Abstract
Phospholipids (PL) are increasingly analyzed by matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry (MS). As in the case of polar molecules, however, the careful selection of the matrix is crucial for optimum results. 9-Aminoacridine (9-AA) was recently suggested as the matrix of choice to analyze PL mixtures because of (a) the improved sensitivity and (b) the reduction of suppression effects compared to other matrices. However, the distinction of phosphatidylcholine (PC) and phosphatidylethanolamine (PE) in the negative ion mode is obscured as PC is also detectable as –CH +3 ion if 9-AA is used as matrix. This may result in the erroneous assignment of PC as a PE species. Using an organic extract from hen egg yolk as example it will be shown that the contribution of PC must be taken into consideration if the negative ion mass spectra are used to evaluate the fatty acyl compositions of PE mixtures. 9-AA can as well be used in hyphenated thin-layer chromatography (TLC)-MALDI-TOF MS where PC and PE are chromatographically well separated for unequivocal assignments.

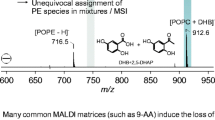
Comparison of negative ion MALDI-TOF mass spectra of isolated 1-palmitoyl-2-oleoyl-sn-phosphatidylcholine (POPC) and 1-palmitoyl-2-oleoyl-sn-phosphatidylethanolamine (POPE) using either DHB (blue) or 9-AA (red) as matrix. The spectra differ significantly as a function of the matrix used. In case of 9-AA, POPC is detectable as negative ion subsequent to the loss of a -CH3 group, which complicates peak assignments when complex mixtures are analyzed
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phospholipids (PLs) are important constituents of all cellular membranes and changes of the PL compositions, particularly the generation of lysophospholipids (lacking one fatty acyl residue in comparison to the original PL), are increasingly considered to correlate with different pathological, for instance, inflammatory conditions. Thus, PLs and their analysis are attracting considerable medical as well as diagnostic interest [1].
Because of this significant interest, different methods of lipid analysis are established and comprise chromatographic (HPLC or high-performance thin-layer chromatography (HPTLC)), spectroscopic (e.g., nuclear magnetic resonance (NMR)), and mass spectrometric (MS) methods [2]. Due to its high sensitivity, MS is regarded as one of the most powerful methods of lipid analysis [3] and the term "lipidomics" has been recently introduced [4]. Using "soft ionization" techniques, basically all lipid classes can be analyzed without a major extent of fragmentation. Although electrospray ionization (ESI) MS [5] was primarily used so far, there is growing evidence that matrix-assisted laser desorption and ionization time-of-flight (MALDI-TOF) MS represents a useful alternative [6, 7]. The interest in PL analysis by MALDI MS has additionally increased since MALDI MS "imaging" methods have been established: PLs present in the cellular membranes of tissues are particularly easily ionized and, thus, more sensitively detectable than, e.g., proteins [8, 9] and other polar molecules.
Although the ionization process of MALDI is only poorly understood so far, it is commonly accepted that the careful choice of the matrix has a tremendous effect on the achievable quality of MALDI-TOF mass spectra [10]. While 2,5-dihydroxybenzoic acid (DHB) was predominantly used as matrix in lipid studies [11], the search for more efficient matrices continuous: trihydroxyacetophenone (THA) [12], p-nitroaniline (PNA) [13] or ionic liquid matrices [14] were introduced.
Although 9-aminoacridine (9-AA) is known to represent a particularly useful matrix for smaller molecules due to its moderate background [15], the advantages of this matrix in the lipid field were only recently recognized. Sun et al. [16] have demonstrated that 9-AA provides higher sensitivity than DHB and that protonated molecular ions are formed without interference by sodiated species. Additionally, in 9-AA preparations, uncharged lipids such as cholesterol or triacylglycerols are not detectable [16], which reduces the complexity of PL mass spectra of mixtures.
Due to its basicity (9-AA, pK = 9.99; DHB, pK = 2.97 [11]), 9-AA is also highly suitable for negative ion MALDI MS. This allows the analysis of phosphatidylethanolamines (PEs) as negative ions avoiding the suppression of PE in the presence of phosphatidylcholine (PC) as observed in positive ion mode [7]. The same approach using the basic matrix para-nitroaniline (PNA) [13], suffered from high volatility, short MALDI observation times, and signal stability [17].
This paper has two different aims: the first aim is to demonstrate that PC is detectable as negative ion if 9-AA is used as matrix. Organic extracts from biological samples normally contain PC in excess over PE and thus a potential contribution of PC in negative ion mass spectra of other lipids in complex mixtures may occur. So far, only very few reports are dealing with the negative ion MALDI MS detection of PC. In the first study, Marto et al. [18] have shown that PC is detectable as negative ion only if trans-4-hydroxy-3-methoxycinnamic acid (TCA) is used as matrix. The loss of a methyl group [M-15]−, loss of a quaternary amine [M-60]− and loss of the choline headgroup [M-86]− were also observed to be very much alike fast atom bombardment (FAB) spectra [19]. Our group has recently shown that use of a cinnamic acid derivate (Cl-CCA) [20] provides for the detection of PC as [M-H]− whereas with DHB matrix, PC is detected as low abundant negatively charged matrix cluster ion [21].
In this work, we established that the 9-AA matrix provides intense [M-CH3]− ions of PC and that the analysis of the negative ion MALDI mass spectra of PL mixtures with a significant PC content must consider the erroneous assignment of PC as a PE species.
The second aim of this paper is to evaluate the capabilities of 9-AA as matrix for the direct analysis of thin-layer chromatography plates by MALDI MS [22]. Although the use of 9-AA results in excellent negative ion mass spectra of acidic lipids, it was not possible to detect compounds with quaternary ammonium groups, such as PC, as negative ions directly from the TLC plate. Although this behavior is not yet completely clear, some potential explanations will be provided and future investigations discussed.
Materials and methods
Chemicals
All chemicals for sample preparation and all solvents (chloroform, methanol, acetonitrile, and isopropanol) were obtained in the highest commercially available purity from Fluka Feinchemikalien GmbH (part of Sigma-Aldrich Chemie GmbH, Taufkirchen, Germany) and used as supplied. The applied matrices were either from Fluka (DHB) or from ACROS Organics, Morris Plains, NJ, USA (9-AA).
Phospholipid standards 1-palmitoyl-2-oleoyl-sn-phosphatidylcholine (POPC) and 1-palmitoyl-2-oleoyl-sn-phosphatidylethanolamine (POPE) were obtained from Avanti Polar Lipids (Alabaster, AL, USA) as 10 mg/ml solutions in CHCl3. They were dried in a vacuum centrifuge ("Speedvac", Jouan, Germany) and re-dissolved in a mixture of isopropanol/acetonitrile (60:40, v/v) for best results [16].
The total PL concentration was in all cases fixed at 1 mg/ml and the POPE contribution was varied between 0 and 40%. The residual PL was POPC. This excess of POPC was used because it is very typical of biological systems and mimics, for instance, the composition of hen egg yolk that will be used in this study as a typical PL mixture [23].
Lipid extraction from hen egg yolk
Hen eggs were purchased from local supermarkets. Twelve different eggs were extracted and the individual extracts were pooled in order to minimize inter-individual deviations.
The egg white and the egg yolk were separated, and the egg yolk was extracted according to the Bligh and Dyer method, i.e., a ratio between chloroform, methanol, and the aqueous phase of 1:1:1 (v/v/v) was used [24]. Further details including the typical composition of the egg yolk extract are available from our previous work [23].
High-performance thin-layer chromatography
Lipid extracts were subjected to HPTLC prior to MALDI-TOF MS. Five-microliter samples (overall amount of lipids about 35 µg) of the total extract was applied on HPTLC silica gel 60 plates (200 µm layer thickness, 10 × 10 cm in size on glass backs, Merck, Darmstadt, Germany) and developed in vertical ascending TLC chambers using chloroform, ethanol, water, triethylamine (35:35:7:35, v/v/v/v) as the solvent system.
Lipids were visualized by spraying with a solution of primuline (Direct Yellow) according to White et al. [25]. Upon excitation by UV light (366 nm), individual lipids become detectable as colored spots. These spots were assessed by using a digital image system in combination with the program Argus X1 delivered by BioStep (Jahnsdorf, Germany).
The identified individual spots were carefully scratched off and the PL eluted by the addition of a mixture of 75 µl CHC13, 75 µl CH3OH, and 75 µl 0.9% NaCl in water and intense vortexing. Afterwards, samples were centrifuged (2,500 revolutions per minute) to allow phase separation. The organic layer was removed and mixed 1:1 (v/v) with the matrix solution. One microliter of that mixture was applied to the MALDI target and directly used for MALDI-TOF MS. In addition, lipids were also characterized directly on the TLC plate as essentially described in [23].
MALDI-TOF mass spectrometry
All spectra were recorded with either DHB (0.5 M in methanol) or 9-AA (10 mg/ml in isopropanol/acetonitrile (60:40, v/v)) and all samples were diluted 1:1 (v/v) with the corresponding matrix solutions. The mixture was subsequently directly applied onto a gold-coated MALDI target.
All MALDI-TOF mass spectra were acquired on an Autoflex I mass spectrometer (Bruker Daltonics, Bremen, Germany) with ion reflector. The system utilizes a pulsed 50 Hz nitrogen laser, emitting at 337 nm. The extraction voltage was 20 kV and gated matrix suppression was applied to prevent the saturation of the detector by matrix ions. All spectra were acquired in reflector mode using delayed extraction. A more detailed description of MALDI-TOF MS—in particular of PLs—is available in [6, 7]. Spectral mass resolutions, signal-to-noise ratios, and peak intensities were determined by the instrument software "Flex Analysis 3.0" (Bruker Daltonics). The mass spectrometer was calibrated using the molecular ions of a standard lipid mixture desorbed from a standard DHB (positive ions) or 9-AA (negative ions) preparation applied next to the spots of interest. Post source decay (PSD) mass spectra were recorded in selected cases as described in [26].
Results and discussion
In Fig. 1, selected positive and negative ion MALDI-TOF mass spectra of isolated POPC and POPE are shown in order to illustrate the effect of different matrices.
Comparison of the positive (+) and negative (−) ion MALDI-TOF mass spectra of POPC and POPE prepared either in DHB (left) or 9-AA (right). The headgroups of both PLs are indicated on the right and DHB matrix-related peaks are marked by asterisks [11]. For details, see the text
These two PLs were selected because they occur in significant amounts in cells, tissues and body fluids. Spectra were obtained using DHB or 9-AA, in negative or positive ion mode. It is obvious from these spectra that the influence of matrix selection is profound and can be summarized as follows:
-
1.
POPC spectra in DHB (positive ion mode) provide intense MH+ and MNa+ ions and some weak fragment ions (1a) [27]. In 9-AA (1b), the spectrum is even stronger dominated by MH+ ion formation in agreement with reference data [16].
-
2.
The positive ion spectra of POPE are dominated by MNa+ and [M-H + 2Na]+ molecular ions, with MH+ (m/z = 718.5) only visible in case of DHB. The PE headgroup neutral ion losses at m/z = 577.5 and 603.5 [26] seem to originate from MH+, but not from MNa+ as MH+ is not detectable in 9-AA preparations. A similar behavior was already observed for triacylglycerols [28].
-
3.
In the negative ion spectra of POPE, [POPE-H]− at m/z = 716.5 is a rather weak peak amongst many others if DHB is used as matrix (1e), accompanied by a POPE + DHB cluster ion at m/z = 892.5 [29]. The intense signals at m/z = 675, 681, and 857 (marked by asterisks) are DHB-derived ions that obscure the negative ion mode spectra [11] due to their considerable intensities. In contrast, 9-AA (1f) provides a single intense [POPE-H]− signal at m/z = 716.5 and no other ion species are generated. This difference can be easily explained by the high basicity of 9-AA in contrast to DHB.
-
4.
The most significant difference is observed if the negative ion spectra of POPC recorded in the presence of 9-AA and DHB are compared. In the presence of DHB (1g), only m/z = 912.6, a DHB + POPC cluster ion [21], is related to POPC. [POPC-H]− (m/z = 758.6) or [POPC-CH3]− (m/z = 744.6) are not detectable. In contrast, an intense [POPC-CH3]− peak is present in the 9-AA preparation (1 h). The identity of this ion was confirmed by MS/MS: an intense PSD signal at m/z = 168.0 indicates the deprotonated phosphorylcholine (data not shown). Although the signals at m/z = 754.6 and 821.5 cannot be assigned so far, the fragment ion formation of m/z = 673.5 and 699.5 is suggested in Fig. 2.
POPC, with its quaternary amine, stabilizes a positive charge quite strongly [30]. Therefore, while POPE is readily detectable as a negative ion, the negative ion generation of POPC occurs only after elimination of CH +3 or other cations.
In order to compare the negative ion yields of POPC- and POPE-related ions, PL mixtures of known compositions were prepared and negative ion mode spectra recorded (Fig. 3).
Negative ion MALDI-TOF mass spectra of defined mixtures between POPC and POPE. The total lipid concentration was in all cases 1 mg/ml and mixed 1:1 with 10 mg/ml 9-AA (in 60% isopropanol, 40% acetonitrile, v/v) prior to measurements. Trace (a) represents pure POPC, while the POPE content was 5% in (b), 10% (c), 15% (d), 20% (e), 25% (f), 30% (g), and 35% (h). The peak at m/z = 716.5 corresponds to [PE 16:0/18:1-H]− and the peak at m/z = 744.6 to [PC 16:0/18:1 -CH3]−
An excess of POPC in comparison to POPE was used in all cases because (a) the detectability of POPE as negative ion is obviously much higher in comparison to POPC and (b) organic extracts of biological samples are normally characterized by a higher PC than PE content [22]. Thus, a total PL concentration of 1 mg/ml was used in all cases and the POPE moiety varied between 0 (3a, 1 mg/ml POPC) and 35% (3 h, 0.65 mg/ml POPC and 0.35 mg/ml POPE).
It is obvious from Fig. 3 that the ratio between the signal intensities of [POPC-CH3]− (m/z = 744.6 as the most intense one) and [POPE-H]− (m/z = 716.5) is about 1:1 if the POPE accounts for as little as 5% indicating that POPE is about 20× more sensitively detectable as negative ion than POPC, and a quantitative evaluation of the data is given in Fig. 4 showing a nearly linear relationship between the POPE content of the mixture and the ratio of the signal intensities.
Plot generated from the spectra shown in Fig. 3. The intensity ratios of the peaks at m/z = 716.5 and 744.6 are plotted against the relative POPE content (in percent) in a POPE/POPC mixture. Linear regression was performed as a guide to the eye and slope (A), intercept (B), and regression coefficient (R) are provided
The standard deviations of the peak ratios determined from three different samples were always of the order of ±8%. This confirms a previous study where the achievable accuracies of PL concentrations determined either by MALDI or ESI were compared: No significant differences could be found [16] confirming that ESI and MALDI provide comparably reliable data. These data show the huge difference of the ionization efficiency between POPE and POPC and certainly permit an intra-spectral relative quantitative estimate, but are actually not intended to open a general quantitative discussion of TLC-MALDI.
The detectability of PC as negative fragment ion is of practical interest because many biologically relevant lipid mixtures are characterized by a huge excess of PC in comparison to PE as well as all further lipids. Such a typical PL mixture is the organic extract from hen egg yolk [23], which is further characterized below. Nevertheless, no fundamental differences are expected if clinically more relevant lipid samples (e.g. cellular extracts) would be investigated.
It was recently demonstrated that 9-AA allows the estimation of the relative fatty acyl compositions of the individual PL classes in complex mixtures [16]. However, the potential contribution of negative ions derived from PC was so far not considered at all. In order to evaluate whether this effect plays a significant role, the negative ion MALDI-TOF mass spectra of the total egg yolk extract (5a) and the isolated PE fraction (5b) are compared in Fig. 5.
Both spectra were exclusively recorded in the presence of 9-AA as matrix because DHB is by far less suitable for the detection of PEs as negative ions. It is obvious from the comparison of both spectra that there are three major differences:
-
1.
The lipid extract (5a) contains various PE signals: PE 16:0/18:1 (m/z = 716.5), PE 18:0/18:2 (m/z = 742.5), PE 18:0/18:1 (m/z = 744.5), and PE 18:0/20:4 (m/z = 766.5). In addition, the two major signals at m/z = 885.5 and 861.5 correspond to phosphatidylinositols PI 18:0/20:4 and PI 18:0/18:2, respectively. The PIs are negatively charged PL and, therefore, provide intense [M-H]− peaks even at concentrations much lower than the PE concentration [23]. PI is absent from the PE fraction (5b).
-
2.
The signals at m/z = 673.5 and 699.5 in the total extract derive from POPC (cf. trace 1 h), the most abundant PC species of hen egg yolk [23]. PC is absent from the PE fraction.
-
3.
The intensity ratios of the peaks at m/z = 742.5 (PE 18:0/18:2) or 744.5 (PE 18:0/18:1) vs. m/z = 766.5 (PE 18:0/20:4) are much higher in the case of the total extract (5a) compared to the PE fraction (5b) due to overlap with negative ions of PC species such as PC 16:0/18:2 (m/z = 742.5) and PC 16:0/18:1 (m/z = 744.5).
Therefore, the contribution of PC-derived negative ions must be taken into account if the fatty acyl compositions of PEs are estimated from the negative ion spectra of the total organic extract. As the masses of [PE 18:0/18:1-H]− (m/z = 744.5) and [PC 16:0/18:1-CH3]− (m/z = 744.5) are isobaric, a clear discrimination is not possible by increased mass accuracy and resolution but rather by lipid group or chromatographic separation.
However, the fatty acyl profile of PE obviously differs prior and subsequent to chromatography as is evident from the reduced POPE (m/z = 716.5) intensity in the PE fraction (Fig. 5b). A superposition with PC 16:0/16:1 is impossible in this case due to its absence from egg yolk extract [23]. This "effect of chromatography" [31] describes that saturated lipids stick more efficiently to silica gel and are only partially released by extraction with organic solvents. Thereby, highly unsaturated lipids are enriched chromatographically and this effect is obviously also relevant for moderately unsaturated lipid species. This effect applies for HPLC and TLC alike [31]. Because a detailed quantitative investigation of this effect was beyond the scope of this investigation, only qualitative data are provided here.
Finally, it should be noted that PC is not detectable as negative ion species if PNA instead of 9-AA is used (data not shown). This is an obvious advantage of this matrix, although PNA sublimates considerably under high vacuum conditions [17] reducing possible analysis time.
Though the acquisition of negative ion mass spectra is by far less popular than positive ion spectra, negative ion spectra are normally less complex and simpler to interpret as adducts are less abundant and less frequent. For instance, if a peak at m/z = 782.6 is observed in the positive ion spectrum from a biological sample, MH+ of PC 16:0/20:4 and MNa+ of PC 16:0/18:1 could not be distinguished [32]. Although this problem can be resolved by MS/MS, the negative ion spectrum would easily distinguish both PC species (m/z = 744.6 vs. 766.6).
Thus, we have finally tried to use 9-AA as matrix for the direct monitoring of the fatty acyl compositions of different PL classes on a developed TLC plate. This particularly provides the advantage that no loss of lipids by extraction can occur as the TLC plate is directly investigated. A typical TLC plate of hen egg yolk stained with primuline is shown in Fig. 6 and the detected PL classes are assigned [23].
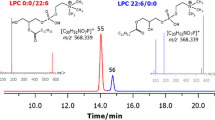
Primuline stained TLC plate of hen egg yolk extract (left) and negative ion MALDI-TOF mass spectra of the spots indicated on the TLC plate (right). 9-AA was added in dried droplet preparation onto the stained spots. Different fatty acyl compositions were monitored by spatially resolved MALDI measurements indicated by "PE (1)" and "PE (2)" [23]. LPC lyso-phosphatidylcholine, SM sphingomyelin, PC phosphatidylcholine, LPE lyso-phosphatidylethanolamine, PI phosphatidylinositol, PE phosphatidylethanolamine
Negative ion MALDI mass spectra (from the positions labeled by "a", "b", "c", and "d") were recorded in the same way as recently described [33] but 9-AA was used as matrix. All PL species containing a quaternary ammonium group with a permanent positive charge (LPC, SM, and PC) are only detectable as positive ions [34]. Corresponding negative ions, even the fragments that we observed in the model studies from MALDI targets (Fig. 1) were not detected from the TLC plate.
Possibly, the acidic surface of silica gel on the TLC plate [35] attenuates and even prevents the detection of LPC, SM and PC as negative ion species, while the more acidic PLs (LPE, PI, and PE) are readily ionized (Fig. 6).
Adding small amounts of aqueous NaOH to the matrix solution did not enhance the detection of these negative ion species but rather the hydrolysis of the PLs of interest. Although more alkaline matrices ("proton sponges") such as 1,8-bis(dimethylamino)naphthalene (TMDAN) with a pK value of 12.1 [36] were very effective for the detection of free fatty acids [37], we failed to detect PLs directly from the TLC plate as TMDAN is obviously a poor matrix for the detection of PLs.
Besides the choice of matrix for PLs, the influence of different solvent systems on spectra quality is striking. As shown by Sun et al. [16] the quality of MALDI mass spectra recorded in the presence of 9-AA strongly depends on the solvent system. Among the tested solvents, such as methanol, isopropanol, and acetonitrile, and various mixtures of these solvents, a mixture of isopropanol and acetonitrile provided the best results (see Fig. 3).
Similar trends were also observed on the TLC surface although it seems not very likely that the solvent composition is the only important parameter. Though further studies are clearly required, it can be recommended to use different matrix systems such as DHB and 9-AA in order to obtain maximum information.
As the contribution of PC species must be taken into account if the fatty acyl composition of the PE moiety of complex lipid mixtures is determined from the total extract in negative ion mode using 9-AA, previous separation into the individual PL classes is recommended.
The direct readout of PEs from TLC-MALDI MS datasets [23, 33] is greatly facilitated in negative ion mode using 9-AA as the spectra are straightforward to interpret and PCs are entirely separated. Therefore, any overlap of spectral assignments from PEs and PCs is eliminated providing a possible high resolution platform for PL quantification in the future.
References
Fuchs B, Schiller J (2009) Lysophospholipids: their generation, physiological role and detection. Are they important disease markers? Mini-Rev Med Chem 9:368–378
Peterson BL, Cummings BS (2006) A review of chromatographic methods for the assessment of phospholipids in biological samples. Biomed Chromatogr 20:227–243
Watson AD (2006) Thematic review series: systems biology approaches to metabolic and cardiovascular disorders. Lipidomics: a global approach to lipid analysis in biological systems. J Lipid Res 47:2101–2111
Wenk MR (2005) The emerging field of lipidomics. Nat Rev Drug Discov 4:594–610
Pulfer M, Murphy RC (2003) Electrospray mass spectrometry of phospholipids. Mass Spectrom Rev 22:332–364
Schiller J, Süß R, Arnhold J, Fuchs B, Leßig J, Müller M, Petković M, Spalteholz H, Zschörnig O, Arnold K (2004) Matrix-assisted laser desorption and ionization time-of-flight (MALDI-TOF) mass spectrometry in lipid and phospholipid research. Prog Lipid Res 43:449–488
Schiller J, Süß R, Fuchs B, Müller M, Zschörnig O, Arnold K (2007) MALDI-TOF MS in lipidomics. Front Biosci 12:2568–2579
Puolitaival SM, Burnum KE, Cornett DS, Caprioli RM (2008) Solvent-free matrix dry-coating for MALDI imaging of phospholipids. J Am Soc Mass Spectrom 19:882–886
Murphy RC, Hankin JA, Barkley RM (2009) Imaging of lipid species by MALDI mass spectrometry. J Lipid Res 50:S317–S322
Hillenkamp F, Peter-Katalinić J (2007) MALDI MS - A practical guide to instrumentation, methods and application. Wiley, Weinheim
Schiller J, Süß R, Fuchs B, Müller M, Petković M, Zschörnig O, Waschipky H (2007) The suitability of different DHB isomers as matrices for the MALDI-TOF MS analysis of phospholipids: which isomer for what purpose? Eur Biophys J 36:517–527
Stubiger G, Belgacem O (2007) Analysis of lipids using 2,4,6 trihydroxy-acetophenone as a matrix for MALDI mass spectrometry. Anal Chem 79:3206–3213
Estrada R, Yappert MC (2004) Alternative approaches for the detection of various phospholipid classes by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J Mass Spectrom 39:412–422
Li YL, Gross ML, Hsu FF (2005) Ionic-liquid matrices for improved analysis of phospholipids by MALDI-TOF mass spectrometry. J Am Soc Mass Spectrom 16:679–682
Vaidyanathan S, Goodacre R (2007) Quantitative detection of metabolites using matrix-assisted laser desorption/ionization mass spectrometry with 9-aminoacridine as the matrix. Rapid Commun Mass Spectrom 21:2072–2078
Sun G, Yang K, Zhao Z, Guan S, Han X, Gross RW (2008) Matrix-assisted laser desorption/ionization time-of-flight mass spectrometric analysis of cellular glycerophospholipids enabled by multiplexed solvent dependent analyte-matrix interactions. Anal Chem 80:7576–7585
Astigarraga E, Barreda-Gómez G, Lombardero L, Fresnedo O, Castaño F, Giralt MT, Ochoa B, Rodríguez-Puertas R, Fernández JA (2008) Profiling and imaging of lipids on brain and liver tissue by matrix-assisted laser desorption/ionization mass spectrometry using 2-mercaptobenzothiazole as a matrix. Anal Chem 80:9105–9114
Marto JA, White FM, Seldomridge S, Marshall AG (1995) Structural characterization of phospholipids by matrix-assisted laser desorption/ionization Fourier transform ion cyclotron resonance mass spectrometry. Anal Chem 67:3979–3984
Zirrolli JA, Clay KL, Murphy RC (1991) Tandem mass spectrometry of negative ions from choline phospholipid molecular species related to platelet activating factor. Lipids 26:1112–1116
Jaskolla T, Fuchs B, Karas M, Schiller J (2009) The new matrix 4-chloro-alpha-cyanocinnamic acid allows the detection of phosphatidylethanolamine chloramines by MALDI-TOF mass spectrometry. J Am Soc Mass Spectrom 20:867–874
Schiller J, Süß R, Petković M, Zschörnig O, Arnold K (2002) Negative-ion matrix-assisted laser desorption and ionization time-of-flight mass spectra of complex phospholipid mixtures in the presence of phosphatidylcholine: a cautionary note on peak assignment. Anal Biochem 309:311–314
Fuchs B, Süß R, Nimptsch A, Schiller J (2009) Matrix-assisted laser desorption and ionization time-of-flight mass spectrometry (MALDI-TOF MS) directly combined with thin-layer chromatography (TLC)—a review of the current state. CHROMATOGRAPHIA 69:95–105
Fuchs B, Schiller J, Süss R, Schürenberg M, Suckau D (2007) A direct and simple method of coupling matrix-assisted laser desorption and ionization time-of-flight mass spectrometry (MALDI-TOF MS) to thin-layer chromatography (TLC) for the analysis of phospholipids from egg yolk. Anal Bioanal Chem 389:827–834
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37:911–917
White T, Bursten S, Frederighi D, Lewis RA, Nudelman E (1998) High-resolution separation and quantification of neutral lipid and phospholipid species in mammalian cells and sera by multi-one-dimensional thin-layer chromatography. Anal Biochem 10:109–117
Fuchs B, Schober C, Richter G, Süss R, Schiller J (2007) MALDI-TOF MS of phosphatidylethanolamines: different adducts cause different post source decay (PSD) fragment ion spectra. J Biochem Biophys Methods 70:689–692
Fuchs B, Schiller J, Süß R, Nimptsch A, Schürenberg M, Suckau D (2009) Capabilities and disadvantages of combined matrix-assisted laser-desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) and high-performance thin-layer chromatography (HPTLC): analysis of egg yolk lipids. J Planar Chromatogr 22:35–42
Gidden J, Liyanage R, Durham B, Lay JO Jr (2007) Reducing fragmentation observed in the matrix-assisted laser desorption/ionization time-of-flight mass spectrometric analysis of triacylglycerols in vegetable oils. Rapid Commun Mass Spectrom 21:1951–1957
Petković M, Schiller J, Müller M, Süß R, Arnold K, Arnhold J (2009) Detection of adducts with matrix clusters in the positive and negative ion mode MALDI-TOF mass spectra of phospholipids. Z Naturforsch 64B:35–42
Cevć G (1993) Phospholipid handbook. Marcel-Dekker, New York
DeLong CJ, Baker PR, Samuel M, Cui Z, Thomas MJ (2001) Molecular species composition of rat liver phospholipids by ESI-MS/MS: the effect of chromatography. J Lipid Res 42:1959–1968
Schiller J, Süß R, Petković M, Hilbert N, Müller M, Zschörnig O, Arnhold J, Arnold K (2001) CsCl as an auxiliary reagent for the analysis of phospholipid mixtures by matrix-assisted laser desorption and ionization time-of-flight mass spectrometry (MALDI-TOF MS). Chem Phys Lipids 113:123–131
Fuchs B, Schiller J, Süß R, Zscharnack M, Bader A, Müller P, Schürenberg M, Becker M, Suckau D (2008) Analysis of stem cell lipids by offline HPTLC-MALDI-TOF MS. Anal Bioanal Chem 392:849–860
Petković M, Schiller J, Müller M, Benard S, Reichl S, Arnold K, Arnhold J (2001) Detection of individual phospholipids in lipid mixtures by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry: phosphatidylcholine prevents the detection of further species. Anal Biochem 289:202–216
Schiller J, Müller K, Süß R, Arnhold J, Gey C, Herrmann A, Leßig J, Arnold K, Müller P (2003) Analysis of the lipid composition of bull spermatozoa by MALDI-TOF mass spectrometry—a cautionary note. Chem Phys Lipids 126:85–94
Staab HA, Saupe T (1988) "Proton Sponges" and the geometry of hydrogen bonds: aromatic nitrogen bases with exceptional basicities. Angew Chem Int ed 27:865–879
Shroff R, Svatoš A (2009) 1, 8-Bis(dimethylamino)naphthalene: a novel superbasic matrix for matrix-assisted laser desorption/ionization time-of-flight mass spectrometric analysis of fatty acids. Rapid Commun Mass Spectrom 23:2380–2382
Acknowledgments
This work was supported by the German Research Council (DFG Schi 476/5-1, TR 67 subproject A2 and Fu 771/1-1) as well as the Federal Ministry of Education and Research (BMBF Grant 0313836).
We thank Dr. Thorsten Jaskolla (University of Frankfurt) for discussions and many useful hints on optimum matrix selection.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fuchs, B., Bischoff, A., Süß, R. et al. Phosphatidylcholines and -ethanolamines can be easily mistaken in phospholipid mixtures: a negative ion MALDI-TOF MS study with 9-aminoacridine as matrix and egg yolk as selected example. Anal Bioanal Chem 395, 2479–2487 (2009). https://doi.org/10.1007/s00216-009-3032-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-009-3032-1