Abstract
Fucosylation of N-glycoproteins has been implicated in various diseases, such as hepatocellular carcinoma (HCC). However, few studies have performed site-specific analysis of fucosylation in liver-secreted proteins. In this study, we characterized the fucosylation patterns of liver-secreted proteins in HCC plasma using a workflow to identify site-specific N-glycoproteins, where characteristic B- and/or Y-ion series with and without fucose in collision-induced dissociation were used in tandem mass spectrometry. In total, 71 fucosylated N-glycopeptides from 13 major liver-secreted proteins in human plasma were globally identified by LC-MS/MS. Additionally, 37 fucosylated N-glycopeptides were newly identified from nine liver-secreted proteins, including alpha-1-antichymotrypsin, alpha-1-antitrypsin, alpha-2-HS-glycoprotein, ceruloplasmin, alpha-1-acid glycoprotein 1/2, alpha-2-macroglobulin, serotransferrin, and beta-2-glycoprotein 1. Of the fucosylated N-glycopeptides, bi- and tri-antennary glycoforms were the most common ones identified in liver-secreted proteins from HCC plasma. Therefore, we suggest that this analytical method is effective for characterizing fucosylation in liver-secreted proteins.

A global map of fucosylated and non-fucosylated glycopeptides from 13 liver-secreted glycoproteins in hepatocellular carcinoma plasma
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protein glycosylation is involved in adhesion, metastasis, and signaling through cell–cell interactions. Glycosylation is divided into O-glycosylation and N-glycosylation based on binding groups (i.e., hydroxyl serine (S) or threonine (T) side chains and carboxyl-amido asparagine (N) residue side chains, respectively) [1]. N-Glycosylation, which is classified into three glycan types, high-mannose, hybrid, and complex [2], occurs at the consensus tri-peptide sequence NXS/T, where X can be any amino acid except proline, in the ER and Golgi apparatus [3]. Aberrant glycosylation of the complex type, including fucosylation by fucosyltransferases and additional branching by N-acetylglucosaminyltransferase V (GnTV), occurs in cancer cells and is involved in adhesion to other cells, invasion into the extracellular matrix, and metastasis [4]. The fucosylation of N-glycoproteins in hepatocellular carcinoma (HCC) tissue was reported to increase the transcript and protein levels of the fucosyltransferase genes Fut 6 and Fut 8, which are involved in glycan fucosylation [5–7].
Anderson and Anderson first reviewed 289 proteins with a dynamic range in concentrations of more than 10 orders of magnitude [8]; since then, a total of 1929 non-redundant proteins with a <1 % false discovery rate have been reported in the PeptideAtlas [9]. A number of these proteins are synthesized and secreted by the liver and function in hemostasis and fibrinolysis and as carriers of various small molecules, lipids, and hormones [10]. Additionally, most liver-enriched proteins were recently classified by the human protein atlas (HPA) project as secreted proteins using microarray-based immunohistochemistry [11]. Alpha feto-protein (AFP), a well-known HCC biomarker approved by the FDA, is synthesized in liver tissue and secreted into plasma [12]. In particular, fucosylated AFP (AFP-L3), an aberrant N-glycoprotein created via fucosylation that exhibits an increased affinity for the fucose-specific lectin Lens Culinaris Agglutinin (LCA), was reported as a better biomarker for HCC than AFP [13–17]. Fucosylated AFP is increased in human serum and plasma from patients with liver cancer [18, 19]. Aberrant glycoproteins in the blood stream may reflect abnormal states in patients with cancer and could be developed as serological cancer biomarkers [20–24].
Mass spectrometry (MS), a powerful technology in proteomic fields, can play a central role in identifying aberrant glycoproteins from various disease-related biological media. Recently, the use of high-resolution tandem MS combined with nano-LC led to identification of the site-specific N-glycopeptides of major proteins, such as haptoglobin (HP) and vitronectin (VTN), in human HCC plasma by the targeted protein enrichment method [25–28] as well as HILIC enrichment methods [29, 30]. The antibody method for the Lewis Y structure and glycan level analysis of purified glycoforms confirmed that the fucosylated glycan level of HP is increased in HCC plasma [26, 31, 32]. Tang and colleagues reported an automatic analysis and label-free quantification method, where several N-glycopeptides from four proteins (HP, VTN, complement C3 protein, and hemopexin) were found to be increased in HCC sera [33]. However, they had previously identified 103 N-glycopeptides from 33 serum proteins [34] and reported that only four fucosylated N-glycopeptides were increased in HCC sera [33]. This discrepancy led us to more closely examine the different types of fucosylated proteins from human HCC plasma.
In this study, we selected 13 liver-secreted proteins to identify fucosylated glycoproteins in HCC plasma. We identified N-glycopeptides and their fucosylated forms, which were analyzed in previous research [28, 35–38]. We report herein the analysis workflow for liver-secreted protein fucosylation in human HCC plasma using LC-MS/MS.
Materials and methods
Materials
K2EDTA-treated human plasma from patients with HCC was prepared at Severance Medical Center, Yonsei University (Seoul, Korea), with the consent of the blood donors and in accordance with Institutional Review Board guidelines (Electronic Supplementary Material (ESM) Table S1); it was stored at −80 °C until use. A multiple affinity removal system (MARS) column was purchased from Agilent (Wilmington, DE, USA). For glycopeptide enrichment, a ZIC-HILIC kit (ProteoExtract Glycopeptide Enrichment Kit) was obtained from EMD Millipore (Billerica, MA). Trypsin (mass spectrometry grade) for protein digestion was obtained from Promega (Madison, WI). Additionally, 1,4-dithiothreitol (DTT), iodoacetamide (IAA), trifluoroacetic acid (TFA), and formic acid (FA) were purchased from Sigma-Aldrich (St. Louis, MO). HPLC-grade acetonitrile (ACN) was also purchased (J.T. Baker, Phillipsburg, NJ).
Methods
Digestion of depleted and non-depleted plasma proteins
A MARS (Agilent) with an HP1100LC system (Agilent) was used to deplete the six most abundant proteins (albumin, transferrin, Ig gamma chain C region (IgG), Ig alpha chain C region (IgA), HP, and alpha-1-antitrypsin (AAT)) in plasma collected from 10 patients with HCC. Based on the manufacturer’s instructions, the human plasma samples were diluted by a factor of five in buffer A, which contained protease inhibitors, before centrifuging through 0.22-μm filters. Diluted plasma samples were injected at a flow rate of 0.25 ml/min (0 % B buffer) for 9 min. The bound fractions were eluted with buffer B (100 %) at a flow rate of 1 ml/min for 3.5 min. The column was regenerated by equilibrating in buffer B (0 %) for 7.5 min. Flow-through fractions collected from MARS-depleted plasma were stored at −20 °C until use. A pooled plasma sample was prepared by mixing 10 MARS-depleted plasmas from the HCC samples. The pooled plasma sample was desalted and concentrated by centrifugal filtration using 10-kDa molecular weight cutoff (MWCO) VIVASPIN filters (Sartorius, Göttingen, Germany). The plasma protein concentrations were determined by Bradford protein assay after filtering for molecular weight. The pooled sample (100 μg) was reduced with 5 mM DTT, alkylated with 12.5 mM IAA, and digested with trypsin (the substrate/enzyme ratio of 10:1) overnight at 37 °C. The digested sample was dried in a SpeedVac for HILIC enrichment.
N-Glycopeptide enrichment by HILIC
N-Glycopeptide enrichment with a ZIC-HILIC kit (ProteoExtract Glycopeptide Enrichment Kit) was performed according to the manufacturer’s instructions (EMD Millipore). This kit includes ZIC Glycocapture Resin, ZIC Binding Buffer, ZIC Wash Buffer, and ZIC Elution Buffer. Briefly, 10 μl plasma digests (3 μg/μl concentration) was prepared by adding 50 μl ZIC Binding Buffer mixed with 50 μl homogenous ZIC Glycocapture Resin by pipetting up and down and incubated for 20 min. The incubated resin was then transferred to a new 1.5-ml tube and centrifuged (2 min, 250×g). The supernatant was discarded. The centrifuged resin was washed three times with 150 μl ZIC Wash Buffer. After adding 75 μl ZIC Elution Buffer for N-glycopeptide elution, the tube was incubated for 5 min with agitation and centrifuged for 2 min at 2500×g. The supernatant, containing the N-glycopeptides, was transferred to a new 1.5-ml tube and centrifuged for 2 min at 10,000×g. The N-glycopeptide supernatant was transferred to a new 1.5-ml tube, avoiding the transfer of resin particles. For LC/MSMS analysis, the HILIC-enriched peptide solution from plasma was dried in a SpeedVac and reconstituted in 100 μl 0.1 % FA/99.95 % ddH2O.
Analysis using nano-liquid chromatography/ESI-MS/MS
The HILIC-enriched tryptic digest was separated with a nanoACQUITY UPLC system (Waters, Milford, MA) and analyzed using an LTQ Orbitrap Elite mass spectrometer (Thermo Scientific, Waltham, MA) equipped with a nano-electrospray source. Each 5 μl aliquot of the peptide solution was loaded into a nanoACQUITY UPLC Symmetry C18 column of i.d.180 μm, length 20 mm, and particle size 5 μm (Waters) using an autosampler. The homemade microcapillary separation column was packed with C18 material (Michrom Bioresources) in a fused-silica capillary [39]. The peptides trapped on the trap column after 10 min at a flow rate of 4 μl/min for desalting were back-flushed on a homemade microcapillary column (i.d. 100 μm and length 200 mm, C18, 3 μm particle size, 100 Å pore size) for separation. Mobile phase A consisted of 100 % water containing 0.1 % FA. Mobile phase B was 100 % ACN containing 0.1 % FA. The LC gradient was 5 % B maintained from 0 to 15 min; then, the mobile phase B was increased to 15 % for 5 min, to 50 % B for 115 min, and to 95 % B for 21 min (95 % B was continued for 13 min). Phase B was then decreased to 5 % for 1 min. The column was re-equilibrated with 5 % B for 10 min before the next run. The electrospray voltage was set to 2.2 kV. The LTQ Orbitrap Elite mass spectrometer (Thermo Scientific) was operated in data-dependent mode during the chromatographic separation. The MS acquisition parameters for full-scan resolution was 60,000 in the Orbitrap for each sample; five data-dependent MS/MS scans were acquired by collision-induced dissociation (CID) and higher-energy collision dissociation (HCD) in each full scan. CID and HCD scans were acquired in linear trap quadrupole (LTQ) mode with a 30-ms activation time and in Orbitrap at resolution 15,000 with a 20-ms activation time, respectively. A 35 % normalized collision energy (NCE) was used for CID and HCD analyses. Previously fragmented ions were excluded for 300 s in all MS/MS scans.
N-Glycopeptide identification
We manually performed systematic identification using CID and HCD spectra, as shown in Fig. 1 [28]. Briefly, manual N-glycopeptide identification included the following steps. First, a target N-glycopeptide database was constructed from a combination of the tryptic peptides including 50 N-glycosylation sites without miscleavage referenced in the Uniprot database (http://www.uniprot.org) and the 331 N-linked glycans that are biologically possible [40]. Next, tandem MS spectra of the N-glycopeptides were selected from HCD spectra based on the presence of at least four unique oxonium ions from glycan fragmentation; these consisted of GlcNAc (m/z 138.1, 168.1, and 204.1), NeuAc (m/z 274.1, 292.1), and GlcNAc-Hex (m/z 366.1) [41, 42]. Tandem MS spectra of targeted N-glycopeptides were selected from HCD and CID spectra based on the presence of y- and b-ions from the fragmentation of the peptide backbone and the Y-series ions from N-glycopeptides. We defined the minimum requirement criteria using two or more fragments of y- and b-ions from the peptide backbone, as well as Y0, Y1, Y2, and Y3-ions from the core N-glycopeptide glycan structure (Y0 = peptide_0_0_0_0 (Hex_HexNAc_Fuc_Sia), Y1 = peptide_0_1_0_0, Y2 = peptide_0_2_0_0 and _0_1_1_0, Y3 = peptide_0_2_1_0, _1_2_0_0 and _0_2_0_0). The y- and b-ions have been used with a relative intensity of more than 2 % (average S/N >3) and a mass accuracy with a tolerance less than 0.05 Da in HCD spectra. By comparing the theoretical monoisotopic ions in DB with the experimental monoisotopic ions in MS1, the target candidates were obtained from the HCD spectra of the selected N-glycopeptides. The N-glycopeptide candidates were then identified from the CID spectra by confirming the Y-series ions (Fig. 1b).
Results and discussion
Selection of target proteins and N-glycopeptides database construction
We previously examined the major proteins in HCC plasma, including AAT, HP, fetuin A (FETUA), alpha-1-antichymotrypsin (AACT), ceruloplasmin (CP), VTN, and alpha-1-acid glycoprotein 1/2 (AGP 1/2), and found indirect evidence of aberrant glycosylation using a specific lectin-coupled MRM assay [35–37]. Recent analyses of a single targeted protein showed that the N-glycopeptides of HP, kininogen-1 (KNG1), VTN, and immunoglobulin gamma isoforms (IgG1, 2, 3, and 4) were identified in HCC plasma [26–28, 43]. Additionally, large-scale analyses using tandem mass spectrometry have also been performed; however, few fucosylated N-glycopeptides were found in human sera [34, 44].
In this study, we selected 13 target proteins based on their abundance in human plasma [45] (Table 1). The protein concentration range of these 13 target proteins was 1–45 μmol/l [45].
In our recent study of vitronectin, a plasma protein, we provided a scheme for site-specific N-glycopeptide analysis of a single target protein [28]. To analyze various fucosylated glycoproteins in human plasma in this study, we modified the strategy for mapping the site-specific N-glycosylation of target proteins to include database construction, HILIC enrichment, LC-MS/MS analysis, and identification of N-glycopeptides (Fig. 1). For site-specific identification of various fucosylated N-glycoeptides from 13 highly abundant plasma proteins, we constructed, based on prior literature, a theoretical database of the mono-isotope ions of tryptic 16,550 N-glycopeptides from a combination of 50 tryptic peptides from 13 target proteins and 331 biologically possible N-linked glycans [40] (Fig. 1a).
Sample preparation and LC-MS/MS analysis
In this study, a pooled HCC plasma sample was prepared, tryptic-digested, and enriched using the HILIC method. The HILIC-enriched samples were analyzed in triplicate using a data-dependent acquisition mode in HCD/CID tandem mass spectrometry. The enriched samples obtained from depleted and non-depleted plasma were separated by reverse-phase nano-LC-MS/MS (Fig. 2a and ESM Fig. S1). The tryptic N-glycopeptides of target proteins were eluted in the range of 60–120 min. The glycopeptides identified from LC-MS/MS runs were further evaluated by their retention times. The relative standard deviation of the retention time was less than 1.2 %. The extracted ion chromatogram (XIC) showed different retention times for different glycopeptides from depleted plasma samples (Fig. 2b). The extracted ion chromatograms of six N-glycopeptides from AALAAFNAQNN176GSNFQLEEISR in FETUA (Fig. 2c) showed retention times that correlated with the type of glycan. Two sialylated N-glycopeptides, AALAAFNAQNN176GSNFQLEEISR_5_4_0_1 (5Hex_4HexNAc_0Fuc_1Sia) and AALAAFNAQNN176GSNFQLEEISR_5_4_0_2 (5Hex_4HexNAc_0Fuc_2Sia), had different retention times, at 81.29 and 83.87 min, respectively, which were dependent on the number of sialic acids. The more sialylated the N-glycopeptide, the longer it was retained in RP-LC [46]. However, the fucosylated N-glycopeptides, including AALAAFNAQNN176GSNFQLEEISR_5_4_1_2 (5Hex_4HexNAc_1Fuc_2Sia) and AALAAFNAQNN176GSNFQLEEISR_6_5_1_3 (6Hex_5HexNAc_1Fuc_3Sia), showed the same retention times (83.69 and 85.36 min, respectively) as their corresponding non-fucosylated N-glycopeptides, AALAAFNAQNN176GSNFQLEEISR_5_4_0_2 (5Hex_4HexNAc_0Fuc_2Sia) and AALAAFNAQNN176GSNFQLEEISR_6_5_0_3 (6Hex_5HexNAc_0Fuc_3Sia). This is because these isoforms have an equal number of sialic acids. When the peptide backbones in N-glycopeptides were identical, the N-glycopeptides, with one exception, were eluted at similar retention times in reverse-phase liquid chromatography. When the glycopeptides had sialic acids, we considered the retention time to be a direct reflection of the number of sialic acids.
LC/MS chromatograms of tryptic peptides from a depleted plasma sample. (a) Base peak chromatogram (BPC) of total peptides from plasma in triplicate runs. (b) Extracted ion chromatogram (XIC) of identified N-glycopeptides from target proteins. (c) XIC of two fucosylated and four non-fucosylated N-glycopeptides with N176 glycosylation site from fetuin a. As an example, pep_5_4_0_2 represents peptide_5Hex_4HexNAc_0Fuc_2Sia. pep = AALA…R (AALAAFNAQNN176GSNFQLEEISR); green circle, mannose; yellow circle, galactose; blue square, N-acetylglucosamine; red triangle, fucose; purple diamond, sialic acid
Identification of target N-glycopeptides in plasma
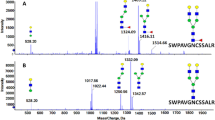
Using the systematic workflow to interpret HCD and CID spectra [28] (Fig. 1b), 189 unique N-glycopeptides were identified from human plasma samples (ESM Table S2). We manually assigned all the y-, b-, Y-, B-, and oxonium ions in the HCD and CID spectra of the target N-glycopeptides for positive identification. The Y-ion series are fragments from glycopeptides containing a glycan structure on a peptide backbone. The total MW of the Y-ion series from N-glycopeptides consists of glycans and peptides. Because we know the theoretical MWs of the 331 N-linked glycans that are biologically possible, we can match the theoretical MWs of peptides using the Uniprot database, which can then be compared with the experimental MWs of the peptides (= (Total MW of Y-ion series) − (MW of glycans)) using a fingerprinting method. Therefore, we can use the Y-ion series for the identification of N-glycopeptides in the HCD and CID spectra using the limited database. The following example illustrates the step-by-step assignments of tandem MS spectra from fucosylated N-glycopeptides. FETUA has two N-glycosylation sites, at N176 and N156, with tryptic glycopeptide sequences of AALAAFNAQNN176GSNFQLEEISR and VCQDCPLLAPLN156DTR, respectively. A recent study reported two N-glycopeptides after a site-specific analysis using acetone enrichment and LC/MS [44]. They identified only one FETUA N-glycopeptide, which consisted of only a bi-antennary type, in sera [44]. However, we found nine bi- and tri-antennary-type N-glycopeptides (ESM Table S2). Figure 3 shows the typical tandem MS spectra of FETUA fucosylated N-glycopeptides containing AALAAFNAQNN176GSNFQLEEISR_5_4_0_2 (5Hex_4HexNAc_0Fuc_2Sia) and AALAAFNAQNN176GSNFQLEEISR_5_4_1_2 (5Hex_4HexNAc_1Fuc_2Sia) in plasma. The candidate N-glycopeptides were first selected using oxonium ions from the glycan fragmentation in the HCD spectra. The peptide backbone of the N-glycopeptides was confirmed by the existence of y- and b-series ions from the peptide fragmentation in the HCD spectra. The N-glycopeptides with the same peptide backbone usually showed similar HCD spectra because they share the b- and y-series ions in HCD mode (Fig. 3a).
LC/MS/MS spectra of non-fucosylated and fucosylated N-glycopeptides, AALAAFNAQNNGSN176FQLEEISR_5_4_0_2 (5Hex_4HexNAc_0Fuc_2Sia) and AALAAFNAQNNGSN176FQLEEISR_ 5_4_1_2, containing N176 site of fetuin A from Fig. 2C. a HCD spectra of non-fucosylated (top) and fucosylated (bottom) N-glycopeptides. b CID spectra of non-fucosylated (top) and fucosylated (bottom) N-glycopeptides. The black arrow indicates oxonium ions, and the purple arrow indicates y- and b-ions fragmented from the peptide. The blue arrow indicates Y-series fragment ions from N-glycopeptides, the red arrow indicates Y-series fragment ions containing fucose, and the black square box indicates a pair of fragment ions containing fucose and non-fucose. As an example, pep_5_4_0_2 represents peptide_5Hex_4HexNAc_0Fuc_2Sia. pep = AALA…R (AALAAFNAQNN176GSNFQLEEISR); green circle, mannose; yellow circle, galactose; blue square, N-acetylglucosamine; red triangle, fucose; purple diamond, sialic acid
To find fucosylated N-glycopeptides, we used B-series ions at m/z 512.36 (1Hex_1HexNAc_1Fuc_0Sia) and m/z 803.53 (1Hex_1HexNAc_1Fuc_1Sia) and/or Y-series ion pairs with/without fucose in CID mode (Fig. 1b). The CID spectrum of fucosylated N-glycopeptides, such as AALAAFNAQNN176GSNFQLEEISR_5_4_1_2 (5Hex_4HexNAc_1Fuc_2Sia), showed B-series ions at m/z 512.36 and 803.53 and nine different pairs of Y-series ions with/without fucose (Fig. 3b).
By systematically interpreting the CID and HCD tandem mass spectra, we profiled target proteins of N-glycopeptides in human plasma. In total, 189 N-glycopeptides from 32 glycosylated sites on 13 major proteins were identified in plasma samples (ESM Table S2). The molecular ions of identified N-glycopeptides were confirmed with less than 10 ppm mass error.
The identified N-glycopeptides were classified as bi-antennary- (42.8 %), tri-antennary- (39.7 %), tetra-antennary- (9.5 %), mono-antennary- (6.3 %), hybrid- (2.1 %), or high-mannose- (0.5 %) type glycoforms in HCC plasma (Fig. 4a). Non-fucosylated and fucosylated N-glycopeptides were then split into two groups (Fig. 4b and c, respectively). The glycoform pattern of the non-fucosylated N-glycopeptides was similar to those of total N-glycopeptides (Fig. 4b). However, the fucosylated N-glycopeptides showed different tri-, bi-, and tetra-antennary glycoform percentages (48, 42, and 10 %, respectively) (Fig. 4c). Recently, Wuhrer et al. reviewed glycomic studies of major human plasma glycoproteins, including alpha-1-acid glycoprotein, alpha-1-antitrypsin, fetuin A, alpha-2-macroglobulin, ceruloplasmin, haptoglobin, kininogen-1, vitronectin, hemopexin, serotransferrin, and beta-2-glycoprotein 1 [47]. In their report, bi-antennary and tri-antennary complex-type glycoforms with/without fucose were the dominant glycoforms of the target plasma proteins, which is similar to the results of this study.
In contrast, mono-antennary glycoforms with only three N-acetylhexoseamine residues were more common in our study than in that performed by Clerc et al. (ESM Table S3). More recently, Reusch et al. reported that in-source decay may occur in all MS-based methods [48]. In their report, the mono-antennary glycoforms were relatively quantified at less than 10 %, based on various MS-based methods, and the loss of an antenna from bi-antennary glycoforms generally occurred with a charge reduction in ESI-source-based MS analysis [48]. Therefore, it is possible that the mono-antennary glycoforms found in the present study were partly formed by in-source decay.
The glycoprotein fucosylation level increases with the progression of liver cancer [49], and this idea has received significant attention in biomarker studies. In this study, we identified 71 fucosylated N-glycopeptides from 13 major human plasma proteins (Table 2, ESM Fig. S2). Additionally, we identified 37 new fucosylated N-glycopeptides from AACT, AAT, FETUA, CP, AGP 1/2, alpha-2-macroglobulin (A2M), serotransferrin (TF), and beta-2-glycoprotein 1 (APOH) in plasma. We focused primarily on the major abundant proteins in human plasma using our systematic workflow, which rigorously searches fucose-related B-series ions (m/z 512.36 and m/z 803.53) as well as Y-series ion pairs with/without fucose from among possible candidate glycopeptide spectra; this approach detected more fucosylated N-glycopeptides from target proteins than any other method to date [26, 33, 34, 43, 44].
Of the 13 liver-secreted glycoproteins in HCC plasma, all N-glycopeptides from 32 N-glycosylation sites were classified as site-specific glycoforms of fucosylated and non-fucosylated glycopeptides (Fig. 5). All the N-glycosylation sites were fully determined from the N-glycopeptides of FETUA, KNG, and VTN, whereas those of AACT, AAT, HP, CP, AGP 1/2, A2M, hemopexin (HPX), TF, and APOH were partially determined.
Most HCC plasma N-glycopeptides were identified as bi- and tri-antennary types, whereas tri- and tetra-antennary types were mainly identified for AACT and AGP. Previous studies have shown that tetra-antennary glycoforms are common for AGP [47, 50]. All the N-glycopeptides identified from HPX, TF, and APOH were bi- and tri-antennary glycoforms. A previous study showed that bi- and tri-antennary glycoforms of the N162 site in APOH were mainly identified in the serum from patients with antiphospholipid syndrome [51]. The fucosylated N-glycopeptides from KNG, FETUA, CP, TF, and APOH were classified as bi- and tri-antennary types. The N241 site of HP was determined to form bi- and tri-antennary-type glycoforms. In a previous study, Goldman et al. identified similar bi- and tri-antennary-type glycoform patterns for HP, KNG, and HPX after neuraminidase treatment [26]. The N432 and N630 sites of TF formed bi- and tri-antennary-type glycoforms; however, no alteration of the N491 site was detected, and this site had been previously shown to have low levels of glycosylation [47, 52]. Hybrid-type glycoforms were identified at N169 of VTN and N869 of A3M. High-mannose-type glycoforms were detected at N869 of A2M [53], and hybrid-type glycoforms were detected at N169 of VTN [28, 47]. These results demonstrated that site-specific N-glycopeptide analysis is important in the study of glycoproteins.
Goldman et al. concluded that the relative distribution of fucosylated N-glycopeptides is protein- and site-specific [26]. The site-specific micro-heterogeneity of fucosylation depends on the activity and stability of fucosyltransferases, such as Fut 8 and/or Fut 6 [5–7], the sequence of glycosyltransferase action, and the protein structure [26]. The limitations of a glycomics approach are apparent in the research with complex samples. Therefore, site-specific N-glycopeptide analysis is necessary for the study of fucosylated glycoproteins in complex samples, such as human HCC plasma.
Conclusions
In this study, we identified site-specific fucosylation of N-glycopeptides from HCC plasma samples. In total, 189 N-glycopeptides from 13 liver-secreted proteins, including 71 fucosylated N-glycopeptides, were identified. The fucosylated N-glycopeptides consisted of tri- (48 %), bi- (42 %), and tetra- (10 %) antennary glycoforms. In this study, 37 fucosylated N-glycopeptides, mostly consisting of bi- and tri-antennary glycoforms, were newly identified in HCC plasma samples. Of particular interest were the tetra- and tri-antennary glycoforms from AACT and AGP. We obtained a list of various fucosylated N-glycopeptides from liver-secreted proteins in human HCC plasma. These fucosylated N-glycopeptides could be used for the quantitation of site-specific fucosylation in HCC plasma proteins. As a next step, using multiple reaction monitoring, we will attempt to analyze differential fucosylation in liver-secreted proteins in various plasma samples from patients with various types of cancer.
Abbreviations
- A2M:
-
Alpha-2-macroglobulin
- AACT:
-
Alpha-1-antichymotrypsin
- AAT:
-
Alpha-1-antitrypsin
- AGP1:
-
Alpha-1-acid glycoprotein 1
- AGP2:
-
Alpha-1-acid glycoprotein 2
- APOH:
-
Beta-2-glycoprotein 1
- CID:
-
Collision-induced dissociation
- CP:
-
Ceruloplasmin
- FETUA:
-
Fetuin A
- Fuc:
-
Fucose
- HCC:
-
Hepatocellular carcinoma
- HCD:
-
Higher-energy collision dissociation
- Hex:
-
Hexose
- HexNAc:
-
N-Acetyl hexosamine
- HILIC:
-
Hydrophilic interaction liquid chromatography
- HP:
-
Haptoglobin
- HPX:
-
Hemopexin
- KNG1:
-
Kininogen-1
- MS:
-
Mass spectrometry
- N:
-
Asparagine
- Sia:
-
Sialic acid
- TF:
-
Serotransferrin
- VTN:
-
Vitronectin
References
Walsh G, Jefferis R. Post-translational modifications in the context of therapeutic proteins. Nat Biotechnol. 2006;24(10):1241–52.
Kornfeld R, Kornfeld S. Assembly of asparagine-linked oligosaccharides. Annu Rev Biochem. 1985;54(1):631–64.
An HJ, Peavy TR, Hedrick JL, Lebrilla CB. Determination of N-glycosylation sites and site heterogeneity in glycoproteins. Anal Chem. 2003;75(20):5628–37.
Fuster MM, Esko JD. The sweet and sour of cancer: glycans as novel therapeutic targets. Nat Rev Cancer. 2005;5(7):526–42.
Cheng L, Luo S, Jin C, Ma H, Zhou H, Jia L. FUT family mediates the multidrug resistance of human hepatocellular carcinoma via the PI3K/Akt signaling pathway. Cell Death Dis. 2013;4(11):e923. doi:10.1038/cddis.2013.450.
Kang X, Wang N, Pei C, Sun LU, Sun R, Chen JIE, et al. Glycan-related gene expression signatures in human metastatic hepatocellular carcinoma cells. Exp Ther Med. 2012;3(3):415–22.
Nakagawa T, Uozumi N, Nakano M, Mizuno-Horikawa Y, Okuyama N, Taguchi T, et al. Fucosylation of N-glycans regulates the secretion of hepatic glycoproteins into bile ducts. J Biol Chem. 2006;281(40):29797–806.
Anderson NL, Anderson NG. The human plasma proteome: history, character, and diagnostic prospects. Mol Cell Proteomics. 2002;2(1):847–67.
Farrah T, Deutsch EW, Omenn GS, Campbell DS, Sun Z, Bletz JA, et al. High-confidence human plasma proteome reference set with estimated concentrations in PeptideAtlas. Mol Cell Proteomics. 2011;10(9), M110.006353.
Boron WF, Boulpaep EL. Medical physiology: a cellular and molecular approach. Philadelphia: Elsevier/Saunders; 2003. p. 1300.
Uhlén M, Fagerberg L, Hallström BM, Lindskog C, Oksvold P, Mardinoglu A, et al. Proteomics. Tissue-based map of the human proteome. Science. 2015;6220(6220):1260419.
Ludwig JA, Weinstein JN. Biomarkers in cancer staging, prognosis and treatment selection. Nat Rev Cancer. 2005;5(11):845–56.
Miyaaki H, Nakashima O, Kurogi M, Eguchi K, Kojiro M. Lens culinaris agglutinin-reactive alpha-fetoprotein and protein induced by vitamin K absence II are potential indicators of a poor prognosis: a histopathological study of surgically resected hepatocellular carcinoma. J Gastroenterol. 2007;42(12):962–8.
Khien VV, Mao HV, Chinh TT, Ha PT, Bang MH, Lac BV, et al. Clinical evaluation of lentil lectin-reactive alpha-fetoprotein-L3 in histology-proven hepatocellular carcinoma. Int J Biol Markers. 2001;16(2):105–11.
Breborowicz J, Mackiewicz A, Breborowicz D. Microheterogeneity of alpha-fetoprotein in patient serum as demonstrated by lectin affino-electrophoresis. Scand J Immunol. 1981;14(1):15–20.
Okuda H, Saito A, Shiratori K, Yamamoto M, Takasaki K, Nakano M. Clinicopathologic features of patients with primary malignant hepatic tumors seropositive for alpha-fetoprotein-L3 alone in comparison with other patients seropositive for alpha-fetoprotein-L3. J Gastroenterol Hepatol. 2005;20(5):759–64.
Naitoh A, Aoyagi Y, Asakura H. Highly enhanced fucosylation of serum glycoproteins in patients with hepatocellular carcinoma. J Gastroenterol Hepatol. 1999;14(5):436–45.
Li D, Mallory T, Satomura S. AFP-L3: a new generation of tumor marker for hepatocellular carcinoma. Clin Chim Acta. 2001;313(1-2):15–9.
Taylor AD, Hancock WS, Hincapie M, Taniguchi N, Hanash SM. Towards an integrated proteomic and glycomic approach to finding cancer biomarkers. Genome Med. 2009;1(6):57.
Zhao J, Qiu W, Simeone DM, Lubman DM. N-linked glycosylation profiling of pancreatic cancer serum using capillary liquid phase separation coupled with mass spectrometric analysis. J Proteome Res. 2007;6(3):1126–38.
Pedersen JW, Blixt O, Bennett EP, Tarp MA, Dar I, Mandel U, et al. Seromic profiling of colorectal cancer patients with novel glycopeptide microarray. Int J Cancer. 2011;128(8):1860–71.
Barrabés S, Pagès-Pons L, Radcliffe CM, Tabarés G, Fort E, Royle L, et al. Glycosylation of serum ribonuclease 1 indicates a major endothelial origin and reveals an increase in core fucosylation in pancreatic cancer. Glycobiology. 2007;17(4):388–400.
Comunale MA, Lowman M, Long RE, Krakover J, Philip R, Seeholzer S, et al. Proteomic analysis of serum associated fucosylated glycoproteins in the development of primary hepatocellular carcinoma. J Proteome Res. 2006;5(2):308–15.
Schiel JE, Au J, He H-J, Phinney KW. LC-MS/MS biopharmaceutical glycoanalysis: identification of desirable reference material characteristics. Anal Bioanal Chem. 2012;403(8):2279–89.
Pompach P, Brnakova Z, Sanda M, Wu J, Edwards N, Goldman R. Site-specific glycoforms of haptoglobin in liver cirrhosis and hepatocellular carcinoma. Mol Cell Proteomics. 2013;12(5):1281–93.
Pompach P, Ashline DJ, Brnakova Z, Benicky J, Sanda M, Goldman R. Protein and site specificity of fucosylation in liver-secreted glycoproteins. J Proteome Res. 2014;13(12):5561–9.
Lee H-J, Cha H-J, Lim J-S, Lee SH, Song SY, Kim H, et al. Abundance-ratio-based semiquantitative analysis of site-specific N-linked glycopeptides present in the plasma of hepatocellular carcinoma patients. J Proteome Res. 2014;13(5):2328–38.
Hwang H, Lee JY, Lee HK, Park GW, Jeong HK, Moon MH, et al. In-depth analysis of site-specific N-glycosylation in vitronectin from human plasma by tandem mass spectrometry with immunoprecipitation. Anal Bioanal Chem. 2014;406(30):7999–8011.
Kuo C-W, Wu I-L, Hsiao H-H, Khoo K-H. Rapid glycopeptide enrichment and N-glycosylation site mapping strategies based on amine-functionalized magnetic nanoparticles. Anal Bioanal Chem. 2012;402(9):2765–76.
Zhao Y, Yu L, Guo Z, Li X, Liang X. Reversed-phase depletion coupled with hydrophilic affinity enrichment for the selective isolation of N-linked glycopeptides by using Click OEG-CD matrix. Anal Bioanal Chem. 2011;399(10):3359–65.
Zhu J, Lin Z, Wu J, Yin H, Dai J, Feng Z, et al. Analysis of serum haptoglobin fucosylation in hepatocellular carcinoma and liver cirrhosis of different etiologies. J Proteome Res. 2014;13(6):2986–97.
Asazawa H, Kamada Y, Takeda Y, Takamatsu S, Shinzaki S, Kim Y, et al. Serum fucosylated haptoglobin in chronic liver diseases as a potential biomarker of hepatocellular carcinoma development. Clin Chem Lab Med. 2015;53(1):95–102.
Mayampurath A, Song E, Mathur A, Yu C-y, Hammoud Z, Mechref Y, et al. Label-free glycopeptide quantification for biomarker discovery in human sera. J Proteome Res. 2014;13(11):4821–32.
Mayampurath A, Yu C-Y, Song E, Balan J, Mechref Y, Tang H. Computational framework for identification of intact glycopeptides in complex samples. Anal Chem. 2014;86(4):453–63.
Ahn YH, Shin PM, Oh NR, Park GW, Kim H, Yoo JS. A lectin-coupled, targeted proteomic mass spectrometry (MRM MS) platform for identification of multiple liver cancer biomarkers in human plasma. J Proteomics. 2012;75(17):5507–15.
Ahn YH, Shin PM, Kim Y-S, Oh NR, Ji ES, Kim KH, et al. Quantitative analysis of aberrant protein glycosylation in liver cancer plasma by AAL-enrichment and MRM mass spectrometry. Analyst. 2013;138(21):6454–62.
Ahn YH, Shin PM, Ji ES, Kim H, Yoo JS. A lectin-coupled, multiple reaction monitoring based quantitative analysis of human plasma glycoproteins by mass spectrometry. Anal Bioanal Chem. 2012;402(6):2101–12.
Lee JY, Kim JY, Park GW, Cheon MH, Kwon K-H, Ahn YH, et al. Targeted mass spectrometric approach for biomarker discovery and validation with nonglycosylated tryptic peptides from N-linked glycoproteins in human plasma. Mol Cell Proteomics. 2011;10(12):M111.009290.1–13.
Link AJ, Eng J, Schieltz DM, Carmack E, Mize GJ, Morris DR, et al. Direct analysis of protein complexes using mass spectrometry. Nat Biotechnol. 1999;17(7):676–82.
Kronewitter SR, An HJ, de Leoz ML, Lebrilla CB, Miyamoto S, Leiserowitz GS. The development of retorosynthetic glycan libraries to profile and classify the human serum N-linked glycome. Proteomics. 2009;9(11):2986–94.
Ozohanics O, Krenyacz J, Ludanyi K, Pollreisz F, Vekey K, Drahos L. GlycoMiner: a new software tool to elucidate glycopeptide composition. Rapid Commun Mass Spectrom. 2008;22(20):3245–54.
Mayampurath AM, Wu Y, Segu ZM, Mechref Y, Tang H. Improving confidence in detection and characterization of protein N-glycosylation sites and microheterogeneity. Rapid Commun Mass Spectrom. 2011;25(14):2007–19.
Yuan W, Sanda M, Wu J, Koomen J, Goldman R. Quantitative analysis of immunoglobulin subclasses and subclass specific glycosylation by LC-MS-MRM in liver disease. J Proteome. 2015;116:24–33.
Takakura D, Harazono A, Hashii N, Kawasaki N. Selective glycopeptide profiling by acetone enrichment and LC/MS. J Proteomics. 2014;101:17–30.
Hortin GL, Sviridov D, Anderson NL. High-abundance polypeptides of the human plasma proteome comprising the top 4 logs of polypeptide abundance. Clin Chem. 2008;54(10):1608–16.
Song E, Mayampurath A, Yu C-Y, Tang H, Mechref Y. Glycoproteomics: identifying the glycosylation of prostate specific antigen at normal and high isoelectric points by LC-MS/MS J. Proteome Res. 2014;13(12):5570–80.
Clerc F, Reiding KR, Jansen BC, Kammeijer GSM, Bondt A, Wuhrer M. Human plasma protein N-glycosylation. Glycoconj J. 2015. doi:10.1007/s10719-015-9626-2.
Reusch D, Haberger M, Falck D, Peter B, Maier B, Gassner J, et al. Comparison of methods for the analysis of therapeutic immunoglobulin G Fc-glycosylation profiles—Part 2: mass spectrometric methods. mAbs. 2015;7(4):732–42.
Gomaa AI, Khan SA, Leen ELS, Waked I, Taylor-Robinson SD. Diagnosis of hepatocellular carcinoma. World J Gastroenterol. 2009;15(11):1301–14.
Imre T, Kremmer T, Heberger K, Molnar-Szollosi E, Ludanyi K, Pocsfalvi G, et al. Mass spectrometric and linear discriminant analysis of N-glycans of human serum alpha-1-acid glycoprotein in cancer patients and healthy individuals. J Proteome. 2008;71(2):186–97.
Kondo A, Miyamoto T, Yonekawa O, Giessing AM, Osterlund EC, Jensen ON. Glycopeptide profiling of beta-2glycoprotein I by mass spectrometry reveals attenuated sialylation in patients with antiphospholipid syndrome. J Proteome. 2009;73(1):123–33.
Satomi Y, Shimonishi Y, Hase T, Takao T. Site-specific carbohydrate profiling of human transferrin by nano-flow liquid chromatography/electrospray ionization mass spectrometry. Rapid Commun Mass Spectrom. 2004;18(24):2983–8.
Arnold JN, Wallis R, Willis AC, Harvey DJ, Royle L, Dwek RA, et al. Interaction of mannan binding lectin with alpha 2 macroglobulin via exposed oligomannose glycans. J Biol Chem. 2006;281(11):6955–63.
Acknowledgments
This research was supported by the National Research Council of Science and Technology (NTM2371511, the Creative Allied Project (CAP)), the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI); it was funded by the Ministry of Health &Welfare, Republic of Korea (grant number: HI13C2098), and by the Ministry of Trade, Industry & Energy (MOTIE, 1415139249) through the Osong Academy–Industry Convergence (BAIO) fostering project.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Ethics approval and consent to participate
This study was performed in accordance with ethical standards. Plasma samples were obtained from voluntary blood donors with informed consent and approval in accordance with IRB guidelines from Yonsei University College of Medicine (Seoul, Korea).
Additional information
Eun Sun Ji and Heeyoun Hwang contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 5758 kb)
Rights and permissions
About this article
Cite this article
Ji, E.S., Hwang, H., Park, G.W. et al. Analysis of fucosylation in liver-secreted N-glycoproteins from human hepatocellular carcinoma plasma using liquid chromatography with tandem mass spectrometry. Anal Bioanal Chem 408, 7761–7774 (2016). https://doi.org/10.1007/s00216-016-9878-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-016-9878-0