Abstract
The search for bacteria-labeling agents that are more efficient and less toxic compared to existing staining dyes is ongoing. Fluorescent quantum dots and carbon dots (CDs) have been extensively researched for various bioimaging applications. Priority is given to CDs due to several advantages, including lower toxicity, versatility in tuning their properties, and better photostability compared to metal-based quantum dots. Although significant progress is still needed to replace existing dyes with CDs for bacteria labeling, they offer promising potential for further improvement in efficiency. Surface charges and functional groups have been reported as decisive factors for bacterial discrimination and live/dead assays; however, a complete guideline for preparing CDs with optimum properties for efficient staining and predicting their labeling performance is lacking. In this review, we discuss the application of fluorescent CDs for bacterial labeling and the underlying mechanisms and principles. We primarily focus on the application and mechanism of CDs for Gram differentiation, live imaging, live/dead bacteria differentiation, bacterial viability testing, biofilm imaging, and the challenges associated with application of CDs. Based on proposed mechanisms of bacterial labeling and ambiguous results reported, we provide our view and guidelines for the researchers in this field to overcome the challenges associated with bacteria labeling using fluorescent CDs.
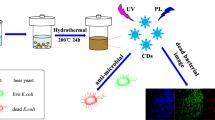
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pathogenic bacteria are classified as Gram-positive and Gram-negative based on the cell surface composition. Gram-positive bacteria are characterized by a thick layer of peptidoglycans without any outer membrane, while Gram-negative bacteria possess a thin peptidoglycan layer and an outer membrane with lipopolysaccharides. Accurate identification of bacterial strains, differentiation, and live/dead differentiation play a crucial role in both theranostics of bacterial infections and the advancement of antibiotic development. Various methods have been employed to detect bacteria, including polymerase chain reaction (PCR), Gram staining, immunological techniques, and Raman spectroscopy [1]. PCR utilizes amplified nucleic acids to identify DNA sequences, immunological techniques involve the interaction between antibodies and antigens, while Raman spectroscopy distinguishes bacteria based on light-scattering differences. However, these approaches are expensive and time-consuming, involve intricate procedures, and may yield false positive results. Fluorescence-based dyes are most widely used for bacterial labeling and they enable real-time monitoring of bacteria with high sensitivity and selectivity, and have high quantum yield [2]. Propidium iodide and propidium monoazide are widely used for lived/dead microbial assay; however, their main limitations are toxicity upon long-time exposure, poor photostability, and high cost. Photodegradation of the dyes imposes constraints on both the duration and time resolution of the experiments. In addition, poor dye penetration and monochromatic properties are unfavorable factors that prompt the search for alternative staining agents. Fluorescent nanoparticles in combination with specific recognition elements, such as aptamer, antibody, bacteriophage, and antimicrobial peptide are used for the labeling of bacteria [3,4,5].
Among various fluorescent nanomaterials, fluorescent carbon dots (CDs) have emerged as a favorable staining agent for bacterial labeling due to their superior biocompatibility and biosafety, ease of preparation, and broad excitation/emission spectral range [6]. They are less toxic when compared to other nanoparticle imaging agents, and are more compliant to in situ bacterial analysis. Their interaction with bacterial cell surfaces allows for precise identification and differentiation of various strains, which has significant implications in medical diagnostics, environmental monitoring, and food safety, where rapid and accurate detection of bacterial pathogens is crucial. The fluorescent properties of CDs are influenced by the molecular precursors used for the preparation of CDs, synthesis methods, surface state, carbon-core and molecular state, surface functionalization, quantum size effect, and conjugate effect [7]. CDs can be tuned to achieve multi-color emission, which enables multi-color labeling of bacteria [8]. In addition, fluorescent CDs have a size in the range of 1.9 to 5 nm, which is very suitable for interaction and internalization into bacterial cells. Antibiotic-modified CDs are also reported for the specific labeling of bacteria [9]. Many reviews are available for the synthesis methods of CDs, structure, origin of fluorescence properties, functional properties, photostability, and toxicity [10,11,12]. A recent review by Lin et al. discussed the application of CDs functionalized with aptamers, antibodies, DNA, and peptides for bacteria sensing [13]. A review article published in 2019 reviewed the application of CDs with a special focus on sensing and killing microorganisms including bacteria, fungi, and viruses [14]. We are of the view that an update of publications since 2019 is important, particularly to have a better understanding of how the surface charges, functional groups, and hydrophilic-hydrophobic properties of CDs influence the labeling mechanism, and to provide an overview of the latest developments, challenges, and future outlook. In this review, we have reviewed the labeling mechanism of fluorescent CDs for their use in Gram differentiation, live imaging, distinguishing live and dead bacteria, assessing bacterial viability, imaging biofilms, and the challenges inherent in the application of CDs. A combination of zeta potential and surface functional groups plays a major role in the selective interaction and labeling of bacteria by the CDs as illustrated in Fig. 1.
Fluorescent carbon dots
The term “carbon dots” has a broad meaning with a variety of carbon nanomaterials coming under this category. Based on the structure, size, and functional properties, CDs are categorized into graphene quantum dots (GQDs), carbon quantum dots (CQDs), carbon nanodots (CNDs), and carbonized polymer dots (CPDs) [15, 16]. Among them, only the CDs that have fluorescence properties can be applied for bacterial imaging. Multi-color CDs can be prepared by controlling their sp3-hybridized carbon cores and sp2-hybridized domains, which enable multi-color bioimaging (Fig. 2A–C) [8]. The emission properties of the CDs can be tuned by using different combinations of precursors and controlling their ratios as illustrated in Fig. 2D [17]. The ultrasmall size and high photostability enable multi-color imaging of Staphylococcus aureus (S. aureus) and Escherichia coli (E. coli) under different excitation bands (Fig. 2E) [18]. Comprehensive reviews on the origin and mechanism of fluorescence of CDs are available [19, 20]. Depending upon the surface charge, functional groups, and pH-dependent behavior, CDs have been widely studied for specific bioimaging of cells and organelles, and our group reported a comprehensive review on this topic with future outlook and challenges [21]. Rather than the fundamental fluorescence characteristics of the CDs, their surface charge and the type of functional groups have been found to be crucial for selective interaction with bacteria for labeling and discrimination. In addition, the size of the functional groups and the quantum yield also play a role in bacteria imaging [22]. Doping of CDs with heteroatoms like boron, nitrogen, phosphorus, and sulfur changes the intrinsic electronic properties of CDs to provide new active sites [7]. Heteroatom doping of CDs enables the selective labeling of dead bacteria rather than live ones, enabling the evaluation of bacterial viability [23]. However, bacteria-derived fluorescent CDs label the dead cells and not the live bacteria [24]. The surface group of CDs may be modified to couple with specific recognition elements such as antibodies, peptides, or aptamers that can stain the bacteria for indirect monitoring [25, 26].
Multi-color emission properties of CDs. A Photographs of dispersions of different CDs, synthesized from o-phenylenediamine and citric acid by hydrothermal method, under daylight (upper) and UV light (bottom). B Emission spectra of CDs dispersion displayed in A under excitation at 365 nm and C time-resolved fluorescence spectra of selected samples. Reproduced from [8] with permission from Wiley‐VCH GmbH. D Schematic representation of synthesis and tuning of multiple color emission CDs based on the precursor ratio by solvothermal method. Reproduced from [17] with permission from Elsevier. E Fluorescence microscopic images of multi-color imaging of S. aureus and E. coli DH5α with surface passivated CDs under different excitation wavelengths: blue filter (UV-2A, 330–380 nm), green filter (B-2A, 450–490 nm), and red filter (G-2A, 510–560 nm). Reproduced from [18] with permission from the American Chemical Society
Gram differentiation of bacteria
Gram-positive bacteria labeling
The cell walls of Gram-positive and Gram-negative bacteria have different compositions and it is mainly exploited for Gram staining. The same principle is also applicable to CDs, which can selectively interact with bacteria enabling Gram differentiation. The bacterial cell walls mainly differ in the thickness of peptidoglycan layers, the presence of outer membranes, and electrical charge. Gram-positive bacteria are more negatively charged compared to Gram-negative bacteria due to the presence of teichoic acid on their thick peptidoglycan layer. Whereas Gram-negative bacteria have a thin peptidoglycan layer and an additional lipopolysaccharide layer. Therefore, positively charged CDs have more affinity toward Gram-positive bacteria than toward Gram-negative bacteria, enabling selective labeling as illustrated in Fig. 3A [27, 28]. CDs have stable fluorescence, which enables them to be used for tracking division and viability assessment of Gram-positive bacteria, for example, lactic acid bacteria [29]. Polarity-sensitive emission properties of the CDs also have been utilized for bacterial differentiation [30]. In this method, CDs having hydrophobic hydrocarbon and quaternary amine functional groups selectively interact with Gram-positive bacteria, resulting in enhanced fluorescence and bacterial differentiation. Not only positively charged CDs, but negatively charged CDs also can stain Gram-positive bacteria. For example, N-doped CDs cross-linked by genipin (N-CDs-GP) synthesized from L-tryptophan and chlorhexidine acetate with a negative zeta potential of −17.3 mV have been reported to stain only Gram-positive S. aureus; however, the interaction between the CDs and the bacteria is not through the negative charge of the CDs, but through the protonated amide functional groups and the negatively charged surface of the bacteria [31]. The study suggests that the zeta potential of the N-CDs-GP has no significant role in interaction with the bacteria. Similarly, Wang et al. reported negatively charged S-doped CDs with a zeta potential of −28 mV that could selectively label Gram-positive bacteria [32]. The mechanism of selectivity has been reported to be due to the presence of sulfate groups on the surface of the CDs. However, further investigation is needed to understand how such highly negatively charged CDs bind with negatively charged cell membranes of live Gram-positive bacteria and why they could not bind with less negatively charged Gram-negative bacteria. Some CDs have been reported to stain both Gram-positive and Gram-negative bacteria by difference in their excitation emission [33, 34]; however, the mechanism of interaction between CDs and bacteria has not been clearly explained. CDs having very mild positive charge (zeta potential = +0.32 mV) and having plenty of amine groups have been reported to distinguish drug-resistant S. aureus and normal S. aureus [35]. CDs synthesized from cyanine 7 dyes (Cy7-CH3) and passivated with 3-hydroxytyramine (CyCDs) stain only the cell wall of the methicillin-resistant S. aureus (MRSA), whereas, it stains the whole cells of normal S. aureus (Fig. 3B). The mechanism of distinction between the drug-resistant and normal strain is due to the higher thickness of the cell wall of the former, which rapidly excretes the CyCDs by the efflux pump, causing illumination only at the cell wall. The report reveals that the imaging effect is the same for drug-resistant E. coli and its normal strain.
Gram differentiation of bacteria. A Schematic representation of interaction between positively charged CDs and the negatively charged peptidoglycans of the cell wall of Gram-positive bacteria. Reproduced from [28] with permission from Elsevier; B rapid labeling and identification drug-resistant bacteria, MRSA and normal strain, S. aureus (SA) using CyCDs. Reproduced from [35] with permission from Elsevier; C confocal fluorescence images of three Gram-positive and three Gram-negative bacteria treated with quaternized CDs showing contact-enhanced emission toward Gram-positive strains. Reproduced from [36] with permission from Elsevier
Other than preferential interaction based on charges, bacterial contact-enhanced fluorescence emission has been reported as an efficient method for the distinction of bacteria by CDs. In this mechanism, the fluorescence of the CDs is enhanced upon interaction with the Gram-positive bacteria (Fig. 3C) [36]. However, the mechanism of interaction is via both electrostatic and hydrophobic between the positively charged quaternized CDs (zeta potential = +33 mV) and the negatively charged bacteria, but the exact mechanism of the contact-enhanced fluorescence emission of the CDs has not been explained. The specificity of the CDs toward a particular bacteria can be enhanced by their surface modification with a specific molecule such as vancomycin. CDs modified with vancomycin interact with the cell walls of Gram-positive bacteria through ligand-receptor interactions, leading to detection down to 9.40 × 104 CFU mL−1 [9]. Vancomycin on the CDs forms hydrogen bonding with the terminal peptide of D-Ala-D-Ala on the cell wall enabling the detection of S. aureus with high specificity.
Some CDs with a zeta potential of −14 mV and having plenty of surface functional groups such as carboxyl, amino, and hydroxyl have been reported to exhibit antibacterial properties only under visible light irradiation (for 60 min) via reactive oxygen species generation [37]. Such CDs show selective labeling of Gram-positive bacteria such as S. aureus and L. monocytogenes, and not Gram-negative, enabling Gram differentiation and multi-color imaging.
Gram-negative bacteria labeling
Conventional Gram staining agents label only Gram-positive bacteria by exploiting differences in their cell wall composition. Conversely, CDs have been reported to stain both Gram-positive and Gram-negative bacteria, which provides an advantage for investigating both types of stains. Staining Gram-negative bacteria solely based on differences in electrostatic interaction is more challenging compared to Gram-positive bacteria. This is attributed to their thin glucan layer and strong outer membrane with higher lipid content. Therefore, selective interaction of CDs with the outer membrane of the Gram-negative bacteria is important. CDs having a high negative zeta potential of −21.6 mV and −31.2 mV have been reported to stain the Gram-negative bacteria, E. coli by simple incubation [38, 39]. The mechanism of labeling has been reported to be the interaction between the carboxylic acid groups on the CDs with the peptides, proteins, and amino acids on the bacterial surface. This mechanism is exactly the reverse of Gram-positive staining, in terms of charges and functional groups. Certain sugars such as mannose have affinity toward the pili of certain bacteria such as wild-type E. coli, opening another method of improving selectivity [40, 41]. In our previous reports, CQDs synthesized from ammonium citrate by a solid-state heating method and functionalized with mannose ligands (Man–CQDs) were found to exhibit specificity toward E. coli, enabling efficient bacteria labeling [40, 41]. These two studies revealed that mannose ligands on the Man-CQDs exhibit multivalent interaction with the FimH lectin units in wild-type pili on the surface of E. coli. Another study also reported a similar interaction for the selective sensing of E. coli, where the authors utilized single-use plastics to prepare biocompatible and highly fluorescent CDs [42]. The intensity of fluorescence from the plastic-derived CDs decreased with an increase in the concentration of bacteria. This decrease was attributed to the association of bacteria with the surface functional groups of CDs, which affected the rate of radiative recombination of photo-generated electrons and holes in the CQDs, thereby enabling the detection of bacteria. This method is primarily suitable for bacterial quantification rather than bacterial imaging.
While most of the CDs used for live imaging of bacteria emit in the short wavelength (<600 nm) region, they may induce photodamage to tissues [43]. Excitation of CDs at a longer wavelength (near-infrared) region reduces the phototoxicity and avoids self-fluorescence by the target tissues [44]. The report demonstrates the conversion of cyanine dye, Cy-COOH, to Cy7-CDs, which retain the optical properties of the dye while exhibiting improved water solubility and quantum yield. The introduction of abundant hydroxyl, carboxyl, and benzene-rich functional groups on the surface enables bacterial identification and quick monitoring of bacterial viability through fluorescence imaging. Specifically, Cy7-CDs do not interact quickly with Gram-positive bacteria, however get reduced upon incubation for 5 min and the fluorescence intensity was enhanced within 40 min of co-incubation, which allows for the discriminative imaging and assessment of the survival rate of bacteria.
As mentioned above, negatively charged CDs can easily stain Gram-negative bacteria, and if they possess antibacterial effects, they may also stain Gram-positive bacteria after a longer incubation period. For example, N and F co-doped CDs with zeta potential −27.7 mV stain Gram-negative E. coli within 20 min, whereas they stain Gram-positive S. aureus in 60 min [45]. However, despite being negatively charged, they exhibit bactericidal effects via the downregulation of gene expressions related to energy uptake and metabolism. Therefore, the incubation time is an important factor for Gram differentiation.
Since CDs do not have species-specific affinity toward different bacterial species, they can be modified with different receptors, which will enable them to bind with all species with different affinity, allowing for quantification of different species by linear discrimination analysis. For example, Zheng et al. fabricated a multiplex assay with fluorescent CDs modified with different receptors, boronic acid, polymyxin, or vancomycin, separately [46]. These three differently functionalized CDs on the sensor array will interact with all bacteria with different affinities. The difference in the affinity toward different species is then evaluated by linear discrimination analysis. Recently, Li et al. reported a similar assay using linear discrimination analysis to distinguish six different bacteria species employing multi-color-emissive CDs at different pH [47]. The authors reported that the assay is rapid and 100% accurate. Amphiphilic CDs having hydrocarbon chain functional groups have been reported to label and distinguish different bacteria species based on the shift in the emission wavelength upon interaction with bacteria [48]. The report suggests that the amphiphilic CDs have different affinity with different species of bacteria, and the interactions lead to species-specific shifts in the excitation-dependent emission wavelengths, visualizing different species under different emission wavelengths. Since the CDs are biocompatible, they enable live imaging to monitor bacterial cell division.
Change in the fluorescence lifetime (FLT) of the CDs upon interaction with Gram-negative bacteria has been reported as a revolutionary mechanism for their selective detection of E. coli [49]. The work reports that when colistin-passivated CDs interact with E. coli, the average FLT (τavg) significantly decreases from 3.91 to 1.59 ns, and the bacterial concentration can be quantified by fluorescence lifetime imaging microscopy and automated confocal laser scanning microscopy with a limit of detection of 3.68–4.89 × 104 CFU mL−1. The selectivity mechanism is the interaction of the colistin having plenty of cationic sites, which replace Ca2+ and Mg2+ ions and integrate with the lipopolysaccharides of the outer membrane of the Gram-negative bacteria. Another mechanism for the identification of bacteria is by targeting the sugar-metabolism-triggered change in the fluorescence of pH-sensitive CDs. Different bacteria have different capabilities for sugar metabolism and their acidic by-products induce different pH to the solution. Thus, the fluorescence of the pH-dependent CDs differs in different bacteria solutions containing sugar such as glucose or lactose; for example, E. coli and S. aureus, enabling their identification and detection as low as 21 and 33 CFU mL−1, respectively [50].
Biofilm imaging
Selective labeling of biofilm is also as important as bacteria labeling and plays a major role in the fundamental understanding of bacterial activities and interactions. Bacterial biofilms are complex communities of bacteria in a self-produced matrix of exopolysaccharide (EPS) composed of proteins, polysaccharides, and extracellular DNA. Biofilms are protective barriers against environmental stress and about 80% of bacterial infections are associated with biofilm formation, leading to life-threatening conditions. Imaging of biofilm and associated areas is often challenging due to the heterogenicity and complexity of biofilm structure [51]. The dense and amphiphilic nature of the EPS matrix restricts the penetration of the imaging agents. Fluorescence microscopy, coupled with functionalized probes, enables the monitoring of biofilms to provide fundamental information about their structure and behavior. Current approaches for analyzing and visualizing EPS in bacterial biofilms commonly involve the use of fluorescent dyes covalently linked to carbohydrate recognition elements, especially lectins, typically carried out through confocal laser scanning microscopy. However, protected by a resilient and sticky EPS matrix, microorganisms embedded in biofilms pose challenges to the penetration of fluorescence dyes for staining. Biofilms not only impose stringent restrictions on dye penetration but also encompass areas with widely varying environmental conditions that can impact the function of dyes [52]. In addition, the dyes are often associated with limitations, such as susceptibility to biofilm damage, short shelf lives, high costs, demanding storage requirements (e.g., low temperature and protection from light), and photobleaching, making them less favorable options. Alternative techniques employ surface-modified semiconductor quantum dots, utilizing biological or synthetic complex substances that are toxic to bacterial cells, thereby hindering in situ analysis and potentially disrupting assembled biofilms. Other methodologies used for biofilm analysis, such as atomic force microscopy (AFM), scanning electron microscopy (SEM), magnetic resonance imaging (MRI), and Raman spectroscopy, are not universally applicable in situ, and therefore, necessitate sample modulation, and may provide only partial structural details in many cases [53].
Cationic fluorescent CDs with a high positive charge surface (+33.1 mV) and ultrasmall size (3.3 nm) facilitate the penetration of the CDs in the bacterial biofilms allowing for fluorescent imaging [51]. Highly positively charged CDs at low concentrations allow for the imaging of biofilm, at medium concentrations they inhibit the biofilm, and when the concentration of CDs is very high the biofilm is eradicated. Furthermore, the excitation-dependent emission properties of fluorescent CDs can enable multi-color fluorescence imaging of the biofilms. CDs as fluorescent probes have the potential to be used for selectively imaging Gram-positive and Gram-negative bacterial biofilms [54].
The bacterial biofilm imaging provides information on the life-cycle of biofilm such as growth, reproduction, and formation of mature biofilm [52, 55]. CDs synthesized from L. plantarum have been reported for the imaging of biofilm-encased E. coli at different stages of biofilm (24–120 h) [52] thereby enabling the understanding of the morphology and physiological state of bacteria in a biofilm. Conversely, amphiphilic CDs bind to the hydrophobic regions in the EPS matrix allowing for the imaging of biofilm [6]. In addition to imaging of biofilm, amphiphilic CDs are also employed for monitoring the effect of external factors such as temperature on the kinetics of EPS growth. Due to the difficulty in staining biofilms, CDs are not well researched for biofilm imaging, and therefore, further investigation is needed for improving the biofilm penetration properties and developing new types of CDs, with less toxicity. A high positive charge on the CDs is highly toxic against bacteria, and is mostly applied for antibacterial and antibiofilm activities, rather than biofilm imaging.
Live bacteria imaging
Biocompatibility of the CDs is crucial for the bioimaging of live bacteria. Although positively charged CDs are easily adsorbed on bacterial membranes, a high positive charge on the CDs could damage the negatively charged bacterial cell membrane by disrupting its structural integrity via strong electrostatic interaction [56]. Yan et al. demonstrated negatively charged peptidoglycan-targeting CDs have low toxicity and are suitable for live bacteria bioimaging as long as 24 h with one-step staining [57]. In their report, triple excitation wavelengths and single-color emission carbon quantum dots (T-SCQDs) synthesized from glucose, glycine, and L-tryptophan possess a net negative charge with a zeta potential of −12.37 mV and plenty of amino functional groups. Although the T-SCQDs carry a net negative charge, the authors conducted a competitive binding assay to prove that the cationic amino groups on them easily bind to the comparatively more negatively charged peptidoglycans (zeta potential = −7.70 mV) of the Gram-positive bacteria, than that with less negative lipopolysaccharides (zeta potential = −3.84 mV) in Gram-negative bacteria, enabling a clear differentiation (Fig. 4A, B). Peptidoglycans also have hydrophobic regions due to the presence of N-acetylglucosamine and N-acetylmuramic acid, allowing for hydrophobic interactions with benzopyrrole regions of the T-SCDs. In addition, the report proves that the T-SCDs can specifically stain S. aureus colonies in a mixture of S. aureus and E. coli colonies. The efficiency of the live bacteria imaging by CDs is such high that they can be used to track division and also viability assessment in bacteria (Fig. 4C) [29].
A Graphical illustration of the synthesis of T-SCQDs from glucose, L-tryptophan, and glycine, and three wavelength excitation and single emission of the T-SCQDs and B principle of selective live imaging of Gram-positive bacteria by T-SCQDs. Reproduced from [57] with permission from the American Chemical Society. C Tracking of cell division in Lactobacillus plantarum using phosphorus and nitrogen-doped CDs at different excitation and emission wavelengths. Reproduced from [29] with permission from Elsevier
Nanosized (1–3 nm), N-doped oxygenated fluorescent CDs with crystalline characteristics prepared from a colloidal system with deep eutectic solvent were reported for labeling both Gram-positive and Gram-negative bacteria [58]. The colloidal CDs were reported to have electrostatic interaction with the bacteria and readily internalized into the cells exhibiting good fluorescence and displaying diverse light emission ranging from bluish to red, depending on the excitation wavelength, accompanied by an exceptionally high quantum yield of approximately 82%. A detailed investigation is needed to understand the interaction of the colloidal CDs with both Gram strains of bacteria.
Live/dead bacteria differentiation
Live/dead bacteria differentiation with fluorescent CDs is achieved due to the difference in the cell membrane transportation of live and dead bacteria caused by the high negative charge of the CDs. Reports demonstrated that only highly negatively charged CDs can selectively label dead bacteria [59] irrespective of their Gram strain, and the differentiation efficiency is lost if the zeta potential of the CDs is decreased [23, 24, 60, 61]. Specific labeling of dead bacteria has been reported by Song et al. with nitrogen, phosphorus, and sulfur co-doped CDs (NPSCDs) [23] to specifically stain dead bacteria (Bacillus aryabhattai) and they do not stain the live bacteria due to their high negative zeta potential (−41.9 mV) and the electrostatic repulsion of the negative charged bacterial cell wall. The report suggests that the NPSCDs enter dead bacteria cells because the cell loses control of the action of carrier proteins, enabling bacterial viability evaluation. Functionalizing CDs with specific molecules such as ampicillin and having moderate negative charge (zeta potential = −12.5 mV) also can target the membrane of dead bacteria only, enabling live/dead differentiation [62]. The selectivity toward dead bacteria or yeast staining is due to the electrostatic repulsion of the live microbes (E. coli, S. aureus, and C. podzolicus) having high negative surface charge (zeta potential = −43.63, −30.5, and −36.4 mV, respectively). A similar phenomenon was observed for bacteria-derived CDs (zeta potential = −23.30 mV) with excitation-dependent emission characteristics [63]. The entry of small molecular substances into the cells necessitates ion channels or transmembrane proteins and hence these CDs face limitations in labeling live microbial cells. In the case of dead cells, the destruction of cell walls or membranes results in reduced selective permeability. This weakening of electrostatic interactions between bacteria and the CDs enables the successful labeling of dead cells. Ultrasmall-sized CDs (1.91 nm, zeta potential = −15 mV) with carboxyl moieties on the surface penetrate the dead cells, avoiding the live ones of both Gram-positive and Gram-negative bacteria as illustrated in Fig. 5A [64]. Ultrasmall sulfur-doped CDs synthesized from rose bengal and 1,4-dimercaptobenzene by hydrothermal method have been reported to distinguish dead cells from live cells [65]. The selective staining mechanism has been determined to be the difference in the uptake pathway of the dead and live cells, in which the dead cells permit the uptake of the ultrasmall CDs (1.6 nm) by passive diffusion followed by interaction with DNA and RNA. Whereas the CDs could not penetrate into the live cells. Other than the live/dead differentiation mechanism based on strong electrostatic repulsion, difference in the affinity of the CDs with the cell wall of live bacteria and entry into the cell of dead bacteria can also be employed. For example, P- and N-co-doped CDs (PN-CDs) with a positive zeta potential of +2.34 mV have been reported for live/dead differentiation, in which they stain both live and dead bacteria with a difference [29]. In this method the PN-CDs label only the cell wall of live bacteria within 1 min, enabling monitoring bacterial viability and also tracking division. Whereas they light up the whole cell of the dead bacteria, by entering the cell, enabling live/dead differentiation. Reports suggest that CDs label dead bacteria irrespective of their method of killing, such as heating, ethanol, formaldehyde, and microwave, and irrespective of Gram strain (Fig. 5B, C) [29, 61], and hence this method can be employed in a wide range of antibacterial studies. A recent work by Liu et al., demonstrated that the nature of the functional groups, their size, and quantum yield influence the imaging of bacteria [22]. CDs prepared from spermine could stain the dead bacteria, and showed a quantum yield of 66.46% along with high fluorescent bleaching resistance (even 3 h irradiation could maintain 70% fluorescence). Modifying the CDs with ethylenediamine imparted primary amine groups on the surface of CDs, which enabled the staining of both dead and live bacteria. The authors reported that the primary amine group played a significant role in interacting with the bacteria. Enriching the surface of CDs with abundant functional groups enables the shift in excitation wavelength to a longer wavelength region (640 nm), which minimizes the fluorescence interference and simultaneously distinguishes live-dead bacteria in the same channel (680–760 nm) [44].
A Principle of live/dead bacteria imaging with CDs: strong electrostatic repulsion between the highly negatively charged CDs and negatively charged bacterial cell walls. Reproduced from [64] with permission from Elsevier; B confocal fluorescence microscopic images of Lactobacillus plantarum killed by different methods and stained by CDs Reproduced from [29] with permission from Elsevier; C fluorescence images of live/dead bacteria differentiation by CDs irrespective of Gram stain. Reproduced from [61] with permission from Elsevier
Viability assessment
Bacterial viability assessment is performed as a part of laboratory research and also in bacterial monitoring in the fields of clinical, environmental, food, and drug development. Fluorescence staining followed by microscopy or flow cytometry analysis is widely used for viability assessment. The limitations of the fluorescent dyes mentioned earlier are also applicable here and hence, CDs could be a viable alternative. However, the biocompatibility of the CDs is essential for viability assessment. CDs with no antibacterial properties and can specifically stain either live or dead bacteria in a mixture of live and dead bacteria can be used for bacterial viability assessment. Therefore, many of the CDs discussed in the above sections for Gram-positive, Gram-negative, live imaging, and live/dead imaging are efficient in bacterial viability assessment [23, 29, 44]. Some CDs have been reported to label live bacterial cell walls quickly, allowing for viability assessment within 1 min [29]. A higher incubation time will cause the CDs to enter the cells of dead bacteria and light up the whole cell, thus differentiating live and dead bacteria.
Before the application of CDs for viability assessment, their antibacterial activity assessment is important. For example, exopolysaccharide-derived CDs with a zeta potential of −24 mV, with no toxicity toward microbes even at concentrations of 3 mg mL−1, enable multi-color imaging capabilities for microbial viability assessment including Gram-negative bacteria, Gram-positive bacteria, and fungus [66]. Some reports on CDs for bacterial viability assessment performed cytotoxicity assessment in the HeLa cell line [67]; however, additionally, assessing the antibacterial capability of the CDs would be more accurate in predicting their appropriateness. Concentration- and time-dependent toxicity against bacteria must be determined before applying CDs for viability assessment and tracking division.
Challenges, guidelines, and future prospects
Improvements are needed for bacterial labeling with CDs, especially in balancing the mechanism of selectivity and antibacterial effect. The zeta potential and types of functional groups on the CDs are important for Gram and live/dead differentiation of bacteria. For example, positive zeta potential is suitable for selective labeling of Gram-positive bacteria due to electrostatic interaction. However, a high positive charge could lead to an antibacterial effect by disrupting membrane function and will be problematic for live imaging or tracking division. Conversely, a very low zeta potential close to zero could result in poor interaction with bacteria. In addition to charge, the amount of amino functional groups on the CDs facilitates attachment with the peptidoglycans of bacteria. Therefore, CDs having mild positive charges below the toxicity threshold and having sufficient amino groups would be suitable for Gram staining. Even CDs with net negative charge and plenty of amino groups can stain Gram-positive bacteria, due to the interaction between amino groups and teichoic acid. Therefore, it can be concluded that the presence of amino groups on the CDs is important for Gram staining and live imaging, and the zeta potential is less significant [31]. Future research on the bacteria labeling with CDs having a net negative charge and containing amino functional groups must include quantification of the amino groups. For a better understanding of the influence of zeta potential, functional groups, types of CDs, and carbon source for CD synthesis on the bacterial labeling mechanism, summarized information is provided in Table 1.
Some negatively charged CDs (N- and F-co-doped CDs) can be used for Gram differentiation due to their difference in duration for staining [45]. They do not exert antibacterial activity via electrostatic interaction; however, they downregulate certain important genes involved in cell division and survival, causing antibacterial effects. Hence, all CDs used for bacterial labeling in future research must be investigated for various antibacterial mechanisms, even though they possess a net negative charge. CDs with negative zeta potential and plenty of carboxyl groups have been reported as suitable staining agents for Gram-negative bacteria. CDs with a high negative zeta potential in the order of > −40 mV will experience a repulsion from the negative charge from the live bacteria, and can stain only dead bacteria, which is utilized as a differentiation mechanism for live/dead assay.
Although charge is important for bacterial labeling, future research should also focus on quantifying cationic and hydrophobic functional groups present on the CDs to better predict bioimaging efficiency and mechanisms. CDs for biofilm imaging have been rarely reported, although there are some reports on cationic and hydrophilic CDs. Highly penetrating CDs may disrupt the biofilm in a concentration-dependent manner; therefore, maintaining a low concentration of CDs is essential in biofilm imaging. More in-depth studies are needed in this regard to develop CDs for biofilm imaging. Some reports indicate that CDs can stain both Gram-positive and Gram-negative bacteria; however, the underlying mechanism remains unestablished [68]. It is advisable to conduct a thorough investigation in all research related to bacterial labeling with CDs to understand the exact mechanism by which such CDs can stain both strains.
Conclusions
Fluorescent CDs have promising potential for bacterial labeling, including quantification by fluorescence measurement or by microscopy analysis. Bacterial labeling with CDs still needs further advancement compared to the bioimaging of cells, organelles, and tissues with fluorescent CDs. Owing to their multi-color emission properties, CDs have a high potential for the biolabeling of bacteria. The main limitation of CDs for bacterial labeling is that the selectivity mechanism is by tuning surface charge and functional groups, which often gives contradictory results due to the imbalance in the charge and functional groups. CDs with mild positive charge or a net negative charge and having cationic amino groups have been reported to be suitable for bioimaging of live Gram-positive bacteria. In addition to charge, the presence of functional groups, such as amino, benzopyrrole, hydrocarbon chain, and hydroxyl, is important to specifically bind to the corresponding sites in the bacterial cell membrane. Therefore, quantification of functional groups on the CDs is essential for all future studies. CDs with antibacterial properties have been observed to label bacteria easily, due to strong interaction and internalization. However, such antibacterial CDs cannot be used for live bacteria imaging and bacterial viability assessment, as they kill the bacteria and label the dead bacteria. CDs having the capacity to label either live or dead bacteria selectively can be employed for bacterial viability assessment. Some reports do not provide the zeta potential values of CDs or the mechanism of interaction between CDs and bacteria, which leaves the reader with incomplete information. Therefore, precursor for CD synthesis, type of CDs, their zeta potential values, surface functional groups and quantity, size, toxicity toward bacteria, and a thorough investigation of the mechanism of labeling must be included in all future work.
References
Franco-Duarte R, Černáková L, Kadam S, Kaushik KS, Salehi B, Bevilacqua A, Corbo MR, Antolak H, Dybka-Stępień K, Leszczewicz M, Relison TS, de Souza AVC, Sharifi-Rad J, Coutinho HDM, Martins N, Rodrigues CF. Advances in chemical and biological methods to identify microorganisms-from past to present. Microorganisms. 2019;7:130. https://doi.org/10.3390/microorganisms7050130.
Yoon SA, Park SY, Cha Y, Gopala L, Lee MH. Strategies of detecting bacteria using fluorescence-based dyes. Front Chem. 2021;9:743923. https://doi.org/10.3389/fchem.2021.743923.
Shen H, Wang J, Liu H, Li Z, Jiang F, Wang F-B, Yuan Q. Rapid and selective detection of pathogenic bacteria in bloodstream infections with aptamer-based recognition. ACS Appl Mater Interfaces. 2016;8:19371–8. https://doi.org/10.1021/acsami.6b06671.
Rotem Edgar MM, Jeeseong Hwang, Amos B. Oppenheim, Richard A. Fekete, Gary Giulian, Carl Merril, Kunio Nagashima, Sankar Adhya. High-sensitivity bacterial detection using biotin-tagged phage and quantum-dot nanocomplexes. Appl Biol Sci. 2006;103:4841–5. https://doi.org/10.1073/pnas.0601211103.
He X, Yang Y, Guo Y, Lu S, Du Y, Li J-J, Zhang X, Leung NLC, Zhao Z, Niu G, Yang S, Weng Z, Kwok RTK, Lam JWY, Xie G, Tang BZ. Phage-guided targeting, discriminative imaging, and synergistic killing of bacteria by AIE bioconjugates. J Am Chem Soc. 2020;142:3959–69. https://doi.org/10.1021/jacs.9b12936.
Ritenberg M, Nandi S, Kolusheva S, Dandela R, Meijler MM, Jelinek R. Imaging Pseudomonas aeruginosa biofilm extracellular polymer scaffolds with amphiphilic carbon dots. ACS Chem Biol. 2016;11:1265–70. https://doi.org/10.1021/acschembio.5b01000.
Cui F, Ye Y, Ping J, Sun X. Carbon dots: current advances in pathogenic bacteria monitoring and prospect applications. Biosens Bioelectron. 2020;156:112085. https://doi.org/10.1016/j.bios.2020.112085.
Wang B, Yu J, Sui L, Zhu S, Tang Z, Yang B, Lu S. Rational design of multi-color-emissive carbon dots in a single reaction system by hydrothermal. Adv Sci. 2021;8:2001453. https://doi.org/10.1002/advs.202001453.
Zhong D, Zhuo Y, Feng Y, Yang X. Employing carbon dots modified with vancomycin for assaying Gram-positive bacteria like Staphylococcus aureus. Biosens Bioelectron. 2015;74:546–53. https://doi.org/10.1016/j.bios.2015.07.015.
Ozyurt D, Kobaisi MA, Hocking RK, Fox B. Properties, synthesis, and applications of carbon dots: a review. Carbon Trends. 2023;12:100276. https://doi.org/10.1016/j.cartre.2023.100276.
Cui L, Ren X, Sun M, Liu H, Xia L. Carbon dots: synthesis, properties and applications. Nanomaterials (Basel). 2021;11:3419. https://doi.org/10.3390/nano11123419.
Wang Y, Li X, Zhao S, Wang B, Song X, Xiao J, Lan M. Synthesis strategies, luminescence mechanisms, and biomedical applications of near-infrared fluorescent carbon dots. Coord Chem Rev. 2022;470:214703. https://doi.org/10.1016/j.ccr.2022.214703.
Lin L, Fang M, Liu W, Zheng M, Lin R. Recent advances and perspectives of functionalized carbon dots in bacteria sensing. Microchim Acta. 2023;190:363. https://doi.org/10.1007/s00604-023-05938-1.
Lin F, Bao Y-W, Wu F-G. Carbon dots for sensing and killing microorganisms. C. 2019;5:33. https://doi.org/10.3390/c5020033.
Xu W, Zeng F, Han Q, Peng Z. Recent advancements of solid-state emissive carbon dots: a review. Coord Chem Rev. 2024;498:215469. https://doi.org/10.1016/j.ccr.2023.215469.
Liu J, Li R, Yang B. Carbon dots: a new type of carbon-based nanomaterial with wide applications. ACS Cent Sci. 2020;6:2179–95. https://doi.org/10.1021/acscentsci.0c01306.
Wang C, Huang J, He Y, Ran G, Song Q. Preparation of multicolor carbon dots with thermally turn-on fluorescence for multidimensional information encryption. Chin Chem Lett. 2024;35:108420. https://doi.org/10.1016/j.cclet.2023.108420.
Pal T, Mohiyuddin S, Packirisamy G. Facile and green synthesis of multicolor fluorescence carbon dots from curcumin: in vitro and in vivo bioimaging and other applications. ACS Omega. 2018;3:831–43. https://doi.org/10.1021/acsomega.7b01323.
Zhu P, Tan K, Chen Q, Xiong J, Gao L. Origins of efficient multiemission luminescence in carbon dots. Chem Mater. 2019;31:4732–42. https://doi.org/10.1021/acs.chemmater.9b00870.
Alafeef M, Srivastava I, Aditya T, Pan D. Carbon dots: from synthesis to unraveling the fluorescence mechanism. Small. 2024;20:2303937. https://doi.org/10.1002/smll.202303937.
Unnikrishnan B, Wu R-S, Wei S-C, Huang C-C, Chang H-T. Fluorescent carbon dots for selective labeling of subcellular organelles. ACS Omega. 2020;5:11248–61. https://doi.org/10.1021/acsomega.9b04301.
Liu Y, Zhong D, Yu L, Shi Y, Xu Y. Primary amine functionalized carbon dots for dead and alive bacterial imaging. Nanomaterials. 2023;13:437. https://doi.org/10.3390/nano13030437.
Song Y, Li H, Lu F, Wang H, Zhang M, Yang J, Huang J, Huang H, Liu Y, Kang Z. Fluorescent carbon dots with highly negative charges as a sensitive probe for real-time monitoring of bacterial viability. J Mater Chem B. 2017;5:6008–15. https://doi.org/10.1039/C7TB01092C.
Hua X-W, Bao Y-W, Wang H-Y, Chen Z, Wu F-G. Bacteria-derived fluorescent carbon dots for microbial live/dead differentiation. Nanoscale. 2017;9:2150–61. https://doi.org/10.1039/C6NR06558A.
Yang L, Deng W, Cheng C, Tan Y, Xie Q, Yao S. Fluorescent immunoassay for the detection of pathogenic bacteria at the single-cell level using carbon dots-encapsulated breakable organosilica nanocapsule as labels. ACS Appl Mater Interfaces. 2018;10:3441–8. https://doi.org/10.1021/acsami.7b18714.
Gao R, Zhong Z, Gao X, Jia L. Graphene oxide quantum dots assisted construction of fluorescent aptasensor for rapid detection of Pseudomonas aeruginosa in food samples. J Agric Food Chem. 2018;66:10898–905. https://doi.org/10.1021/acs.jafc.8b02164.
Liu S, Quan T, Yang L, Deng L, Kang X, Gao M, Xia Z, Li X, Gao D. N, Cl-codoped carbon dots from Impatiens balsamina L. Stems and a deep eutectic solvent and their applications for Gram-positive bacteria identification, antibacterial activity, cell imaging, and clo– sensing. ACS Omega. 2021;6:29022–36. https://doi.org/10.1021/acsomega.1c04078.
Sumaiyah, Hasibuan PAZ, Tanjung M, Lianto W, Gea S, Piliang A, Situmorang SA. Hydrothermally nitrogen-doped carbon dots (N-C-dots) from isolated lignin of oil palm empty fruit bunch for bacterial imaging of Staphylococcus aureus. Case Stud Chem Environ Eng. 2023;8:100455. https://doi.org/10.1016/j.cscee.2023.100455.
Fu T, Wan Y, Jin F, Liu B, Wang J, Yin X, Fu X, Tian B, Feng Z. Efficient imaging based on P- and N-codoped carbon dots for tracking division and viability assessment of lactic acid bacteria. Colloids Surf B Biointerfaces. 2023;223:113155. https://doi.org/10.1016/j.colsurfb.2023.113155.
Yang J, Zhang X, Ma Y-H, Gao G, Chen X, Jia H-R, Li Y-H, Chen Z, Wu F-G. Carbon dot-based platform for simultaneous bacterial distinguishment and antibacterial applications. ACS Appl Mater Interfaces. 2016;8:32170–81. https://doi.org/10.1021/acsami.6b10398.
Chu X, Wu F, Sun B, Zhang M, Song S, Zhang P, Wang Y, Zhang Q, Zhou N, Shen J. Genipin cross-linked carbon dots for antimicrobial, bioimaging and bacterial discrimination. Colloids Surf B Biointerfaces. 2020;190:110930. https://doi.org/10.1016/j.colsurfb.2020.110930.
Wang Z, Xu K-F, Wang G, Durrani S, Lin F, Wu F-G. “One stone, five birds”: ultrabright and multifaceted carbon dots for precise cell imaging and glutathione detection. Chem Eng J. 2023;457:140997. https://doi.org/10.1016/j.cej.2022.140997.
Baig MMF, Chen Y-C. Bright carbon dots as fluorescence sensing agents for bacteria and curcumin. J Colloid Interface Sci. 2017;501:341–9. https://doi.org/10.1016/j.jcis.2017.04.045.
Prakash A, Yadav S, Tiwari P, Saxena P, Srivastava A, Tilak R. Facile synthesis of highly fluorescent nitrogen-doped carbon quantum dots and their role in bioimaging of some pathogenic microorganisms. J Nanopart Res. 2023;25. https://doi.org/10.1007/s11051-023-05893-1.
Liu W, Wu B, Sun W, Liu W, Gu H, Du J, Fan J, Peng X. Near-infrared II fluorescent carbon dots for differential imaging of drug-resistant bacteria and dynamic monitoring of immune system defense against bacterial infection in vivo. Chem Eng J. 2023;471:144530. https://doi.org/10.1016/j.cej.2023.144530.
Yang J, Gao G, Zhang X, Ma Y-H, Chen X, Wu F-G. One-step synthesis of carbon dots with bacterial contact-enhanced fluorescence emission: fast gram-type identification and selective Gram-positive bacterial inactivation. Carbon. 2019;146:827–39. https://doi.org/10.1016/j.carbon.2019.02.040.
Gao Z, Yang D, Wan Y, Yang Y. One-step synthesis of carbon dots for selective bacterial inactivation and bacterial differentiation. Anal Bioanal Chem. 2020;412:871–80. https://doi.org/10.1007/s00216-019-02293-0.
Das P, Bose M, Ganguly S, Mondal S, Das AK, Banerjee S, Das NC. Green approach to photoluminescent carbon dots for imaging of Gram-negative bacteria Escherichia coli. Nanotechnology. 2017;28:195501. https://doi.org/10.1088/1361-6528/aa6714.
Yuan M, Zhong R, Gao H, Li W, Yun X, Liu J, Zhao X, Zhao G, Zhang F. One-step, green, and economic synthesis of water-soluble photoluminescent carbon dots by hydrothermal treatment of wheat straw, and their bio-applications in labeling, imaging, and sensing. Appl Surf Sci. 2015;355:1136–44. https://doi.org/10.1016/j.apsusc.2015.07.095.
Weng CI, Chang HT, Lin CH, Shen YW, Unnikrishnan B, Li YJ, Huang CC. One-step synthesis of biofunctional carbon quantum dots for bacterial labeling. Biosens Bioelectron. 2015;68:1–6. https://doi.org/10.1016/j.bios.2014.12.028.
Lai IP-J, Harroun SG, Chen S-Y, Unnikrishnan B, Li Y-J, Huang C-C. Solid-state synthesis of self-functional carbon quantum dots for detection of bacteria and tumor cells. Sens Actuators B Chem. 2016;228:465–70. https://doi.org/10.1016/j.snb.2016.01.062.
Kumari M, Chaudhary S. Modulating the physicochemical and biological properties of carbon dots synthesised from plastic waste for effective sensing of E. coli. Colloids Surf B. 2020;196:111333. https://doi.org/10.1016/j.colsurfb.2020.111333.
Nie X, Jiang C, Wu S, Chen W, Lv P, Wang Q, Liu J, Narh C, Cao X, Ghiladi RA, Wei Q. Carbon quantum dots: a bright future as photosensitizers for in vitro antibacterial photodynamic inactivation. J Photochem Photobiol B Biol. 2020;206:111864. https://doi.org/10.1016/j.jphotobiol.2020.111864.
Liu W, Gu H, Liu W, Lv C, Du J, Fan J, Peng X. NIR-emitting carbon dots for discriminative imaging and photo-inactivation of pathogenic bacteria. Chem Eng J. 2022;450:137384. https://doi.org/10.1016/j.cej.2022.137384.
Hua J, Hua P, Qin K. Highly fluorescent N, F co-doped carbon dots with tunable light emission for multicolor bio-labeling and antibacterial applications. J Hazard Mater. 2023;459:132331. https://doi.org/10.1016/j.jhazmat.2023.132331.
Zheng L, Qi P, Zhang D. Identification of bacteria by a fluorescence sensor array based on three kinds of receptors functionalized carbon dots. Sens Actuators B Cheml. 2019;286:206–13. https://doi.org/10.1016/j.snb.2019.01.147.
Li H, Yao S, Wang X, Xu H, Zhao C, Li J, Wang J. Rapid statistical discrimination of fluorescence images of multiple foodborne pathogens using multicolor-emissive carbon dots. Microchem J. 2024;197:109719. https://doi.org/10.1016/j.microc.2023.109719.
Nandi S, Ritenberg M, Jelinek R. Bacterial detection with amphiphilic carbon dots. Analyst. 2015;140:4232–7. https://doi.org/10.1039/C5AN00471C.
Pathak A, Navaneeth P, Gupta M, Pradeep A, Nair BG, Suneesh PV, Elangovan R, Sundberg L-R, Marjomäki V, Babu TGS. Revolutionizing Gram-negative bacteria detection: flim and multicolor imaging based selective interaction study using colistin passivated carbon dots. Sens Actuators B Chem. 2023;395:134433. https://doi.org/10.1016/j.snb.2023.134433.
Zhao M, Gao X, Tao Z, Wang X, Lin X, Wang S, Liu Y. Sugar-metabolism-triggered pathogenic bacteria identification based on pH-sensitive fluorescent carbon dots. Sens Actuators B Chem. 2020;316:128063. https://doi.org/10.1016/j.snb.2020.128063.
Ran H-H, Cheng X, Bao Y-W, Hua X-W, Gao G, Zhang X, Jiang Y-W, Zhu Y-X, Wu F-G. Multifunctional quaternized carbon dots with enhanced biofilm penetration and eradication efficiencies. J Mater Chem B. 2019;7:5104–14. https://doi.org/10.1039/C9TB00681H.
Lin F, Li C, Dong L, Fu D, Chen Z. Imaging biofilm-encased microorganisms using carbon dots derived from L. plantarum. Nanoscale. 2017;9:9056–64. https://doi.org/10.1039/C7NR01975K.
van Hoogstraten SWG, Kuik C, Arts JJC, Cillero-Pastor B. Molecular imaging of bacterial biofilms—a systematic review. Crit Rev Microbiol. 1–22. https://doi.org/10.1080/1040841X.2023.2223704.
Lin J, Xu L, Zheng Y, Wu D, Yue J. Imitation-mussel fluorescent silicon quantum dots for selective labeling and imaging of bacteria and biofilms. Front bioeng biotechnol. 2022;10. https://doi.org/10.3389/fbioe.2022.971682.
Zhao D, Zhang R, Liu X, Li X, Xu M, Huang X, Xiao X. Screening of chitosan derivatives-carbon dots based on antibacterial activity and application in anti-Staphylococcus aureus biofilm. Int J Nanomedicine. 2022;17:937–52. https://doi.org/10.2147/ijn.S350739.
Jian H-J, Wu R-S, Lin T-Y, Li Y-J, Lin H-J, Harroun SG, Lai J-Y, Huang C-C. Super-cationic carbon quantum dots synthesized from spermidine as an eye drop formulation for topical treatment of bacterial keratitis. ACS Nano. 2017;11:6703–16. https://doi.org/10.1021/acsnano.7b01023.
Yan C, Wang C, Hou T, Guan P, Qiao Y, Guo L, Teng Y, Hu X, Wu H. Lasting tracking and rapid discrimination of live Gram-positive bacteria by peptidoglycan-targeting carbon quantum dots. ACS Appl Mater Interfaces. 2021;13:1277–87. https://doi.org/10.1021/acsami.0c19651.
Damarla K, Mehra S, Kang TS, Yadav S, Mishra A, Kumar A. DES-N-doped oxygenated carbon dot colloidal solutions for light harvesting and bio-imaging applications. Mater Adv. 2020;1:3476–82. https://doi.org/10.1039/D0MA00621A.
Gao Z, Zhao CX, Li YY, Yang Y. Beer yeast-derived fluorescent carbon dots for photoinduced bactericidal functions and multicolor imaging of bacteria. Appl Microbiol Biotechnol. 2019;103:4585–93.
Lu F, Song Y, Huang H, Liu Y, Fu Y, Huang J, Li H, Qu H, Kang Z. Fluorescent carbon dots with tunable negative charges for bio-imaging in bacterial viability assessment. Carbon. 2017;120:95–102. https://doi.org/10.1016/j.carbon.2017.05.039.
Tu Y, Wang S, Yuan X, Xiang Y, Qin K, Wei Y, Zhang Q, Chen X, Ji X. Facile hydrothermal synthesis of nitrogen, phosphorus-doped fluorescent carbon dots for live/dead bacterial differentiation, cell imaging and two nitrophenols detection. Dyes Pigm. 2021;184:108761. https://doi.org/10.1016/j.dyepig.2020.108761.
Yuan X, Tu Y, Chen W, Xu Z, Wei Y, Qin K, Zhang Q, Xiang Y, Zhang H, Ji X. Facile synthesis of carbon dots derived from ampicillin sodium for live/dead microbe differentiation, bioimaging and high selectivity detection of 2,4-dinitrophenol and Hg(II). Dyes Pigm. 2020;175:108187. https://doi.org/10.1016/j.dyepig.2020.108187.
Ji X, Wang S, Luo Y, Yuan X, Wei Y, Zhang Q, Qin K, Tu Y. Green synthesis of weissella-derived fluorescence carbon dots for microbial staining, cell imaging and dual sensing of vitamin B12 and hexavalent chromium. Dyes Pigm. 2021;184:108818. https://doi.org/10.1016/j.dyepig.2020.108818.
Liu Y, Xu Y, Wen Q. Carbon dots for staining bacterial dead cells and distinguishing dead/alive bacteria. Anal Biochem. 2024;687:115432. https://doi.org/10.1016/j.ab.2023.115432.
Yu X-W, Liu X, Jiang Y-W, Li Y-H, Gao G, Zhu Y-X, Lin F, Wu F-G. Rose bengal-derived ultrabright sulfur-doped carbon dots for fast discrimination between live and dead cells. Anal Chem. 2022;94:4243–51. https://doi.org/10.1021/acs.analchem.1c04658.
Lin F, Li C, Chen Z. Exopolysaccharide-derived carbon dots for microbial viability assessment. Front Microbiol. 2018;9: https://doi.org/10.3389/fmicb.2018.02697.
Qie X, Zan M, Li L, Gui P, Chang Z, Ge M, Wang R-S, Guo Z, Dong W-F. High photoluminescence nitrogen, phosphorus co-doped carbon nanodots for assessment of microbial viability. Colloids Surf B Biointerfaces. 2020;191:110987. https://doi.org/10.1016/j.colsurfb.2020.110987.
Sun L, Zhang H, Wang Y, Xiong Z, Zhao X, Xia Y. Chitosan-derived N-doped carbon dots for fluorescent determination of nitrite and bacteria imaging. Spectrochim Acta A Mol Biomol Spectrosc. 2021;251:119468. https://doi.org/10.1016/j.saa.2021.119468.
Funding
This work was supported by the National Science and Technology Council of Taiwan under Contract No. 112-2811-M-019-008 and the Center of Excellence for the Oceans, National Taiwan Ocean University, Taiwan (R.O.C.).
Author information
Authors and Affiliations
Contributions
Anisha Anand: writing—original draft; review and editing. Chih-Ching Huang: conceptualization; writing—review and editing. Jui-Yang Lai: conceptualization; writing—review and editing. Darakhshan Bano: writing—original draft. Helen Indah Pardede: writing—original draft. Amina Hussain: writing—original draft. Sehresh Saleem: writing—original draft. Binesh Unnikrishnan: conceptualization; writing—review and editing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper. Chih-Ching Huang is guest editor of Analytical and Bioanalytical Chemistry for the topical collection featuring Luminescent Nanomaterials but was not involved in the peer review of this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Published in the topical collection Luminescent Nanomaterials for Biosensing and Bioimaging with guest editors Li Shang, Chih-Ching Huang, and Xavier Le Guével.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Anand, A., Huang, CC., Lai, JY. et al. Fluorescent carbon dots for labeling of bacteria: mechanism and prospects—a review. Anal Bioanal Chem 416, 3907–3921 (2024). https://doi.org/10.1007/s00216-024-05300-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-024-05300-1