Abstract
Since its introduction in the 1980s, matrix-assisted laser desorption/ionization mass spectrometry (MALDI MS) has gained a prominent role in the analysis of high molecular weight biomolecules such as proteins, peptides, oligonucleotides, and polysaccharides. Its application to low molecular weight compounds has remained for long time challenging due to the spectral interferences produced by conventional organic matrices in the low m/z window. To overcome this problem, specific sample preparation such as analyte/matrix derivatization, addition of dopants, or sophisticated deposition technique especially useful for imaging experiments, have been proposed. Alternative approaches based on second generation (rationally designed) organic matrices, ionic liquids, and inorganic matrices, including metallic nanoparticles, have been the object of intense and continuous research efforts. Definite evidences are now provided that MALDI MS represents a powerful and invaluable analytical tool also for small molecules, including their quantification, thus opening new, exciting applications in metabolomics and imaging mass spectrometry. This review is intended to offer a concise critical overview of the most recent achievements about MALDI matrices capable of specifically address the challenging issue of small molecules analysis.

An ideal Book of matrices for MALDI MS of small molecules
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Although matrix-assisted laser desorption/ionization mass spectrometry (MALDI MS) [1] has revolutionized the analysis of large, nonvolatile, and labile molecules (such as proteins, peptides, oligonucleotides, and polysaccharides), its application to low-molecular weight compounds (LMWC) (otherwise denoted as “small molecules”) has been regarded, for long time, not appropriate or even useless. One of the main reasons was certainly represented by the large background generated by matrix related ions in the m/z range of interest for small molecules (vide infra). A second reason was the concurrent explosive growth of electrospray ionization (ESI) source [2], easily interfaced to liquid chromatography (LC), which has made LC-ESI-MS one of the most versatile and powerful analytical tools presently available. However, specific advantages of MALDI (e.g., high tolerance towards contaminants and buffers, uncomplicated spectra due to the almost exclusive generation of single charged ions, typical sub-femtomole sensitivity, and high throughput analysis) coupled to the most recent improvements in related instrumentations, has provided strong foundations to reassess the role this technique could play in the LMWC scenario. Indeed, the application of MALDI MS to small molecules is continuously growing, driven by the resurgence of interest in metabolomics studies and, more recently, by the overwhelming growth of mass spectrometry imaging (MSI) techniques that make the LMWC scenario absolutely relevant. The successful application of MALDI MS is strictly dependent on the choice of the matrix that should meet specific requirements, including analyte incorporation into an excess of matrix [3], high absorption at the irradiation laser wavelength (via electronic or vibrational excitation), efficient analyte ionization, and null or minimal and controllable analyte fragmentation. Commonly used matrices typically feature linear conjugated π systems and/or aromatic rings for photons absorption in the UV region and acid functional groups for proton transfer to analyte. In general, derivatives of benzoic acid, cinnamic acid, and related aromatic compounds are recognized as elective matrices for proteins and peptides [4], while picolinic and succinic acids have been used as matrices for the MALDI MS analysis of oligonucleotides [5].
Since the ‘first generation’ of MALDI matrices (mostly selected on largely empirical criteria) are, themselves, small organic molecules, having molecular weights typically below 300 Da, it is immediately evident why most of them could have been considered, in the recent past, unsuitable for MALDI analysis of LMW analytes. Considering, for instance, the archetype matrix 4-hydroxy-α-cyanocinnamic acid (CHCA), a plethora of matrix-related ions are generated during laser irradiation. Apart from the matrix (M) protonated adduct [M+H]+ at m/z 190.0499 and main fragment ions at m/z 172.0393 ([M-H2O+H]+) and m/z 146.0600 ([M-CO2+H]+), sodiated and potassiated adducts are also observed together with several matrix cluster ions covering the range from m/z 379.0925 ([2M+H]+) up to m/z 1287.0864 ([5M+4K-3H]+) [6]. It is also worth noting that the shift of the laser wavelength from 337 nm (N2 laser) to 355 nm, provided by the recently introduced Nd:YAG solid-state lasers, results in even more intense CHCA matrix clusters. Although specific pretreatments of the applied samples (e.g., washing of previously crystallized sample spots with ammonium-containing buffers) could significantly reduce the spectral abundance of matrix clusters, the matrix-related background ions and the low accuracy/resolution of the old generation MALDI instruments no doubt have discouraged the use of conventional matrices for LMWC analysis. Apart from significant instrumental improvements, several efforts have been made and several approaches have been proposed to partially suppress or even avoid matrix-related peaks such as, for instance, the use of matrix additives [7,8,9], and the synthesis of novel organic and inorganic matrices [10,11,12,13,14]. A separate mention is deserved for matrix-less approaches such as desorption ionization on silicon (DIOS), pioneered by Siuzdak’s group, which opened new horizons for small molecules analysis by laser desorption/ionization (LDI) MS [15].
Different deposition procedures of conventional matrices have also been exploited to boost MALDI MSI of LMW peptides, lipids, drugs and metabolites in tissues, cells, and microorganisms. Recent advances in deposition strategies together with development of high spatial resolution, high sensitivity MS instruments, and novel matrices for MALDI MSI have been reviewed elsewhere [16,17,18,19,20]. Some recent “fundamental books” focusing on basic principles, instrumentation, methods, and applications of MALDI MS, to which the interested reader should refer, remain an invaluable source of information [21,22,23,24,25]. The mechanisms of ion formation in MALDI MS, still a matter of debate, has been the object of several dedicated reviews [26,27,28,29,30,31,32] that need to be continuously updated as new insights in this topic are gathered [33,34,35,36,37].
The present review aims to provide a critical overview of the most recent achievements in the specific field of MALDI MS analysis of LMW analytes. The first section deals with the applications of classic MALDI matrices to LMWC through strategies such as the use of matrix additives, specific sample pretreatments, and matrix/analyte derivatization. New matrix deposition strategies, even if specifically devised for MALDI MSI experiments, are also briefly described since target analytes are typically LMW primary and secondary metabolites. The second (main) section deals with second generation organic matrices and is structured into sub-sections, each focusing on a class/function of emerging matrices for LMW analytes, namely: reactive matrices; matrices for negative ion mode; binary, hybrid (organic-inorganic) and nanomaterial based matrices; ionic liquids and electron transfer matrices.
First generation matrices for LMWC analysis
Representative examples of first generation matrices for UV MALDI (see Fig. 1) are benzoic acid derivatives (e.g., DHB or 2,5-dihydroxybenzoic acid), cinnamic acid derivatives (e.g., CHCA), ferulic acid, sinapinic acid (SA), and 2,4,6-trihydroxyacetophenone (THAP). Interestingly, some of these matrices, originally developed for the analysis of proteins, peptides, and oligonucleotides, are “lucky survivors” even for LMWC (despite the presence of potentially interfering matrix-related ions) mostly because of the introduction of last generation MALDI instruments. Indeed, as first demonstrated by Persike et al. [38], matrix signals can be conveniently used for the internal calibration, thus significantly improving mass accuracy (e.g., <5 ppm for acetylcholine). As a result of accurate mass measurement and high resolution offered by modern time-of-flight (TOF) instruments, the CHCA fragment ion [M+H-CO2]+ at m/z 146.061 could be resolved from the protonated acetylcholine (ACh) ion ([C7NH16O2]+) at m/z 146.117. ACh and Ch were successfully determined in mouse brain microdialysis samples. Persike and Karas [39] further confirmed that the sensitivity, resolution, and mass accuracy of a last generation MALDI-TOF MS system permit to overcome the limitations posed by the matrix (CHCA) background demonstrating the simultaneous quantification of 10 phenothiazines in spiked plasma samples. An atmospheric pressure MALDI high-resolution mass spectrometry (quadrupole/orbital trap hybrid analyzer) method has been described [40] for the simultaneous detection/quantitation of 26 triazines and triazoles in grapes using CHCA as matrix. MS and MS/MS spectra were acquired at a resolution of 35,000 FWHM (full width at half-maximum) and mass accuracy was typically within 5 ppm. The protonated adduct of atrazine-desethyldesisopropyl and cyprazine could be detected at m/z 146.0228 and 228.1011, respectively, in spite of the presence of matrix ions [M+H-CO2]+ and [M+K]+ with monoisotopic mass 146.0600 and 228.0058, respectively.
Among first generation matrices, DHB seems to be the most appropriate for lipid analysis [41] since, contrary to cinnamic and sinapinic acid, it gives a low yield of photochemically generated matrix ions. Porcari et al. [42] reported the characterization of lipid extracts of Atlantic sturgeon caviar using a DHB matrix in combination with a high resolution mass spectrometer. Yet, unless some special precautions are taken, for many other LMWC these conventional matrices do not work as successfully as in the previous examples owing to matrix effects, signal interference, or suppression of the analyte signal.
Matrix additives/dopants, supported matrices, and new deposition strategies
One of the simplest way to minimize or even remove the matrix background spectral interferences is to exploit the well-known but often overlooked matrix suppression effect (MSE) as suggested by McCombie and Knochenmuss [43]. Since laser intensity and the matrix/analyte molar ratio are the most easily governable factors strongly influencing MSE, laser power and analyte concentration were systematically varied for many LMW (from 150 to 550 Da) test analytes using two conventional matrices (DHB and CHCA) and finally evaluating a properly defined MSE score. Since high values of analyte-to-matrix molar ratios are required for MSE to be effective, an inconvenience that can be encountered (when more than one analyte is present) is the simultaneous occurrence of analyte suppression effects (ASE).
Matrix background could be suppressed more easily in positive ion mode by adding CHCA with a quaternary ammonium (QA) surfactant [44, 45] such as cetyltrimethylammonium bromide (CTAB) as first demonstrated by Guo et al. [46], who also introduced the acronym “matrix suppressed laser desorption/ionization” (MSLDI). Since effective MSE can be achieved only above certain analyte/matrix molar ratios, it is suggested that in MSLDI, CTAB could act as an analyte during the LDI process. Within the two-step model of MALDI ionization, this implies a depletion of primary matrix ions by neutral analytes, via secondary ion-molecule reactions. In MSLDI a slight improvement in resolution was also achieved, probably due to cool ions (i.e., small plume) generation; however, MSLDI has a poorer detection limit, which should make it most useful for screening of abundant LMWC.
One problem encountered in using QA salts as additive is that not only the matrix ions but also the analyte signals could be unacceptably suppressed if the ratio of matrix/QA salt/analyte is not carefully optimized – an issue that is further complicated by the possibility that the optimal ratio might be dependent on the nature of matrix, analyte, and QA salt. Mechanistic aspects of matrix and analyte suppression effects induced by QA salts have been investigated [47]. It was suggested that the observed MSE and ASE could be rationalized by a cluster ionization model assuming that analytes and matrix ions, coexisting in the cluster, compete for the limited number of net charges available. In the presence of a sufficient number of QA ions, the net charges will be removed, thus resulting in the MSE and ASE. According to the authors ‘the cluster ionization model’ was not conflicting with the ionization via secondary gas-phase reactions and, ultimately, the final ions observed are the combined results of both mechanisms. Later on, Lou et al. [48], using different model compounds and two different sample deposition, i.e., standard dried-droplets and modified thin layer methods, demonstrated that generation of gas-phase ions from charged matrix/analyte clusters is the ionization step likely responsible for matrix/analyte ion suppression effects. Ion suppression effects in multi-analyte mixture can be significantly reduced by caesium chloride addition to a DHB matrix as demonstrated by Popkova and Schiller [49] for lipid analysis in crude adipose tissue extracts where, in absence of added salts, phosphatidylcholines and phosphatidylethanolamines are suppressed by the most abundant triacylglycerols (TAG). Comparison with other alkali chlorides provided evidence that the size of the alkali ion plays a role in ion formation yield and analyte fragmentation as well.
A different strategy to reduce/eliminate matrix interference relies on the incorporation of the matrix (e.g., DHB) into a sol-gel polymeric structure as reported by Lin and Chen [50] or on matrices supported onto different organic/inorganic materials. An example is provided by 2′,4′,6′-trihydroxyacetophenone monohydrate (THAP) matrix supported on cyclodextrin that was successfully applied for the analysis of testosterone and diazepam [51]. In the presence of cyclodextrin, matrix fragment ions and alkali adduct ions were significantly suppressed in favor of [THAP+H]+, thus reducing the background and amplifying the analyte signals. The same matrix was supported on lithium-substituted mordenite [52] and used for acetylsalicylic acid and phenobarbital analysis. The binding energies of Li+ and H+ to THAP, estimated by quantum-chemical models around 236 and 927 kJ/mol, respectively, account for Li+ detachment from [THAP+Li]+ producing [analyte+Li]+ ions. The tested molecules were then preferentially ionized as lithium adducts, avoiding the concurrent formation of protonated or other alkali metal adducts that would adversely affect sensitivity.
A similar mechanism can be invoked when neutral NaDHB and NH4DHB salts were used as matrices for lipids analysis [53]. The cationization efficiency was improved, avoiding the separation of lipid species into several ionization states. This allowed for simplified lipid spectra interpretation and detection of sterols and steryl esters not affordable by standard acidic DHB.
Lithium 2,5-dihydroxybenzoate (LiDHB) [54], lithium α-cyano-4-hydroxycinnamate, lithium sinapate, and lithium salts of other organic aromatic acids [55] were synthesized and tested as potential matrices for MALDI MS of hydrocarbons, di- and tri-acylglycerols, and wax esters.
Lithium vanillate offered the most remarkable increase in sensitivity (compared with LiDHB) for long-chain hydrocarbons and wax esters; moreover, the formation of homogeneous spots makes this matrix a potential candidate for MALDI MSI applications.
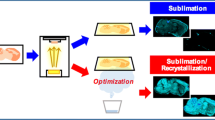
Novel deposition methods of conventional matrices have been essentially developed for MALDI MSI where thin tissue slices (prepared by cryo-section) are mounted on sample plates and fixed; then matrix is applied on the entire tissue or on specific regions of interest [56]. Gemperline et al. [57] compared three application techniques (namely airbrush, automatic sprayer, and sublimation) of DHB and CHCA matrices for the metabolome analysis of Medicago truncatula root and nodule tissues. A comparison of mass spectra provides straightforward evidence that the peak patterns of matrix are greatly influenced by the application technique; airbrush deposition generated the most intense matrix-related background that could significantly hinder metabolites detection. Moreover, when sample was analyzed, the sublimation and automatic sprayer methods produced less pronounced analyte delocalization and an increased number of metabolites detected especially when using DHB.
Recently, an electric field-assisted scanning-spraying matrix coating system [58] was described. The generation of very small crystals (<10 μm) of common matrices resulted in an enhancement of detection sensitivity and a number of LMWC (fatty acids, nucleosides, monophosphate, nucleosides, and N-acetylneuraminic acid) could be detected in cancerous tissues. The same method was employed using other matrices (i.e., quercetin for positive and negative ion detection and 9-aminoacridine (9AA) for negative ion detection) for the analysis of endogenous compounds in the cancerous and non-cancerous regions of human prostate cancer tissues; 152 metabolites, out of the 1091 overall detected and localized, exhibited differential distributions between the two tissue regions [59].
Small and homogeneous spots can be generated by electrowetting-assisted drying [60] on special functionalized e-MALDI target plates. This deposition method proved successful for the analysis of LMW drugs such as paracetamol, quinine, penicillin V, aspirin, ibuprofen, and fenofibrate. As is clear from scanning electron microscopy (SEM) images (Fig. 2), e-MALDI drop drying leads to very uniform crystallization of DHB, which results in improved spot quality, enhanced sensitivity, and reduction of the analytes’ lateral diffusion in MSI experiments. Very small and regular crystals were generated by electrospray deposition, which allowed for an intimate incorporation of analyte throughout the matrix layer. This method was applied for the analysis of peptides [61] and for MALDI FT ICR imaging of lipids in mouse brain and liver cancer tissue section [62]. Interestingly, the electrospray deposition method was suitable for either various organic matrices or metallic nanoparticles (NPs), which made it unique for MALDI MSI analysis. A recently proposed sample deposition protocol exploits the inkjet technology to automatically drop cells and matrix solutions onto ITO glass plates, the surfaces of which were made hydrophobic by treating them with octadecyltrimethoxysilane. Membrane phospholipids (e.g., phosphatidylcholines) of printed cells were detected by MALDI MS using DHB as a matrix [63]. Similarly, a subcellular detailed investigation was achieved by a dual-positioner nanomanipulator workstation, which permitted a concomitant extraction of organelle content and the co-deposition of analyte and matrix solution for MALDI analysis [64].
Top row: SEM images of dried samples of DHB matrix and paracetamol (10:1 molar ratio) with conventional (a), and with e-MALDI (b) drop drying (Thick scale bars: 0.5 mm). Bottom row: same comparison for DHB matrix with quinine (10:1 molar ratio): (c) conventional drop drying; (d) e-MALDI (Thick scale bars: 0.25 mm). Images were obtained side by side on the same target plate. Reproduced with permission from [60]. Copyright American Chemical Society, 2016
Since high quality MALDI MS images are heavily dependent on the matrix deposition step, an open-source software and hardware device has been recently developed [65, 66] capable of delivering uniform and reproducible coatings even by inexperienced operators.
Solvent-free one-step automatic matrix deposition permitting homogeneous coverage of mouse brain tissue with 2–4 μm-sized crystals of CHCA has been described by Trimpin et al [67].
The advantages of these deposition strategies, over those employing additives, stems from their adaptability to various kinds of matrices and the absence of further interfering signals due to their pure physical character. However, they are specifically designed for MALDI MSI experiments and often require a dedicated instrumentation.
High molecular weight matrices, matrix/analyte derivatization
One feasible alternative approach to elude the problem of matrix background in the low m/z range is to search for high molecular weight (HMW) matrices (see Fig. 3). An example is given by 2,3,4,5-tetrakis(3′,4′-dihydroxylphenyl)thiophene (DHPT), which allows detection of a wide range of LMW amines (i.e., amino acids, peptides, vitamin B, β-agonists, alkaloid and aromatic amines) at femtomole level (Chen et al. [68]). Compared with CHCA, DHPT gives a mass spectrum almost devoid of peaks in the low mass range, the sole matrix ions detected being the radical cation at m/z 516.087 (positive ion mode) and the deprotonated molecule at m/z 515.081 (negative ion mode). Also worth mentioning in this framework is a group of synthetic porphyrins having a high enough molecular weight to ensure absence of spectral interference for small molecules. A further advantage is that their laser absorption and proton affinity can be fine-tuned by changing the side chains on the central structure. Ayorinde et al. [69] first tested meso-tetrakis(pentafluorophenyl)porphyrin (F20TPP) as a candidate MALDI matrix for the analysis of alkylphenol ethoxylates; F20TPP was also successfully employed for several other applications such as the analysis of the fatty acid composition of vegetable oils [70], detection of sugars, ascorbic acid, citric acid, and sodium-benzoate in beverages [71] and HIV protease inhibitor drugs in cell extracts [72]. The goal of shifting the matrix mass towards higher values could also be achieved by synthesizing suitable derivatives of a conventional matrix. Porta et al. [73], for instance, produced the CHCA derivatives (E)-2-cyano-3-(naphthalen-2-yl)acrylic acid (NpCCA) and (2E)-3-(anthracen-9-yl)-2-cyanoprop-2-enoic acid (AnCCA) pursuing a two-fold purpose: (1) minimize the matrix spectral interference in the low m/z range, and (2) obtaining more favorable proton affinity values. Indeed, AnCCA has no matrix ions below m/z 230 whereas NpCCA has only a few ions below this m/z value; an improvement per se only marginal compared with CHCA that, however, is compensated for by significantly improved signal-to-noise ratio (compared with CHCA) for 30% of the LMWC tested.
A feasible alternative to HMW matrices is to shift the analytes mass towards higher m/z values using appropriate derivatizing agents; charged groups can be eventually introduced, thus increasing the analyte’s ionization efficiency. A representative example is given by N-phosphorylation reaction of natural amino acids and small peptides as proposed by Gao et al. [74]. Under mild conditions, the N-terminal and ɛ-amino group of lysine can be fully labeled with a neutral phosphoryl group originating a reduced number of side products and permitting the analysis of these target molecules in serum sample as N-phosphoryl derivatives. Compounds containing primary or secondary amine groups were easily derivatized by tris(2,4,6-trimethoxyphenyl)phosphonium acetic acid and N-hydroxysuccinimide esters. This derivatization reaction, allowing a limit of detection in the low femtomole range, was applied to quantify a mixture of antibiotics by investigating isotopically coded light and heavy derivatives [75]. Girard’s reagents T and P have been employed for the derivatization of oxosteroids [76], steroids [77], and small oligosaccharides [78]. Hailat and Helleur [79] converted some selected sterols into their corresponding picolinyl esters, N-methylpyridyl ethers, and sulphated esters before their analysis in mussels’ extracts by MALDI MS using DHB and THAP conventional matrices. A nonreductive amination reaction using aminopyrazine as co-matrix of DHB and derivatizing agent has been proposed for the characterization of oligosaccharides released from selected N-glycoproteins [80].
One drawback common to the above described labeling methods is that additional tedious purification steps are required, implying that one of the strengths of MALDI (i.e., reduced sample manipulation) is lost. To prevent this problem, Rohmer et al. [81] suggested performing the derivatization reaction directly on the target plate using 3-aminoquinoline (3-AQ) as both labeling reagent and MALDI matrix. However, derivatization rates were found to be critically dependent on the amount of organic solvents required to guarantee high yields in the spot. To avoid this issue, a modified protocol was proposed introducing an ionic liquid matrix; mixing 3-AQ and CHCA allowed for high-sensitivity detection of glycans detached from oligosaccharides [82]. A distinctive approach, de facto involving analyte derivatization, is represented by reactive MALDI matrices that will be specifically treated later (vide infra).
Second generation matrices for small molecules analysis
An attractive strategy in the development of novel MALDI matrices consists in their rational design, whereby the core structure of classic matrices is purposely modified to achieve specific physo-chemical properties [14]. This represents the step forward towards the replacement of the mostly empirical approach to the matrix choice, typical of the early times of MALDI, with a rationally based one. The pioneers in the field of rationally designed (RD) matrices were Jaskolla and co-workers, who synthesized numerous CHCA and DHB derivatives aiming to tune their proton affinity and improve analyte ionization efficiency. It was demonstrated that the PA could be reduced by incorporating electron withdrawing moieties, e.g., halogens, into the CHCA core structure [83]. Among the screened matrices, 4-chlorocyanocinnamic acid (ClCCA), obtained by replacing the hydroxyl group of CHCA with a chlorine atom, was effective in increasing the sensitivity of tryptic digest of standard proteins. Indeed, because of their reduced PA, the halogenated matrices exhibit an improved protonation of weakly basic, neutral, and acidic peptides. However, the addition of multiple electron withdrawing moieties [84] was not satisfactory due to a blueshift of the absorption wavelength and to morphological changes (needle-like crystals were formed). A possible method to address these issues could be the mixture of two halogen-substituted compounds with similar PA or the addition of CHCA; in this case one matrix can be excited by the laser and can guarantee the proton transfer to the more halogenated matrix and finally to the analytes. Here, it is worth mentioning that CClCA matrix has found interesting applications in the identification of lipids [85] and vitamin B [86, 87]. The previously mentioned [73] NpCCA and (AnCCA) give a further example of RD matrices aimed to tune the PA of CHCA with a concomitant shift towards higher m/z values. These novel matrices showed a better S/N ratio and crystallization behavior compared with CHCA. 4-Phenyl-α-cyanocinnamic acid amide matrix was recently described for negative ion mode MALDI MSI of various lipid classes displaying a superior sensitivity and reproducibility compared to e.g., 9AA [88]. Fukuyama et al. developed alkylated DHB for analysis of hydrophobic peptides [89] that was later applied for the MALDI MSI of phospholipids in brain tissue [90].
Very recently, a library of 59 structurally related cinnamic acid derivatives was synthesized and potential MALDI matrices were assessed by analyzing sulfatides, a class of anionic lipids abundant in brain [91]. The cinnamic acids currently used are E-cinnamic isomers; Salum et al. [92] reported the use of Z-sinapinic acid (SA) for MALDI MS of peptides. Compared with E-SA and CHCA, a smaller number of clusters was observed in the low m/z region allowing the rapid and sensitive detection of very short hydrophilic and hydrophobic peptides.
Reactive matrices
The so called “reactive matrices” can work simultaneously as MALDI matrices and as a derivatization agent to improve ionization yield, gain structural information, or enable analysis of small molecules by lowering their volatility or increasing their final m/z value. Reactive MALDI was first described by Zhang and Gross [93]; a mixture of anthranilic acid (AA), nicotinic acid, and diammonium citrate (2:1:0.003) formed a Schiff base, namely an “in situ” MALDI matrix reacting with the basic sites of modified oligodeoxynucleotides. A mixture composed of 2,4-dinitrophenylhydrazine (DNPH), a well-known derivatizing agent, and DHB was reported by Brombacher et al. [94] for the analysis of seven corticosteroids separated by capillary liquid chromatography coupled (off-line) to MALDI MS. Such a mixture was reported as a “reactive matrix”, which can be somewhat misleading since DHB played the usual role of a conventional MALDI matrix whereas reactivity (i.e., formation of steroid-dinitrophenylhydrazone derivatives) was merely ensured by the added DNPH [95]. Similarly, DNPH and 4-dimethylamino-6-(4-methoxy-1-naphthyl)-1,3,5-triazine-2-hydrazine (DMNTH) have been successfully used as reactive matrices for the analysis of carbonyl-containing compounds [96]. Note, however, that significant improvements in sensitivity and limits of detection can only be achieved using prolonged (48 h) reaction time and the assistance of a conventional CHCA matrix. Conversely, Schiller’s group has demonstrated that DNPH can be used as a genuine reactive matrix (i.e., no assistance by a conventional matrix) for the MALDI MS analysis of aldehydes after conversion into their corresponding hydrazones [97]. By such an approach, lipid oxidation products could be easily detected; at the same time, it was demonstrated that DNPH also enabled the detection of non-oxidized phospholipids. Other hydrazide and hydrazine reagents [98, 99] were applied for the detection at parts per million (ppm) levels of small, hazardous gases possessing aldehyde (formaldehyde, acetaldehyde, propionaldehyde) or ketone (acetone, methyl ethyl ketone, methyl isobutyl ketone) moieties.
The pyrylium cation is a six-membered heterocyclic ion consisting of five carbon atoms and one positively charged oxygen atom. A family of pyrylium salts was tested as derivatizing agents for the analysis of small molecules containing primary amine groups [100]. Among these, 2,4-diphenyl-pyranylium (DPP) was used as an effective reactive matrix to map the distribution of dopamine and amphetamine and to quantify a neurotoxin in brain tissue sections [100]. The aromatic compounds 2,4-dihydroxybenzaldehyde and 2,5-dihydroxyacetophenone were employed for the study of polyamines [101], whereas 1,2-phenylenediamine was selected for the analysis of ketocarboxylic acids as pyruvate [102]. Oligosaccharides were derivatized with a 2-hydrazinopyrimidine reactive matrix that can facilitate their ionization due to the presence of an electron-withdrawing N-heterocycle. This approach was applied to the analysis of neutral and sialylated oligosaccharides released from glycoproteins of biological samples. Moreover, their improved fragmentation allowed a detailed structural study by MS/MS [103].
A modified version of this strategy, named label-assisted LDI, employed a pyrene-based boronic acid for the detection of various cis-1,2-diols. The boronic functional group could act as a molecular recognition probe towards vic-diols, allowing their selective detection via LDI-MS in a single experiment. The strategy was applied for catecholamine neurotransmitters dopamine and epinephrine detection [104].
An example of RD reactive matrix, aimed to confer molecular recognition properties to the archetype matrix CHCA, was recently described [105]. 4-Formyl-phenylboronic acid (FPhBA) was condensed with cyanoacetic acid by a readily accessible standard Knoevenagel condensation, resulting in the formation of (E)-4-(2-cyano-2-carboxyvinyl) phenyl]boronic acid (CCPBA); the α-cyanocinnamic acid core was preserved whereas the hydroxyl moiety was replaced by a boronic acid residue able to act as a molecular recognition probe. Although the UV-vis absorption spectrum of CCPBA showed a slight bathochromic shift compared with CHCA, a similar molar absorption coefficient value at the laser wavelength (337 nm) was observed. It was demonstrated that this matrix selectively recognizes vic-diols, α-hydroxyacids, and amino alcohol compounds. For instance, small diols such as glycerol (MW = 92), 1,2-ethandiol (MW = 62), or 1,3-propanediol (MW = 76), that are metabolic markers of chronic pathologies or contaminants of food commodities, could be readily detected by MALDI MS at level as low as 50 pmol/spot. Moreover, the proposed matrix first allowed the selective detection of fluoride anions even in the presence of equimolar amounts of other halides. To show the effectiveness of the CCPBA matrix, a human urine sample was simply diluted (1:10 v/v) with water and directly analyzed obtaining a highly informative spectrum where several metabolites could be tentatively assigned. It is worth noting that, under the same conditions, a conventional matrix as 9-AA gave a spectrum containing only matrix-related peaks.
An unpredicted matrix reactivity towards some analytes has also been described [106, 107]. For instance, thiosalicylic acid (TSA), known for its reducing properties, behaves as a reactive matrix towards disulfide bond producing TSA-adducted peptides. Comparison with MALDI mass spectra obtained using conventional matrices permits counting of disulfide bonds in the investigated peptides [106]. An unusual case of “unwanted reactive MALDI” has been described by Chendo et al. [107] using standard acidic matrices (e.g., 2,5-DHB, DCTB and dithranol among others) to assist ionization of nitroxide-terminated poly(4-vinylpyridine). Either covalent or non-covalent adducts with some of the investigated matrices were formed depending on the MALDI source pressure; a mechanism was also proposed to explain the matrix/polymer covalent adduct formation.
Matrices for negative ion mode
9-Aminoacridine (9AA) was introduced for the analysis of LMWC (as phenols, carboxylic acids, sulfonates, and alcohols) [108] in negative ion mode where proton transfer is typically dictated by differences of the gas-phase basicities of deprotonated analyte and deprotonated matrix [109]. Then 9AA has found extensive use for the analysis of phytohormones [110], bile acids in urine [111] and plasma [112], oligosaccharides [113], and phospholipids [114, 115]; worth mentioning are also metabolomic studies [116, 117] and the analysis of various natural resins and varnish samples from cultural heritage objects [118]. In some cases, however, matrices other than 9AA provided even better results in negative ion mode MALDI. Norharmane, for instance, permitted a 100-fold improvement, compared with 9AA, in the limit of detection of the bacterial endotoxin lipid A and phospholipids in tissues infected with Francisella novicida [119].
Other new matrices purposely designed for negative ion MALDI MS analysis of small molecules have been described, such as 2-mercaptobenzothiazole [120], N-(1-naphthyl)ethylenediamine dinitrate [121], 1,5-diaminonaphthalene [122], 2,5-diaminonaphthalene [123], 4-phenyl-α-cyanocinnamic acid amide [88], and quercetin [124]. 1,5-Diaminonaphthalene (DAN) has been reported for the analysis of phospholipids (PL), in both negative and positive ion modes [125], and LMW metabolites from corn leaf sections [122]. As far as PLs are concerned, it seems that DAN provided significantly higher signals for phosphatidylethanolamines, whereas 9AA seems to preferentially ionize phosphoinositol (PI) species. DAN hydrochloride salt was used in MALDI imaging of liver, brain, and kidney tissues from mice, permitting to visualize the distribution and change of small metabolites, including metal ions, amino acids, carboxylic acids, nucleotide derivatives, and lipids [126]. DAN exhibits an interesting behavior acting as a reduction agent, leading to the cleavage of disulphide bonds and thus facilitating in-source decay fragmentation of proteins [127]. DAN generates mainly c- and z-series ions by N–Cα bond cleavage, whereas 3-hydroxy-4-nitrobenzoic acid, a recently proposed oxidizing matrix, generated numerous a- and d-series ions [128] giving a complete picture of peptide pattern fragmentation. It was recently demonstrated that DAN can also function as electron transfer matrix (see below).
Several other matrices for negative ion MALDI-MS analysis of small molecules have been described such as 1-naphthylhydrazine hydrochloride [129], 2-(2-aminoethylamino)-5-nitropyridine [130], and nifedipine (2,6-dimethyl-3,5-dicarbomethoxy-4-(2-nitrophenyl)-1,4-dihydropyridine) [131], but their use is limited to very specific applications.
Ammonia-treated N-(1-naphthyl) ethylenediamine dihydrochloride has been employed as a matrix in negative ion mode to directly quantify free fatty acids (FFAs) in serum samples. By constructing multiple point calibration curves, using a C17:0 FA as internl standard, it was possible to demonstrate that FFAs in hyperglycemic patient sera were significantly higher than in healthy controls [132]. A similar matrix, N-(1-naphthyl) ethylenediamine dihydrochloride, was helpful in quantifying sulfated oligosaccharides such as chondroitin-4-sulfate in biological samples using deuterium-labeled ISs [133].
A significant breakthrough in this field is, no doubt, represented by the introduction of the so called “proton sponges” as MALDI matrices. The Alder’s proton sponge [134] (see Fig. 4), 1,8-bis (dimethylamino)naphthalene (DMAN) was rediscovered and first proposed by Svatos’ group in the more general framework of a rational protocol for matrix selection based on Brønsted-Lowry acid-base theory [135, 136]. A distinctive feature of a proton sponge (B) is the different behavior compared with classic MALDI matrices; being a very strong base (pKBH+ ≈ 18 in acetonitrile) the proton sponge is supposed to be capable of deprotonating an acidic analyte AH in the liquid-phase, thus forming an ion pair [A-H]–/[B+H]+. The acid base-driven equilibrium between the ion pair and the AH/B complex in the liquid phase is mirrored in the crystal phase and is responsible for the amounts of observed [A-H]– ions in the gas phase upon laser irradiation. To highlight this mechanism, the acronym “matrix-assisted ionization/laser desorption” (MAILD) was then proposed [136]. As DMAN itself does not undergo deprotonation in the gas phase, a further distinctive feature of such a matrix is that it is entirely ionless (in negative ion mode), thus representing a paradigm for MALDI MS analysis of small molecules as deprotonated anions. Indeed, DMAN proved to be particularly effective for negative ion mode analysis of several LMW acidic metabolites (e.g., carboxylic acids, fatty acids, amino acids, vitamins, plant and animal hormones) involved in important metabolic pathways such as Krebs cycle, fatty acid, and glucosinolate biosynthesis.
In the field of lipidomics, DMAN revealed to be effective for lipid fingerprinting of Gram-positive (Lactobacillus sanfranciscensis and Lactobacillus plantarum) microorganisms by intact cells MALDI MS [137]; free fatty acids, mono-, di-, and triglycerides, phospholipids, glycolipids, and cardiolipins (CLs), spanning a m/z range from 187 (deprotonated azelaic acid) to 1496 (CL 75:5 –H]-) were readily detected. The most noticeable drawback of DMAN is represented by its high sublimation rate under high-vacuum conditions, which makes it poorly suitable for long-lasting experiments such as MALDI MSI. Other proton sponges, stronger than DMAN, such as 1,8-bis(tetramethylguanidino)naphthalene (pKBH+ ≈ 25 in acetonitrile) were proposed for the determination of perfluorooctane sulfonates and perfluorooctanoic acid in water samples [138]. Twelve azahelicenes, displaying proton affinities comparable to those of classic proton sponges, have been tested as MAILD/MALDI matrices for the analysis of fatty acids and organic acids in a wide range of samples [139]. At least one of the azahelicenes investigated operates as a MALDI matrix (i.e., analyte deprotonation in the gas phase), whereas the most promising compound, i.e., 1,14-diaza[5]helicene (see Fig. 4), was found to be a MAILD matrix better than DMAN.
Novel, rationally designed matrices, retaining the 1,8-substituted naphthalene ring of DMAN, could be synthetized aimed to outperform the closely related “predecessors”. A representative example is provided by 1,8-di(piperidinyl)naphthalene (DPN), a low vapor pressure and ionless matrix, differing from the parent DMAN for the presence of piperidinyl substituents that increase the distance between the di-nitrogen center-chelated proton and a deprotonated anion, and steric repulsion [140]. According to the MAILD mechanism an increased distance is assumed to facilitate the dissociation of the ion pair, thus increasing the analyte signal intensity. Since the basicity of DPN and DMAN are roughly the same, it is supposed that the higher ionization efficiency exhibited by DPN could result from kinetic effects. Indeed, proton shielding in [DPN+H]+ by the two piperidinyl groups should reduce the rate at which (in the gas-phase) the proton is exchanged back to the analyte anion, resulting in an improved ionization. It is worth noting that DPN is capable of ionizing, with very low sensitivity, some compounds lacking an acidic proton such as glucose that can be detected as deprotonated molecule. Being vacuum stable, DPN can be proficiently used for imaging experiments.
A further example of vacuum stable proton sponge is provided by 3-(4,5-bis(dimethylamino)napthalen-1-yl)furan-2,5-dione (4-maleicanhydridoproton sponge or “MAPS” – see Fig. 4) [141] synthesized by reacting, at room temperature, DMAN with bromomaleic anhydride according to the Tyler´s reaction [142]. Despite structural modification due to maleic anhydride moiety introduction in the DMAN structure, the pKBH+ of MAPS (18.0 in acetonitrile) was comparable to that of DMAN (18.6); at the same time the ionless property of DMAN was retained. However, the higher vacuum stability of MAPS, compared with DMAN, makes it more suitable for MALDI MSI experiments of small molecules. MAPS was thus successfully employed to obtain ionic maps of lactate, 2-hydroxyglutarate and chloride anions (m/z 89, 147, 35, respectively) in an aggressive brain tumor tissue (Glioblastoma multiforme) as shown in Fig. 5.
MALDI Imaging results of a human diffuse glioma tissue section using the novel matrix MAPS. (a) Optical (left) and H&E stained (right) sections were graded by a pathologist. Subsequent MALDI-Imaging showed the spatial distribution and regions of interesting spectra corresponding to chloride ions (b), lactate (c) and 2-hydroxyglutarate (d) in the tissue sections. Reproduced from [141] with permission from The Royal Society of Chemistry
A noteworthy advance in the field of rationally designed matrices possessing outstanding basicity has been recently provided by our group [143]. Superbasic proton sponges having a 1,8-bisphosphazenylnaphthalene (PN) core unit and diverse P-amino and -alkyl substituents such as dimethylamino, pyrrolidino, tris-(pyrrolidino)phosphazenylbis(pyrrolidino), methyl, n-butyl, isopropyl, and cyclo-pentyl have been recently synthesized and characterised [144]. Such proton sponges display exceedingly high pKBH+ values ranging approximately from 30 to 32, in acetonitrile, depending on the substituents. Such outstanding basicity makes PNs potential candidates as negative ion MAILD/MALDI matrices offering a promising tool for producing deprotonated molecules from otherwise hardly ionizable small compounds. Indeed, the possibility of producing deprotonated species, with virtually absent fragmentation, from compounds lacking acidic protons such as steroids, sterols, fatty alcohols, and neutral saccharides was recently demonstrated. For instance, using a TPPN 1,8-bis(trispyrrolidinophosphazenyl)] matrix (see Fig. 4), cholesterol could be detected as intact deprotonated molecule at m/z 385.348 and quantified at levels as low as 3 pmol/spot or 6 nM. Other representative small molecules such as stearyl alcohol, β-estradiol, 17-α-ethinylestradiol, nomegestrol, ergocalciferol, cholecalciferol, β-sitosterol, campesterol, and sucrose could be deprotonated as well. Cholesterol, fatty acids, lysophospholipids, and phospholipids could also be analyzed in complex samples such as egg yolk and brain tissue extracts. Contrary to DMAN, which is completely ionless in negative ion mode, the tested superbasic proton sponges produce few clearly recognizable, matrix-related fragment ions, indicating that they are not entirely photostable under laser irradiation. Moreover, TPPN was experimentally proven to be, contrary to DMAN, a vacuum-stable matrix suitable for MSI experiments. Among superbasic proton sponges, TPPN proved also to be capable of deprotonating neutral carbohydrates, providing an efficient and simple way to produce gas-phase anions [145]. Note that negative ion mass spectra tend to be much more informative than their positive ion counterpart because of predominant cross-ring cleavage ions. To overcome the inability of conventional matrices to produce deprotonated molecules, several approaches, often unsatisfactory for neutral saccharides, have been proposed such as the use of matrix additives (chloride salt, sulphuric acid [146] and benzenesulfonic/p-toluene sulfonic acids [147]) and chemical derivatization using 2-aminobenzoic acid [148] or 2- aminoacridone [149]. A further approach is represented by the use of nonconventional matrices (β-carboline alkaloids) such as nor-harman working well for neutral cyclomaltooligosaccharides [150] but not for linear oligosaccharides [151] or harmine doped with ammonium chloride [152]. In this framework, the use of TPPN opens the way for structural characterization of neutral mono-, di-, tri-, and tetra-saccharides, cyclodextrins, and saccharide alditols by MALDI-MS/MS.
Binary, hybrid (organic-inorganic), and nanomaterial-based matrices
Binary matrices were first proposed by Solouki et al. [153]; 4-nitroaniline/coumarin and fructose/DHB matrix mixtures were investigated aimed at improving ion yield in MALDI Fourier-transform ion cyclotron resonance (FTICR) MS for the analysis of biomolecules such as bovine insulin and lysozyme (use of melittin as internal calibrant also improved mass accuracy). A binary matrix specifically intended for MALDI MS analysis of LMWC was described by Guo and He [154]; indeed a CHCA/9AA mixture produced fewer background interferences compared with single matrix in both positive and negative ion mode. After a screening of many LMWC, it was concluded that only analytes with pKa values outside the range of pKa of the binary components could be proficiently detected. As a result, other binary matrices were developed for the analysis of small biomolecules. Shanta et al. [155] tested a combination of 3-hydroxycoumarin and 6-aza-2-thiothymine for the analysis of drugs or a combination of DHB, CHCA with a mixture of trifluoroacetic acid and piperidine, for the identification of phospholipids in rat brain tissue sections [156]. The binary matrix composed of 2,5-DHB and 2,6-DHB was shown to improve the detection of glycans released from ovalbumin by PNGase F. Sixteen glycans could be detected compared with 10 and 9 glycans detected using 2,5-DHB and 2,6-DHB single matrix, respectively [157]. A binary matrice composed of DMAN and 9-AA was used for the direct analysis of whole bacterial cells in Gram-positive and Gram-negative microorganisms [158]. About 50 major membrane components, including free fatty acids, acylglycerols, phospholipids, and glycolipids were easily detected in negative ion mode. Ionizing properties of N-butyl-4-hydroxy-1,8-naphthalimide (BHN) were evaluated using model compounds such as O-acetyl-L-carnitine hydrochloride, oxytocin, and small peptides. BHN was found to improve co-crystallization of target analytes when mixed with DHB; compared with CHCA the BHN/DHB binary matrix provided a lower background and improved signal-to-noise ratio in the determination of small molecules desorbed from rat brain tissue slices by positive ion MALDI MS [159]. Carbon dots (CDs)-9AA binary matrix was used for nucleosides, amino acids, oligosaccharides, peptides, and drugs analysis in human urine, exhibiting excellent reproducibility compared with 9AA matrix; it was argued that CDs likely act as a matrix additive capable of suppressing 9AA ionization [160].
Hybrid organic-inorganic matrices consisting of e.g., organic matrix and metal nanoparticles have been proposed with the aim to attain a synergic effect and outperform other MS methods for small molecule analysis. Lin et al. [161] first proposed this type of strategy combining conventional matrices (DHB and CHCA) with polymeric metal nanoparticles (MNP) for the detection of small drugs such as salicylamide, mefenamic acid, ketoprofen, and prednisolone. The hybrid matrix provided high ionization efficiency with low background noise for the analysis of small target molecules in a complex mixture. A hybrid composed of immobilized silica and DHB on iron oxide magnetic nanoparticles for MALDI MS analysis of diverse LMW organic and inorganic compounds has been reported [162]. A hybrid material obtained by covalent attachment of an ionic liquid, composed of CHCA and 3-aminopropyltriethoxysilane (APTES), to TiO2 nanoparticles has been reported as a novel MALDI matrix for LMWC such as amino acids and dopamine. The rationale behind such an approach is an expected synergic effect exhibited by the ionic liquid (improved ionization efficiency) and the nanomaterial (desorption/ionization process facilitated by the large surface area). Indeed, compared with CHCA, the hybrid matrix provided reduced background and improved sensitivity [163]. A mixture of DHB and Fe3O4 nanoparticles was successfully used to alleviate the issue of triacylglycerol (TG) ion suppression by phosphatidylcholine (PC) [164] in MALDI MSI of maize seed cross-sections. A hybrid organic-inorganic material composed of CHCA covalently attached to amorphous silica (prepared by standard sol-gel chemistry) was characterized as a matrix for LDI analysis (as protonated and cationized adducts) of diverse synthetic peptide mixtures covering a 550–1300 Da mass range [165]. A hybrid matrix composed of CHCA covalently bound (trough APTES) to mesoporous silica- was synthesized and used to analyze 12 quinolone antibiotics and three pesticides [166].
Inorganic (e.g., silicon- and metal-based) nanomaterials are largely studied in the framework of LMWC analysis by surface assisted laser desorption ionization (SALDI) techniques (and variants) that are outside the scope of this review. Silicon-based matrices include silicon nanopost arrays [167, 168], silicon nanowires [169], and silicon nanopillars [170], while a huge number of metal matrices is reported based on gold [171,172,173,174,175] and silver substrates [176, 177] or metal oxides as titanium oxide [178,179,180], zinc oxide [181], and lithium oxide [182]. For further information, the reader should refer to recent dedicated reviews [10,11,12,13, 176, 183]. A recent overview of desorption/ionization mechanisms in SALDI MS is also available [184].
Graphene deserves particular mention as matrix for the analysis of LMWC (amino acids, polyamines, nucleosides, anticancer drugs, and steroids) by MALDI MS [185]. The graphene matrix functions as analytes trapping substrate and energy receptacle for laser radiation, providing highly efficient desorption/ionization free of matrix-related ions and without significant fragmentation. Graphene as MALDI matrix was also employed for nitropolycyclic aromatic hydrocarbons determination in PM2.5 samples using 9-nitroanthracene-d9 as the internal standard [186]. To overcome limitations to spatial resolution imposed by traditional matrix applications in MALDI MSI, a two-dimensional sheet of graphene was deposited directly on top of soybean leaves and rat brain tissue samples via a “dry transfer” process that ensures absence of analyte migration. Several classes of LMWC (small peptides, lipids, glycosylated metabolites) are desorbed and ionized, with minimal background interference from the matrix [187].
Graphene oxide (GO) has been employed as matrix for negative ion mode MALDI MSI of mouse brain tissue sections [188]; 190 lipids and 22 LMW metabolites were overall detected, whereas in situ MS/MS provided structural identification of 69 lipids. Compared with 9AA and N-(1-naphthyl) ethylenediamine dihydrochloride matrices, GO revealed to be especially effective for fatty acids and sphingolipids. Metabolite heterogeneity in viable and necrotic tumor regions of mouse breast cancer tissue could also be highlighted. Interestingly, the signal-to-noise (S/N) ratios of low-abundance phospholipid classes that are frequently suppressed by the presence of most abundant and easy to ionize phosphatidylcholines can be selectively improved, in positive mode MALDI MSI, using GO matrix [189]. Functionalized GO was used as a matrix for the detection of dopamine in cerebrospinal fluid [190] and for lipids analysis by TOF-SIMS [191]; GO modified with 4-vinylphenylboronic acid was used to selectively capture compounds with vicinal diols [192].
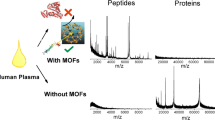
A brief mention is deserved for hybrid nanomaterials such as metal–organic frameworks (MOFs), namely coordination bonded networks consisting of metal ions as connectors and organic ligands as linkers [193]. An interesting aspect of MOF is that they can be designed (through the appropriate choice of the organic ligand) to act both as matrices for LDI MS of small molecules and adsorption material for simultaneous analyte enrichment as demonstrated for phosphopeptides [194] and environmental pollutants such as benzo(a)pyrene and perfluorinated compounds [195].
Ionic liquids matrices
Ionic liquid matrices (ILMs) for MALDI MS stem from general research on ionic liquids defined as non-molecular salts having a melting point below 100 °C or, in the case of the so-called room temperature ILs (RTILs), at or below room temperature. ILs possess negligible vapor pressure and good electrical conductivity; they are thermally stable, nonexplosive, nonflammable, and are capable of dissolving a wide range of analytes. Armstrong et al. [196] first investigated the potential use of a number of ILs as MALDI matrices. ILMs have the potential to overcome many of the limitations of solid matrices; in particular they do not require co-crystallization with the analyte and produce homogeneous solutions, thus avoiding search for “hot spots” and greatly improving quantitative analysis [197]. A second generation of ILMs was introduced by Crank and Armstrong [198], who examined over 100 cation/anion pairs aimed at designing ILs possessing different properties. The most common examples of ILMs consist of equimolar mixtures of conventional MALDI matrix compounds (e.g., DHB, CHCA, SA) together with organic bases (e.g., pyridine, tributylamine, aniline, just to name a few). ILMs have been successfully applied to a variety of molecules, including peptides [199, 200], proteins [201], phosphopeptides [202], oligosaccharides [203, 204], and phospholipids [205]. Some review papers [9, 206] and a recent book chapter [207] are available.
Here, representative applications of ILMs in the framework of MALDI MS detection, imaging, and quantitation of LMWC will be highlighted. Indeed, an outstanding property of several ILMs is their suitability for LMWC since, compared with the classic solid matrix, background signals are significantly suppressed by the charged organic base. For instance, CHCA/aniline ILMs revealed a highly versatile matrix for amino acids, peptides, proteins, lipids, phospholipids, synthetic polymers, and sugars [208]. ILMs have been employed as sample probe and matrix for liquid–liquid microextraction and ionization of phospholipids in soybean [209] or vegetable oils [210]. MALDI MS analysis of phosphopeptides in the low femtomole range was achieved using a solidified ionic liquid matrix consisting of 3-AQ and CHCA [211]. The ILMs formed by CHCA/triethylammonium (or diisopropylammonium) were used to analyze, with minimum sample preparation, 14 pharmaceutical drugs in different tablets, capsules, and solutions, providing a simple procedure for detecting drug counterfeits [212].
Some examples of applications of ILMs for LMWC imaging are worth mentioning. CHCA-based ionic liquids were employed for the neuronal single cell profiling of lipid species by matrix enhanced secondary ion mass spectrometry (ME-SIMS) [213], whereas CHCA/butylamine and DHB/butylamine ILMs were successfully used for MALDI MSI of lipids in mouse liver and cerebellum tissue sections [214].
Three ionic liquid matrices based on 2,5-DHB (i.e., 2,5-DHB/Aniline, 2,5-DHB/pyridine, and 2,5-DHB/3-acetylpyridine) were tested for MALDI MSI of lipids in human ovarian cancer biopsies [215]. RTILMs consisting of equimolar mixture of mefenamic acid and different bases (aniline, dimethyl aniline, pyridine, and 2-methyl picoline) were tested for their ability to ionize different and wide classes of compounds such as drugs, carbohydrates, and aminoacids, and found useful for MALDI MS analysis of lysates of pathogenic bacteria [216].
Recently, MALDI MS quantitative analysis of fructo-oligosaccharide mixtures in rice noodles, using two ILMs consisting of equimolar mixture of DHB with N-methylaniline or N-ethylaniline has been reported [217].
A detailed investigation of 27 ILMs for analyte quantification by MALDI MS was reported by Moon et al. [218], who found that some specific requirements should be met in order to ensure spot-to-spot spectral reproducibility.
Electron transfer matrices
Common matrices can putatively form radicals, but they scarcely exchange electrons with analyte, being the competing proton transfer the favored process. The electron transfer (ET) mechanism can become important when compounds with low ionization potentials are examined. The choice of matrices with ionization energies (IE) higher than analyte could promote electron transfer secondary reactions, thus avoiding the occurrence of proton transfers that usually induce the formation of matrix clusters. Accordingly, ET matrices (Fig. 6) could be potentially useful for LMWC analysis. McCarley et al. [219] found that anthracene and terthiophene could work as ET matrices for metallocenes as ferrocene. A two-photon ionization of matrix was proposed, followed by ET from the analyte to the matrix radical cation. This process is possible only if the ionization potential (IP) of the analyte is lower than the matrix one (in this example IPanthracene = 7.44 eV versus IPferrocene = 6.71 eV). Anthracene, pyrene, and acenaphthene were utilized as nonpolar matrices for the analysis of polybutadiene, polyisoprene, and polystyrene samples of various average molecular weights [220] .9,10-diphenylanthracene was used as ET secondary reaction matrix for the analysis of retinol and chlorophyll [221], and tetrathiafulvalene was used for pigments detection [222].
Non-protic 2-(2 E)-3-(4-tert-butylphenyl)-2-methylprop-2-enylidene] malononitrile (DCTB) has proven to be an excellent ET matrix for labile compounds such as substituted fullerenes, due to a very low onset of ion production [223]. Such a nonpolar and aprotic matrix was also used in MALDI MS analysis of coordination compounds, organometallics, porphyrins, phthalocyanines, carbohydrates, calixarenes, and macrocycles. A comparison of DCTB performance with that of dithranol, 2,5-DHB, and 2,4,6-THAP matrices is provided [224].
Eight phenylenevinylene oligomers (PVs) were synthesized and characterized as potential ET matrices for UV-MALDI. Indeed molar absorptivities (at 355 nm) are similar or higher than those of CHCA and DCTB, calculated IP values range from 6.88 to 7.96 eV, and positive ion mode LDI MS shows the presence of PV radical cation-free of clusters, adducts, or protonated species. Four selected PVs were tested as ET MALDI matrices for porphyrins, phthalocyanines, and polyaromatic compounds, achieving in the most favorable case, LOD in the low fmol range [225].
Effective ET matrices are typically aprotic compounds; however, it has recently been demonstrated that even matrices with basic residues as DAN can work as ET matrices for the analysis of chlorophylls [226] and bacteriochlorophylls [227]. Indeed, DAN drastically outperformed other ET matrices since it avoided demetallation and phytol-ester linkage fragmentation usually observed in the previous MALDI MS analysis of chlorophylls.
Conclusions and outlooks
Recent achievements in the design and applications of conventional and novel matrices specifically addressing the challenging issue of MALDI MS analysis of LMWC are presented and discussed. The outlook for the future is promising. Definite evidence is provided that MALDI MS represents a powerful and invaluable analytical tool also for small molecules analysis, including their quantification, thus opening new and exciting applications in most disparate fields, including food and environmental chemistry, biomedicine, metabolomics, and imaging mass spectrometry to name just a few (see Table 1).
The large degree of molecular heterogeneity of LMWC (paradigmatic is the field of lipidomics) precludes the search for a universal matrix; on the other hand, demands on quantitative MALDI MS and MALDI MSI applications ask for matrices possessing, in addition to the classic requisites, precise physicochemical features such as homogeneous crystallization, small crystal sizes, and high vacuum stability.
First generation archetype matrices (e.g., CHCA and DHB) have been adapted to LMWC analysis through the use of additive/dopants or matrix/analyte derivatization but they tend to be replaced by second generation (rationally designed) matrices. Rationally designed matrices possessing molecular recognition properties have been proposed that are expected to find interesting new applications for targeted MALDI MS analysis.
Ionic liquid matrices, the properties of which can be tailored for specific applications by varying the nature of the acidic and basic counterparts, can improve spot homogeneity, spot-to-spot reproducibility, and analyte quantitation. Furthermore, many liquid matrices can be operated in both positive and negative ion modes, thus helping profiling complex mixtures such those encountered in metabolomics/lipidomics studies. RTILs are promising matrices also for MALDI MSI since spatial resolution is not limited by the size of the matrix crystals and their enhanced durability allows for a high number of repetitive shots on one tissue position.
Matrices for negative ion mode have significantly contributed to extend MALDI MS to LMWC analysis, and research efforts on this specific topic are far from being considered exhaustive. Prototypical is the discovery of completely ionless proton sponge matrices and superbasic proton sponges capable of deprotonating compounds lacking acidic protons such as steroids, sterols, fatty alcohols, and neutral saccharides. Moreover, vacuum stability of such matrices makes them potential candidates for MALDI MSI experiments. Proton sponges provide an exemplary case of compounds originally intended as catalyst for organic synthesis that have found surprising applications as MALDI matrices. It is not the sole example; indeed many inorganic, organic, and hybrids nanomaterials, originally designed and developed for applications completely outside the field of MS, are more and more considered for their applications as MALDI matrices for LMWC analysis and imaging. Carbon-based nanomaterials (such as graphene, two-dimensional graphene, graphene oxide) appear to be particularly promising even for imaging experiments. Phenylenevinylene oligomers provide the most recent example of a class of materials originally intended for applications in the field of organic electronics and the properties of which can be rationally designed, that has been discovered as ET matrices for LMWC. The continuous development of new and advanced organic/inorganic materials presumably will continue to drive the attention of mass spectrometrists towards their possible applications in MALDI MS to address continuously challenging analytical problems. It is safe to foresee that the search for new MALDI matrices will likely be an endless story.
References
Karas M, Bachmann D, Hillenkamp F. Influence of the wavelength in high-irradiance ultraviolet laser desorption mass spectrometry of organic molecules. Anal Chem. 1985;57(14):2935–9.
Yamashita M, Fenn JB. Electrospray ion source. Another variation on the free-jet theme. J Phys Chem. 1984;88(20):4451–9.
Krüger R, Anja Pfenninger IF, Glückmann M, Karas M. Analyte incorporation and ionization in matrix-assisted laser desorption/ionization visualized by pH indicator molecular probes. Anal Chem. 2001;73:5812–21.
Beavis RC, Chaudhary T, Chait BT. α-Cyano-4-hydroxycinnamic acid as a matrix for matrixassisted laser desorption mass spectromtry. Org Mass Spectrom. 1992;27(2):156–8.
Tang W, Nelson CM, Zhu L, Smith LM. Positive ion formation in the ultraviolet matrix-assisted laser desorption /ionization analysis of oligonucleotides by using 2,5-dihydroxybenzoic acid. J Am Soc Mass Spectrom. 1997;8(3):218–24.
Smirnov IP, Zhu X, Taylor T, Huang Y, Ross P, Papayanopoulos IA, Martin SA, Pappin DJ. Suppression of α-cyano-4-hydroxycinnamic acid matrix clusters and reduction of chemical noise in MALDI-TOF mass spectrometry. Anal Chem. 2004;76(10):2958–65.
Bergman N, Shevchenko D, Bergquist J. Approaches for the analysis of low molecular weight compounds with laser desorption/ionization techniques and mass spectrometry. Anal Bioanal Chem. 2014;406(1):49–61.
Dreisewerd K. Recent methodological advances in MALDI mass spectrometry. Anal Bioanal Chem. 2014;406(9/10):2261–78.
Abdelhamid HN. Organic matrices, ionic liquids, and organic matrices@nanoparticles assisted laser desorption/ionization mass spectrometry. TrAC - Trends Anal Chem. 2017;89:68–98.
Shi CY, Deng CH. Recent advances in inorganic materials for LDI-MS analysis of small molecules. Analyst. 2016;141(10):2816–26.
Lu M, Yang X, Yang Y, Qin P, Wu X, Cai Z. Nanomaterials as assisted matrix of laser desorption/ionization time-of-flight mass spectrometry for the analysis of small molecules. Nanomaterials. 2017;7(4):87–107.
Zhang X, Niu J, Lu M, Cai Z. Recent advances of nanomaterials assisted negative ion laser desorption/ionization-time-of-flight mass spectrometry in the analysis of small molecules. Chinese J Chromatogr. 2016;34(11):1017.
Wang J, Liu Q, Liang Y, Jiang G. Recent progress in application of carbon nanomaterials in laser desorption/ionization mass spectrometry. Anal Bioanal Chem. 2016;408(11):2861–73.
Kiss A, Hopfgartner G. Laser-based methods for the analysis of low molecular weight compounds in biological matrices. Methods. 2016;104:142–53.
Wei J, Buriak JM, Siuzdak G. Desorption-ionization mass spectrometry on porous silicon. Nature. 1999;399(6733):243–6.
Baker TC, Han J, Borchers CH. Recent advancements in matrix-assisted laser desorption/ionization mass spectrometry imaging. Curr Opin Biotechnol. 2017;43:62–9.
Trim PJ, Snel MF. Small molecule MALDI MS imaging: Current technologies and future challenges. Methods. 2016;104:127–41.
Schwamborn K, Kriegsmann M, Weichert W. MALDI imaging mass spectrometry — From bench to bedside. Biochim Biophys Acta – Proteins Proteomics. 2017;1865(7):776–83.
Gessel MM, Norris JL, Caprioli RM. MALDI imaging mass spectrometry: Spatial molecular analysis to enable a new age of discovery. J Proteomics. 2014;107:71–82.
Wang P, Giese RW. Recommendations for quantitative analysis of small molecules by matrix-assisted laser desorption ionization mass spectrometry. J Chromatogr A. 2017;1486:35–41.
Hillenkamp F, Peter-Katalinić J. MALDI MS: a practical guide to instrumentation, methods and applications, Weinheim: Wiley-VCH Verlag GmbH & Co. KGaA; 2007.
Cramer R, editor. Advances in MALDI and Laser-Induced Soft Ionization Mass Spectrometry, Springer International Publishing Switzerland; 2016.
Cole RB. Electrospray and MALDI mass spectrometry: fundamentals, instrumentation, practicalities, and biological applications. Hoboken: Wiley; 2010.
Shah HN, Gharbia S. MALDI-TOF and tandem MS for clinical microbiology. 1st ed. Wiley, Chichester, West Sussex; 2017.
Hosseini S, Martínez Chapa SO. Fundamentals of MALDI-ToF-MS analysis : applications in bio-diagnosis, tissue engineering, and drug delivery. Singapore: Springer; 2017.
Knochenmuss R. The coupled chemical and physical dynamics model of MALDI. Annu Rev Anal Chem. 2016;9(1):365–85.
Moon JH, Yoon S, Bae YJ, Kim MS. Formation of gas-phase peptide ions and their dissociation in MALDI: Insights from kinetic and ion yield studies. Mass Spectrom Rev. 2015;34(2):94–115.
Knochenmuss R. Ion formation mechanisms in UV-MALDI. Analyst. 2006;131(9):966.
Karas M, Krüger R. Ion formation in MALDI: the cluster ionization mechanism. Chem Rev. 2003;103:427–40.
Knochenmuss A, Zenobi R. MALDI ionization: the role of in-plume processes. Chem Rev. 2003;103:441–52.
Dreisewerd K. The desorption process in MALDI. Chem Rev. 2003;103:395–426.
Zenobi R, Knochenmuss R. Ion formation in MALDI mass spectrometry. Mass Spectrom Rev. 1998;17(5):337–66.
Mirabelli MF, Zenobi R. Observing proton transfer reactions inside the maldi plume: experimental and theoretical insight into MALDI gas-phase reactions. J Am Soc Mass Spectrom. 2017;28(8):1676–86.
Niehaus M, Schnapp A, Koch A, Soltwisch J, Dreisewerd K. New insights into the wavelength dependence of MALDI mass spectrometry. Anal Chem. 2017;89(14):7734–41.
Alonso E, Zenobi R. Non-linear photoelectron effect contributes to the formation of negative matrix ions in UV-MALDI. Phys Chem Chem Phys. 2016;18(29):19574–87.
Jaskolla TW, Karas M. Compelling evidence for lucky survivor and gas phase protonation: the unified MALDI analyte protonation mechanism. J Am Soc Mass Spectrom. 2011;22(6):976–88.
Karas M, Gluckmann M, Schafer J. Ionization in matrix-assisted laser desorption/ionization: singly charged molecular ions are the lucky survivors. J Mass Spectrom. 2000;35(1):1–12.
Persike M, Zimmermann M, Klein J, Karas M. Quantitative determination of acetylcholine and choline in microdialysis samples by MALDI-TOF MS. Anal Chem. 2010;82(3):922–9.
Persike M, Karas M. Rapid simultaneous quantitative determination of different small pharmaceutical drugs using a conventional matrix-assisted laser desorption/ionization time-of-flight mass spectrometry system. Rapid Commun Mass Spectrom. 2009;23(22):3555–62.
Mahale V, Singh A, Phadke GS, Ghanate AD, Oulkar DP, Banerjee K, Panchagnula V. Determination of triazines and triazoles in grapes using atmospheric pressure matrix-assisted laser desorption/ionization high-resolution mass spectrometry. J AOAC Int. 2017;100(3):640–6.
Schiller J, Arnhold J, Benard S, Müller M, Reichl S, Arnold K. Lipid analysis by matrix-assisted laser desorption and ionization mass spectrometry: a methodological approach. Anal Biochem. 1999;267(1):46–56.
Porcari AM, Fernandes GD, Belaz KA, Schwab NV, Santos VG, Alberici RM, Varvara Gromova A, Eberlin MN, Lebedev AT, Tata A. Analytical methods high throughput MS techniques for caviar lipidomics high throughput MS techniques for caviar lipidomics. Anal Methods. 2014;6(8):2413–792.
McCombie G, Knochenmuss R. Small-Molecule MALDI using the matrix suppression effect to reduce or eliminate matrix background interferences. Anal Chem. 2004;76(17):4990–7.
Grant DC, Helleur RJ. Surfactant-mediated matrix-assisted laser desorption/ionization time-of-flight mass spectrometry of small molecules. Rapid Commun Mass Spectrom. 2007;21(6):837–45.
Kosevich M V, Boryak OA, Chagovets VV, Pashynska VA, Orlov VV, Stepanian SG, Shelkovsky VS. “Wet chemistry” and crystallochemistry reasons for acidic matrix suppression by quaternary ammonium salts under matrix-assisted laser desorption/ionization conditions. Rapid Commun Mass Spectrom. 2007;21(11):1813–9.
Guo Z, Zhang Q, Zou H, Guo B, Ni J. A method for the analysis of low-mass molecules by MALDI-TOF mass spectrometry. Anal Chem. 2002;74:1637–41.
Lou X, van Dongen JLJ, Vekemans JAJM, Meijer EW. Matrix suppression and analyte suppression effects of quaternary ammonium salts in matrix-assisted laser desorption/ionization time-of-flight mass spectrometry: an investigation of suppression mechanism. Rapid Commun Mass Spectrom. 2009;23(19):3077–82.
Lou X, van Dongen JLJ, Milroy L-G, Meijer EW. Generation of gas-phase ions from charged clusters: an important ionization step causing suppression of matrix and analyte ions in matrix-assisted laser desorption/ionization mass spectrometry. Rapid Commun Mass Spectrom. 2016;30(24):2628–34.
Popkova Y, Schiller J. Addition of CsCl reduces ion suppression effects in the matrix-assisted laser desorption/ionization mass spectra of triacylglycerol/phosphatidylcholine mixtures and adipose tissue extracts. Rapid Commun Mass Spectrom. 2017;31(5):411–8.
Lin Y-S, Chen Y-C. Laser desorption/ionization time-of-flight mass spectrometry on sol−gel-derived 2,5-dihydroxybenzoic acid film. Anal Chem. 2002;74:5793–8.
Yonezawa T, Asano T, Fujino T, Nishihara H. Cyclodextrin-supported organic matrix for application of MALDI-MS for forensics. Soft-ionization to obtain protonated molecules of low molecular weight compounds. Chem Phys. 2013;419:17–22.
Suzuki J, Sato A, Yamamoto R, Asano T, Shimosato T SH, Kondo J, Yamashita KI, Hashimoto K FT. Matrix-assisted laser desorption ionization using lithium-substituted mordenite surface. Chem Phys Lett. 2012;546:159–63.
Jaskolla TW, Onischke K, Schiller J. 2,5-Dihydroxybenzoic acid salts for matrix-assisted laser desorption/ionization time-of-flight mass spectrometric lipid analysis: simplified spectra interpretation and insights into gas-phase fragmentation. Rapid Commun Mass Spectrom. 2014;28:1353–63.
Cvacka J, Svatos A. Matrix-assisted laser desorption/ionization analysis of lipids and high molecular weight hydrocarbons with lithium 2,5-dihydroxybenzoate matrix. Rapid Commun Mass Spectrom. 2003;17(19):2203–7.
Horká P, Vrkoslav V, Hanus R, Pecková K, Cvačka J. New MALDI matrices based on lithium salts for the analysis of hydrocarbons and wax esters. J Mass Spectrom. 2014;49(7):628–38.
Schwartz SA, Reyzer ML, Caprioli RM. Direct tissue analysis using matrix-assisted laser desorption/ionization mass spectrometry: practical aspects of sample preparation. J Mass Spectrom. 2003;38(7):699–708.
Gemperline E, Rawson S, Li L. Optimization and comparison of multiple MALDI matrix application methods for small molecule mass spectrometric imaging. Anal Chem. 2014;86:10030–5.
Guo S, Wang Y, Zhou D, Li Z. Electric field-assisted matrix coating method enhances the detection of small molecule metabolites for mass spectrometry imaging. Anal Chem. 2015;87:5860–5.
Wang, X, Han, J, Hardie, DB, Yang, J, Pan, J, Borchers C. Metabolomic profiling of prostate cancer by matrix assisted laser desorption/ionization-Fourier transform ion cyclotron resonance mass spectrometry imaging using matrix coating assisted by an electric field (MCAEF). Biochim Biophys Acta – Proteins Proteomics. 2017;1865(7):755–67.
Kudina O, Eral B, Mugele F. E-MALDI: an electrowetting-enhanced drop drying method for MALDI mass spectrometry. Anal Chem. 2016;88(9):4669–75.
Malys BJ, Owens KG. Improving the analyte ion signal in matrix-assisted laser desorption/ionization imaging mass spectrometry via electrospray deposition by enhancing incorporation of the analyte in the matrix. Rapid Commun Mass Spectrom. 2017;31(9):804–12.
Li S, Zhang Y, Liu J, Han J, Guan M, Yang H, Lin Y, Xiong S, Zhao Z. Electrospray deposition device used to precisely control the matrix crystal to improve the performance of MALDI MSI. Sci Rep. 2016;6:37903.
Korenaga A, Chen F, Li H, Uchiyama K, Lin J-M. Inkjet automated single cells and matrices printing system for matrix-assisted laser desorption/ionization mass spectrometry. Talanta. 2017;162:474–8.
Phelps MS, Sturtevant D, Chapman KD, Verbeck GF. Nanomanipulation-Coupled matrix-assisted laser desorption/ ionization-direct organelle mass spectrometry: a technique for the detailed analysis of single organelles. J Am Soc Mass Spectrom. 2016;27(2):187–93.
Stoeckli M, Staab D, Wetzel M, Brechbuehl M. iMatrixSpray: a free and open source sample preparation device for mass spectrometric imaging. Chim Int J Chem. 2014;68(3):146–9.
Stoeckli M, Staab D. Reproducible matrix deposition for MALDI MSI based on open-source software and hardware. J Am Soc Mass Spectrom. 2015;26(6):911–4.
Trimpin S, Keune S, Räder HJ, Müllen K. Solvent-free MALDI-MS: developmental improvements in the reliability and the potential of MALDI in the analysis of synthetic polymers and giant organic molecules. J Am Soc Mass Spectrom. 2006;17(5):661–71.
Chen S, Chen L, Wang J, Hou J, He Q, Liu J, Wang J, Xiong S, Yang G, Nie Z. 2,3,4,5-Tetrakis(3′,4′-dihydroxylphenyl)thiophene: a new matrix for the selective analysis of low molecular weight amines and direct determination of creatinine in urine by MALDI-TOF MS. Anal Chem. 2012;84:10291–7.
Ayorinde FO, Hambright P, Porter TN, Keith QL. Use of meso- tetrakis(pentafluorophenyl)porphyrin as a matrix for low molecular weight alkylphenol ethoxylates in laser desorption/ ionization time-of-flight mass spectrometry. Rapid Commun Mass Spectrom. 1999;13(24):2474–9.
Ayorinde FO, Garvin K, Saeed K. Determination of the fatty acid composition of saponified vegetable oils using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Rapid Commun Mass Spectrom. 2000;14(7):608–15.
Ayorinde FO, Bezabeh DZ, Delves IG. Preliminary investigation of the simultaneous detection of sugars, ascorbic acid, citric acid, and sodium benzoate in non-alcoholic beverages by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Rapid Commun Mass Spectrom. 2003;17(15):1735–42.
van Kampen JJA, Luider TM, Ruttink PJA, Burgers PC. Metal ion attachment to the matrix meso-tetrakis(pentafluorophenyl)porphyrin, related matrices and analytes: an experimental and theoretical study. J Mass Spectrom. 2009;44(11):1556–64.
Porta T, Grivet C, Knochenmuss R, Varesio E, Hopfgartner G. Alternative CHCA-based matrices for the analysis of low molecular weight compounds by UV-MALDI-tandem mass spectrometry. J Mass Spectrom. 2011;46:144–52.
Gao X, Bi X, Wei J, Peng Z, Liu H, Jiang Y, Wei W, Cai Z. N-phosphorylation labeling for analysis of 20 natural amino acids and small peptides by using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Analyst. 2013;138:2632–9.
Lee PJ, Weibin Chen A, Gebler JC. Qualitative and quantitative analysis of small amine molecules by MALDI-TOF mass spectrometry through charge derivatization. Anal Chem. 2004;76:4888–93.
Wang Y, Hornshaw M, Alvelius G, Bodin K, Liu S, Sjövall J, Griffiths WJ. Matrix-assisted laser desorption/ionization high-energy collision-induced dissociation of steroids: analysis of oxysterols in rat brain. Anal Chem. 2006;78:164–73.
Galesio M, Rial-Otero R, Capelo-Martínez J-L. Comparative study of matrices for their use in the rapid screening of anabolic steroids by matrix-assisted laser desorption/ionisation time-of-flight mass spectrometry. Rapid Commun Mass Spectrom. 2009;23(12):1783–91.
Gouw JW, Burgers PC, Trikoupis MA, Terlouw JK. Derivatization of small oligosaccharides prior to analysis by matrix-assisted laser desorption/ionization using glycidyltrimethylammonium chloride and Girard’s reagent T. Rapid Commun Mass Spectrom. 2002;16(10):905–12.
Hailat I, Helleur RJ. Direct analysis of sterols by derivatization matrix-assisted laser desorption/ionization time-of-flight mass spectrometry and tandem mass spectrometry. Rapid Commun Mass Spectrom. 2014;28(2):149–58.
Cai Y, Zhang Y, Yang P, Lu H. Improved analysis of oligosaccharides for matrix-assisted laser desorption/ionization time-of-flight mass spectrometry using aminopyrazine as a derivatization reagent and a co-matrix. Analyst. 2013;138:6270–6.
Rohmer M, Meyer B, Mank M, Stahl B, Bahr U, Karas M. 3-Aminoquinoline acting as matrix and derivatizing agent for MALDI MS analysis of oligosaccharides. Anal Chem. 2010;82(9):3719–26.
Kaneshiro K, Fukuyama Y, Iwamoto S, Sekiya S, Tanaka K. Highly sensitive MALDI analyses of glycans by a new aminoquinoline-labeling method using 3-aminoquinoline/α-cyano-4-hydroxycinnamic acid liquid matrix. Anal Chem. 2011;15:3663–7.
Jaskolla TW, Lehmann W-D, Karas M. 4-Chloro-alpha-cyanocinnamic acid is an advanced, rationally designed MALDI matrix. Proc Natl Acad Sci USA. 2008;105(34):12200–5.
Soltwisch J, Jaskolla TW, Hillenkamp F, Karas M, Dreisewerd K. Ion yields in UV-MALDI mass spectrometry as a function of excitation laser wavelength and optical and physico-chemical properties of classical and halogen-substituted MALDI matrixes. Anal Chem. 2012;84(15):6567–76.
van der Werf ID, Calvano CD, Germinario G, Cataldi TRI, Sabbatini L. Chemical characterization of medieval illuminated parchment scrolls. Microchem J. 2017;134:146–53.
Calvano CD, Ventura G, Palmisano F, Cataldi TRI. 4-Chloro-α-cyanocinnamic acid is an efficient soft matrix for cyanocobalamin detection in foodstuffs by matrix-assisted laser desorption/ionization mass spectrometry (MALDI MS). J Mass Spectrom. 2016;51:841–8.
Ventura G, Arnesano F, Calvano CD, Palmisano F, Cataldi TRI. Cyanocobalamin conjugates of cisplatin and diaminocyclohexane-platinum(II): matrix-assisted laser desorption ionization mass spectrometry characterization using 4-chloro-α-cyanocinnamic acid as the matrix. RSC Adv. 2017; 7(85):53658–666.
Fülöp A, Porada MB, Marsching C, Blott H, Meyer B, Tambe S, Sandhoff R, Junker H-D, Hopf C. Alkylated dihydroxybenzoic acid as a MALDI matrix additive for hydrophobic peptide analysis. Anal Chem. 2012;84(9):4237–43.
Fukuyama Y, Tanimura R, Maeda K, Watanabe M, Kawabata S-I, Iwamoto S, Izumi S, Tanaka K. Improved spatial resolution of matrix-assisted laser desorption/ionization imaging of lipids in the brain by alkylated derivatives of 2,5-dihydroxybenzoic acid. Rapid Commun Mass Spectrom. 2014;28(5):403–12.
Stoyanovsky DA, Sparvero LJ, Amoscato AA, He RR, Watkins S, Pitt BR, Bayir H, Kagan VE. Improved spatial resolution of matrix-assisted laser desorption/ionization imaging of lipids in the brain by alkylated derivatives of 2,5-dihydroxybenzoic acid. Rapid Commun Mass Spectrom. 2014;28(5):403–12.
Tambe S, Blott H, Fülöp A, Spang N, Flottmann D, Bräse S, Hopf C, Junker HD. Structure-performance relationships of phenyl cinnamic acid derivatives as MALDI-MS matrices for sulfatide detection. Anal Bioanal Chem. 2017;409:1569–80.
Salum ML, Giudicessi SL, Schmidt De León T, Camperi SA, Erra-Balsells R. Application of Z-sinapinic matrix in peptide MALDI-MS analysis. J Mass Spectrom. 2017;52(3):182–6.
Zhang LK Gross ML. Location of abasic sites in oligodeoxynucleotides by tandem mass spectrometry and by a chemical cleavage initiated by an unusual reaction of the ODN with MALDI matrix. J Am Soc Mass Spectrom. 2002;13(12):1418–26.
Brombacher S, Owen SJ, Volmer DA. Automated coupling of capillary-HPLC to matrix-assisted laser desorption/ionization mass spectrometry for the analysis of small molecules utilizing a reactive matrix. Anal Bioanal Chem. 2003;376(6):773–9.
Vogel M, Büldt A, Karst U. Hydrazine reagents as derivatizing agents in environmental analysis--a critical review. Fresenius J Anal Chem. 2000;366(8):781–91.
Flinders B, Morrell J, Marshall PS, Ranshaw LE, Clench MR. The use of hydrazine-based derivatization reagents for improved sensitivity and detection of carbonyl containing compounds using MALDI-MSI. Anal Bioanal Chem. 2015;407(8):2085–94.
Teuber K, Fedorova M, Hoffmann R, Schiller J. 2,4-Dinitrophenylhydrazine as a New Reactive Matrix to Analyze Oxidized Phospholipids by MALDI-TOF Mass Spectrometry. Anal Lett. 2012;45(9):968–76.
Shigeri Y, Ikeda S, Yasuda A, Ando M, Sato H, Kinumi T. Hydrazide and hydrazine reagents as reactive matrices for MALDI-MS to detect gaseous aldehydes. J Mass Spectrom. 2014;49(8):742–49.
Shigeri Y, Kamimura T, Ando M, Uegaki K, Sato H, Tani F, Arakawa R, Kinumi T. 2-Hydrazinoquinoline: a reactive matrix for matrix-assisted laser desorption/ionization mass spectrometry to detect gaseous carbonyl compounds. Eur J Mass Spectrom. 2016;22(2):83.
Shariatgorji M, Nilsson A, Källback P, Karlsson O, Zhang X, Svenningsson P, Andren PE. Pyrylium Salts as Reactive Matrices for MALDI-MS Imaging of Biologically Active Primary Amines. J Am Soc Mass Spectrom. 2015;26(6):934–939.
Zaikin V, Borisov R, Polovkov N, Slyundina M. Reactive matrices for matrix-assisted laser desorption/ionization mass spectrometry of primary amines. Eur J Mass Spectrom. 2015;21(3):403–11.
Tholey A, Wittmann C, Kang M-J, Bungert D, Hollemeyer K, Heinzle E. Derivatization of small biomolecules for optimized matrix-assisted laser desorption/ionization mass spectrometry. J Mass Spectrom. 2002;37(9):963–973.
Jiang K, Aloor A, Qu J, Xiao C, Wu Z, Ma C, Zhang L, Wang PG. Rapid and sensitive MALDI MS analysis of oligosaccharides by using 2-hydrazinopyrimidine as a derivative reagent and co-matrix. Anal Bioanal Chem. 2017;409(2):421–9.
Addy PS, Bhattacharya A, Mandal SM, Basak A. Label-assisted laser desorption/ionization mass spectrometry (LA-LDI-MS): an emerging technique for rapid detection of ubiquitous cis-1,2-diol functionality. RSC Adv. 2014;4(87):46555–60.
Monopoli A, Calvano CD, Nacci A, Palmisano F. Boronic acid chemistry in MALDI MS: a step forward in designing a reactive matrix with molecular recognition capabilities. Chem Commun 2014;50:4322–4.
Asakawa D, Osaka I. Direct MALDI-MS analysis of the disulfide bonds in peptide using thiosalicylic acid as a reactive matrix. J Mass Spectrom. 2017;52(2):127–31.
Chendo C, Phan TNT, Marque SRA, Gigmes DLC. Mass spectrometry of nitroxide-terminated poly(4-vinylpyridine): A case of unwanted reactive MALDI. Int J Mass Spectrom. 2016;405:50–8.
Vermillion-Salsbury RL, Hercules DM. 9-Aminoacridine as a matrix for negative mode matrix-assisted laser desorption/ionization. Rapid Commun Mass Spectrom. 2002;16(16):1575–81.
Dashtiev M, Wäfler E, Röhling U, Gorshkov M, Zenobi R. Positive and negative analyte ion yield in matrix-assisted laser desorption/ionization. Int J Mass Spectrom. 2007;268(2–3):122–30.
Shroff R, Muck A, Svatoš A. Analysis of low molecular weight acids by negative mode matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Rapid Commun Mass Spectrom. 2007;21(20):3295–300.
Mims D, Hercules D. Quantification of bile acids directly from urine by MALDI–TOF–MS. Anal Bioanal Chem. 2003;375(5):609–16.
Mims D, Hercules D. Quantification of bile acids directly from plasma by MALDI-TOF-MS. Anal Bioanal Chem. 2004;378(5):1322–26.
Becher J, Muck A, Mithöfer A, Svatoš A, Boland W. Negative ion mode matrix-assisted laser desorption/ionisation time-of-flight mass spectrometric analysis of oligosaccharides using halide adducts and 9-aminoacridine matrix. Rapid Commun Mass Spectrom. 2008;22:1153–8.
Fuchs B, Bischoff A, Sü R, Teuber K, Schürenberg M, Suckau D, Schiller J. Phosphatidylcholines and -ethanolamines can be easily mistaken in phospholipid mixtures: A negative ion MALDI-TOF MS study with 9-aminoacridine as matrix and egg yolk as selected example. Anal Bioanal Chem. 2009;395:2479–87.
Calvano CD, Italiano F, Catucci L, Agostiano A, Cataldi TRI, Palmisano F, Trotta M. The lipidome of the photosynthetic bacterium Rhodobacter sphaeroides R26 is affected by cobalt and chromate ions stress. BioMetals 2014;27(1):65–73.
Vaidyanathan S, Goodacre R. Quantitative detection of metabolites using matrix-assisted laser desorption/ionization mass spectrometry with 9-aminoacridine as the matrix. Rapid Commun Mass Spectrom. 2007;21(13):2072–78.
Edwards JL, Kennedy RT. Metabolomic Analysis of Eukaryotic Tissue and Prokaryotes Using Negative Mode MALDI Time-of-Flight Mass Spectrometry. Anal Chem. 2005;77(7):2201–09.
Teearu A, Vahur S, Rodima T, Herodes K, Bonrath W, Netscher T, Tshepelevitsh S, Trummal A, Lõkov M, Leito I. Method development for the analysis of resinous materials with MALDI-FT-ICR-MS: novel internal standards and a new matrix material for negative ion mode. J Mass Spectrom. 2017;52(9):603–17.
Scott AJ, Flinders B, Cappell J, Liang T, Pelc RS, Tran B, Kilgour DPA, Heeren RMA, Goodlett DR, Ernst RK. Norharmane matrix enhances detection of endotoxin by MALDI-MS for simultaneous profiling of pathogen, host and vector systems. Pathog Dis. 2016;74(8):ftw097.
Teuber K, Schiller J, Fuchs B, Karas M, Jaskolla TW. Significant sensitivity improvements by matrix optimization: a MALDI-TOF mass spectrometric study of lipids from hen egg yolk. Chem Phys Lipids. 2010;163(6):552–60.
Chen R, Chen S, Xiong C, Ding X, Wu CC, Chang HC, Xiong S, Nie Z. N-(1-naphthyl) ethylenediamine dinitrate: A new matrix for negative ion MALDI-TOF MS analysis of small molecules. J Am Soc Mass Spectrom. 2012;49:737–41.
Korte AR, Lee YJ. MALDI-MS analysis and imaging of small molecule metabolites with 1,5-diaminonaphthalene (DAN). J Mass Spectrom. 2014;49(8):737–41.
Garate J, Fernández R, Lage S, Bestard-Escalas J, Lopez DH, Reigada R, Khorrami S, Ginard D, Reyes J, Amengual I, Barceló-Coblijn G, Fernández JA. Imaging mass spectrometry increased resolution using 2-mercaptobenzothiazole and 2,5-diaminonaphtalene matrices: application to lipid distribution in human colon. Anal Bioanal Chem. 2015;407(16):4697–708.
Wang X, Han J, Pan J, Borchers CH. Comprehensive Imaging of Porcine Adrenal Gland Lipids by MALDI-FTMS Using Quercetin as a Matrix. Anal Chem. 2014;86(1):638–46.
Dong W, Shen Q, Baibado JT, Liang Y, Wang P, Huang Y, Zhang Z, Wang Y, Cheung HY. Phospholipid analyses by MALDI-TOF/TOF mass spectrometry using 1,5-diaminonaphthalene as matrix. Int J Mass Spectrom. 2013;343–344:15–22.
Liu H, Chen R, Wang J, Chen S, Xiong C, Wang J, Hou J, He Q, Zhang N, Nie Z, Mao L. 1,5-diaminonaphthalene hydrochloride assisted laser desorption/ionization mass spectrometry imaging of small molecules in tissues following focal cerebral ischemia. Anal Chem. 2014;86:10114–21.
Molin L, Seraglia R, Dani FR, Moneti G, Traldi P. The double nature of 1,5-diaminonaphthalene as matrix-assisted laser desorption/ionization matrix: some experimental evidence of the protonation and reduction mechanisms. Rapid Commun Mass Spectrom. 2011;25(20):3091–96.
Fukuyama Y, Izumi S, Tanaka K. 3-Hydroxy-4-nitrobenzoic Acid as a MALDI Matrix for In-Source Decay. Anal Chem. 2016;88(16):8058–63.
He Q, Chen S, Wang J, Hou J, Wang J, Xiong S, Nie Z. 1-Naphthylhydrazine hydrochloride: A new matrix for the quantification of glucose and homogentisic acid in real samples by MALDI-TOF MS. Clin Chim Acta 2013;420:94–98.
Lorkiewicz P, Yappert MC. 2-(2-Aminoethylamino)-5-nitropyridine as a basic matrix for negative-mode matrix-assisted laser desorption/ionization analysis of phospholipids. J Mass Spectrom. 2009;44:137–43.
Nguyen H-N, Tanaka M, Komabayashi G, Matsui T. The photobase generator nifedipine as a novel matrix for the detection of polyphenols in matrix-assisted laser desorption/ionization mass spectrometry. J Mass Spectrom. 2016;51(10):938–46.
Zhang Y, Wang Y, Guo S, Guo Y, Liu H, Li Z. Ammonia-treated N-(1-naphthyl) ethylenediamine dihydrochloride as a novel matrix for rapid quantitative and qualitative determination of serum free fatty acids by matrix-assisted laser desorption/ionization-Fourier transform ion cyclotron resonance mass spectrometry. Anal Chim Acta. 2013;794:82–9.
Hsieh C-C, Guo JY, Hung S-U, Chen R, Nie Z, Chang H-C, Wu C-C. Quantitative analysis of oligosaccharides derived from sulfated glycosaminoglycans by nanodiamond-based affinity purification and matrix-assisted laser desorption/ionization mass spectrometry. Anal Chem. 2013;85(9):4342–9.
Alder RW, Bowman PS, Steele WRS, Winterman DR. The remarkable basicity of 1,8-bis(dimethylamino)naphthalene. Chem Commun 1968;(13):723–4.
Shroff R, Svatos A. Proton sponge: a novel and versatile MALDI matrix for the analysis of metabolites using mass spectrometry. Anal Chem. 2009;81:7954–9.
Shroff R, Rulísek L, Doubsky J, Svatos A. Acid-base-driven matrix-assisted mass spectrometry for targeted metabolomics. Proc Natl Acad Sci USA. 2009;106(25):10092–6.
Calvano CD, Zambonin CG, Palmisano F. Lipid fingerprinting of Gram-positive lactobacilli by intact cells matrix-assisted laser desorption/ionization mass spectrometry using a proton sponge based matrix. Rapid Commun Mass Spectrom. 2011;25(12):1757–64.
Cao D, Wang Z, Han C, Cui L, Hu M, Wu J, Liu Y, Cai Y, Wang H, Kang Y. Quantitative detection of trace perfluorinated compounds in environmental water samples by matrix-assisted laser desorption/ionization-time of flight mass spectrometry with 1,8-bis(tetramethylguanidino)-naphthalene as matrix. Talanta. 2011;85(1):345–52.
Napagoda M, Rulíšek L, Jančařík A, Klívar J, Šámal M, Stará IG, Starý I, Šolínová V, Kašička V, Svatoš A. Azahelicene superbases as MAILD matrices for acidic analytes. ChemplusChem. 2013;78(9):937–42.
Weißflog J, Svatoš A. 1,8-Di(piperidinyl)-naphthalene – rationally designed MAILD/MALDI matrix for metabolomics and imaging mass spectrometry. RSC Adv. 2016;6(79):75073–81.
Giampà M, Lissel MB, Patschkowski T, Fuchser J, Hans VH, Gembruch O, Bednarz H, Niehaus K. Maleic anhydride proton sponge as a novel MALDI matrix for the visualization of small molecules (<250 m/z) in brain tumors by routine MALDI ToF imaging mass spectrometry. Chem Commun. 2016;52(63):9801–4.
Swor CD, Zakharov LN, Tyler DR. A colorimetric proton sponge. J. Org Chem. 2010;75(20):6977–9.
Calvano CD, Cataldi TRI, Kögel JF, Monopoli A, Palmisano F, Sundermeyer J. Superbasic alkyl-substituted bisphosphazene proton sponges: a new class of deprotonating matrices for negative ion matrix-assisted ionization/laser desorption mass spectrometry of low molecular weight hardly ionizable analytes. Rapid Commun Mass Spectrom. 2016;30(14):1680–6.
Kögel JF, Xie X, Baal E, Gesevičius D, Oelkers B, Kovačević B, Sundermeyer J. Superbasic alkyl-substituted bisphosphazene proton sponges: synthesis, structural features, thermodynamic and kinetic basicity, nucleophilicity and coordination chemistry. Chem A Eur J. 2014;20(25):7670–85.
Calvano CD, Cataldi TRI, Kögel JF, Monopoli A, Palmisano F, Sundermeyer J. Structural characterization of neutral saccharides by negative ion MALDI mass spectrometry using a superbasic proton sponge as deprotonating matrix. J Am Soc Mass Spectrom. 2017;28(8):1666–75.
Wong AW, Cancilla MT, Voss LR, Lebrilla CB. Anion dopant for oligosaccharides in matrix-assisted laser desorption/ionization mass spectrometry. Anal Chem. 1998;71:205–11.
Wong AW, Wang H, Lebrilla CB. Selection of anionic dopant for quantifying desialylation reactions with MALDI-FTMS. Anal Chem. 2000;72:1419–25.
Thaysen-Andersen M, Mysling S, Højrup P. Site-specific glycoprofiling of N-Linked glycopeptides using MALDI-TOF MS: strong correlation between signal strength and glycoform quantities. Anal Chem. 2009;81(10):3933–43.
North S, Birrell H, Camilleri P. Positive and negative ion matrix-assisted laser desorption/ionization time-of-flight mass spectroh2-aminoacridone. Rapid Commun Mass Spectrom. 1998;12(7):349–56.
Nonami H, Tanaka K, Fukuyama Y, Erra-Balsells R. β-Carboline alkaloids as matrices for UV-matrix-assisted laser desorption/ionization time-of-flight mass spectrometry in positive and negative ion modes. Analysis of proteins of high molecular mass, and of cyclic and acyclic oligosaccharides. Rapid Commun. Mass Spectrom. 1998;12(6):285–96.
Yamagaki T, Nakanishi H. Negative-mode matrix-assisted laser desorption/ionization mass spectrometry of maltoheptaose and cyclomaltooligosaccharides. J Mass Spectrom Soc Jpn. 2002;50(4):204–7.
Yang C, Yanjie Jiang A, Cole RB. Anionic adducts of oligosaccharides by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Anal Chem. 2003;75:1638–44.
Solouki T, Gillig KJ, Russell DH. Mass measurement accuracy of matrix-assisted laser desorbed biomolecules: a Fourier-transform ion cyclotron resonance mass spectrometry study. Rapid Commun. Mass Spectrom. 1994;8(1):26–31.
Guo Z, He L. A binary matrix for background suppression in MALDI-MS of small molecules. Anal Bioanal Chem. 2007;387:1939–44.
Shanta SR, Kim TY, Hong JH, Lee JH, Shin CY, Kim KPK-H, Kim YH, Kim SK, Kim KPK-H. A new combination MALDI matrix for small molecule analysis: application to imaging mass spectrometry for drugs and metabolites. Analyst. 2012;137:5757–62.
Shanta SR, Zhou LH, Park YS, Kim YH, Kim Y, Kim KP. Binary matrix for MALDI imaging mass spectrometry of phospholipids in both ion modes. Anal Chem. 2011;15:1252–9.
Kim Y, Kim T, Lee J, Im H, Kim J. Enhanced detection of glycans by MALDI-TOF mass spectrometry using a binary matrix of 2,5-dihydroxybenzoic acid and 2,6-dihydroxybenzoic acid. Mass Spectrom Lett. 2013;4(2):38–40.
Calvano CD, Monopoli A, Ditaranto N, Palmisano F. 1,8-bis(Dimethylamino)naphthalene/9-aminoacridine: a new binary matrix for lipid fingerprinting of intact bacteria by matrix assisted laser desorption ionization mass spectrometry. Anal Chim Acta. 2013;798:56–63.
Cheng X, Ye X, Liu D, Zhao N, Gao H, Wang P, Ge G, Zhang X. N-butyl-4-hydroxy-1,8-naphthalimide: a new matrix for small molecule analysis by matrix-assisted laser desorption/ionization mass spectrometry. Rapid Commun Mass Spectrom. 2017;31(21):1779–84.
Chen Y, Gao D, Bai H, Liu H, Lin S, Jiang Y. Carbon dots and 9AA as a binary matrix for the detection of small molecules by matrix-assisted laser desorption/ionization mass spectrometry. J Am Soc Mass Spectrom. 2016;27(7):1227–35.
Lin P-C, Tseng M-C, Su A-K, Chen Y-J, Lin C-C. Functionalized magnetic nanoparticles for small-molecule isolation, identification, and quantification. Anal Chem. 2007;79:3401–8.
Tseng MC, Obena R, Lu YW, Lin PC, Lin PY, Yen YS, Lin JT, Huang LD, Lu KL, Lai LL, Lin CC, Chen YJ. Dihydrobenzoic Acid Modified Nanoparticle as a MALDI-TOF MS Matrix for Soft Ionization and Structure Determination of Small Molecules with Diverse Structures. J Am Soc Mass Spectrom. 2010;21(11):1930–9.
Liu H, Dai J, Zhou J, Huang H, Fei Chena ZL. A hybrid ionic liquid–matrix material, TiO2–Si–NH3+[CHC−], as a novel matrix for the analysis of small molecules by MALDI-TOF MS. Int J Mass Spectrom. 2015;376:85–9.
Feenstra AD, O’Neill KC, Yagnik GB, Lee YJ. Organic-inorganic binary mixture matrix for comprehensive laser-desorption ionization mass spectrometric analysis and imaging of medium-size molecules including phospholipids, glycerolipids, and oligosaccharides. RSC Adv. 2016;6(101):99260–8.
Fleith C, Cantel S, Subra G, Mehdi A, Ciccione J, Martinez J, Enjalbal C. Laser desorption ionization mass spectrometry of peptides on a hybrid CHCA organic-inorganic matrix. Analyst. 2014;139(15):3748–54.
Tang HZ, Ma YL, Liu F, Liu F, Liu ZW, Li JW, Zhou HY, Gao ZX. Detection of small molecules using SBA-15 modified CHCA as a novel matrix of MALDI-TOF MS. Int J Mass Spectrom. 2017;417:34–39.
Morris NJ, Anderson H, Thibeault B, Vertes A, Powell MJ, Razunguzwa TT. Laser desorption ionization (LDI) silicon nanopost array chips fabricated using deep UV projection lithography and deep reactive ion etching. RSC Adv. 2015;5(88):72051–7.
Korte AR, Stopka SA, Morris N, Razunguzwa T, Vertes A. Large-scale metabolite analysis of standards and human serum by laser desorption ionization mass spectrometry from silicon nanopost arrays. Anal Chem. 2016;88(18):8989–96.
Picca RA, Calvano CD, Lo Faro MJ, Fazio B, Trusso S, Ossi PM, Neri F, D’Andrea C, Irrera A, Cioffi N. Functionalization of silicon nanowire arrays by silver nanoparticles for the laser desorption ionization mass spectrometry analysis of vegetable oils. J Mass Spectrom. 2016;51(9):849–56.
Alhmoud HZ, Guinan TM, Elnathan R, Kobus H, Voelcker NH. Surface-assisted laser desorption/ionization mass spectrometry using ordered silicon nanopillar arrays. Analyst. 2014;139:5999–6009.
Marsico ALM, Duncan B, Landis RF, Tonga GY, Rotello VM, Vachet RW. Enhanced laser desorption/ionization mass spectrometric detection of biomolecules using gold nanoparticles, matrix, and the coffee ring effect. Anal Chem. 2017;89(5):3009–14.
Abdelhamid HN, Wu H-F. Gold nanoparticles assisted laser desorption/ionization mass spectrometry and applications: from simple molecules to intact cells. Anal Bioanal Chem. 2016;408(17):4485–502.
Dufresne M, Masson J-F, Chaurand P. Sodium-doped gold-assisted laser desorption ionization for enhanced imaging mass spectrometry of triacylglycerols from thin tissue sections. Anal Chem. 2016;88(11):6018–25.
Marsico ALM, Creran B, Duncan B, Elci SG, Jiang Y, Onasch TB, Wormhoudt J, Rotello VM, Vachet RW. Inkjet-printed gold nanoparticle surfaces for the detection of low molecular weight biomolecules by laser desorption/ionization mass spectrometry. J Am Soc Mass Spectrom. 2015;26(11):1931–7.
Sekula, J, Niziol, J, Rode, W, Ruman T. Gold nanoparticle-enhanced target (AuNPET) as universal solution for laser desorption/ionization mass spectrometry analysis and imaging of low molecular weight compounds. Anal Chim Acta. 2015;875:61–72.
Sekuła J, Nizioł J, Rode W, Ruman T. Silver nanostructures in laser desorption/ionization mass spectrometry and mass spectrometry imaging. Analyst. 2015;140(18):6195–209.
Schnapp A, Niehoff AC, Koch A, Dreisewerd K. Laser desorption/ionization mass spectrometry of lipids using etched silver substrates. Methods. 2016;104:194–203.
Popović IA, Nešić M, Vranješ M, Šaponjić Z, Petković M. SALDI-TOF-MS analyses of small molecules (citric acid, dexasone, vitamins E and A) using TiO2 nanocrystals as substrates. Anal Bioanal Chem. 2016;408(26):7481–90.
Wu Q, Chu JL, Rubakhin SS, Gillette MU, Sweedler J V. Dopamine-modified TiO2 monolith-assisted LDI MS imaging for simultaneous localization of small metabolites and lipids in mouse brain tissue with enhanced detection selectivity and sensitivity. Chem Sci. 2017;8(5):3926–38.
Popović I, Milovanović D, Miletić J, Nešić M, Vranješ M, Šaponjić Z, Petković M. Dependence of the quality of SALDI TOF MS analysis on the TiO2 nanocrystals’ size and shape. Opt Quantum Electron. 2016;48(2):113.
Yang M-R, Wang M, Xiao-Yan T, Zhou Jian MX-F. Analysis of small molecule compounds by matrix assisted laser desorption ionization time-of-flight mass spectrometry with ZnO, CuO, and NiO nanoparticles as matrix. Chinese J Anal Chem. 2015;43:1058–62.
Li Z, Zhang Y-W, Xin Y-L, Bai Y, Zhou H-H, Liu H-W. A lithium-rich composite metal oxide used as a SALDI-MS matrix for the determination of small biomolecules. Chem Commun. 2014;50(50):15397–9.
Unnikrishnan B, Chang C-Y, Chu H-W, Anand A, Huang C-C. Functional gold nanoparticles coupled with laser desorption ionization mass spectrometry for bioanalysis. Anal Methods. 2016;8(46):8123–33.
Picca RA, Calvano CD, Cioffi N, Palmisano F. Mechanisms of nanophase-induced desorption in LDI-MS. A SHORT REVIEW. Nanomater (Basel, Switzerland). 2017;7(4):75–81.
Dong X, Cheng J, Li J, Wang Y. Graphene as a novel matrix for the analysis of small molecules by MALDI-TOF MS. Anal Chem. 2010;82(14):6208–14.
Ma N, Bian W, Li R, Geng H, Zhang J, Dong C, Shuang S, Cai Z. Quantitative analysis of nitro-polycyclic aromatic hydrocarbons in PM 2.5 samples with graphene as a matrix by MALDI-TOF MS. Anal Methods. 2015;7(9):3967–71.
Friesen WL, Schultz BJ, Destino JF, Alivio TEG, Steet JR, Banerjee S, Wood TD. Two-dimensional graphene as a matrix for MALDI imaging mass spectrometry. J Am Soc Mass Spectrom. 2015;26(11):1963–6.
Zhou D, Guo S, Zhang M, Liu Y, Chen T, Li Z. Mass spectrometry imaging of small molecules in biological tissues using graphene oxide as a matrix. Anal Chim Acta. 2017;962:52–9.
Wang Z, Cai Y, Wang Y, Zhou X, Zhang Y, Lu H. Improved MALDI imaging MS analysis of phospholipids using graphene oxide as new matrix. Sci Rep. 2017;7:44466.
Zheng X, Zhang J, Wei H, Chen H, Tian Y, Zhang J. Determination of dopamine in cerebrospinal fluid by MALDI-TOF mass spectrometry with a functionalized graphene oxide matrix. Anal Lett. 2016;49(12):1847–61.
Cai L, Sheng L, Xia M, Li Z, Zhang S, Zhang X, Chen H. Graphene oxide as a novel evenly continuous phase matrix for TOF-SIMS. J Am Soc Mass Spectrom. 2017;28(3):399–408.
Zhang J, Zheng X, Ni Y. Selective enrichment and MALDI-TOF MS analysis of small molecule compounds with vicinal diols by boric acid-functionalized graphene oxide. J Am Soc Mass Spectrom. 2015;26(8):1291–8.
Shih Y-H, Chien C-H, Singco B, Hsu C-L, Lin C-H, Huang H-Y. Metal-organic frameworks: new matrices for surface-assisted laser desorption–ionization mass spectrometry. Chem Commun. 2013;49(43):4929.
Chen L, Ou J, Wang H, Liu Z, Ye M, Zou H. Tailor-made stable Zr(IV)-based metal-organic frameworks for laser desorption/ionization mass spectrometry analysis of small molecules and simultaneous enrichment of phosphopeptides. ACS Appl Mater Interfaces. 2016;8:20292–300.
Wang S, Niu H, Zeng T, Zhang X, Cao D, Cai Y. Rapid determination of small molecule pollutants using metal-organic frameworks as adsorbent and matrix of MALDI-TOF-MS. Microporous Mesoporous Mater. 2017;239:390–5.
Armstrong DW, Zhang L-K, He L, Gross ML. Ionic liquids as matrixes for matrix-assisted laser desorption/ionization mass spectrometry. Anal Chem. 2001;73:3679–86.
Li YL, Gross ML. Ionic-liquid matrices for quantitative analysis by MALDI-TOF mass spectrometry. J Am Soc Mass Spectrom. 2004;15(12):1833–7.
Crank JA, Armstrong DW. Towards a second generation of ionic liquid matrices (ILMs) for MALDI-MS of peptides, proteins, and carbohydrates. J Am Soc Mass Spectrom. 2009;20(10):1790–800.
Cramer R, Corless S. Liquid ultraviolet matrix-assisted laser desorption/ionization - mass spectrometry for automated proteomic analysis. Proteomics. 2005;5(2):360–70.
Towers MW, Mckendrick JE, Cramer R. Introduction of 4-chloro-α-cyanocinnamic acid liquid matrices for high sensitivity UV-MALDI MS. J Proteome Res. 2010;9(4):1931–40.
Mank M, Stahl A, Boehm G. 2,5-Dihydroxybenzoic acid butylamine and other ionic liquid matrixes for enhanced MALDI-MS analysis of biomolecules. Anal Chem. 2004;76:2938–50.
Tholey A. Ionic liquid matrices with phosphoric acid as matrix additive for the facilitated analysis of phosphopeptides by matrix-assisted laser desorption/ionization mass spectrometry. Rapid Commun Mass Spectrom. 2006;20(11):1761–8.
Laremore T, Murugesan S, Park, TJ, Avci, FY, Zagorevski D, Linhardt R. Matrix-Assisted Laser Desorption/Ionization Mass Spectrometric Analysis of Uncomplexed Highly Sulfated Oligosaccharides Using Ionic Liquid Matrices. Anal Chem 2006;78:1774–9.
Fukuyama Y, Nakaya S, Yamazaki Y, Tanaka K. Ionic liquid matrixes optimized for MALDI-MS of sulfated/sialylated/neutral oligosaccharides and glycopeptides. Anal Chem. 2008;80:2171–9.
Li YL, Gross ML, Hsu F-F. Ionic-liquid matrices for improved analysis of phospholipids by MALDI-TOF mass spectrometry. J Am Soc Mass Spectrom. 2005;16(5):679–82.
Tholey A, Heinzle E. Ionic (liquid) matrices for matrix-assisted laser desorption/ionization mass spectrometry-applications and perspectives. Anal Bioanal Chem. 2006;386:24–37.
Towers MW, Cramer R. Ionic liquids and other liquid matrices for sensitive MALDI MS analysis. Advances in MALDI and Laser-Induced Soft Ionization Mass Spectrometry. Springer International Publishing, Cham; 2016. p. 51–64.
Calvano CD, Carulli S, Palmisano F. Aniline/α-cyano-4-hydroxycinnamic acid is a highly versatile ionic liquid for matrix-assisted laser desorption/ ionization mass spectrometry. Rapid Commun Mass Spectrom. 2009;23:1659–68.
Shrivas K, Tapadia K. Ionic liquid matrix-based dispersive liquid–liquid microextraction for enhanced MALDI–MS analysis of phospholipids in soybean. J Chromatogr B. 2015;1001:124–30.
Calvano CD, Ceglie CD, D’Accolti L, Zambonin C. MALDI-TOF mass spectrometry detection of extra-virgin olive oil adulteration with hazelnut oil by analysis of phospholipids using an ionic liquid as matrix and extraction solvent. Food Chem. 2012;134(2):1192–8.
Mukherjee G, Röwer C, Koy C, Protzel C, Lorenz P, Thiesen H-J, Hakenberg OW, Glocker MO. Ultraviolet Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry for Phosphopeptide Analysis with a Solidified Ionic Liquid Matrix. Eur J Mass Spectrom. 2015;21(2):65–77.
Bronzel JL, Milagre CDF, Milagre HMS. Analysis of low molecular weight compounds using MALDI- and LDI-TOF-MS: Direct detection of active pharmaceutical ingredients in different formulations. J Mass Spectrom. 2017;52(11):752–8.
Do TD, Comi TJ, Dunham SJB, Rubakhin SS, Sweedler J V. Single Cell Profiling Using Ionic Liquid Matrix-Enhanced Secondary Ion Mass Spectrometry for Neuronal Cell Type Differentiation. Anal Chem. 2017;89(5):3078–86.
Shrivas K, Hayasaka T, Goto-Inoue N, Sugiura Y, Zaima N, Setou M. Ionic Matrix for Enhanced MALDI Imaging Mass Spectrometry for Identification of Phospholipids in Mouse Liver and Cerebellum Tissue Sections. Anal Chem. 2010; 82(21):8800–6.
Meriaux C, Franck J, Wisztorski M, Salzet M, Fournier I. Liquid ionic matrixes for MALDI mass spectrometry imaging of lipids. J Proteomics 2010;73(6):1204–18.
Abdelhamid HN, Khan MS, Wu H.-F. Design, characterization and applications of new ionic liquid matrices for multifunctional analysis of biomolecules: A novel strategy for pathogenic bacteria biosensing. Anal Chim Acta 2014;823:51–60.
Cheng X, Ye X, Liu D, Zhao N, Gao H, Wang P, Ge GZX. Novel ionic liquid matrices for qualitative and quantitative detection of carbohydrates by matrix assisted laser desorption/ionization mass spectrometry. Anal Chim Acta. 2017;985:114–20.
Moon JH, Park KM, Ahn SH, Lee SH, Kim MS. Investigations of Some Liquid Matrixes for Analyte Quantification by MALDI. J Am Soc Mass Spectrom. 2015;26(10):1657–64.
McCarley TD, McCarley RL, Limbach PA. Electron-Transfer Ionization in Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry. Anal Chem. 1998;70:4376–9.
Macha SF, Limbach PA, Savickas P. Application of nonpolar matrices for the analysis of low molecular weight nonpolar synthetic polymers by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J Am Soc Mass Spectrom. 2000;11(8):731–7.
Nazim Boutaghou M, Cole RB. 9,10-Diphenylanthracene as a matrix for MALDI-MS electron transfer secondary reactions. J Mass Spectrom. 2012;47:995–1003.
Asakawa D, Lee CC, Hiraoka K. Negative-mode MALDI mass spectrometry for the analysis of pigments using tetrathiafulvalene as a matrix. J Mass Spectrom. 2008;43:1494–501.
Ulmer L, Mattay J, Torres-Garcia HG, Luftmann H. Letter: The Use of 2-[(2 E )-3-(4- Tert -Butylphenyl)-2-Methylprop-2-Enylidene]Malononitrile as a Matrix for Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry. Eur J Mass Spectrom. 2000;6(1):49–52.
Wyatt MF, Stein BK, Brenton AG. Characterization of various analytes using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry and 2-[(2E)-3-(4-tert- butylphenyl)-2-methylprop-2-enylidene]malononitrile matrix. Anal Chem. 2006;78:199–206.
Castellanos-García LJ, Agudelo BC, Rosales HF, Cely M, Ochoa-Puentes C, Blanco-Tirado C, Sierra CA, Combariza MY. Oligo p-Phenylenevinylene Derivatives as Electron Transfer Matrices for UV-MALDI. J Am Soc Mass Spectrom. 2017;28(12):2548–60.
Calvano CD, Ventura G, Cataldi TRI, Palmisano F. Improvement of chlorophyll identification in foodstuffs by MALDI ToF/ToF mass spectrometry using 1,5-diaminonaphthalene electron transfer secondary reaction matrix. Anal Bioanal Chem. 2015;407(21):6369–79.
Calvano CD, Ventura G, Trotta M, Bianco G, Cataldi TRI, Palmisano F. Electron-Transfer Secondary Reaction Matrices for MALDI MS Analysis of Bacteriochlorophyll a in Rhodobacter sphaeroides and Its Zinc and Copper Analogue Pigments. J Am Soc Mass Spectrom. 2016;28:125–35.
Acknowledgments
This work was supported by the project PONa3 00395/1 “BIOSCIENZE & SALUTE (B&H)” of the Ministero Italiano per l’Istruzione, l’Università e la Ricerca (MIUR).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest. This article does not contain any studies with human or animal subjects.
Rights and permissions
About this article
Cite this article
Calvano, C.D., Monopoli, A., Cataldi, T.R.I. et al. MALDI matrices for low molecular weight compounds: an endless story?. Anal Bioanal Chem 410, 4015–4038 (2018). https://doi.org/10.1007/s00216-018-1014-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-018-1014-x