Abstract
Accurate measurement and understanding of therapeutic uptake and metabolism is key in the drug development process. This work examines the amount of doxorubicin that can penetrate into spheroids after being encapsulated in a liposomal configuration in comparison with free drug. Through a process known as serial trypsinization, three distinct cellular populations of a spheroid were successfully separated and a small molecule extraction was used to isolate the chemotherapeutic. Doxorubicin showed a time-dependent permeability into spheroids with the most drug accumulating in the core at 24 h of treatment. Entrapment of the chemotherapeutic delayed the permeability of the drug and resulted in reduced amounts quantified at the earlier time points. These findings validate the claim that liposomal therapeutics have the ability to alter the pharmacokinetics and pharmacodynamics profiles of a drug while also demonstrating the combined power of mass spectrometry and three-dimensional cell cultures to evaluate drug penetration and metabolism.

Graphical abstract
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lack of understanding regarding drug uptake and metabolism within tissues is one of the leading causes of failure during the drug development pipeline [1,2,3]. It is critical to understand the adsorption, distribution, metabolism, and excretion (ADME) of a putative therapeutic in the human body in order to develop one viable drug candidate to the Federal Drug Administration. To produce biologically relevant information from in vitro assays, it is necessary to manipulate cells in a manner that would most accurately represent cells in vivo [4]. Spatial analysis of novel therapeutics can unlock key information regarding drug distribution and metabolism as well as changes in the localization and relative abundance of proteins in response to treatment [5,6,7,8,9,10,11,12]. Spatial analysis can allow one to detect small changes over time and differences between the drug configurations.
Two-dimensional cell culture is conventionally used during drug development because of its ease and lower cost, relative to other model systems. However, these cultures lack many aspects of an in vivo tumor and do not accurately mimic what is found in a tumor cellular microenvironment. Recent studies have shown that cells grown in three-dimensions display vastly different chemical and cellular responses than cells grown in the traditional two-dimensional fashion due to the oxygen and nutrient gradients that form during development [13,14,15,16,17,18]. Spheroids, a type of three-dimensional cell culture, display pathophysiological gradients that are more complex than two-dimensional cell culture and can also be used in a high throughput manner. These cultures develop cellular populations that are comparable with in vivo tumors. These cellular populations form in response to decreasing nutrient and oxygen concentrations from the exterior of the spheroid to the necrotic core [14, 19,20,21]. These layers consist of an outer region of proliferating cells, a middle layer of quiescent cells, and a necrotic core [15, 17, 18]. These different cellular populations can be harvested by serial trypsinization.
Serial trypsinization is a process in which dilute trypsin is introduced to the spheroids for a short period of time before being quenched. Then the cell suspension is collected and spheroids are washed to allow further trypsin proteolysis [22,23,24,25]. Through this process, cells from progressively deeper regions in the spheroids can be collected.
Nanotechnology has been at the cutting edge of novel therapeutic development. Nanocarriers have the ability to positively alter pharmacokinetic profiles and have the capability for co-encapsulation of therapeutics [26]. Nano-drug delivery vehicles include polymeric nanoparticles, micelles, dendrimers, carbon nanotubes, and liposomes [27]. Since 2017, there have been 50 different nano-drug delivery vehicles approved by the Federal Drug Administration (FDA) [28]. It has been suggested that nano-drugs may show tumor specificity due to the enhanced permeability and retention (EPR) effect [29, 30]. Briefly, in tumor tissues, leaky blood vessels allow nanocarriers to penetrate and preferentially accumulate in the tumorous tissue [31]. However, reproducing the EPR effect in an in vitro system remains a challenge [32, 33]. Spheroids can provide a first step in therapeutic testing but additional testing must be done prior to the clinical setting as drugs inherently behave differently in vivo. It has also been shown that the EPR effect occurs differently between murine and human model systems so it is critical to understand the tumor microenvironment to produce a viable novel therapeutic [34,35,36].
To date, the liposomal encapsulation of therapeutics is considered the most successful nano-drug delivery vehicle [37]. Liposomes are spherical vesicles made up of lipids and possess several advantages over other drug delivery vehicles. They have been shown to have low toxicity and high biocompatibility due to the fact that they are composed of naturally occurring phospholipids [24, 25, 38]. Additionally, liposomes have the ability for self-assembly and depending on their size can deliver a high concentration of the entrapped contents directly to a site of interest while avoiding elimination by the body [23]. Therefore, liposomes have the potential to facilitate greater drug penetration into tissue samples and could theoretically make treatments more efficient. Studies have also shown that liposomal drugs have profound effects on the pharmacokinetic and pharmacodynamics profiles of the entrapped contents [39].
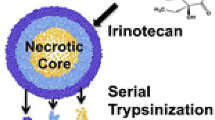
In this work, we aimed to quantitatively characterize the amount of doxorubicin in the three distinct populations of cells present in our spheroid model system. Our laboratory has previously developed a liposomal doxorubicin configuration [8]. This liposomal configuration of doxorubicin was used to dose the spheroids for a range between 2 and 72 h. A separate group of spheroids was also dosed with free doxorubicin for the same time course. The distinct populations of cells within the spheroid were isolated through serial trypsinization. Figure 1 shows the experimental schematic.
Schematic of experimental workflow. Spheroids are dosed with one of the three forms of doxorubicin for a range of 2–72 h. Serial trypsinization is then performed to isolate the different cellular populations. A small molecule extraction is done to remove the drug molecule from the cells for analysis
As liposomal drugs have been shown to alter the pharmacokinetics and pharmacodynamics profile of the entrapped therapeutic, we expected to see varying amounts of doxorubicin present with our spheroid system. By performing the serial trypsinization, we spatially quantified the amount of the chemotherapeutic and its metabolites in the different regions of the spheroid and inferred the rate in which it enters the cells.
Materials and methods
Chemicals and materials
HPLC grade acetonitrile (ACN) and HPLC grade water were purchased from B & J Brand (Honeywell Burdick & Jackson, Muskegon, MI). Chloroform and Tris were all obtained from Fisher Scientific (Fair Lawn, NJ).
Cell culture and spheroid formation
The colon carcinoma cell line HCT 116 was purchased from ATCC (Manassas, VA) and maintained in McCoy’s 5A cell culture media (Life Technologies, Grand Island, NY) supplemented with 10% fetal bovine serum (Thermo Scientific, Gaithersburg, MD) and 1% L-glutamine (Invitrogen, San Diego, CA). Cells were grown in 5% CO2 at 37 °C and passed every 4 days. Cell lines were used within 3 months after resuscitation of frozen aliquots thawed from liquid nitrogen. The cells were verified by short tandem repeat (STR) sequencing in 2016.
Spheroids were prepared in an agarose-coated 96-well plate as previously described [40,41,42]. Cells were seeded into each well at a density of 7000 cells/well, incubated at 37 °C and 50% of the volume of the cell culture media was changed every 48 h after 4 days in culture. After 13 days in culture, spheroids reach their maximum growth and were dosed with liposomal doxorubicin, free doxorubicin, or control media for 2, 12, 24, 48, or 72 h. Liposomal doxorubicin was created as previously described [8]. Spheroids were dosed with either 30 μM free doxorubicin or liposomal doxorubicin for 2, 6, 12, 24, 48, and 72 h.
Serial trypsinization
Cells from spheroids were separated using serial trypsinization to collect the three distinct physiological populations. Spheroids were washed in 1× PBS briefly to wash away media in a cell culture dish. A solution of 0.05% trypsin/EDTA at room temperature was added, and the spheroids were gently rotated for 3 min on a rotary shaker at 90 rpm. Chilled complete medium was then added to inhibit the trypsin and was gently rotated for 3 min. The cell suspension was carefully collected from the spheroids, and medium without FBS was added to wash serum out of the dish (3-min rotation) to allow further trypsin proteolysis. By repeating the trypsinization, complete medium washing, and serum-free medium washing steps repeatedly, cells from progressively deeper layers in the spheroid were harvested. The first three fractions were combined as a single sample representing the outer proliferative cells of the spheroid. Cells from the next three trypsin treatments were collected representing the intermediate region. The seventh fraction contained the remaining spheroid necrotic core, which was dissociated completely by pipetting. Cell suspensions from each of the populations were centrifuged at 2000g for 10 min and cell pellets were washed twice with water. Cell pellets were stored at − 80 °C for small molecule extraction. Forty spheroids were used for each condition at each time point [20].
Small molecule extraction
Small molecules were extracted by adding 500 μL of an extraction solution (0.01 M HCl:methanol, 2:3, v/v) containing 200 nM daunorubicin. After mixing by vortex, the cellular extracts were sonicated and centrifuged at 15,000g for 15 min at 4 °C. The supernatants were transferred into clean tubes and evaporated to dryness under vacuum using a SpeedVac. The dried extracts were redissolved by vortexing in 100 μL of 20:80 (v/v) methanol:water solution [20]. To remove protein contaminants, samples were centrifuged with Amicon Ultra-0.5 Filter Devices with a 30,000 Da molecular weight cutoff. Flow-through was collected and 2 μL of sample was injected into the LC-MS/MS for quantitation.
Calibration curve
A calibration curve of doxorubicin containing 200 nM daunorubicin (internal standard) was constructed. 2.5 nM, 25 nM, 50 nM, 100 nM, 250 nM, 300 nM, 500 nM, and 1 μM concentration solutions of doxorubicin were used to construct the curve.
Mass spectrometer parameters
A nanoAcquity ultra performance LC system (2 × 100 mm, 2.7 μM particle size C18 column) (Thermo Fisher Scientific, Milford, MA, USA) was used for separation of the small molecules. Samples were eluted with a binary solvent system with 0.1% formic acid (A) and 0.1% formic acid in methanol (B) using the following linear gradient separation: buffer B was increased from 15 to 100% in 20 min, washed with 100% B for 3 min, and equilibrated for 7 min with 15% buffer B before the next injection. The eluted small molecules were analyzed using a Thermo Quantiva Triple Quadrupole HPLC MS/MS system (Thermo Fisher Scientific, Waltham, MA, USA). Instrument parameters were set as follows: H-ESI source, positive ion voltage, 4500 V, spray voltage was static, sheath gas, 10 Arb, auxiliary gas, 5 Arb, sweep gas, 5Arb, ion transfer tube temperature 250 °C, vaporizer temperature 100 °C. Transitions were made for doxorubicin, daunorubicin (internal standard), doxorubicinone, 7-deoxydoxorubicinone, and 7-deoxydoxrubicinolone (Table 1). Structures of these compounds can be seen in the Electronic Supplementary Material (ESM). Q1 and Q3 resolution (FWHM) was set to 2, CID gas was 1.5 mTorr. Samples were run in duplicate injections. The data was analyzed and quantified by Xcalibur software (Thermo Fisher Scientific, Waltham, MA, USA).
Results and discussion
Spheroids serve as an effective in vitro platform for testing drug delivery and metabolism. As nano-drug carriers continue to increase in popularity, it is important to quantitatively analyze how much drug is reaching the intended target. In this work, we aimed to quantitatively characterize the amount of doxorubicin delivered in two different ways to the different cellular populations present in our spheroid model system. Doxorubicin was delivered in its free drug form as well as a liposomal configuration. As liposomal drugs have been shown to alter the pharmacokinetics and pharmacodynamics profile of an entrapped therapeutic, we expected to see observe differential gradients of doxorubicin present with our spheroid system. To obtain more spatial information as to where the doxorubicin was located in our system, serial trypsinization was performed to isolate the distinct cellular populations of a spheroid, outer proliferative cells, quiescent cells in the middle, and necrotic cells in the core. A small molecule extraction was then performed to extract the doxorubicin as well as its metabolites from the cells.
Quantitation of doxorubicin in spheroids
Data was processed and peak area ratios of doxorubicin compared with daunorubicin were calculated by using Quan Browser of the Xcalibur software. Based on the slope and intercept of the doxorubicin calibration curve, the concentration of doxorubicin extracted from cells originating from the outer proliferative region, the quiescent middle, and necrotic core was calculated. Exact amounts of the drug in each region were then determined based on the total volume of each sample (100 μL). Figure 2 shows the amount of doxorubicin that was detected in each of the cellular populations for six different time points for the two delivery methods. As expected, the total amount of doxorubicin present in the spheroid increases with the drug incubation time.
In the first 6 h of treatment, doxorubicin penetrates into the outer layer of cells with some drug also detected in the middle quiescent cells. Doxorubicin was detected at minimal levels in the core of the spheroids in both treatment conditions after 6 h of treatment. Differences between the two drug delivery methods become noticeable after 6 h of treatment. The results observed after 6 h are statistically significantly between the free doxorubicin and the liposomal doxorubicin (p value < 0.05). Free doxorubicin is detected at much larger levels in the spheroids after 12 h of treatment when compared with the liposomal doxorubicin penetration. On average, after 12 h of treatment, free doxorubicin is quantified at 37 times greater concentration than the liposomal doxorubicin (7.45 μM vs. 0.20 μM) in all three cellular populations combined. As the time course progresses, more doxorubicin penetrated in the spheroid from the liposomal configuration; however, free doxorubicin was always detected at a higher concentration in the layers.
After 72 h of treatment, the concentration of free doxorubicin detected is still 2.9 times greater than that of the liposomal doxorubicin (25.64 μM vs. 8.92 μM) in all cellular layers combined. This result was an expected outcome as liposomal therapeutics are known to alter the pharmacokinetics and pharmacodynamics profiles of a drug.
One of the substantial benefits of liposomal drug delivery is prolonged exposure on the entrapped therapeutic. The trend in this data shows this phenomenon exactly. By the 72-h time point, the free drug concentrations are almost at its maximum capacity and the spheroids were dosed at 30 μM for both conditions. However, the liposomal configuration had not yet reached its dosage capacity so a prolonged therapeutic effect may have occurred if this time course would have continued. This prolonged therapeutic effect is due to the fact the doxorubicin must first be released from the liposome through the lipid bilayer components binding to cellular bilayers before it has the ability to passively diffuse into cells. The doxorubicin entrapped in the liposome is essentially inactive until it is released.
Metabolite detection
Metabolism directly influences the amount of parent drug quantified in the cells. Doxorubicin is part of the anthracycline family and there are two main metabolic routes for anthracycline metabolism: electron reduction and deglycosidation [43]. It is believed that electron reduction is carried out by several oxidoreductases while the enzymes and candidate genes for deglycosidation metabolism are less well characterized. It is thought that deglycosidation accounts for 1–3% of doxorubicin metabolism [30]. Recent studies have implicated NADPH in this metabolism mechanism [44]. However, a large proportion of doxorubicin (~ 50%) is unchanged as it is eliminated from the body [44, 45].
In a previous MALDI MSI study in our laboratory, we observed three doxorubicin metabolites after liposomal entrapment of the chemotherapeutic and treatment in spheroids: doxorubicinone, 7-deoxydoxorubicinone, and 7-deoxydoxorubicinolone [8]. These metabolites were quantified with the same approach as doxorubicin. Figure 3 shows the quantitation of doxorubicinone. This metabolite was detected after 6 h of treatment in the outer and middle cells of spheroids when dosed with free doxorubicin. Similar to the trend of the parent drug, doxorubicinone is always found at a higher concentration in the free drug dosed spheroid in comparison with the liposomal-treated ones, but by the 72-h time point, the gap between the samples has drastically closed. The concentration of the free doxorubicin dosed spheroids was determined to be 1.7 times greater than that of the PC liposome doxorubicin (3.5 μM vs. 2.1 μM) at this last time point. However, the difference between the two different dosing schemes is still statistically significant at the 72-h time point (p value < 0.05).
Doxorubicinone quantified in spheroids that had been serial trypsinized. This metabolite was first detected at the 6-h time point. Total amount of doxorubicinone detected increases over the time course. Doxorubicinone metabolized from free doxorubicin (yellow) is seen at higher concentrations at all time points in comparison with liposomal doxorubicin (green)
7-Dexoydoxorubicinone was quantified within this experiment (Fig. 4). Following the trend of doxorubicin and doxorubicinone, this metabolite was also first detected in free drug dosed spheroids. This metabolite does not appear until 24 h of treatment and then constant levels are maintained for the rest of the time course. 7-Dexoydoxorubicinone could be quantified in the liposomal configurations of doxorubicin, however, not until 72 h of treatment for liposomal doxorubicin. The detected amount of this metabolite is constant for both drug delivery types and is much lower than what was detected for doxorubicin and doxorubicinone. 7-Deoxydoxorubicinolone was not able to be quantified in this experiment due to the levels of this metabolite being below the level of detection of the instrument.
These detected metabolism products match previous results by the Arriaga group [46]. The metabolism pathway is depicted in Fig. 5. Using laser-induced fluorescence mass spectrometry (LIF-MS) and micellar electrokinetic chromatography laser-induced fluorescence (MEKC-LIF), they determined that doxorubicinone was the most abundant metabolite detected, validating the results in this present study. However, doxorubicinone forms as a result of an acid-catalyzed cleavage of the ether bond over the sugar ring during sample preparation not because of metabolism within the cells. This acid-catalyzed cleavage explains why this is the predominate metabolite detected in both studies. This study as well as the study performed by the Arriaga group detected 7-deoxydoxorubicinone as a main metabolite formed. Both studies detected levels of 7-deoxydoxorubicinone at constant levels after an initial incubation period with doxorubicin. 7-Deoxydoxorubicinolone and doxol were not detected in this present study.
In addition, the Arriaga group observed 7-deoxydoxorubicinolone using MEKC-LIF but not using an HPLC approach. MEKC-LIF is capable of analyzing small volumes of sample and has high separation efficiency providing the resolution needed in order to deal with high sample complexity [46]. Doxol was also not detected as a metabolite which matches previous studies that indicate hepatocytes and liver enzymes necessary for this conversion, which are not present in the spheroid model system [47, 48].
Conclusions
By utilizing spheroids, serial trypsinization, and mass spectrometry, one can use this methodology for an accurate evaluation of drug delivery and metabolism in an in vitro system. This is a quantitative approach that can be easily used to spatially locate therapeutics and the metabolites created. By looking at different delivery configurations for doxorubicin, we quantified the spatially specific differences that correlate to different cellular populations in drug concentrations. We observed that the liposomal configurations altered the pharmacokinetic and pharmacodynamics profile of doxorubicin for the distinct populations of cells. Delayed permeation of the drug and the appearance of metabolites were quantified from 24 to 72 h. The production of metabolites between free drug and the liposomal configuration was greatly diminished by the 72-h time point. Accurate assessment of drug concentrations is key to understanding the effectiveness and can potentially point towards areas of improvement for future novel therapeutics.
References
Trédan O, Galmarini CM, Patel K, Tannock IF. Drug resistance and the solid tumor microenvironment. J Natl Cancer Inst. 2007;99:1441–54. https://doi.org/10.1093/jnci/djm135.
Lammers T, Kiessling F, Hennink WE, Storm G. Drug targeting to tumors: principles, pitfalls and (pre-) clinical progress. J Control Release. 2012;161:175–87. https://doi.org/10.1016/j.jconrel.2011.09.063.
Mehta G, Hsiao AY, Ingram M, Luker GD, Takayama S. Opportunities and challenges for use of tumor spheroids as models to test drug delivery and efficacy. J Control Release. 2012;164:192–204. https://doi.org/10.1016/j.jconrel.2012.04.045.
Pampaloni F, Reynaud EG, Stelzer EHK. The third dimension bridges the gap between cell culture and live tissue. Nat Rev Mol Cell Biol. 2007;8:839–45. https://doi.org/10.1038/nrm2236.
LaBonia GJ, Ludwig KR, Mousseau CB, Hummon AB. iTRAQ quantitative proteomic profiling and MALDI–MSI of colon cancer spheroids treated with combination chemotherapies in a 3D printed fluidic device. Anal Chem. 2018;90:1423–30. https://doi.org/10.1021/acs.analchem.7b04969.
Schroll MM, LaBonia GJ, Ludwig KR, Hummon AB. Glucose restriction combined with autophagy inhibition and chemotherapy in HCT 116 spheroids decreases cell clonogenicity and viability regulated by tumor suppressor genes. J Proteome Res. 2017;16:3009–18. https://doi.org/10.1021/acs.jproteome.7b00293.
Schroll M, Ludwig K, Bauer K, Hummon A. Calcitriol supplementation causes decreases in tumorigenic proteins and different proteomic and metabolomic signatures in right versus left-sided colon cancer. Metabolites. 2018;8:5. https://doi.org/10.3390/metabo8010005.
Lukowski JK, Weaver EM, Hummon AB. Analyzing liposomal drug delivery systems in three-dimensional cell culture models using MALDI imaging mass spectrometry. Anal Chem. 2017;89:8453–8. https://doi.org/10.1021/acs.analchem.7b02006.
Schroll MM, Liu X, Herzog SK, Skube SB, Hummon AB. Nutrient restriction of glucose or serum results in similar proteomic expression changes in 3D colon cancer cell cultures. Nutr Res. 2016;36:1068–80. https://doi.org/10.1016/j.nutres.2016.08.002.
Feist PE, Sidoli S, Liu X, Schroll MM, Rahmy S, Fujiwara R, et al. Multicellular tumor spheroids combined with mass spectrometric histone analysis to evaluate epigenetic drugs. Anal Chem. 2017;89:2773–81. https://doi.org/10.1021/acs.analchem.6b03602.
Liu X, Hummon AB. Chemical imaging of platinum-based drugs and their metabolites. Sci Rep. 2016;6:38507. https://doi.org/10.1038/srep38507.
Liu X, Flinders C, Mumenthaler SM, Hummon AB. MALDI mass spectrometry imaging for evaluation of therapeutics in colorectal tumor organoids. J Am Soc Mass Spectrom. 2018;29:516–26. https://doi.org/10.1007/s13361-017-1851-4.
Yue X, Lukowski JK, Weaver EM, Skube SB, Hummon AB. Quantitative proteomic and phosphoproteomic comparison of 2D and 3D colon cancer cell culture models. J Proteome Res. 2016;15:4265–76. https://doi.org/10.1021/acs.jproteome.6b00342.
Sutherland RM, Sordat B, Bamat J, Gabbert H, Bourrat B, Mueller-Klieser W. Oxygenation and differentiation in multicellular spheroids of human colon carcinoma. Cancer Res. 1986;46:5320–9.
Freyer JP, Sutherland RM. Selective dissociation and characterization of cells from different regions of multicell tumor spheroids. Cancer Res. 1980;40:3956–65.
Sutherland RM. Cell and environment interactions in tumor microregions: the multicell spheroid model. Science. 1988;240:177–84.
Weigelt B, Ghajar CM, Bissell MJ. The need for complex 3D culture models to unravel novel pathways and identify accurate biomarkers in breast cancer. Adv Drug Deliv Rev. 2014;69–70:42–51. https://doi.org/10.1016/j.addr.2014.01.001.
Vidi P-A, Bissell MJ, Lelièvre SA. Three-dimensional culture of human breast epithelial cells: the how and the why. Methods Mol Biol. 2013;945:193–219. https://doi.org/10.1007/978-1-62703-125-7_13.
Hirschhaeuser F, Menne H, Dittfeld C, West J, Mueller-Klieser W, Kunz-Schughart LA. Multicellular tumor spheroids: an underestimated tool is catching up again. J Biotechnol. 2010;148:3–15. https://doi.org/10.1016/j.jbiotec.2010.01.012.
Liu X, Hummon AB. Quantitative determination of irinotecan and the metabolite SN-38 by nanoflow liquid chromatography-tandem mass spectrometry in different regions of multicellular tumor spheroids. J Am Soc Mass Spectrom. 2015;26:577–86. https://doi.org/10.1007/s13361-014-1071-0.
Liu X, Weaver EM, Hummon AB. Evaluation of therapeutics in three-dimensional cell culture systems by MALDI imaging mass spectrometry. Anal Chem. 2013;85:6295–302. https://doi.org/10.1021/ac400519c.
Allen TM, Cullis PR. Liposomal drug delivery systems: from concept to clinical applications. Adv Drug Deliv Rev. 2013;65:36–48. https://doi.org/10.1016/j.addr.2012.09.037.
Lombardo D, Calandra P, Barreca D, Magazù S, Kiselev M. Soft interaction in liposome nanocarriers for therapeutic drug delivery. Nanomaterials. 2016;6:125. https://doi.org/10.3390/nano6070125.
Sercombe L, Veerati T, Moheimani F, Wu SY, Sood AK, Hua S. Advances and challenges of liposome assisted drug delivery. Front Pharmacol. 2015. https://doi.org/10.3389/fphar.2015.00286.
Nagayasu A, Uchiyama K, Kiwada H. The size of liposomes: a factor which affects their targeting efficiency to tumors and therapeutic activity of liposomal antitumor drugs. Adv Drug Deliv Rev. 1999;40:75–87. https://doi.org/10.1016/S0169-409X(99)00041-1.
Chowdhury S, Yusof F, Salim WWAW, Sulaiman N, Faruck MO. An overview of drug delivery vehicles for cancer treatment: nanocarriers and nanoparticles including photovoltaic nanoparticles. J Photochem Photobiol B Biol. 2016;164:151–9. https://doi.org/10.1016/j.jphotobiol.2016.09.013.
Felice B, Prabhakaran MP, Rodríguez AP, Ramakrishna S. Drug delivery vehicles on a nano-engineering perspective. Mater Sci Eng C. 2014;41:178–95. https://doi.org/10.1016/j.msec.2014.04.049.
Ventola CL. Progress in nanomedicine: approved and investigational nanodrugs. P T. 2017;42:742–55.
Tekade RK, Maheshwari R, Soni N, Tekade M, Chougule MB. Nanotechnology for the development of nanomedicine. In: Nanotechnology-based approaches target. Deliv. Drugs Genes. Elsevier, pp 3–61, 2017.
van Vlerken LE, Duan Z, Little SR, Seiden MV, Amiji MM. Biodistribution and pharmacokinetic analysis of paclitaxel and ceramide administered in multifunctional polymer-blend nanoparticles in drug resistant breast cancer model. Mol Pharm. 2008;5:516–26. https://doi.org/10.1021/mp800030k.
Danhier F. To exploit the tumor microenvironment: since the EPR effect fails in the clinic, what is the future of nanomedicine? J Control Release. 2016;244:108–21. https://doi.org/10.1016/j.jconrel.2016.11.015.
Danhier F, Feron O, Préat V. To exploit the tumor microenvironment: passive and active tumor targeting of nanocarriers for anti-cancer drug delivery. J Control Release. 2010;148:135–46. https://doi.org/10.1016/j.jconrel.2010.08.027.
Huang X, Peng X, Wang Y, Wang Y, Shin DM, El-Sayed MA, et al. A reexamination of active and passive tumor targeting by using rod-shaped gold nanocrystals and covalently conjugated peptide ligands. ACS Nano. 2010;4:5887–96. https://doi.org/10.1021/nn102055s.
Tang Y, Soroush F, Sheffield JB, Wang B, Prabhakarpandian B, Kiani MF. A biomimetic microfluidic tumor microenvironment platform mimicking the EPR effect for rapid screening of drug delivery systems. Sci Rep. 2017;7:9359. https://doi.org/10.1038/s41598-017-09815-9.
Seok J, Warren HS, Cuenca AG, Mindrinos MN, Baker HV, Xu W, et al. Genomic responses in mouse models poorly mimic human inflammatory diseases. Proc Natl Acad Sci. 2013;110:3507–12. https://doi.org/10.1073/pnas.1222878110.
Prabhakarpandian B, Shen M-C, Nichols JB, Garson CJ, Mills IR, Matar MM, et al. Synthetic tumor networks for screening drug delivery systems. J Control Release. 2015;201:49–55. https://doi.org/10.1016/j.jconrel.2015.01.018.
Farokhzad OC, Langer R. Impact of nanotechnology on drug delivery. ACS Nano. 2009;3:16–20. https://doi.org/10.1021/nn900002m.
Lu H, Utama RH, Kitiyotsawat U, Babiuch K, Jiang Y, Stenzel MH. Enhanced transcellular penetration and drug delivery by crosslinked polymeric micelles into pancreatic multicellular tumor spheroids. Biomater Sci. 2015;3:1085–95. https://doi.org/10.1039/C4BM00323C.
Liu Z, Bi Y, Sun Y, Hao F, Lu J, Meng Q, et al. Pharmacokinetics of a liposomal formulation of doxorubicin in rats. Saudi Pharm J. 2017;25:531–6. https://doi.org/10.1016/j.jsps.2017.04.019.
Friedrich J, Seidel C, Ebner R, Kunz-Schughart LA. Spheroid-based drug screen: considerations and practical approach. Nat Protoc. 2009;4:309–24. https://doi.org/10.1038/nprot.2008.226.
Ahlf Wheatcraft DR, Liu X, Hummon AB. Sample preparation strategies for mass spectrometry imaging of 3D cell culture models. J Vis Exp. 2014. https://doi.org/10.3791/52313.
Li H, Hummon AB. Imaging mass spectrometry of three-dimensional cell culture systems. Anal Chem. 2011;83:8794–801. https://doi.org/10.1021/ac202356g.
Thorn CF, Oshiro C, Marsh S, Hernandez-Boussard T, McLeod H, Klein TE, et al. Doxorubicin pathways: pharmacodynamics and adverse effects. Pharmacogenet Genomics. 2011;21:440–6. https://doi.org/10.1097/FPC.0b013e32833ffb56.
Mordente A, Meucci E, Silvestrini A, Martorana GE, Giardina B. New developments in anthracycline-induced cardiotoxicity. Curr Med Chem. 2009;16:1656–72.
Minotti G. Reactions of adriamycin with microsomal iron and lipids. Free Radic Res Commun. 1989;7:143–8.
Wang Y, Katzenmeyer JB, Arriaga EA. Combination of micellar electrokinetic and high-performance liquid chromatographies to assess age-related changes in the in vitro metabolism of Fischer 344 rat liver. J Gerontol Ser A. 2011;66A:935–43. https://doi.org/10.1093/gerona/glr074.
Vrignaud P, Londos-Gagliardi D, Robert J. Hepatic metabolism of doxorubicin in mice and rats. Eur J Drug Metab Pharmacokinet. 11:101–5. https://doi.org/10.1007/BF03189834.
Pollak N, Dölle C, Ziegler M. The power to reduce: pyridine nucleotides--small molecules with a multitude of functions. Biochem J. 2007;402:205–18. https://doi.org/10.1042/BJ20061638.
Acknowledgments
The authors thank Matt Bernier and the Campus Chemical Instrument Center at Ohio State University for help with the analysis of the samples. A.H. and J.L. were supported by the National Science Foundation (CAREER Award, No. CHE-1351595).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 631 kb)
Rights and permissions
About this article
Cite this article
Lukowski, J.K., Hummon, A.B. Quantitative evaluation of liposomal doxorubicin and its metabolites in spheroids. Anal Bioanal Chem 411, 7087–7094 (2019). https://doi.org/10.1007/s00216-019-02084-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-02084-7