Abstract
Honeybees are important pollinators, having an essential role in the ecology of natural and agricultural environments. Honeybee colony losses episodes reported worldwide and have been associated with different pests and pathogens, pesticide exposure, and nutritional stress. This nutritional stress is related to the increase in monoculture areas which leads to a reduction of pollen availability and diversity. In this study, we examined whether nutritional stress affects honeybee gut microbiota, bee immunity, and infection by Nosema ceranae, under laboratory conditions. Consumption of Eucalyptus grandis pollen was used as a nutritionally poor-quality diet to study nutritional stress, in contraposition to the consumption of polyfloral pollen. Honeybees feed with Eucalyptus grandis pollen showed a lower abundance of Lactobacillus mellifer and Lactobacillus apis (Firm-4 and Firm-5, respectively) and Bifidobacterium spp. and a higher abundance of Bartonella apis, than honeybees fed with polyfloral pollen. Besides the impact of nutritional stress on honeybee microbiota, it also decreased the expression levels of vitellogenin and genes associated to immunity (glucose oxidase, hymenoptaecin and lysozyme). Finally, Eucalyptus grandis pollen favored the multiplication of Nosema ceranae. These results show that nutritional stress impacts the honeybee gut microbiota, having consequences on honeybee immunity and pathogen development. Those results may be useful to understand the influence of modern agriculture on honeybee health.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Western honeybees Apis mellifera are important managed pollinators worldwide, having an essential role in the ecology of natural environments and agricultural production [1, 2]. In recent years, large-scale colony losses have been reported in different countries [3,4,5,6]. These losses have been associated with the infection by multiple pests and pathogens (such as the mite Varroa destructor, the microsporidia Nosema ceranae, and different RNA viruses), intoxication with pesticides, and nutritional stress [7,8,9]. Nutritional stress is linked to the increase of monoculture areas which leads to a reduction of pollen availability and diversity for honeybees [7, 9].
Adult honeybee nutrition relies on the intake of honey and pollen. Pollen provides proteins, lipids, and vitamins necessary for the healthy growth and development [reviewed at 10, 11]. Its composition, quality, or quantity influences honeybee physiology (protein concentrations, macromolecule metabolism, hypopharyngeal glands size, tissue growth and development, glutathione S-transferase activity, immunocompetence) [12,13,14,15], longevity [16], and resistance to pathogens including Nosema apis and Nosema ceranae [17,18,19].
Previous studies have shown that honeybee colonies subjected to nutritional stress, located in a Eucalyptus grandis plantation during the flowering period in autumn, became rapidly infected by N. ceranae, decreasing brood and adult population [20, 21] and undergoing high colony losses [21, 22]. These losses may be related with the poor-nutritional quality of Eucalyptus spp. pollen, since protein percentage decreases during the flowering period to values under 20% [20, 21], they have low content of lipids [20, 23] and essential fatty acid omega 3 [24], and they have deficiency in the essential amino acid isoleucine [25, 26].
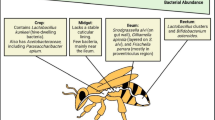
Besides that, it has been proposed that environmental landscape affects the honeybee gut microbiota, influencing the relative abundance of some of its members [27]. This microbiota is dominated by eight core bacterial clusters, which comprise 95–99% of bacteria in the gut [28,29,30,31], including Gilliamella apicola [32], Snodgrasssella alvi [32], Lactobacillus Firm-4 [33], Lactobacillus Firm-5 [33], Bifidobacterium spp. [33], Frischella perrara [34], Bartonella apis [35], and Parasaccharibacter apium [36]. It has been involved in the defense against pathogens, metabolism, growth, development, and immunity [30]. Disruption of the microbiota, named dysbiosis, may have consequences on the bee development and immunity, affecting the ability of bees to respond to environmental stressors [30].
The honeybee immune system is composed of a complex network of mechanisms, including cellular and humoral immune defenses reviewed in [37]. In particular, humoral defenses rely on the production of antimicrobial peptides (apidaecins, abaecin, hymenoptaecin, and defensins), which act generating leaks in prokaryotic membranes; and inhibiting bacterial protein translation or folding [38]. Besides that, honeybees have evolved a social immunity, a collective defense that arises from the behavioral cooperation among individuals, such as grooming, hygienic behavior or food sterilization [37, 39]. Doublet et al. [40], identified a common set of genes that respond to the infection by different pathogens (V. destructor, Nosema apis, N. ceranae, and RNA viruses), including hymenoptaecin, defensin, and abaecin, among others.
We hypothesize that nutritional stress shapes the composition of the honeybee gut microbiota having consequences on honeybee immunity and favoring the infection by pathogens. Therefore, we evaluated the effect of two different pollen diets (E. grandis or polyfloral pollen) in the composition of the gut microbiota, on the expression of different genes involved in the humoral and cellular immune response, and in the infection by N. ceranae.
Methods
Pollen Samples
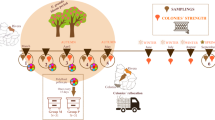
E. grandis monofloral stored pollen samples (bee bread) were manually collected from frames of colonies located in E. grandis plantations during its flowering period (Rivera, Uruguay), in April and May (autumn) 2014, for trial 1 or 2, respectively. Polyfloral stored pollen (bee bread) was manually collected from colonies located in the experimental apiary at the Instituto Nacional de Investigación Agropecuaria (INIA “La Estanzuela”, Colonia, Uruguay) in April (autumn) and October (spring), 2014 for trials 1 and 2, respectively. In all cases, one frame per colony, from five colonies, were collected and mixed. Pollen samples were homogenized making a paste, and no water or syrup was added. Samples were kept at −20 °C until used or analyzed. Palynological analyses were performed to validate their botanical composition [41]. At least 1200 pollen grains per sample were identified [42]. Although there is a low risk of pesticide pollen contamination, we cannot completely rule out this possibility.
Two independent trials were performed in 2014 (trials 1 and 2), and different pollen samples were used in each trial (Fig. 1A suppl. data).
Trial 1
In spring, three healthy A. mellifera colonies, local hybrids between A. m. scutellata, A. m. ligustica, and A. m. mellifera, were randomly selected from the experimental apiary at INIA “La Estanzuela”, Colonia, Uruguay. Colonies had been treated against the mite V. destructor in the previous autumn (with Amitraz in stripes) and did not show clinical symptoms of diseases.
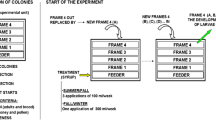
One frame of brood containing last-stage pupae was removed from each hive, taken to the laboratory, and incubated at 34 ± 1 °C, 60% relative humidity in the darkness until honeybee emergence, as recommended by Williams et al. [43]. Immediately after emergence, bees from different frames were collected. Only bees emerged within the first 24 h were used, in order to standardize the age. After that time, bees from different colonies were mixed, placed in cages, and incubated at 30 ± 1 °C, 60% relative humidity in the darkness. Six cages of bees (n = 40 per cage) were fed with E. grandis stored pollen and other six with polyfloral stored pollen. About 3 g of pollen was served per cage in a sterile plastic recipient, and it was replaced every 48 h. A sucrose solution was also provided to bees in Pasteur pipettes (ad libitum, 50% w/w in water). Every day, bees were monitored, dead individuals were counted and removed, and sucrose solution was replaced (Fig. 1B suppl. data).
Trial 2
An independent second trial was performed using three healthy colonies from the experimental apiary of the Universidad Nacional de Mar del Plata (Mar del Plata, Argentina), local hybrids between A. m. ligustica and A. m. mellifera. As in trial 1, recommendations by Williams et al. [43] were followed. Six cages of bees (n = 80 per cage) were fed with E. grandis stored pollen while other six were fed with polyfloral pollen, ad libitum. About 3 g of pollen was served per cage in a sterile plastic recipient, and it was replaced every 48 h. Three days post-emergence, honeybees from three cages per nutritional regimen were individually infected with 100,000 spores of N. ceranae in 10 μl of sucrose solution [44] and returned to the same cage with the same diet. Honeybees from the other cages were individually fed with the same volume of sugar syrup, as controls. Every day, bees were monitored, dead individuals were removed, pollen consumption was estimated (by weight), and sucrose solution was replaced (Fig. 1C suppl. data).
Nosema ceranae spores used for infection were obtained from foraging honeybees from a naturally infected colony and purified by the triangulation method [45]. The spore suspension was quantified using a hemocytometer [46] and immediately used for experimental infection (Fig. 1C. suppl. data). The species identification was carried out by multiplex PCR according to Martin-Hernández et al. [47] (Table 1 suppl. data).
Microbiota Analyses
DNA Extraction from Pollen
Three replicates of E. grandis and polyfloral pollen samples (0.2 g/sample) used in trials 1 and 2 were processed according to Anderson et al. [48]. DNA was quantified using a NanoDrop1000 spectrophotometer (Thermo Scientific™) and concentrations were normalized to 10 ng/μl.
DNA Extraction from Honeybees
Ten days after emergence, 20 honeybees/cage from 3 cages/treatment from trials 1 and 2 were collected. Honeybees were externally sterilized using a chlorine solution 1% [48]. Guts were extracted, pooled (n = 20/cage), and homogenized using a glass rod and a plastic tube. DNA was extracted using the SDS-CTAB method [49], quantified using a NanoDrop1000 spectrophotometer (Thermo Scientific™), and concentrations were normalized to 20 ng/μl.
PCR for Detection of Pathogens
DNAs obtained from stored pollen and honeybees were subjected to PCR for the detection of N. ceranae, N. apis, and Lotmaria passim, as previously described [47, 50] (Table 1 suppl. data). PCR reactions were carried out using a MultiGeneOptiMax Thermal Cycler (Labnet International, USA). Positive controls (using DNA from N. ceranae, N. apis, or L. passim) and negative controls were included. Amplified products were analyzed by electrophoresis in a 2% agarose gel in Tris/borate/EDTA (TBE) at 120 V for 30 min, stained with GelRed (Biotum, USA), and visualized by UV light.
Quantitative PCR
The quantity of 16S rRNA genes (bacterial load) in honeybees was determined through ratio calculations using universal 16S rRNA gene [51] and a honeybee gene (RPS5 gene [52], Table 1 suppl. data). To quantify the total bacterial number in pollen, a standard curve from DNA obtained from an Escherichia coli XL1 Blue culture was used. Cycle threshold (Ct) values of each sample were then compared to the standard curve to approximate the number of bacteria per sample, according to Ott et al. [53]. In both cases, PCR were carried out in a final volume of 20 μl, comprising 1× SYBR (Power SYBR ® Green PCR Master Mix, Applied Biosystems), 0.3 μM of each primer, 40 ng of DNA and RNAse-free water, and PCR cycling described before. All qPCR reactions were performed in triplicate, in a BIO-RAD CFX96™ Real-Time system, and two negative controls were included in each run.
16S rRNA Amplicon Sequencing
DNA obtained from stored pollen and honeybees was analyzed by sequencing of V4 region of 16S rRNA gene using an Illumina MiSeq platform and 250 paired-end (PE) cycles (University of Texas at Austin, USA).
Immune Gene Expression Analyses
RNA Extraction and cDNA Synthesis
Ten days after emergence, ten honeybees per nutritional regimen from trial 1 were collected and stored at −80 °C. Individual honeybees were homogenized in RLT buffer (Qiagen) and subjected to RNA extraction using the RNAeasy Plus minikit (Qiagen), according to the manufacturer’s instructions. One microliter of total RNA was treated with DNAse and used to generate first-strand cDNA using the Quantitec Reverse Transcription kit, also according to the manufacturer’s instructions (Qiagen).
Quantitative PCR
Relative expression of abaecin, defensin, lysozyme, glucose dehydrogenase, glucose oxidase, hymenoptaecin, and vitellogenin genes were assessed using previously reported primers. Ribosomal protein S5 (RPS5) and ß-actin were used as housekeeping genes to normalize the variation of cDNA levels ([52, 54, 55]; Table 1 suppl. data). The reaction mix consisted of 1X QuantiTect SYBR Green PCR MasterMix (Qiagen), 0.5 μM of each primer, RNAse-free water and 5 μl of 1:10 diluted cDNA in a final volume of 25 μl. PCR reactions were carried out using a BIO-RAD CFX96™ Real-Time system and the cycling program consisted of an initial 95 °C for 15 min, and 40 cycles of three-step PCR at 94 °C for 15 s, 52 °C for 30 s and 72 °C for 30 s. Specificity of the reaction was checked by analysis of the melting curve of the final amplified product (from 65 to 95 °C, with increments of 0.5 °C every 0.05 s.).
Fluorescence was measured in the elongation step and negative controls (without DNA) were included in each reaction run.
Quantification of N. ceranae Spores
At 4, 7, and 12 days post infection, five honeybees per cage from trial 2 were sampled. Midguts were removed, and spore number/honeybees (intensity of parasite infection) were quantified by using a hemocytometer (Neubauer improved) [46].
Data Analysis
Honeybee Survival
The impact of nutrition and infection by N. ceranae on honeybee survival was analyzed using the Kaplan-Meier method, and statistical differences were compared using the Log-rank test.
16S rRNA Amplicon Sequencing
For analysis of bacterial community data, the protocol suggested by Engel et al. [56] was followed. Paired-end reads were joined using the fastq-join method and demultiplexed with QIIME software package (Qiime.org) [57]. Sequences were analyzed using the QIIME software tool with default parameters for each step. Reads were screened for chimeras using the software program USEARCH 6.1. De novo operational taxonomic unit (OTU) picking was performed with the uclust option in QIIME [58]. Assignment of taxonomy to representative OTUs was carried out with the Greengenes database classifier [59] at the default 97% sequence identity. Sequences matching plant chloroplast or mitochondrial 16S rRNA were filtered from the dataset. The optimal sampling depth was determined through examination of exploratory rarefaction curves of observed OTUs plotted against sampling depth, and the dataset was rarefied.
To evaluate the impact of nutrition/Nosema spp. infection on the composition of the gut microbiota, dissimilarity matrices Unifrac weighted (by OTUs abundance), Unifrac unweighted (presence/absence of OTUs), and Bray-Curtis were built. Matrices were used to produce classical multidimensional scaling (principal coordinates analysis). Statistical differences between groups were tested using PERMANOVA (using 1000 permutations). The Shannon diversity index was also calculated, and differences between groups were assessed using the Student t test or Mann-Whitney Test.
Besides that, differences between abundance of the different OTUs were examined using the DESeq2 software [60], as described by Jones et al. [27]. Statistical analyses were performed using R software [61].
Gene Expression Analyses
The geometric mean of the Ct (threshold cycle number) of reference genes (RPS5 and β-actin) was calculated and used for normalization. The expression ratio was analyzed as described by Pfaffl [62]. The data corresponding to each gene were analyzed to determine if they fitted a normal distribution (Kolmogorov-Smirnov test) and whether there was a homogeneous variance (Levene test).
Gene transcript levels, bacterial abundance, and N. ceranae infection between different groups were evaluated by Student t test in those variables which fit parametric assumptions, and by the Mann-Whitney test when the variables did not. P values below 0.05 were considered significant. Statistical analyses were performed using Past 3x version 2.17c [63, 64].
Results
Pollen Diets
In order to evaluate the impact of nutritional stress on honeybees, newly emerged individuals were fed with two different diets: E. grandis monofloral stored pollen and polyfloral stored pollen (bee bread) (Fig. 1 suppl. data). Two independent trials were performed; Eucalyptus grandis stored pollen samples used in both trials had a purity of 99%, while polyfloral stored pollen samples were composed by 18 or 23 different species (trial 1 and 2, respectively; Table 2 suppl. Data).
Both pollen types showed similar bacterial loads and composition, although subtle differences were found (Table 3 and Figs. 2 and 3, suppl. data). E. grandis pollen showed a lower OTU diversity than polyfloral pollen, according to rarefaction curves (Fig. 4 suppl. data) and Shannon diversity index (t-test, t = −15.65; t = −8.28, for trials 1 and 2, respectively, p < 0.01 in both cases).
According to DESeq2, abundance of Lactobacillus Firm-4 and Firm-5 was similar in both pollen samples in trial 1 and trial 2, and Bifidobacterium spp. and Bartonella apis were not detected.
Besides that, N. apis, N. ceranae, and L. passim were not detected in any pollen sample (data not shown).
Those results indicate that both groups of honeybees received pollen with different botanical species compositions and subtle differences on associated microbiota and absence (or undetectable level) of the studied pathogens were observed.
Impact of Nutritional Stress on Honeybee Survival and Pollen Consumption
Honeybees fed on E. grandis and polyfloral pollen survived almost 60 days under laboratory conditions, and survival curves were similar in both groups. No difference was observed after 30 (Log-rank test; Statistic = 0.02; p = 0.88, Fig. 5 suppl. data) or 60 days (Log-rank test; Statistic = 5.56; p = 0.35).
Impact of Nutritional Stress on the Gut Microbiota
The impact of different diets on the honeybee gut microbiota was assessed through the analysis of the bacterial community size and community composition, by 16S rRNA gene amplicon sequencing. No significant changes in the bacterial community size were found in the gut microbiota honeybees fed with E. grandis monofloral stored pollen and polyfloral stored pollen (Table 4 suppl. data). Regarding OTUs diversity, in trial 1, polyfloral pollen intake increased the diversity according to rarefaction curves and Shannon diversity index, meanwhile in trial 2, no significant difference was observed (MW test, U = 0.00 p = 0.05; U = 0.00 p = 0.14 for trials 1 and 2 (control bees), respectively (Fig. 1; Fig. 6 suppl. data).
Relative abundance of bacterial genera and Shannon diversity index of gut bacterial community of honeybees analyzed through 16S rRNA amplicon sequencing (Illumina MySeq). a Trial 1, honeybees were fed with E. grandis (Eg) or polyfloral (PF) stored pollen. A single MiSeq PE run of 250 cycles resulted in 232,345 reads, of which 221,576 (95%) passed stringent quality thresholds. The data set was rarefied to 29,372 sequences, and after alignment and clustering, we identified a total of 221,560 unique sequences and 159 OTUs (at 97% identity) across the entire data set. b Trial 2, honeybees were fed with E. grandis or polyfloral stored pollen and infected with Nosema spp. spores. In total, 712,706 reads were obtained of which 712,600 passed the quality filters. Subsequently, the data set was normalized to 23,399 sequences per sample, with remaining 112 OTUs
The gut microbiota was composed by Lactobacillus Firm-4 and Firm-5, Bifidobacterium spp., Snodgrassella alvi, Pasteurellales/Orbales, Bartonella apis, and Parasaccaribacter apium.
The diet did not alter the bacterial general composition of the gut microbiota (trial 1, Unifrac weighted analysis PERMANOVA test statistic = 5.22, p = 0.09; Unifrac unweighted analysis PERMANOVA test statistic = 2.21, p = 0.10; Bray-Curtis PERMANOVA analysis test statistic = 4.57, p = 0.11; non-infected honeybees (control) from trial 2, Unifrac weighted analysis PERMANOVA test statistic = 3.44, p = 0.1; Unifrac unweighted analysis PERMANOVA test statistic = 2.08, p = 0.1; Bray-Curtis PERMANOVA analysis test statistic = 2.24, p = 0.1; Fig. 2).
However, significant differences at OTU level were observed according to DESeq2 results. Consumption of E. grandis pollen (model of nutritional stress) decreased the abundance of Lactobacillus mellifer and L. apis (Lactobacillus Firm-4 and Firm-5) compared with polyfloral pollen in trial 1 (Fig. 3). E. grandis pollen also decreased the abundance of Bifidobacterium asteroids and Bifidobacterium corineforme compared to polyfloral pollen, in trial 2 (non-infected honeybees). On the other hand, E. grandis pollen increased the abundance of Bartonella apis in both trials.
Comparison of abundance of different OTUs between honeybees fed with E. grandis and polyfloral stored pollen, by using DESeq2 analysis. A log2-fold change of >0 indicates that abundance was higher in honeybees fed on E. grandis pollen. a Samples from trial 1; b samples from non-infected honeybees (control) from trial 2; c samples from N. ceranae-infected honeybees from trial 2. OTUs with less than 60 reads in total were not considered
Impact of Nutritional Stress on Immune Gene Expression
Diets altered the expression of genes involved in honeybee physiology, according to quantitative PCR results (Fig. 4). E. grandis pollen consumption decreased the expression level of vitellogenin, glucose oxidase, hymenoptaecin, and lysozyme (MW test, U = 13 p = 0.03; U = 6 p = 0.05, U = 2 p = 0.02, and U = 11 p = 0.01; respectively), in comparison to polyfloral pollen. On the other hand, expression of abaecin, defensin, and glucose dehydrogenase genes was not affected (p ≥ 0.05 in all cases).
Relative expression of glucose oxidase, hymenoptaecin, lysozyme, and vitellogenin genes in honeybees fed with E. grandis (Eg) or polyfloral (PF) stored pollen. Quantification of gene expression was carried by qPCR. Ten individual honeybees per treatment were used. Asterisks indicate significant differences (p values under 0.05)
Impact of Nutritional Stress on N. ceranae and the Gut Microbiota
To evaluate the impact of nutritional stress on pathogen infections, honeybees fed on different diets were infected with N. ceranae spores. A higher level of N. ceranae spores was observed in honeybees fed with E. grandis pollen compared to bees fed on polyfloral (4 days post-infection 5.9 × 104 ± 1.2 × 104 and 2.4 × 104 ± 8.6 × 103 spores/bee, t-test t = −4.01 p = 0.02; 7 days post-infection 2.5 × 106 ± 9.8 × 105 and 6.2 × 105 ± 4.95 × 105 spores/bee, t-test t = −3.02 p = 0.04, Fig. 5). Twelve days after infection, the number of spores was similar in both groups, reaching 2.6 × 107 ± 2.9 × 106 and 2.3 × 107 ± 2.2 × 106 spores/bee in honeybees fed with E. grandis or polyfloral pollen, respectively. No Nosema spp. spores were detected in the honeybees that belonged to the control groups (non-infected honeybees).
Nosema ceranae infection together with diet generated significant alterations on the composition of the gut microbiota (Figs. 1 and 2, Unifrac weighted analysis PERMANOVA test statistic = 4.99, p = 0.001; Unifrac unweighted analysis PERMANOVA test statistic = 2.56, p = 0.004; Bray-Curtis PERMANOVA analysis test statistic = 5.50, p = 0.001).
Infection increased the abundance of two OTUs of Acetobacteriaceae in honeybees fed on polyfloral pollen (Fig. 6). However, in the case of honeybees fed on E. grandis pollen, the impact was higher since the abundance of five different OTUs increased (Acetobacteraceae, Enterobacteraceae, Bifidobacterium spp., Bartonella apis, and G. apicola).
Comparison of abundance of different OTUs between honeybees fed with E. grandis (a) or polyfloral stored pollen (b) and infected by N. ceranae, by using DESeq2 analysis. A log2-fold change of >0 indicates that abundance was higher in infected honeybees. OTUs with less than 60 reads in total were not considered
Interestingly, N. ceranae-infected honeybees fed with E. grandis pollen showed a lower abundance of Lactobacillus helsingborgensis (Lactobacillus Firm-5) and a higher abundance of Bartonella apis than infected honeybees fed on polyfloral pollen (Fig. 3).
Discussion
Nutritional stress has been proposed as an important driver of honeybee colony losses [7, 9]. In a previous study, we demonstrated that under field conditions, nutritional stress (feeding of colonies mainly on E. grandis pollen) promoted the reproduction of Nosema spp. and weakened the colonies, decreasing the adult honeybee and brood populations [20]. However, the mechanisms underlying this interaction remain elusive.
In this study, we show that nutritional stress alters the composition of the honeybee gut microbiota and immunity, favoring the infection by N. ceranae.
Eucalyptus grandis pollen was used as a model to study nutritional stress, since it has a low lipid content, the protein percentage decreases during the flowering period to values under 20% and it is deficient in isoleucine [20, 21].
Two independent trials using newly emerged honeybees under controlled laboratory conditions were performed. Although the natural establishment of the microbiota can take about 4 days after emergence [63, 64], those bees were able to develop a typical microbiota [28,29,30,31]. Bacteria could be acquired by direct contact with the frames after emergence or with pollen ingestion [63, 64].
Nutritional stress (consumption of E. grandis pollen) induced the decrease of the abundance of Lactobacillus mellifer, Lactobacillus apis, or Bifidobacterium spp. and increased the abundance of Bartonella apis, in healthy bees. Those results were consistent when the microbiota of N. ceranae-infected honeybees was assessed.
Pollen bacteria did not explain the differential abundance of these bacterial species in the honeybee gut, since species of Lactobacillus Firm-4 and Firm-5 abundances were similar in both pollens from each trial, and Bifidobacterium spp. and Bartonella apis were not detected.
Analysis of complete genomes of Lactobacillus spp. revealed that these bacteria possess numerous phosphotransferase systems involved in the uptake of sugars, while Bifidobacterium spp. has abundant genes for carbohydrate utilization [33, 65, 66]. Both genera are all able to utilize glucose and fructose, the most abundant sugars in the honeybee diet. These bacteria have large putative surface proteins which may be related to adhesion or degradation of plant compounds, and gene clusters for biosynthesis and utilization of trehalose, a disaccharide used for energy storage in insects [33, 65, 66]. Besides that, it has been proposed that Lactobacillus spp. and Bifidobacterium spp. have in vitro inhibitory effect against P. larvae and Melissococcus plutonius, causative agents of American and European foulbrood, respectively [67,68,69].
Bartonellaceae is a bacterial family that includes facultative intracellular pathogens present in a variety of mammals, usually acquired by vector transmission or by animal bites or scratches [70, 71]. Hubert et al. [72] showed that V. destructor could act as a reservoir of these bacteria, but their significance in the bee microbiota is still unknown.
Those changes in the honeybee gut microbiota in bees subjected to nutritional stress may represent an important disadvantage regarding the bee physiology and defense against pathogens.
Although under laboratory conditions consumption of E. grandis pollen did not affect bee lifespan, it decreased the expression level of vitellogenin, compared to polyfloral pollen. Vitellogenin has an important role in the division of labor, foraging specialization, queen longevity, and resistance to oxidative stress [55, 73,74,75]. A low vitellogenin level is associated with precocious foraging, low protection against oxidative stress and a shorter lifespan [74]. Previous studies have also evidenced that vitellogenin expression changes in response to diet, suggesting it could be an interesting marker of the honeybee nutritional status on healthy honeybees [13, 76,77,78]. The low vitellogenin expression level found on honeybees fed on E. grandis pollen confirmed that honeybees are under nutritional stress.
Eucalytpus grandis pollen consumption also generated a decrease in the glucose oxidase expression, in accordance with Alaux et al. [15]. This enzyme catalyzes the oxidation of β-d-glucose to gluconic acid and hydrogen peroxide, which has antiseptic properties. Those products reach the larval food and honey, contributing to food sterilization and prevention of contamination with pathogenic microorganisms [39]. For this reason, glucose oxidase activity is considered as a marker of social immunity [15].
Furthermore, in this study, we confirmed that nutritional stress also decreased the expression of genes involved in individual immunity (hymenoptaecin and lysozyme). Hymenoptaecin is an antimicrobial peptide [79] and lysozyme is a non-specific immunity factor that hydrolyzes (1, 4)-glycosidic bonds in the peptidoglycan layer of the bacterial cell wall [80]. Lysozyme also promotes the expression of antimicrobial peptides, optimizing the immune response [81]. The decrease in the expression of those peptides in honeybees under nutritional stress may be associated with a diminished defensive ability against pathogens.
Finally, nutritional stress promoted the multiplication of N. ceranae, during the first stages of infection. Although previous studies had proposed that rich pollen diets stimulated the development of Nosema spp. [18, 19, 82], comparisons were performed between honeybees feed with sugar syrup and honeybees fed with different amounts of pollen, but not using honeybees fed with pollen with different botanical composition. In this case, nutritional stress may accelerate N. ceranae reproduction, although at the end of the experiment all bees reached were infected by 2 × 107 spores/bee.
The increase of N. ceranae spores in honeybees under nutritional stress may be associated with an alteration of the honeybee gut microbiota and a depression of the immune system, which might accelerate the microsporidium multiplication.
In this regard, previous studies have reported that dietary supplementation with Lactobacillus spp. or Bifidobacterium spp. (or their metabolites) can reduce the infection level of this microsporidium [82,83,84].
Infection by N. ceranae seems to generate a stronger impact on the gut microbiota of nutritional stressed honeybees, compared to honeybees fed on polyfloral pollen. In particular, N. ceranae infection increased the abundance of five different OTUs, including G. apicola. The association between G. apicola and N. ceranae has previously been reported by Rubanov et al. [85], and further studies should be carried out to explain this interaction.
Results obtained in the present study contribute to the understanding of the influence of agriculture intensification on honeybee colony health. Vast monoculture areas might cause honeybee’s nutritional stress, decreasing the abundance of potentially beneficial microorganisms of the honeybee gut microbiota, decreasing the expression of honeybee immune-related genes, and favoring the multiplication of pathogens like N. ceranae. Besides that, N. ceranae infection can also depress the honeybee immune system [86], subtly alter the honeybee gut microbiota [85], cause energetic stress [87] and digestive problems affecting the nutritional status of honeybees [47].
Those results evidence the existence of a complex network between nutrition, gut microbiota, immunity, and pathogen infection. These links, which have been profusely described in other animal species and humans, encourage the design of strategies for the improvement of honeybee health through nutritional approaches or modulation of the gut microbiota using beneficial microbes.
References
Klein AM, Vaissiere BE, Cane JH, Steffan-Dewenter I, Cunningham SA, Kremen C, Tscharntke T (2007) Importance of pollinators in changing landscapes for world crops. Proc R Soc B Biol Sci 274:303–313. https://doi.org/10.1098/rspb.2006.3721
Potts SG, Imperatriz-Fonseca V, Ngo HT, Aizen MA, Biesmeijer JC, Breeze TD, Dicks LV, Garibaldi LA, Hill R, Settele J, Vanbergen AJ (2016) Safeguarding pollinators and their values to human well-being. Nature 540:220–229. https://doi.org/10.1038/nature20588
Neumann P, Carreck NL (2010) Honey bee colony losses. J Apic Res 49:1–6. https://doi.org/10.3896/IBRA.1.49.1.01
Kulhanek K, Steinhauer N, Rennich K, Caron DM, Sagili RR, Pettis JS, Ellis JD, Wilson ME, Wilkes JT, Tarpy DR, Rose R, Lee K, Rangel J, vanEngelsdorp D (2017) A national survey of managed honey bee 2015-2016 annual colony losses in the USA. J Apic Res 56:328–340. https://doi.org/10.1080/00218839.2017.1344496
Gray A, Brodschneider R, Adjlane N, Ballis A, Brusbardis V, Charrière J-D, Chlebo R, Coffey M F, Cornelissen B, Amaro da Costa C, Csáki T, Dahle B, Danihlík J, Maja Dražić M, Evans G, Fedoriak M, Forsythe I, de Graaf D, Gregorc A, Johannesen J, Kauko L, Kristiansen P, Martikkala M, Martín-Hernández R, Medina-Flores C A, Mutinelli F, Patalano S, Petrov P, Raudmets A, Ryzhikov VA, Simon-Delso N, Stevanovic J, Topolska G, Uzunov A, Vejsnaes F, Williams A, Zammit-Mangion M, Soroker V (2019) Loss rates of honey bee colonies during winter 2017/18 in 36 countries participating in the COLOSS survey, including effects of forage sources, J Apic Res, 58:4, 479–485. https://doi.org/10.1080/00218839.2019.1615661
Requier F, Antúnez K, Morales CL, Aldea Sánchez P, Castilhos D, Garrido M, Giacobino A, Reynaldi FJ, Rosso Londoño JM, Santos E, Garibaldi LA (2018) Trends in beekeeping and honey bee colony losses in Latin America. J Apic Res 57:657–662. https://doi.org/10.1080/00218839.2018.1494919
Goulson D, Nicholls E, Botias C, Rotheray EL (2015) Bee declines driven by combined stress from parasites, pesticides, and lack of flowers. Science 347. https://doi.org/10.1126/science.1255957
Steinhauer N, Kulhanek K, Antúnez K, Human H, Chantawannakul P, Chauzat MP, vanEngelsdorp D (2018) Drivers of colony losses. Curr Opin Insect Sci 26:142–148. https://doi.org/10.1016/j.cois.2018.02.004
Naug D (2009) Nutritional stress due to habitat loss may explain recent honeybee colony collapses. Biol Conserv 142:2369–2372. https://doi.org/10.1016/j.biocon.2009.04.007
Keller I, Fluri P, Imdorf A (2005) Pollen nutrition and colony development in honey bees: part I. Bee World 86:3–10. https://doi.org/10.1080/0005772X.2005.11099641
Brodschneider R, Crailsheim K (2010) Nutrition and health in honey bees. Apidologie 41:278–294. https://doi.org/10.1051/apido/2010012
DeGrandi-Hoffman G, Chen Y, Huang E, Huang MH (2010) The effect of diet on protein concentration, hypopharyngeal gland development and virus load in worker honey bees (Apis mellifera L.). J Insect Physiol 56:1184–1191. https://doi.org/10.1016/j.jinsphys.2010.03.017
Alaux C, Dantec C, Parrinello H, Le Conte Y (2011) Nutrigenomics in honey bees: digital gene expression analysis of pollen's nutritive effects on healthy and varroa-parasitized bees. BMC Genomics 12:496. https://doi.org/10.1186/1471-2164-12-496
Di Pasquale G, Salignon M, Le Conte Y, Belzunces LP, Decourtye A, Kretzschmar A, Suchail S, Brunet JL, Alaux C (2013) Influence of pollen nutrition on honey bee health: do pollen quality and diversity matter? PLoS ONE 8:e72016. https://doi.org/10.1371/journal.pone.0072016
Alaux C, Ducloz F, Crauser D, Le Conte Y (2010) Diet effects on honeybee immunocompetence. Biol Lett 6:562–565. https://doi.org/10.1098/rsbl.2009.0986
Schmidt JO, Thoenes SC, Levin MD (1987) Survival of honey bees, Apis mellifera (Hymenoptera: Apidae), fed various pollen sources. J Econ Entomol 80:176–183. https://doi.org/10.1093/aesa/80.2.176
Rinderer TE, Elliott KD (1977) Worker honey bee response to infection with Nosema apis. J Econ Entomol 70:431–433. https://doi.org/10.1093/jee/70.4.431
Porrini MP, Sarlo E, Medici SM, Garrido PM, Porrini DP, Damiani N, Eguaras MJ (2011) Nosema ceranae development in Apis mellifera: influence of diet and infective inoculums. J Apic Res 50:35–41. https://doi.org/10.3896/IBRA.1.50.1.04
Basualdo M, Barragán S, Antúnez K (2014) Bee bread increases honeybee haemolymph protein and promote better survival despite of causing higher Nosema ceranae abundance in honeybees. Environ Microbiol Rep 6:396–400. https://doi.org/10.1111/1758-2229.12169
Branchiccela B, Castelli L, Corona M, Díaz-Cetti S, Invernizzi C, de la Escalera MG, Mendoza Y, Santos E, Silva C, Zunino P, Antúnez K (2019) Impact of nutritional stress on the honeybee colony health. Sci Rep 12:9:10156. https://doi.org/10.1038/s41598-019-46453-9
Invernizzi C, Santos E, García E, Daners G, Di Landro R, Saadoun A, Cabrera C (2011) Sanitary and nutritional characterization of honeybee colonies in Eucalyptus grandis plantations. Arch zootec 60:1303–1314. https://doi.org/10.4321/S0004-05922011000400045
Mendoza Y, Díaz S, Ramallo G, Invernizzi C (2012) Incidence of Nosema ceranae during winter in honey bees colonies removed from Eucaliptus grandis plantations. Veterinaria 48(188):13–19
Roulston TH, Cane JH (2000) Pollen nutritional content and digestibility for animals. Plant Syst Evol 222:187–209. https://doi.org/10.1007/BF00984102
Arien Y, Dag A, Zarchin S, Masci T, Shafira S (2015) Omega-3 deficiency impairs honey bee learning. Proc Natl Acad Sci of the U S A 51:15761–15766. https://doi.org/10.1073/pnas.1517375112
Groot D (1953) Protein and amino acid requirements of the honey bee (Apis mellifera L.). Physiol Comp Oecol 3:197–285
Somerville DC (2001) Nutritional value of bee collected pollens. Rural Industries Research and Development Corporation 1–166
Jones JC, Fruciano C, Hildebrand F, Al Toufalilia H, Balfour NJ, Bork P, Engel P, Ratnieks FLW, Hughes WOH (2018) Gut microbiota composition is associated with environmental landscape in honey bees. Ecol Evol 8:441–451. https://doi.org/10.1002/ece3.3597
Sabree ZL, Hansen AK, Moran NA (2012) Independent studies using deep sequencing resolve the same set of core bacterial species dominating gut communities of honey bees. PLoS One 7:e41250. https://doi.org/10.1371/journal.pone.0041250
Moran NA, Hansen AK, Powell JE, Sabree ZL (2012) Distinctive gut microbiota of honey bees assessed using deep sampling from individual worker bees. PLoS One 7:e36393. https://doi.org/10.1371/journal.pone.0036393
Raymann K, Moran NA (2018) The role of the gut microbiome in health and disease of adult honey bee workers. Curr Opin Insect Sci 26:97–104. https://doi.org/10.1016/j.cois.2018.02.012
Engel P, Martinson VG, Moran NA (2013) Functional diversity within the simple gut microbiota of the honey bee. Proc Natl Acad Sci U S A 109:11002–11007. https://doi.org/10.5281/zenodo.147966
Kwong WK, Moran NA (2013) Cultivation and characterization of the gut symbionts of honey bees and bumble bees: description of Snodgrassella alvi gen. nov., sp. nov., a member of the family Neisseriaceae of the Betaproteobacteria, and Gilliamella apicola gen. nov., sp. nov., a member of Orbaceae fam. nov., Orbales ord. nov., a sister taxon to the order ‘Enterobacteriales’ of the Gammaproteobacteria. Int J Syst Evol Microbiol 63:2008–2018. https://doi.org/10.1099/ijs.0.044875-0
Ellegaard KM, Tamarit D, Javelind E, Olofsson TC, Andersson SG, Vasquez A (2015) Extensive intra-phylotype diversity in lactobacilli and bifidobacteria from the honeybee gut. BMC Genomics 16:284. https://doi.org/10.1186/s12864-015-1476-6
Engel P, Kwong WK, Moran NA (2013) Frischella perrara gen. Nov., sp. nov., a gammaproteobacterium isolated from the gut of the honeybee, Apis mellifera. Int J Syst Evol Microbiol 63:3646–3651. https://doi.org/10.1099/ijs.0.049569-0
Kesnerova L, Moritz R, Engel P (2016) Bartonella apis sp. nov., a honey bee gut symbiont of the class Alphaproteobacteria. Int J Syst Evol Microbiol 66:414–421. https://doi.org/10.1099/ijsem.0.000736
Corby-Harris V, Snyder LA, Schwan MR, Maes P, McFrederick QS, Anderson KE (2014a) Origin and effect of alpha 2.2 Acetobacteraceae in honey bee larvae and description of Parasaccharibacter apium gen. Nov., sp. nov. Appl Environ Microbiol 80:7460–7472. https://doi.org/10.1128/AEM.02043-14
Evans JD, Spivak M (2010) Socialized medicine: individual and communal disease barriers in honey bees. J Invertebr Pathol 103:S62–S72. https://doi.org/10.1016/j.jip.2009.06.019
Danihlík J, Aronstein K, Petřivalský M (2016) Antimicrobial peptides: a key component of honey bee innate immunity. J Apic Res 54:123–136. https://doi.org/10.1080/00218839.2015
White JWJ, Subers MH, Schepartz AI (1963) The identification of inhibine, antibacterial factor in honey, as hydrogen peroxide, and its origin in a honey glucose oxidase system. Biochim Biophys Acta 73:57–70. https://doi.org/10.1016/0926-6569(63)90108-1
Doublet V, Poeschl Y, Gogol-Döring A, Alaux C, Annoscia D, Aurori C, Barribeau SM, Bedoya-Reina OC, Brown MJ, Bull JC, Flenniken ML, Galbraith DA, Genersch E, Gisder S, Grosse I, Holt HL, Hultmark D, Lattorff HM, Le Conte Y, Manfredini F, McMahon DP, Moritz RF, Nazzi F, Niño EL, Nowick K, van Rij RP, Paxton RJ, Grozinger CM (2017) Unity in defense: honeybee workers exhibit conserved molecular responses to diverse pathogens. BMC Genomics 18:207. https://doi.org/10.1186/s12864-017-3597-6
Faegri K, Iversen J (1975) Textbook of modern pollen analysis. T Munksgaard Copenhagen, p 423
Louveaux J, Mauricio A, Vorwohl G (1978) Methods of melissopalynology. Bee World 59:139–157. https://doi.org/10.1080/0005772X.1978.11097714
Wiliams GR, Alaux C, Costa C, Csákit T, Doublet V, Eisenhardt D, Fries I, Kuhn R, Mcmahon DP, Medrzycki P, Murray TE, Natsopoulou ME, Neumann, P, Oliver R, Paxton RJ, Pernal SF, Shutler D, Tanner G, Van der steen JJM, Brodschneider R (2013) Standard methods for maintaining adult Apis mellifera in cages under in vitro laboratory conditions. In V Dietemann, JD Ellis, P Neumann (Eds) the COLOSS BEEBOOK, volume I: standard methods for Apis mellifera research. J Apic Res 52(1). https://doi.org/10.3896/IBRA.1.52.1.04
Porrini MP, Garrido PM, Eguaras MJ (2013) Individual feeding of honeybees:modification of the Rinderer technique. J Apic Res 52:194–195
Fries I, Chauzat MP, Chen YP, Doublet V, Genersch E, Gisder S, Higes M, McMahon DP, Martín-Hernández R, Natsopoulou M, Paxton RJ, Tanner G, Webster TC, Williams GR (2013) Standard methods for Nosema research. J Apic Res 52:1–28. https://doi.org/10.3896/IBRA.1.52.1.14
Human H, Beodschneider R, Dietemann V, Dively G, Ellis J, Forsgren E, Fries I, Hatjina F, Hu F-L, Jaffé R, Köhler A, Pirk CWW, Rose R, Strauss U, Tanner G, Van der Steen JJM, Vejsnaes F, Williams GR, Zheng H-Q (2013) Miscellaneous standard methods for Apis mellifera research. In V Dietemann; J D Ellis; P Neumann (Eds) the COLOSS BEEBOOK, volume I: standard methods for Apis mellifera research. J Apic Res 52(4). https://doi.org/10.3896/IBRA.1.52.4.10
Martin-Hernandez R, Meana A, Prieto L, Salvador AM, Garrido-Bailon E, Higes M (2007) Outcome of colonization of Apis mellifera by Nosema ceranae. Appl Environ Microbiol 73:6331–6338. https://doi.org/10.1128/AEM.00270-07
Anderson KE, Carroll MJ, Sheehan T, Lanan MC, Mott BM, Maes P, Corby-Harris V (2014) Hive-stored pollen of honey bees: many lines of evidence are consistent with pollen preservation, not nutrient conversion. Mol Ecol 23:5904–5917. https://doi.org/10.1111/mec.12966
Zhou J, Bruns MA, Tiedje JM (1996) DNA recovery from soils of diverse composition. Appl Environ Microbiol 62:316–322 0099-2240/96/$04.0010
Arismendi N, Bruna A, Zapata N, Vargas M (2016) PCR-specific detection of recently described Lotmaria passim (Trypanosomatidae) in Chilean apiaries. J Invertebr Pathol 134:1–5. https://doi.org/10.1016/j.jip.2015.12.008
Denman SE, McSweeney CS (2006) Development of a real-time PCR assay for monitoring anaerobic fungal and cellulolytic bacterial populations within the rumen. FEMS Microbiol Ecol 58:572–582. https://doi.org/10.1111/j.1574-6941.2006.00190.x
Evans JD (2006) Beepath: an ordered quantitative-PCR array for exploring honey bee immunity and disease. J Invertebr Pathol 93:135–139. https://doi.org/10.1016/j.jip.2006.04.004
Ott SJ, Musfeldt M, Ullmann U, Hampe J, Schreiber S (2004) Quantification of intestinal bacterial populations by real-time PCR with a universal primer set and minor groove binder probes: a global approach to the enteric Flora. J Clin Microbiol 42:2566–2572. https://doi.org/10.1128/JCM.42.6.2566-2572.2004
Yang X, Cox-Foster DL (2005) Impact of an ectoparasite on the immunity and pathology of an invertebrate: evidence for host immunosuppression and viral amplification. Proc Natl Acad Sci U S A 102:7470–7475. https://doi.org/10.1073/pnas.0501860102
Corona M, Velarde R, Remolina S, Moran-Lauter A, Wang Y, Hughes KA, Robinson GE (2007) Vitellogenin, juvenile hormone, insulin signalling, and queen honey bee longevity. Proc Natl Acad Sci U S A 104:7128–7133. https://doi.org/10.1073/pnas.0701909104
Engel P, James RR, Koga R, Kwong WK, McFrederick QS, Moran NA (2013) Standard methods for research on Apis mellifera gut symbionts. J Apic Res 52(4):1–24. https://doi.org/10.3896/IBRA.1.52.4.07
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Lan Y, Wang Q, Cole JR, Rosen GL (2012) Using the RDP classifier to predict taxonomic novelty and reduce the search space for finding novel organisms. Public Library of Science ONE 7:e32491. https://doi.org/10.1371/journal.pone.0032491
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
R Core Team (2013) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29(e45):387–409. https://doi.org/10.1051/apido:2000130
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: Paleontological statistics software package for education and data analysis. Palaeontol Electron:4 http://palaeo-electronica.org/2001_2001/past/issue2001_2001.htm
Powell JE, Martinson VG, Urban-Mead K, Moran NA (2014) Routes of Acquisition of the gut microbiota of the honey bee Apis mellifera. Appl Environ Microbiol 80:7378–7387. https://doi.org/10.1128/AEM.01861-14
Bottacini F, Milani C, Turroni F, Sánchez B, Foroni E, Duranti S, Serafini F, Viappiani A, Strati F, Ferrarini A, Delledonne M, Henrissat B, Coutinho P, Fitzgerald GF, Margolles A, van Sinderen VM (2012) Bifidobacterium asteroides PRL2011 genome analysis reveals clues for colonization of the insect gut. PLoS One 7(9):e44229. https://doi.org/10.1371/journal.pone.0044229
Kwong WK, Moran NA (2016) Gut microbial communities of social bees. Nat Rev Microbiol 14:374–384. https://doi.org/10.1038/nrmicro.2016.43
Forsgren E, Olofsson TC, Vasquez A, Fries I (2010) Novel lactic acid bacteria inhibiting Paenibacillus larvae in honey bee larvae. Apidologie 41:99–108. https://doi.org/10.1051/apido/2009065
Vásquez A, Forsgren E, Fries I, Paxton RJ, Flaberg E, Szekely L, Olofsson TC (2012) Symbionts as major modulators of insect health: lactic acid Bacteria and honeybees. PLoS One 7(7):e33188. https://doi.org/10.1371/journal.pone.0033188
Killer J, Dubná S, Sedláček I, Švec P (2014) Lactobacillus apis sp. nov., from the stomach of honeybees (Apis mellifera), having an in vitro inhibitory effect on the causative agents of American and European foulbrood. Int J Syst Evol Microbiol 64:152–157. https://doi.org/10.1099/ijs.0.053033-0
Chomel BB, Kasten RW (2010) Bartonellosis, an increasingly recognized zoonosis. J Appl Microbiol 109:743–750. https://doi.org/10.1111/j.1365-2672.2010.04679.x
Kopecky J, Nesvorna M, Hubert J (2014) Bartonella-like bacteria carried by domestic mite species. Exp Appl Acarol 64:21–32. https://doi.org/10.1007/s10493-014-9811-1
Hubert J, Erban T, Kamler M, Kopecky J, Nesvorna M, Hejdankova S, Titera D, Tyl J, Zurek L (2015) Bacteria detected in the honeybee parasitic mite Varroa destructor collected from beehive winter debris. J Appl Microbiol 119:640–654. https://doi.org/10.1111/jam.12899
Amdam GV, Norberg K, Hagen A, Omholt SW (2003) Social exploitation of vitellogenin. Proc Natl Acad Sci USA 100: 1799-1802. https://doi.org/10.1073/pnas.0333979100
Seehuus SC, Norberg K, Gimsa U, Krekling T, Amdam GV (2006) Reproductive protein protects functionally sterile honey bee workers from oxidative stress. Proceedings of the National Academy of Sciences USA 103:962–967. https://doi.org/10.1073/pnas.0502681103
Nelson CM, Ihle KE, Fondrk MK, Page Jr RE, Amdam GV (2007) The gene vitellogenin has multiple coordinating effects on social organization. PLoS Biology 5: 0673–0677. https://doi.org/10.1371/journal.pbio.0050062
Bitondi MMG, Simões ZLP (1996) The relationship between level of pollen in the diet, vitellogenin and juvenile hormone titres in Africanized Apis mellifera workers. J Apic Res 35:27–36. https://doi.org/10.1080/00218839.1996.11100910
Basualdo M, Barragán S, Vanagas L, García C, Solana H, Rodríguez E, Bedascarrasbure E (2013) Conversion of high and low pollen protein diets into protein in worker honey bees (Hymenoptera: Apidae). J Econ Entomol 106:1553–1558. https://doi.org/10.1603/EC12466
Corby-Harris V, Jones BM, Walton A, Schwan MR, Anderson KE (2014) Transcriptional markers of sub-optimal nutrition in developing Apis mellifera nurse workers. BMC Genomics 15:134. https://doi.org/10.1186/1471-2164-15-134
Casteels P, Ampe C, Jacobs F, Tempst P (1993) Functional and chemical characterization of Hymenoptaecin, an antibacterial polypeptide that is infection-inducible in the honeybee (Apis mellifera). J Biol Chem 268:7044–7054
Gillespie JP, Kanost M (1997) Biological mediators of insect immunity. Annu Rev Entomol 42:611–643. https://doi.org/10.1146/annurev.ento.42.1.611
Imler JL, Bulet P (2005) Antimicrobial peptides in Drosophila: structures, activities and gene regulation. Chem Immunol Allergy 86:1–21. https://doi.org/10.1159/000086648
Maggi M, Negri P, Plischuk S, Szawarski N, DePiano F, De Feudis L, Eguaras M, Audisio C (2013) Effects of the organic acids produced by a lactic acid bacterium in Apis mellifera colony development, Nosema ceranae control and fumagillin efficiency. Vet Microbiol https://doi.org/10.1016/j.vetmic.2013.07.030
Baffoni L, Gaggìa F, Alberoni D, Cabbri R, Nanetti A, Biavati B, Di Gioia D (2015) Effect of dietary supplementation of Bifidobacterium and Lactobacillus strains in Apis mellifera L. against Nosema ceranae. Benef Microbes 7:1–8. https://doi.org/10.3920/BM2015.0085
Arredondo D, Castelli L, Porrini M, Garrido M, Eguaras M, Zunino P, Antúnez K (2017) Lactobacillus kunkeei strains decreased the infection by honey bee pathogens Paenibacillus larvae and Nosema ceranae. Benef Microbes 9:1–12. https://doi.org/10.3920/BM2017.0075
Rubanov A, Russell KA, Rothman JA, Nieh JC, McFrederick QS (2019) Intensity of Nosema ceranae infection is associated with specific honey bee gut bacteria and weakly associated with gut microbiome structure. Sci Rep 9:3820. https://doi.org/10.1038/s41598-019-40347-6
Antúnez K, Martín-Hernández R, Prieto L, Meana A, Zunino P, Higes M (2009) Immune suppression in the honey bee (Apis mellifera) following infection by Nosema ceranae (Microsporidia). Environ Microbiol 11:2284–2290. https://doi.org/10.1111/j.1462-2920.2009.01953.x
Mayack C, Naug D (2009) Energetic stress in the honeybee Apis mellifera from Nosema ceranae infection. J Invertebr Pathol 100:185–188. https://doi.org/10.1016/j.jip.2008.12.001
Acknowledgments
This work was supported by “Comisión Sectorial de Investigación Científica” (CSIC 6406) and “Agencia Nacional de Investigación e Innovación” (POS_NAC_2013_1_12228), Uruguay. Authors thank researchers and technicians from the Instituto Nacional de Investigación Agropecuaria (Yamandú Mendoza, Gustavo Ramallo, Carlos Silva and Sebastián Díaz) for their help with sample collection. Finally, we thank the anonymous reviewers for their valuable comments.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(PDF 631 kb)
Rights and permissions
About this article
Cite this article
Castelli, L., Branchiccela, B., Garrido, M. et al. Impact of Nutritional Stress on Honeybee Gut Microbiota, Immunity, and Nosema ceranae Infection. Microb Ecol 80, 908–919 (2020). https://doi.org/10.1007/s00248-020-01538-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-020-01538-1