Abstract
One of the distinguishing morphological features of conchiferan mollusks is the presence of a calcified exterior shell. Despite being conchiferans, most extant cephalopods have lost, internalized, or degraded their shells, with the exception of the basally divergent nautiloids. We compiled the shell matrix protein (SMP) data from several Nautilus pompilius studies and compared them with publicly available conchiferan SMP data, including those of Sepia pharaonis and Spirula spirula, two decapodiform cephalopods with internalized and partially internalized calcified shells, respectively. The recompilation of N. pompilius data revealed the presence of 85 distinct SMPs. Reciprocal homology searches suggested that N. pompilius shares 27 proteins known for their significant roles in shell formation and biomineralization, such as Pif/Pif-like, with other conchiferans. This is in agreement with our previous results, which suggested that the main functional domains of the SMPs in N. pompilius were also found in the conchiferans, suggesting that the domains were present in the ancestral conchiferans, with some recruited ancestrally, and some taxon-specifically. Meanwhile, 16 N. pompilius proteins are shared among the conchiferans, with only six proteins present in the cephalopods. indicating that some proteins were lost or co-opted in the course of their evolution. Three proteins shared among the conchiferans (CD109 antigen, Chitinase, and Tyrosinase) and one shared among the cephalopods (SOUL Domain-containing protein) apparently have dual functions of immunity and shell biomineralization. Intriguingly, four proteins are shared only between the two decapodiforms, indicating that they were co-opted in the decapodiform lineage with internalized shells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Presently, members of two major lophotrochozoan phyla, Mollusca (the mollusks) and Brachiopoda (the brachiopods) form extremely biomineralized (calcified) external tissue called the shell (Crenshaw 1989; Cowen 2013; Wernström et al. 2022). For the animals still retaining this structure, the shell is important, because it mainly functions by giving structural support and protection of soft body parts (Lowenstam and Weiner 1989; Simkiss and Wilbur 1989). The presence of the shell might have given the mollusks an edge to survive, contributing to their biodiversity (ca. 85,000–200,000 extant known species) and success in adaptation to various environmental conditions, from the deep seas to the shallow seas and freshwater, from terrestrial to subterranean environments (Lindberg 2001; Ponder et al. 2019; Prié 2019). As the group’s major synapomorphy (Kocot et al. 2017), shells had evolved early during the evolution of mollusks in Early Cambrian (ca. 525 million years ago) (Budd 2003; Jackson et al. 2010; Vinther 2015). Of the presently recognized seven classes of extant mollusks (Aplacophora, Polyplacophora, Gastropoda, Bivalvia, Scaphopoda, Monoplacophora, and Cephalopoda) (Kocot et al. 2020), six of them (i.e., besides Aplacophora) are shelled (Sigwart and Sutton 2007). Interestingly, however, the spicules on the skin of Aplacopora are thought to form through the precipitation of calcium carbonates on the skin/mantle tissue-secreted proteins (Woodland 1907; Beedham and Trueman 1968), suggesting a possibility that they are homologous to the shell (e.g., Scheltema 1993; Todt and Wanninger 2010; McDougall and Degnan 2018; Wanninger and Wollesen 2019), but probably extremely reduced during its evolution (Kocot et al. 2019). Meanwhile, some members of the shelled six classes have reduced or lost their shells, such as the teredinid shipworms (Bivalvia) and the nudibranch sea slugs (Gastropoda). The non-shell-forming mollusks, however, apparently retain their shell matrix protein-coding genes in their genome (Setiamarga et al. 2021; Yoshida et al. 2022).
Although exact details of the mechanisms of shell formation, biomineralization, and their evolution in mollusks are still unclear (Furuhashi et al. 2009; Pomar 2020), recent studies have provided some key insights. First, the molluscan shell is composed mainly of calcium carbonate crystals, such as aragonite and calcite, arranged in several superimposed layers in different arrangements with bioorganic molecules mixed in (Spann et al. 2010; Frenzel and Harper 2011; Marin et al. 2012). This is because the molluscan shell is formed through an organic matrix-mediated process (Weiner et al. 1983), where cells and tissues secrete biomolecules, such as proteins, forming organic layers that later act as a framework in which inorganic crystal nucleation and mineral salt precipitation occurs (Lowenstam 1981; Isowa et al. 2012). The involvement of proteins in biomineralization and shell formation, including morphogenesis at both the micro, macro, and molecular levels suggests that they are genetically controlled (Carter 1990). Although present only in trace amounts inside the shell, the proteins (called shell matrix proteins; SMPs) apparently play essential roles in shell formation and structural maintenance (Addadi et al. 2006; Marin et al. 2013). For example, Pif, a major SMP, was shown to be crucially involved in nacre formation (Suzuki et al. 2009), while the highly acidic protein Aspein (Isowa et al. 2012) promotes calcite precipitation (Takeuchi et al. 2008). Recent development in technology in large-scale analysis of biomolecules using multiomics approaches, such as genome sequencing, transcriptomics, and proteomics, have allowed for the identification of relatively comprehensive lists of SMPs in various species of mollusks (e.g., Marie et al. 2011, 2012, 2017; Mann and Edsinger 2014; Liu et al. 2015; Feng et al. 2017; Liao et al. 2015; Zhang et al. 2018; Zhao et al. 2018; Ishikawa et al. 2020).
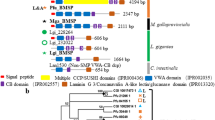
Except for the early branching Nautiloidea (the nautiloids), all members of extant Cephalopoda (the cephalopods; e.g., vampire squids, octopuses, squids, and cuttlefishes) had evolutionarily internalized, degenerated, or completely lost their shells (Kröger et al. 2011; Setiamarga et al. 2021; Setiamarga 2021) (Fig. 1A). Phylogenetically, the cephalopods are included in the subphylum Conchifera (the conchiferans), a monophyletic group composed of five of the seven classes of calcified external shell-forming mollusks (i.e., Gastropoda, Bivalvia, Scaphopoda, Monoplacophora, and Cephalopoda) (Kocot et al. 2011, 2020; Smith et al. 2011). Many extinct members of the cephalopods, such as the ammonites (Fig. 1B) and some extinct nautiloids (Fig. 1C), also had prominent biomineralized shells (Kröger et al. 2011; Ponder et al. 2019; Setiamarga 2021). Thus, to obtain a more complete general picture of the SMPs and their evolution in mollusks, information and insights from diverse cephalopods are important. Moreover, because of the various degrees of shell degeneration in extant cephalopods, a comparative study would allow for the elucidation of gene loss and recruitments during shell evolution, and the identification of key genes in molluscan shell formation.
The nautiloids (Nautiloidea sensu stricto) is an early branching clade of the cephalopods (Saunders 1981). There are eight presently recognized extant nautiloid species in a single family (Nautilidae): Nautilus pompilius, N. belauensis, N. macromphalus, N. stenomphalus, N. vitiensis, N. samoaensis, N. vanuatuensis, and Allonautilus scrobiculatus (Wray et al. 1995; Barord et al. 2023). Extant nautiloids are considered as living fossils, because they retain many ancestral traits of the cephalopods including the external, chambered shell (Fig. 1D) (Ward et al. 1984; Fortey 2011; Kröger et al. 2011; Setiamarga 2021). Previously, we conducted a multiomics study on the SMPs of one of the eight surviving species of the nautiloids, N. pompilius, by conducting transcriptomics of the mantle tissue and proteomics of the shell, to obtain their SMPs (Setiamarga et al. 2021). By comparing obtained N. pompilius SMPs with those of other conchiferans, we found that most protein domains were conserved and present in the putative ancestral conchiferans. At around the same time when our study was concluded, two draft genomes of N. pompilius were reported (Zhang et al. 2021; Huang et al. 2022), with the former study also including the identification of 78 SMPs. Since the two N. pompilius SMPs studies used different methods, there are slight differences in the number of overlapping SMPs identified (Fig. 2A). Therefore, the two SMP data sets must be compiled and curated, to obtain a more complete view of the proteins and their evolution, as well as their functions during shell formation. Comparisons with the SMPs of other conchiferans including other cephalopods will help to shed light on another piece of information regarding the evolution of biomineralized shells in mollusks at the molecular and genetic levels, and how such an important morphological character was lost in the evolution of the cephalopods.
Newly compiled data of N. pompilius shell matrix proteins (SMPs). A Comparison between the number of identified SMPs in Setiamarga et al. (2021) and Zhang et al. (2021). The newly compiled data re-identified a total of 85 proteins. B The schematic view of the domain structures in the re-identified 85 SMPs of N. pompilius. C Sequences of structural proteins with repeat motifs rich in Gly/Ala rich. Gly, Ala, and Asp are shown in red, blue, and green letters, respectively. Signal peptides are underlined
Materials and methods
Nautilus pompilius sequence data acquisition and compilation
The shell matrix proteins (SMPs) of N. pompilius were data-mined from two previously published data sets (Setiamarga et al. 2021; Zhang et al. 2021). We re-identified the full sequences of SMPs from both studies by mapping the sequences on the draft genome sequence reported by Huang et al. (2022) using BLASTp searches, and afterward compared the data sets. Afterward, domain structures of identified sequences were predicted using the online version of SMART (Letunic et al. 2021; http://smart.embl-heidelberg.de/smart/change_mode.pl; accessed in November 2022) and InterProscan (Jones et al. 2014; https://www.ebi.ac.uk/interpro/search/sequence/; accessed in November 2022). Signal peptide domain was predicted using SignalP 5.0 (Nielsen et al. 2019; Almagro Armenteros et al. 2019; https://services.healthtech.dtu.dk/service.php?SignalP-5.0; accessed in November 2022).
Comparative analysis of shell matrix proteins
Sequence data of shell matrix proteins (SMPs) of 13 mollusks were collected from previously published data, including those of two cephalopods, the ramhorn squid Spirula spirula (Oudot et al. 2020) and the pharaoh cuttlefish Sepia pharaonis (Liu et al. 2023). The complete list of the molluscan species compared in this study is shown in Table S1. Reciprocal local BLASTp was conducted to identify shared proteins among these species. A genomic search using local BLASTp on validated gene models was conducted to look for the availability of the homologs of molluscan SMP-coding genes in the genome of N. pompilius. Reciprocal local BLASTp searches were also conducted for the three cephalopod species compared in this study. All searches were conducted under the e value threshold score of < 1e-20. Homologous proteins among species were visualized using Circos v 0.69 (Krzywinski et al. 2009). It is to be noted that the sequence data of some of the proteins and/or genes identified by Oudot et al. (2020) are not published. For such sequences, we based our analyses only on the reported name of the sequences, by comparing them to the name assigned by the annotation to our sequences.
Phylogenetic analyses
Phylogenetic analyses were conducted on a select major SMPs for which the homologs are present in multiple mollusks compared in this study (SOUL Domain-containing protein, CD109 antigen, Chitinase, Tyrosinase, EGF-ZP Domain-containing protein). To do so, amino acid sequences were collected from Ensembl Metazoa (https://metazoa.ensembl.org/index.html; accessed in November 2022), MolluscDB (Liu et al. 2021; http://mgbase.qnlm.ac/home; accessed in November 2022), published SMPs data collected from various repositories (Table S2), Genbank (NCBI; https://www.ncbi.nlm.nih.gov/; accessed in November 2022), using HMM (HMMER v3.3.2; Eddy 2009) and BLASTp searches. Multiple sequence alignments were performed in MAFFT v7 server (Katoh and Standley 2013; https://mafft.cbrc.jp/alignment/server/; accessed in November 2022). Trimming to exclude poorly aligned regions was conducted in trimAl v1.3 (Capella-Gutiérrez et al. 2009), which was implemented in Phylemon 2 (Sánchez et al. 2011; http://phylemon.bioinfo.cipf.es; accessed in November 2022). Amino acid substitution model selections were conducted in MEGA X (Kumar et al. 2018) using Akaike information criterion (AICc) (Stecher et al. 2020). Maximum-likelihood (ML) phylogenetic tree was inferred using RAxML v.8.2.12 with 1,000 bootstrap replicates (Stamatakis 2014).
Whenever possible, full amino acid sequences were utilized in the phylogenetic inferences. However, when impossible, sequences of one or more representative domains of the proteins/genes were used. For example, due to the diversity of domain structures of Chitinases, a reliable alignment using full sequences was not possible. Therefore, for phylogenetic analysis, only the GH18 domains were used in both alignments and phylogenetic inferences.
Microstructure observations of Nautilus pompilius shell
The observation of shell microstructure of N. pompilius was conducted in the VE-8800 Scanning Electron Microscope (SEM) (Keyence, Osaka, Japan). Sample preparation was as follows. Samples were fragmented into small pieces (ca. 1 cm2) using a precision cutoff machine TS-45 (Maruto Testing Machine Co., Tokyo, Japan). Observation surface was polished using the polisher FG-18 (Ryobi Ltd., Hiroshima, Japan). Afterward, the observation surface was then treated with 1 M NaOH for 10 min and etched with Mutvei’s solution for 5 min (Schöne et al. 2005). After ultrasonic cleaning using US Cleaner USD-2R (AS ONE Corp., Osaka, Japan) for 5 min, the sample was left to air-dry overnight, sections were coated with osmium using Osmium coater HPC-1SW (Vacuum Device Inc., Ibaraki, Japan). The SEM image was visualized under the accelerating voltage of 10–12 kV.
Results
Nautilus pompilius sequence data acquisition and compilation
Previously, we identified 61 unique shell matrix protein (SMP) sequences in the shell of N. pompilius through multiomics analysis (mantle tissue transcriptomics vs. shell proteomics) (Setiamarga et al. 2021). However, because of the lack of sequencing depth, 14 out of 61 sequences showed frameshifts most likely caused by sequencing errors, while of the 47 in-frame sequences 41 were fragmented. Meanwhile, in their report about the full draft genome of N. pompilius, Zhang et al. (2021) identified 78 SMPs. When data of the two studies were compared, only 29 sequences overlapped (Fig. 2A).
In this study, we mapped the 61 unique sequences of Setiamarga et al. (2021) to the high-quality draft genome sequence of N. pompilius (Huang et al. 2022). The sequences were identified as 46 SMP-coding genes, and their full sequences obtained. The data set was then merged with the SMP data set of Zhang et al. (2021). As a result, we obtained a final set consisting of the full sequences of 85 SMPs (Fig. 2A, B). Among them, more than half (45 SMPs) had a signal peptide domain. The number of SMPs without any functional domain was nine. The amino acid sequence of some (MSTRG.749, MSTRG.5336, MSTRG.8937, and MSTRG.11770) are found to be enriched in Gly/Ala (Fig. 2C). The annotation result of the 85 sequences of the N. pompilius SMPs of this study is shown in Table S3.
Comparative analysis of conchiferan and cephalopod shell matrix proteins
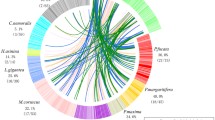
In the present study, we compared the more exhaustive data of N. pompilius SMPs we compiled in this study (Fig. 2A, B; Table S3) with the SMP data sets of several conchiferans to look into the evolution of molluscan biomineralization toolkit proteins (Fig. 3A; Tables S1, S2). The result indicated that the degree of homology between N. pompilius and other conchiferan SMPs was limited to around 10% or less (Fig. 3B). These 10% of common SMPs include 11 different types of proteins, i.e., Pif/Pif-like, Peroxidase, BPTI/Kunitz family of Serine Protease Inhibitors, Tyrosinase, Chitinase, Collagen-like, L-Amino-acid Oxidase, CD109 antigen, EGF-ZP Domain-containing protein (EGF-ZP), Carbonic Anhydrase, Chitin Binding Domain-containing protein (ChBD) (Fig. 3C; Fig. S1). This result confirmed our previous result (Setiamarga et al. 2021), but added more shared proteins between N. pompilius and the other conchiferans. Some of these common SMPs are conserved as multiple unigenes on the N. pompilius genome (e.g., unigenes with differing domain structures but identified as similar proteins based on the similarity of their main functional domain(s), such as Pif/Pif-like protein contains two types of unigenes (Pif and Pif-like); Chitinase and Tyrosinase contain two types of unigenes each; ChBD-containing protein contains three types of unigenes). If they are counted as individual genes/proteins, the number of genes shared among the conchiferans including N. pompilius becomes 27.
Common shell matrix proteins (SMPs) conserved among the conchiferans. A The list of published SMPs data sets mapped on the cladogram of the organisms. A corresponding list is also presented in Table S1. B A Circos diagram depicting sequence homologies between the SMPs of N. pompilius SMPs and other conchiferans (E value score < 1.0e−20). C Common SMPs among N. pompilius and the conchiferans. D Common SMPs among the cephalopods. Protein names written in red indicate SMPs matched in both venn diagrams. Because sequence data set of S. spirula (Oudot et al. 2020) is incomplete, some sequences were identified only based on their annotated names (showed in parentheses)
Two SMPs (Chitinase and Tyrosinase) were shared between two cephalopods, N. pompilius and S. pharaonis, and the conchiferans. These proteins are well-known as major SMPs in the conchiferans (Fig. 3B–D; Fig. S1; e.g., Arivalagan et al. 2016; Feng et al. 2017; Shimizu et al. 2022). Meanwhile, N. pompilius and S. spirula share Actin and Peptidyl-prolyl Cis–trans Isomerase B. These proteins were also detected in the SMPs of the gastropods (Marie et al. 2013; Mann et al. 2018; Shimizu et al. 2019). Two types of proteins are shared among the three species of the cephalopods presently studied (N. pompilius, S. pharaonis, and S. spirula) and the other conchiferans: CD109 antigen and VWA Domain-containing proteins (Pif/Pif-like/Collagen-like) (Figs. 3D; Fig. 4; Figs. S1, S2).
The Pif, Pif-like, and Collagen-like proteins contain von Willebrand Factor Type A (VWA) domain(s), but with different arrangements and numbers (Fig. 4). However, while the VWA Domain-containing protein of N. pompilius could be recognized as Pif/Pif-like, based on the structure of the protein (such as the presence, numbers, and arrangements of the VWA and ChBD domains) because of its complete sequences, the proteins of S. pharaonis and S. spirula were annotated as Collagen-like on NCBI, because both sequences only have the VWA domain(s) as their identified functional domain(s) (two for S. pharaonis and one for S. spirula. We classified Pif/Pif-like proteins and Collagen-like proteins as VWA Domain-containing proteins (Pif/Pif-like/Collagen-like), because we cannot rule out the possibility that the registered sequences of S. pharaonis and S. spirula were incomplete and/or fragmented.
While all VWA Domain-containing proteins and CD109 antigen are also found in the conchiferans, the SOUL Domain-containing protein (SOUL) was discovered in all cephalopods, but not in the other conchiferans (Fig. 3; Fig. 5; Fig. S2). Meanwhile, the number of SMPs shared only between the decapodiforms S. pharaonis and S. spirula are four: Hexosaminidase, Neurofilament, Calmodulin, and Arginine Kinase.
Molecular phylogeny of SOUL Domain-containing proteins (SOUL). A Domain structure of the SOUL Domain-containing protein (SOUL) in the SMPs. B Number of SOUL proteins with SOUL Domain (PF04832), including both SMP and non-SMP types, conserved in published metazoan genomes and SMPs. Length of bars indicates the number of proteins in each genome. C Maximum likelihood tree of the amino acid sequences, inferred using the WAG + Γ + I model with 1000 bootstrap replicates. Bootstraps values of major clades are shown (‘-’ means less than 30%). Abbreviations: Nam: Necator americanus, Cel: Caenorhabditis elegans, Hro: Helobdella robusta, Cte: Capitella teleta, Lan: Lingula anatina, Npo: Nautilus pompilius, Sph: Sepia pharaonis, Ssp: Spirula spirula, Aar: Architeuthis dux, Aar: Argonauta argo, Omi: Octopus minor, Obi: Octopus bimaculoides, Pca: Pomacea canaliculata, Bgl: Biomphalaria glabrata, Aca: Aplysia californica, Lgi: Lottia gigantea, Gae: Gigantopelta aegis, Hla: Haliotis laevigata, Pma: Pinctada fucata martensii, Pye: Patinopecten yessoensis, Mph: Modiolus philippinarum, Bpl: Bathymodiolus platifrons, Cgi: Crassostrea gigas, Pfu: Pinctada fucata
Molecular phylogenetics of some N. pompilius SMPs
In this study, we selected five major molluscan SMPs (SOUL Domain-containing protein, CD109 antigen, Chitinase, Tyrosinase, and EGF-ZP), and conducted molecular phylogenetic analyses to understand their evolutionary history.
SOUL domain-containing protein
We found a copy of SOUL Domain-containing (SOUL) protein in the shell matrices of the three cephalopods compared in this study, namely, N. pompilius, S. pharaonis, and S. spirula (Fig. 5A), but not in other mollusks (Fig. 5B). Phylogenetic and copy number analyses suggest the presence of multiple copies of SOUL-coding genes in the genomes of the animals surveyed in this study, including in the brachiopod Lingula anatina (Fig. 5B, C).The SMP-type SOUL found in the cephalopods form a monophyletic clade together with some of the non-SMP type, with high support (BS = 100%). The sequences were also found in the genome of N. pompilius (Fig. 5B).
CD109 antigen
Although various studies have identified a Thioester-containing protein (TEP) the CD109 antigen (Fig. 6A), as a major SMP, the actual type of this TEP has not been addressed and thus unclear. Annotations have been based mainly only on sequence similarities of the domains. In this study, we performed phylogenetic analysis to classify 205 TEPs of conchiferan SMPs (Fig. 6B). The resulting tree supports the monophyly of all TEP types reported previously (e.g., Sekiguchi et al. 2012; Duval et al. 2020; Marquez et al. 2022), the topology recovered the two subfamilies, C3/C4/C5 and A2M, with full statistical supports (bootstrap values (BS) = 100%). The A2M subfamilies further diverged into six monophyletic clades: the CPAMD8 (BS = 100%), A2M (BS = 100%), MCR (BS = 100%), iTEP (BS = 100%), CD109 (BS = 50%), and molluscan TEP (mTEP) (BS = 56%).
Molecular phylogeny of Thioester-containing proteins (TEPs), including CD109 antigen. A Domain structures of Thioester-containing protein (TEP) family. B Maximum likelihood tree of the amino acid sequences, inferred using the LG + Γ + I model with 1000 bootstrap replications. Bootstraps values of major clade are shown. Background colors of the topology corresponds to the protein family name, all monophyletic in this tree. Outgroup sequences were collected from Sekiguchi et al. (2012), Duval et al. (2020), and Marquez et al. (2022). The domain structures of SMP-type TEPs are shown in Fig. S3
Non-SMP molluscan (bivalves, gastropods, and cephalopods) TEPs formed five monophyletic clades (the C3/C4/C5, A2M, CD109, mTEP, and MCR clades). Besides them, arthropods specific CD109s form the monophyletic iTEP clade (Sekiguchi et al. 2012), with the monophyletic CD109 clade containing several animal TEPs (e.g., mouse, humans, and the bobtail squid) forming its sister clade. Sister to the two clades was the monophyletic mTEP clade (BS = 56%), which was composed exclusively of the molluscan TEPs. The SMPs of all non-cephalopod conchiferans studied here are included in this clade. We found that only the N. pompilius SMP TEP (which were annotated as CD109 antigen in Setiamarga et al. 2021) and the non-SMP ones were also included in the mTEP clade. The TEPs of the other cephalopods studied here, S. pharaonis and S. spirula, were included in the A2M clade.
Chitinase
Molluscan Chitinases are known to have complex and highly diverse domain structures, with one or multiple GH18 domains (PF00704) present in all Chitinases, while in some, one or several Chitin-binding domains (ChBD) are also present besides the GH18 (Table S4). Such domain structure patterns are also seen in those reported as SMPs (Fig. 7A). We found both types of Chitinases (MSTRG20089 and MSTRG20090) in the SMPs of N. pompilius, with MSTRG20090 having two GH18 domains, a signal peptide, and a ChBD domain, while the former had only a single GH18 domain (Fig. 7A).
Molecular phylogeny of Chitinases. A Domain structure of Chitinase of molluscan SMPs. B Maximum likelihood tree of the amino acid sequences, inferred using the LG + Γ + I model with 1000 bootstrap replicates. Bootstraps values of major clades are shown (‘-’ means less than 30%). Outgroup sequences were obtained from Nematoda, Annelida, and Brachiopoda (Table S2). Clades with high bootstrap support (Chitinase without ChBDs groups I–VI) are highlighted gray
The phylogenetic tree inferred using the amino acid sequences of the GH18 domains of Chitinases collected from 536 lophotrochozoans suggested that the enzyme did not form any monophyletic bifurcation corresponding the two types of domain structures mentioned above (Fig. 7B). Six monophyletic clades of some non-SMP Chitinases with only GH18 from all molluscan lineages (bivalve, gastropod, cephalopod; denoted as I–VI in Fig. 7B) were found (BS = 49–100%), but none of the Chitinases reported as SMPs are included in the six clades. Our phylogenetic analysis also recovered a clade containing some of the pteriomorph bivalvian SMPs together with other seemingly non-SMP sequences (BS = 100%), as previously reported (Shimizu et al. 2020). Other molluscan SMP Chitinases (including those of the bivalves) are not monophyletic, but apparently formed taxon-specific clades (bivalve, gastropod, and cephalopod clades) with the non-SMP ones (albeit statistically very weakly-supported). This is also the case for the Chitinases of N. pompilius (MSTRG20089, containing only the GH18 domain; MSTRG20090, containing both GH18 and ChBD domains), which were found to be included in a clade together with other cephalopod Chitinases.
Tyrosinase
Tyrosinase, an oxidoreductase metalloenzyme containing a single Tyrosinase domain (PF00264), was found in all of the 22 Lophotrochozoan genomes included in this study (Fig. 8A). In mollusks, the number of Tyrosinase gene copies present in the genomes showed considerable variability, with more number observed in bivalves than in other mollusks (Bivalvia = 24–53; Gastropoda = 1–12; Cephalopoda = 4–15). Despite the limited number of species analyzed for SMPs (Small Secreted Proteins), the number of Tyrosinases identified as SMPs was also variable and probably included isoforms. Although not found in all of the 21 Tyrosinases identified as conchiferan SMPs, a signal peptide was apparently conserved in addition to the single Tyrosinase domain (Fig. 8B). Two copies of the Tyrosinase gene were discovered in all cephalopods studied here (Fig. 8B).
Molecular phylogeny of Tyrosinases. A Number of proteins with a single Tyrosinase domain (PF00264) conserved in the genomes of animals studied here, plotted on a phylogenetic tree of the organisms compared. Length of bars indicates the number of proteins in each genome. B Domain structures of the SMP-type Tyrosinases in mollusks. (C) Maximum likelihood tree of the amino acid sequences, inferred using the LG + Γ + I model with 1000 bootstrap replicates. Bootstraps values of major clades are shown (‘-’ means less than 30%). Representative clades with gene number expansions in the genome are highlighted by the gray color. Outgroup sequences were obtained from Nematoda, Annelida, and Brachiopoda (Table S2). Nam: Necator americanus, Cel: Caenorhabditis elegans, Nam: Necator americanus, Nam: Necator americanus, Nam: Necator americanus, Nam: Necator americanus, Hro: Helobdella robusta, Cte: Capitella teleta, Lan: Lingula anatina, Npo: Nautilus pompilius, Aar: Architeuthis dux, Aar: Argonauta argo, Omi: Octopus minor, Obi: Octopus bimaculoides, Pca: Pomacea canaliculata, Bgl: Biomphalaria glabrata, Aca: Aplysia californica, Lgi: Lottia gigantea, Gae: Gigantopelta aegis, Hla: Haliotis laevigata, Pma: Pinctada fucata martensii, Pye: Patinopecten yessoensis, Mph: Modiolus philippinarum, Bpl: Bathymodiolus platifrons, Cgi: Crassostrea gigas, Pfu: Pinctada fucata. Mytilidae consist of B. platifrons and M. philippinarum. Pteriidae consist of P. fucata, P. fucata martensii
Our phylogenetic tree did not reveal a monophyletic SMP Tyrosinase, which is consistent with our previous report (Setiamarga et al. 2021) (Fig. 8C). Only a certain degree of phylogenetic clustering of the 344 Tyrosinase protein sequences analyzed in this study was observed in the tree, with almost no taxon-specificity, and in most cases, with relatively low statistical support for the nodes. For example, some of the cephalopod Tyrosinases formed two monophyletic clades, each containing both SMP and non-SMP copies of the gene (BS = 100% and 68%). Meanwhile, the heavily expanded copies of bivalve Tyrosinases formed multiple clades, with some containing the SMP and non-SMP copies of the gene.
EGF-ZP domain-containing protein
A copy of the EGF-ZP Domain-containing protein (EGF-ZP) (Fig. 9A) was identified in N. pompilius SMPs (Setiamarga et al. 2021). However, no EGF-ZP and other proteins thought to form complexes with EGF-ZP in biomineralization (e.g., Shematrin, Pif, and Serine Protease Inhibitor) were detected in the SMPs of the decapodiforms S. pharaonis and S. spirula SMPs. Similar to Tyrosinase, our phylogenetic tree, which includes only the 63 protein sequences annotated as EGF-ZP from mollusks (with one sequence of the annelids as outgroups), also indicated that the EGF-ZP proteins employed as SMPs do not form a monophyletic clade, with a possible taxon-specific clusterings (Fig. 9B). For example, the SMP and non-SMP EGF-ZP proteins in the cephalopods formed a monophyletic clade (BS = 100%), while those of the bivalves formed two monophyletic clades (BS = 100% and 95%).
Phylogenetic tree of EGF-ZP Domain-containing proteins (EGF-ZP). A Domain structures of EGF-ZP Domain-containing proteins (EGF-ZP) in mollusks. B Maximum likelihood tree of EGF-ZP amino acid sequences, inferred using the LG + Γ + I model with 1000 bootstrap replicates. Bootstraps values of major clades are shown (‘-’ means less than 30%). An annelid sequence was used as the outgroup
Discussion
Features and evolution of the shell matrix proteins in mollusks
Since Pieter Harting, one of the pioneers in the study of biomineralization, identified the organic matrix as an essential calcareous component, numerous researchers have been examining its roles in biomineralized structures (Harting 1872; Sommerdijk and Cölfen 2010; Dauphin 2023). The nautiloids have fascinated researchers for being the only extant taxon of the cephalopods still retaining its calcified external shell with distinct structural traits like the presence of a nacreous layer, a characteristic shared with the shells of other mollusks. Because the nautiloids are early-branching cephalopods, examining their shells would allow scientists to reconstruct presumed ancestral features in the cephalopods and study their evolution. Macrostructurally, the equiangular spiral shell of the nautiloids has several distinct parts: the wall, septum, callus, septal neck, and siphuncle. Meanwhile, the main microstructures of the shell are composed of the mineralized layers of granular, prismatic, and nacreous aragonite (Lowenstam et al. 1984; Saunders and Landman 2009; Velázquez-Castillo et al. 2006; Fig. 1D) and organic layers composed of Beta Chitin and other SMPs (Weiner and Traub 1980).
Recent technological and analytical method developments in genomics, transcriptomics, and proteomics have allowed for relatively exhaustive identifications of various matrix proteins embedded in the organic layers of the shell, called shell matrix proteins, in mollusks (Fig. 3; Fig. S1; Table S1). One of the main aim of analyzing SMPs is to identify the genetic tool kits and the key players in molluscan biomineralization, which will allow for the elucidation of the actual mechanism of biomineralization, shell formation, and their evolution at the molecular level, besides opening up possibilities of genetic manipulation of the identified and characterized proteins for applicable purposes (such as pearl production). These studies had thus far revealed that the number of shared SMPs among different conchiferans are relatively low, especially at higher taxonomy, such as between Bivalvia and Gastropoda (Marie et al. 2012; Mann and Jackson 2014; Feng et al. 2017; Song et al. 2019). Those studies also suggested that even the degree of homologies among shared SMPs is arguably low even at the inter-species level in the two molluscan classes (Bivalvia and Gastropoda). This high level of diversity shown by the SMPs was thought to be caused by rapid evolution of the genes, probably because of the differing selective pressures to adapt to the different details of niches in the living environment of each species. Besides that, being close to the actual expressed phenotypes and thus most likely downstream of the gene expression networks, the selective pressures on the SMPs are relatively low, that is, as long as they maintain the necessary spatial structures needed to get the job done, they do not have to strictly retain their primary structures. This would also allow organisms to co-opt non-homologous proteins as SMPs, as long as they have the necessary spatial and functional configurations and structures (e.g., Nakashima et al. 2019), which could probably be caused by structural and functional convergence (Pearson and Sierk 2005). Intriguingly, such differences in recruited SMPs apparently correspond to crystallographic differences, such as the microstructures (e.g., Jackson et al. 2010; Kocot et al. 2016). Deep homology, that is, when core sets of homologous proteins get recruited to form morphologically homologous structures in different species (Zaquin et al. 2021), might also explain why some representative proteins thought to be crucial for biomineralization, such as Nacrein, Pif, and EGF-ZP, have been identified in various bivalves and gastropods (Kocot et al. 2016). Sometimes, key proteins are so important that they get re-recruited to form similar but morphologically non-homologous structures, such as the shell-like calcified eggcase of the argonauts (e.g., Yoshida et al. 2022).
The shell matrix proteins of N. pompilius and the cephalopods provide insights to shell evolution in mollusks at the molecular level
To understand the evolution and diverse mechanisms of biomineralization in mollusks, information from the cephalopods is crucial, because the lineage has evolved the structure, but has extant members with the biomineralized shells internalized (e.g., the cuttlefish), degenerated (e.g., the squids), lost (e.g., the octopuses), and even reinvented (e.g., the argonauts). Therefore, studying the SMPs in the nautiloids is a crucial first step in elucidating the evolution of mollusk shells.
Previously, we showed that the presence of eight out 47 SMPs of N. pompilius SMPs are shared among the conchiferans (Pif/BMSP-like, CD109 antigen, Tyrosinase, EGF-like domain-containing protein, Chitinase, Peroxidase, BPTI/Kunitz domain-containing protein, Uncharacterized LOTGI_169029; Setiamarga et al. 2021). From the updated N. pompilius SMPs in the present study, we found an additional four (Carbonic Anhydrase (CA), L-amino-acid Oxidase, Collagen-like, and Peroxidase), making the total number of homologous SMPs in the conchiferans to be 11 out of 85. We did not find one protein identified in our previous study, Uncharacterized LOTGI_169029, probably because the protein was an artifact caused by sequencing and/or assembly errors and was undetected anymore when mollusk genome sequences were updated. The presence of this gene repertoire, which was probably acquired in the common ancestor of the cephalopods, gastropods, and bivalves, confirmed the presence of a core set of shell matrix proteins conserved among the three conchiferan classes. Five polypeptide fragments were identified SMPs in N. macromphalus: Mucoperlin-like (which exhibited some characteristics of Mucin; Marin et al. 2000), MSI31/MSI60-like (which is enriched in Gly/Ala, Sudo et al. 1997), N14/N16/Pearlin-like (which contain Heparin-binding motif, acidic regions, and Gly/Asn-rich region, Samata et al. 1999; Montagnani et al. 2011), Nacrein-like (also known as Carbonic Anhydrase, Miyamoto et al. 2003), and Tyrosinase-like. Comparison between the two Nautilus species showed that N. pompilius and N. macromphalus share four out of the five proteins (except for N14/N16/Pearlin-like). This indicates that these major SMPs are probably involved in the nacreous shell formation in the two Nautilus species, or even possibly, in all nautiloids. Meanwhile, at present, we cannot decisively determine if the lack of N14/N16/Pearlin-like in N. pompilius was because the protein is a species-specific protein to N. macromphalus or an artifact.
We also conducted a comparison of the SMPs of three cephalopods with biomineralized shells, namely, N. pompilius, S. pharaonis, and S. spirula. While the shells of S. pharaonis and S. spirula are internalized and degenerated, they are considered to be true (morphological) homologs of the shell of the conchiferans, and retain some of the morphological structures seen in both the nautiloid and ammonite shells, such as the phragmocone (Kröger et al. 2011). Our findings suggest that some core matrix proteins (CD109 antigen, Tyrosinase, and Chitinase) are conserved among the cephalopods, although Tyrosinase and Chitinase were not found in the S. spirula SMPs. It should be noted that the three cephalopods studied here have different shell microstructures (Dauphin et al. 2020; Checa et al. 2022). As previously mentioned, this could suggest that the differences in the utilized SMPs among the three species are reflected in the differing shell microstructures (Jackson et al. 2010; Kocot et al. 2016). In addition, we also found that four proteins (Hexosaminidase, Neurofilament, Calmodulin, and Arginine Kinase) were shared only by the decapodiforms S. pharaonis and S. spirula. No studies on conchiferan SMPs have reported their presence thus far, suggesting that these proteins were probably recruited by the ancestral lineage leading to decapodiforms during or after the internalization of the calcified shell. Further research is still necessary to verify this hypothesis.
Nevertheless, it must be noted that not only were S. spirula sequences probably fragmented (not full sequences), but also one previously published data set used in this study (Oudot et al. 2020) was also incomplete, because not all sequences were published. This prompted us to analyze some proteins by reported annotated names rather than by sequence-based analytical methods, which may have caused a biased result. Therefore, a re-examination involving a more complete data set of S. spirula SMP sequences must be conducted in the future.
Cephalopod shell matrix proteins with repeat motifs
Studies on the amino acid compositions of N. pompilius and its congener, N. belauensis, have shown that the organic matrix is extremely abundant in the residues of Gly/Ala/Asx (Asp or Asn), and Ser (Weiner and Hood 1975; Weiner and Traub 1980; Lowenstam et al. 1984; Keith et al. 1993). This property was consistent with other conchiferan shells despite the different microstructures, and it has been suggested that these amino acid sequences probably play roles in shell formation. (e.g., Mercenaria mercenaria Hare and Abelson 1965; Crassostrea virginica Meenakshi et al. 1975; Mytilus californianus Keith et al. 1993). Weiner and Hood (1975) assumed that these proteins most likely consisted of a series of (Asp-Y)n (Y: Serine or Glycine) forming an antiparallel-sheet conformation, and suggested that the negatively charged Asp has a Ca2+ binding capacity (Weiner 1979). Subsequently, sequences with properties similar to what was predicted were identified from several bivalve species (e.g., Patinopecten yessoensis (MSP-1; Sarashina and Endo 1998), Pinctada fucata (Aspein; Tsukamoto et al. 2004), Atrina rigida (Asprich; Gotliv et al. 2005), Pinna nobilis (Caspartin; Marin et al. 2005), Crassostrea nippona (MPP1; Samata et al. 2008). Marie et al. (2009) also reported a possible presence of similar Gly/Ala/Asx (Asp or Asn) rich proteins in the nacreous layer of the congener of N. pompilius, N. macromphalus. The identification of MSI31/MSI60-like (Sudo et al. 1997) and N14/N16/Pearlin-like (Samata et al. 1999; Montagnani et al. 2011) in the same species explains the amino acid compositional bias.
Nautilin-63, which was characterized as an acidic phosphorylated protein of 63 kDa band extracted from the matrix protein of the nacreous layer in N. macromphalus (Marie et al. 2011), is also enriched in Gly/Asx/Thr residues and predicted to have a similar Ca2+ binding capacity (Marie et al. 2011). However, previous multiomics/phenomics studies on N. pompilius SMPs failed to find any protein with similar characteristics (Setiamarga et al. 2021; Zhang et al. 2021). This was probably because either Nautilin-63 is a species-specific protein, or because it is difficult to obtain a complete gene and/or protein sequence with highly repetitive motifs/domains. In this study, we succeeded in identifying possible full sequences with similar features, when we mapped some fragmented sequences on the genome. MSTRG.8937 was identified from the SMPs of N. pompilius, and is composed of repeat motifs rich in Gly/Ala/Asp residues. Previous studies have shown that the sequence similarities among possible orthologs of highly acidic proteins with highly repetitive acidic amino acid residues are very low, even among sequences from congeners (Isowa et al. 2012; McDougall et al. 2013). Nautilin-63 and MSTRG.8937 may be homologous, although this homology cannot be confirmed because of low similarities between the two sequences.
Three Gly/Ala rich proteins (MSTRG.749, MSTRG.5336, and MSTRG.11770) were identified from the SMPs and genome of N. pompilius. However, the three proteins do not contain acidic amino acids. Although reciprocal BLAST searches failed to find any homolog to these proteins, similar structural proteins have been previously identified and characterized as Silk Fibroin-like protein in the nacreous layer of Pinctada fucata (Sudo et al. 1997). It is to be noted that the shell protein was considered as Silk Fibroin-like protein based only on its Ala-rich content and function as a structural protein. These unique repeat sequences are thought to structurally strengthen the shell in mollusks (Yano et al. 2006), because its primary structure resembles the spider silk protein (which is thought to cause the strength of the spider web), and thus most likely functions similarly (Gatesy et al. 2001; Numata 2020). Another similar structural protein in plants, the Glycine-rich protein (GRP), is also thought to have been the cause of mechanical strength of plant cell walls (Lei and Wu 1991). Interestingly, MSTRG.5336 is the only N. pompilius SMP consisting of not only Gly/Ala, but also Gln/Gly/Arg in its repeat motif.
In this study, we succeeded in confirming the presence of several structural proteins containing orderly repeat motifs in N. pompilius, suggesting the presence of possibly few types of structural proteins in the shell matrix, probably involved in shell biomineralization. Our observation here also agrees with previous suggestions on the non-conservation of proteins with repeat motifs. Previous studies have shown that the repeat-motif proteins in mollusk SMPs are not sequence conserved (Weiner and Hood 1975), but the abundance of the same residue(s) is probably critical for them to function as structural proteins (Sarashina and Endo 1998; Isowa et al. 2012).
Remarks on the evolution of some N. pompilius SMPs
Pif/Pif-like
Pif was discovered from the SMPs of P. fucata and its structure consists of one VWA domain, two Chitin-binding domains, and a LamininG-like motif at the C-terminus (Suzuki et al. 2009, 2013; Fig. 4). Later, two types of the orthologous proteins with different domain arrangements, BMSP and LamininG3 (sometimes grouped together as Pif-like), were also reported. The former is apparently specific to bivalve SMPs, and the latter was found in gastropods and cephalopods (Suzuki et al. 2011; Ishikawa et al. 2020; Setiamarga et al. 2021; Yoshida et al. 2022). Pif and Pif-likes apparently play a role in forming a thin organic film layer of the nacreous layers and in controlling the growth direction of aragonites in the shell of Pinctada fucata (Suzuki et al. 2009). Other than the three types of Pif orthologs, recent studies have revealed the presence of multiple SMPs containing one or several VWA and ChBD domains, and a low complexity region (which is a signature of Pif/Pif-like) in various configurations, but with low sequence similarities to Pif (Zhao et al. 2020; Ishikawa et al. 2020). The low complexity region is located, and probably derived from, the LamG3 domain which remains intact in the LamininG3 protein (Suzuki et al. 2013; Setiamarga et al. 2021). Thus, in this study, an unidentified protein containing ChBD domain(s) and a low complexity region has been broadly classified as a “Pif-like” protein.
We found five Pif homologs in the genome and SMPs of N. pompilius: one Pif, one LamininG3, and three Pif-like proteins (Fig. 4). Pif possibly plays similar functions in biomineralization as its paralogs in P. fucata, including but not limited to, the formation of the nacreous layer. Meanwhile, LamininG3 is usually conserved in the cephalopod genomes, despite being found only in the genomes of a limited number of conchiferan species (e.g., Albertin et al. 2015; Kim et al. 2018; Da Fonseca et al. 2020; Huang et al. 2022; Yoshida et al. 2022). While both Pif and LamininG3 show their standard domain configurations (Fig. 4), Pif-like has a distinct configuration with two VWA, a single ChBD, and a low complexity region. This is similar to what was previously found in P. fucata (Zhao et al. 2020).
SOUL domain-containing protein
SOUL Domain-containing protein (SOUL) belongs to the Heme-binding Protein (HBP)/SOUL family of proteins with the SOUL Domain (PF04832) as the main functional domain (Fig. 5A). SOUL represents a group of evolutionarily conserved proteins with members in animals, plants, and even bacteria (Goodfellow et al. 2021). Their recognized representative function is to bind heme to its prosthetic group (Taketani et al. 1998). In mammals, two paralogs, HEBP1 and HEBP2, have been functionally analyzed. HEBP1 binds heme to iron, while HEBP2 does not bind to iron but has other multiple functions related to heme, such as binding to free heme and heme transport (Fortunato et al. 2016). Heme consists of a small cyclic tetrapyrrole with a centrally chelated Fe atom (Layer et al. 2010).
The phylogenetic analysis and copy number survey suggest that the presence of multiple copies of the SOUL-coding genes in metazoan genomes may have been caused by gene duplications and losses, followed by specialization of paralog functions, along with further copy number expansions in the molluscan lineage (Fortunato et al. 2016) (Fig. 5B, C). SOUL has not previously been reported as an SMP in mollusks. Nevertheless, this study identified SOUL proteins in the shell matrices of the three cephalopods examined, forming a monophyletic clade with high support (Fig. 5C). This finding suggests that they were likely recruited and co-opted in the lineage leading to cephalopods for biomineralized shell formation. A high percentage of sequence similarity at the C-terminus of the SMP-type cephalopod sequences may also indicate selection pressure to maintain protein structure.
Oudot et al. (2020) suggested that SOUL may be involved in biomineralization by storing and transporting iron. Ferritin, which is conserved in the SMPs of bivalves and gastropods, also has the ability to bind iron. It is possible that cephalopods have co-opted SOUL to perform a similar function instead of Ferritin. However, the relationship between iron and CaCO3-based biomineralization remains unclear, and thus the potential functional similarity between SOUL and Ferritin requires further investigation. In addition, SOUL may participate in binding calcium ions or other biomineralization-related prosthetic groups or in calcium ion-dependent interactions between proteins (Mikasa et al. 2018).
CD109 antigen
CD109 antigen belongs to the Thioester-containing proteins (TEPs) superfamily (Nonaka and Kimura 2006). TEPs are classified into two subfamilies: Complement Factors (C3/C4/C5) and Alpha-2 Macroglobulins (A2Ms). The A2M subfamily comprises several closely related molecules: Alpha-2 Macroglobulins (A2M), CD109 antigen (CD109), Macroglobulin Complement-related protein (MCR), Pregnancy Zone protein (PZP), C3 And PZP Like Alpha-2-Macroglobulin Domain-containing protein 8 (CPAMD8), and insect TEP (iTEP). These proteins share a set of domains as their basic signature: the signal peptide, A2M-N, A2M-N2, A2M, Alpha-Macroglobulin Thiol-ester Bond-forming Region, A2M Complement, and A2M Receptor domains. Several members of the A2M superfamily can be further distinguished by their additional domains added to the basic common domains. For example, C3/C4/C5 has the NTR (C345C) domain at the C-terminus, CPAMD8 has the KAZAL domain at the C-terminus, and MCR has the LDLa domain between its A2M-N2 and A2M domains (Fig. 6A). Some functional analyses have suggested that the protein must undergo proteolytic cleavage, causing most proteins identified through proteomics to be incomplete (Portet et al. 2018).
We found the non-SMP molluscan (bivalves, gastropods, and cephalopods) TEPs classified into five monophyletic clades (C3/C4/C5, A2M, CD109, mTEP, and MCR) in the genomes of the respective mollusks. These TEPs were probably conserved in the most recent common ancestor of mollusks. The iTEP clade, which consists solely of CD109 specific to arthropods (Sekiguchi et al. 2012), probably expanded from a CD109 included in the animal TEP clade (the CD109 clade), in a lineage that eventually gave rise to arthropods. It is also worth noting that only the SMP and non-SMP TEPs of N. pompilius were included in the mTEP clade, whereas those of S. pharaonis and S. spirula belonged to the A2M clade. This suggests that the expansion of mTEP probably occurred in the molluscan lineage after the split between mTEP and the CD109+iTEP clades, which might have happened at the same time as the Ecdysozoa-Lophotrochozoa split, an event that could have contributed to the development of the ability to form a calcified shell. Furthermore, the switch to A2M in the two decapodiforms possibly coincided with the internalization of their mineralized shells.
Chitinase
Chitinase is a representative conchiferan SMP, found in all studies on conchiferan shell biomineralization. The enzyme belongs to the Glycoside Hydrolase family 18 (GH18), which members are characterized by the presence of the GH18 domain (PF00704). In shell formation, it is thought that the enzyme works by catalyzing the degradation of the Chitin framework during CaCO3 biomineralization. SMP Chitinases show various domain structures (Fig. 7A), but can be artificially grouped into two types: Those consisting of only the GH18 Domain, and those with the Chitin-binding (ChBD) and GH18 Domains. The ChBDs could be located either before (on the N terminus side) or after (on C terminus side) the GH18, but apparently never both, and the number is not fixed (one or two ChBD domains are usually observed). Some mollusk Chitinases have complex domain structures, where several ChBD and GH18 domains are present (Table S4), indicating that both ChBD and GH18 domains seemed to undergo frequent tandem duplication and a possible concerted evolution among them.
The six monophyletic clades (I-VI; Fig. 7B) of Chitinases that consist only of the GH18 domain are noteworthy, because they include non-SMP Chitinases from all molluscan lineages (bivalve, gastropod, cephalopod), indicating possible functional constraints at the primary sequence level. However, Chitinases with the ChBD, including those found in SMPs, were not monophyletic. These proteins seem to have diversified in each taxon by acquiring or losing domains through domain shuffling and tandem duplication. For instance, our phylogenetic tree indicates that bivalves had probably doubled their number of SMPs (8–15) in at least the common ancestor of Pteriomorphia.
We also acknowledged that the variety of domain structures shown by some of the Chitinase sequences included in this study, such as the lack of certain domains, could be experimental artifacts caused by, for example, sequencing errors, or other technical difficulties that could have prevented the complete sequencing of intact sequences. Therefore, the absence of domains in the sequences analyzed in this study may not indicate their actual absence. For example, the signal peptide domain was not always present in all SMP Chitinases observed in this study.
Tyrosinase
Tyrosinase, which is a member of the Type-3 Copper-containing Metalloprotein superfamily, plays a critical role in physiological processes including oxygen transport, pigmentation and innate immunity as the key rate-limiting enzyme in melanin synthesis (Cerenius et al. 2008; Cieslak et al. 2011). In mollusks, Tyrosinase is also involved in shell coloration through melanin synthesis (Yu et al. 2018; Zhu et al. 2021). SMP Tyrosinases contain only a single Tyrosinase domain (PF00264). Regardless of its involvement in the shell biomineralization process, the signal peptide is not necessarily included in a Tyrosinase. A survey on various mollusk draft genomes on Tyrosinases with a single Tyrosinase domain showed that the copy numbers of the gene varies significantly among different molluscan species. For example, the bivalves apparently underwent multiple tandem duplications of their Tyrosinases (Cephalopoda: 4–15; Gastropoda: 3–12; Bivalve: 24–53) (Fig. 8A) (Aguilera et al. 2014).
Since the general support values of our trees are low, we cannot discuss the phylogenetic evolution of this gene based on with certainty. However, the topology of the tree suggests a possibility that after clade-specific tandem duplications, different copies of Tyrosinase were recruited in each clade for shell formation (Setiamarga 2021). Similar phenomena have also been observed for several SMPs, such as Chitinase and EGF-ZP, also surveyed and discussed in this study.
EGF-ZP domain-containing protein
EGF-ZP Domain-containing protein (EGF-ZP) is a type of protein composed of one or two EGF domain(s) and one ZP domain (Fig. 7A). In studies on bivalves and gastropods, EGF-ZP were often identified as SMPs. Meanwhile, a copy of the protein was also identified in N. pompilius SMPs (Setiamarga et al. 2021). The EGF domain functions by helping crystals to aggregate, forming multiple columnar prisms inside the crystals (Iwamoto et al. 2020). The ZP domain forms complexes and functions by assisting protein–protein interaction, forming complexes with other proteins which functions are biomineralization-related, such as Shematrin, Pif, and Serine Protease Inhibitor (Jain et al. 2018; Shimizu et al. 2022). Pif and Serine Protease Inhibitor are also present in the N. pompilius SMPs, suggesting the formation of similar complexes and crystal aggregates also probably occur during the shell formation of N. pompilius. Meanwhile, we could not detect the presence of EGF-ZP, Pif, and Serine Protease Inhibitor in the SMPs of the decapodiforms S. pharaonis and S. spirula. Although we cannot rule out experimental artifacts causing these proteins to be undetected in these species, there is also a possibility that the absence of these proteins indicate that the two decapodiform species do not employ the shell formation mechanism via this protein complexes.
Our EGF-ZP phylogenetic tree also suggested that the proteins employed as SMPs do not form a monophyletic clade (Fig. 7B). It seems that after an expansion of the copy number of the protein-coding gene in the genomes of mollusks, each clade recruited different copies of the gene for shell formation. This result is also in agreement with the result of a previous study by Shimizu et al. (2022).
Presumed functions of cephalopod shell matrix proteins and the mechanism of shell formation
In this study, three of the cephalopod SMPs shared with the conchiferans were reportedly produced by the hemocytes (CD109 antigen, Chitinase, and Tyrosinase) (e.g., Lin et al. 2002; Badariotti et al. 2007; Huang et al. 2010; Terwilliger 2007). A protein shared only among the cephalopods, SOUL, is also expressed by the hemocytes (Goodfellow et al. 2021). This is intriguing, because one of the suggested mechanisms of CaCO3 crystallization in mollusks is hemocyte-mediated biomineralization (Mount et al. 2004). It is well-known that pearl grafting (Schmitt et al. 2018) would trigger a significant hemocyte response, causing a high level of hemocyte accumulation during pearl-sac development (Kishore and Southgate 2015). The level of the response corresponds to the degree of damage to host tissue, as a part of the immunoreaction by the host oyster to recognize foreign materials (Kishore and Southgate 2015; Shen et al. 2020). While molecular and cellular studies on immunoreaction and tissue repair in cephalopods are still limited (Imperadore et al. 2022), proteomic analysis of hemocytes obtained from protozoan parasite-infected octopods indicates an elevated immune response and increased hemocyte activity (Castellanos-Martínez et al. 2014). Furthermore, infection-derived inflammations, which might also show possible color and morphological changes of the damaged tissues, have been observed in the regenerating soft tissues of octopods (Zullo and Imperadore 2019). These observations, taken altogether with those observed in pearl-sac development and pearl grafting, might suggest a potential involvement of the hemocytes and immunoreactions in the regeneration of damaged soft tissues, although further studies are still necessary to confirm this hypothesis.
Similar increase in hemocytes has been also observed in the shell repairing process in oysters (Ruddell 1971; López et al. 1997), suggesting the involvement of the hemocytes and SMPs in oyster’s shell formation and/or repair (Johnstone et al. 2008). Li et al. (2016) also showed evidence that circulating hemocytes can directly participate in the CaCO3 crystal formation in two species of pearl oyster (Pinctada fucata and Crassostrea virginica). Meanwhile, a recent study by Liu et al. (2023) reported the detection of a large amount of Hemocyanin in S. pharaonis’s SMPs, indicating the presence of hemocytes in the shell matrix of this species.
Interestingly, besides being consistently identified and categorized as SMPs, the three proteins shared among the conchiferans and the one shared solely among the cephalopods have been reported to play a role in the function of hemocytes related to immunity. This suggests that these proteins have multiple functions in both immunity and shell formation (A2M and CD109 antigen: Wyatt et al. 2014; Yazzie et al. 2015; Chitinase: Badariotti et al. 2007; Zhou et al. 2020; Tyrosinase: Zhang et al. 2006; Mao et al. 2018; Zhu et al. 2021; Xiong et al. 2022; SOUL: Goodfellow et al. 2021), as well as soft tissue regeneration. This could also explain the mechanism behind shell repair after damage, as soft tissue injuries may occur concurrently with damaged shells, for example, when caused by predation (Tsujino and Shigeta 2012). Therefore, we suggest that the proteins were co-opted and used in shell biomineralization-related processes in at least the ancestral lineage leading to the shelled mollusks. Future studies focusing on their actual functions in shell formation and biomineralization must be conducted to confirm this hypothesis.
Conclusions
In this study, we updated our previous findings on a set of shell matrix proteins (SMPs) of the living fossil N. pompilius, which is a member of an early branching group of extant cephalopods still retaining their external calcified shell, the nautiloids (Setiamarga et al. 2021). A set of 85 proteins was recovered through in silico analyses comparing our previously identified set of proteins and those of Zhang et al. (2021), and by mapping the sequences on the draft genome of the species (Huang et al. 2022). We succeeded in identifying several proteins with repeat motifs but with low sequence similarities to their putative homologs in other organisms, such as Nautilin-63 and a highly acidic protein. These proteins are thought to function by providing a framework for calcium carbonate crystallizations. By comparing the re-identified N. pompilius SMPs with those of a wide range of mollusk taxa, we identified a set of core proteins commonly shared in the conchiferans. We also identified the SMPs shared among three cephalopods (N. pompilius, S. pharaonis, and S. spirula), and between the latter two decapodiforms. The common SMPs shared between the two decapodiforms probably co-opted after the internalization of their shells.
Our phylogenetic analyses on a select five of N. pompilius SMPs, CD109 antigen, Chitinase, Tyrosinase, EGF-ZP, and SOUL, revealed their possible evolutionary scenarios. The five proteins were selected, because while they are known to be expressed in the hemocytes and play a role in immunity, studies in pearl and shell formation indicated their involvement in biomineralization. The phylogenetic analysis of CD109 antigen (TEPs), which was found in conchiferan SMPs, indicated that the different types of TEPs, mTEP and A2M, were utilized in external shell N. pompilius and internal shell S. pharaonis and S. spirula, respectively. Their phylogeny also suggests recruitments of different orthologs to carry out a similar function, which in this case is shell formation. The recruitments probably related to the internalization of the shell in the lineage. Meanwhile, phylogenetic analyses of Tyrosinase and Chitinase revealed that the two genes were probably recruited for shell formation independently in each molluscan lineage. Although low bootstrap values, probably caused by rapid evolution, have prevented a straightforward discussion about their evolution, it seems that extreme expansion of copy numbers through tandem duplications and losses of domains and genes leading to lineage-specific subfunctionalizations probably happened to these two genes. Meanwhile, SOUL was detected as an SMP only in the cephalopods. The cephalopod SMPs were recovered as a monophyletic clade with a high statistical support. This probably indicates that the protein was recruited for shell formation in the lineage leading to the cephalopods, and underwent a functional constraint causing them to maintain their sequences to be similar, suggesting that these proteins are probably crucial in cephalopod shell formation.
Data availability
In this study, we conducted bioinformatics analyses on publicly available data. The results of our analyses are either presented as the main tables and figures, or appended to this paper as supplementary materials. Newick files (.tre files) of all phylogenetic trees will be made available upon request.
References
Addadi L, Joester D, Nudelman F, Weiner S (2006) Mollusk shell formation: a source of new concepts for understanding biomineralization processes. Eur J Chem 12(4):980–987. https://doi.org/10.1002/chem.200500980
Aguilera F, McDougall C, Degnan BM (2014) Evolution of the tyrosinase gene family in bivalve molluscs: independent expansion of the mantle gene repertoire. Acta Biomater 10(9):3855–3865. https://doi.org/10.1016/j.actbio.2014.03.031
Albertin CB, Simakov O, Mitros T, Wang ZY, Pungor JR, Edsinger-Gonzales E, Brenner S, Ragsdale CW, Rokhsar DS (2015) The octopus genome and the evolution of cephalopod neural and morphological novelties. Nature 524(7564):220–224. https://doi.org/10.1038/nature14668
Almagro Armenteros JJ, Tsirigos KD, Sønderby CK, Petersen TN, Winther O, Brunak S, von Heijne G, Nielsen H (2019) SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat Biotechnol 37(4):420–423. https://doi.org/10.1038/s41587-019-0036-z
Arivalagan J, Marie B, Sleight VA, Clark MS, Berland S, Marie A (2016) Shell matrix proteins of the clam, Mya truncata: roles beyond shell formation through proteomic study. Mar Genom 27:69–74. https://doi.org/10.1016/j.margen.2016.03.005
Badariotti F, Lelong C, Dubos MP, Favrel P (2007) Characterization of Chitinase-like proteins (Cg-Clp1 and Cg-Clp2) involved in immune defence of the mollusc Crassostrea gigas. FEBS Lett 274(14):3646–3654. https://doi.org/10.1111/j.1742-4658.2007.05898.x
Barord GJ, Combosch DJ, Giribet G, Landman N, Lemer S, Veloso J, Ward PD (2023) Three new species of Nautilus Linnaeus, 1758 (Mollusca, Cephalopoda) from the Coral Sea and South Pacific. Zookeys 1143:51–69. https://doi.org/10.3897/zookeys.1143.84427
Beedham GE, Trueman ER (1968) The cuticle of the Aplacophora and its evolutionary significance in the Mollusca. J Zool 154(4):443–451. https://doi.org/10.1111/j.1469-7998.1968.tb01676.x
Budd GE (2003) The Cambrian fossil record and the origin of the phyla. Integr Comp Biol 43(1):157–165. https://doi.org/10.1093/icb/43.1.157
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25(15):1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Carter JG (1990) Evolutionary significance of shell microstructure in the Palaeotaxodonta, Pteriomorphia and Isofilibranchia (Bivalvia: Mollusca). Skelet Biominer Patt Process Evol Trends 1:135–296
Castellanos-Martínez S, Diz AP, Álvarez-Chaver P, Gestal C (2014) Proteomic characterization of the hemolymph of Octopus vulgaris infected by the protozoan parasite Aggregata octopiana. J Proteomics 105:151–163. https://doi.org/10.1016/j.jprot.2013.12.008
Cerenius L, Lee BL, Söderhäll K (2008) The proPO-system: pros and cons for its role in invertebrate immunity. Trends Immunol 29(6):263–271. https://doi.org/10.1016/j.it.2008.02.009
Checa AG, Grenier C, Griesshaber E, Schmahl WW, Cartwright JH, Salas C, Oudot M (2022) The shell structure and chamber production cycle of the cephalopod Spirula (Coleoidea, Decabrachia). Mar Biol 169(10):132. https://doi.org/10.1007/s00227-022-04120-0
Cieslak M, Reissmann M, Hofreiter M, Ludwig A (2011) Colours of domestication. Biol Rev Biol 86(4):885–899. https://doi.org/10.1111/j.1469-185X.2011.00177.x
Cowen R (2013) History of life, 5th edn. John Wiley & Sons, p 312 (ISBN: 978-0470671726)
Crenshaw MA (1989) Biomineralization mechanisms. Skelet Biominer Patt Process Evol Trends 5:1–9
Da Fonseca RR, Couto A, Machado AM, Brejova B, Albertin CB, Silva F, Gardner P, Baril T, Hayward A, Campos A, Ribeiro ÂM (2020) A draft genome sequence of the elusive giant squid, Architeuthis dux. GigaScience 9(1):giz152. https://doi.org/10.1093/gigascience/giz152
Dauphin Y (2023) A brief history of biomineralization studies. ACS Biomater Sci 9(4):1774–1790. https://doi.org/10.1021/acsbiomaterials.1c01385
Dauphin Y, Luquet G, Percot A, Bonnaud-Ponticelli L (2020) Comparison of embryonic and adult shells of Sepia officinalis (Cephalopoda, Mollusca). Zoomorphology 139:151–169. https://doi.org/10.1007/s00435-020-00477-2
Duval D, Pichon R, Lassalle D, Laffitte M, Gourbal B, Galinier R (2020) A new assessment of thioester-containing proteins diversity of the freshwater snail Biomphalaria glabrata. Genes 11(1):69. https://doi.org/10.3390/genes11010069
Eddy SR (2009) A new generation of homology search tools based on probabilistic inference. Genome Inform 23:205–211. https://doi.org/10.1142/9781848165632_0019
Feng D, Li Q, Yu H, Kong L, Du S (2017) Identification of conserved proteins from diverse shell matrix proteome in Crassostrea gigas: characterization of genetic bases regulating shell formation. Sci Rep 7(1):1–12. https://doi.org/10.1038/srep45754
Fortey R (2011) Survivors: the Animals and plants that time has left behind. HarperCollins, p 336 (ISBN: 978-0007209873)
Fortunato AE, Sordino P, Andreakis N (2016) Evolution of the SOUL Heme-Binding protein superfamily across eukarya. J Mol Evol 82:279–290. https://doi.org/10.1007/s00239-016-9745-9
Furuhashi T, Schwarzinger C, Miksik I, Smrz M, Beran A (2009) Molluscan shell evolution with review of shell calcification hypothesis. Comp Biochem Physiol B Biochem Mol Biol 154(3):351–371. https://doi.org/10.1016/j.cbpb.2009.07.011
Frenzel M, Harper EM (2011) Micro-structure and chemical composition of vateritic deformities occurring in the bivalve Corbicula fluminea (Müller, 1774). J Struct Biol 174(2):321–332. https://doi.org/10.1016/j.jsb.2011.02.002
Gatesy J, Hayashi C, Motriuk D, Woods J, Lewis R (2001) Extreme diversity, conservation, and convergence of spider silk fibroin sequences. Science 291(5513):2603–2605. https://doi.org/10.1126/science.1057561
Goodfellow BJ, Freire F, Carvalho AL, Aveiro SS, Charbonnier P, Moulis JM, Delgado L, Ferreira GC, Rodrigues JE, Poussin-Courmontagne P, Birck C, McEwen A, Macedo AL (2021) The SOUL family of heme-binding proteins: structure and function 15 years later. Coord Chem Rev 448:214189. https://doi.org/10.1016/j.ccr.2021.214189
Gotliv BA, Kessler N, Sumerel JL, Morse DE, Tuross N, Addadi L, Weiner S (2005) Asprich: a novel aspartic acid-rich protein family from the prismatic shell matrix of the bivalve Atrina rigida. ChemBioChem 6(2):304–314. https://doi.org/10.1002/cbic.200400221
Hare PE, Abelson PH (1965) Amino acid composition of some calcified proteins. Carnegie Inst Wash Yearbk 64:223–232
Harting P (1872) On the artificial production of some of the principal organic calcareous formations. J Cell Sci 12(46):118–123. https://doi.org/10.1242/jcs.s2-12.46.118
Huang QS, Yan JH, Tang JY, Tao YM, Xie XL, Wang Y, Wei XQ, Yan QH, Chen QX (2010) Cloning and tissue expressions of seven chitinase family genes in Litopenaeus vannamei. Fish Shellfish Immunol 29(1):75–81. https://doi.org/10.1016/j.fsi.2010.02.014
Huang Z, Huang W, Liu X, Han Z, Liu G, Boamah GA, Wang Y, Yu F, Gan Y, Xiao Q, Luo X (2022) Genomic insights into the adaptation and evolution of the nautilus, an ancient but evolving “living fossil.” Mol Ecol Resour 22(1):15–27. https://doi.org/10.1111/1755-0998.13439
Imperadore P, Galli R, Winterhalder MJ, Zumbusch A, Uckermann O (2022) Imaging arm regeneration: label-free multiphoton microscopy to dissect the process in octopus vulgaris. Front Cell Dev Biol 10:18. https://doi.org/10.3389/fcell.2022.814746
Ishikawa A, Shimizu K, Isowa Y, Takeuchi T, Zhao R, Kito K, Fujie M, Satoh N, Endo K (2020) Functional shell matrix proteins tentatively identified by asymmetric snail shell morphology. Sci Rep 10(1):1–12. https://doi.org/10.1038/s41598-020-66021-w
Isowa Y, Sarashina I, Setiamarga DH, Endo K (2012) A comparative study of the shell matrix protein aspein in pterioid bivalves. J Mol Evol 75(1):11–18. https://doi.org/10.1007/s00239-012-9514-3
Iwamoto S, Shimizu K, Negishi L, Suzuki N, Nagata K, Suzuki M (2020) Characterization of the chalky layer-derived EGF-like domain-containing protein (CgELC) in the pacific oyster, Crassostrea gigas. J Struct Biol 212(1):107594. https://doi.org/10.1016/j.jsb.2020.107594
Jackson DJ, McDougall C, Woodcroft B, Moase P, Rose RA, Kube M, Reinhardt R, Rokhsar DS, Montagnani C, Joubert C, Piquemal D, Degnan B (2010) Parallel evolution of nacre building gene sets in molluscs. Mol Biol Evol 27(3):591–608. https://doi.org/10.1093/molbev/msp278
Jain G, Pendola M, Huang YC, Gebauer D, Koutsoumpeli E, Johnson S, Evans JS (2018) Selective synergism created by interactive nacre framework-associated proteins possessing EGF and vWA motifs: implications for mollusk shell formation. Biochem 57(18):2657–2666. https://doi.org/10.1021/acs.biochem.8b00119
Johnstone MB, Ellis S, Mount AS (2008) Visualization of shell matrix proteins in hemocytes and tissues of the Eastern oyster, Crassostrea virginica. J Exp Zool B Mol 310(3):227–239. https://doi.org/10.1002/jez.b.21206
Jones P, Binns D, Chang HY, Fraser M, Li W, McAnulla C, McWilliam H, Maslen J, Mitchell A, Nuka G, Pesseat S, Quinn AF, Sangrador-Vegas A, Scheremetjew M, Yong SY, Lopez R, Hunter S (2014) InterProScan 5: genome-scale protein function classification. Bioinformatics 30(9):1236–1240. https://doi.org/10.1093/bioinformatics/btu031
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Keith J, Stockwell S, Ball D, Remillard K, Kaplan D, Thannhauser T, Sherwood R (1993) Comparative analysis of macromolecules in mollusc shells. Comp Biochem Physiol B Biochem Mol Biol 105(3–4):487–496. https://doi.org/10.1016/0305-0491(93)90078-J
Kim BM, Kang S, Ahn DH, Jung SH, Rhee H, Yoo JS, Lee JE, Lee S, Han YH, Ryu KB, Cho SJ (2018) The genome of common long-arm octopus Octopus minor. Gigascience 7(11):giy119. https://doi.org/10.1093/gigascience/giy119
Kishore P, Southgate PC (2015) Haemocyte persistence after grafting for pearl production in Pinctada margaritifera (Linnaeus, 1758). Fish Shellfish Immunol 42(2):530–532. https://doi.org/10.1016/j.fsi.2014.11.035
Kocot KM, Cannon JT, Todt C, Citarella MR, Kohn AB, Meyer A, Santos SR, Schander C, Moroz LL, Lieb B, Halanych KM (2011) Phylogenomics reveals deep molluscan relationships. Nature 477(7365):452–456. https://doi.org/10.1038/nature10382
Kocot KM, Aguilera F, McDougall C, Jackson DJ, Degnan BM (2016) Sea shell diversity and rapidly evolving secretomes: insights into the evolution of biomineralization. Front Zool 13:1–10. https://doi.org/10.1186/s12983-016-0155-z
Kocot KM, McDougall C, Degnan BM (2017) Developing perspectives on molluscan shells, part 1: Introduction and molecular biology. Physiology of molluscs: a collection of selected reviews Apple Academic Press, pp 1–41. https://doi.org/10.1201/9781315207483
Kocot KM, Todt C, Mikkelsen NT, Halanych KM (2019) Phylogenomics of Aplacophora (Mollusca, Aculifera) and a solenogaster without a foot. Proc R Soc B 286(1902):20190115. https://doi.org/10.1098/rspb.2019.0115
Kocot KM, Poustka AJ, Stöger I, Halanych KM, Schrödl M (2020) New data from Monoplacophora and a carefully-curated dataset resolve molluscan relationships. Sci Rep 10(1):1–8. https://doi.org/10.1038/s41598-019-56728-w
Kröger B, Vinther J, Fuchs D (2011) Cephalopod origin and evolution: a congruent picture emerging from fossils, development and molecules: extant cephalopods are younger than previously realised and were under major selection to become agile, shell-less predators. BioEssays 33(8):602–613. https://doi.org/10.1002/bies.201100001
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19(9):1639–1645. https://doi.org/10.1101/gr.092759.109
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Layer G, Jahn D, Deery E, Lawrence AD, Warren MJ (2010) 7.13 - Biosynthesis of heme and vitamin B12. In: Liu HW, Mander L (eds) Comprehensive natural products II – chemistry and biology, vol 7. Elsevier, pp 445–499 (ISBN: 978-0080453828)
Lei M, Wu R (1991) A novel glycine-rich cell wall protein gene in rice. Plant Mol Biol 16:187–198. https://doi.org/10.1007/BF00020551
Letunic I, Khedkar S, Bork P (2021) SMART: Recent updates, new developments and status in 2020. Nucleic Acids Res 49(D1):D458–D460. https://doi.org/10.1093/nar/gkaa937
Li S, Liu Y, Liu C, Huang J, Zheng G, Xie L, Zhang R (2016) Hemocytes participate in calcium carbonate crystal formation, transportation and shell regeneration in the pearl oyster Pinctada fucata. Fish Shellfish Immunol 51:263–270. https://doi.org/10.1016/j.fsi.2016.02.027
Liao Z, Bao LF, Fan MH, Gao P, Wang XX, Qin CL, Li XM (2015) In-depth proteomic analysis of nacre, prism, and myostracum of Mytilus shell. J Proteomics 122:26–40. https://doi.org/10.1016/j.jprot.2015.03.027
Lin M, Sutherland DR, Horsfall W, Totty N, Yeo E, Nayar R, Wu XF, Schuh AC (2002) Cell surface antigen CD109 is a novel member of the α2 Macroglobulin/C3, C4, C5 family of thioester-containing proteins. Blood 99(5):1683–1691. https://doi.org/10.1182/blood.V99.5.1683
Lindberg DR (2001) Molluscs. In: Scheiner S (ed) Encyclopedia of biodiversity, 2nd edn. Elsevier, pp 373–383. https://doi.org/10.1016/B978-0-12-384719-5.00096-4 (ISBN: 9780123847201)
Liu C, Li S, Kong J, Liu Y, Wang T, Xie L, Zhang R (2015) In-depth proteomic analysis of shell matrix proteins of Pinctada fucata. Sci Rep 5(1):1–14. https://doi.org/10.1038/srep17269
Liu F, Li Y, Yu H, Zhang L, Hu J, Bao Z, Wang S (2021) MolluscDB: an integrated functional and evolutionary genomics database for the hyper-diverse animal phylum Mollusca. Nucleic Acids Res 49(D1):D988–D997. https://doi.org/10.1093/nar/gkaa918
Liu C, Ji X, Huang J, Wang Z, Liu Y, Hincke MT (2023) Proteomics of shell matrix proteins from the cuttlefish bone reveals unique evolution for cephalopod biomineralization. ACS Biomater Sci Eng 9(4):1796–1807. https://doi.org/10.1021/acsbiomaterials.1c00693
Lowenstam HA (1981) Minerals formed by organisms. Science 211(4487):1126–1131. https://doi.org/10.1126/science.7008198
Lowenstam HA, Traub W, Weiner S (1984) Nautilus hard parts: a study of the mineral and organic constitutents. Paleobiology 10(2):268–279. https://doi.org/10.1017/S0094837300008198
Lowenstam HA, Weiner S (1989) On biomineralization. Oxford University Press, p 336 (ISBN: 978-0195049770)
López C, Carballal MJ, Azevedo C, Villalba A (1997) Morphological characterization of the hemocytes of the clam, Ruditapes decussatus (Mollusca: Bivalvia). J Invertebr Pathol 69(1):51–57. https://doi.org/10.1006/jipa.1996.4639
Mann K, Edsinger E (2014) The Lottia gigantea shell matrix proteome: re-analysis including MaxQuant iBAQ quantitation and phosphoproteome analysis. Proteome Sci 12(1):1–12. https://doi.org/10.1186/1477-5956-12-28
Mann K, Jackson DJ (2014) Characterization of the pigmented shell-forming proteome of the common grove snail Cepaea nemoralis. BMC Genom 15:1–16. https://doi.org/10.1186/1471-2164-15-249
Mann K, Cerveau N, Gummich M, Fritz M, Mann M, Jackson DJ (2018) In-depth proteomic analyses of Haliotis laevigata (greenlip abalone) nacre and prismatic organic shell matrix. Proteome Sci 16(1):1–25. https://doi.org/10.1186/s12953-018-0139-3
Mao J, Zhang W, Zhang X, Tian Y, Wang X, Hao Z, Chang Y (2018) Transcriptional changes in the Japanese scallop (Mizuhopecten yessoensis) shell infested by Polydora provide insights into the molecular mechanism of shell formation and immunomodulation. Sci Rep 8(1):17664. https://doi.org/10.1038/s41598-018-35749-x
Marie B, Marin F, Marie A, Bédouet L, Dubost L, Alcaraz G, Milet C, Luquet G (2009) Evolution of nacre: biochemistry and proteomics of the shell organic matrix of the cephalopod Nautilus macromphalus. ChemBioChem 10(9):1495–1506. https://doi.org/10.1002/cbic.200900009
Marie B, Zanella-Cléon I, Corneillat M, Becchi M, Alcaraz G, Plasseraud L, Luquet G, Marin F (2011) Nautilin-63, a novel acidic glycoprotein from the shell nacre of Nautilus macromphalus. FEBS J 278(12):2117–2130. https://doi.org/10.1111/j.1742-4658.2011.08129.x
Marie B, Joubert C, Tayalé A, Zanella-Cléon I, Belliard C, Piquemal D, Cochennec-Laureau N, Marin F, Gueguen Y, Montagnani C (2012) Different secretory repertoires control the biomineralization processes of prism and nacre deposition of the pearl oyster shell. Proc Natl Acad Sci USA 109(51):20986–20991. https://doi.org/10.1073/pnas.1210552109
Marie B, Jackson DJ, Ramos-Silva P, Zanella-Cléon I, Guichard N, Marin F (2013) The shell-forming proteome of Lottia gigantea reveals both deep conservations and lineage-specific novelties. FEBS J 280(1):214–232. https://doi.org/10.1111/febs.12062
Marie B, Arivalagan J, Mathéron L, Bolbach G, Berland S, Marie A, Marin F (2017) Deep conservation of bivalve nacre proteins highlighted by shell matrix proteomics of the Unionoida Elliptio complanata and Villosa lienosa. J R Soc Interface 14(126):20160846. https://doi.org/10.1098/rsif.2016.0846
Marin F, Corstjens P, de Gaulejac B, de Vrind-De JE, Westbroek P (2000) Mucins and molluscan calcification: molecular characterization of mucoperlin, a novel mucin-like protein from the nacreous shell layer of the fan mussel Pinna nobilis (Bivalvia, Pteriomorphia). J Biol Chem 275(27):20667–20675. https://doi.org/10.1074/jbc.M003006200
Marin F, Amons R, Guichard N, Stigter M, Hecker A, Luquet G, Layrolle P, Alcaraz G, Riondet C, Westbroek P (2005) Caspartin and calprismin, two proteins of the shell calcitic prisms of the Mediterranean fan mussel Pinna nobilis. J Biol Chem 280(40):33895–33908. https://doi.org/10.1074/jbc.M506526200
Marin F, Le Roy N, Marie B (2012) The formation and mineralization of mollusk shell. Front Biosci 4(3):1099–1125
Marin F, Marie B, Hamada SB, Ramos-Silva P, Le Roy N, Guichard N, Wolf S, Montagnani C, Joubert C, Piquemal D, Saulnier D, Gueguen Y (2013) “Shellome”: Proteins involved in mollusc shell biomineralization–diversity, functions. In: Watabe S, Maeyama K, Nagasawa H (eds) Recent advances in pearl research–Proceedings of the International Symposium on Pearl Research 2011. Terrapub, pp 149–166 (ISBN: 978-4887041585)
Marquez J, Dinguirard N, Gonzalez A, Kane AE, Joffe NR, Yoshino TP, Castillo MG (2022) Molecular characterization of thioester-containing proteins in Biomphalaria glabrata and their differential gene expression upon Schistosoma mansoni exposure. Front Immunol. https://doi.org/10.3389/fimmu.2022.903158
McDougall C, Aguilera F, Degnan BM (2013) Rapid evolution of pearl oyster shell matrix proteins with repetitive, low-complexity domains. J R Soc Interface 10(82):2013004. https://doi.org/10.1098/rsif.2013.0041
McDougall C, Degnan BM (2018) The evolution of mollusc shells. Wiley Interdiscip Rev Dev Biol 7(3):e313. https://doi.org/10.1002/wdev.313
Meenakshi VR, Blackwelder P, Hare PE, Wilbur KM, Watabe N (1975) Studies on shell regeneration—i. matrix and mineral composition of the normal and regenerated shell of Pomacea paludosa. Comp Biochem Physiol Mol Integr Physiol 50(2):347. https://doi.org/10.1016/0300-9629(75)90024-9
Mikasa T, Kugo M, Nishimura S, Taketani S, Ishijima S, Sagami I (2018) Thermodynamic characterization of the Ca2+-dependent interaction between SOUL and ALG-2. Int J Mol Sci 19(12):3802. https://doi.org/10.3390/ijms19123802
Miyamoto H, Yano M, Miyashita T (2003) Similarities in the structure of Nacrein, the shell-matrix protein, in a bivalve and a gastropod. J Molluscan Stud 69(1):87–89. https://doi.org/10.1093/mollus/69.1.87
Montagnani C, Marie B, Marin F, Belliard C, Riquet F, Tayalé A, Zanella-Cléon I, Fleury E, Gueguen Y, Piquemal D, Cochennec-Laureau N (2011) Pmarg-Pearlin is a matrix protein involved in nacre framework formation in the pearl oyster Pinctada margaritifera. ChemBioChem 12(13):2033–2043. https://doi.org/10.1002/cbic.201100216
Mount AS, Wheeler AP, Paradkar RP, Snider D (2004) Hemocyte-mediated shell mineralization in the eastern oyster. Science 304(5668):297–300. https://doi.org/10.1126/science.109050
Nakashima K, Iwashita S, Suzuki T, Kato C, Kohno T, Kamei Y, Sasaki M, Urayama O, Ohno-Iwashita Y, Dohmae N, Song SY (2019) A spatial similarity of stereochemical environments formed by amino acid residues defines a common epitope of two non-homologous proteins. Sci Rep 9(1):1–10. https://doi.org/10.1038/s41598-019-51350-2
Nielsen H, Tsirigos KD, Brunak S, von Heijne G (2019) A brief history of protein sorting prediction. Protein J 38(3):200–216. https://doi.org/10.1007/s10930-019-09838-3
Nonaka M, Kimura A (2006) Genomic view of the evolution of the complement system. Immunogenetics 58:701–713. https://doi.org/10.1007/s00251-006-0142-1
Numata K (2020) How to define and study structural proteins as biopolymer materials. Polym J 52(9):1043–1056. https://doi.org/10.1038/s41428-020-0362-5
Oudot M, Neige P, Shir IB, Schmidt A, Strugnell JM, Plasseraud L, Broussard C, Hoffmann R, Lukeneder A, Marin F (2020) The shell matrix and microstructure of the ram’s horn squid: Molecular and structural characterization. J Struct Biol 211(1):107507. https://doi.org/10.1016/j.jsb.2020.107507
Pearson WR, Sierk ML (2005) The limits of protein sequence comparison? Curr Opin Struct Biol 15(3):254–260. https://doi.org/10.1016/j.sbi.2005.05.005
Pomar L (2020) Carbonate systems. In: Scarselli N, Adam J, Chiarella D (eds) Regional geology and tectonics principles of geologic analysis (second edition), vol 1. Elsevier, pp 235–311. https://doi.org/10.1016/B978-0-444-64134-2.00013-4
Ponder WF, Lindberg DR, Ponder JM (2019) Biology and evolution of the mollusca, vol 1. CRC Press, p 924 (ISBN: 978-0815361695)
Portet A, Galinier R, Pinaud S, Portela J, Nowacki F, Gourbal B, Duval D (2018) BgTEP: an antiprotease involved in innate immune sensing in Biomphalaria glabrata. Front Immunol 9:1206. https://doi.org/10.3389/fimmu.2018.01206
Prié V (2019) Molluscs. In: White W, Culver D, Pipan T (eds) Encyclopedia of Caves, 3rd edn. Academic Press, Cham, pp 725–731
Ruddell CL (1971) The fine structure of oyster agranular amebocytes from regenerating mantle wounds in the Pacific oyster. Crassostrea Gigas J Invertebr Pathol 18(2):260–268
Samata T, Hayashi N, Kono M, Hasegawa K, Horita C, Akera S (1999) A new matrix protein family related to the nacreous layer formation of Pinctada fucata. Febs Lett 462(1–2):225–229. https://doi.org/10.1016/S0014-5793(99)01387-3
Samata T, Ikeda D, Kajikawa A, Sato H, Nogawa C, Yamada D, Yamazaki R, Akiyama T (2008) A novel phosphorylated glycoprotein in the shell matrix of the oyster Crassostrea nippona. The FEBS J 275(11):2977–2989. https://doi.org/10.1111/j.1742-4658.2008.06453.x
Sánchez R, Serra F, Tárraga J, Medina I, Carbonell J, Pulido L, De María A, Capella-Gutíerrez S, Huerta-Cepas J, Gabaldón T, Dopazo J, Dopazo H (2011) Phylemon 2.0: a suite of web-tools for molecular evolution, phylogenetics, phylogenomics and hypotheses testing. Nucleic Acids Res 39(suppl 2):W470–W474. https://doi.org/10.1093/nar/gkr408
Sarashina I, Endo K (1998) Primary structure of a soluble matrix protein of scallop shell; implications for calcium carbonate biomineralization. Am Min 83(11–12 part 2):1510–1515. https://doi.org/10.2138/am-1998-11-1239
Saunders WB (1981) The species of living Nautilus and their distribution. Veliger 24:8–17
Saunders WB, Landman N (2009) Nautilus: the biology and paleobiology of a living fossil, reprint with additions, vol 6. Springer Science & Business Media, pp 463–486
Schmitt N, Marin F, Thomas J, Plasseraud L, Demoy-Schneider M (2018) Pearl grafting: tracking the biological origin of nuclei by straightforward immunological methods. Aquac Res 49(2):692–700. https://doi.org/10.1111/are.13499
Scheltema AH (1993) Aplacophora as progenetic aculiferans and the coelomate origin of mollusks as the sister taxon of Sipuncula. Biol Bull 184(1):57–78. https://doi.org/10.2307/1542380
Schöne BR, Dunca E, Fiebig J, Pfeiffer M (2005) Mutvei’s solution: an ideal agent for resolving microgrowth structures of biogenic carbonates. Palaeogeogr Palaeoclimatol Palaeoecol 228(1–2):149–166. https://doi.org/10.1016/j.palaeo.2005.03.054
Sekiguchi R, Fujito NT, Nonaka M (2012) Evolution of the thioester-containing proteins (TEPs) of the arthropoda, revealed by molecular cloning of TEP genes from a spider, Hasarius Adansoni. Dev Comp Immunol 36(2):483–489. https://doi.org/10.1016/j.dci.2011.05.003
Setiamarga DH, Hirota K, Yoshida MA, Takeda Y, Kito K, Ishikawa M, Shimizu K, Isowa Y, Ikeo K, Sasaki T, Endo K (2021) Hydrophilic shell matrix proteins of Nautilus pompilius and the identification of a core set of conchiferan domains. Genes 12(12):1925. https://doi.org/10.3390/genes12121925
Setiamarga DH (2021) Evolutionary saga of the cephalopod shell: A molecular paleontological approach. Aquabiology 43(3):270–279 (In Japanese with English abstract)
Shen W, Hu Y, He Z, Xu S, Xu X, Zhang G, Ren G (2020) Histological and comparative transcriptome analyses provide insights into the immune response in pearl sac formation of Hyriopsis cumingii. Front Mar Sci 7:256. https://doi.org/10.3389/fmars.2020.00256
Shimizu K, Kimura K, Isowa Y, Oshima K, Ishikawa M, Kagi H, Kito K, Hattori M, Chiba S, Endo K (2019) Insights into the evolution of shells and love darts of land snails revealed from their matrix proteins. Genome Biol Evol 11(2):380–397. https://doi.org/10.1093/gbe/evy242
Shimizu K, Kintsu H, Awaji M, Matumoto T, Suzuki M (2020) Evolution of biomineralization genes in the prismatic layer of the pen shell Atrina pectinata. J Mol Evol 88:742–758. https://doi.org/10.1007/s00239-020-09977-7
Shimizu K, Negishi L, Ito T, Touma S, Matsumoto T, Awaji M, Kurumizaka H, Yoshitake K, Kinoshita S, Asakawa S, Suzuki M (2022) Evolution of nacre-and prisms-related shell matrix proteins in the pen shell, Atrina pectinata. Comp Biochem Physiol D Genom Proteom 44:101025. https://doi.org/10.1016/j.cbd.2022.101025
Sigwart JD, Sutton MD (2007) Deep molluscan phylogeny: synthesis of palaeontological and neontological data. Proc R Soc b: Biol Sci 274(1624):2413–2419. https://doi.org/10.1098/rspb.2007.0701
Simkiss K, Wilbur KM (1989) Biomineralization. Academic Press, p 337 (ISBN: 9780080925844)
Smith SA, Wilson NG, Goetz FE, Feehery C, Andrade SC, Rouse GW, Giribet G, Dunn CW (2011) Resolving the evolutionary relationships of molluscs with phylogenomic tools. Nature 480(7377):364–367. https://doi.org/10.1038/nature10526
Sommerdijk NA, Cölfen H (2010) Lessons from nature—biomimetic approaches to minerals with complex structures. MRS Bull 35(2):116–121. https://doi.org/10.1557/mrs2010.630
Song X, Liu Z, Wang L, Song L (2019) Recent advances of shell matrix proteins and cellular orchestration in marine molluscan shell biomineralization. Front Mar Sci 6:41. https://doi.org/10.3389/fmars.2019.00041
Spann N, Harper EM, Aldridge DC (2010) The unusual mineral vaterite in shells of the freshwater bivalve Corbicula fluminea from the UK. Naturwissenschaften 97(8):743–751. https://doi.org/10.1007/s00114-010-0692-9
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stecher G, Tamura K, Kumar S (2020) Molecular evolutionary genetics analysis (MEGA) for macOS. Mol Biol Evol 37(4):1237–1239. https://doi.org/10.1093/molbev/msz312
Sudo S, Fujikawa T, Nagakura T, Ohkubo T, Sakaguchi K, Tanaka M, Nakashima K, Takahashi T (1997) Structures of mollusc shell framework proteins. Nature 387(6633):563–564. https://doi.org/10.1038/42391
Suzuki M, Saruwatari K, Kogure T, Yamamoto Y, Nishimura T, Kato T, Nagasawa H (2009) An acidic matrix protein, Pif, is a key macromolecule for nacre formation. Science 325(5946):1388–1390 https://www.science.org/doi/10.1126/science.1173793
Suzuki M, Iwashima A, Tsutsui N, Ohira T, Kogure T, Nagasawa H (2011) Identification and characterisation of a calcium carbonate-binding protein, blue mussel shell protein (BMSP), from the nacreous layer. Chembiochem 12(16):2478–2487. https://doi.org/10.1002/cbic.201100317
Suzuki M, Iwashima A, Kimura M, Kogure T, Nagasawa H (2013) The molecular evolution of the pif family proteins in various species of mollusks. Mar Biotechnol 15:145–158. https://doi.org/10.1007/s10126-012-9471-2
Taketani S, Adachi Y, Kohno H, Ikehara S, Tokunaga R, Ishii T (1998) Molecular characterization of a newly identified heme-binding protein induced during differentiation of urine erythroleukemia cells. J Biol Chem 273(47):31388–31394. https://doi.org/10.1074/jbc.273.47.31388
Takeuchi T, Sarashina I, Iijima M, Endo K (2008) In vitro regulation of CaCO3 crystal polymorphism by the highly acidic molluscan shell protein Aspein. FEBS Lett 582(5):591–596. https://doi.org/10.1016/j.febslet.2008.01.026
Terwilliger NB (2007) Hemocyanins and the immune response: defense against the dark arts. Integr Comp Biol 47(4):662–665. https://doi.org/10.1093/icb/icm039
Todt C, Wanninger A (2010) Of tests, trochs, shells, and spicules: development of the basal mollusk Wirenia argentea (Solenogastres) and its bearing on the evolution of trochozoan larval key features. Front Zool 7(1):1–17. https://doi.org/10.1186/1742-9994-7-6
Tsujino Y, Shigeta Y (2012) Biological response to experimental damage of the phragmocone and siphuncle in Nautilus pompilius Linnaeus. Lethaia 45(3):443–449. https://doi.org/10.1111/j.1502-3931.2012.00306.x
Tsukamoto D, Sarashina I, Endo K (2004) Structure and expression of an unusually acidic matrix protein of pearl oyster shells. Biochem Biophys Res Commun 320(4):1175–1180. https://doi.org/10.1016/j.bbrc.2004.06.072
Velázquez-Castillo R, Reyes-Gasga J, García-Gutierrez DI, Jose-Yacaman M (2006) Nanoscale characterization of nautilus shell structure: an example of natural self-assembly. J Mater Res 21(6):1484–1489. https://doi.org/10.1557/jmr.2006.0190
Vinther J (2015) The origins of molluscs. Palaeontology 58(1):19–34. https://doi.org/10.1111/pala.12140
Wanninger A, Wollesen T (2019) The evolution of molluscs. Biol Rev 94(1):102–115. https://doi.org/10.1111/brv.12439
Ward P, Carlson B, Weekly M, Brumbaugh B (1984) Remote telemetry of daily vertical and horizontal movement of Nautilus in Palau. Nature 309(5965):248–250. https://doi.org/10.1038/309248a0
Weiner S (1979) Aspartic acid-rich proteins: major components of the soluble organic matrix of mollusk shells. Calcif Tissue Int 29:163–167. https://doi.org/10.1007/BF02408072
Weiner S, Hood L (1975) Soluble protein of the organic matrix of mollusk shells: a potential template for shell formation. Science 190(4218):987–989. https://doi.org/10.1126/science.1188379
Weiner S, Traub W (1980) X-ray diffraction study of the insoluble organic matrix of mollusk shells. FEBS Lett 111(2):311–316. https://doi.org/10.1016/0014-5793(80)80817-9
Weiner S, Talmon Y, Traub W (1983) Electron diffraction of mollusc shell organic matrices and their relationship to the mineral phase. Int J Biol Macromol 5(6):325–328. https://doi.org/10.1016/0141-8130(83)90055-7
Wernström JV, Gąsiorowski L, Hejnol A (2022) Brachiopod and mollusc biomineralisation is a conserved process that was lost in the phoronid–bryozoan stem lineage. EvoDevo 13:17. https://doi.org/10.1186/s13227-022-00202-8
Woodland W (1907) Studies in spicule formation. VI. The scleroblastic development of the spicules in some Mollusca and in one genus of colonial ascidians. J Cell Sci 51(201):45–54. https://doi.org/10.1242/jcs.s2-51.201.45
Wray CG, Landman NH, Saunders WB, Bonacum J (1995) Genetic divergence and geographic diversification in Nautilus. Paleobiology 21(2):220–228. https://doi.org/10.1017/S009483730001321X
Wyatt AR, Kumita JR, Mifsud RW, Gooden CA, Wilson MR, Dobson CM (2014) Hypochlorite-induced structural modifications enhance the chaperone activity of human α2-macroglobulin. PNAS 111(20):E2081–E2090
Xiong X, Cao Y, Li Z, Huang R, Jiao Y, Zhao L, Du X, Zheng Z (2022) Insights from tyrosinase into the impacts of modified morphology of calcium carbonate on the nacre formation of pearl oysters. Front Mar Sci. https://doi.org/10.3389/fmars.2022.935609
Yano M, Nagai K, Morimoto K, Miyamoto H (2006) Shematrin: a family of glycine-rich structural proteins in the shell of the pearl oyster Pinctada fucata. Comp Biochem Physiol B Biochem Mol Biol 144(2):254–262. https://doi.org/10.1016/j.cbpb.2006.03.004
Yazzie N, Salazar KA, Castillo MG (2015) Identification, molecular characterization, and gene expression analysis of a CD109 molecule in the Hawaiian bobtail squid Euprymna scolopes. Fish Shellfish Immunol 44(1):342–355. https://doi.org/10.1016/j.fsi.2015.02.036
Yoshida MA, Hirota K, Imoto J, Okuno M, Tanaka H, Kajitani R, Toyoda A, Itoh T, Ikeo K, Sasaki T, Setiamarga DH (2022) Gene recruitments and dismissals in the argonaut genome provide insights into pelagic lifestyle adaptation and shell-like eggcase reacquisition. Genome Biol Evol 14(11):evac140. https://doi.org/10.1093/gbe/evac140
Yu F, Pan Z, Qu B, Yu X, Xu K, Deng Y, Liang F (2018) Identification of a tyrosinase gene and its functional analysis in melanin synthesis of Pteria penguin. Gene 656:1–8. https://doi.org/10.1016/j.gene.2018.02.060
Zaquin T, Malik A, Drake JL, Putnam HM, Mass T (2021) Evolution of protein-mediated biomineralization in scleractinian corals. Front Genet 12:618517. https://doi.org/10.3389/fgene.2021.618517
Zhang C, Xie L, Huang J, Chen L, Zhang R (2006) A novel putative tyrosinase involved in periostracum formation from the pearl oyster (Pinctada fucata). Biochem Biophys Res Commun 342(2):632–639. https://doi.org/10.1016/j.bbrc.2006.01.182
Zhang M, Wang Y, Li Y, Li W, Li R, Xie X, Wang S, Hu X, Zhang L, Bao Z (2018) Identification and characterization of neuropeptides by transcriptome and proteome analyses in a bivalve mollusc Patinopecten yessoensis. Front Genet 9:197. https://doi.org/10.3389/fgene.2018.00197
Zhang Y, Mao F, Mu H, Huang M, Bao Y, Wang L, Wong NK, Xiao S, Dai H, Xiang Z, Ma M, Xiong Y, Zhang Z, Zhang L, Song X, Wang F, Mu X, Li J, Ma H, Zhang Y, Zheng H, Simakov O, Yu Z (2021) The genome of Nautilus pompilius illuminates eye evolution and biomineralization. Nat Ecol Evol 5(7):927–938. https://doi.org/10.1038/s41559-021-01571-4
Zhao R, Takeuchi T, Luo YJ, Ishikawa A, Kobayashi T, Koyanagi R, Villar-Briones A, Yamada L, Sawada H, Iwanaga S, Nagai K, Satoh N, Endo K (2018) Dual gene repertoires for larval and adult shells reveal molecules essential for molluscan shell formation. Mol Biol Evol 35(11):2751–2761. https://doi.org/10.1093/molbev/msy172
Zhao R, Takeuchi T, Koyanagi R, Villar-Briones A, Yamada L, Sawada H, Ishikawa A, Iwanaga S, Nagai K, Che Y, Satoh N (2020) Phylogenetic comparisons reveal mosaic histories of larval and adult shell matrix protein deployment in pteriomorph bivalves. Sci Rep 10(1):22140. https://doi.org/10.1038/s41598-020-79330-x
Zhou Y, Yan Y, Yang D, Zheng G, Xie L, Zhang R (2020) Cloning, characterization, and functional analysis of chitinase-like protein 1 in the shell of Pinctada fucata. Acta Biochim Biophys Sin (shanghai) 52(9):954–966. https://doi.org/10.1093/abbs/gmaa076
Zhu Y, Li Q, Yu H, Liu S, Kong L (2021) Shell biosynthesis and pigmentation as revealed by the expression of tyrosinase and tyrosinase-like protein genes in Pacific oyster (Crassostrea gigas) with different shell colors. Mar Biotechnol 23:777–789. https://doi.org/10.1007/s10126-021-10063-2
Zullo L, Imperadore P (2019) Regeneration and healing. In: Gestal C, Pascual S, Guerra Á, Fiorito G, Vieites JM (eds) Handbook of pathogens and diseases in cephalopods. Springer, pp 193–199. https://doi.org/10.1007/978-3-030-11330-8
Acknowledgements
We would like to thank Dr. Masataka Kusube (National Institute of Technology, Wakayama College) for their support through the KOSEN GEAR 5.0 project. KH and DHES would like to also thank members of Setiamarga Lab. for their constant support during the course of this study. Computations were partially performed on the NIG supercomputer at ROIS National Institute of Genetics, Shizuoka, Japan. We also sincerely thank the two anonymous reviewers for their invaluable suggestions to improve the quality of this manuscript.
Funding
This study was partially supported by Takeda Science Foundation Life Science Research Grants 2022 and Grants-in-aid for Basic Research (KIBAN-C) No. 19K12424, both awarded to DHES, and Grants-in-aid for Basic Research (KIBAN-C) No. 22K06340 awarded to MAY and DHES. KH was supported by KOSEN GEAR 5.0 of National Institute of Technology: Agriculture and Fisheries Project awarded to DHES.
Author information
Authors and Affiliations
Contributions
The study was conceived by DHES. Data analyses were conducted by KH, NT, MS, and DHES. TS, MAY, and DHES supervised the study. DHES wrote the first draft of the manuscript. KH and DHES prepared figures and tables. KH, MAY, and DHES edited subsequent versions of the manuscript. All authors read, edited, and confirmed the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to disclose.
Ethics approval
This study is an in silico study conducted on previously published data. Therefore, no ethical approval was required for this study.
Additional information
Responsible Editor: M. Kuba.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Primary Corresponding Author: Davin H. E. Setiamarga.
Supplementary Information
Below is the link to the electronic supplementary material.
227_2023_4244_MOESM1_ESM.pdf
Supplementary file1 Fig. S1 The number of shared shell matrix proteins (SMPs) among the conchiferans (details in Table S1). Numbers indicated the actual number of sequences deemed as homologs as obtained through reciprocal local BLASTp searches under (E-value score < 1.0e-20). In Fig. 3B–C, the numbers were crunched into “types”: a single type could contain several sequences. Numbers in parentheses denote proteins identified as homologs by the reciprocal local BLASTp searches, but with clearly different domain structures. (PDF 42 KB)
227_2023_4244_MOESM2_ESM.pdf
Supplementary file2 Fig. S2 Common shell matrix proteins (SMPs) among the cephalopods. (A) A Circos diagram depicting sequence homologies among N. pompilius SMPs and other cephalopods (E-value score < 1.0e-20). The link showed the result of three patterns of reciprocal BLASTp searches; N. pompilius vs S. pharaonis; N. pompilius vs S. spirula; and S. pharaonis vs S. pharaonis. Lines in darker color (red, green, black) denote proteins conserved in three species. (B) Number of domains shared among the cephalopod SMPs. (C) The domain structures of three common SMPs shared among the cephalopods. (PDF 312 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hirota, K., Tochino, N., Seto, M. et al. Comparative proteomics of the shell matrix proteins of Nautilus pompilius and the conchiferans provide insights into mollusk shell evolution at the molecular level. Mar Biol 170, 106 (2023). https://doi.org/10.1007/s00227-023-04244-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00227-023-04244-x