Abstract
In organisms with complex life histories, dramatic changes in microbial community structure may occur with host development and immune system maturation. Amphibian host susceptibility to diseases such as chytridiomycosis may be affected by the reorganization of skin microbial community structure that occurs during metamorphosis. We tracked changes in the bacterial communities inhabiting skin of Korean fire-bellied toads (Bombina orientalis) that we infected as tadpoles with different strains of Batrachochytrium dendrobatidis (Bd), the pathogenic fungus that causes chytridiomycosis. We found that B. orientalis undergoes a major change in skin bacterial community composition between 5 and 15 days following metamorphosis. Richness indices and phylogenetic diversity measures began to diverge earlier, between aquatic and terrestrial stages. Our results further reveal differences in skin bacterial community composition among infection groups, suggesting that the effect of Bd infection on skin microbiome composition may differ by Bd strain. Additional studies are needed to further investigate the structural and temporal dynamics of microbiome shifts during metamorphosis in wild and captive amphibian populations. Analyses of the ontogeny of microbiome shifts may contribute to an understanding of why amphibians vary in their susceptibility to chytridiomycosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hosts and the microbial communities with which they live have multifarious interactions [1]. The microbial communities can vary temporally and spatially, especially on the skin [2, 3]. Resident skin microbiota can effectively shield hosts from, or alternatively make them more vulnerable to, environmental disturbances and pathogens [4, 5]. Thus, studies of temporal patterns of skin microbiota diversity may provide clues as to varying disease susceptibility. These changes may be especially dramatic in organisms with complex life histories that transition between habitats with fundamentally different environmental and ecological conditions.
Amphibians undergo radical organizational transformation as they metamorphose from an aquatic to largely terrestrial life form. For example, larval skin rapidly degenerates and is remodeled into a multilayered glandular epidermis including keratinized and cornified stratum corneum [6,7,8]. Most physiological functions, including the immune system, are reorganized during metamorphosis. Before metamorphosis, tadpoles are immunocompetent despite producing less diverse antibodies than adults and expressing major histocompatibility complex (MHC) class II but not class I molecules on certain epithelial cell surfaces [9]. During metamorphosis, the adaptive immune system shuts down, perhaps to prevent larval lymphocytes from attacking newly emerging antigens on adult tissues, so individuals may be especially susceptible to pathogens [9]. Within 1 week of metamorphosis, however, class I and II MHC molecules, which guide T-lymphocyte recognition of antigens, rapidly appear on all tissues (class I), and on antigen-presenting cells and lymphocytes (class II), enhancing adaptive immune responses [10, 11]. Although some amphibians produce anti-microbial peptides as larvae [12], the innate immune system typically becomes fully functional only several weeks after metamorphosis [9, 13].
Amphibian skin microbiomes should be influenced by physiological and immunological changes accompanying metamorphosis. Indeed, recent field studies have shown that the skin bacterial community of Rana cascadae frogs and Anaxyrus boreas toads shifts during development, as alpha diversity increases from larval to metamorphic stages [14, 15]. Differences in bacterial community structure and diversity were also found between juveniles and adults in the direct-developing frog Eleutherodactylus coqui [16]. However, studies of subjects sampled in the field, such as these, do not follow the ontogeny of bacteria communities borne by specific individuals, so they provide little insight into the timing of this skin bacterial community shift.
Skin bacteria contribute to the defenses of their amphibian hosts against Batrachochytrium dendrobatidis (Bd) [17,18,19], a fungal pathogen infecting keratinized epithelial cells of amphibians. Bd causes chytridiomycosis, a disease that is associated with worldwide amphibian population declines and species extinctions [20, 21]. Innate immune defenses against Bd may be enhanced by skin bacteria [22] that produce anti-Bd metabolite mixtures [23]. Skin bacterial community changes after metamorphosis may increase host susceptibility to infection [24,25,26,27]. Clearly, a better understanding of how amphibian skin microbiome dynamics are influenced by Bd infection is needed to develop effective disease mitigation strategies.
Here, we track changes in bacterial community structure on the skin of fire-bellied toads (Bombina orientalis) as they metamorphose from tadpoles to toadlets in the laboratory. This species, like other Asian amphibians, appears resilient to Bd infection [28,29,30,31], probably because of their historical association with native Bd strains [29]. Native Korean Bd strains are phylogenetically very different from the invasive global pandemic lineage (Bd-GPL) associated with global amphibian population declines [32]. Thus, they are likely to differ from Bd-GPL strains in important characteristics linked to virulence [33] and their effects on skin bacterial community structure [27]. In this study, we experimentally inoculate subjects during their development with either an endemic Bd strain isolated from Korean B. orientalis or an invasive strain of the Bd-GPL lineage to determine how Bd infection modifies the skin bacterial community during and following metamorphosis.
Material and Methods
Bd Infection Experimental Design
Two egg clutches of B. orientalis were collected directly from mating pairs in Chuncheon, Gangwon Province, South Korea (37° 58.664′ N 127° 36.146′ E), and then hatched in the laboratory. Once tadpoles had a fully developed operculum and no visible gills (Gosner stage 25 [34]), 35 tadpoles were transferred individually into 1-L polypropylene flasks containing 140 mL of UV-treated charcoal-filtered water, which were placed into an incubator at 20 °C on a 12:12 L/D photoperiod. The tadpoles were assigned randomly to three treatments: (i) inoculation with a Korean Bd strain isolated from B. orientalis (strain KBO327T6, N = 12, hereafter denoted “Bd-Korea”), (ii) inoculation with a hypervirulent Bd-GPL strain [32] isolated in Australia (strain AbercrombieNP-L.booroolongensis-09-LB-P7, N = 12, hereafter denoted “Bd-Australia”), and (iii) inoculation with a control sham made of a filtrate from live Korean Bd culture (N = 9).
Zoospore concentration of Bd cultures was assayed before exposure by counting only active zoospores using a hemocytometer. Sterile water was used to dilute the zoospore inocula when necessary. Filtrates were obtained from active Korean Bd culture using a 0.2-μm sterile filter.
Tadpoles were repeatedly inoculated with 10,000 active zoospores every 4 days to coincide with water changes (days 0, 4, 8, and 12). Tadpoles were fed boiled spinach ad libitum every other day. Once forelimbs of a tadpole emerged, the water in its flask was reduced by 70%, the flask tilted to provide both aquatic and terrestrial habitat, and a last inoculation was performed. When an individual’s tail had resorbed to a stub (Gosner stage 45 [34]), the subject was considered to have metamorphosed, and water was removed from the flask and replaced with moist paper towel. Metamorphs were fed with pinhead crickets (Gryllus bimaculatus) ad libitum, and paper towels were changed every 4 days.
Survival was recorded on a daily basis. Difference in survival rates among treatments was tested using a Kaplan-Meier log rank survival analysis with survival days as the response variable. The infection experiment was run for 76 days from the first Bd inoculation. Throughout the experiment, laboratory material and water waste was placed into 10% sodium hypochlorite solution and autoclaved to kill Bd. Surviving individuals were euthanized by immersion in a 5-g/L solution of MS-222 [35].
Subjects were rinsed with sterile water before swabbing. The whole body of each individual was swabbed using sterile cotton swabs (MW100, Medical Wire and Equipment, Corsham, Wiltshire, UK) five times during the experiment: when tadpoles had fully developed, keratinized hindlimbs susceptible to Bd infection (Gosner stage 38) [34, 36], and at 1, 5, 15, and 30 days post-metamorphosis. The mouthparts of tadpoles, which represent the main target of Bd infection during the aquatic stage, were included in the body swab. Swabs were stored in sterile vials at −70 °C until DNA extraction.
Molecular Laboratory Work
DNA was extracted from the swabs using PowerSoil DNA isolation kits (Mo Bio Laboratories, Carlsbad, CA, USA). Blank DNA extractions with a sterile swab were run in parallel to swab samples. These blank extractions were used in subsequent PCR amplification to test for potential contamination during the extraction process. Bacterial skin communities were determined using barcoded Illumina sequencing of the 16S rRNA gene with primers targeting a 250-bp portion of the V4 region. Sample-identifying barcode sequences were included at the 5′ ends on the forward and reverse primers (total of 20 different barcoded primers, Table S1). PCR amplification was prepared in 50-μL reactions using 20–30 ng of template DNA in a laminar flow hood to minimize contamination, using a different set of forward and reverse primers for each sample (see Table S1 for detailed PCR protocol). Each run was accompanied by a negative control (1 μL of sterile water instead of DNA). Each sample was amplified in triplicate, and all PCR amplicons were pooled together at similar concentration (50 ng/μL per sample). These pooled samples were sequenced using a MiSeq Illumina sequencer by Macrogen (Seoul, South Korea).
Bd Screening
A portion of the DNA extracted from swabs was used to screen samples for Bd infection using two different PCR methods to decrease the risk of false negative results [37]. First, we performed a highly sensitive nested PCR method specifically targeting the 5.8S rDNA and the ribosomal internal transcribed spacer regions (ITS) of Bd [38]. The first PCR was run in a volume of 20 μL containing 1 μL of DNA sample, 0.2 μM of forward primer Bd18SF1 (5′-TTTGTACACACCGCCCGTCGC-3′) and reverse primer Bd28SR1 (5′-ATATGCTTAAGTTCAGCGGG-3′), 0.2 mM of each dNTP, 2 mM of MgCl2, and 1.0 unit of TaKaRa Ex Taq DNA Polymerase (Takara Bio, Otsu, Shiga, Japan). The PCR conditions consisted of an initial denaturation at 94 °C for 5 min, followed by 30 cycles of 30 s at 94 °C, 30 s at 50 °C, and 2 min at 72 °C, and a final extension at 72 °C for 7 min. A second PCR was run in a volume of 20 μL containing 1 μL of products from the first PCR, 0.2 μM of forward primer, Bd1a (5′-CAGTGTGCCATATGTCACG-3′) and reverse primer, Bd2a (5′-CATGGTTCATATCTGTCCAG-3′), 0.2 mM of each dNTP, 2 mM of MgCl2, and 1.0 unit of TaKaRa Ex Taq DNA polymerase. The conditions for the second PCR consisted of an initial denaturation at 94 °C for 5 min, followed by 30 cycles of 45 s at 94 °C, 45 s at 60 °C, and 60 s at 72 °C, and a final extension at 72 °C for 7 min. Each sample was run in triplicate with positive (DNA from strain AbercrombieNP-L.booroolongensis-09-LB-P7) and negative (1 μL of sterile water) controls. Status of Bd infection was determined by presence/absence of a band after electrophoresis on an agarose gel. Samples were considered positive when finding at least one positive result from the three replicates.
Intensity of infection was estimated in terms of Bd zoospore genomic equivalents (ZGEs) in all swab samples using a qPCR assay [39]. The qPCR assay was run on an Illumina Eco Real-Time PCR System (Illumina, San Diego, CA, USA) in a volume of 10 μL containing 1× SYBR Green quantitative PCR reagent kit (PhileKorea Technology, Seoul, South Korea), 0.25 mM of both ITS1–3 Chytr (5′-CCTTGATATAATACAGTGTGCCATATGTC-3′) and 5.8S Chytr (5′-AGCCAAGAGATCCGTTGTCAA-3′) primers, and 2 μL of DNA. The PCR conditions consisted of an initial denaturation at 95 °C for 10 min, followed by 50 cycles of 10 s at 95 °C and 1 min at 58 °C. Each sample was assayed in duplicate, along with standards of known Bd quantity (100, 10, 1, and 0.1 zoospores, strain AbercrombieNP-L.booroolongensis-09-LB-P7) and negative controls (5 μL sterile water). We estimated Bd ZGEs per swab, averaged over the two replicates, from threshold cycle (Ct) values after corrections for dilutions following DNA extraction and PCR procedures.
Sequence Data Processing
Illumina sequencing data were pair-assembled using PANDAseq [40] with an assembly quality score of 0.9, the most stringent option to reduce errors. Pair-assembled sequences were trimmed, aligned, and filtered using the mothur pipeline [41]. Alignment was done using the EzTaxon database version 2014 [42]. Additionally, chimeras detected using the chimera.uchime command within mothur were removed. We subsampled the sequence data to 6357 reads per sample, which corresponds to the minimum number of reads obtained across samples, excluding one sample that had 121 reads (AB6M3) which was removed from further analyses (Table S2). Any sequence present only once across samples (singleton) was removed from the dataset to improve the comparability of the sequence data and the resolution of the analyses [43, 44]. This filtered dataset was subsampled again to 6305 reads per sample to have comparable estimates of richness and diversity among samples.
The EzTaxon training set was used with Bayesian classifier algorithm implemented in mothur for taxonomic classification of operational taxonomic units (OTUs), clustered at ≥97% sequence similarity. No sequences matching chloroplast, mitochondria, eukaryote, or archaea were observed in the classified sequences. We estimated richness (i.e., number of OTUs and Chao Index) and diversity indices (i.e., non-parametric Shannon and inverse Simpson) using calculators within mothur [41]. Faith’s phylogenetic diversity (PD) [45] was estimated using the phylo.diversity command within mothur from a phylogenetic tree built using FastTreeMP [46].
Statistical Analyses
All statistical analyses were conducted on the subsampled dataset of 6305 reads per sample. To assess whether richness (number of OTUs and Chao index) and diversity indices (non-parametric Shannon and inverse Simpson index) differ in relation to sampling time and treatment, we used linear or generalized mixed models (GLMM) with sampling time, infection status, survival, treatment, and the interaction of survival with treatment as factors. Infection intensity was not included as a factor because values were low in all infected individuals (see “Results” section). Individual ID nested within clutch was included as a random factor, but clutch was later removed as its inclusion did not reduce variation more than models with only ID as a random factor.
Where necessary to improve the model owing to heterogeneity in the spread of residuals, different variances were allowed among sampling times or treatments for some diversity indices. An observational random effect, fitting a unique value for each observation, was included in generalized mixed models to account for overdispersion. ID was removed from the model if its standard deviation was less than that of the observational term, except in cases where including it showed a better fit of the data to the model. We used likelihood ratio tests (LRTs) to calculate the predictive power of each variable. Non-significant interactions and terms were removed sequentially. In cases of significant effect of sampling time in a model, pairwise differences between sampling times were estimated with Wald z-statistics, correcting for multiple comparisons using the false discovery rate method [47].
We used permutational multivariate analyses (PERMANOVA, Anderson 2001) with 999 permutations to assess whether community composition (beta diversity) differed in relation to sampling time, infection status, treatment, and survival. Bray-Curtis and UniFrac distance matrices were used as measures of community structure for these analyses. Differences in community structure were visualized with non-metric multidimensional scaling (NMDS) plots based on these matrices. All statistical analyses were run with R v3.0.2 [48].
Phyla, Order, and Core Community Analyses
We defined the core community as those OTUs that were present on at least 90% of all the samples [23]. We calculated the relative abundance of these OTUs, of phyla, and of orders within the phyla Proteobacteria and Bacteroidetes (the two most abundant phyla, representing more than 78% of the community in all samples; see “Results” section) by dividing the number of sequences of each OTU/phylum/order by the total number of reads per sample (i.e., 6305). Phyla and orders were retained for further analyses if their mean relative abundance exceeded a threshold of 0.5% at a minimum of one sampling stage within a treatment.
We used GLMM to assess whether the relative abundance of phyla, orders, and core OTUs differed in relation to sampling time, infection status, treatment, and survival, and the interaction between treatment and survival following the same procedure, as described above. P values were corrected for multiple comparisons using the false discovery rate method [47].
Results
Time to metamorphosis did not differ among treatments (control 54.7 ± 2.5 days, Bd-Korea 48.3 ± 1.7 days, Bd-Australia 51.3 ± 1.9 days, LRT2 = 2.34, P = 0.31). Survival was low across treatments, with only two or three individuals in each group surviving until the end of the experiment (Fig. 1). Chances of survival were not significantly different among treatments (Kaplan-Meier log rank = 0.39, df = 2, P = 0.82). Low levels of Bd infection were detected in three subjects in the Bd-Korea treatment and six subjects in the Bd-Australia treatment at one or more sampling times (range 1.5–35.6 ZGEs per swab, Table S2). Bd prevalence did not significantly differ between the Bd-Korea and Bd-Australia treatments (χ 2 1 = 0.01, P = 0.91).
Richness and Diversity
After sequence processing, we obtained 472,875 reads of 75 swab samples, and 993 OTUs (raw data available in the NCBI Sequence Read Archive, accession SRP066198). The number of OTUs and Chao richness index differed in relation to sampling time (GLMMs, OTUs: LRT4 = 17.17, P = 0.002, Chao: LRT4 = 22.08, P = 0.0002; Table S3); they were significantly higher at stage 38 than at later sampling times (Fig. 2, Table S4). Diversity indices did not differ among sampling times, by survival, nor by infection status (Table S3). Faith’s PD differed among sampling times (LRT4 = 21.44, P = 0.0002; Table S3), being significantly higher at stage 38 than at later stages and at 1 and 5 days compared to 15 and 30 days post-metamorphosis (Fig. 2, Table S4).
Examination of individual trajectories of bacterial richness and diversity across sampling times did not reveal any relationship between the initial state of the bacterial alpha diversity in individuals experimentally exposed to a Bd strain and their survival to infection (Fig. S1). For individuals in the control group, levels of bacterial richness and diversity became more similar at 5 days post-metamorphosis, before diverging again at later stages (Fig. S1). This trend was not observed in Bd-infected individuals. For the Bd-Korea group, richness indices appeared more variable from 5 days on (Fig. S1).
Community Structure
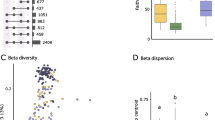
Bacterial community structure (beta diversity) differed among sampling times (PERMANOVA, Bray-Curtis: pseudo-F = 5.04, P = 0.001, UniFrac: pseudo-F = 2.59, P = 0.001) and treatments (Bray-Curtis: pseudo-F = 2.26, P = 0.001, UniFrac: pseudo-F = 1.60, P = 0.001). However, bacterial community structure did not vary with survivorship (Bray-Curtis: pseudo-F = 1.09, P = 0.31, UniFrac: pseudo-F = 1.26, P = 0.06) nor Bd infection status (Bray-Curtis: pseudo-F = 1.06, P = 0.37, UniFrac: pseudo-F = 1.06, P = 0.27). The NMDS plots showed that bacterial community composition differed among a group formed by early sampling at stage 38, 1 and 5 days, and the later sampling at 15 and 30 days on axis 1 of NMDS plots (Fig. 3 and Fig. S2). These two later sampling times shared 77.6% of OTUs, whereas other stages shared between 32.1 and 51.3% of OTUs (Fig. 4). All OTUs identified at 15 and 30 days also were present in earlier stages (Fig. 4).
Venn diagrams showing the proportion of unique and shared OTUs on the skin of B. orientalis at different sampling stages and subject to different Bd infection treatments. Each diagram represents the proportion of a shared OTUs between consecutive sampling stages, b shared and unique OTUs between early (stage 38, 1 day, and 5 days pooled) and late sampling times (15 and 30 days pooled), and c shared and unique OTUs across treatment groups for the early and late sampling times. Bd-Kor, group infected by the Korean native Bd strain; Bd-Aus, group infected by the Australian Bd-GPL strain
Differences in composition among treatments were not evident in the NMDS plots. However, variation in bacterial community structure changed with sampling time, being less variable for the Bd-Korea treatment at earlier sampling times (stage 38, 1–5 days) but most variable for the later sampling times (15–30 days; Fig. 3 and S2). Control group individuals were most similar in bacterial community composition at 15 and 30 days. The three treatment groups shared 31.7% of OTUs at the earlier sampling times (stage 38, 1–5 days data pooled together), whereas 10.5–19.7% of the OTUs were unique to each treatment (Fig. 4). At the later sampling times, shared OTUs decreased to 22.7%, and the proportion of unique OTUs increased to 16.0 and 23.6% for the control and Bd-Australia groups, respectively. This proportion decreased from 19.7 to 16.9% in the Bd-Korea group (Fig. 4).
Relative Abundance of Major Phyla and Orders
Abundance of six major phyla, of six orders within the phylum Proteobacteria, and of five orders within the phylum Bacteroidetes differed among sampling times (Table 1, Figs. S3–S5). Some phyla and orders were significantly more abundant at 5 days than at 15 and 30 days. Others were significantly more abundant at 15 and 30 days, including Pseudomonadales and Flavobacteriales (Tables 1 and S5–S7, Figs. S3–S5).
Composition and Abundance of Core Bacterial Community
The core community was represented by 11 OTUs, which on average correspond to 50 ± 3% of the whole community (Fig. 5). The total relative abundance of core OTUs did not differ among sampling times (LRT4 = 5.77, P = 0.22) or treatments (LRT2 = 0.17, P = 0.92). All but the one most abundant core OTU (EU589321_EU591642) differed significantly among sampling times (Fig. 5, Tables 2 and S8). Some core OTUs, notably those within the order Burkholderiales, were more abundant in the early sampling times. Aeromonas salmonicida (order Aeromonadales) was most abundant at stage 38 (Fig. 5, Tables 2 and S8) and was more abundant in subjects that died before the end of the experiment (LRT1 = 15.75, P = 0.008). Other core OTUs, including those within the order Pseudomonadales, were more abundant at 15 and 30 days (Fig. 5, Tables 2 and S8).
We also observed differences in relative abundance among treatments for some core OTUs. Comamonadaceae_f .2 was most abundant at 5 days in the filtrate and Bd-Korea treatments only (Fig. 5, Tables 2 and S8). The OTU HQ178786 sp. was significantly more abundant in the control treatment (LRT1 = 12.91, P = 0.02).
Discussion
We found that a major reorganization of the skin bacterial community of B. orientalis accompanies other metamorphic changes. Richness and phylogenetic diversity (alpha diversity) started to decrease during metamorphosis, whereas community structure (beta diversity) changes were apparent only later, between 5 and 15 days after metamorphosis. The timing of changes in skin bacteria alpha and beta diversity has not previously been documented in metamorphosing amphibians. We cannot be certain, however, that this asynchronicity between alpha and beta diversity changes occurs in natural conditions. Additional studies are needed to further investigate the structural and temporal dynamics of microbiome shift during metamorphosis in wild and captive amphibian populations.
The reduction of alpha diversity we observed during metamorphosis contrasts with the higher diversity reported in tadpoles, rather than adults, in wild populations of R. cascadae frogs and A. boreas toads [14, 15]. Multiple studies have shown that alpha diversity of amphibian skin bacterial communities can significantly decrease in captive environments with limited bacterial reservoirs [23, 49, 50]. Although the bacterial environment in our captive setting was not characterized, its diversity was most likely reduced as UV-treated charcoal-filtered water was used throughout the experiment.
Mortality was high across treatments, representing a potentially important caveat to our results. Bacterial contamination attributable to husbandry, notably by A. salmonicida, may be at the origin of this issue. Exposure to fungal metabolites inhibiting amphibian lymphocyte responses [51] also may have increased host susceptibility to environmental bacteria. Although this perturbation most likely affected the skin microbiome of the toads, the timing and characteristics of the bacterial community shift were observed across individuals regardless of their health. Indeed, our analyses showed that neither survival nor infection status had any effect on any indices of alpha and beta diversity except in the case of A. salmonicida abundance. Thus, the patterns of community reorganization that we observed appear robust. Still, further studies in optimized experimental settings would help confirm the results obtained.
Larval microbial communities persisted through the transition from an aquatic to a terrestrial environment as subjects underwent massive physiological and morphological changes. Microbiome reorganization was delayed until after metamorphic climax, commensurate with the maturation of the innate and adaptive immune systems [13, 52]. Susceptibility to Bd is affected by immune system function, so it is likely to decrease at this time. As keratinization of the epithelium begins prior to metamorphosis [36], metamorphosing larvae thus may be especially susceptible to chytridiomycosis [22].
We found significant differences in bacteria abundance and community composition among treatments. These results suggest that skin bacterial community composition changes differentially based on Bd infection history and strain specificity. A previous laboratory experiment had already shown that Bd infection causes changes in amphibian skin bacterial community composition [27]. Our control group could have been exposed to fungal metabolites present in the inoculated filtrate [51]. Thus, this treatment potentially includes the effects of Bd factors on the skin bacterial community in the absence of infection by live zoospores. In this case, the results obtained suggest that exposure to factors associated with Bd products affects the skin bacterial community differently than does infection by zoospores. This possibility should be further explored.
The skin microbiota is temporally and spatially highly variable. Especially dramatic changes in its composition occur during host development. We have shown that skin bacterial community reorganization follows physiological and ecological changes that accompany amphibian metamorphosis. Based on our results, we also suggest that development of the skin bacterial community is affected by Bd infection history. Variability in amphibian susceptibility to chytridiomycosis is still poorly understood and represents a major issue for amphibian conservation. This study opens new avenues of research on this topic.
References
McFall-Ngai M, Hadfield MG, Bosch TCG, Carey HV, Domazet-Loso T, Douglas AE, Dubilier N, Eberl G, Fukami T, Gilbert SF, Hentschel U, King N, Kjelleberg S, Knoll AH, Kremer N, Mazmanian SK, Metcalf JL, Nealson K, Pierce NE, Rawls JF, Reid A, Ruby EG, Rumpho M, Sanders JG, Tautz D, Wernegreen JJ (2013) Animals in a bacterial world, a new imperative for the life sciences Proc Natl Acad Sci USA 110:3229–3236
Costello EK, Lauber CL, Hamady M, Fierer N, Gordon JI, Knight R (2009) Bacterial community variation in human body habitats across space and time Science 326:1694–1697
Gao Z, Tseng CH, Pei Z, Blaser MJ (2007) Molecular analysis of human forearm superficial skin bacterial biota Proc Natl Acad Sci USA 104:2927–2932
Rosenthal M, Goldberg D, Aiello A, Larson E, Foxman B (2011) Skin microbiota: microbial community structure and its potential association with health and disease Infect Genet Evol 11:839–848
Grice EA, Segre JA (2011) The skin microbiome Nat Rev Micro 9:244–253
Brown DD, Cai L (2007) Amphibian metamorphosis Dev Biol 306:20–33
Ellison TR, Mathisen PM, Miller L (1985) Developmental changes in keratin patterns during epidermal maturation Dev Biol 112:329–337
Marantelli G, Berger L, Speare R, Keegan L (2004) Distribution of the amphibian chytrid Batrachochytrium dendrobatidis and keratin during tadpole development Pac Conserv Biol 10:173–179
Rollins-Smith LA (1998) Metamorphosis and the amphibian immune system Immunol Rev 166:221–230
Du Pasquier L, Flajnik MF (1990) Expression of MHC class II antigens during Xenopus development Dev Immunol 1:85–95
Salter-Cid L, Nonaka M, Flajnik MF (1998) Expression of MHC class Ia and class Ib during ontogeny: high expression in epithelia and coregulation of class Ia and imp7 genes J Immunol 160:2853–2861
Woodhams DC, Bell SC, Bigler L, Caprioli RM, Chaurand P, Lam BA, Reinert LK, Stalder U, Vazquez VM, Schliep K, Hertz A, Rollins-Smith LA (2016) Life history linked to immune investment in developing amphibians Conserv Physiol 4:cow025
Holden WM, Reinert LK, Hanlon SM, Parris MJ, Rollins-Smith LA (2015) Development of antimicrobial peptide defenses of southern leopard frogs, Rana sphenocephala, against the pathogenic chytrid fungus, Batrachochytrium dendrobatidis Dev Comp Immunol 48:65–75
Kueneman JG, Parfrey LW, Woodhams DC, Archer HM, Knight R, McKenzie VJ (2013) The amphibian skin-associated microbiome across species, space and life history stages Mol Ecol 23:1238–1250
Kueneman JG, Woodhams DC, Van Treuren W, Archer HM, Knight R, McKenzie VJ (2015) Inhibitory bacteria reduce fungi on early life stages of endangered Colorado boreal toads (Anaxyrus boreas) ISME J 10:934–944
Longo AV, Savage AE, Hewson I, Zamudio KR (2015) Seasonal and ontogenetic variation of skin microbial communities and relationships to natural disease dynamics in declining amphibians R Soc Open Sci 2:140377
Becker MH, Harris RN (2010) Cutaneous bacteria of the redback salamander prevent morbidity associated with a lethal disease PLoS One 5:e10957
Harris RN, Brucker RM, Walke JB, Becker MH, Schwantes CR, Flaherty DC, Lam BA, Woodhams DC, Briggs CJ, Vredenburg VT, Minbiole KPC (2009) Skin microbes on frogs prevent morbidity and mortality caused by a lethal skin fungus ISME J 3:818–824
Woodhams DC, Vredenburg VT, Simon M-A, Billheimer D, Shakhtour B, Shyr Y, Briggs CJ, Rollins-Smith LA, Harris RN (2007) Symbiotic bacteria contribute to innate immune defenses of the threatened mountain yellow-legged frog, Rana muscosa Biol Conserv 138:390–398
Berger L, Speare R, Daszak P, Green DE, Cunningham AA, Goggin CL, Slocombe R, Ragan MA, Hyatt AD, McDonald KR, Hines HB, Lips KR, Marantelli G, Parkes H (1998) Chytridiomycosis causes amphibian mortality associated with population declines in the rain forests of Australia and Central America Proc Natl Acad Sci USA 95:9031–9036
Skerratt LF, Berger L, Speare R, Cashins S, McDonald KR, Phillott AD, Hines HB, Kenyon N (2007) Spread of chytridiomycosis has caused the rapid global decline and extinction of frogs EcoHealth 4:125–134
Becker MH, Walke JB, Cikanek S, Savage AE, Mattheus N, Santiago CN, Minbiole KPC, Harris RN, Belden LK, Gratwicke B (2015) Composition of symbiotic bacteria predicts survival in Panamanian golden frogs infected with a lethal fungus Proc R Soc B 282:20142881
Loudon AH, Holland JA, Umile TP, Burzynski EA, Minbiole KPC, Harris RN (2014) Interactions between amphibians symbiotic bacteria cause the production of emergent anti-fungal metabolites Front Microbiol 5:441
Rachowicz LJ, Vredenburg VT (2004) Transmission of Batrachochytrium dendrobatidis within and between amphibian life stages Dis Aquat Org 61:75–83
Tobler U, Schmidt BR (2010) Within- and among-population variation in chytridiomycosis-induced mortality in the toad Alytes obstetricans PLoS One 5:e10927
Walker SF, Bosch J, Gomez V, Garner TWJ, Cunningham AA, Schmeller DS, Ninyerola M, Henk DA, Ginestet C, Arthur C-P, Fisher MC (2010) Factors driving pathogenicity vs. prevalence of amphibian panzootic chytridiomycosis in Iberia Ecol Lett 13:372–382
Jani AJ, Briggs CJ (2014) The pathogen Batrachochytrium dendrobatidis disturbs the frog skin microbiome during a natural epidemic and experimental infection Proc Natl Acad Sci USA 11:5049–5058
Bataille A, Cashins SD, Grogan LF, Skerratt LF, Hunter D, McFadden M, Scheele B, Brannelly LA, Marcis A, Harlow PS, Bell S, Berger L, Waldman B (2015) Susceptibility of amphibians to chytridiomycosis is associated with MHC class II conformation Proc R Soc B 282:20143127
Bai C, Liu X, Fisher MC, Garner TWJ, Li Y (2012) Global and endemic Asian lineages of the emerging pathogenic fungus Batrachochytrium dendrobatidis widely infect amphibians in China Div Distrib 18:307–318
Bataille A, Fong JJ, Cha M, Wogan GOU, Baek HJ, Lee H, Min MS, Waldman B (2013) Genetic evidence for a high diversity and wide distribution of endemic strains of the pathogenic chytrid fungus Batrachochytrium dendrobatidis in wild Asian amphibians Mol Ecol 22:4196–4209
Swei A, Rowley JJL, Rodder D, Diesmos MLL, Diesmos AC, Briggs CJ, Brown R, Cao TT, Cheng TL, Chong RA, Han B, Hero J-M, Hoang HD, Kusrini MD, Le DTT, McGuire JA, Meegaskumbura M, Min M-S, Mulcahy DG, Neang T, Phimmachak S, Rao D-Q, Reeder NM, Schoville SD, Sivongxay N, Srei N, Stock M, Stuart BL, Torres LS, Tran DTA, Tunstall TS, Vieites D, Vredenburg VT (2011) Is chytridiomycosis an emerging infectious disease in Asia? PLoS One 6:e23179
Farrer RA, Weinert LA, Bielby J, Garner TWJ, Balloux F, Clare F, Bosch J, Cunningham AA, Weldon C, du Preez LH, Anderson L, Pond SLK, Shahar-Golan R, Henk DA, Fisher MC (2011) Multiple emergences of genetically diverse amphibian-infecting chytrids include a globalized hypervirulent recombinant lineage Proc Natl Acad Sci USA 108:18732–18736
Fisher MC, Bosch J, Yin Z, Stead DA, Walker J, Selway L, Brown AJP, Walker LA, Gow NAR, Stajich JE, Garner TWJ (2009) Proteomic and phenotypic profiling of the amphibian pathogen Batrachochytrium dendrobatidis shows that genotype is linked to virulence Mol Ecol 18:415–429
Gosner KL (1960) A simplified table for staging anuran embryos and larvae with notes on identification Herpetologica 16:183–190
Torreilles SL, McClure DE, Green SL (2009) Evaluation and refinement of euthanasia methods for Xenopus laevis J Am Assoc Lab Anim Sci 48:512–516
McMahon TA, Rohr JR (2015) Transition of chytrid fungus infection from mouthparts to hind limbs during amphibian metamorphosis EcoHealth 12:188–193
Shin J, Bataille A, Kosch TA, Waldman B (2014) Swabbing often fails to detect amphibian chytridiomycosis under conditions of low infection load PLoS One 9:e111091
Goka K, Yokoyama JUN, Une Y, Kuroki T, Suzuki K, Nakahara M, Kobayashi A, Inaba S, Mizutani T, Hyatt AD (2009) Amphibian chytridiomycosis in Japan: distribution, haplotypes and possible route of entry into Japan Mol Ecol 18:4757–4774
Hyatt AD, Boyle DG, Olsen V, Boyle DB, Berger L, Obendorf D, Dalton A, Kriger K, Heros M, Hines H, Phillott R, Campbell R, Marantelli G, Gleason F, Coiling A (2007) Diagnostic assays and sampling protocols for the detection of Batrachochytrium dendrobatidis Dis Aquat Org 73:175–192
Masella AP, Bartram AK, Truszkowki JM, Brown DG, Neufeld JD (2012) PANDAseq: paired-end assembler for illumina sequences BMC Bioinformatics 13:31
Schloss PD, Westcott SI, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities Appl Environ Microbiol 75:7537–7541
Chun J, Lee HJ, Jung Y, Kim M, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences Int J Syst Evol Microbiol 57:2259–2261
Unterseher M, Jumpponen A, Öpik M, Tedersoo L, Moora M, Dormann CF, Schnittler M (2011) Species abundance distribution and richness estimations in fungal metagenomics—lessons learned from community ecology Mol Ecol 20:275–285
Zhou J, Wu L, Deng Y, Zhi X, Jiang Y-H, Tu Q, Xie J, Van Nostrand JD, He Z, Yang Y (2011) Reproducibility and quantitation of amplicon sequencing-based detection ISME J 5:1303–1313
Faith DP, Baker AM (2006) Phylogenetic diversity (PD) and biodiversity conservation: some bioinformatics challenges Evol Bioinforma 2:121–128
Price MN, Dehal PS, Arkin AP (2009) FastTree: computing large minimum-evolution trees with profiles instead of a distance matrix Mol Biol Evol 26:1641–1650
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing J R Statist Soc B 57:289–300
R Development Core Team (2011) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Bataille A, Lee-Cruz L, Tripathi B, Kim H, Waldman B (2016) Microbiome diversity differs between amphibian skin regions: implications for chytridiomycosis mitigation efforts Microbiol Ecol 71:221–232
Becker MH, Richards-Zawacki CL, Gratwicke B, Belden LK (2014) The effect of captivity on the cutaneous bacterial community of the critically endangered Panamanian golden frog (Atelopus zeteki) Biol Conserv 176:199–206
Rollins-Smith LA, Fites JS, Reinert LK, Shiakolas AR, Umile TP, Minbiole KPC (2015) Immunomodulatory metabolites released by the frog-killing fungus, Batrachochytrium dendrobatidis Infect Immun 83:4565–4570
Rollins-Smith LA, Ramsey JP, Pask JD, Reinert LK, Woodhams DC (2011) Amphibian immune defenses against chytridiomycosis: impacts of changing environments Integr Comp Biol 51:552–562
Acknowledgements
We thank Jaehyub Shin for assistance with laboratory work. The research was supported by the National Research Foundation of Korea (NRF), and funded by the Ministry of Education (2014-063422 to A.B. and 2015R1D1A1A01057282 to B.W) and by the Ministry of Science, ICT, and Future Planning (2010-0002767 and 2012R1A1A2044449 to B.W.) of the Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Animal husbandry and experimental protocols were approved by the Institutional Animal Care and Use Committee (SNU-140827-2) and the Institutional Biosafety Committee (SNUIBC-P120725-2-1) of Seoul National University. Fieldwork was conducted under a permit issued by the mayor’s office, Chuncheon, Gangwon Province. The study species is not threatened or legally protected in South Korea.
Electronic supplementary material
ESM 1
(DOCX 1004 kb)
Rights and permissions
About this article
Cite this article
Bataille, A., Lee-Cruz, L., Tripathi, B. et al. Skin Bacterial Community Reorganization Following Metamorphosis of the Fire-Bellied Toad (Bombina orientalis). Microb Ecol 75, 505–514 (2018). https://doi.org/10.1007/s00248-017-1034-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-1034-7