Abstract
The detection of Salmonella spp. in food samples is regulated by the ISO 6579:2002 standard, which requires that precise procedures are followed to ensure the reliability of the detection process. This standard requires buffered peptone water as a rich medium for the enrichment of bacteria. However, the effects of different brands of buffered peptone water on the identification of microorganisms by Raman spectroscopy are unknown. In this regard, our study evaluated the discrimination between two bacterial species, Salmonella enterica and Escherichia coli, inoculated and analyzed with six of the most commonly used buffered peptone water brands. The results showed that bacterial cells behaved differently according to the brand used in terms of biomass production and the spectral fingerprint. The identification accuracy of the analyzed strains was between 85% and 100% depending on the given brand. Several batches of two brands were studied to evaluate the classification rates between the analyzed bacterial species. The chemical analysis performed on these brands showed that the nutrient content was slightly different and probably explained the observed effects. On the basis of these results, Raman spectroscopy operators are encouraged to select an adequate culture medium and continue its use throughout the identification process to guarantee optimal recognition of the microorganism of interest.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the European Union, Salmonella is the second most frequent bacterial genus involved in gastrointestinal epidemics in humans [1]. The ISO 6579 standard describes the process to identify this pathogen in food samples and proposes a panel of traditional methods able to identify isolated strains by immunological, biochemical, or molecular methods [2]. Over the last decade, new technologies have emerged for the identification of bacteria including surface plasmon resonance, nuclear magnetic resonance, mass spectrometry (especially its variant MALDI-TOF), and many other techniques based on vibrational spectroscopy [3,4,5]. These new techniques are rapid, user-friendly, and highly reliable for food manufacturers. Among these new analytical techniques, Raman spectroscopy has been proposed with the aim of simplifying the analysis process and reducing the investigation time of samples in various fields of application [6, 7]. This optical method is based on the inelastic scattering of light and allows the molecular compositions of several types of samples to be measured quickly and nondestructively [8]. Currently, Raman spectroscopy is used to identify bacteria isolated from medical samples [9,10,11], food matrices [12, 13], and many other domains [14,15,16]. The high sensitivity of this technique provides considerable advantages compared with other vibrational techniques, e.g., the limit of detection of Raman microspectroscopy is at the single-cell level [17, 18]. Overtaking the limitations of infrared spectroscopy, Raman spectroscopy is suitable for analysis in aqueous conditions such as those encountered in industrial food processes. Regardless of the advantages of Raman spectroscopy, some drawbacks remain. For instance, the chemical composition of living samples is continuously changing and depends on the life cycle of the cells, generating changes in the Raman spectra [19, 20]. These changes observed for bacteria make it possible to connect the variations in the intensities of nucleic acid bands to the physiological state of cells [20, 21]. To ensure the highest quality of Raman spectra, the literature recommends analyzing bacteria in the exponential growth phase [22]. However, for colonies that contain a mixture of bacteria in different physiological states, only small colonies containing mostly young cells should be analyzed [23]. To guarantee optimal discrimination between bacterial colonies, only spectra having a high DNA/RNA ratio should be selected [22]. In addition to the impact of bacterial growth stage, the spectral fingerprint of bacteria depends largely on the culture conditions and the procedure used to inoculate and isolate the targeted microorganism [13, 24]. In fact, the temperature, oxygenation, and amount of nutrients present in the growth media affect bacterial development and Raman spectra [25, 26]. Consequently, when spectral fingerprints are used, the identification process must use the same method to isolate the microorganisms [24]. This challenge reported for the discrimination between bacteria is potentially due to the residuals from growth media in the bacterial cells [27]. Nonetheless, the Raman bands responsible for this discrimination are similar to those of the chemical molecules that are naturally present in the cell; additionally, Raman spectra remain unchanged after multiple washing cycles of the bacterial cells [28]. In further studies, Raman spectroscopy was used to identify Salmonella spp. according to the ISO 6579:2002 standards [22]. As reported, this standard indicates the procedure that should be followed to identify the presence of this pathogen in samples and dictates all of the growth media that should be used in the applied process [2]. Buffered peptone water (BPW) is one of the rich growth media recommended by this standard for the investigation of Salmonella spp. in food matrices. However, in food industry practices, several suppliers have proposed various BPW brands for the inoculation of bacteria, and many batches sometimes exist for each brand. The ISO standard does not designate any particular brand that should be used in the investigation process; consequently, the end user has many choices for the same growth medium, and the various media may have different qualities and unknown impacts on the identification of bacteria by spectroscopic methods. The present study aims to evaluate the impact of BPW brands and batches on the quality of the Raman bacterial fingerprint and on the discrimination efficiency between Salmonella enterica and Escherichia coli used as model bacterial cells.
Materials and methods
Bacterial strains and growth conditions
Two bacterial strains were used in our study: S. enterica Paratyphi B (CIP 55.42) and E. coli K12 (ATCC 700926). These two strains were inoculated from cryogenic tubes stored at −80 °C in buffered peptone water with a cryoprotectant (15% sterile glycerol). Precultures were conducted in flasks (100 ml) containing 10 ml of BPW and incubated overnight at 30 °C with stirring at 250 rpm (IKA, KS 4000 ic control, Germany). The precultures were used to inoculate each bacterial strain in triplicate in 250-ml flasks containing 50 ml of BPW. The starting optical density (OD) was equal to 0.1 for triplicates (UV–Vis spectrophotometer, Helios Ɛ, UVE 082917, France). Bacterial growth was followed by measuring the optical density at 620 nm (OD620nm) over time. At the exponential growth phase, 15 ml of each culture was centrifuged at 6000 g for 5 min (Awel, MF 20-R, France), and 10 μl of the obtained biomass was used for Raman analyses.
Buffered peptone water (BPW)
The six principal brands of commercialized BPW used in our study are displayed in Table 1. These different media were prepared following the manufacturer’s instructions. For agar plates, 15 g of bacteriological agar type E (Biokar diagnostics, ref. A1012HA, France) was added to 1 l of prepared liquid media. These media were autoclaved at 120 °C for 20 min and, after sterilization, the BPW with agar was poured into sterile petri dishes and stored at 4 °C until use (1 month maximum).
Chemical analysis
Ten milliliters of each brand was prepared from lyophilized powder following the respective manufacturer’s instructions, and the total organic carbon was measured with a TOC-VCSN analyzer (Shimadzu, France). Eighty grams of each lyophilized brand was sent to Eurofins Scientific (France) for elemental analysis (Table 2). Samples were sent under sterile conditions to avoid degradation during transport. The analysis of the dissolved inorganic particles was conducted by inductively coupled plasma optical emission spectrometry (ICP-OES) according to ISO 11885:2009 [29]. The determination of the amino acid content was conducted by HPLC according to ISO 13903:2005 [30]. The protein analysis was conducted by the Kjeldahl method, and the lipids were analyzed by acid hydrolysis following Soxhlet extraction [31].
Raman microspectroscopy measurements
The Raman spectra were acquired using a Raman spectrometer (Senterra, Bruker Optics, France) driven by Opus software (Bruker Optics GmBH, V 7.2, Germany). This device was equipped with two gratings (400 and 1200 lines/mm), a CCD camera cooled to −60 °C, and a BX51 Olympus microscope with multiple objectives (the objective LCLanN100x/0.85 was used in the analyses, the laser spot = 1.12 μm). The analyses were performed at 785 nm, and the laser power on the sample was approximately 25 mW. The spectral resolution was approximately 8 cm−1. Five acquisitions of 10 s each were necessary for each spectrum (50 s for each measurement). The bacteria were analyzed with a procedure reported in a previous study [22]. Briefly, the strains were grown under the recommended conditions (temperature and growth media) on agar plates or liquid cultures. After incubation, the liquid cultures were centrifuged, and 10 μl of the obtained biomass was deposited onto gold slides, dried for 5–10 min at room temperature, and finally analyzed by Raman spectroscopy. For colonies, bacteria were picked up from separate colonies and spread onto gold surfaces. A total of 15 spectra were acquired for each deposit, and three deposits from three different cultures were analyzed (3 × 3 × 15 = 135 spectra for each brand).
Spectral preprocessing and data analysis
Preprocessing
The Raman spectra were processed using Opus software (Bruker optics GmBH, V 7.2, Germany). The spectral range from 350 to 1750 cm−1 was used in our study for the discrimination and classification of bacteria. An elastic concave (64° and 10 iterations) method was used for the baseline correction in the range of interest. The protein band at 1245 cm−1 was used to normalize the spectra (min–max normalization). To guarantee good quality spectra for discrimination, only spectra from bacteria in the exponential phase were chosen by selecting spectra with a high DNA/RNA ratio at 780–820 cm−1, as reported in a previous study [22]. To determine the chemical compositions of the different brands of BPW, the data were standardized by subtraction of the average and divided by the standard deviation before using principal component analysis (PCA).
Data exploration
MATLAB software (MathWorks, Inc. V 2013, France) with the statistical toolbox was used for data exploration. The SAISIR® package was used to perform the same statistical treatments as previously described [22, 32]. The 2D plot of the spectra was produced using the imagesc function from MATLAB, which is a top-down representation of the Raman intensities. The PCA was conducted on the spectral matrix with the pca function, and the 3D plot of PCA scores was conducted with the first three PCs (PC1, PC2, and PC3), which contain the majority of the data information. The normed PCA was determined with the normed_pca function and used to analyze the chemical compositions between brands. Factorial discriminant analysis (FDA) was performed with the fda2 function of the SAISIR® package and used as a supervised method to evaluate the spectral data sets. The first 10 PC scores were used in the calculation, and the displayed results of FDA were the average of 100-fold iterations. The Kruskal–Wallis test was applied on the first PCA score with the kruskalwallis function in MATLAB. This statistical analysis is a nonparametric version of the classical one-way ANOVA that compares the medians of tested samples. The Multcompare function was combined with the Kruskal–Wallis test to determine which pairs of data were significantly different. Further information regarding the SAISIR® MATLAB procedures can be found elsewhere [33, 34].
Results and discussion
Growth and Raman spectra of S. enterica and E. coli on different BPW brands
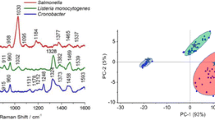
Salmonella enterica Paratyphi B (Sp) was cultivated on six BPW brands mostly used in laboratories (Biokar, Merck, Oxoid, Fisher, Sigma, and Difco) to evaluate their impact on the identification of bacteria. The growth profiles were monitored for 20 h and showed that Salmonella cultures on Biokar had the highest optical density at 620 nm (OD620nm = 2.9 ± 0.02), followed by Merck (OD620nm = 2.3 ± 0.03), Oxoid (OD620nm = 1.9 ± 0.056), and Fisher (OD620nm = 1.5 ± 0.02). The cultures on Sigma (OD620nm = 0.9 ± 0.01) and Difco (OD620nm = 0.7 ± 0.007) had the lowest optical densities and consequently the lowest biomass production (Table 1). The growth profiles of all brands are presented in Electronic Supplementary Material (ESM), Fig. S1. The highest growth rates (μ) were observed for Biokar, Merck, and Oxoid (μ = 0.82, 0.77, 0.81 h−1, respectively). Fisher and Difco had intermediate values (μ = 0.66 and 0.51 h−1), whereas Sigma (μ = 0.48 h−1) had the lowest values. The E. coli cultures had the highest optical density and growth rate on Biokar (OD620nm = 2.73 ± 0.027 and μ = 0.80 ± 0.005), Oxoid (OD620nm = 2.18 ± 0.036 and μ = 0.70 ± 0.009), and Merck (OD620nm = 1.53 ± 0.056 and μ = 0.76 ± 0.004). The lowest values were obtained for Difco (OD620nm = 0.73 ± 0.002 and μ = 0.37 ± 0.002) and Sigma (OD620nm = 0.72 ± 0.013 and μ = 0.41 ± 0.022) (Table 1). The Kruskal–Wallis test was applied to the growth profiles of Salmonella grown on all studied brands, and the results show that the medians of groups were significantly different, notably between the Biokar and Difco brands (p = 0.0065). These results indicate that bacterial cells have a different growth behavior in different brands of the same culture medium. Different bacterial colonies inoculated on these brands were analyzed by Raman spectroscopy to evaluate the ability to discriminate different strains in various brands. The spectral fingerprint of Salmonella Paratyphi B showed many differences in the Raman shifts and the intensities of Raman profiles depending on the brand used. Visual inspection of the 2D plot of the spectra showed the Raman bands of the macromolecules present in the bacterial cells, and the bands were impacted by the growth medium used (Fig. 1a): 480 cm−1 for C–O–C of carbohydrates, 780 cm−1 for uracil ring, 810 cm−1 for C–O–P–O–C of RNA backbone, 1270 cm−1 for C–N of amide III, 1350 cm−1 for C–H stretching, 1455 cm−1 for C–H2 stretching, and 1665 cm−1 for C=O of amide I. The PCA conducted on these spectra (750 spectra) showed the presence of several groups more or less distant depending on their degree of similarity (Fig. 1b). In fact, the spectra of the bacteria inoculated on Difco, Oxoid, Merck, and Fisher were close and formed nearly a single group, whereas the spectra of Sigma and Biokar were spaced apart slightly. The analysis of the first three PCA loadings attributes this discrimination to the more Raman bands shown in the 2D plot (highlighted areas in Fig. 1c). The loadings PC1 (38%) and PC2 (19.8%) contributed equally to separate Biokar from the other BPW brands. The Raman bands at 780, 810, 1000, 1450, and 1665 cm−1, which correlated in the negative part of PC1 and PC2, were responsible for this discrimination. PC3 (13.6%) contributed to the separation of Sigma (bands at 475, 1443, and 1650 cm−1) from Merck (bands at 723, 780, and 814 cm−1). Therefore, the change in the composition of brands affected the entire bacterial cell and did not manifest in one specific location of the Raman fingerprint. The Kruskal–Wallis test confirmed the presence of five distinct groups of brands with a p value of 2 × 10−61 ≪ 0.05, G1, Fisher; G2, Merck; G3, Oxoid and Sigma; G4, Biokar; and G5, Difco (Fig. 1d). Note that Oxoid and Sigma were classified together in the same group because their medians were not significantly different. These results confirm that the influence of BPW brands affects not only bacterial growth but also the spectral fingerprint of microbial cells. From a physiological point of view, this observation could be explained either by the change in the metabolism of bacterial cells according to the provided nutrients or by the presence of chemical interactions between the constituents of different brands and the bacterial membranes (or some organelles).
Evaluation of the impact of the buffered peptone water brands on the spectral fingerprint of S. enterica. a 2D plot of Raman spectra intensities with the brands that were used for the inoculation of bacteria (Y-axis represents the Raman spectra order in the data matrix), Biokar, 120 spectra; Merck, 130 spectra; Fisher, 135 spectra; Difco, 95 spectra; Oxoid, 135 spectra; and Sigma, 125 spectra. b 3D plot of PC1, PC2, and PC3 scores of PCA applied on the Raman spectra. c Loadings on PCA scores (PC1, PC2, and PC3). d Kruskal–Wallis test based on the PC1 score of PCA: G1, G2, G3, G4, and G5 are the observed groups
Determination of chemical composition of BPW brands
Chemical analysis was conducted for all of the brands used to understand the observed impact on the growth and the Raman spectra of the bacteria examined in our study. Table 2 summarizes the results obtained from Eurofins laboratories. These analyses revealed many elements that can widely influence the growth of bacteria, including amino acids, vitamins, or other nutrients (Table 2). Given the difficulty of interpreting the impact of all these nutrients on bacterial cells, the data were explored by chemometric methods. The visualization of the first PCA score and loadings indicated the relative properties of the brands from a chemical point of view. Three groups were apparent. Merck, Difco, and Oxoid formed the first group with negative PC1 scores, Sigma and Fisher formed the second group with high positive PC1 scores, and Biokar formed a third group with an intermediate PC1 score (Fig. 2a). The analysis of the PC1 loading plot showed the correlation between the BPW brands and their chemical compositions (Fig. 2b). The total nitrogen and the amino acids lysine, serine, threonine, phenylalanine, methionine, histidine, leucine, valine, taurine, isoleucine, and glutamic acid had the highest contribution to the PC1 scores to separate Fisher and Sigma from other brands. Magnesium, phosphorus, potassium, organic carbon, dry extract, amino acids such as ornithine, glycine, alanine, tyrosine, and arginine, and total carbohydrates influence the discrimination between Oxoid, Merck, and Difco from other brands. Biokar is mainly different from the other brands in the amount of sodium, calcium, proline, asparagine, cysteine, carbohydrates, proteins, and lipids. These results are consistent with the chemical analysis presented in Table 2. The most important impact is related to the carbon sources available to bacteria. In fact, Biokar contains the highest ratio of carbohydrates (10 g/kg) and the highest ratio of total organic carbon, including proteins and lipids. The food energy provided by 100 g of the Biokar brand is 26 kJ. This high level of carbohydrates provides bacteria a significant carbon source necessary for their development during different growth phases. Hence, S. enterica and E. coli inoculated on Biokar brand had the highest biomass production (Table 1). The Raman spectra were also very different from those of other brands (Fig. 1). In fact, the spectra of Salmonella inoculated on this brand (G4) were different from other Raman spectra of Salmonella inoculated on the other brands. Merck contains 0.3 g/kg carbohydrates and the second highest ratio of proteins (0.7 g/kg), and the food energy of the Merck brand is lower than that of the Biokar brand (17 kJ/100 g). Salmonella had the second highest biomass production on the Merck brand (Table 1). The Raman spectra of Salmonella on the Merck brand were also impacted and were significantly different from those of Salmonella on other brands (Fig. 1). The Oxoid and Fisher brands contain a low concentration of carbohydrates (0.1 g/kg), and Difco and Sigma contain a very low concentration of carbohydrates (< 0.1 g/kg). Unlike the other brands, Difco contains a high ratio of inorganic phosphorus (110 g/kg). The Raman spectra of Salmonella were different on the brands tested except for Oxoid and Sigma. This result shows the remarkable difference between the analyzed brands and explains the divergence observed in the growth and the spectral fingerprint of the same bacterial species inoculated on the brands tested.
Principal component analysis on the standardized data of the chemical compositions obtained for all of the BPW brands. a Plot of the PC1 scores for the brand function. b Plot of the first loading (PC1) showing the contribution of the chemical compositions on the discrimination between the analyzed brands by PCA. Amino acid letter codes: alanine (Ala), cysteine (Cys), aspartic acid (Asp), glutamic acid (Glu), phenylalanine (Phe), glycine (Gly), histidine (His), isoleucine (Iso), lysine (Lys), leucine (Leu), methionine (Met), asparagine (Asp), ornithine (Orn), proline (Prl), glutamine (Glu), arginine (Arg), serine (Ser), threonine (Thr), valine (Val), tryptophan (Try), tyrosine (Tyr), and taurine (Tau). Inorganic compound codes: potassium (Pot), calcium (Cal), phosphorus (Pho), magnesium (Mag), and sodium (Sod). Proteins (Pro), total nitrogen (To-N), lipids (Lip), total carbohydrate (T-Ca), total organic carbon (To-C), dry extract (Dry), food energy (Foo), ash (Ash)
Impact of growth media composition on discrimination between S. enterica and E. coli
Raman spectra of Salmonella Paratyphi B and E. coli colonies inoculated on the six brands were analyzed by statistical methods to evaluate their discrimination. Different scenarios of statistical comparisons could be considered to evaluate the ability of the models to discriminate between Salmonella and Escherichia despite the BPW brands. The first strategy was to compare all of the obtained spectra (1360 spectra) to evaluate the global discrimination between these two strains regardless of the brand used. The 3D projection of the first three principal components of PCA (PC1, 38%; PC2, 19%; and PC3, 14%)—a confidence ellipse with significance level = 0.05—allows the visualization of the spatial distribution of colony spectra inoculated on the Biokar, Difco, Merck, Oxoid, Fisher, and Sigma brands (Fig. 3a). This observation shows the absence of any clear separation between Sp and Ec depending on the analyzed PC. The factorial discriminant analysis was used to confirm the PCA observation and to evaluate the percentage of classification between spectra separated into data for model training (3/4 of data) and validation (1/4 of data). The validation results show 92% good classification for Salmonella and 95% good classification for Escherichia. In terms of error, 8% of Salmonella spectra (108 spectra) were classified in the Escherichia group, and 5% of Escherichia spectra (68 spectra) were classified in the Salmonella group (Fig. 3b). The high number of errors in the classification of spectra may be the consequence of an overall effect or due to a single (or more) brand. The second strategy was to compare spectra of bacteria inoculated only on the same brand. The different PCA representations show a heterogeneous distribution of the spectra according to the studied brand (ESM, Fig. S2). The variability inside the same ellipse (same bacterial strain) and the distance between the ellipses of Sp and Ec are also different according to the brands. Nine regions of Raman spectra that are potentially responsible for this observation were identified by the analysis of the first PC loading: 446–497, 626–677, 716–857, 972–1037, 1088–1140, 1178–1230, 1306–1326, 1422–1500, and 1557–1609 cm−1 (ESM, Fig. S3). In parallel, factorial discriminant analysis was used to evaluate the discrimination between these two bacterial species and to calculate the error rates for each brand. The results show that the classification of Salmonella is almost stable when the same brand is used (Fig. 4). A perfect classification was observed for Biokar (99% of classification), Difco (100%), Fisher (99%), Sigma (98%), and Merck (99%), while the classification was very low for Oxoid (85%). The BPW impacts the growth and Raman fingerprints of bacteria as a result of the difference in the chemical composition of the commercial brands. Normally, culture media dedicated to the same application should contain equivalent amounts and types of nutrients and provide a similar effect with respect to the identification of a bacterial strain. For more consideration, standards with precise compositions (and concentrations) of products should be used in bacterial research, but not brands. The ISO 6579:2002 standards, used as an example in our study, do not specify any brands of BPW because the classical methods for Salmonella identification are not affected by changes in brand composition. In contrast, the Raman signal is sensitive to the molecular environment and the physiochemical changes of cells. Thus, any variations in the composition of the culture medium may cause disturbances in the growth of bacterial cells and probably in their molecular composition. For routine applications, errors in Salmonella or Escherichia classification may have a dramatic consequence on the safety of consumers. For identification, Raman spectroscopy users must use suitable brands for their applications and preferably keep using the same brand to obtain the best classification rates.
Evaluation of impact of batch on discrimination between S. enterica and E. coli
For the studied brands, the discrimination between Sp and Ec was evaluated on two different batches from Biokar and four different batches from Merck. No additional batches were obtained for the remaining brands in this study. For Salmonella cultures, the first batch of Biokar (B1) had an OD620nm of 2.9 ± 0.02 and μ of 0.82 ± 0.017 (h−1). The second Biokar batch (B2) had an OD620nm of 2.8 ± 0.011 and μ of 0.86 ± 0.009 (h−1). The optical densities and growth rates of the Merck batches were as follows: M1 (2.3 ± 0.03, μ = 0.77 ± 0.004), M2 (2.2 ± 0.02, μ = 0.78 ± 0.023), M3 (2.8 ± 0.01, μ = 0.76 ± 0.014), and M4 (2.2 ± 0.02, μ = 0.80 ± 0.012). A similar tendency was observed for Escherichia cultures inoculated on these batches (data provided in Table 1). The Kruskal–Wallis test was applied to the growth data with different Merck and Biokar batches and showed no significant difference (p values were significantly higher than 0.05: p for Merck = 0.3916 and p for Biokar = 0.318); consequently, different batches had no impact on the growth of bacteria.
Raman data obtained from these batches were evaluated by factorial discriminant analysis. The comparison between spectra of Sp and Ec inoculated on the second Biokar batch (Sp and Ec on B2) gave 95% good discrimination (the classification rate of Sp vs Ec on B1 was 99%). This result shows a remarkable difference in the rate of discriminating Sp between these two batches (4.8% decrease in the classification rate). The spectra of the second batch (B2) were used to predict the bacterial type from the model performed only on B1 spectra, and 87% of Salmonella good classification was obtained in this case (13% error). This result reveals the difference in the rate of classification between these two batches. The grouping of all Ec and Sp spectra inoculated on the two Biokar batches in the same database increased the recognition of Sp to 96% of the total good classification (4% error). The same comparison was performed on the Merck batches. The FDA was applied separately on bacteria inoculated on the same batches and showed an excellent classification (100%) for all cultures of Sp and Ec (M1, 176 spectra; M2, 253; M3, 212; and M4, 220). Nevertheless, the classification was strongly impacted when spectra of each batch were compared to the set of spectra created after the four batches were pooled: M1, 85%; M2, 92%; M3, 87%; and M4, 99%. As with the Biokar batch, the grouping of all Ec and Sp spectra inoculated on all batches in the same database increased the recognition of Sp on Merck to 99%. This result confirms that the impact of the batch must be considered in the model to ensure better recognition of bacteria.
The different rates of discrimination obtained on the different batches suggest that there is some effect of batch on the classification of bacteria by Raman spectroscopy on these two brands. The model is much more efficient if batch-related variability is considered. In this regard, the diversity observed between the brands in terms of biomass production and Raman spectra adds an additional challenge for the use of Raman spectroscopy in the analysis of living samples without the establishment of statistical models able to eliminate the impact of the culture media. The possibility of eliminating this impact was reported for some techniques, such as mass spectrometry [35]. However, the filtering of the data is easier with mass spectra, in which the variables are related to a specific fragment of a molecule. Therefore, the fragments characterizing the chemical compounds of the growth media may be removed, leaving the characteristics of all other variables of the bacteria in the remaining mass spectrum. Moreover, Raman data consist of a mixture of vibrational signals and are not a collection of discrete m/z intensities; therefore, the elimination of the bands responsible for the brand features is much more difficult than with mass spectra. Consequently, it is very difficult to separate the signal produced by the molecules constituting the bacteria from the signal exclusively due to the culture medium. This effect forces the end user to define and always use the same brand of culture media to ensure the best Raman spectra necessary for the identification process.
Conclusion
The food standards establish the exact procedure that should be followed to guarantee reliable identification of microorganisms. According to ISO 6579:2002, buffered peptone water is required for the enrichment of Salmonella present in food samples. Nevertheless, this medium is provided by many manufacturers, and these products are dedicated to the same application. The present study evaluated the use of six brands of BPW to discriminate S. enterica from E. coli by Raman spectroscopy. The analysis of the growth profiles in these brands showed that the bacteria behaved differently. Biokar provided the highest biomass production, followed by Merck, Oxoid, and Fisher, whereas Sigma and Difco had the lowest biomass production. The observed difference was due to the various ratios of nutrients contained in the different culture media, as demonstrated by chemical analysis. These changes in the composition impacted the spectral fingerprint of the bacteria differently and subsequently affected their recognition. As shown in our study, the success rates of discrimination by Raman spectroscopy are dependent on the brand used: 100% for Difco, 99% for Biokar, Merck, and Fisher, 98% for Sigma, and 85% for Oxoid. Four batches of Merck and two batches of Biokar were used to evaluate the effect of batches on the discrimination between bacteria. Some differences were found between batches of the same brands; 96% of global good classification was observed for Salmonella inoculated on Biokar and 99% on Merck batches. The different rates of discrimination obtained on the different batches suggest that there is some effect of batch on the classification of bacteria by Raman spectroscopy, and the statistical models are much more efficient if batch-related variability is considered. The operator can obtain different classification rates leading to either the correct or incorrect identification of the analyzed strains depending on the brand used. Given the complexity needed to eliminate this impact from Raman signals, end users are encouraged to carefully select an adequate culture medium suitable for the identification process. A new data algorithm able to remove the impact of the culture medium may be an important solution to ensure the high performance of microorganism recognition.
References
Paniel N, Noguer T. Detection of Salmonella in food matrices, from conventional methods to recent aptamer-sensing technologies. Foods. 2019;8:371. https://doi.org/10.3390/foods8090371.
International Organization for Standardization. ISO 6579:2002. Horizontal method for the detection of Salmonella spp. Microbiology of food and animal feeding stuffs. Geneva: ISO.
Bhandari D, Chen F-C, Bridgman RC. Detection of Salmonella typhimurium in romaine lettuce using a surface plasmon resonance biosensor. Biosensors. 2019;9. https://doi.org/10.3390/bios9030094.
Cheng K, Chui H, Domish L, Hernandez D, Wang G. Recent development of mass spectrometry and proteomics applications in identification and typing of bacteria. Proteomics Clin Appl. 2016;10:346–57. https://doi.org/10.1002/prca.201500086.
Chen M, Shpirt AM, Guo X, Shashkov AS, Zhuang Y, Wang L, et al. Identification serologically, chemically and genetically of two Escherichia coli strains as candidates for new O serogroups. Microbiol Read Engl. 2015;161:1790–6. https://doi.org/10.1099/mic.0.000136.
Ho C-S, Jean N, Hogan CA, Blackmon L, Jeffrey SS, Holodniy M, et al. Rapid identification of pathogenic bacteria using Raman spectroscopy and deep learning. Nat Commun. 2019;10:1–8. https://doi.org/10.1038/s41467-019-12898-9.
Su W-H, He H-J, Sun D-W. Non-destructive and rapid evaluation of staple foods quality by using spectroscopic techniques: a review. Crit Rev Food Sci Nutr. 2017;57:1039–51. https://doi.org/10.1080/10408398.2015.1082966.
Baker MJ, Byrne HJ, Chalmers J, Gardner P, Goodacre R, Henderson A, et al. Clinical applications of infrared and Raman spectroscopy: state of play and future challenges. Analyst. 2018;143:1735–57. https://doi.org/10.1039/c7an01871a.
Kirschner C, Maquelin K, Pina P, Thi NAN, Choo-Smith LP, Sockalingum GD, et al. Classification and identification of enterococci: a comparative phenotypic, genotypic, and vibrational spectroscopic study. J Clin Microbiol. 2001;39:1763–70. https://doi.org/10.1128/JCM.39.5.1763-1770.2001.
Boardman AK, Wong WS, Premasiri WR, Ziegler LD, Lee JC, Miljkovic M, et al. Rapid detection of bacteria from blood with surface-enhanced Raman spectroscopy. Anal Chem. 2016;88:8026–35. https://doi.org/10.1021/acs.analchem.6b01273.
Pence I, Mahadevan-Jansen A. Clinical instrumentation and applications of Raman spectroscopy. Chem Soc Rev. 2016;45:1958–79. https://doi.org/10.1039/c5cs00581g.
Mizrach A, Schmilovitch Z, Korotic R, Irudayaraj J, Shapira R. Yeast detection in apple juice using Raman spectroscopy and chemometric methods. Trans ASABE. 2007;50:2143–9.
Meisel S, Stöckel S, Rösch P, Popp J. Identification of meat-associated pathogens via Raman microspectroscopy. Food Microbiol. 2014;38:36–43. https://doi.org/10.1016/j.fm.2013.08.007.
Rae A, Stosch R, Klapetek P, Hight Walker AR, Roy D. State of the art Raman techniques for biological applications. Methods. 2014;68:338–47. https://doi.org/10.1016/j.ymeth.2014.02.035.
Smith R, Wright KL, Ashton L. Raman spectroscopy: an evolving technique for live cell studies. Analyst. 2016. https://doi.org/10.1039/C6AN00152A.
Conrad AO, Bonello P. Application of infrared and Raman spectroscopy for the identification of disease resistant trees. Front Plant Sci. 2015;6:1152. https://doi.org/10.3389/fpls.2015.01152.
Rosch P, Harz M, Schmitt M, Peschke KD, Ronneberger O, Burkhardt H, et al. Chemotaxonomic identification of single bacteria by micro-Raman spectroscopy: application to clean-room-relevant biological contaminations. Appl Environ Microbiol. 2005;71:1626–37. https://doi.org/10.1128/AEM.71.3.1626-1637.2005.
Baritaux J-C, Simon A-C, Schultz E, Emain C, Laurent P, Dinten J-M. A study on identification of bacteria in environmental samples using single-cell Raman spectroscopy: feasibility and reference libraries. Environ Sci Pollut Res Int. 2016;23:8184–91. https://doi.org/10.1007/s11356-015-5953-x.
Harz A, Roesch P, Popp J. Vibrational spectroscopy—a powerful tool for the rapid identification of microbial cells at the single-cell level. Cytometry A. 2009;75A:104–13. https://doi.org/10.1002/cyto.a.20682.
Choo-Smith L-P, Maquelin K, van Vreeswijk T, Bruining HA, Puppels GJ, Thi NAN, et al. Investigating microbial (micro)colony heterogeneity by vibrational spectroscopy. Appl Environ Microbiol. 2001;67:1461–9. https://doi.org/10.1128/AEM.67.4.1461-1469.2001.
Liu T-T, Lin Y-H, Hung C-S, Liu T-J, Chen Y, Huang Y-C, et al. A high speed detection platform based on surface-enhanced Raman scattering for monitoring antibiotic-induced chemical changes in bacteria cell wall. PLoS One. 2009:4. https://doi.org/10.1371/journal.pone.0005470.
Assaf A, Cordella CBY, Thouand G. Raman spectroscopy applied to the horizontal methods ISO 6579:2002 to identify Salmonella spp. in the food industry. Anal Bioanal Chem. 2014;406:4899–910. https://doi.org/10.1007/s00216-014-7909-2.
Maquelin K, Choo-Smith LP, van Vreeswijk T, Endtz HP, Smith B, Bennett R, et al. Raman spectroscopic method for identification of clinically relevant microorganisms growing on solid culture medium. Anal Chem. 2000;72:12–9. https://doi.org/10.1021/ac991011h.
Meisel S, Stoeckel S, Elschner M, Roesch P, Popp J. Assessment of two isolation techniques for bacteria in milk towards their compatibility with Raman spectroscopy. Analyst. 2011;136:4997–5005. https://doi.org/10.1039/c1an15761b.
Hutsebaut D, Maquelin K, De Vos P, Vandenabeele P, Moens L, Puppels GJ. Effect of culture conditions on the achievable taxonomic resolution of Raman spectroscopy disclosed by three Bacillus species. Anal Chem. 2004;76:6274–81. https://doi.org/10.1021/ac049228l.
Mlynáriková K, Samek O, Bernatová S, Růžička F, Ježek J, Hároniková A, et al. Influence of culture media on microbial fingerprints using Raman spectroscopy. Sensors. 2015;15:29635–47. https://doi.org/10.3390/s151129635.
Marotta NE, Bottomley LA. Surface-enhanced Raman scattering of bacterial cell culture growth media. Appl Spectrosc. 2010;64:601–6.
Premasiri WR, Gebregziabher Y, Ziegler LD. On the difference between surface-enhanced Raman scattering (SERS) spectra of cell growth media and whole bacterial cells. Appl Spectrosc. 2011;65:493–9. https://doi.org/10.1366/10-06173.
International Organization for Standardization. ISO 11885:2009. Water quality—determination of selected elements by inductively coupled plasma optical emission spectrometry (ICP-OES). Geneva: ISO.
International Organization for Standardization. ISO 13903:2005. Animal feeding stuffs—determination of amino acids content. Geneva: ISO.
Srigley CT, Mossoba MM. Current analytical techniques for food lipids. In: Spizzirri UM, Cirillo G, editors. Food safety: innovative analytical tools for safety assessment. Hoboken: Wiley-Blackwell; 2016. p. 33–64.
Cordella CBY, Bertrand D. SAISIR: a new general chemometric toolbox. TrAC Trends Anal Chem. 2014;54:75–82. https://doi.org/10.1016/j.trac.2013.10.009.
Slutsky B, et al. Handbook of chemometrics and qualimetrics: part A by D. L. Massart, B. G. M. Vandeginste, L. M. C. Buydens, S. De Jong, P. J. Lewi, and J. Smeyers-Verbeke. Data handling in science and technology volume 20A. Elsevier: Amsterdam, 1997, xvii 867 pp. ISBN 0-444-89724-0. J Chem Inf Comput Sci. 1998;38:1254–4. https://doi.org/10.1021/ci980427d.
Cordella CBY. PCA: the basic building block of chemometrics. In: Krull IS, editor. Analytical chemistry. InTech; 2012. https://doi.org/10.5772/51429.
Valentine N, Wunschel S, Wunschel D, Petersen C, Wahl K. Effect of culture conditions on microorganism identification by matrix-assisted laser desorption ionization mass spectrometry. Appl Environ Microbiol. 2005;71:58–64. https://doi.org/10.1128/AEM.71.1.58-64.2005.
Acknowledgments
This study was supported by the OSEO-ISI research grant program and the AgriFoodGPS project. The authors would like to thank the European Community, the European Funds for Regional Development (FEDER), Eurofins Scientific, the Region of Pays de la Loire, the University of Nantes, and the Paris Institute for Life, Food and Environmental Sciences (AgroParisTech) for their contributions to this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest declared for any of the authors of this manuscript. No conflicts, informed consent, or human or animal rights were applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 335 kb)
Rights and permissions
About this article
Cite this article
Assaf, A., Grangé, E., Cordella, C.B.Y. et al. Evaluation of the impact of buffered peptone water composition on the discrimination between Salmonella enterica and Escherichia coli by Raman spectroscopy. Anal Bioanal Chem 412, 3595–3604 (2020). https://doi.org/10.1007/s00216-020-02596-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-020-02596-7