Abstract
Pore-forming proteins/toxins (PFPs/PFTs) are the distinct class of membrane-damaging proteins. They act by forming oligomeric pores in the plasma membranes. PFTs and PFPs from diverse organisms share a common mechanism of action, in which the designated pore-forming motifs of the membrane-bound protein molecules insert into the membrane lipid bilayer to create the water-filled pores. One common characteristic of these pore-forming motifs is that they are amphipathic in nature. In general, the hydrophobic sidechains of the pore-forming motifs face toward the hydrophobic core of the membranes, while the hydrophilic residues create the lining of the water-filled pore lumen. Interestingly, pore-forming motifs of the distinct subclass of PFPs/PFTs share very little sequence similarity with each other. Therefore, the common guiding principle that governs the sequence-to-structure paradigm in the mechanism of action of these PFPs/PFTs still remains an enigma. In this article, we discuss this notion using the examples of diverse groups of membrane-damaging PFPs/PFTs.
Graphic Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
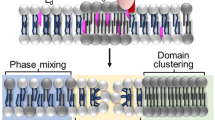
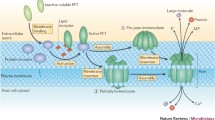
Membrane pore-formation is one of the most efficient mechanisms of cell-killing employed by the diverse life forms. Such pathophysiological function is achieved by a unique group of protein toxins known as the pore-forming toxins (PFTs). PFTs form pores of specific size and stoichiometry in the lipid bilayer of the target cell membranes. This allows unrestricted movement of ions, water, and other molecules through the cell membranes, which in turn disrupts the cellular homeostasis leading to the cell-death (Bischofberger et al. 2012; Dal Peraro and van der Goot 2016; Mondal et al. 2018). PFTs are generally produced as soluble monomers. Upon encountering the target plasma membranes, they assemble into oligomeric complexes, which are finally converted into the transmembrane pores (Gonzalez et al. 2008). Presence of the PFTs is ubiquitously found in diverse organisms that include bacteria, fungi, sea anemones, plants, and even higher vertebrates such as humans. These proteins are produced for different purposes in different organisms. Pathogenic bacteria utilize PFTs as their virulence factors, whereas eukaryotic organisms usually use them for the defense mechanisms. Higher vertebrates produce pore-forming proteins (PFPs) to kill the pathogenic bacteria, and to destroy the pathogen-infected cells (Bayly-Jones et al. 2017; Dal Peraro and van der Goot 2016; Los et al. 2013; Spicer et al. 2017).
Although the PFTs from diverse organisms have different functions, they share remarkable similarities in their structure and mechanism of pore-formation. PFTs bind to the plasma membranes of the target cells, either via specific association with the components of the cell membranes or receptor(s), or through non-specific association with the membrane lipid bilayer. Upon membrane-binding, PFT monomers assemble into the oligomeric pores. This involves membrane-insertion of the specific pore-forming motif(s) that creates the transmembrane water-filled pore (Gonzalez et al. 2008). Depending on the number of protomers involved in the pore-formation process, pore size can vary in different PFTs. The stoichiometric assembly can range from tetramer for the thermostable direct hemolysin (TDH) from Vibrio parahaemolyticus, to the heptameric assembly for Staphylococcus aureus α-hemolysin and Vibrio cholerae cytolysin (VCC), where pore diameters are in the range of 1–2 nm (De and Olson 2011; Kundu et al. 2017; Song et al. 1996). In contrast, more than 30–50 protomers are known to be involved in the pore-formation by the cholesterol-dependent cytolysins (CDCs), such as Perfringolysin O (PFO) from Clostridium perfringens (Rossjohn et al. 1997).
PFTs are classified into two distinct categories, based on the secondary structural motifs that PFTs utilize to generate the oligomeric pore. PFTs, in which the membrane-spanning elements are composed of the α-helices, are called α-PFTs (Dal Peraro and van der Goot 2016; Mondal et al. 2018). Some of the well-characterized α-PFTs include colicins (Colicin Ia from Escherichia coli), Cytolysin A (ClyA; produced by E. coli, Salmonella species, and Shigella flexneri) and Actinoporins (for example, Fragaceatoxin from Actinia fragacea) (Mechaly et al. 2009; Mueller et al. 2009; Parker et al. 1992). PFTs that utilize β-barrels for the membrane-spanning regions are called the β-PFTs (Mondal and Chattopadhyay 2020). β-PFTs have two subfamilies: (i) small β-PFTs that form pores of ~ 1–2 nm diameter, with ~ 7–9 protomers in the pore assembly (for example, α-hemolysin from S. aureus) and (ii) large β-PFTs such as CDCs that generate pores of ~ 30–50 nm diameter consisting of ~ 30–50 protomers in the pore assembly (for example, Listeriolysin O from L. monocytogenes) (Koster et al. 2014; Song et al. 1996). Both the families of α-PFTs and β-PFTs include multiple well-explored eukaryotic PFPs. Actinoporins are the group of one such eukaryotic α-PFTs that are secreted by the different species of sea anemones (Kristan et al. 2009). Membrane attack complex and perforin family (MACPF) proteins are produced by the cells of the immune system of the higher vertebrates (Reboul et al 2016). These proteins form β-barrel pores on the plasma membranes of the pathogen-infected cells (Tavares et al. 2014). Apart from the MACPFs, gasdermins represent another group of eukaryotic PFPs. Gasdermins form β-barrel pores ‘from-inside’ of the cells, that in turn mediate the release of the inflammatory cytokines and trigger pyroptosis (Shi et al. 2017). β-PFTs are more extensively characterized at the molecular levels, as compared to the α-PFTs. Numerous studies on the well-orchestrated structure–function mechanisms of the PFTs have revealed in-depth molecular insights regarding their modes of action. However, the intricate details of the pore-formation mechanism of the PFTs still remain elusive in most of the cases.
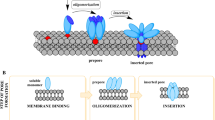
One of the crucial steps in the pore-formation mechanism of the PFTs is the insertion of the pore-forming motif into the lipid bilayer of the target cell membranes (Mondal and Chattopadhyay 2020). The insertion step is usually linked with the oligomerization of the membrane-associated toxin protomers. In one of the models of pore-formation mechanism, PFTs first oligomerize on the membrane surface, and then subsequently insert their pore-forming motifs into the membranes to form the transmembrane pores. Small β-PFTs such as S. aureus α-hemolysin and Vibrio cholerae cytolysin (VCC) follow this model (Rai and Chattopadhyay 2014; Valeva et al. 1995). Some of the large β-PFTs such as CDCs follow another model, where the sequential oligomerization of the toxin protomers is associated with the concomitant membrane-insertion process (Bakrac et al. 2008).
In most of the cases, the pore-forming motifs of the PFTs remain folded against the central domain of the protein, when they are in the water-soluble monomeric form. In α-PFTs and small β-PFTs, the pore-forming region(s) already contain the pre-formed α-helical segments and β-strands, respectively, which constitute the transmembrane motif (Mueller et al. 2009; Olson and Gouaux 2005). When these PFTs interact with the lipid bilayer, these pre-formed motifs insert into the membranes (Dal Peraro and van der Goot 2016; Degiacomi et al. 2013; Mondal and Chattopadhyay 2020). Interestingly, the pore-forming element(s) in the large β-PFTs like CDCs remain as α-helical form in the water-soluble monomeric state (Heuck et al. 2010). During the pore-formation process, these elements undergo structural rearrangements to convert into the β-hairpins to form the components of the β-barrel pore (Mondal and Chattopadhyay 2020; Reboul et al. 2016). Studies have also revealed that the pore-formation process of many β-PFTs involve formation of the intermediates that are termed as the pre-pore, in which the pore-forming motifs remain partially collapsed and are not fully inserted into the membranes (Yamashita et al. 2014). Once the pre-pore oligomeric assembly is created, each of the protomers fully extends their pore-forming region into the membrane lipid bilayer to form the β-barrel stem of the transmembrane pore (Iacovache et al. 2016; Paul and Chattopadhyay 2014).
One remarkable feature of the pore-forming motif(s) of the PFTs is their structural compatibility within the physicochemical environment of the membrane lipid bilayer so that they can form the water-filled transmembrane pores. Overall amphipathic nature of the pore-forming motifs allows them to anchor into the membrane environment. In the transmembrane form, hydrophobic residues of the pore-forming segment face toward the hydrophobic core of the membrane lipid bilayer, while the hydrophilic residues line the water-filled lumen of the pore, in general (De and Olson 2011; Mondal and Chattopadhyay 2020; Song et al. 1996). It is documented that the physicochemical environment of the membrane lipid bilayer itself plays critical roles to shape the structural architecture of both the α-helical and β-barrel transmembrane motifs (White et al. 2001; Wimley et al. 1998). Also, it is obvious that the structural disposition of the pore-forming motif of any PFT must be guided by the inherent pattern of the amino acid sequences of such regions. Various computational approaches have explored to predict the β-barrel structures in the membrane proteins (Waldispuhl et al. 2006). For example, one such study has attempted to identify the potential membrane-interacting surfaces in such motifs, and their possible positioning with respect to the membrane bilayer mid-plane (Wimley 2002). Similarly, computational prediction of the transmembrane α-helical segments has also made considerable progress in the recent years (Fleishman and Ben-Tal 2006). Interestingly, the pore-forming motifs of the diverse PFT family members show large variations in their sequence patterns. Previous literatures have shown that within a specific subclass of the PFTs, pore-forming motifs display noticeable sequence similarities. For example, membrane-interacting regions of the CDC family of β-PFTs and transmembrane domains of the ClyA-like α-PFTs reveal presence of conserved signature sequences at some places (Brauning and Groll 2018; Savinov and Heuck 2017). However, very little sequence similarities are observed across the different subclass of the α-PFTs and β-PFTs, even though they adopt overall similar structural disposition, particularly for their transmembrane scaffolds. Therefore, it remains an enigma that how a similar structural disposition of the pore-forming element(s) of the structurally-related PFTs is generated from the sequence motifs that are not so similar. In this article, we discuss this notion, using some of the prototype examples of the PFT family members. This review article attempts to provide a consolidated overview of the sequence diversities in the pore-forming motifs of the distinct class of PFTs and PFPs.
Small Pore-Forming β-PFTs
β-PFTs that form small oligomeric pores of 1–2 nm diameter, composed of ~ 7 protomers, are one of the most well-characterized PFTs. Some of the archetypical examples of such β-PFTs include S. aureus α-hemolysin, Leukocidin F, and V. cholerae cytolysin (VCC) (De and Olson 2011; Olson et al. 1999; Song et al. 1996) (Fig. 1). Transmembrane oligomeric pores formed by these β-PFTs have a characteristic mushroom-shaped architecture, in which the β-barrel stem traverses through the depth of the membrane lipid bilayer (Fig. 1a). For the formation of the β-barrel stem, each of the toxin protomers contributes a pore-forming β-strand pair constituted from ~ 40–45 amino acid residues (De and Olson 2011; Savva et al. 2013). Sequence alignment of the pore-forming motifs of some of the prominent small pore-forming β-PFTs shows presence of prominent sequence similarities at many places (Fig. 1c).
Small pore-forming β-PFTs. a Structural model of the oligomeric pore of S. aureus α-hemolysin (generated using the PDB ID 7AHL). One protomer is highlighted in green color. b Structural model of the oligomeric pore of Aeromonas hydrophila Aerolysin (generated using the PDB ID 5JZT). One of the protomers is highlighted in green. c Sequence alignment of the pore-forming motifs of some of the prototype small pore-forming β-PFTs. LeukF, leukocidin F from S. aureus; NetB, necrotic enteritis toxin B-like of Clostridium perfringens; α-Hly, S. aureus α-hemolysin; VCC; Vibrio cholerae cytolysin. d Sequence alignment of the pore-forming motifs of some of the prototype small pore-forming β-PFTs having an elongated β-barrel structure. Iota toxin, from Clostridium perfringens; Epsilon toxin, from Clostridium perfringens; Aerolysin, from Aeromonas hydrophila; Lysenin, from Eisenia fetida. Protective antigen, from Bacillus anthracis (Color figure online)
There is another group of β-PFTs that form small β-barrel pores, having distinctly different structural architecture. Aeromonas hydrophila Aerolysin, anthrax toxin protective antigen pore, lysenin are some of the prominent examples of this category (Iacovache et al. 2016; Jiang et al. 2015; Podobnik et al. 2016, 2017). The oligomeric transmembrane pores formed by these β-PFTs show highly elongated β-barrel stems (Fig. 1b). Here, the β-barrel stem is constituted from ~ 70–75 residue-long β-strand pairs contributed by ~ 7–9 toxin protomers (Savva et al. 2019; Yamada et al. 2020). Pore-forming motifs of these elongated β-barrel-containing PFTs show some degree of sequence similarity at places (Fig. 1d), extent of which appears to be less as compared to that observed in the other category of small pore-forming β-PFTs mentioned above.
It is important to note that these two distinct categories of small pore-forming β-PFTs do not share any detectable sequence similarity in their pore-forming motifs. Therefore, the sequence comparison of the pore-forming motifs of these small pore-forming β-PFTs possibly suggest distinct guiding principle for the structural disposition of their transmembrane β-barrel elements.
Cholesterol-Dependent Cytolysins (CDCs)
Cholesterol-dependent cytolysins (CDCs) are another distinct class of β-PFTs, produced mostly by the Gram-positive pathogenic bacteria. CDCs are secreted by the pathogens as the water-soluble monomers, which form oligomeric pores on the cholesterol-containing cellular membrane. CDCs make large pores of diameter in the range of ~ 30–50 nm. ~ 30–50 toxin protomers are involved in such pore assembly (Reboul et al. 2016; Sonnen et al. 2014; van Pee et al. 2017). Studies on several CDCs have revealed a remarkably conserved structural architecture. In the CDC monomers, pore-forming motifs are retained in the form of two spatially distant, ~ 35 residue-long, α-helices (Fig. 2a). In the process of membrane pore-formation, these two α-helical motifs convert into two amphipathic transmembrane β-hairpins (TMH-1 and TMH-2) that insert into the membranes to form the β-barrel pores (Heuck et al. 2010, 2001; Tweten 2005) (Fig. 2b–d).
CDC family of large pore-forming β-PFTs. a Structural model of the Listeriolysin O monomer (generated using the PDB ID 4CDB). Two of the α-helical motifs that form the transmembrane β-hairpins (TMH-1 and TMH-2) are shown in red. b Structural model of a Pneumolysin protomer (generated using the PDB ID 5LY6) in its pore-forming state. c Structural map of the Pneumolysin pore (generated using the map obtained from the electron microscopy data bank (EMDB) corresponding to the EMD ID 4118). The model was visualized using the UCSF Chimera (Pettersen et al. 2004). d Sequence alignment of the pore-forming motifs [transmembrane β-hairpin (TMH)-1 and 2] of some of the prototype CDC family members (Color figure online)
Sequence alignment of the pore-forming motifs of some of the prototype CDCs shows presence of highly conserved/similar residues at a number of positions. Presence of conserved/similar residues appears to be more prominent in TMH-2 of these CDCs (Fig. 2d). Therefore, these observations suggest that the CDC family of large pore-forming β-PFTs have possibly evolved with a common guiding principle that is encoded in the sequence of their pore-forming motifs.
Perforin and Membrane Attack Complex
Membrane attack complex of the complement system and perforin produced by the T lymphocytes constitute a distinct class of eukaryotic pore-forming proteins, commonly defined as the MACPF. They play pivotal roles in the vertebrate immune response generation and host defense mechanism (Rosado et al. 2008). In their mode of action, they display remarkable similarity with the CDC family of β-PFTs (Bayly-Jones et al. 2017; Reboul et al. 2016). In the monomeric state, MACPF pore-forming domain harbors two α-helical motifs, each being ~ 50-residue-long (Law et al. 2010; Lovelace et al. 2011). During oligomeric pore-formation, these motifs convert into transmembrane β-hairpins that insert into the membranes to create the β-barrel (Reboul et al. 2016) (Fig. 3a). In spite of sharing an overall similar structural mechanism of membrane pore-formation, the pore-forming motifs of the MACPFs do not display any sequence similarity with those of the CDCs (Rosado et al. 2008). However, within the MACPF category, perforin and the membrane attack complex share some detectable sequence similarity in their pore-forming motifs (Fig. 3b).
MACPF and gasdermin family of eukaryotic β-PFPs. a Structural model of the pore assembly of a perforin family member, perforin-2 (generated using the PDB ID 6SB5). b Sequence alignment of the pore-forming motifs of mouse perforin and human complement component C8α. c Structural model of the gasdermin A3 protomer in its pore-forming configuration (based on the PDB ID 6CB8). d Sequence alignment of the pore-forming motifs of the gasdermin family members of human (h) and mouse (m) origin
Gasdermins
Gasdermin family of PFPs are implicated in the host immune functions that include inflammation and pyroptosis. Functionality of the gasdermins is attributed to their ability to form β-barrel pores in the target membranes. In the oligomeric β-barrel pore, each of the 26–28 protomers contributes two anti-parallel β-strand pairs to create the transmembrane β-barrel stem (Ruan et al. 2018) (Fig. 3c). Such a structural design of the gasdermin β-barrel pores is similar to those observed with the pores formed by the CDC family of β-PFTs, as well as MACPFs of the vertebrate immune system (Liu et al. 2019). However, the pore-forming motifs of the structurally-characterized gasdermins do not share any similarity with those of the MACPF/CDCs. However, the individual members within the gasdermin family show similarities with each other in their pore-forming motifs’ sequence patterns (Fig. 3d).
α-PFTs
As compared to the β-PFTs, α-PFTs are much less characterized. One of the most prominent α-PFTs is the cytolysin A (ClyA). ClyA is mostly an α-helical protein, and it forms homo-dodecameric pores in the target membranes. The transmembrane pore is constituted from the α-helical motifs of the ClyA protomers (Mueller et al. 2009) (Fig. 4a). Pore-forming motifs of ClyA from different bacterial species show remarkable sequence identity (Hunt et al. 2010) (Fig. 4a). Yersinia YaxAB is another example of the α-PFT. Recent structural study has revealed that the heterodimer of YaxA and YaxB forms decameric pore assembly, in which the transmembrane regions are constituted from the α-helical motifs (Brauning et al. 2018) (Fig. 4b). Interestingly, the α-helical pore-forming motifs of YaxA and YaxB adopt distinct structural disposition in the pore state (Fig. 4b), and they do not share any sequence similarity with the pore-forming region of ClyA. Therefore, YaxAB appears to represent a distinct subgroup in the α-PFT family.
ClyA and YaxAB families of α-PFTs. a Structural model of ClyA pore assembly (PDB ID 2WCD). One of the protomers is shown in cyan color, and two of its pore-forming α-helical segments are colored in dark red. Sequence alignment of the two pore-forming α-helical regions of the two ClyA-family members is shown in the right side. b Structural model of YaxAB pore (PDB ID 6EL1). Two neighboring protomers of YaxA and YaxB are shown in cyan and pale red, respectively. Pore-forming regions of YaxA and YaxB are shown in dark red and blue, respectively. Sequence alignment of the pore-forming regions of YaxA and YaxB is shown in the right side (Color figure online)
Actinoporins
Actinoporins are the distinct group of eukaryotic PFTs secreted by the sea anemones. Actinoporins possess unique structural organization composed of a central β-sandwich domain flanked by an N-terminal α-helical motif (Kristan et al. 2009; Ramirez-Carreto et al. 2020; Rojko et al. 2016) (Fig. 5a). The most well-characterized actinoporins are equinatoxin II (EqtII) from Actinia equina, fragaceatoxin C (FraC) from Actinia fragacea, and sticholysin I and II (Stn I and Stn II) from Stichodactyla helianthus (Kristan et al. 2007; Mancheno et al. 2003; Mechaly et al. 2009; Morante et al. 2015). Upon binding to the target cell membranes, actinoporins transform into the oligomeric pores, in which ~ 30 residue-long N-terminal α-helical region of each protomer inserts into the membrane to create the pore (Gutierrez-Aguirre et al. 2004; Kristan et al. 2007; Ros et al. 2015) (Fig. 5b). Therefore, based on their pore-formation mechanism, actinoporins are commonly considered to act as α-PFTs. Nevertheless, actinoporins have distinct overall structural organization, as compared to the conventional α-PFTs such as ClyA. Sequence alignment of the pore-forming motifs of some of the well-characterized actinoporins shows high sequence similarities, thus suggesting relatedness in these PFTs (Fig. 5c).
Actinoporin family of eukaryotic α-PFTs. a Structural model of the Fragaceatoxin C monomer (PDB ID 3VWI). N-terminal 30-residue-long region is colored in dark red. b Four of the protomers of the Fragaceatoxin C pore (PDB ID 4TSY). N-terminal 30-residue-long α-helical region (that acts as the pore-forming motif) of one protomer is shown in dark red. c Sequence alignment of the N-terminal pore-forming motifs of the actinoporin family members (Color figure online)
Conclusion
Large numbers of studies on wide varieties of PFTs have provided detail insights regarding the structural mechanisms associated with their mode of actions. It is clear that the diverse class of PFTs employ distinct structural mechanisms to achieve one single purpose of damaging the target membranes via formation of well-defined transmembrane pores. There is little or no sequence similarities observed in the pore-forming motifs of the PFTs that belong to the distinct sub-categories. Nevertheless, such pore-forming motifs from the distinct sub-categories of α-PFTs and β-PFTs still retain the propensities to adopt either α-helical or β-barrel structures in their membrane-inserted configuration. Such a propensity is possibly guided by the inherent amino acid composition of these motifs, and further shaped up by the physicochemical environment of the membrane lipid bilayer. It is not surprising that similar secondary structural motifs can be generated from diverse sequences. What is surprising for the case of PFTs/PFPs that their pore-forming motifs adopt similar structural architecture that can create water-filled pores in the context of the hydrophobic environment of the membrane lipid bilayer. It remains a long-standing puzzle associated with the structure–function mechanisms of these class of protein toxins. Notably, distinct PFTs within the same sub-category show some degree of sequence identities/similarities in their pore-forming motifs, in many cases. Exact implications of such conserved residues within the pore-forming motifs still remain unexplored in the most cases. Future research endeavors exploring the precise structural and functional roles of any conserved residue(s) within the pore-forming motifs would provide crucial information regarding the structure–function mechanisms in this class of protein toxins. Future structural and bioinformatics studies in this direction will possibly shed more detail insights regarding the origin and evolution of the distinct PFT families, and the structural mechanisms associated with their membrane pore-forming properties.
Methods
Amino acid sequence alignments of the pore-forming motifs of distinct PFTs were generated using the ClustalW (available online at https://www.genome.jp/tools-bin/clustalw), and were visualized with ESPript 3.0 (available online at https://espript.ibcp.fr/) (Robert and Gouet 2014). Strictly conserved residues are shown as white characters in red background, and similar residues are shown as red characters in white background. Strictly conserved and similar residues are shown with blue frames. Amino acid sequences corresponding to the pore-forming motifs were retrieved either from the protein data bank (PDB) (https://www.rcsb.org/), or from the NCBI server (https://www.ncbi.nlm.nih.gov/protein/). Start and end residues of the pore-forming motifs are numbered accordingly. Mean %identity and mean %similarity scores for the aligned sequences were obtained from the ESPript 3.0.
Protein structure coordinates were obtained from the protein data bank (PDB). The protein structural models were visualized using PyMOL (The PyMOL Molecular Graphics System, Version 2.3.2 Schrödinger, LLC.).
References
Bakrac B, Gutierrez-Aguirre I, Podlesek Z, Sonnen AF, Gilbert RJ, Macek P, Lakey JH, Anderluh G (2008) Molecular determinants of sphingomyelin specificity of a eukaryotic pore-forming toxin. J Biol Chem 283:18665–18677
Bayly-Jones C, Bubeck D, Dunstone MA (2017). The mystery behind membrane insertion: a review of the complement membrane attack complex. Philos Trans R Soc Lond B 372
Bischofberger M, Iacovache I, van der Goot FG (2012) Pathogenic pore-forming proteins: function and host response. Cell Host Microbe 12:266–275
Brauning B, Groll M (2018) Structural and mechanistic features of ClyA-like alpha-pore-forming toxins. Toxins (Basel) 10
Brauning B, Bertosin E, Praetorius F, Ihling C, Schatt A, Adler A, Richter K, Sinz A, Dietz H, Groll M (2018) Structure and mechanism of the two-component alpha-helical pore-forming toxin YaxAB. Nat Commun 9:1806
Dal Peraro M, van der Goot FG (2016) Pore-forming toxins: ancient, but never really out of fashion. Nat Rev Microbiol 14:77–92
De S, Olson R (2011) Crystal structure of the Vibrio cholerae cytolysin heptamer reveals common features among disparate pore-forming toxins. Proc Natl Acad Sci USA 108:7385–7390
Degiacomi MT, Iacovache I, Pernot L, Chami M, Kudryashev M, Stahlberg H, van der Goot FG, Dal Peraro M (2013) Molecular assembly of the aerolysin pore reveals a swirling membrane-insertion mechanism. Nat Chem Biol 9:623–629
Fleishman SJ, Ben-Tal N (2006) Progress in structure prediction of alpha-helical membrane proteins. Curr Opin Struct Biol 16:496–504
Gonzalez MR, Bischofberger M, Pernot L, van der Goot FG, Freche B (2008) Bacterial pore-forming toxins: the (w)hole story? Cell Mol Life Sci 65:493–507
Gutierrez-Aguirre I, Barlic A, Podlesek Z, Macek P, Anderluh G, Gonzalez-Manas JM (2004) Membrane insertion of the N-terminal alpha-helix of equinatoxin II, a sea anemone cytolytic toxin. Biochem J 384:421–428
Heuck AP, Tweten RK, Johnson AE (2001) Beta-barrel pore-forming toxins: intriguing dimorphic proteins. Biochemistry 40:9065–9073
Heuck AP, Moe PC, Johnson BB (2010) The cholesterol-dependent cytolysin family of gram-positive bacterial toxins. Subcell Biochem 51:551–577
Hunt S, Green J, Artymiuk PJ (2010) Hemolysin E (HlyE, ClyA, SheA) and related toxins. Adv Exp Med Biol 677:116–126
Iacovache I, De Carlo S, Cirauqui N, Dal Peraro M, van der Goot FG, Zuber B (2016) Cryo-EM structure of aerolysin variants reveals a novel protein fold and the pore-formation process. Nat Commun 7:12062
Jiang J, Pentelute BL, Collier RJ, Zhou ZH (2015) Atomic structure of anthrax protective antigen pore elucidates toxin translocation. Nature 521:545–549
Koster S, van Pee K, Hudel M, Leustik M, Rhinow D, Kuhlbrandt W, Chakraborty T, Yildiz O (2014) Crystal structure of listeriolysin O reveals molecular details of oligomerization and pore formation. Nat Commun 5:3690
Kristan K, Viero G, Macek P, Dalla Serra M, Anderluh G (2007) The equinatoxin N-terminus is transferred across planar lipid membranes and helps to stabilize the transmembrane pore. FEBS J 274:539–550
Kristan KC, Viero G, Dalla Serra M, Macek P, Anderluh G (2009) Molecular mechanism of pore formation by actinoporins. Toxicon 54:1125–1134
Kundu N, Tichkule S, Pandit SB, Chattopadhyay K (2017) Disulphide bond restrains the C-terminal region of thermostable direct hemolysin during folding to promote oligomerization. Biochem J 474:317–331
Law RH, Lukoyanova N, Voskoboinik I, Caradoc-Davies TT, Baran K, Dunstone MA, D'Angelo ME, Orlova EV, Coulibaly F, Verschoor S, Browne KA, Ciccone A, Kuiper MJ, Bird PI, Trapani JA, Saibil HR, Whisstock JC (2010) The structural basis for membrane binding and pore formation by lymphocyte perforin. Nature 468:447–451
Liu Z, Wang C, Yang J, Zhou B, Yang R, Ramachandran R, Abbott DW, Xiao TS (2019) Crystal structures of the full-length murine and human gasdermin D reveal mechanisms of autoinhibition, lipid binding, and oligomerization. Immunity 51(43–49):e4
Los FC, Randis TM, Aroian RV, Ratner AJ (2013) Role of pore-forming toxins in bacterial infectious diseases. Microbiol Mol Biol Rev 77:173–207
Lovelace LL, Cooper CL, Sodetz JM, Lebioda L (2011) Structure of human C8 protein provides mechanistic insight into membrane pore formation by complement. J Biol Chem 286:17585–17592
Mancheno JM, Martin-Benito J, Martinez-Ripoll M, Gavilanes JG, Hermoso JA (2003) Crystal and electron microscopy structures of sticholysin II actinoporin reveal insights into the mechanism of membrane pore formation. Structure 11:1319–1328
Mechaly AE, Bellomio A, Morante K, Gonzalez-Manas JM, Guerin DM (2009) Crystallization and preliminary crystallographic analysis of fragaceatoxin C, a pore-forming toxin from the sea anemone Actinia fragacea. Acta Crystallogr Sect F 65:357–360
Mondal AK, Chattopadhyay K (2020) Taking toll on membranes: curious cases of bacterial beta-barrel pore-forming toxins. Biochemistry 59:163–170
Mondal AK, Sreekumar A, Kundu N, Kathuria R, Verma P, Gandhi S, Chattopadhyay K (2018) Structural basis and functional implications of the membrane pore-formation mechanisms of bacterial pore-forming toxins. Adv Exp Med Biol 1112:281–291
Morante K, Caaveiro JM, Tanaka K, Gonzalez-Manas JM, Tsumoto K (2015) A pore-forming toxin requires a specific residue for its activity in membranes with particular physicochemical properties. J Biol Chem 290:10850–10861
Mueller M, Grauschopf U, Maier T, Glockshuber R, Ban N (2009) The structure of a cytolytic alpha-helical toxin pore reveals its assembly mechanism. Nature 459:726–730
Olson R, Gouaux E (2005) Crystal structure of the Vibrio cholerae cytolysin (VCC) pro-toxin and its assembly into a heptameric transmembrane pore. J Mol Biol 350:997–1016
Olson R, Nariya H, Yokota K, Kamio Y, Gouaux E (1999) Crystal structure of staphylococcal LukF delineates conformational changes accompanying formation of a transmembrane channel. Nat Struct Biol 6:134–140
Parker MW, Postma JP, Pattus F, Tucker AD, Tsernoglou D (1992) Refined structure of the pore-forming domain of colicin A at 2.4 A resolution. J Mol Biol 224:639–657
Paul K, Chattopadhyay K (2014) Pre-pore oligomer formation by Vibrio cholerae cytolysin: insights from a truncated variant lacking the pore-forming pre-stem loop. Biochem Biophys Res Commun 443:189–193
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Podobnik M, Savory P, Rojko N, Kisovec M, Wood N, Hambley R, Pugh J, Wallace EJ, McNeill L, Bruce M, Liko I, Allison TM, Mehmood S, Yilmaz N, Kobayashi T, Gilbert RJ, Robinson CV, Jayasinghe L, Anderluh G (2016) Crystal structure of an invertebrate cytolysin pore reveals unique properties and mechanism of assembly. Nat Commun 7:11598
Podobnik M, Kisovec M, Anderluh G (2017). Molecular mechanism of pore formation by aerolysin-like proteins. Philos Trans R Soc Lond B 372
Rai AK, Chattopadhyay K (2014) Trapping of Vibrio cholerae cytolysin in the membrane-bound monomeric state blocks membrane insertion and functional pore formation by the toxin. J Biol Chem 289:16978–16987
Ramirez-Carreto S, Miranda-Zaragoza B, Rodriguez-Almazan C (2020). Actinoporins: from the structure and function to the generation of biotechnological and therapeutic tools. Biomolecules 10
Reboul CF, Whisstock JC, Dunstone MA (2016) Giant MACPF/CDC pore forming toxins: a class of their own. Biochim Biophys Acta 1858:475–486
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucl Acids Res 42:W320–W324
Rojko N, Dalla Serra M, Macek P, Anderluh G (2016) Pore formation by actinoporins, cytolysins from sea anemones. Biochim Biophys Acta 1858:446–456
Ros U, Rodriguez-Vera W, Pedrera L, Valiente PA, Cabezas S, Lanio ME, Garcia-Saez AJ, Alvarez C (2015) Differences in activity of actinoporins are related with the hydrophobicity of their N-terminus. Biochimie 116:70–78
Rosado CJ, Kondos S, Bull TE, Kuiper MJ, Law RH, Buckle AM, Voskoboinik I, Bird PI, Trapani JA, Whisstock JC, Dunstone MA (2008) The MACPF/CDC family of pore-forming toxins. Cell Microbiol 10:1765–1774
Rossjohn J, Feil SC, McKinstry WJ, Tweten RK, Parker MW (1997) Structure of a cholesterol-binding, thiol-activated cytolysin and a model of its membrane form. Cell 89:685–692
Ruan J, Xia S, Liu X, Lieberman J, Wu H (2018) Cryo-EM structure of the gasdermin A3 membrane pore. Nature 557:62–67
Savinov SN, Heuck AP (2017). Interaction of cholesterol with perfringolysin O: what have we learned from functional analysis? Toxins (Basel) 9
Savva CG, Fernandes da Costa SP, Bokori-Brown M, Naylor CE, Cole AR, Moss DS, Titball RW, Basak AK (2013) Molecular architecture and functional analysis of NetB, a pore-forming toxin from Clostridium perfringens. J Biol Chem 288:3512–3522
Savva CG, Clark AR, Naylor CE, Popoff MR, Moss DS, Basak AK, Titball RW, Bokori-Brown M (2019) The pore structure of Clostridium perfringens epsilon toxin. Nat Commun 10:2641
Shi J, Gao W, Shao F (2017) Pyroptosis: gasdermin-mediated programmed necrotic cell death. Trends Biochem Sci 42:245–254
Song L, Hobaugh MR, Shustak C, Cheley S, Bayley H, Gouaux JE (1996) Structure of staphylococcal alpha-hemolysin, a heptameric transmembrane pore. Science 274:1859–1866
Sonnen AF, Plitzko JM, Gilbert RJ (2014) Incomplete pneumolysin oligomers form membrane pores. Open Biol 4:140044
Spicer BA, Conroy PJ, Law RHP, Voskoboinik I, Whisstock JC (2017) Perforin-A key (shaped) weapon in the immunological arsenal. Semin Cell Dev Biol 72:117–123
Tavares J, Amino R, Menard R (2014) The role of MACPF proteins in the biology of malaria and other apicomplexan parasites. Subcell Biochem 80:241–253
Tweten RK (2005) Cholesterol-dependent cytolysins, a family of versatile pore-forming toxins. Infect Immun 73:6199–6209
Valeva A, Palmer M, Hilgert K, Kehoe M, Bhakdi S (1995) Correct oligomerization is a prerequisite for insertion of the central molecular domain of staphylococcal alpha-toxin into the lipid bilayer. Biochim Biophys Acta 1236:213–218
van Pee K, Neuhaus A, D'Imprima E, Mills DJ, Kuhlbrandt W, Yildiz O (2017). CryoEM structures of membrane pore and prepore complex reveal cytolytic mechanism of Pneumolysin. Elife 6
Waldispuhl J, Berger B, Clote P, Steyaert JM (2006) Predicting transmembrane beta-barrels and interstrand residue interactions from sequence. Proteins 65:61–74
White SH, Ladokhin AS, Jayasinghe S, Hristova K (2001) How membranes shape protein structure. J Biol Chem 276:32395–32398
Wimley WC (2002) Toward genomic identification of beta-barrel membrane proteins: composition and architecture of known structures. Protein Sci 11:301–312
Wimley WC, Hristova K, Ladokhin AS, Silvestro L, Axelsen PH, White SH (1998) Folding of beta-sheet membrane proteins: a hydrophobic hexapeptide model. J Mol Biol 277:1091–1110
Yamada T, Yoshida T, Kawamoto A, Mitsuoka K, Iwasaki K, Tsuge H (2020) Cryo-EM structures reveal translocational unfolding in the clostridial binary iota toxin complex. Nat Struct Mol Biol 27:288–296
Yamashita D, Sugawara T, Takeshita M, Kaneko J, Kamio Y, Tanaka I, Tanaka Y, Yao M (2014) Molecular basis of transmembrane beta-barrel formation of staphylococcal pore-forming toxins. Nat Commun 5:4897
Funding
We acknowledge funding under the Centre of Excellence (COE) in the Frontier Areas of Science and Technology (FAST) program of the Ministry of Human Resource Development, Govt. of India, in the area of protein science, design, and engineering. We also thank Indian Institute of Science Education and Research Mohali for support; Research fellowships from the Council of Scientific and Industrial Research (CSIR), India (to A.K.M. and K.L.), University Grants Commission, India (to P.V. and S.C.), and Department of Biotechnology, India (to M.S.) are also acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that there is no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mondal, A.K., Verma, P., Lata, K. et al. Sequence Diversity in the Pore-Forming Motifs of the Membrane-Damaging Protein Toxins. J Membrane Biol 253, 469–478 (2020). https://doi.org/10.1007/s00232-020-00141-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-020-00141-2