and weighted UniFrac distance were calculated in Phyloseq. Principal Coordinates Analysis (PCoA) plots were produced in Phyloseq in combination with ggplot2 in R.
Relative Abundances of Prominent Taxa
Relative abundances of the five most prevalent families identified from each study and food source were calculated for all food source groups and studies (Fig. S1, Table S2) to compare whether our bioinformatics pipeline reproduced the identification of major bacterial families and quantification of these microbiota. Reads assigned to these families accounted for over 95% of all reads analyzed in the study. In addition, relative abundances were calculated for the ten most abundant phyla across studies within the Unfed group and the six most abundant phyla from the Sugar group, respectively (Table 2). Relative abundances of families within the four most abundant phyla identified from the Unfed group accounting for greater than 91% of reads (Proteobacteria, Firmicutes, Actinobacteria, Bacteroidetes) and the two identified from the Sugar group accounting for greater than 98% of reads (Proteobacteria, Actinobacteria) across samples were visualized with heatmaps constructed in ggplot2 (Fig. S2). Presence-absence surveys were performed for taxa from the class Clostridia and the genera Asaia, Enterobacter, Pseudomonas, Elizabethkingia, and Serratia by counting the number of experimental samples harboring detectable levels of these bacteria across studies and food groups and subsequently calculating the total percentage of samples containing these taxa, respectively (Table 3; Table S3).
Statistical Analyses
A linear mixed effects model accounting for a random effect of each study was constructed with the lme4 package version 1.1–23 in R [52] to test whether the food source impacted mean Simpson and Shannon diversity values (Table 1). Statistical significance was tested with type II Wald F tests with Kenward-Roger degrees of freedom using the car package version 3.0–9 in R [53]. Post-hoc pairwise comparisons across food sources were conducted with simultaneous tests for general linear hypotheses using the multcomp package version 1.4–15 in R (54; Table S1), and P-values were adjusted with the Bonferroni method. Wilcoxon rank sum tests were performed on samples from the Unfed and Sugar groups to test for a study-specific effect on mean alpha diversity measures and mean relative abundances of prominent bacterial phyla (Tables 1 and 2). Pairwise analysis of similarities (ANOSIM; 55) was conducted to compare beta diversity metrics using similar methodology as Muturi et al. 2019 across food sources with PAST version 4.01 [56] with 9999 permutations (Table 1; Table S1) and P-values were adjusted with the Bonferroni method. Permutational multivariate analysis of variance (PERMANOVA; 57) was conducted to test for differences across studies for the Unfed and Sugar groups with 9999 permutations (Table 1) using the vegan package version 2.5–6 in R [58]. Both ANOSIM and PERMANOVA are sensitive to differential within-group beta diversity variation, which can lead to false positives output from these tests [59]. Permutational statistical tests for the homogeneity of group dispersions [34] were performed with 9999 permutations to detect significant differences in beta diversity variation [35] across groups using vegan (Table 1; Table S1). All statistical analyses were conducted at an α of 0.05.
Data Availability
Locations of the data sets in publicly available repositories are provided in Table 4.
References
Gaio ADO, Gusmão DS, Santos AV, Berbert-Molina MA, Pimenta PFP, Lemos FJA (2011) Contribution of midgut bacteria to blood digestion and egg production in Aedes aegypti (Diptera: Culicidae). Parasites Vectors 4:105
Coon KL, Brown MR, Strand MR (2016) Gut bacteria differentially affect egg production in the anautogenous mosquito Aedes aegypti and facultatively autogenous mosquito Aedes atropalpus (Diptera: Culicidae). Parasites Vectors 9:375
Chouaia B, Rossi P, Epis S, Mosca M, Ricci I, Damiani C, Ulissi U, Crotti E, Daffonchio D, Bandi C, Favia G (2012) Delayed larval development in Anopheles mosquitoes deprived of Asaia bacterial symbionts. BMC Microbiol. 12:S2
Coon KL, Vogel KJ, Brown MR, Strand MR (2014) Mosquitoes rely on their gut microbiota for development. Mol. Ecol. 23:2727–2739
Correa MA, Matusovsky B, Brackney DE, Steven B (2018) Generation of axenic Aedes aegypti demonstrate live bacteria are not required for mosquito development. Nat. Commun 9:4464
Guégan M, Zouache K, Démichel C, Minard G, Tran Van V, Potier P, Mavingui P, Valiente Moro C (2018) The mosquito holobiont: fresh insight into mosquito-microbiota interactions. Microbiome 6:49
Scolari F, Casiraghi M, Bonizzoni M (2019) Aedes spp. and their microbiota: a review. Front Microbiol 10:2036
Strand MR (2018) Composition and functional roles of the gut microbiota in mosquitoes. Curr Opin Insect Sci 28:59–65
Coon KL, Brown MR, Strand MR (2016) Mosquitoes host communities of bacteria that are essential for development but vary greatly between local habitats. Mol. Ecol 25:5806–5826
Dickson LB, Jiolle D, Minard G, Moltini-Conclois I, Volant S, Ghozlane A, Bouchier C, Ayala D, Paupy C, Moro CV, Lambrechts L (2017) Carryover effects of larval exposure to different environmental bacteria drive adult trait variation in a mosquito vector. Sci. Adv 3:e1700585
Muturi EJ, Dunlap C, Ramirez JL, Rooney AP, Kim CH (2019) Host blood-meal source has a strong impact on gut microbiota of Aedes aegypti. FEMS Microbiol. Ecol 95:213
Wang Y, Gilbreath III TM, Kukutla P, Yan G, Xu J (2011) Dynamic gut microbiome across life history of the malaria mosquito Anopholes gambiae in Kenya. PLoS One 6:e24767
Ramirez JL, Souza-Neto J, Cosme RT, Rovira J, Ortiz A, Pascale JM, Dimopoulos G (2012) Reciprocal tripartite interactions between the Aedes aegypti midgut microbiota, innate immune system and dengue virus influences vector competence. PLoS Negl. Trop. Dis 6:e1561
Charan SS, Pawar KD, Severson DW, Patole MS, Shouche YS (2013) Comparative analysis of midgut bacterial communities of Aedes aegypti mosquito strains varying in vector competence to dengue virus. Parasitol. Res 112:2627–2637
Dennison NJ, Jupatanakul N, Dimopoulos G (2014) The mosquito microbiota influences vector competence for human pathogens. Curr Opin Insect Sci 3:6–13
Jupatanakul N, Sim S, Dimopoulos G (2014) The insect microbiome modulates vector competence for arboviruses. Viruses 6:4294–4313
Hegde S, Rasgon JL, Hughes GL (2015) The microbiome modulates arbovirus transmission in mosquitoes. Curr Opin Virol 15:97–102
Molina-Henao EH, Graffe MY, Elsa P, Serrato IM, Correa A, Romero LV, Caicedo PA, Ocampo CB (2020) Culturable microbial composition in the midgut of Aedes aegypti strains with different susceptibility to dengue-2 virus infection. Symbiosis 80:85–93
Gusmão DS, Santos AV, Marini DC, Bacci M, Berbert-Molina MA, Lemos FJA (2010) Culture-dependent and culture-independent characterization of microorganisms associated with Aedes aegypti (Diptera: Culicidae) and dynamics of bacterial colonization in the midgut. Acta Trop 115:275–281
Terenius O, Lindh JM, Eriksson-Gonzales K, Bussière L, Laugen AT, Bergquist H, Titanji K, Faye I (2012) Midgut bacterial dynamics in Aedes aegypti. FEMS Microbiol. Ecol 80:556–565
David MR, Dos Santos LMB, Vicente ACP, Maciel-de-Freitas R (2016) Effects of environment, dietary regime and ageing on the dengue vector microbiota: evidence of a core microbiota throughout Aedes aegypti lifespan. Mem. Inst. Oswaldo Cruz 111:577–587
Dickson LB, Ghozlane A, Volant S, Bouchier C, Ma L, Vega-Rúa A, Dusfour I, Jiolle D, Paupy C, Mayanja MN, Kohl A, Lutwama JJ, Duong V, Lambrechts L (2018) Diverse laboratory colonies of Aedes aegypti harbor the same adult midgut bacterial microbiome. Parasites Vectors 11:207
Saab SA, Zu Dohna H, Nilsson LK, Onorati P, Nakhleh J, Terenius O, Osta MA (2020) The environment and species affect gut bacteria composition in laboratory co-cultured Anopholes gambiae and Aedes albopictus mosquitoes. Sci. Rep 10:3352
Nearing JT, Douglas GM, Comeau AM, Langille MGI (2018) Denoising the denoisers: an independent evaluation of microbiome sequence error-correction approaches. PeerJ 6:e5364
Callahan BJ, McMurdie PJ, Holmes SP (2017) Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J 11:2639–2643
Balvočiute M, Huson DH (2017) SILVA, RDP, Greengenes, NCBI and OTT - how do these taxonomies compare? BMC Genomics 18:114
Salter SJ, Cox MJ, Turek EM, Calus ST, Cookson WO, Moffatt MF, Turner P, Parkhill J, Loman NJ, Walker AW (2014) Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol 12:87
Schloss PD (2018) Identifying and overcoming threats to reproducibility, replicability, robustness, and generalizability in microbiome research. mBio 9:e00525-18
Hegde S, Khanipov K, Albayrak L, Golovko G, Pimenova M, Saldaña MA, Rojas MM, Hornett EA, Motl GC, Fredregill CL, Dennett JA, Debboun M, Fofanov Y, Hughes GL (2018) Microbiome interaction networks and community structure from laboratory-reared and field-collected Aedes aegypti, Aedes albopictus, and Culex quinquefasciatus mosquito vectors. Front. Microbiol 9:2160
Frankel-Bricker J, Buerki S, Feris KP, White MM (2020) Influences of a prolific gut fungus (Zancudomyces culisetae) on larval and adult mosquito (Aedes aegypti)-associated microbiota. Appl. Environ. Microbiol 86:e02334–e02319
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13:581–583
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J, Glöckner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35:7188–7196
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596
Anderson MJ (2006) Distance-based tests for homogeneity of multivariate dispersions. Biometrics 62:245–253
Anderson MJ, Ellingsen KE, McArdle BH (2006) Multivariate dispersion as a measure of beta diversity. Ecol. Lett 9:683–693
Mancini MV, Damiani C, Accoti A, Tallarita M, Nunzi E, Cappelli A, Bozic J, Catanzani R, Rossi P, Valzano M, Serrao A (2018) Estimating bacteria diversity in different organs of nine species of mosquito by next generation sequencing. BMC Microbiol 18:126
Segata N, Baldini F, Pompon J, Garrett WS, Truong DT, Dabiré RK, Diabaté A, Levashina EA, Catteruccia F (2016) The reproductive tracts of two malaria vectors are populated by a core microbiome and by gener-and swarm-enriched microbial biomarkers. Sci. Rep 6:24207
Sharma P, Sharma S, Maurya RK, Das De T, Thomas T, Lata S, Singh N, Pandey KC, Valecha N, Dixit R (2014) Salivary glands harbor more diverse microbial communities than gut in Anopheles culicifacies. Parasit. Vectors 7:235
Foggie T, Achee N (2009) Standard operating procedures : rearing Aedes aegypti for the HITSS and box laboratory assays training manual
Sullam KE, Pichon S, Schaer TMM, Ebert D (2017) The combined effect of temperature and host clonal line on the microbiota of a planktonic crustacean. Microb. Ecol 76:506–517
Frankel-Bricker J, Song MJ, Benner MJ, Schaack S (2020) Variation in the microbiota associated with Daphnia magna across genotypes, populations, and temperature. Microb. Ecol 79:731–742
Duguma D, Hall MW, Rugman-Jones P, Stouthamer R, Terenius O, Neufeld JD, Walton WE (2015) Developmental succession of the microbiome of Culex mosquitoes. BMC Microbiol 15:140
Bascuñán P, Niño-Garcia JP, Galeano-Castañeda Y, Serre D, Correa MM (2018) Factors shaping the gut bacterial community assembly in two main Colombian malaria vectors. Microbiome 6:148
Rodríguez-Ruano SM, Juhaňáková E, Vávra J, Nováková E (2020) Methodological insight into mosquito microbiome studies. Front. Cell. Infect. Microbiol 10:86
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. PNAS 108:4516–4522
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1
Core Team R (2020) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria https://www.R-project.org/
Schliep KP (2011) Phangorn: phylogenetic analysis in R. Bioinformatics 27:592–593
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217
Davis NM, Proctor D, Holmes SP, Relman DA, Callahan BJ (2018) Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome 6:226
Wickham H (2011) Ggplot2. Wiley Interdiscip Rev Comput Stat 3:180–185
Bates D, Maechler M, Bolker B, Walker S, Maechler M, Walker S (2015) Package “lme4”: linear mixed-effects models using “Eigen” and S4. J. Stat. Softw https://CRAN.R-project.org/package=lme4 67
Fox J, Weisberg S, Price B, Adler D, Bates D, Baud-Bovy G (2018) Package “car”. https://CRAN.R-project.org/package=car
Hothorn T, Bretz F, Westfall P, Heiberger RM (2016) Package “multcomp”: simultaneous inference in general parametric models. Project for Statistical Computing, Vienna, Austria
Clarke KR (1993) Non-parametric multivariate analyses of changes in community structure. Aust. J. Ecol 18:117–143
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol. Electron 4:9
Anderson MJ (2017) Permutational multivariate analysis of variance (PERMANOVA). Wiley StatsRef, Statistics Reference Online, pp 1–15
Dixon P (2003) VEGAN, a package of R functions for community ecology. J. Veg. Sci 14:927–930
Anderson MJ, Walsh DCI (2013) PERMANOVA, ANOSIM, and the mantel test in the face of heterogeneous dispersions: what null hypothesis are you testing? Ecol. Monogr 83:557–574
Acknowledgements
We thank Michael Wojahn for collaboration and assistance regarding construction of the phylogenetic tree used in phylogenetic-based analyses. We also thank Trevor Caughlin for providing insights on community analyses. We thank 3 anonymous reviewers and the handling editor for giving constructive critiques and suggested revisions that greatly improved the quality of the manuscript. Finally, we thank the research groups performing significant research analyzing mosquito-associated microbiomes that produced the high-quality data sets used in this analysis.
Code Availability
Reproducible code used for this analysis is provided in supplemental File S1.
Author information
Authors and Affiliations
Contributions
JFB conceived the project, performed all analyses, and was the primary author of the manuscript. LKF was a major contributor in writing and revising the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
The publisher has the authors’ permission to publish the content presented herein.
Competing Interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Frankel-Bricker, J., Frankel, L.K. Re-Analysis of 16S rRNA Gene Sequence Data Sets Uncovers Disparate Laboratory-Specific Microbiomes Associated with the Yellow Fever Mosquito (Aedes aegypti). Microb Ecol 83, 167–181 (2022). https://doi.org/10.1007/s00248-021-01739-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-021-01739-2