Abstract
Alternative oxidases (AOXs) are mitochondrial cyanide-resistant membrane-bound metallo-proteins catalyzing the oxidation of ubiquinol and the reduction of oxygen to water bypassing two sites of proton pumping, thus dissipating a major part of redox energy into heat. Here, the structure of Arabidopsis thaliana AOX 1A has been modeled using the crystal structure of Trypanosoma brucei AOX as a template. Analysis of this model and multiple sequence alignment of members of the AOX family from all kingdoms of Life indicate that AOXs display a high degree of conservation of the catalytic core, which is formed by a four-α-helix bundle, hosting the di-iron catalytic site, and is flanked by two additional α-helices anchoring the protein to the membrane. Plant AOXs display a peculiar covalent dimerization mode due to the conservation in the N-terminal region of a Cys residue forming the inter-monomer disulfide bond. The multiple sequence alignment has also been used to infer a phylogenetic tree of AOXs whose analysis shows a polyphyletic origin for the AOXs found in Fungi and a monophyletic origin of the AOXs of Eubacteria, Mycetozoa, Euglenozoa, Metazoa, and Land Plants. This suggests that AOXs evolved from a common ancestral protein in each of these kingdoms. Within the Plant AOX clade, the AOXs of monocotyledon plants form two distinct clades which have unresolved relationships relative to the monophyletic clade of the AOXs of dicotyledonous plants. This reflects the sequence divergence of the N-terminal region, probably due to a low selective pressure for sequence conservation linked to the covalent homo-dimerization mode.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alternative oxidases (AOXs) are mitochondrial cyanide—insensitive membrane-bound proteins involved in redox reactions. These metallo-proteins, recalling the structural organization of members of the “ferritin” family, display a four-α-helix bundle, a common structural element for binuclear metal centers. In general, the two iron atoms are ligated by four conserved Glu or Asp residues and two His side chains, which form two universally conserved Glu–X–X–His motifs (Siedow et al. 1995; Berthold et al. 2000; McDonald 2008; Albury et al. 2009; Moore et al. 2013; Neimanis et al. 2013).
AOX branches from the cytochrome pathway and catalyzes the oxidation of ubiquinol and the four-electron reduction of oxygen to water. In contrast to cytochrome c oxidase, the main mitochondrial terminal oxidase, AOX does not pump protons. Bypassing two sites of proton pumping, at complexes III and IV, AOX activity dissipates a major part of the redox energy into heat and thus results in a lower synthesis of ATP (Vanlerberghe and McIntosh 1997; Berthold et al. 2000; Millenaar and Lambers 2003; McDonald and Vanlerberghe 2004; Rasmusson et al. 2008; McDonald et al. 2009; Moore et al. 2013; Vishwakarma et al. 2014).
Although AOX is an ubiquitous enzyme within the Plant kingdom, it is also found in many other Eukaryote kingdoms, including fungi, protists, and few animal species, as well as Bacteria but not Archaea (Chaudhuri and Hill 1996; Van Hellemond et al. 1998; Ajayi et al. 2002; Stenmark and Nordlund 2003; Veiga et al. 2003; Roberts et al. 2004; Stechmann et al. 2008; Albury et al. 2009; McDonald et al. 2009). AOX is encoded by a small gene family, consisting of two distinct subfamilies termed AOX1 and AOX2. In the model plant Arabidopsis thaliana, AOX is encoded by five genes, namely AOX1a, AOX1b, AOX1c, AOX1d, and AOX2 (Supplementary Fig. 1) (Clifton et al. 2006). The expression of AOX is primarily induced when the mitochondrial respiration is impaired. Moreover, the expression of AOX gene(s) is tissue-specific and evolutionarily regulated (Thirkettle-Watts et al. 2003). AOX seems to play a role in controlling the levels of reactive oxygen species (ROS) generated by the respiratory chain (Purvis and Shewfelt 1993). In fact, AOX could act to reduce ROS generation and could prevent the over-reduction of the ubiquinone pool, not only in plants but also in fungi and lower invertebrates (Maxwell et al. 1999; Parsons et al. 1999; McDonald et al. 2009; Van Aken et al. 2009; Li et al. 2011). Then, the functions of AOX include thermogenesis, stress tolerance, and the maintenance of mitochondrial and cellular homeostasis (Finnegan et al. 2004; Vanlerberghe 2013).
Two plant AOX molecular models have been built (Umbach and Siedow 1993; Moore et al. 1995; Siedow et al. 1995; Andersson and Nordlund 1999; Berthold et al. 2000) and the crystal structure of the Trypanosoma brucei AOX (TAO) has been solved (Shiba et al. 2013). However, the plant AOX molecular models are conflicting.
In the first model published in 1995, AOX was postulated to show a covalent homo-dimeric assembly through the occurrence of a disulfide bridge, each monomer displaying a hydroxo-bridged di-iron center residing within a four-α-helix bundle. The two di-iron binding Glu–X–X–His motifs were predicted to be located on the first and fourth α-helix of the bundle. Moreover, it was proposed that AOX is anchored to the mitochondrial membrane by two transmembrane α-helices (Umbach and Siedow 1993; Moore et al. 1995; Siedow et al. 1995). The second model published in 1999 predicted a different length of the α-helices and postulated that the Glu–X–X–His motifs involved in di-iron binding were located on the second and the fourth α-helix of the bundle. A putative membrane-binding motif formed by two short hydrophobic helices, which would insert in a single leaflet of the lipid bilayer, was also proposed (Andersson and Nordlund 1999; Berthold et al. 2000).
Very recently, the crystal structure of the Trypanosoma brucei alternative oxidase (TAO) has been solved (Shiba et al. 2013). The catalytic center is formed by two His residues, each coordinating one iron atom on the same side of the bi-metal cluster, two Glu residues connecting the two ferric ions, and two additional Glu residues each coordinating one iron ion. In addition to the di-iron active site, TAO contains a hydrophobic cavity for ubiquinol binding. TAO is a non-covalent homo-dimer, the N-terminal region of each monomer being pivotal for assembly in a domain-swapping fashion (Shiba et al. 2013).
The availability of the TAO crystal structure provides for the first time the opportunity to analyze the structural details of AOXs from different organisms through homology modeling. In addition, structural phylogenomic analyses can provide important insights into AOX function and molecular evolution. Here, the structural model of Arabidopsis thaliana AOX 1A (AtAOX), built by a combination of ab initio/threading strategy is reported. Furthermore, in order to put the available structural information in an evolutionary context, 111 protein sequences homologous to A. thaliana AOX 1A were collected and assembled in a multiple amino acid sequence alignment. This allowed to map the conservation of residues essential for AOXs structure and function along the evolutionary ladder and to infer a phylogenetic framework for the structural evolution of AOXs.
Results and Discussion
The Three-Dimensional Model of AOX 1A
Plant AOXs contain a mitochondrial targeting pre-sequence (residues 1–62), which is cleaved after translocation of the protein on the mitochondrial membrane (Tanudji et al. 1999). We thus removed the first 62 residues of the AtAOX amino acid sequence and used the mature sequence to build the structural model shown in Fig. 1.
Analysis of the model shows that the catalytic core of AtAOX is formed by a four-α-helix bundle. As expected, likewise TAO, the AtAOX active site is located in a hydrophobic cavity and contains four strictly conserved Glu residues at positions 183, 222, 273, and 324, which coordinate the two iron atoms. The stability of the di-iron center is also due to the presence of two strictly conserved His residues at positions 225 and 327 (Fig. 2). All these conserved residues form the two Glu–X–X–His motifs (i.e., Glu222–X–X–His225 and Glu324–X–X–His327), located on the second and the fourth helix of the four α-helix bundle, as previously predicted (Andersson and Nordlund 1999). The two His residues are too far away from the di-iron center to coordinate the two metal ions, but they form hydrogen bonds with Glu183, Glu222, Glu273, and with Asn221, another conserved residue within the catalytic center (Moore et al. 2013; Shiba et al. 2013). Like in TAO, Asn221 forms a hydrogen bond network with conserved residues Tyr304 and Asp323 (Shiba et al. 2013). In TAO, a second putative hydrophobic cavity connects the di-iron active site with a single leaflet of the lipid bilayer facing the mitochondrial matrix (Shiba et al. 2013). This groove is present also in the AtAOX model, being formed by the highly conserved residues Arg164, Asp168, Arg178, Ala186, Leu182, Leu272, Glu183, Glu275, and Ala276. Moreover, three Tyr residues, pivotal in TAO ubiquinol binding and catalysis (Shiba et al. 2013), are also conserved in AtAOX as Tyr258, Tyr280, and Tyr304. As in TAO, a fourth tyrosine residue, Tyr271, is present in AtAOX. However, this Tyr residue is not strictly conserved across all AOXs sequences (see below; Shiba et al. 2013).
Plant AOXs have been reported to be covalent homo-dimers (Umbach and Siedow 1993; Umbach et al. 1994); accordingly, the N-terminal arm of AtAOX appears to be involved in a disulfide bond-mediated homo-dimeric assembly. Indeed, the modeled N-terminal region of AtAOX contains highly conserved amino acid residues (Met191, Met195, His198, Ser201, Leu202, Arg203, Leu216, Arg223, Leu226, Arg240, and Gln247), which in TAO mediate the monomer–monomer contacts (Shiba et al. 2013). Interestingly, in the AtAOX dimer model Cys127 of each monomer is perfectly positioned to form a disulfide bond, despite the non-covalent assembly of the template TAO structure (see below). This residue is conserved in all plant AOXs and together with a second conserved cysteine, Cys177 in AtAOX, it is well known to play a role in the post-translational regulation of AOX activity (Polidoros et al. 2009; Moore et al. 2013; Neimanis et al. 2013).
The N-terminal 31 residues of AtAOX (residues 63–93) are missing from the structural model as the corresponding region of TAO is unstructured and not resolved in the crystal structure of the latter protein. Therefore, to get insight into the structure and function of this region, structure prediction was carried out using two alternative ab initio protein structure prediction programs (Rosetta ab initio and QUARK; Kim et al. 2004; Xu and Zhang 2012). In both cases, the N-terminal 31 residues of AtAOX are predicted to form a single α-helix displaying an amphi-ionic character with four lysine residues lining one side of the helix (Supplementary Fig. 2). Therefore, this helix could contribute to stabilize the interaction of AtAOX with the mitochondrial membrane through interaction with the negatively charged phosphate groups of phospholipids.
In order to put the structural information derived from the TAO three-dimensional structure and the AtAOX molecular model in an evolutionary context, protein sequences homologous to A. thaliana AOX 1A were collected by exhaustive BLASTP searches and assembled in a multiple amino acid sequence alignment (see below). The retrieved AOXs protein sequences dataset contained 111 homologous protein sequences from 37 Land Plants, three Rhodophyta, 27 Fungi, three Metazoa, four Euglenozoa, five Stramenopiles, one Alveolate, two Mycetozoa, one Heterolobosea, and 28 Eubacteria (see Supplementary Materials Table 1 for details). The AtAOX molecular model was then used to map the conservation of residues essential for AOXs structure and function (Moore et al. 2013) along the evolutionary ladder (Fig. 3). Panel A of Fig. 3 shows the location of the three strictly conserved Tyr residues (Tyr258, Tyr280, Tyr304), plus that of a fourth, almost strictly conserved tyrosine residue (Tyr271). Given the proximity to the iron cluster, Tyr280 (Tyr220 in TAO) is the most likely candidate for the amino acid radical involved in catalysis, while Tyr304 (Tyr246 in TAO) supports the hydrogen bonding network which stabilizes the iron cluster ligands (Moore et al. 2013). The strict conservation of Tyr258 (Tyr198 in TAO) is less straightforward to explain. However, both in TAO three-dimensional structure and in AtAOX structural model, this Tyr residue forms a hydrogen bond with a His residue (His206 in TAO, His266 in AtAOX) which links helices α5 and α6 and contributes to shape the cavity that connects the active site to the inner leaflet of the membrane. The almost strict conservation of Tyr271 (Tyr211 in TAO) is apparently linked to his role in dictating the conformation of the N-terminal region which folds back onto the protein helical core. In fact, Tyr271 links helix α4 to the unstructured N-terminal region via an hydrogen bond with the protein backbone. Interestingly, in our dataset, the only substitutions of Tyr 271 observed are with a His (in Ceriporiopsis subvermispora, Sphaerulina musiva, Zymoseptoria tritici, and Nannochloropsis gaditana) and a Cys (in Dacryopinax sp.), residues which retain the hydrogen bonding capability.
Conservation of residues important for the structure/function of AOXs mapped onto the AtAOX structural model. a Active site cavity. b Second hydrophobic cavity. Strictly conserved residues are colored in blue, residues not strictly conserved in red. Residues forming the catalytic site, shown as a reference, are colored by atom type (Color figure online)
Panel B of Fig. 3 shows the residues essential for AOXs structure and function which build up the second hydrophobic cavity observed in the vicinity of the AOX active site (Moore et al. 2013). All the residues are conserved or conservatively substituted. In particular, the three hydrophobic residues closest to the iron cluster (Leu182, Ala186 and Ala276; Leu122, Ala126 and Ala216 in TAO, respectively) are strictly conserved in all the dataset members. Leu272 (Leu212 in TAO) is conservatively substituted by a Phe residue in 20 bacterial AOX sequences, while Glu275 (Glu215 in TAO) conservative substitution into Gln is observed only in Acidovorax sp. (Bacteria) and Angomonas deanei (Trypanosomatidae). Interestingly, Asp168 and Arg178 (Asp100 and Arg118 in TAO), which participate to an extensive hydrogen bonding network centered around the strictly conserved Arg164 (Arg96 in TAO) and involving also Glu275, are substituted into residues able to preserve the above-mentioned network of electrostatic interactions. In fact, Asp168 is substituted by a His in Naegleria gruberi and in Chondrus crispus, and by an Asn in Dictyostelium purpureum and Polysphondylium pallidum (Eukarya), while Arg178 is substituted by a His in Naegleria gruberi and Chondrus crispus, and by a Tyr in Dictyostelium purpureum and Polysphondylium pallidum. Thus, the Asp168–Arg178 interaction becomes a His–His interaction in Naegleria gruberi and in Chondrus crispus, and an Asn–Tyr interaction in Dictyostelium purpureum and Polysphondylium pallidum.
Phylogenesis and Molecular Evolution of AOXs
The full-length alignment of the AOXs protein sequences forming our dataset includes 556 or 585 positions (depending on the alignment algorithm), of which 222 positions are included in the catalytic site alignment. The alignment trimmed by poorly aligned positions with the Gblock analysis was 225 in length and was identical to the catalytic site alignment plus three additional residues at its C-terminus. Therefore, while the catalytic residues show a high degree of conservation across the taxa analyzed, the remaining region of the AOX protein proves difficult to align as a consequence of the high evolutionary rate both in terms of amino acid substitutions and insertions or deletions. These poorly aligned regions had no impact on the phylogenetic reconstruction as their removal produced a phylogenetic tree which has identical supported nodes to that based on the full alignment (Figs. 4, 5). Moreover, phylogenetic reconstructions were consistent irrespective of the algorithm used to build multiple sequence alignments and of the number of seeds used during tree searches (Supplementary Fig. 3).
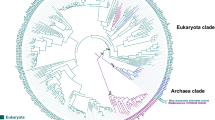
Maximum likelihood (ML) phylogenetic tree based on the full-length sequence of AOX proteins from member of Eubacteria and Eukaryote kingdoms. Multiple sequence alignment was calculated using Clustal W. The ML tree search implemented the LG + G + I substitution model selected by ProtTest under the AIC criterion. Bootstrap values over 100 replicates are reported in correspondence to the nodes
Maximum likelihood (ML) phylogenetic tree based on the catalytic core sequence of AOX proteins from member of Eubacteria and Eukaryotes kingdoms. Multiple sequence alignment was calculated using Clustal W. The ML tree search implemented the LG + G substitution model selected by ProtTest under the AIC criterion. Bootstrap values over 100 replicates are reported in correspondence to the nodes
The inferred Maximum Likelihood (ML) trees show that AOXs of Eubacteria, Land Plants, Metazoa, Mycetozoa, and Euglenozoa form five derived and monophyletic clades with high bootstrap support (BS > 79; Fig. 4). Thus, the main isoform of AOX in each of these organism groups likely evolved from the ancestral AOX carried by their common ancestor. This result is in good agreement with the phylogenetic analysis of Suzuki et al. (2005). Compared to this latter study, our phylogeny includes an extended sampling of AOXs across both the bacterial and eukaryote domains, thus uncovering additional phylogenetic relationships and clades, especially within Eubacteria, Land plants, and Fungi. In the Fungi kingdom, at least two independent AOX lineages are present. One lineage includes the reciprocally monophyletic AOX clades of Ascomycota and Basidiomycota, the other one including the monophyletic AOX clades of Zygomycota, Chytridiomycota, and Microsporidia (Fig. 4). However, the support for these two main lineages is not very strong in the ML trees (Figs. 4, 5). Previous studies supported the monophyly of Ascomycota, Basidiomycota, and Microsporidia AOX clades (Williams et al. 2010; Sun et al. 2007), but with no support for their sister relationships (Williams et al. 2010). Thus, the possibility that the AOX variants of each of these Fungi phyla may have had an independent origin cannot be ruled out. Within Stramenopiles, AOXs of Oomycota and Phaeophytes form two monophyletic groups (BS > 84; Figs. 4, 5) with poorly resolved phylogenetic relationships.
Although the AOX lineages corresponding to Eubacteria and most Eukaryote kingdoms and phyla are well supported, their relationships are poorly resolved. By applying a midpoint rooting, we observed two main clusters in the ML trees, one containing AOXs of Ascomycota and Basidiomycota Fungi, Metazoa, Euglenozoa, Stramenopiles, and Alveolata, and the other including AOXs of Eubacteria, Land Plants, Rhodophyta, Mycetozoa, Heterolobosea, and Zygomycota, Microsporidia, and Chytridiomycota Fungi (Figs. 4, 5). An interesting relationship is found between Florideophyceae red algae and Land Plants AOXs suggesting a common ancestral AOX for Archaeoplastida. However, the lack of support for this node and other basal nodes in the ML trees prevents clear-cut conclusions about the phylogenetic relationships among the main AOX lineages.
AOX, initially considered to be limited to plant species, is expressed in all kingdoms, except Archaebacteria (McDonald et al. 2009). Besides, since 1971, cyanide-tolerant O2 consumption has been reported in animal mitochondria (Hall et al. 1971). AOX was found for the first time in animals which belong to the phyla Mollusca, Nematoda, and Chordata (McDonald and Vanlerberghe 2004), but the taxonomic distribution in this kingdom is broad. The phylogenetic analysis in this work evidenced the presence of a Metazoa monophyletic clade grouping marine benthic organisms, i.e., Crassostrea gigas, Aplysia californica, and Strongylocentrotus purpuratus. The presence of AOX in these species is probably related to the fact that marine organisms may be confronted with rapidly varying oxygen concentrations, hypoxic conditions, and increase of hydrogen sulfide concentration (Turrens 2003; Abele et al. 2007; Sussarellu et al. 2013). Interestingly, Eubacteria AOXs identified in this study all come from sea-water bacterial species in which AOX likely exerts a protective role toward oxidative stress and hydrogen sulfide-mediated inhibition of cytochrome c oxidase.
In the Fungi kingdom, the two AOX clades of Ascomycota and Basidiomycota include species which are mammalian parasites and plant pathogens (e.g., Arthroderma otae and Sphaerulina musiva); in Fungi, one of the putative roles of AOX is to decrease ROS generation (Yukioka et al. 1998; Li et al. 2011). Moreover, in these pathogens, AOXs may have evolved as a defense against the host toxic metabolites, such as carbon monoxide, nitric oxide, cyanide, and hydrogen sulfide that inhibit cytochrome c oxidase (Cooper and Brown 2008).
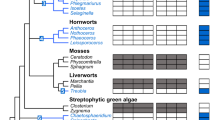
The AOX lineage of Land Plants is structured in three monophyletic clusters with AOXs of the Poaceae Monocots either lying outside a clade comprising AOXs of Araceae Monocots and Eudicots (Figs. 4, 5, and Supplementary Fig. 3A) or in a sister relationship with the AOXs clade of Araceae Monocots (Supplementary Fig. 3B). The support for two distinct clades of Monocots AOXs is high in all the ML trees either based on the full sequence or on the catalytic residues (Figs. 4, 5, Supplementary Figs. 3A, B). In order to analyze in a deeper detail the differences between these two clades, a multiple sequence alignment of full-length AOX proteins from Land Plants was constructed (Fig. 6). This analysis shows that AOXs of Poaceae possess a characteristic portion of the N-terminal sequence which differs from the corresponding region of the other Land Plants.
Sequence and structural features of land plant AOXs N-terminal region. a Multiple amino acid sequence alignment of the 94–128 region of Land Plants AOXs (numbering refers to A. thaliana AOX). The position of the conserved Cys residue involved in the covalent homo-dimer formation (Cys127 in A. thaliana AOX) is indicated by the black star on top of the alignment. Eudicots, Araceae, and Poaceae sequence blocks are indicated by the letters E, A, and P on the right side of the alignment. For clarity, the Araceae and Poaceae sequence blocks are enclosed in a semi-transparent green and yellow rectangle, respectively. b Schematic representation of the structural model of the A. thaliana AOX homo-dimer. The 94–127 regions of each monomer, mediating monomer–monomer interactions, are highlighted by a green border. Cys127 residue of each monomer, mediating covalent dimer formation through a disulfide bridge, is represented in spacefill and colored in yellow. Plant AOXs sequence identity is mapped onto the backbone by coloring from red (high sequence conservation) to blue (low sequence conservation). Yellow circles highlight the low sequence conservation regions corresponding to the N-terminal segment and contacting regions (Color figure online)
The inter-monomers disulfide bond mediating covalent dimerization (see above) is shared by all three Land Plants clades as demonstrated by analysis of the structural models built for two representative species of Araceae and Poaceae, Arum maculatum and Zea mays, respectively (not shown). This result provides some clues for the divergence of the N-terminal sequence of Poaceae with respect to Araceae and Eudicots. In fact, the covalent nature of the plant AOXs homo-dimer likely releases the selective pressure for sequence conservation of the N-terminal region of plant AOXs, leading to the accumulation of mutations in the relatively recent Poaceae group. This is not limited to the N-terminal region involved in the domain-swapping interactions but involves also positions in the catalytic core region that are involved in monomer–monomer contacts (Fig. 6). This finding is in agreement with the separation of Araceae and Poaceae also in the phylogenetic trees based on the multiple sequence alignment of the catalytic site region (Fig. 5, and Supplementary Fig. 3A). An alternative explanation is that, despite the conservation of the disulfide bond, Poaceae may have developed a different pattern of interactions mediated by the N-terminal region with respect to the other Land Plants for a different requirement of physiological regulation. In fact, AOX activity depends on the redox state of the disulfide bridge, with the reduced form being significantly more active than the oxidized form (Umbach et al. 1994; Rhoads et al. 1998). Indeed, in the Poaceae present in our dataset, a specific regulation of AOX activity may be related to the peculiar physiology acquired during their evolution from primitive grasses, living near forest margins, to plants growing in open dry habitats and resisting to drought and other environmental stress (Kellogg 2001).
Conclusion
The recent availability of the crystal structure of TAO (Shiba et al. 2013) has allowed for the first time to analyze the structural details of Plant AOXs through molecular modeling of A. thaliana AOX 1A. Furthermore, the integration of structure prediction with the analyses of amino acid sequences of AOXs from all kingdoms of Life (except Archaea, which do not encode AOXs) allowed to identify conserved elements of AOXs and to infer a phylogenetic tree for the molecular evolution of this protein family.
A very high degree of conservation of the catalytic core of AOXs has been observed, with main differences restricted to the highly variable N-terminal region and contacting residues involved in the homo-dimer formation. In this regard, it is worthwhile to mention the peculiar covalent homo-dimerization mode of Plant AOXs due to the conservation of a cysteine residue which participates to the formation of the intermolecular disulfide bond.
Phylogenetic relationships of AOXs suggest that AOXs proteins have a monophyletic origin in most Eukaryotes kingdoms and in Eubacteria, while AOXs of Fungi seems to have had an independent origin in most phyla. Within plants, AOXs of monocotyledons form two distinct clades with Poaceae AOXs showing loose sequence conservation in the N-terminal region and in positions of the catalytic core region involved in monomer–monomer contacts (Fig. 5). This may be due to the fact that the covalent nature of the plant AOXs homo-dimer releases the selective pressure for sequence conservation or rather to structural requirements for physiological regulation related to environmental stress, evolved during evolution of Poaceae in dry open habitats.
Methods
Structural Model and Amino Acid Sequence Searches
The A. thaliana AOX 1A, the Arum maculatum AOX 1B, and the Zea mays AOX 1 amino acid sequences were obtained from the National Center for Biotechnology Information (NCBI) website (http://www.ncbi.nlm.nih.gov/). These sequences were used to build the corresponding structural models using I-TASSER (Roy et al. 2010) (http://zhanglab.ccmb.med.umich.edu/). The UCSF Chimera program (Pettersen et al. 2004) was used to analyze the structural models and to compare them with the three-dimensional structure of TAO (PDB code 3VV9) (Shiba et al. 2013).
The structural model of the N-terminal 31 residues of AtAOX (residues 63–93) were modeled using the ab initio protein structure prediction programs Rosetta ab initio and QUARK (Kim et al. 2004; Xu and Zhang 2012). Electrostatic potential calculations have been carried out using Swiss PDB viewer (Guex and Peitsch 1997).
Amino acid sequences from the different taxa were obtained using the A. thaliana AOX 1A amino acid sequence as a bait in BLASTP (Altschul et al. 1997) searches. Searches were carried out in the NCBI non-redundant amino acid sequences (NR) database using an E-value threshold of 1 × 10−10. The searches were conducted with a maximum target of 50 aligned sequences. Besides, subsets of the NR database were used to retrieve AOX sequences from specific taxa: Bacteria (taxid:2), Trypanosomatidae (taxid:5654), Fungi (taxid:4751), Monocotyledoneae (taxid:4447), Dicotyledoneae (taxid:71240), as well as non-Plant and non-Fungi Eukarya (taxid:2759).
Phylogenetic Analyses
Phylogenetic analyses were conducted using the Maximum Likelihood (ML) method. Since multiple sequences alignment and tree search parameters have a great impact in phylogenetic reconstructions, we tested for consistency of phylogenetic reconstructions based on different datasets, alignment algorithms, and starting trees.
First, to assess whether highly variable regions in the alignment introduced noise in the phylogenetic reconstruction, we built three distinct amino acid sequence alignments using Clustal W 2.0.1 (Thompson et al. 1994). A first alignment included the entire length sequences of AOXs (full-length alignment); then the full-length alignment was trimmed by (1) removing poorly aligned positions (conserved alignment) with the software Gblocks 0.91b (Castresana 2000) using a relaxed selection of blocks (Talavera and Castresana 2007), and (2) selecting only the portion of the alignment corresponding to the catalytic site (catalytic site alignment). These trimmed alignments resulted identical for 99 % of the positions (see “Results” section), therefore downstream analyses were performed only on the full-length alignment and on the catalytic site alignment. Second, we performed multiple sequence alignment of the full-length and catalytic site datasets using Clustal Omega 1.2.1 (Sievers et al. 2011), which has a similar performance to that of high-quality aligners on small datasets, while it outperforms most packages, in terms of execution time and quality, on large datasets.
The Pairwise Homoplasy Index (PHI) test, implemented in SPLITSTREE v. 4 (Huson and Bryant 2006), did not find statistically significant evidence for recombination in our alignments (p > 0.05). For each of the four alignments, the fitness among 112 models of protein evolution was estimated trough PhyML 3.0 (Le and Gascuel 2008) by ProtTest 2.0 (Abascal et al. 2005) and the optimal model was selected under the corrected Akaike Information Criterion (Akaike 1974). The LG model (Le and Gascuel 2008) with gamma distributed rates across site (+G) and a proportion of invariant sites (+I) was selected for the full-length alignments and the LG + G model for the catalytic site alignments (with the observed amino acid frequencies (+F) for the alignment based on Clustal W).
ML tree searches were performed via PhyML using the model selected by ProtTest, the best swapping strategy between the Nearest Neighbor Interchanges (NNI) and the Subtree Pruning and Regrafting (SPR), with either a single starting tree or multiple (100) random starting trees. Nodes support for the resulting phylogenetic tree was evaluated by 100 bootstrap replications in single starting-tree searches and by Shimodaria–Hasegawa approximate likelihood ratio tests of bipartition support (SH-aLRT) (Guindon et al. 2010) for multiple-seed searches. Phylogenetic analyses were carried out in the PALM integrated framework for phylogenetic reconstruction with automatic likelihood model selectors (Chen et al. 2009) and in the Lifeportal computational resource (https://lifeportal.uio.no).
Phylogenetic analyses based on different multiple sequence alignment algorithms and with single- or multiple-seed searches gave identical results in terms of supported nodes; therefore we discuss in detail trees based on the PALM pipeline using Clustal W alignment algorithm, single starting tree, and 100 bootstrap replicates (Figs. 3, 4) and we provide in Supplementary Fig. 3, the trees calculated in the Lifeportal, based on Clustal Omega, 100 starting tree, and SH-aLRT test of support.
Abbreviations
- Δ9:
-
Desaturase
- Stearoyl:
-
Acyl carrier desaturase
- AOX:
-
Alternative oxidase
- ML:
-
Maximum Likelihood
- MMO:
-
Methane–monooxygenase
- RNR:
-
R2 subunit from ribonucleotide reductase
- ROS:
-
Reactive oxygen species
- TAO:
-
Trypanosomal alternative oxidase
- AtAOX:
-
Arabidopsis thaliana AOX 1A
References
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21:2104–2105
Abele E, Philip E, Gonzalez PM, Puntarulo S (2007) Marine invertebrate mitochondria and oxidative stress. Front Biosci 12:933–946
Ajayi WU, Chaudhuri M, Hill GC (2002) Site-directed mutagenesis reveals the essentiality of the conserved residues in the putative diiron active site of the trypanosome alternative oxidase. J Biol Chem 277:8187–8193
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Automat Contr AC-19:716–723
Albury MS, Elliott C, Moore AL (2009) Towards a structural elucidation of the alternative oxidase in plants. Physiol Plant 137:31–327
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucl Acids Res 25:3389–3402
Andersson ME, Nordlund P (1999) A revised model of the active site of alternative oxidase. FEBS Lett 449:17–22
Berthold DA, Andersson ME, Nordlund P (2000) New insight into the structure and function of the alternative oxidase. Biochim Biophys Acta 1460:241–254
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Chaudhuri M, Hill GC (1996) Cloning, sequencing and functional activity of the Trypanosoma brucei brucei alternative oxidase. Mol Biochem Parasitol 83:125–129
Chen SH, Su SY, Lo CZ, Chen KH, Huang TJ, Kuo BH, Lin CY (2009) PALM: a paralleled and integrated framework for phylogenetic inference with automatic likelihood model selectors. PLoS One 4:e8116
Clifton R, Millar AH, Whelan J (2006) Alternative oxidase in Arabidopsis: a comparative analysis of differential expression in gene family provides new insights into function of non-phosphorylating bypasses. Biochim Biophys Acta 1757:730–741
Cooper CE, Brown GC (2008) The inhibition of mitochondrial cytochrome oxidase by the gases carbon monoxide, nitric oxide, hydrogen cyanide and hydrogen sulfide: chemical mechanism and physiological significance. J Bioenerg Biomembr 40:533–539
Finnegan PM, Soole KL, Umbach AL (2004) Alternative mitochondrial electron transport proteins in higher plants. In: Day DA, Millar AH, Whelan J (eds) Plant mitochondria: from gene to function, vol 17., Advances in photosynthesis and respirationKluwer, Dordrecht, pp 163–230
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of phyml 3.0. Syst Biol 59:307–321
Hall FR, Hollingworth RM, Shankland DL (1971) Cyanide tolerance in millipedes: the biochemical basis. Comp Biochem Physiol 38:723–737
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267
Keeling P, eander BS, Simpson A (2009) Eukaryotes. Eukaryota, organisms with nucleated cells. Version 28 Oct 2009. http://tolweb.org/Eukaryotes/3/2009.10.28 in The Tree of Life Web Project, http://tolweb.org/
Kellogg EA (2001) Evolutionary history of the grasses. Plant Physiol 125:1198–1205
Kim DE, Chivian D, Baker D (2004) Protein structure prediction and analysis using the Robetta server. Nucl Acids Res 32:526–531
Le SQ, Gascuel O (2008) An improved general amino acid replacement matrix. Mol Biol Evol 25:1307–1320
Li Q, Bai Z, O’Donnell A, Harvey LM, Hoskisson PA, McNeil B (2011) Oxidative stress in fungal fermentation processes: the roles of alternative respiration. Biotechnol Lett 33:457–467
Maxwell DP, Wang Y, McIntosh L (1999) The alternative oxidase lowers mitochondrial reactive oxygen production in plant cells. Proc Natl Acad Sci USA 96:8271–8276
McDonald AE (2008) Alternative oxidase: an inter-kingdom perspective on the function and regulation of this broadly distributed ‘cyanide resistant’ terminal oxidase. Funct Plant Biol 35:535–552
McDonald A, Vanlerberghe G (2004) Branched mitochondrial electron transport in the animalia: presence of alternative oxidase in several animal phyla. IUBMB Life 56:333–341
McDonald AE, Vanlerberghe GC, Staples JF (2009) Alternative oxidase in animals: unique characteristics and taxonomic distribution. J Exp Biol 212:2627–2634
Millenaar FF, Lambers H (2003) The alternative oxidase: in vivo regulation and function. Plant Biol 5:2–15
Moore AL, Umbach AL, Siedow JN (1995) Structure-function relationship of the alternative oxidase of plant mitochondria: a model of the active site. J Bioenerg Biomembr 27:367–377
Moore AL, Shiba T, Young L, Harada S, Kita K, Ito K (2013) Unraveling the heater: new insights into the structure of the alternative oxidase. Annu Rev Plant Biol 64:637–663
Neimanis K, Staples JF, Hüner NP, McDonald AE (2013) Identification, expression, and taxonomic distribution of alternative oxidases in non-angiosperm plants. Gene 526:275–286
Parsons HL, Yip JY, Vanlerberghe GC (1999) Increased respiratory restriction during phosphate-limited growth in transgenic tobacco cells lacking alternative oxidase. Plant Physiol 121:1309–1320
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Polidoros AN, Mylona PV, Arnholdt-Schmitt B (2009) AOX gene structure, transcript variation and expression in plants. Physiol Plant 137:342–353
Purvis AC, Shewfelt RL (1993) Does the alternative pathway ameliorate chilling injury in sensitive plant tissue? Physiol Plant 88:712–718
Rasmusson AG, Geisler DA, Moller IM (2008) The multiplicity of dehydrogenases in the electron transport chain of plant mitochondria. Mitochondrion 8:47–60
Rhoads DM, Umbach AL, Sweet CR, Lennon AM, Rauch GS, Siedow JN (1998) Regulation of the cyanide-resistant alternative oxidase of plant mitochondria. Identification of the cysteine residue involved in α-keto acid stimulation and intersubunit disulfide bond formation. J Biol Chem 273:30750–30756
Roberts CW, Roberts F, Henriquez FL, Akiyoshi D, Samuel BU, Richards TA, Mihous W, Kyle D, McIntosh L, Hill GC, Chaudhuri M, Tzipori S, McLeod R (2004) Evidence for mitochondrial-derived alternative oxidase in the apicomplexan parasite Cryptosporidium parvum: a potential anti-microbial agent target. Int J Parasitol 34:297–308
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5:725–738
Shiba T, Kido Y, Sakamoto K, Inaoka DK, Tsuge C (2013) Structure of the trypanosome cyanide insensitive alternative oxidase. Proc Natl Acad Sci USA 110:4580–4585
Siedow JN, Umbach AL, Moore AL (1995) The active site of the cyanide-resistant oxidase from plant mitochondria contains a binuclear iron center. FEBS Lett 362:10–14
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Stechmann A, Hamblin K, Pérez-Brocal V, Gaston D, Richmond GS, van der Giezen M, Clark CG, Roger AJ (2008) Organelles in Blastocystis that blur the distinction between mitochondria and hydrogenosomes. Curr Biol 18:580–585
Stenmark P, Nordlund P (2003) A prokaryotic alternative oxidase present in the bacterium Novosphingobium aromaticivorans. FEBS Lett 552:189–192
Sun J, Lu X, Rinas U, Ping Zeng A (2007) Metabolic peculiarities of Aspergillus niger disclosed by comparative metabolic genomics. Genome Biol 8(9):R182
Sussarellu R, Dudognon T, Fabioux C, Soudant P, Moraga D, Kraffe E (2013) Rapid mitochondrial adjustments in response to short-term hypoxia and re-oxygenation in the Pacific oyster, Crassostrea gigas. J Exp Biol 216:1561–1569
Suzuki T, Hashimoto T, Yabu Y, Majiwa PA, Ohshima S, Suzuki M, Lu S, Hato M, Kido Y, Sakamoto K, Nakamura K, Kita K, Ohta N (2005) Alternative oxidase (AOX) genes of African trypanosomes: phylogeny and evolution of AOX and plastid terminal oxidase families. J Eukaryot Microbiol 52(4):374–381
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577
Tanudji M, Sjöling S, Glaser E, Whelan J (1999) Signals required for the import and processing of the alternative oxidase into mitochondria. J Biol Chem 15:1286–1293
Thirkettle-Watts D, McCabe TC, Clifton R, Moore C, Finnegan PM, Day DA, Whelan J (2003) Analysis of the alternative oxidase promoters from soybean. Plant Physiol 133:1158–1169
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignments through sequence weighting, position specific gap penalties and weight matrix choice. Nucl Acids Res 22:4673–4680
Turrens JF (2003) Mitochondrial formation of reactive oxygen species. J Physiol 552:335–344
Umbach AL, Siedow JN (1993) Covalent and noncovalent dimers of the cyanide-resistant alternative oxidase protein in higher plant mitochondria and their relationship to enzyme activity. Plant Physiol 103:845–854
Umbach AL, Wiskich JT, Siedow JN (1994) Regulation of alternative oxidase kinetics by pyruvate and intermolecular disulfide bond redox status in soybean seedling mitochondria. FEBS Lett 348:181–184
Van Aken O, Giraud E, Clifton R, Whelan J (2009) Alternative oxidase: a target and regulator of stress responses. Physiol Plant 137:354–361
Van Hellemond JJ, Simons B, Millenaar FF, Tielens AG (1998) A gene encoding the plant-like alternative oxidase is present in Phytomonas but absent in Leishmania spp. J Eukaryot Microbiol 45:426–430
Vanlerberghe GC (2013) Alternative oxidase: a mitochondrial respiratory pathway to maintain metabolic and signaling homeostasis during abiotic and biotic stress in plants. Int J Mol Sci 14:6805–6847
Vanlerberghe GC, McIntosh L (1997) Alternative oxidase: from gene to function. Annu Rev Plant Physiol Plant Mol Biol 48:703–734
Veiga A, Arrabaça JD, Loureiro-Dias MC (2003) Cyanide–resistant respiration, a very frequent metabolic pathway in yeasts. FEMS Yeast Res 3:239–245
Vishwakarma A, Bashyam L, Senthilkumaran B, Scheibe R, Padmasree K (2014) Physiological role of AOX1a in photosynthesis and maintenance of cellular redox homeostasis under high light in Arabidopsis thaliana. Plant Physiol Biochem 81:44–53
Williams BAP, Elliot C, Burri L, Kido Y, Kita K et al (2010) A broad distribution of the alternative oxidase in microsporidian parasites. PLoS Pathog 6(2):e1000761
Xu D, Zhang Y (2012) Ab initio protein structure assembly using continuous structure fragments and optimized knowledge-based force field. Proteins 80:1715–1735
Yukioka H, Inagaki S, Tanaka R, Katoh K, Miki N, Mizutani A, Masuko M (1998) Transcriptional activation of the alternative oxidase gene of the fungus Magnaporthe grisea by a respiratory-inhibiting fungicide and hydrogen peroxide. Biochim Biophys Acta 1442:161–169
Acknowledgments
D.S. is supported by FCT post-doctoral fellowship SFRH/BPD/105274/2014 from the European Social Fund and Portuguese Ministério da Educação e Ciência, and by the project “Genomics and Evolutionary Biology” cofinanced by North Portugal Regional Operational Programme 2007/2013 (ON.2-O Novo Norte), under the National Strategic Reference Framework (NSRF), through the European Regional Development Fund (ERDF).
Author Contributions
RP collected the AOXs sequences, carried out the molecular modeling studies and drafted the manuscript. DS carried out the phylogenetic analysis and helped to draft the manuscript. VB carried out ab initio molecular modeling studies and electrostatic potential calculations. RA participated in the interpretation of the results and helped to draft the manuscript. PA and FP conceived the study, participated in its design and coordination and helped to draft the manuscript. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Rosa Pennisi and Daniele Salvi contributed equally to this work.
Database: AtAOX model data are available in the PMDB database under the accession number PM0080189.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pennisi, R., Salvi, D., Brandi, V. et al. Molecular Evolution of Alternative Oxidase Proteins: A Phylogenetic and Structure Modeling Approach. J Mol Evol 82, 207–218 (2016). https://doi.org/10.1007/s00239-016-9738-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-016-9738-8