Abstract
Chile is topographically and climatically diverse, with a wide array of diverse undisturbed ecosystems that include native plants that are highly adapted to local conditions. However, our understanding of the diversity, activity, and role of rhizobacteria associated with natural vegetation in undisturbed Chilean extreme ecosystems is very poor. In the present study, the combination of denaturing gradient gel electrophoresis and 454-pyrosequencing approaches was used to describe the rhizobacterial community structures of native plants grown in three representative Chilean extreme environments: Atacama Desert (ATA), Andes Mountains (AND), and Antarctic (ANT). Both molecular approaches revealed the presence of Proteobacteria, Bacteroidetes, and Actinobacteria as the dominant phyla in the rhizospheres of native plants. Lower numbers of operational taxonomic units (OTUs) were observed in rhizosphere soils from ATA compared with AND and ANT. Both approaches also showed differences in rhizobacterial community structures between extreme environments and between plant species. The differences among plant species grown in the same environment were attributed to the higher relative abundance of classes Gammaproteobacteria and Alphaproteobacteria. However, further studies are needed to determine which environmental factors regulate the structures of rhizobacterial communities, and how (or if) specific bacterial groups may contribute to the growth and survival of native plants in each Chilean extreme environments.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Significant evidence indicates that rhizosphere microbiomes play an essential role in the performance and survival of plants in nature [6, 7]. Extreme environments are not the exception, and microbial communities may also exert beneficial effects on adaptation and survival of native plants to local harsh conditions. However, relatively, little is known about the diversity, distribution, and roles of bacteria in soil under natural vegetation from extreme environments. Much of our knowledge on taxonomy and functionality of bacterial communities in the rhizosphere has come from studies on agroecosystems (i.e., pasture, crop, and forest tree species).
Chile is a narrow and long country that includes a wide variety of climates and undisturbed extreme ecosystems, such as the Atacama Desert, Antarctica, Andes Mountains, Patagonia, etc. The Atacama Desert is considered to be one of the oldest and driest deserts on the Earth, and its soil has been compared to that of Mars [39]. The presence of bacteria in the extreme hyperarid core of the Atacama Desert has been explored by culture-dependent and -independent approaches, revealing low contents of culturable heterotrophic bacteria (in some samples, the equivalent of only 102~105 CFU g−1 of soil), low DNA content (<1.2 μg DNA g−1 of soil), and low bacterial diversity (<152 observed operational taxonomic units (OTUs) g−1) [3, 33, 39, 40]. However, the structures of bacterial communities associated with roots of native desert plants in the Atacama Desert are understudied.
Azua-Bustos et al. [3] mentioned that the diversities of soil bacteria in the Atacama Desert are similar to those reported in other extreme environments, such as Dry Valley in Antarctica. Antarctica is also considered to be a pristine extreme environment with a high vulnerability to global climate change. Antarctica harbors low nutrient soils with low soil moisture contents and high levels of salinity and solar radiation [9]. Studies in Antarctica have generally focused on the diversity of macroorganisms and the genesis, properties, and classification of soils. With respect to microbiology, most recent studies have focused on evaluation of the biotechnological potential of secondary metabolites (antibiotics, antitumor agents, enzymes, etc.) produced by culturable bacteria (particularly Actinobacteria), and few studies have explored microbial diversity and beneficial plant-microbe interactions. In this context, the application of the 16S ribosomal RNA (rRNA) gene-based molecular approaches has revealed similar patterns of bacterial diversity between the only two native vascular plant species (Deschampsia antarctica and Colobanthus quitensis) found in Antarctic ecosystems [56]. However, no correlation between bacterial community structure and native plant species was reported by Roesch et al. [47], who concluded that soil pH was the best predictor for microbial community structure in ice free zones in Antarctica.
Bacterial community structures associated with roots of native plants are being studied across agroecosystems and undisturbed ecosystems of northern, central, and southern of Chile [27]; however, our understanding of the diversity and distribution of rhizobacterial communities in Chilean extreme environments is still very poor. In microbial ecology, the selection of molecular techniques is often influenced by the expertise, costs, and equipment available in the laboratory [52], and the combination of two or three techniques is frequently applied to obtain a more reliable and complete overview of qualitative and quantitative information regarding bacterial communities in the environmental samples [14, 35]. Therefore, to obtain a more reliable and complete overview of qualitative and quantitative information of bacterial communities, a combination of denaturing gradient gel electrophoresis (DGGE) and 454-pyrosequencing approaches was used in the present study to describe the bacterial community structures present in the rhizospheres of native plants grown in three representative Chilean extreme environments: the Atacama Desert, the Andes Mountains, and the Antarctica.
Materials and Methods
Sampling
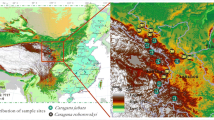
A total of 18 rhizosphere samples (three per plant species) were collected from three Chilean extreme environments (Fig. 1), including samples from the shrubs Atriplex sp. (Amaranthaceae family; ATA_A1, ATA_A2, and ATA_A3) and Stipa sp. (Poaceae family; ATA_B1, ATA_B2, and ATA_B3) grown in the Atacama Desert (ATA; 22° S, 68° W); herbaceous perennial shrubs Equisetum arvense (Equisetaceae family; AND_A1, AND_A2, and AND_A3) and Blechnum chilense (Blechnaceae family AND_B1, AND_B2, and AND_B3) grown in the Huilo-Huilo Ecological Reserve in the Andes Mountain (AND; 39° S, 71° W); and vascular plants C. quitensis (Caryophyllaceae family; ANT_A1, ANT_A2, and ANT_A3) and D. antarctica (Poaceae family; ANT_B1, ANT_B2, and ANT_B3) grown in Southern Shetland Islands from Antarctica (ANT; 62° S; 58° W). The selection of plant species in ATA and AND was based on their dominance at sampling sites, whereas only two vascular plants are present at the ANT sampling sites.
From 50 to 100 g of rhizosphere samples (including roots and adhering soil) were collected from random locations in plant rhizospheres using a clean spade to excavate intact roots from soil to a depth of 0 to 20 cm. Samples were transferred to sterile plastic 50-ml Falcon tubes and immediately transported to the laboratory on ice for soil and microbiological analyses. Rhizosphere soils were removed from roots by shaking, thoroughly mixed, and stored at −80 °C until analysis.
Chemical Properties of Rhizosphere Soils
Rhizosphere samples (~50 g) were subjected to chemical analysis to determine rhizosphere soil properties. Soil pH was measured in 1:2.5 soil/deionized water suspensions. Available phosphorus (POlsen) was extracted using 0.5 M Na-bicarbonate and analyzed using the molybdate blue method [38]. Organic matter was estimated by the Walkley-Black method [15]. Inorganic N was extracted with 2 M KCl and NO3 −-N and determined by the Devarda alloy distillation method [44]. Exchangeable cations of potassium (K+), calcium (Ca2+), magnesium (Mg2+), and sodium (Na+) were extracted with 1 M ammonium acetate at pH 7.0 and analyzed by flame atomic adsorption spectrophotometry (FAAS) [60]. Exchangeable aluminum (Al3+) was extracted with 1 M KCl and analyzed by FAAS [8].
DNA Extraction
Rhizosphere samples (1 to 2 g) were vortexed for 1 h with 2 mL of sodium phosphate buffer 0.1 M, pH 8, and centrifuged at 16,000×g × 10 min [24]. The supernatant was discarded, and pellet was subjected to cell disruption by bead-beating for 30 s with a Powerlyzer® 24 homogenizer (Mo-Bio Laboratories, CA, USA), and the solution was subjected to DNA purification using a Power Soil® DNA Isolation Kit (Mo-Bio Laboratories) according to the manufacturer’s instructions. Insufficient concentrations of DNA were obtained from ATA samples by this method, such that ATA samples were processed as follows: 6 g of rhizosphere soil were suspended in 10 ml of sterile distilled water and dispersed by sonication for 1 min at 130 W (20 kHz). The suspension was transferred to new sterile plastic tubes, centrifuged (13,400×g for 5 min), and the supernatant discarded. This procedure was repeated three times, and DNA was extracted from the resulting pellet by using the Power Soil® DNA Isolation Kit.
The quality and quantity of DNA extracts were measured using a microplate spectrophotometer (Multiskan GO, Thermo Fisher Scientific, Inc., MA, USA). The DNA purity was assessed by determination of the A280/A260 absorbance ratios, and only DNA extracts with absorbance ratios of ~1.8 were used for bacterial community analyses.
Denaturing Gradient Gel Electrophoresis
Bacterial community structures were evaluated by DGGE as follows. Partial 16S rRNA genes were amplified by PCR with DNA template concentrations of ~20 ng μL−1. Fragments of 16S rRNA genes (regions V6–V8) were amplified by hot start touchdown PCR using primers EUB f933-GC (5′-CGC CCG CCG CGC GCG GCG GGC GGG GCG GGG GCA CGG GGG GCA CAA GCG GTG GAG CAT GTG G’-3) and EUB r1387 (5′-GCC CGG GAA CGT ATT CAC CG-3′) designed by Iwamoto et al. [25]. The PCR conditions included a hot start of 95 °C for 10 min, followed by annealing at an initial temperature of 65 °C, followed by 20 cycles for 1 min each with a 0.5 °C decrease each cycle to 55 °C, and extension at 72 °C for 3 min. Then, ten additional cycles were carried out at 55 °C annealing and 94 °C denaturation for 1 min each, primer extension at 72 °C for 3 min, and a final extension step of 7 min at 72 °C.
The DGGE runs were performed using a DCode system (Bio-Rad Laboratories, Inc., CA, USA). Twenty microliters of PCR products (~500 ng μL−1) were loaded onto 6 % (w/v) polyacrylamide gels over a urea gradient of between 35 and 65 % (urea and formamide). Electrophoresis was run for 12 h at 100 V. The gels were then stained with SYBR Gold (Invitrogen™, Thermo Fisher Scientific, Inc.) for 30 min and photographed on a UV transilluminator (GelDoc-It®TS2 Imager, UVP, CA, USA). The DGGE image analysis of the band profiles was carried out using Phoretix 1D Pro Gel Analysis Software (TotalLab Ltd., Newcastle, UK; http://totallab.com/). Based on the matrix that was given by Phoretix 1D analysis, changes in the bacterial communities between rhizosphere samples were calculated by similarity profile analysis (SIMPROF test) with Bray-Curtis similarity index, 5 % significance level, and <0.1 stress values [12, 13], and visualized by non-metric multidimensional scaling (NMDS) analysis using Primer 6 software (Primer-E Ltd., Ivybridge, UK; http://www.primer-e.com/).
Dominant bands present in DGGE gels were chosen for sequencing to determine common and specific bacterial groups in rhizosphere samples. Dominant bands in the DGGE gels were excised, purified, and reamplified using the primers and cycling parameters described above and electrophoresed a second time in DGGE gels to avoid sequencing of multiple amplicons due to close proximity. PCR products were then sequenced by Macrogen, Inc. (Seoul, Korea). The sequences obtained were compared with those present in the GenBank database from the National Center for Biotechnology Information (NCBI) using megablast tool (http://blast.ncbi.nlm.nih.gov/) for taxonomic assignment (>70 % identity and 0.01 cutoff for E-values).
Four hundred fifty-four-pyrosequencing
The purified DNA samples were submitted to Macrogen, Inc. for 454-pyrosequencing. The 16S rRNA genes were amplified by PCR using the universal bacterial primer UNI_AMP-27 F (5′-Zxxx GAG TTT GAT CMT GGC TCA G-3′ and UNI_AMP-518R (5′-K WTT ACC GCG GCT GCT GG-3′) [31], where Z and K represent two pyrosequencing primers (CCA TCT CAT CCC TGC GTG TCT CCG ACT CAG and CCT ATC CCC TGTG TGC CTT GGC AGT CTC AG), and xxx was designed for the sample identification barcoding key. After a hot start at 95 °C for 3 min, PCR amplification was carried out for 35 cycles at 94 °C for 15 s, 55 °C for 45 s, and 72 °C for 1 min. A final extension step was carried out at 72 °C for 8 min. The 16S rRNA gene libraries were sequenced by with Roche 454 GS-FLX System using Titanium Chemistry (Roche Diagnostics Corporation, Life Sciences, Branford, CT, USA).
The sequences were analyzed by using Mothur software package [49], version 1.34.4 (http://www.mothur.org/), as described in previous reports [22, 29]. In brief, the low-quality sequences were filtered and barcode trimmed according to standard Mothur operational procedures (http://www.mothur.org/wiki/Schloss_SOP). The sequences were then aligned and chimeras filtered using standard procedures. The sequences were aligned against SILVA database [43] (release 119), using the kmer search tool to find template sequences and aligned by the Needleman-Wunsch algorithm. Sequences were assigned and clustered into OTUs at the 3 % dissimilarity cutoff, and the amount of sequences in each group was rarified according to the group with lowest quantity. Each OTU was assigned using a naive Bayesian classifier provided by the SILVA 16S rRNA database at a confidence threshold of 0.8. The sequence coverage was calculated using Good’s coverage [23]. Relative abundances, rarefaction curves and differences between bacterial communities were computed using the appropriate Mothur applications. Relative abundances were determined by dividing the number of reads for any given OTU for the total number of reads obtained. To determine the level of similarity between bacterial communities, a distance matrix was constructed using Bray-Curtis calculator and visualized as an NMDS plot. In addition, a color heat map was built using R software (https://www.r-project.org/) to visualize the similarities between rhizosphere communities assigned to class levels.
The sequences obtained in this study were deposited in NCBI Sequence Read Archive (http://trace.ncbi.nlm.nih.gov/Traces/sra/) under accession number SRP064983.
Results
Chemical Properties of Rhizosphere Soils
Large differences in soil chemical properties were observed between the rhizosphere soils (Table 1). Low pHs were observed in rhizosphere soils from AND and ANT, whereas alkaline pHs were observed in rhizosphere soils from the ATA. Higher P concentrations were found in the rhizosphere soils from ATA and ANT, compared with rhizosphere soils from AND. As expected, the organic matter contents were low in all rhizosphere soil samples, except in E. arvense rhizospheres from AND. The N contents were lower in the rhizosphere soils from AND and ATA, compared with rhizosphere soils from ANT.
Denaturing Gradient Gel Electrophoresis
The DGGE banding profiles of bacterial communities differed between the different rhizosphere soil samples (Fig. 2a). The majority of representative 16S rRNA gene sequences obtained from excised DGGE bands showed high similarities to members of the phyla Proteobacteria and Bacteroidetes (Table 2). In rhizosphere soils from ATA, the majority of excised DGGE bands showed high similarity with members of families Halomonadaceae, Pseudomonadaceae, and Moraxellaceae (Table 2; Fig. 2b). In AND, the majority of excised DGGE bands showed high similarity with “unclassified bacteria” (73~100 % identity); however, a few sequences were assigned as members of families Sphingobacteriaceae, Bradyrhizobiaceae, Bacillaceae, and Moraxellaceae. In ANT rhizosphere soils, most bands were assigned as members of Chitinophagaceae and Actinomycetaceae. A dendrogram based on image analysis of DGGE profiles showed three main clusters (0.6 distance value) according to extreme environments studied (Fig. 3a). Similar results were observed at 20 % similarity with NMDS analysis (Fig. 3b), which also revealed differences in bacterial community structures among plant species at 40 % similarity in AND and ANT samples but not for ATA rhizosphere soils.
a Denaturing gradient gel electrophoresis (DGGE) banding profiles of bacterial communities associated rhizosphere of plants grown in Chilean extreme environments (Atacama Desert, Andes Mountain, and Antarctica). Arrowheads and numbers indicate representative bands that were excised for DNA sequence analysis. Atriplex sp.: ATA_A1, ATA_A2, and ATA_A3; Stipa sp.: ATA_B1, ATA_B2, and ATA_B3; Equisetum arvense: AND_A1, AND_A2, and AND_A3; Blechnum chilense: AND_B1, AND_B2, and AND_B3; Colobanthus quitensis: ANT_A1, ANT_A2, and ANT_A3; Deschampsia antarctica: ANT_B1, ANT_B2, and ANT_B3. b Phylogenetic tree showing the affiliation of representative bands that were excised from DGGE profile gel. The neighbor-joining tree was constructed using MEGA6 software (http://www.megasoftware.net/), and a bootstrap analysis was performed with 1000 trials. The bar indicates 5 % sequence divergence
a Dendrogram of DGGE profiles generated by Phoretix 1D Pro Gel Analysis Software (http://totallab.com/). b Non-metric multidimensional scaling (NMDS) analysis of DGGE profiles generated by Primer 6 software (http://www.primer-e.com/) with the Bray-Curtis similarity index, 5 % significance level, and <0.1 stress values [12, 13]. Atriplex sp.: ATA_A1, ATA_A2, and ATA_A3; Stipa sp.: ATA_B1, ATA_B2, and ATA_B3; Equisetum arvense: AND_A1, AND_A2, and AND_A3; Blechnum chilense: AND_B1, AND_B2, and AND_B3; Colobanthus quitensis: ANT_A1, ANT_A2, and ANT_A3; Deschampsia antarctica: ANT_B1, ANT_B2, and ANT_B3
Four hundred fifty-four-pyrosequencing
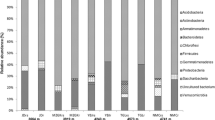
Four hundred fifty-four-pyrosequencing data analysis also revealed differences between extreme environments studied. Higher sequence reads and OTUs were observed in ANT rhizosphere soils (8150 to 16,320 reads and 2829 to 4905 OTUs) followed by AND (5414 to 10,960 reads and 2041 to 3698 OTUs) and ATA (7832 to 9578 reads and 195 to 1663 OTUs) rhizosphere soils (Table 3). Proteobacteria were the dominant taxa in all rhizosphere soil samples, ranging between 34 to 54, 50 to 57, and 22 to 96 % for ANT, AND, and ATA, respectively (Fig. 4a); followed by Actinobacteria phyla, ranging between 12 to 40, 2 to 23, and 7 to 17 % for ANT, ATA, and AND, respectively; and Bacteroidetes phyla, ranging between 3 to 1, <1 to 38, and 3 to 10 % for ANT, ATA, and AND, respectively. It is noteworthy that Acidobacteria was also one of the dominant phyla in ANT (3~10 %) and AND (9~19 %) rhizosphere soils, but low relative abundance (0 to 0.01 %) was observed in ATA rhizosphere soils. At the class level (85 % similarity), the relative abundance of bacterial groups clearly differed between the sites. Gammaproteobacteria was the dominant class (11 to 92 %) in the rhizosphere soils from ATA (Fig. 4b). In contrast, Alphaproteobacteria (25 to 43 %) and Betaproteobacteria (8 to 18 %) were the dominant classes in the rhizosphere soils from AND, whereas, Actinobacteria (10 to 31 %), Alphaproteobacteria (12 to 22 %), Betaproteobacteria (3 to 21 %), and Gammaproteobacteria (2 to 31 %) were the dominant classes in the rhizosphere soils from ANT. At the genus level (90 % of similarity), most sequences were classified as “others” (67 to 93 %) and Burkholderia (2 to 20 %) in the majority of rhizosphere soil samples. It is noteworthy that 4 of 6 samples from ATA showed a high abundance relative of Stenotrophomonas (56 to 84 %).
Relative abundances in percent of the total bacterial community at the phyla (a) and class (b) level in the rhizosphere soil samples from plants grown in the Atacama Desert, Andes Mountain, and Antarctica. Sequences that could not be classified into any known group are assigned as “unclassified,” and sequences of which the abundances are too low are assigned as “others”. Atriplex sp.: ATA_A1, ATA_A2, and ATA_A3; Stipa sp.: ATA_B1, ATA_B2, and ATA_B3; Equisetum arvense: AND_A1, AND_A2, and AND_A3; Blechnum chilense: AND_B1, AND_B2, and AND_B3; Colobanthus quitensis: ANT_A1, ANT_A2, and ANT_A3; Deschampsia antarctica: ANT_B1, ANT_B2, and ANT_B3
Similarity analyses between bacterial community structures visualized by NMDS are shown in Fig. 5a. Similar to the NMDS analysis from DGGE banding profiles, the NMDS plot from pyrosequence data showed three groups according to the environments studied. Also, the bacterial communities were separately grouped among both plant species, except in ATA rhizosphere soils. The heat map confirms our observations, showing the three clusters and revealing that differences in bacterial communities are mainly attributed to Gammaproteobacteria, Alphaproteobacteria, Acidobacteria, Betaproteobacteria, Actinobacteria, unclassified bacteria, and others (Fig. 5b).
a NMDS analysis of bacterial communities generated by Mothur package (http://www.mothur.org/) with the Bray-Curtis similarity index. b Heat map of the bacterial communities at the class level generated by R software (https://www.r-project.org/). Atriplex sp.: ATA_A1, ATA_A2, and ATA_A3; Stipa sp.: ATA_B1, ATA_B2, and ATA_B3; Equisetum arvense: AND_A1, AND_A2, and AND_A3; Blechnum chilense: AND_B1, AND_B2, and AND_B3; Colobanthus quitensis: ANT_A1, ANT_A2, and ANT_A3; Deschampsia antarctica: ANT_B1, ANT_B2, and ANT_B3
Discussion
Both 16S rRNA-based approaches showed differences in bacterial community structures between rhizosphere soil samples from native plants grown in studied Chilean extreme environments. Bacterial communities in the rhizospheres of plants differ in their abundances, compositions, and activities according to diverse abiotic and biotic factors. Among abiotic factors, the shifts in rhizosphere bacterial communities have commonly been related to soil type [28, 50]. A continental-scale study reported a large-scale biogeography pattern of soil microorganisms, mainly attributing the differences in diversity and richness of soil bacterial communities to soil pH under natural conditions [20]. Soil properties (soil pH and carbon content) have also been correlated with changes in the diversities of rhizosphere bacterial communities in extreme environments, such as arid deserts [2]. Soil metagenomic analyses of pristine environments also suggest the effects of ecosystem type, such as soil pH and altitude, on composition, diversity, and biosynthesis potential of soil microorganisms [37]. Our results showed a great variation in chemical properties between the studied rhizosphere soil samples, particularly in pH, available P, and organic matter contents. A recent study also suggest that elevated nutrient (nitrogen and phosphorus) inputs induce predictable shifts in the taxonomy and functionality of soil microbial communities impacting belowground ecosystems worldwide [32]. In Arctic soils, warming-induced shifts in the structures of microbial communities have been observed [18], whereas, variations in the distribution and structures of bacterial communities have been attributed to K, C, Ca, and moisture contents in Antarctic soils [53]. In this context, based on the diversity and distribution of soil bacterial taxa across an Antarctic mountainside (environment with low complexity and oligotrophic soils), Okie et al. [41] predicted that (i) lower temperatures should reduce taxon niche widths due to slower growth rates; (ii) the changing niche widths should lead to contrasts between alpha- and beta-diversity; and (iii) beta-diversity increases with elevation and decreases with temperature. In addition, this theory also predicted a relationship between alpha-diversity and soil pH, and a negative relationship between alpha-diversity and soil salinity.
Our analysis also revealed differences in bacterial community structures in the rhizospheres between plant species, even in plant species grown in the same extreme environment, particularly in AND and ANT. Changes in the structures of rhizosphere bacterial communities have mainly been attributed to plant species or genotypes [42, 51]. Under natural vegetation, the variations in soil microbial community structures between and within plant species have been strongly correlated with plant identity, followed by soil chemistry, spatial location, and plant genus [1, 10]. Similarly, a field-scale study showed a selective effect of plant species (grasses and legumes) on soil Streptomyces assemblages in an ecological protected reserve [4]. Plant species-specific selection of rhizosphere bacterial communities has also been observed in extreme environments, such as high-altitude alpine ecosystem characterized by oligotrophic and harsh environmental conditions [11]. However, the changes in microbial community structures were not only attributed to plant species but also a combined effect of plant species and soil fractions (rhizosphere or bulk soil) in cold hostile climates with poorly developed soils [36]. In contrast to our findings, a study on rhizosphere soils showed similar bacterial diversity patterns between the same Antarctic vascular plants (D. Antarctica and C. quitensis) from different locations [56].
In relation to bacterial community composition, the sequencing of excised DGGE bands showed the occurrence of members of the phyla Proteobacteria and Bacteroidetes. By conventional 16S rRNA-based approaches (DGGE, automated ribosomal intergenic spacer analysis [ARISA], and cloning), the occurrence of Proteobacteria and Bacteroidetes in rhizosphere and bulk soils from high mountains, Arctic, and Antarctic environments have previously been reported [11, 18, 53]. In this context, the presence of Proteobacteria and Bacteroidetes have also been recently reported in the rhizosphere and bulk soils from Chilean extreme environments, such as Atacama Desert and Andes Mountain, as revealed by DGGE [3, 27]. Similar to our results in ANT rhizosphere soils, Actinobacteria have also been found in soils from Arctic and Antarctic environments [18, 53]. It is noteworthy that several sequenced DGGE bands from AND were phylogenetically assigned as unclassified bacteria. Recently, Saleem et al. [48] reported that plant roots (Nicotania tabacum) select rare bacterial rhizosphere taxa as dominant rhizosphere microbiome colonizers.
In general, representatives of the Proteobacteria, Actinobacteria, and Bacteroidetes phyla were the dominant taxa in rhizosphere soil samples. Through high-throughput sequencing (HTS) of 16S rRNA genes, the same taxa have been reported as common inhabitants in rhizosphere microbiomes under natural conditions [1, 4]. In Antarctic environments, Proteobacteria, Actinobacteria, and Bacteroidetes were also dominant phyla in bulk soils determined by pyrosequencing [47, 57]. In this context, the Firmicutes were reported as the most abundant phylum (~40 %) in the rhizosphere soils of Antarctic vascular plants [56]; however, Firmicutes only represent a relative abundance of 3 to 10 % in rhizosphere soils from the Antarctic in our study. In relation to desert environments, Proteobacteria and Actinobacteria were identified as the dominant bacterial groups in rhizosphere and bulk soils of the Sonoran Desert by a recent pyrosequencing project [2]. In the Atacama Desert, the phylum-level composition of soil bacterial communities is dominated by Actinobacteria and Chloroflexi, followed by the Acidobacteria and Proteobacteria [17, 40]. It is necessary to mention that a different protocol was used to extract total DNA from ATA samples; therefore, we cannot discard possible selectivity in releasing microbial cells of different bacterial taxa from ATA soil samples compared with AND and ANT, because it has been reported that wide differences in total extractable DNA as derived by the use of different extraction protocols for the same soil sample [24, 59]. However, our previous study showed similar dominant bacterial taxa (Acidobacteria and Proteobacteria) present in rhizosphere soil samples from the Atacama Desert [27].
At the class level, greater differences between bacterial communities were found between rhizosphere soils from ATA compared with rhizosphere soils from AND and ANT. These differences could be driven by the lower numbers of OTUs observed in ATA combined with a higher relative abundance of certain bacterial classes, such as Gammaproteobacteria (particularly Stenotrophomonas spp.). Atacama Desert is characterized by low species richness (69 to 330 OTUs g−1) [17, 39], whereas Stenotrophomonas spp. are found in a wide variety of environments, and particularly, in close association with plants [46]. Some Stenotrophomonas strains can produce antimicrobial compounds that protect plants against opportunist phytopathogen attacks, as well as generate factors (auxins) that can promote plant growth. In this sense, Alphaproteobacteria was the dominant class in the rhizosphere soils from AND and ANT. Alphaproteobacteria contain diverse symbionts of plants, many of which fix atmospheric nitrogen (N2) (e.g., Rhizobium, Bradyrhizobium, Mesorhizobium, Sinorhizobium). At the genus level, Burkholderia (a Betaproteobacteria member) was the dominant group in AND and ANT. Burkholderia spp. are also widely distributed in the environment, especially in the rhizospheres of plants, and some may act as plant growth promoting rhizobacteria (PGPR), such as legume symbionts (beta-rhizobia), and those involved in phytopathogen biocontrol and as bioremediation agents [55, 58]. Native Brazilian Mimosa spp. grown in sub-tropical regions with acidic soils are primarily nodulated by Burkholderia strains [30, 45], whereas, native Indian Mimosa spp. grown in arid regions with alkaline soils are primarily nodulated by Sinorhizobium strains; therefore, local soil factors may play an important role in the occurrence of bacteria nodulating native legumes [21]. Moreover, the presence of Burkholderia spp. in rhizosphere soils from AND and ANT is not surprising because this bacterial genus is known to be highly tolerant of acidic soils local soil [54, 55]. It is known that bacteria in the rhizosphere help plant to tolerate environmental stresses, such as drought and soil salinity [61]. PGPR are commonly found in the rhizospheres of plants in arid environments, contributing to the ability to maintain plant functional homeostasis despite environmental pressures [5, 19, 26, 34]. Studies have also suggested that microbial phyla, particularly Proteobacteria and Cyanobacteria, may significantly contribute to the total carbon and nitrogen fixed in Antarctic desert soils [16]. Putative PGPR have also been isolated from Chilean extreme environments [27].
Based on our results, we conclude that specific bacterial community structures in the rhizospheres of native plants in Chilean extreme environments are governed by a combination of local soil conditions and plant identity, and native plants may attract, select, and conserve specific bacterial groups with multiple plant growth promoting traits to sustain the growth and tolerance to local harsh conditions. However, further studies are required to validate this statement.
Conclusions
Both molecular approaches showed differences in rhizobacterial community structures between native plants grown in studied Chilean extreme environments. Our results also revealed differences in rhizobacterial community structures between plant species, even in plant species grown in the same extreme environment, particularly in AND and ANT. In relation to rhizobacterial community composition, Proteobacteria, Actinobacteria, and Bacteroidetes phyla were the dominant taxa in all rhizosphere soil samples, as revealed by both molecular methods. Rhizosphere soils from ATA harbored fewer OTUs than rhizosphere soils from AND and ANT. Alphaproteobacteria was the dominant class in the rhizosphere soils from AND and ANT, whereas, Gammaproteobacteria was the dominant class in the rhizosphere soils from ATA. At the genus level, Burkholderia was the dominant genus in AND and ANT, and Stenotrophomonas was the dominant genus in ATA. Further studies are needed to elucidate which environmental factors (e.g., soil properties such as pH and salinity) regulate the diversity and structure of bacteria in the rhizosphere, and how (or if) specific bacterial groups affect the growth, survival, and tolerance of native plants in Chilean extreme environments.
References
Aleklett K, Leff JW, Fierer N, Hart M (2015) Wild plant species growing closely connected in a subalpine meadow host distinct root-associated bacterial communities. PeerJ 3:e804. doi:10.7717/peerj.804
Andrew DR, Fitak RR, Munguia-Vega A, Racolta A, Martinson VG, Dontsovag K (2012) Abiotic factors shape microbial diversity in Sonoran Desert soils. Appl Environ Microbiol 78:7527–7537
Azua-Bustos A, Caro-Lara L, Vicuña R (2015) Discovery and microbial content of the driest site of the hyperarid Atacama Desert, Chile. Environ Microbiol Rep. doi:10.1111/1758-2229.12261
Bakker MG, Bradeen JM, Kinkel LL (2013) Effects of plant host species and plant community richness on streptomycete community structure. FEMS Microbiol Ecol 83:596–606
Basil AJ, Strap JL, Knotek-Smith HM, Crawford DL (2004) Studies on the microbial populations of the rhizosphere of big sagebrush (Artemisia tridentata). J Ind Microbiol Biotechnol 31:278–288
Berendsen RL, Pieterse CMJ, Bakker PA (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17:478–486
Berg G, Grube M, Schloter M, Smalla K (2014) Unraveling the plant microbiome: looking back and future perspectives. Front Microbiol 5:148. doi:10.3389/fmicb.2014.00148
Bertsch PM, Bloom PR (1996) Aluminum. In: Bigham JM (ed) Methods of Soil Analysis, Part 3—Chemical Methods. Soil Science Society of America, Madison, pp 526–527
Bockheim JG (2015) The Soils of Antarctica. Springer International Publishing, Switzerland. doi:10.1007/978-3-319-05497-1, 273 pp
Burns JH, Anacker BL, Strauss SY, Burke DJ (2015) Soil microbial community variation correlates most strongly with plant species identity, followed by soil chemistry, spatial location and plant genus. AoB Plants 7:plv030. doi:10.1093/aobpla/plv030
Ciccazzo S, Esposito A, Rolli E, Zerbe S, Daffonchio D, Brusetti L (2014) Different pioneer plant species select specific rhizosphere bacterial communities in a high mountain environment. SpringerPlus 3:391. doi:10.1186/2193-1801-3-391
Clarke KR (1993) Non-parametric multivariate analyses of changes in community structure. Aust J Ecol 18:117–143
Clarke KR, Somerfield PJ, Gorley RN (2008) Testing null hypotheses in exploratory community analyses: similarity profiles and biota-environmental linkage. J Exp Mar Biol Ecol 366:56–69
Cleary DFR, Smalla K, Mendonça-Hagler LCS, Gomes NCM (2012) Assessment of variation in bacterial composition among microhabitats in a mangrove environment using DGGE fingerprints and barcoded pyrosequencing. PLoS ONE 7(1):e29380. doi:10.1371/journal.pone.0029380
Combs SM, Nathan MV (1998) Soil organic matter. In: Brown JR (ed) Recommended Chemical Soil Test Procedures for the North Central Region. NCR Publication No. 221. Missouri Agricultural Experiment Station, Columbia, pp 31–33
Cowan DA, Makhalanyane TP, Dennis PG, Hopkins DW (2014) Microbial ecology and biogeochemistry of continental Antarctic soils. Front Microbiol 5:154. doi:10.3389/fmicb.2014.00154
Crits-Christoph A, Robinson CK, Barnum T, Fricke WF, Davila AF, Jedynak B, McKay CP, DiRuggiero J (2013) Colonization patterns of soil microbial communities in the Atacama Desert. Microbiome 1:28. doi:10.1186/2049-2618-1-28
Deslippe JR, Hartmann M, Simard SW, Mohn WW (2012) Long-term warming alters the composition of Arctic soil microbial communities. FEMS Microbiol Ecol 82:303–315
Ferjani R, Marasco R, Rolli E, Cherif H, Cherif A, Gtari M, Boudabous A, Daffonchio D, Ouzari H-I (2015) The date palm tree rhizosphere is a niche for plant growth promoting bacteria in the oasis ecosystem. BioMed Res Int 2015:153851. doi:10.1155/2015/153851
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci U S A 103:626–631
Gehlot HS, Panwar D, Tak N, Tak A, Sankhla IS, Poonar N, Parihar R, Shekhawat NS, Kumar M, Tiwari R, Ardley J, James EK, Sprent JI (2012) Nodulation of legumes from the Thar desert of India and molecular characterization of their rhizobia. Plant Soil 357:227–243
Goda A, Maruyama F, Michi Y, Nakagawa I, Harada K (2014) Analysis of the factors affecting the formation of the microbiome associated with chronic osteomyelitis of the jaw. Clin Microbiol Infect 20:O309–O317
Good IJ (1953) The population frequencies of species and the estimation of population parameters. Biometrika 40:237–264
He J, Xu Z, Hughes J (2005) Pre-lysis washing improves DNA extraction from a forest soil. Soil Biol Biochem 37:2337–2341
Iwamoto T, Tani K, Nakamura K, Suzuki Y, Kitagawa M, Eguchi M, Nasu M (2000) Monitoring impact of in situ biostimulation treatment on ground-water bacterial community by DGGE. FEMS Microbiol Ecol 32:129–141
Jorquera MA, Shaharoona B, Nadeem SM, Mora ML, Crowley DE (2012) Plant growth-promoting rhizobacteria associated with ancient clones of creosote bush (Larrea tridentata) 64: 1008–1017
Jorquera MA, Inostroza NG, Lagos LM, Barra PJ, Marileo LG, Rilling JI, Campos DC, Crowley DE, Richardson AE, Mora ML (2014) Bacterial community structure and detection of putative plant growth-promoting rhizobacteria associated with plants grown in Chilean agro-ecosystems and undisturbed ecosystems. Biol Fertil Soils 50:1141–1153
Kuramae EE, Yergeau E, Wong LC, Pij AS, van Veen JA, Kowalchuk GA (2012) Soil characteristics more strongly influence soil bacterial communities than land-use type. FEMS Microbiol Ecol 79:12–24
Lagos LM, Navarrete OU, Maruyama F, Crowley DE, Cid FP, Mora ML, Jorquera MA (2014) Bacterial community structures in rhizosphere microsites of ryegrass (Lolium perenne var. Nui) as revealed by pyrosequencing. Biol Fertil Soils 50:1253–1266
Lammel DR, Cruz LM, Carrer H, Cardoso EJBN (2013) Diversity and symbiotic effectiveness of beta-rhizobia isolated from sub-tropical legumes of a Brazilian Araucaria Forest. World J Microbiol Biotechnol 29:2335–2342
Lee TK, Van Doan T, Yoo K, Choi S, Kim C, Park J (2010) Discovery of commonly existing anode biofilm microbes in two different wastewater treatment MFCs using FLX Titanium pyrosequencing. Appl Microbiol Biotechnol 87:2335–2343
Leff JW, Jones SE, Prober SM, Barberán A, Borer ET, Firn JL, Harpole WS, Hobbie SE, Hofmockel KS, Knops JM, McCulley RL, La Pierre K, Risch AC, Seabloom EW, Schütz M, Steenbock C, Stevens CJ, Fierer N (2015) Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe. Proc Natl Acad Sci U S A 112:10967–10972
Maier RM, Drees KP, Neilson JW, Henderson DA, Quade J, Betancourt JL, Navarro-González R, Rainey FA, McKay CP (2010) Microbial life in the Atacama Desert. Science 306:1289–1290
Marasco R, Rolli E, Ettoumi B, Vigani G, Mapelli F, Borin S, Abou-Hadid AF, El-Behairy UA, Sorlini C, Cherif A, Zocchi G, Daffonchio D (2012) A drought resistance-promoting microbiome is selected by root system under desert farming. PLoS ONE 7:e48479. doi:10.1371/journal.pone.0048479
Marzorati M, Wittebolle L, Boon N, Daffonchio D, Verstraete W (2008) How to get more out of molecular fingerprints: practical tools for microbial ecology. Environ Microbiol 10:1571–1581
Massaccesi L, Benucci GMN, Gigliotti G, Cocco S, Corti G, Agnelli A (2015) Rhizosphere effect of three plant species of environment under periglacial conditions (Majella Massif, central Italy). Soil Biol Biochem 89:184–195
McCarthy CB, Colman DI (2015) Soil metagenomes from different pristine environments of northwest Argentina. Genome Announc 3:e00926–15
Murphy J, Riley JP (1962) A modified single solution method for the determination of phosphate in natural waters. Anal Chim Acta 27:31–36
Navarro-González R, Rainey FA, Molina P, Bagaley DR, Hollen BJ, de la Rosa J, Small AM, Quinn RC, Grunthaner FJ, Cáceres L, Gomez-Silva B, McKay CP (2003) Mars-like soils in the Atacama Desert, Chile, and the dry limit of microbial life. Science 302:1018–1021
Neilson JW, Quade J, Ortiz M, Nelson WM, Legatzki A, Tian F, LaComb M, Betancourt JL, Wing RA, Soderlund CA, Maier RM (2012) Life at the hyperarid margin: novel bacterial diversity in arid soils of the Atacama Desert, Chile. Extremophiles 16:553–566
Okie JG, Van Horn DJ, Storch D, Barrett JE, Gooseff MN, Kopsova L, Takacs-Vesbach CD (2015) Niche and metabolic principles explain patterns of diversity and distribution: theory and a case study with soil bacterial communities. Proc R Soc B 282:20142630. doi:10.1098/rspb.2014.2630
Peiffer JA, Spor A, Koren O, Jin Z, Tringe SG, Dangl JL, Buckler ES, Ley RE (2013) Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci U S A 110:6548–6553
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig WG, Peplies J, Glockner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35:7188–7196
Radojević M, Bashkin VN (1999) Practical Environmental Analysis. The Royal Society of Chemistry, Cambridge, 464 pp
Reis FB, Simon MF, Gross E, Boddey RM, Elliott GN, Neto NE, Loureiro MF, Queiroz LP, Scotti MR, Chen WM, Norén A, Rubio MC, de Faria SM, Bontemps C, Goi SR, Young JPW, Sprent JI, James EK (2010) Nodulation and nitrogen fixation by Mimosa spp. in the Cerrado and Caatinga biomes of Brazil. New Phytol 186:934–946
Ryan RP, Monchy S, Cardinale M, Taghavi S, Crossman L, Avison MB, Berg G, van der Lelie D, Dow JM (2009) The versatility and adaptation of bacteria from the genus Stenotrophomonas. Nat Rev Microbiol 7:514–525
Roesch LFW, Fulthorpe RR, Pereira AB, Pereira CK, Lemos LN, Barbosa AD, Suleiman AKA, Gerber AL, Pereira MG, Loss A, da Costa EM (2012) Soil bacterial community abundance and diversity in ice-free areas of Keller Peninsula, Antarctica. Appl Soil Ecol 61:7–15
Saleem M, Law AD, Moe LA (2015) Nicotiana roots recruit rare rhizosphere taxa as major root-inhabiting microbes. Microb Ecol 71:469–472
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing Mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Schreiter S, Ding G-C, Heuer H, Neumann G, Sandmann M, Grosch R, Kropf S, Smalla K (2014) Effect of the soil type on the microbiome in the rhizosphere of field-grown lettuce. Front Microbiol 5:144. doi:10.3389/fmicb.2014.00144
Schweitzer JA, Bailey JK, Fischer DG, LeRoy CJ, Lonsdorf EV, Whitham TG, Hart SC (2008) Plant–soil–microorganism interactions: heritable relationship between plant genotype and associated soil microorganisms. Ecology 89:773–781
Smalla K, Oros-Sichler M, Milling A, Heuer H, Baumgarte S, Becker R, Neuber G, Kropf S, Ulrich A, Tebbe CC (2007) Bacterial diversity of soils assessed by DGGE, T-RFLP and SSCP fingerprints of PCR-amplified 16S rRNA gene fragments: do the different methods provide similar results? J Microbiol Methods 69:470–479
Stomeo F, Makhalanyane TP, Valverde A, Pointing SB, Stevens MI, Cary CS, Tuffin MI, Cowan DA (2012) Abiotic factors influence microbial diversity in permanently cold soil horizons of a maritime-associated Antarctic Dry Valley. FEMS Microbiol Ecol 82:326–340
Stopnisek N, Bodenhausen N, Frey B, Fierer N, Eberl L, Weisskopf L (2014) Genus-wide acid tolerance accounts for the biogeographical distribution of soil Burkholderia populations. Environ Microbiol 16:1503–1512
Suarez-Moreno ZR, Caballero-Mellado J, Coutinho BG, Mendonça-Previato L, James EK, Venturi V (2012) Common features of environmental and potentially beneficial plant-associated Burkholderia. Microb Ecol 63:249–266
Teixeira LCRS, Peixoto RS, Cury JC, Sul WJ, Pellizari VH, Tiedje J, Rosado AS (2010) Bacterial diversity in rhizosphere soil from Antarctic vascular plants of Admiralty Bay, maritime Antarctica. ISME J 4:989–1001
Tytgat B, Verleyen E, Obbels D, Peeters K, De Wever A, D’hondt S, De Meyer T, Van Criekinge W, Vyverman W, Willems A (2014) Bacterial diversity assessment in Antarctic terrestrial and aquatic microbial mats: a comparison between bidirectional pyrosequencing and cultivation. PLoS One 9:e97564. doi:10.1371/journal.pone.0097564
Vial L, Chapalain A, Groleau M-C, Déziel E (2011) The various lifestyles of the Burkholderia cepacia complex species: a tribute to adaptation. Environ Microbiol 13:1–12
Wagner AO, Praeg N, Reitschuler C, Illmer P (2015) Effect of DNA extraction procedure, repeated extraction and ethidium monoazide (EMA)/propidium monoazide (PMA) treatment on overall DNA yield and impact on microbial fingerprints for bacteria, fungi and archaea in a reference soil. Appl Soil Ecol 93:56–64
Warncke D, Brown JR (1998) Potassium and other basic cations. In: Brown JR (ed) Recommended Chemical Soil Test Procedures for the North Central Region. NCR Publication No. 221. Missouri Agricultural Experiment Station, Columbia, pp 31–33
Yang J, Kloepper JW, Ryu CM (2009) Rhizosphere bacteria help plants tolerate abiotic stress. Trends Plant Sci 14:1–4
Acknowledgments
The authors wish to thank the two anonymous referees for their helpful and constructive criticism. This study was financed by International Cooperation Projects Conicyt-USA (code USA2013-0010), Conicyt-MEC (no. 80140015), and Fondecyt no. 1120505. J.J. Acuña thanks the Fondecyt Postdoctoral Project (no. 3140620).
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article can be found at http://dx.doi.org/10.1007/s00248-016-0830-9.
Rights and permissions
About this article
Cite this article
Jorquera, M.A., Maruyama, F., Ogram, A.V. et al. Rhizobacterial Community Structures Associated with Native Plants Grown in Chilean Extreme Environments. Microb Ecol 72, 633–646 (2016). https://doi.org/10.1007/s00248-016-0813-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-016-0813-x