Abstract
Macroecological studies have primarily focused on investigating the relationships between body size and geographic distribution on large scales, including regional, continental, and even global levels. While the majority of these studies have been conducted on terrestrial species, a limited number of studies have been carried out on aquatic species, and even fewer have considered the importance of phylogeny in the observed patterns. Cephalopods provide a good model for examining these macroecological patterns due to their large geographic and bathymetric ranges, wide range of body sizes, as well as diverse fin sizes and shapes. In this study, we assess the relationships between mantle length, fin size, and hatchling size with the geographic and bathymetric distribution of 30 squid species from the worldwide distributed family Loliginidae. To test a macroecological hypothesis, we evaluated the phylogenetic signal and correlated evolution to assess the role of biological traits in squid distribution, using a molecular phylogeny based on two mitochondrial and one nuclear genes. Biological traits (mantle length and fin size) exhibit high phylogenetic signals, while distribution demonstrates low signal. The correlation analyses revealed the existence of a relationship between adult mantle length and fin size with geographic and bathymetric distribution, but not with hatchling size. The geographic distribution of loliginid squids evolved in relation to mantle length, where larger squids with large fins (e.g. Sepioteuthis) have wide distributions, while small-finned species (e.g. Pickfordiateuthis) have narrow distributions. This study paves the way for exploring similar relationships in other squid families or other marine swimming animals.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The geographic distribution of species is determined by ecological and evolutionary factors, as well as the dispersal abilities of each species, and environmental conditions (Brown et al. 1996). The locomotion mode aids species in migration and dispersal to colonize new habitats, thereby expanding their geographic ranges. Species that have evolved specialized anatomical structures for long-distance movement, such as wings or fins, exhibit wide geographic distributions (e.g. birds, bats, fishes) (Böhning-Gaese et al. 2006; Laube et al. 2013; Luo et al. 2019). Recently, Luo et al. (2019) demonstrated that bat species with larger wings have larger distribution ranges compared to those with smaller wings and that the size of geographic ranges was associated with wing aspect ratio. That study highlights the relationship between specialized anatomical structures and distributions, suggesting the significant role of dispersal capacity in shaping species’ geographic distributions (Luo et al. 2019). In aquatic species, such as fishes and cephalopods, that utilize fins for locomotion, it is possible that the distribution patterns are going to be similar to those reported for birds and bats (Laube et al. 2013; Luo et al. 2019). On the other hand, in marine invertebrates, dispersal capabilities and consequently geographic distributions are influenced by larval size and movement capacities (Hansen 1980; Brown et al. 1996; Cowen and Sponaugle 2009; Villanueva et al. 2016; Ibáñez et al. 2018). Therefore, it is pertinent to investigate whether body size, fins, or larval dispersal influences the size of geographic ranges in marine invertebrates.
Cephalopods are exclusively marine animals that include nautilids, cuttlefishes, squids, and octopods, with approximately 860 species distributed across 50 families and 174 genera (Hoving et al. 2014; Luna et al. 2021). Some species reach large sizes as adults, enabling them to have wide geographic distributions spanning over 5000 km (Rosa et al. 2008a, b; Ibáñez et al. 2009, 2019; Rosa et al. 2012, 2019). They primarily inhabit the first 1000 m depth and have daily vertical migrations (Boyle and Rodhouse 2005). The combination of several characteristics, such as wide distribution, jet propulsion, high dispersal, and wide range of adult body size, makes cephalopods a good model for testing biogeographic hypotheses.
Since all cephalopods share funnel and jet propulsion, the distinct size and shape of fins could be determining factors in their movement capabilities (Clarke 1988). There are nine major fin shapes among squids, all of which contribute to fast swimming and orientation control (following Clarke 1988; Boyle and Rodhouse 2005). These fins are muscular hydrostats with an intramuscular network of crossed connective tissue fibres that provide support for fin movements (Johnsen and Kier 1993). Among pelagic squids, those belonging to the Loliginidae family display diverse fin morphologies, a wide amplitude of body sizes in both juveniles and adults, and high variability of geographic and bathymetric distribution (Anderson 2000a; Jereb et al. 2010). These traits make them an excellent study model to understand the factors that explain their distribution. Loliginidae contains species which can reach a rather large size (at least 900 mm of mantle length, ML, in Loligo forbesii), along with dwarf species like Pickfordiateuthis, where females can mature up to 7.9 mm ML (Brakoniecki 1996). These benthopelagic squids have a pelagic paralarval stage, form schools, are active swimmers, and are chasing predators (Nesis 1980). Loliginid squids have elongated flapping fins that produce large-amplitude waves for economical, gentle swimming and hovering, as well as for controlling stability and aiding jet escape (Clarke 1988). By combining finning and jetting, cephalopods can generate different swimming gaits (Anderson and DeMont 2000; Stewart et al. 2010).
To understand the historical and ecological processes that influence the distributions of species, it is necessary to integrate comparative-quantitative biogeographic and phylogenetic studies (Brown et al. 1996; Hernández et al. 2013). In this study, we aim to test predictions of macroecology based on data about the distribution, fin characteristics (shape and size), and mantle length of both hatchling and adult loliginid squids worldwide. To achieve this, we collected data on distribution (latitudinal range, area, and depth) and biological traits (mantle length of young and adults and fin size) of loliginids. Additionally, with inferred the phylogenetic relationships of the loliginids to estimate the phylogenetic correlation between these traits. This study aims to test whether the geographic distribution can be predicted by dispersal capacity, inferred from mantle length and fin size, among loliginids within a phylogenetic comparative framework.
Materials and methods
Database
In this study, we included 30 species of loliginid squids from ten genera out of the 47 reported by Jereb et al. (2010). The data collected included squid mantle lengths (maximum mantle length, ML, mm, Fig. S1) as indicator of body size, hatchling size (ML), fin length (FL, mm, Fig. S1), and distribution (latitudinal range, bathymetric range, and area distribution). Fin shapes were classified as: rhomboid, round, or elliptic. In several loliginid species, males exhibit larger ML than females, while rarely females are larger (Jereb et al. 2010). This indicates different jet swimming capacities between genders. Additionally, some species, the maximum ML corresponds to only one gender. To explore the effect of mantle length gender on distribution, we conducted analyses separating the data based on gender when enough data were available. To avoid allometric effects, we transformed the fin length into fin length index (FLI, Roper and Voss 1983) (see supplementary material, Table S1). The distribution areas for all different species were estimated using maps from the literature (Jereb et al. 2010). To ensure accuracy, we georeferenced these maps using the open-source GIS software QGIS 3.0.1 (QGIS Development Team 2018) with the plugin Georeferencer GDAL 3.1.9. This process assigned spatial information to each pixel in a map, relating it to a coordinate in geographic space. From the georeferenced maps, we recreated the distribution area as a polygon shapefile, from which we obtained its latitudinal range (in degrees) and area (km2) with the plugin Calculate Geometry 0.3.2. We also recorded maximum depth of each species from the literature, to assess the bathymetric distribution range (m). This database was combined with a new molecular phylogeny reconstructed with data from GenBank (Table S2).
To assess the relationship between distributional data (latitudinal ranges, bathymetric range, and area distribution) and fin shape (rhomboid, round, and elliptic), we performed one-way ANOVA in R v4.1.2 core (R Development core team 2022), after log transformation of the data.
Phylogenetic analysis
Nucleotide sequences (16S, COI, and RHO) for each gene were separately aligned using Multiple Sequence Comparison by Log-Expectation (MUSCLE) with default parameters for gap insertion and gap extension (Edgar 2004), implemented in the MEGA X software (Kumar et al. 2018). The best-supported substitution model for each gene was identified prior to the phylogenetic analysis using jModelTest2 (Darriba et al. 2012) (Table S3). Once aligned, the sequences for each of the three genes were concatenated into a single partitioned matrix using Mesquite v3.10 (Maddison and Maddison 2016), which allowed for a separate substitution model to be used for each gene (Table S2).
We estimated phylogenies for the partitioned dataset using Bayesian Inference with MrBayes v3.2 software (Ronquist et al. 2012). Bayesian phylogenetic inference was performed using 10,000,000 generations of four heated Markov chain Monte Carlo (MCMC), sampling every 1,000 generations. We discarded the first 10% (1,000,000) of generations as burn-in, resulting in a total of 9,001 trees sampled from the posterior probability distribution. A majority consensus tree (50%) was computed from these 10,000 trees. We also evaluated convergence to the posterior distribution and mixing of the MCMC using Tracer v1.6 (Rambaut et al. 2014). Effective sample size (ESS) > 200 was accepted. The trees were rooted using Spirula spirula (Owen, 1836), Ommastrephes bartramii (Lesueur, 1821), and Sthenoteuthis oualaniensis (Lesson, 1830) as outgroups. Certain cryptic lineages (i.e. Doryteuthis pleii, D. pealeii, Lolliguncula brevis, and Sepioteuthis lessoniana) lacked morphometric and distribution data. Only the lineages with the most data (and their respective distribution) were included in the phylogeny in order to avoid obscuring the results with cryptic species or genetic lineages (Okutani 1984, 2005; Segawa et al. 1993; Cheng et al. 2014).
For comparative purposes, all outgroup species (three tree tips, S. spirula, O. bartramii, and S. oualaniensis) were removed from the tree using the drop.tip function in “APE” package (Paradis et al. 2004), implemented in R v4.1.2 core (R Development Core Team 2022). The new phylogram was transformed into an ultrametric tree using the Grafen’s (1989) method. To explore the association between the standardized database and the phylogeny, a heatmap was employed using the phylo.heatmap function in “Phytools” package (Revell 2012).
Phylogenetic signal
The ultrametric tree was used to estimate the phylogenetic signal of each trait, denoted as Pagel’s λ (Pagel 1999). Lambda (λ) varies between 0 and 1, quantifying the amount of phylogenetic signal in the studied trait. A value of λ = 0 indicates that the trait distribution across species is independent of the phylogeny, while λ = 1 suggests that the distribution of trait values conforms to the expectations of the Brownian motion model (Pagel 1999). The analyses were conducted using the phylosig function, implemented in “Phytools”. The likelihood value of λ estimated for each trait was compared to the likelihood value of λ equal to 0 through likelihood ratio test (LRT).
To determine whether fin shape exhibits a phylogenetic signal, we employed the “phylo.signal.disc” algorithm, comparing the number of transitions according to unrestrained parsimony against a null distribution obtained by randomizing the species data, effectively disrupting any underlying phylogenetic structure (Rezende and Diniz-Filho 2012). The null distribution was based on 1000 replicates and was implemented in R.
Correlated evolution
We used phylogenetic generalized least squares models (PGLS; Pagel 1999) to establish the existence of a linear relationship between ML (females, males, and hatchlings), FLI, and distribution data (latitudinal, bathymetric, and area). These models were performed using the corPagel function in the “APE” package in R. The correlation structure of PGLS was based on the assumption of Brownian motion model, multiplying the off-diagonal elements (the covariances) by λ. To compare all predictor variables, we calculate IC 95% for coefficients of all PGLS analyses.
Results
Biological data
The mantle length of the 30 loliginid species ranged from 22 mm in Pickfordiateuthis pulchella to 937 mm in L. forbesii, whereas fin length index ranged from 23 in Alloteuthis africana to 90 in Sepioteuthis lessoniana (Table S1). The geographic distribution ranged from 73,729 km2 in Lolliguncula argus up to 28,000,000 km2 in S. lessoniana (Table S1).
Among the distributional data, the only significant relationship was found between the distribution area compared with fin shape (F2,27 = 5.158, P = 0.0127); the species with elliptic shape had the larger distribution (see Fig. 1).
Phylogenetic signal
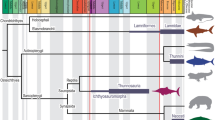
Both squids’ mantle length (ML) (female and male) and fin length index (FLI) were highly structured across the phylogeny in the heatmap (Fig. 2), with both traits exhibiting a high significant phylogenetic signal (λ > 0.75, P < 0.05; Table 1). These results provide evidence of a concordance between loliginid phylogeny and both biological traits evaluated. All distribution data showed a low phylogenetic signal (λ < 0.70, P > 0.5; Table 1) and an absence of phylogenetic structure (Fig. 2).
Fin shape shows a higher correspondence with the phylogeny of loliginids (Fig. 3). The “phylo.signal.disc” algorithm for fin shape shows that five transitions were required to obtain the observed distribution along the loliginid phylogeny (Fig. 3). This result differed significantly (P < 0.001) from the median number of transitions (11; range 7–12) across the 1000 replicates employed to build the null distribution.
Correlated evolution
The largest loliginid squids have wider distribution, indicating a positive correlation with area distribution and bathymetric distribution (Table 2, Fig. 4). However, neither female nor male ML correlated with the latitudinal range (β < 0.12, Table 2, Fig. 4). The PGLS analysis showed a significant correlation between female and male ML with geographic distribution (β > 0.75, Table 2, Fig. 4). Similarly, FLI was positively and significantly correlated with area distribution and bathymetric range (β > 1.0, Table 2, Fig. 5), but not with latitudinal range (β < 0.1, Table 2, Fig. 5). Hatchling size did not show a correlation with the distribution data (Table 2, Fig. 5). Among all predictor traits (body size, hatchling size, and fin length), the best predictor of geographic and bathymetric distribution of loliginid squids was FLI (β = 1.4–2.4, Table 2, Fig. 6). Finally, the box-and-whisker plot does not show clear differences between female and male coefficients for all PGLS analyses.
Plots illustrating the relationships between female and male mantle length and distributional traits of loliginid squids, where A female mantle length/latitudinal distribution, B female mantle length/geographic distribution, C female mantle length/bathymetric distribution, D male mantle length/latitudinal distribution, E male mantle length/geographic distribution, and F male mantle length/bathymetric distribution. Dashed lines represent regressions from PGLS
Plots depicting the relationships between biological and distributional traits of loliginid squids, where A fin length index/latitudinal distribution, B fin length index/geographic distribution, C fin length index/bathymetric distribution, D hatchling mantle length/latitudinal distribution, E hatchling mantle length/geographic distribution, and F hatchling mantle length/bathymetric distribution. Dashed lines represent regressions from PGLS
Discussion
Geographic range size, which refers to the extent of a species’ occurrence, is a fundamental biogeographic variable influenced by several intrinsic and extrinsic factors (Brown et al. 1996; Gaston 2003). In the case of loliginid species, we observed that larger species with larger fin sizes have broader bathymetric and geographic distribution, akin to the positive correlation between wings and distribution observed in birds and bats (Böhning-Gaese et al. 2006; Laube et al. 2013; Luo et al. 2019). This macroecological relationship suggests exploring these patterns in other squid families or marine swimming animals (e.g. fishes) to understand whether the geographic range size is related to these and other biological traits.
The geographic distribution, encompassing area and latitudinal and bathymetric range, does not exhibit a significant phylogenetic signal throughout the family. This contradicts the statements made by other authors (Brakoniecki 1986; Anderson 2000a, b; Ulloa et al. 2017), who posited that geographically close species share common ancestors. However, our data are quantitative (i.e. km2), not qualitative (i.e. biogeographic provinces), and thus do not exhibit significant phylogenetic signal. In contrast, biological traits (i.e. body size and fins) have a strong phylogenetic signal, as suggested by other studies on cephalopods (Ibáñez et al. 2018; 2021; Anderson and Marian 2020).
The geographic distribution of loliginid squids is suggested to be influenced by their environment and ecology (Sales et al. 2013, 2017; Costa et al. 2021). One crucial factor is their dispersion capacity. For certain American Loliginid species from the Doryteuthis and Lolliguncula genera, the Amazon plume Barrier (Floeter et al. 2008) does not impede the dispersion of D. pleii, D. pealeii, and L. brevis above the major influence area of the plume (between Pará and Amapá states, and North Coast of Brazil, Muller-Karger et al. 1988; Hoorn et al. 2010), despite the significant reduction in salinity. However, low-salinity environments can act as barriers for both adults and paralarvae, resulting in high mortality (Hanlon and Messenger 1996; Hanlon 1998; Boyle and Rodhouse 2005). Nevertheless, the considerable capacity of dispersion of loliginid squids is evidenced by that these species do in fact disperse above the plume area of influences (Boyle and Rodhouse 2005; Jereb et al. 2010).
Body size and distribution
Our results show that loliginid squids with larger body size, both females and males, exhibit a larger geographic distribution, while smaller species have a restricted distribution, similar to some marine fishes (Hernández et al. 2013). However, the swimming capacity of squids differs that of fishes. For millions of years, cephalopods roamed the world’s oceans as jet-propelled masters of the pelagic world, until fishes, using highly efficient undulatory locomotion, displaced them from many nektonic habitats (Hoar et al. 1991). According to Nesis (1980), the evolution of squids, such as Loliginidae, has been strongly influenced by competition with fishes. In fact, their biology, distribution, and geographic variation are determined by direct competition between species that occupy different ecologic habitats (depth and thermal preferences) and exhibit inverse circadian activity levels (Martins and Juanicó 2018). Indeed, the directional jet propulsion of the cephalopods provides better acceleration and manoeuvrability than many fishes (Foyle and O’Dor 1988). Combined with lift production by the fins, squids may have more dynamic lift capabilities (Hoar et al. 1991). In this study, the positive relationship between mantle length, fin length, bathymetric range, latitudinal range, and distribution area suggests that dispersal capacity plays a role in shaping species’ geographic distributions. When there are no geographic or ecological barriers preventing it, larger squid will have greater dispersal capacity, enabling them to reach a wider distribution range.
The geographic distribution of loliginids appears to have evolved in relation to body size. Larger squids (Sepioteuthis) have wide distributions, while smaller species (Pickfordiateuthis, Afrololigo) have narrow distributions (Fig. 2). Previous studies in vertebrates have detected a positive relationship between geographic range size and body size (Diniz-Filho and Tôrres 2002; Hernández et al. 2013), which is consistent with the relationships between mantle length and geographic distribution in our results (Fig. 4). However, the existence of significant variation in the sizes of geographic ranges among small species is a common pattern in macroecology (Brown 1995). Species falling below to regression line (ML vs distribution) have a smaller spatial distribution than predicted by its body size, for instance, L. argus, a coastal species characterized by its small size at maturity (Jereb and Roper 2010), which would decrease its potential final size. This species reaches the smallest area of all the studied loliginid species (73,729 km2). A similar case occurs in the small species that inhabits sandy-mud bottoms Afrololigo mercatoris, which shares many morphological traits with Lolliguncula (Jereb et al. 2010), such as the small size.
Size and shape have important implications for the hydrodynamics of marine organisms (DeMont and Hokkanen 1992). The absence of shared common ancestors along geographically close species may be determined by the biology and dispersal ability of each species. Indeed, the high mobility of adult loliginids (Boyle and Rodhouse 2005) makes their dispersal more widespread. A notable case is that of the three neritic species of the genus Loligo, which have a high dispersal ability, and an evident difference between the maximum ML of their males, which could be one of the reasons for habitat choice (L. forbesii, L. vulgaris, and L. reynaudii; Jereb et al. 2010; Iwata et al. 2018). Similarly, in adults of A. africana and Uroteuthis bartschi (maximum ML of 200 mm in males, 150 mm in females; Voss 1963; Jereb et al. 2010), there is a remarkable sexual dimorphism, where males display the characteristic extremely long and spike-like tail (73% of the dorsal ML in A. Africana adults) as secondary sexual trait. However, our results did not reveal a strong difference in distribution related to ML of females and males. In this line, both genders have the same dispersal capacity and sexual dimorphism could be related to other factors (e.g. growth, reproduction).
Fins shape/length and distribution
Fin length and shape exhibit a highly significant phylogenetic signal, demonstrating the existing concordance between the fin size and shape and loliginid phylogeny. Fins shape undergoes changes during ontogenetic growth, with hatching squid having fins proportionally smaller than those of adults. This change in relative size may also reflect a shift in the use of the fins, similar to the different usages of the variously shaped adult fins (Hoar et al. 1991). The shape of the lateral fins of cephalopods varies in accordance with the animal’s size and lifestyle (Packard 1972), with hydrodynamic constraints being primary determinant of fin shape (Daniel 1988). For species such as from the genus Uroteuthis, fins play an efficient role in maintaining position in the water column. As growth occurs, and the animal transitions from a viscosity-dominant to an inertia-dominant system, the development of the fin structure becomes crucial, with shape further influencing friction and pressure drag on animals during movement (Moltschaniwskyj 1995).
Fin size emerges as the most important biological trait for predicting the distribution of loliginid squids (Fig. 6), with species possessing the largest elliptical fins (i.e. Sepioteuthis) exhibiting wider distributions. Sepioteuthis lessoniana, widely distributed throughout the Indo-Pacific and Mediterranean regions, is the species with the broadest range of distribution among these squids. The genus features very large and distinctive fins, broadly oval in outline and with a fin length comprising over 90% up to nearly 100% of ML (Jereb et al. 2010), making them markedly different from other loliginids. In contrast, Sepioteuthis australis, distributed in the southwestern Pacific Ocean, is distinguished from S. lessoniana by the angular lateral margins of its fins, although not by the fin length (Jereb et al. 2010). The distant geographic distribution of both species may explain this contrasting phylogenetic footprint. On the other hand, the loliginids A. africana and L. argus have the smallest FLI and limited distribution. Among the medium-sized, continental shelf species, Uroteuthis (P.) edulis exhibits a broad fin (70% ML in length, 60% ML in width), as does its congener Uroteuthis (P.) chinensis with a fin spreading to two-thirds of the ML. However, both Uroteuthis have a less extensive range of distribution than it would be expected based solely on their fin size.
Another important factor affecting the dispersal capacity of the loliginids is the speed they can achieve in swimming and their manoeuvrability, combining finning and jetting to generate different swimming gaits (Anderson and DeMont 2000; Stewart et al. 2010). For instance, there are differences between the locomotive repertoire and the high degree of manoeuvrability enabled by fin propulsion of Loliguncula brevis (Bartol et al. 2001) than the extensive use of the fins for swimming of the ommastrephid Illex illecebrosus (Harrop et al. 2014). In the first species, its swimming gaits could help them achieve a distributional range similar to that of larger loliginids. The high diel vertical migration behavioural flexibility expressed by L. forbesii could be very advantageous in terms of energy conservation, prey capture, and predator avoidance (Cones et al. 2022), leading to greater evolutionary success and, consequently, a larger dispersal capacity than expected for its size.
Hatchling size and distribution
Marine mollusc species lacking a planktonic larval phase in their life cycle tend to have smaller size ranges compared to species with more readily dispersed planktotrophic larvae (Brown et al. 1996; Villanueva et al. 2016; Ibáñez et al. 2018). In fact, some Neogastropoda snails exhibit wide distributions, extended gene flow, and resistance to isolation, resulting in greater species longevity for species with planktonic larvae (Hansen 1980). Loliginids, on the other hand, have a pelagic paralarval stage during which they spend two to three months in the plankton (Garcia-Mayoral et al. 2020). During this time, they disperse but remain on the continental shelf by controlling their vertical position (Roura et al. 2016, 2019). Consequently, this group tends to have large size ranges, leading to extensive geographic distribution (Villanueva et al. 2016). Some squids exhibit very high dispersal rates, due to lengthy planktonic paralarval stages and highly migratory adult stages, leading some authors to predict panmixia (genetic homogeneity) of squid populations across large geographic areas (e.g. Shaw et al. 2010). Nonetheless, many loliginid squids show structured population at large scales (Brierley et al. 1995; Shaw et al. 1999; Herke and Foltz 2002; Aoki et al. 2008; Ibáñez et al. 2012). This pattern is associated with reduced paralarval dispersion, since eggs are deposited on the seabed or attached on sessile organisms (e.g. kelps, corals), which promotes structure between populations (Carrasco et al. 2021).
Despite all this, our results show that species of loliginid squids (both females and males) falling above the regression line (ML vs distribution, Fig. 4) have larger distribution ranges than predicted by their body size. This pattern could be associated with paralarval dispersion, but our results do not support the idea that paralarvae dispersion explains the wide distribution of some species, as our analyses did not find a correlation between hatchling size and distribution data (Fig. 5). This might indicate a reduction in distribution, or it could suggest that our knowledge of their distribution is still lacking.
In this comparative study, we evaluated the potential effects of body size (young and adults) and fins (shape and size) of squids as predictors of their distribution using a phylogenetic approach. This approach has been scarcely used in macroecological studies on marine animals, particularly in the assessment of phylogenetic signal on geographic distribution (i.e. Hernández et al. 2013; Ulloa et al. 2017; Ibáñez et al. 2018). In this sense, our research is among the first studies correctly address trait comparisons for marine animals, suggesting that further research should incorporate this approach in macroecological studies. New approaches to the knowledge of the distributional range of mobile species, such as the bio-logging tags (Flaspohler et al. 2019; Cones et al. 2022), or eDNA could help in accurate assessments of the real extent of the species, as well as their biological activity and behavioural patterns.
Data availability
All data of this article is in supplementary files.
References
Anderson FE (2000a) Phylogenetic relationships among loliginid squids (Cephalopoda: Myopsida) based on analyses of multiple data sets. Zool J Linnean Soc 130:603–633
Anderson FE (2000b) Phylogeny and historical biogeography of the loliginid squids (Mollusca: Cephalopoda) based on mitochondrial DNA sequence data. Mol Phylogenet Evol 15(2):191–214
Anderson EJ, Demont ME (2000) The mechanics of locomotion in the squid Loligo pealei: locomotory function and unsteady hydrodynamics of the jet and intra mantle pressure. J Exp Biol 203(18):2851–2863
Anderson FE, Marian JEA (2020) The grass squid Pickfordiateuthis pulchella is a paedomorphic loliginid. Mol Phylogenet Evol 147:106801
Aoki M, Imai H, Naruse T, Ikeda Y (2008) Low genetic diversity of oval squid, Sepioteuthis cf. lessoniana (Cephalopoda: Loliginidae), in Japanese waters inferred from a mitochondrial DNA non-coding region. Pac Sci 62:403–411
Bartol IK, Patterson MR, Mann R (2001) Swimming mechanics and behavior of the shallow-water brief squid Lolliguncula brevis. J Exp Biol 204(21):3655–3682
Böhning-Gaese K, Caprano T, Van Ewijk K, Veith M (2006) Range size: disentangling current traits and phylogenetic and biogeographic factors. Am Nat 167:555–567
Boyle P, Rodhouse P (2005) Cephalopods. Ecology and fisheries. Blackwell Publishing, Oxford
Brakoniecki TF (1996) A revision of the genus Pickfordiateuthis Voss, 1953 (Cephalopoda; Myopsida). Bull Mar Sci 58(1):9–28
Brakoniecki TF (1986) A generic revision of the family Loliginidae (Cephalopoda: Myopsida) based primarily on the comparative morphology of the hectocotylus. Ph.D. Dissertation, University of Miami, Coral Gables, FL. p 163.
Brierley AS, Thorpe JP, Pierce GJ, Clarke MR, Boyle PR (1995) Genetic variation in the neritic squid Loligo forbesi (Myopsida: Loliginidae) in the northeast Atlantic Ocean. Mar Biol 122:79–86
Brown JH (1995) Macroecology. University of Chicago Press, Chicago
Brown JH, Stevens GC, Kaufman DM (1996) The geographic range: size, shape, boundaries, and internal structure. Annu Rev Ecol Evol Syst 27:597–423
Carrasco SA, Bravo M, Ibáñez CM, Zapata-Hernández G (2021) Discrete spawning aggregations of the loliginid squid Doryteuthis gahi reveal life-history interactions of a dwarf morphotype at the center of its distribution range. Front Mar Sci 7:1251
Cheng SH, Anderson FE, Bergman A, Mahardika GN, Muchlisin ZA, Dang BT, Calumpong HP, Mohamed KS, Sasikumar G, Venkatesan V, Barber PH (2014) Molecular evidence for co-occurring cryptic lineages within the Sepioteuthis cf. lessoniana species complex in the Indian and Indo-West Pacific Oceans. Hydrobiologia. 725(1):165–188
Clarke MR (1988) Evolution of buoyancy and locomotion in recent cephalopods. paleontology and neontology of Cephalopoda. The Mollusca 12:203–213
Cones SF, Zhang D, Shorter KA, Katija K, Mann DA, Jensen FH, Fontes J, Afonso P, Mooney TA (2022) Swimming behaviors during diel vertical migration in veined squid Loligo forbesii. Mar Ecol Prog Ser 691:83–96
Costa TA, Sales JB, Markaida U, Granados-Amores J, Gales SM, Sampaio I, Vallinoto M, Rodrigues-Filho LFS, Ready JS (2021) Revisiting the phylogeny of the genus Lolliguncula Steenstrup 1881 improves understanding of their biogeography and proves the validity of Lolliguncula argus Brakoniecki & Roper, 1985. Mol Phylogenet Evol 154:106968
Cowen RK, Sponaugle S (2009) Larval dispersal and marine population connectivity. Annu Rev Mar Sci 1(1):443–466
Daniel TL (1988) Forward flapping flight from flexible fins. Can J Zool 66(3):630–638
DeMont ME, Hokkanen JEI (1992) Hydrodynamics of animal movement. In: Biewener AA (ed) Biomechanics (structures and systems): a practical approach. Oxford University Press, New York, pp 263–284
Diniz-Filho JAF, Tôrres NM (2002) Phylogenetic comparative methods and the geographic range size–body size relationship in New World terrestrial Carnivora. Evol Eco. 16:351–367
Flaspohler GE, Caruso F, Mooney TA, Katija K, Fontes J, Afonso P, Shorter KA (2019) Quantifying the swimming gaits of veined squid (Loligo forbesii) using bio-logging tags. J Exp Biol 222(24):jeb198226
Floeter SR, Rocha LA, Robertson DR, Joyeux JC, Smith-Vaniz WF, Wirtz P, Edwards AJ, Barreiros JP, Ferreira CEL, Gasparini JL, Brito A, Falcón JM, Bowen BW, Bernardi G (2008) Atlantic reef fish biogeography and evolution. J Biogeogr 35:22–47
Foyle TP, ’O’Dor RK, (1988) Predatory strategies of squid (Illex illecebrosus) attacking small and large fish. Mar Freshw Behav Physiol 13(2):155–168
Garcia-Mayoral E, Roura Á, Ramilo A, Gonzalez AF (2020) Spatial distribution and genetic structure of loliginid paralarvae along the Galician coast (NW Spain). Fish Res 222:105406
Gaston KJ (2003) The structure and dynamics of geographic ranges. Oxford University Press
Grafen A (1989) The phylogenetic regression. Philos Trans R Soc Lond B Biol Sci 326:119–157. https://doi.org/10.1098/rstb.1989.0106
Hanlon RT (1998) Mating systems and sexual selection in the squid Loligo: how might commercial fishing on spawning squids affect them? Calif Coop Ocean Fish Investig Rep 39:92–100
Hanlon RT, Messenger JB (1996) Cephalopod Behaviour. Cambridge University Press, New York
Hansen TA (1980) Influence of larval dispersal and geographic distribution on species longevity in neogastropods. Paleobiology 6(2):193–207
Harrop J, Vecchione M, Felley JD (2014) In situ observations on behaviour of the ommastrephid squid genus Illex (Cephalopoda: Ommastrephidae) in the northwestern Atlantic. J Nat Hist 48(41–42):2501–2516. https://doi.org/10.1080/00222933.2014.937367
Herke SW, Foltz DW (2002) Phylogeography of two squid (Loligo pealei and L. plei) in the Gulf of Mexico and northwestern Atlantic Ocean. Mar Biol 140:103–115
Hernández CE, Rodríguez-Serrano E, Avaria-Llautureo J, Inostroza-Michael O, Morales-Pallero B, Boric-Bargetto D et al (2013) Using phylogenetic information and the comparative method to evaluate hypotheses in macroecology. Methods Ecol Evol 4:401–415
Hoar JA, Sim E, Webber DM, O’Dor RK (1991) The role of fins in the competition between squid and fish. In: Maddock L, Bone Q, Rayner JMV (ed) Mechanics and physiology of animal swimming. Pp 27–43. ISSN/ISBN: 0-521-46078-6.
Hoorn C, Wesselingh FP, Hovikoski J, Guerrero J (2010) The development of the iogeogra mega-wetland (Miocene; Brazil, Colombia, Peru, Bolivia). In: Hoorn C, Wesselingh FP (eds) Amazonia, landscape and species evolution: a look into the past. Wiley, New York, pp 123–142. https://doi.org/10.1002/9781444306408.ch8
Ibáñez CM, Camus PA, Rocha R (2009) Diversity and distribution of cephalopod species of the coast off Chile. Mar Biol Res 5:374–384
Ibáñez CM, Argüelles J, Yamashiro C, Adasme L, Céspedes R, Poulin E (2012) Spatial genetic structure and demographic inference of the Patagonian squid Doryteuthis gahi in the Southeastern Pacific Ocean. J Mar Biol Ass UK 92:197–203
Ibáñez CM, Rezende E, Sepúlveda RD, Avaria-Llautureo J, Hernández CE, Sellanes J, Poulin E, Pardo- Gandarillas MC (2018) Thorson’s rule, life history evolution and diversification of benthic octopuses (Cephalopoda: Octopodoidea). Evolution 72–9:1829–1839
Ibáñez CM, Braid H, Carrasco SA, López-Córdova D, Torretti G, Camus PA (2019) Zoogeographic patterns of pelagic oceanic cephalopods along the eastern Pacific Ocean. J Biogeogr 46:1260–1273
Ibáñez CM, Díaz-Santana-Iturrios M, Carrasco SA, Fernández-Álvarez FA, López-Córdova DA, Cornejo CF, Ortiz N, Rocha F, Vidal EAG and Pardo-Gandarillas MC (2021) Macroevolutionary trade-offs and trends in life history traits of cephalopods through a comparative phylogenetic approach. Front Mar Sci 8:707825. https://doi.org/10.3389/fmars.2021.707825
Iwata Y, Sauer WH, Sato N, Shaw PW (2018) Spermatophore dimorphism in the chokka squid Loligo reynaudii associated with alternative mating tactics. J Molluscan Stud 84(2):157–162
Jereb P, Vecchione M, Roper CFE (2010) Family Loliginidae. In: Jereb P, Roper CFE (eds) Cephalopods of the world. An annotated and illustrated catalogue of species known to date Myopsid and Oegopsid Squids. FAO Species Catalogue for Fishery Purposes No 2, vol 2. FAO, Rome, pp 38–117
Johnsen S, Kier WM (1993) Intramuscular crossed connective tissue fibres: skeletal support in the lateral fins of squid and cuttlefish (Mollusca: Cephalopoda). J Zool, London 231:311–338
Laube I, Korntheuer H, Schwager M, Trautmann S, Rahbek C, Böhning-Gaese K (2013) Towards a more mechanistic understanding of traits and range sizes. Global Ecol Biogeogr 22(2):233–241
Luna A, Rocha F, Perales-Raya C (2021) A review of cephalopods (Phylum: Mollusca) of the Canary Current Large Marine Ecosystem (Central-East Atlantic, African coast). J Mar Biol Assoc UK 101:1–25
Luo B, Santana SE, Pang Y, Wang M, Xiao Y, Feng J (2019) Wing morphology predicts geographic range size in vespertilionid bats. Sci Rep 9:4526. https://doi.org/10.1038/s41598-019-41125-0
Martins RS, Juanicó M (2018) Biology, distribution and geographic variation of loliginid squids (Mollusca: Cephalopoda) off southwestern Atlantic. Zoologia (curitiba) 35:1–16
Moltschaniwskyj NA (1995) Multiple spawning in the tropical squid Photololigo sp.: what is the cost in somatic growth? Mar Biol 124:127–135
Muller-Karger FE, McClain CR, Richardson P (1988) The dispersal of Amazon’s water. Nature 333:56–59
Nesis KN (1980) Sepiids and Loliginids: A comparative review of the distribution and evolution of neritic cephalopods. Zool Zhurnal 59(5):677–688
Okutani T (1984) Life history of Aori-ika (Sepioteuthis lessoniana). Saibai Giken 13:69–75
Packard A (1972) Cephalopods and fish: the limits of convergence. Biol Rev 47(2):241–307
Pagel M (1999) Inferring the historical patterns of biological evolution. Nature 401:877–884
R Development Core Team (2022) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Rambaut A, Suchard M, Xie W, Drummond A (2014) Tracer v. 1.6. Institute of Evolutionary Biology, University of Edinburgh, U.K.
Revell LJ (2012) phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol Evol 3:217–223
Rezende EL, Diniz-Filho JA (2012) Phylogenetic analyses: comparing species to infer adaptations and physiological mechanisms. Compr Physiol 2:639–674
Ronquist F, Teslenko M, Van Der Mark P, Ayres DL, Darling A, Höhna S et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3):539–542
Roper CFE, Voss GL (1983) Guidelines for taxonomic descriptions of cephalopod species. Mem Mus Vic 44:48–63
Rosa R, Dierssen HM, González L, Seibel BA (2008a) Ecological biogeography of Cephalopod Molluscs in the Atlantic Ocean: historical and contemporary causes of coastal diversity patterns. Glob Ecol Biogeogr 17:600–610
Rosa R, Dierssen HM, González L, Seibel BA (2008b) Large-scale diversity patterns of cephalopods in the Atlantic open ocean and deep-sea. Ecology 89:3449–3461
Rosa R, González L, Dierssen HM, Seibel BA (2012) Environmental determinants of latitudinal size-trends in cephalopods. Mar Ecol Prog Ser 464:153–165
Rosa R, Pissarra V, Borges FO, Santos C, Xavier J, Gleadall IG, Golikov A, Bello G, Morais L, Lishchenko F, Roura Á, Judkins H, Ibáñez CM, Piatkowski U, Vecchione M, Villanueva R (2019) Global patterns of species richness in coastal cephalopods. Front Mar Scienc 6:469
Roura Á, Álvarez-Salgado XA, González ÁF, Gregori M, Rosón G, Otero J, Guerra Á (2016) Life strategies of cephalopod paralarvae in a coastal upwelling system (NW Iberian Peninsula): insights from zooplankton community and spatio-temporal analyses. Fish Oceanogr 25:241–258
Roura Á, Amor M, González ÁF, Guerra Á, Barton ED, Strugnell JM (2019) Oceanographic processes shape genetic signatures of planktonic cephalopod paralarvae in two upwelling regions. Progr Oceanogr 170:11–27
Sales JB, Shaw PW, Haimovici M, Markaida U, Cunha DB, Ready JS, Figueiredo-Ready WM, Angioletti F, Schneider H, Sampaio I (2013) New molecular phylogeny of the squids of the family Loliginidae with emphasis on the genus Doryteuthis Naef, 1912: mitochondrial and nuclear sequences indicate the presence of cryptic species in the southern Atlantic Ocean. Mol Phylogenet Evol 68(2):293–299
Sales JBL, Rodrigues-Filho LFS, Ferreira YS, Carneiro J, Asp NE, Shaw PW, Haimovici M, Markaida U, Ready J, Schneider H, Sampaio I (2017) Divergenceof cryptic species of Doryteuthis plei Blainville, 1823 (Loliginidae, Cephalopoda) in the Western Atlantic Ocean is associated with the formation of the Caribbean Sea. Mol Phylogenet Evol 106:44–54
Segawa S, Hirayama S, Okutani T (1993) Is Sepioteuthis lessoniana in Okinawa a single species? In: Okutanitodor RK, Kubodera T (eds) Recent advances in Cephalopod fisheries biology. Tokai University Press, Tokyo, pp 513–521
Shaw PW, Pierce G, Boyle PR (1999) Subtle population structuring within a highly vagile marine invertebrate, the veined squid Loligo forbesi (Cephalopoda: Loliginidae) uncovered using microsatellite DNA markers. Mol Ecol 8:407–417
Shaw PW, Hendrickson L, McKeown NJ, Stonier T, Naud MJ, Sauer WHH (2010) Discrete spawning aggregations of loliginid squid do not represent genetically distinct populations. Mar Ecol Prog Ser 408:117–127
Stewart WJ, Bartol IK, Krueger PS (2010) Hydrodynamic fin function of brief squid. Lolliguncula Brevis J Exp Biol 213(12):2009–2024
Ulloa PM, Hernández CE, Rivera RJ, Ibáñez CM (2017) Biogeografía histórica de los calamares de la familia Loliginidae (Teuthoidea: Myopsida). Lat Am J Aquat Res 45:113–129
Villanueva R, Vidal EAG, Fernández-Álvarez FÁ, Nabhitabhata J (2016) Early mode of life and hatchling size in cephalopod molluscs: influence on the species distributional ranges. PLoS ONE 11:e0165334
Voss GL (1963) Cephalopods of the Philippine Islands. Bull US Natl Mus. https://doi.org/10.5479/si.03629236.234.1
Acknowledgements
We would like to thank Francisco Rocha for the comments on the initial stages of the manuscript.
Funding
CMI acknowledges funding grant REG UNAB 04-2020. JBLS thanks Instituto de Ciencias Biologicas from UFPA for the project support number 041/2020/ICB/UFPA.
Author information
Authors and Affiliations
Contributions
CMI and CM-G were involved in conceptualization, compiled data, methodology, formal analysis, writing—original draft, and review and editing. FIT, AL, and JBLS were involved in conceptualization and writing—review and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest and also no financial interests.
Ethical approval
Ethical review and approval were not required for this study because this work does not contain any experimental studies with live animals. Biological, distributional data, as well as sequences, were taken from open sources (i.e. GenBank, FAO books). This study did not need ethical approval since it was based on literature review and published information.
Additional information
Responsible Editor: R. Rosa.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ibáñez, C.M., Luna, A., Márquez-Gajardo, C. et al. Biological traits as determinants in the macroecological patterns of distribution in loliginid squids. Mar Biol 170, 133 (2023). https://doi.org/10.1007/s00227-023-04286-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00227-023-04286-1