Abstract
Glioblastoma (GBM) is an aggressive type IV brain tumor that originates from astrocytes and has a poor prognosis. Despite intensive research, survival rates have not significantly improved. Noncoding RNAs (ncRNAs) are emerging as critical regulators of carcinogenesis, progression, and increased treatment resistance in GBM cells. They influence angiogenesis, migration, epithelial-to-mesenchymal transition, and invasion in GBM cells. ncRNAs, such as long ncRNAs (lncRNAs), microRNAs (miRNAs), and circular RNAs (circRNAs), are commonly dysregulated in GBM. miRNAs, such as miR-21, miR‐133a, and miR‐27a‐3p, are oncogenes that increase cell proliferation, metastasis, and migration by targeting TGFBR1 and BTG2. In contrast, lncRNAs, such as HOXD-AS2 and LINC00511, are oncogenes that increase the migration, invasion, and proliferation of cells. CircRNAs, such as circ0001730, circENTPD7, and circFOXO3, are oncogenes responsible for cell growth, angiogenesis, and viability. Developing novel therapeutic strategies targeting ncRNAs, cell migration, and angiogenesis is a promising approach for GBM. By targeting these dysregulated ncRNAs, we can potentially restore a healthy balance in gene expression and influence disease progression. ncRNAs abound within GBM, demonstrating significant roles in governing the growth and behavior of these tumors. They may also be useful as biomarkers or targets for therapy. The use of morpholino oligonucleotides (MOs) suppressing the oncogene expression of HOTAIR, BCYRN1, and cyrano, antisense oligonucleotides (ASOs) suppressing the expression of ncRNAs such as MALAT1 and miR-10b, locked nucleic acids (LNAs) suppressing miR-21, and peptide nucleic acids (PNAs) suppressing the expression of miR-155 inhibited the PI3K pathway, tumor growth, angiogenesis, proliferation, migration, and invasion. Targeting oncogenic ncRNAs with RNA-interfering strategies such as MOs, ASOs, LNAs, CRISPR-Cas9 gene editing, and PNA approaches may represent a promising therapeutic strategy for GBM. This review emphasizes the critical role of ncRNAs in GBM pathogenesis, as well as the potential for new therapeutic strategies targeting these pathways to improve the prognosis and quality of life for GBM patients.
Graphical Abstract
Noncoding RNAs in glioblastoma play a dual role as both tumor suppressors and oncogenes, intricately regulating gene expression and contributing to the complex molecular landscape of this aggressive brain cancer.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
RNA was once thought of as just a messenger molecule for relaying the code from DNA by ribosomes to synthesize proteins. However, in the past three decades, different types of RNA molecules have been discovered, the most important of which are the noncoding RNAs (ncRNAs). ncRNA constitutes around 98% of RNA transcribed from human DNA, a revelation from whole-genome sequencing, while only approximately 3% comprises protein-coding genes (Song et al. 2024). The rest of the genome encodes mainly ncRNA. With the development of high throughput sequencing methodology, the majority of DNA sequences in the human genome have been elucidated (Bao et al. 2024). The ENCODE (Encyclopedia of DNA Elements) project revealed that less than 2% of the mammalian genome encodes messenger RNA. Still, at least 70% of the genome can produce transcripts of different sizes, mostly ncRNAs (Mattick et al. 2023). ncRNAs play important roles in regulating life activities such as DNA replication, transcription, RNA processing, translation, and protein functions. These ncRNAs are split into two classes: housekeeping and regulatory RNAs. tRNA, rRNA, snRNA, and snoRNA belong to these housekeeping RNA classes responsible for maintaining a constant protein expression level in the cells (Beňačka et al. 2023). The housekeeping ncRNAs are parts of the critical molecular machinery required for basic life activities, including transport RNA carrying amino acids, small nucleolar RNA guiding RNA modification and processing, and ribosomal RNA involved in protein synthesis (Koffler et al. 2023). The regulatory ncRNAs, such as microRNA (miRNA), long noncoding RNA (lncRNA), and circular RNA (circRNA), participate in multiple processes including tumorigenesis and tumor angiogenesis (Ahmadi et al. 2024).
The realm of lncRNAs, characterized by transcripts over 200 nucleotides long, has unveiled its involvement in numerous biological processes. These ncRNAs are theorized to influence gene expression through epigenetic, transcriptional, and post-transcriptional modifications, with their levels intricately linked to various biological traits such as cell survival (Wu et al. 2023). Their dysregulation of ncRNA is increasingly recognized as a factor in numerous human disorders, particularly noteworthy being their distinct expression profiles across diverse tumor types, where they can act as suppressors or promoters of tumor growth (Ismail et al. 2023). Dysregulated ncRNA expression plays a pivotal role in cancer initiation, progression, resistance to therapy, and the dysregulation of genes involved in carcinogenesis, metastasis, and malignancy progression through different tumor stages (Xue et al. 2022). Operating through various cellular processes including transcriptional, post-transcriptional, and epigenetic mechanisms, ncRNAs, affected by genetic changes, are closely linked to cancer development. Additionally, their involvement in the global regulation of cellular processes positions ncRNAs as potential biomarkers in cancer, thus introducing a fresh perspective to cancer diagnosis and treatment (Lu et al. 2023a, Peng et al. 2023). For instance, in GBM, the deletion of miR‐489‐3p, miR‐181a‐5p, and miR‐128‐3p tumor suppressors is associated with the deletion of BDNF and KPNA4 (Lui et al. 2024). Conversely, amplifying specific chromosomal regions has led to the overexpression of oncogenic lncRNAs like FAL1 and PVT1 (Kciuk et al. 2023). Furthermore, recurrent mutations occurring in the promoters of lncRNAs NEAT1 and MIR210HG have been implicated in altered expression patterns and the progression of glial cells (Tang et al. 2023).
In recent decades, ncRNAs have been targeted to reduce glioblastoma (GBM) through various mechanisms. Inhibiting metastasis, angiogenesis, cell proliferation, migration, and invasion are crucial for effective treatment strategies (Dai et al. 2023). Metastasis is a significant obstacle in GBM management, as it leads to extracranial spread and worsens patient outcomes. Angiogenesis, the formation of new blood vessels, supports tumor growth and metastasis by providing necessary nutrients and oxygen (Gu et al. 2023). Cell proliferation and apoptosis are critical for tumor progression and response to therapy. Migration and invasion enable GBM cells to disseminate and colonize distant sites (Wang et al. 2023). By targeting angiogenesis, migration, and cell survival, processes can improve treatment outcomes by reducing the risk of metastasis and enhancing the efficacy of therapies. Targeting these mechanisms can lead to more effective and personalized treatments for GBM patients. Targeting dysregulated ncRNAs in GBM holds promise for restoring normal gene expression patterns and potentially influencing disease progression (Rios et al. 2024). Various therapeutic strategies are under investigation to target ncRNAs effectively. RNA inhibiting strategies such as antisense oligonucleotides (ASOs) are short, single-stranded DNA or RNA molecules designed to bind complementary sequences on target ncRNAs. This binding can lead to degradation of the target ncRNA via RNase H-mediated mechanisms or block its interaction with other molecules, thereby altering its regulatory function. ncRNAs are abundant in GBM and play crucial roles in tumor growth and behavior regulation. They also hold potential as biomarkers or therapeutic targets (Garbo et al. 2022). Emerging methods such as morpholino oligonucleotides (MOs) (Hua et al. 2024), locked nucleic acids (LNAs) (Shivakumar et al. 2024), and peptide nucleic acids (PNAs) (Pradeep et al. 2023) offer precise approaches for targeting ncRNAs in GBM therapy. Another approach involves the use of miRNA mimics or antagonists. Mimics are synthetic versions of miRNAs that can compensate for deficient miRNAs in disease states, while antagonists can inhibit the activity of overexpressed miRNAs.
The World Health Organization (WHO) has defined glioma as a kind of brain cancer known as GBM. It is the most unrelenting primary brain cancer, proves refractory to treatment, and portends a dismal future (Lin et al. 2023). Among human cancers, GBM, or grade IV glioma, is a particularly deadly one. It represents the greatest incidence rate of malignant brain and central nervous system (CNS) malignancies, accounting for 27% of primary brain tumors and an astounding 80% of malignant cases. About 308,102 new cases and 251,329 new fatalities related to GBM are recorded each year (Akhlaghdoust et al. 2024). GBM is distinguished by its aggressive and rapid clinical progression, mutations, epigenetic modifications, prominent vascularization, and infiltrative growth pattern. Age, race, ethnicity, and gender are among the variables that affect the incidence of GBM. The time-of-life annual incidence rate of GBM is 3.20/100,000 individuals. Men are 1.6 times more likely to develop GBM than women (Goel et al. 2022; Tepper et al. 2016). The incidence of GBM is higher in the West than in less developed countries, especially in the case of white people, and it tends to rise with age (Grochans et al. 2022). The median overall survival for those with GBM is a dismal 15 months. Despite employing intensive therapeutic approaches such as surgery and chemoradiotherapy, the median survival duration for individuals with GBM has shown minimal improvement (Huang et al. 2024). Due to several limitations in chemotherapy, radiotherapy, and surgery, recent research has highlighted the involvement of miRNA, lncRNA, and circRNA in GBM; it is exemplified in Fig. 1 sourced from the Scopus database, which elucidates their roles and potential as targets for therapeutic intervention. Figure 2 details the research work conducted on miRNA, lncRNA, and circRNA during a specific period from 2016 to 2022. The recent adoption of these strategies in clinical practice highlights the crucial need for research into novel diagnostic and therapeutic approaches for GBM.
A bibliograph network map of scientific research on glioblastoma with miRNA therapy. It specifies a period of the occurrence of the keyword from 2016 (blue) to 2022 (yellow). This figure is utilized from the Scopus database by keywords “glioblastoma” AND “miRNA” lncRNA” AND “circRNA” and a time range of 2016–2022
However, there is still a lot of research needed to fully understand the regulation of cell processes to make use of new biomarkers and new technologies as tools to improve the treatment of GBM. This review focuses on the recent developments and advances in ncRNA biology pertinent to their role in GBM tumorigenesis and therapy response. Here, we delve into the functional roles of miRNA, lncRNA, and circRNAs, in GBM development, progression, and therapy resistance, thus highlighting the potential and clinical implications of the ncRNAs in GBM. In previous research on GBM, targeting the disease has often led to the reappearance of the tumor due to the development of resistance. To overcome this limitation, a novel approach involves targeting ncRNAs, which have been shown to play critical roles in GBM pathogenesis. Suppressing these ncRNA biomarkers using ASOs, MOs, LNAs, and PNAs can inhibit apoptosis, cell migration, and angiogenesis, thereby preventing the recurrence of metastasis. This targeted strategy can potentially improve treatment outcomes and enhance the quality of life for GBM patients. This study will offer valuable insights to various stakeholders, including researchers, clinicians, patients, neuro-oncologists, oncologists, molecular biologists, geneticists, and healthcare professionals involved in GBM diagnosis and treatment. In this context, this article aims to review the exciting progress of ncRNAs in the development of GBM and highlight the possibility of developing novel therapeutic approaches that target these metastasis and angiogenesis pathways to enhance the prognosis and quality of life for GBM patients.
Noncoding RNA
The classification of ncRNAs is based on various criteria including their structure, function, biogenesis, localization, and interaction with DNA or protein-coding mRNAs. miRNAs, small interfering RNAs (siRNAs), piwi-interacting RNAs (piRNAs), and other types are all small regulatory RNAs. They consist of up to 200 bases. Long noncoding RNAs, on the other hand, are those that are longer than 200 bases. Many small, noncoding RNAs do useful things. Two examples are transfer RNAs (tRNAs) and ribosomal RNAs (rRNAs) (Chaudhary and Banerjee 2024). In eukaryotic cells, ncRNAs make up about 0.002 to 0.2% of the total RNA content, while rRNA and tRNA make up 80 to 90% and 10 to 15% of the total RNA content, respectively. There are two more types of lncRNAs: antisense lncRNAs and long intergenic ncRNAs (lincRNAs). The regulatory regions of these lncRNAs distinguish them from protein-coding genes within the same genomic locus (DeOcesano et al. 2020). Natural antisense transcripts (NATs) are another group (Svoboda 2020). They consist of lncRNAs, whose sequences align with those of other RNA molecules. NATs frequently work as nonprotein-coding antisense RNA partners with protein-coding mRNAs. This suggests that they might help control nonprotein-coding sense RNAs (Širvinskas et al. 2023). To target angiogenesis through dysregulated ncRNAs, it is essential to understand the mechanisms of miRNAs, lncRNAs, and circRNAs individually.
MicroRNAs
Biogenesis of miRNAs
miRNAs are naturally produced and only 18 to 22 nucleotides long, control gene expression by binding to specific sections of messenger RNA molecules. There are multiple processes involved in the production of miRNAs (O’Brien et al. 2018a, 2018b). After RNA polymerase II/III generates primary miRNA in the nucleus, it undergoes a metamorphosis into precursor miRNA (pre-miRNA). Ran, Exportin5, and guanosine triphosphate (GTP) work together to make this complicated journey of pre-miRNAs from the cytoplasm back to the nucleus possible (O’Brien et al. 2018a, 2018b). Pre-miRNAs are cleaved into mature double-stranded miRNAs once they reach the nucleus. Following maturity, these miRNAs split into two independent strands. When one of these strands attaches itself to the intended mRNA, an RNA-induced silencing complex is created (RISC) (Pong and Gullerova 2018). This interaction inhibits the target gene’s expression. One strand, typically the guide strand, directs the RISC complex to the target mRNA sequence, while the other strand, known as the passenger strand, is usually degraded or otherwise discarded (Pong and Gullerova 2018).
In eukaryotes, miRNAs bind to mRNA transcripts and cause mRNA degradation or translational suppression. They can identify mRNA molecules for specific breakdowns, which can directly cause mRNA degradation. With the help of enzymes and protein complexes, the miRNA-induced silencing complex (miRISC) orchestrates the degradation of mRNA (Luo et al. 2018). Argonaute proteins, which are at the center of mRISC, direct miRISCs to target specific mRNAs. Through their interference with the translation process, miRNAs can either entirely stop or reduce the expression of a gene. In mammalian cells, miRNAs can amplify translation inhibition through the de-adenylation and destruction of mRNA (Mattei et al. 2021). Unconventional mechanisms by which miRNAs adversely affect target gene expression have been uncovered in multiple investigations (Zhao et al. 2022). One study by Matsui et al. showed that miR-589 interacts with RNA in the cyclooxygenase-2 (COX-2) promoter region, which increases the transcription of COX-2 (Matsui et al. 2013).
Intricacies of miRNA targeting and functional dynamics
There are two main ways that miRNAs are made: the canonical pathway and the noncanonical pathway. Through the canonical route, RNA polymerase II copies genes to make pri-miRNAs. Starting with a cap structure and a polyadenylation tail, the primary miRNA (pri-miRNA) goes on a trip of change that ends with a stem-loop structure formed by complementary base pairing (Pong and Gullerova 2018). The microprocessor, which is made up of the Drosha enzyme and the DGCR8 protein, keeps cutting up the pri-miRNA after it has been changed into the shape of a hairpin. Pre-miRNAs are hairpin-shaped and usually between 60 and 120 nucleotides long. They are made by this process (Manzoni et al. 2018). Taking the pre-miRNA across the nuclear border and into the cytoplasm is done by the Exportin5/RanGTP complex, which works like a molecular ferry. The pre-miRNA meets Dicer, an RNase III enzyme, in the cytoplasm of the cell. This starts a complicated dance of cleavage that makes mature miRNA pairs that are about 22 nucleotides long (Liu et al. 2018). A protein called Argonaute (AGO) will only bind to the 3p strand from its 3′ end or the 5p strand from its 5′ end after the pre-miRNA hairpin structure unwinds. The chosen strand is then added to the RISC (Medley et al. 2021).
The miRNA-guided RISC can either block translation or break down mRNA. There are two alternative pathways for miRNA synthesis that do not involve the Drosha/DGCR8 complex or Dicer. Mirtrons and m7G-capped pre-miRNAs are two types of pre-miRNAs that are made from introns during splicing and look like Dicer substrates (Martinez et al. 2017). These pre-miRNAs mature independently of the Drosha/DGCR8 complex and are transported to the cytoplasm by Exportin 1, bypassing Drosha cleavage. Small hairpin RNA (shRNA) transcript maturation occurs independently of Dicer (Saloura et al. 2017). Pre-miRNA transcripts embark on a two-stage odyssey, beginning with a meticulous cleavage by the microprocessor complex within the nucleus, followed by a journey to the cytoplasm orchestrated by the Exportin5/RanGTP complex (Ali Syeda et al. 2020). The intermediate miRNA undergoes further processing, where its 3′ end is trimmed after being incorporated into the catalytic component 2 of the argonaute RISC. This trimming takes place after the catalytic core of RISC cleaves the miRNA (Singh 2020) (Chen et al. 2017). This additional trimming step contributes to the generation of the mature miRNA form. Regulation of miRNA biogenesis is vital for cellular development, as illustrated in Fig. 3. However, if this miRNA pathway becomes dysregulated, it can potentially trigger tumorigenesis, providing opportunities for therapeutic targets and biomarkers.
The intricate journey of miRNA biogenesis and its mode of action begins with the transcription of pri-miRNA, an elongated RNA molecule folded into a hairpin structure. This pri-miRNA is then cleaved by a nuclear complex called the microprocessor, composed of Drosha and DGCR8 enzymes, to form a shorter hairpin structure known as pre-miRNA. The pre-miRNA embarks on a journey to the cytoplasm, where it is further processed by Dicer, another enzyme, to generate a mature miRNA duplex. This duplex consists of two complementary miRNA strands, one of which is selected and loaded into Argonaute proteins, forming a miRNA-induced silencing complex (miRISC). Noncanonical pathways deviate from this standard route. Some miRNAs, such as small hairpin RNAs (shRNAs), are cleaved and exported independently of the microprocessor complex and undergo AGO2-dependent processing. Others, like mirtrons and m7G-pre-miRNAs, rely on Dicer for their cytoplasmic maturation and exhibit unique export mechanisms. Regardless of the pathway, all roads lead to the formation of functional miRISC complexes. These complexes initiate translational inhibition, blocking the production of proteins from target mRNAs. In some cases, miRISC also triggers de-adenylation and degradation of the target mRNA, leading to its destruction. Created with biorender.com
A miRISC is formed through canonical or noncanonical pathways, involving an AGO protein family member and a guide strand. This intricate system accurately recognizes its mRNA targets by attaching to the corresponding miRNA response elements (MREs) situated on the targeted mRNA. On average, each human miRNA family may target about 300 conserved mRNAs. A single mammalian miRNA family has the potential to regulate approximately 300 conserved mRNAs on average. The “seed region” (nucleotides 2–8 from the 5′ end) is crucial for miRNA-MRE interaction. Target prediction algorithms often generate numerous anticipated matches due to reliance on conserved sequences with six, seven, or eight nucleotides matching the miRNA’s seed region (Chen et al. 2017; McGeary et al. 2019; Singh 2020). For target mRNA cleavage, a high degree of sequence complementarity is necessary to determine AGO2 endonuclease cleavage or miRISC prevention of mRNA translation and subsequent degradation. Cleavage is initiated with a fully complementary contact between the miRNA and MRE, inducing instability at the 3′ end of the guiding miRNA, leading to its eventual destruction. However, perfect complementarity is uncommon in humans due to central mismatches in most MREs, making AGO2 endonuclease function and destabilization of the miRNA’s 3′ terminus highly improbable (Gu et al. 2018).
The miRISC complex, which includes the GW182 protein family, orchestrates the degradation of mRNA. This complex recruits effector proteins that disassemble the mRNA molecule, making it susceptible to digestion by the enzyme 5′-3′ exoribonuclease-1 (XRN1). While miRNAs are traditionally linked to gene silencing, emerging evidence suggests that they also play a role in gene activation (Iwakawa and Tomari 2022). When serum is not present, AGO2 and FXR1 combine to form a complex that binds to particular mRNA sequences and starts translation. This phenomenon is seen in nondividing cells such as oocytes. miRNAs exert diverse effects within gene regulatory networks, influenced by both the abundance and properties of binding sites on their target RNA molecules. The strength of miRNA binding is affected by factors like mismatches, secondary structure of target RNA, and seed base pairing, potentially leading to a dosage dilution effect (Foster et al. 2021; de Souza et al. 2012). ceRNAs, such as circRNAs or lncRNAs with complementary MREs can impact miRNA binding and hinder transcriptional suppression by sequestering miRNAs. Single mRNAs can have multiple MREs for distinct miRNAs, amplifying the collaborative effect of miRNAs on gene regulation (Komatsu et al. 2023). For instance, the tumor suppressor p21Cip1/Waf1 is targeted by at least 28 miRNAs, indicating the coordinated action of miRNAs in gene regulation. The subcellular localization of the miRISC is crucial beyond MRE load, with various cellular compartments housing miRISC and influencing gene regulation by miRNAs, including the trans-Golgi network, nucleus, mitochondria, rough endoplasmic reticulum, lysosomes, stress granules, processing bodies, and endosomes (Wu et al. 2010).
When miRISC is found in the nucleus, it can interact with DNA to change chromatin structure and impact patterns of gene expression. It can also affect the alternative splicing of recently generated mRNA. The miRISC can encourage translational activation, prevent translation start, or promote the breakdown of mRNA throughout the cytoplasm, along with the mitochondria. Additionally, miRISC can hinder translation by binding to mRNA undergoing translation in the rough endoplasmic reticulum (ER) (Sadakierska-Chudy 2020). Additionally, it can be sent to lysosomes for breakdown or retained in stress granules (Khattri et al. 2015; Saloura et al. 2019). Finally, the miRISC can be internalized by endosomes via the Golgi and released to other cells via exosomes to promote cell–cell contact. MiRNAs have dynamic roles in gene regulation, impacting essential activities including transcription and translation in addition to the stability of mRNA. Consequently, miRNAs act as crucial regulatory molecules that govern the initiation and progression of human cancers (Nabariya et al. 2022). The process of miRNA biogenesis, previously discussed, is crucial for understanding the regulation of miRNAs in GBM. To further elucidate the role of miRNAs in GBM research, it is essential to identify which miRNA oncogenes are upregulated or downregulated. This knowledge is vital for determining the targets of ncRNAs that can significantly impact GBM research. Understanding the regulatory mechanisms of miRNAs can provide valuable insights into the development and progression of GBM, a highly aggressive and invasive brain cancer. By identifying the specific miRNAs involved in GBM, researchers can focus on the key molecular pathways and targets that contribute to the disease’s pathogenesis. This information can be used to develop novel therapeutic strategies, such as miRNA-based therapies, to effectively treat GBM.
miRNAs in GBM
MiRNAs have many functions in the initiation, progression, and treatment of GBM. They support several areas, including improved patient prognosis, treatment development and improvement, and tumor diagnosis. In the context of GBM, miRNAs are essential for controlling treatment resistance, metabolism, metastasis, and tumor growth. The dysregulated miRNA oncogenes and tumor suppressor genes in GBM elucidate the roles of specific miRNAs in processes such as angiogenesis or cell metastasis, paving the way for targeted therapy are detailed in Table 1. The expression levels of dysregulated miRNAs in GBM, ranging from low grade to high grade, are depicted in Fig. 4. To identify biomarkers essential for blocking cell migration and angiogenesis in GBM, these tables and figures are indispensable.
Heatmap of the differentially expressed miRNAs between low-grade GBM (n = 3) and high-grade GBM (n = 3). Each row represents the expression levels of a specific miRNA, and each column represents a single sample: Blue color denotes low miRNA expression, whereas red denotes high miRNA expression. H, high-grade GBM; L, low-grade GBM
Because of their complexity, miRNAs can regulate other genes that code for proteins, making them either tumor suppressors or oncogenes. In GBM, several miRNAs are dysregulated and linked to a low prognosis. A study by Moller et al. revealed a striking alteration in miRNA expression patterns within GBM tumors compared to normal brain tissue (Møller et al. 2013). Ninety-five miRNAs, including miR (137, 128, 34a, and 7), displayed a marked decrease in expression, while 256 miRNAs, including miR (93, 17, 21, and 221/222), exhibited a substantial increase in expression (Mafi et al. 2022). These discoveries suggest that miRNAs could play a role in the development and progression of GBM. These miRNAs exert a profound influence on the defining features of GBM, encompassing angiogenesis, self-renewal, cell proliferation, migration, invasion, and resistance to treatment. Furthermore, these miRNAs can function as biomarkers because they may be released via vesicles. In this context, we explore some well-studied miRNAs that have been linked to the control of important GBM biology pathways.
Cell proliferation and apoptosis
MiR-21, identified early on as a key miRNA associated with glioma aggressiveness, shows elevated expression levels in GBM and aligns with the World Health Organization (WHO) tumor grade classification. It regulates tumor suppressor genes, such as programmed cell death protein 4 (PDCD4); acidic leucine-rich nuclear phosphoprotein 32 (ANP32A); phosphatase and tensin homolog deleted on chromosome 10 (PTEN); SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 4 (SMARCA4); and Sprouty homolog 2 (SPRY2) (Aloizou et al. 2020). Silencing miR-21 markedly restricts tumor cell proliferation and growth in immunodeficient mice. Furthermore, miR-21’s antiapoptotic properties stem from its ability to suppress PDCD4, Tap63, and HNPRK, proteins that promote cell death. The interplay between p53 and miR-221/222 in GBM suggests that p53 is crucial for mediating the tumor-suppressive functions of these miRNAs, consistent with the poor prognosis associated with p53 mutations in GBM. This intricate interplay triggers the interaction of PUMA with Bcl-2 and Bcl-xL, initiating a cascade of events leading to cell death. However, elevated levels of miR-221/222 counteract PUMA’s effects, promoting cell survival (Rhim et al. 2022). Furthermore, reduced expression of miR-34a in GBM directly downregulates receptor tyrosine kinases and critical regulators, disrupting various cellular processes. MiR (58, 26a, 181, 15b, 153, 125b, 148a, 363, 1218, and 184) are among the other miRNAs involved in these processes (Shahzad et al. 2021).
Migration and invasion
MiR-21 stands as a master regulator, orchestrating the growth and survival of GBM cells while simultaneously exerting a crucial influence on their migratory and invasive potential. It inhibits the activity of proteins that break down the extracellular matrix, such as the tissue inhibitor of metalloproteinase 3 (TIMP3), which normally controls the levels of matrix-degrading enzymes. MiR-146b, an additional miRNA, has been observed to suppress matrix metalloproteinase 16 (MMP16), which encourages GBM cell entry. Analogously, miR-10b fosters cell invasiveness by upregulating matrix metalloproteinase 14 (MMP14) expression through HOXD10 suppression (Mazurek et al. 2020).
Angiogenesis
The intricate network of blood vessels that sustains GBM tumors is controlled by a group of miRNAs known as angiomiRs; these molecules play a pivotal role in governing the development and function of these vessels. MiR-296, a prominent angiomiR, shows elevated expression in endothelial cells under the influence of pro-angiogenic factors and is associated with the formation of tumor blood vessels (Annese et al. 2020). Reduction of miR-296 expression diminishes tumor angiogenesis, while increased miR-125b levels promote tumor vascularization through its effect on the transcription factor MAZ, highlighting the importance of miRNAs in regulating angiogenesis (Loubalova et al. (2023). Conversely, miR-218 inhibits GBM tumor neovascularization by targeting HIF-2α. In addition, miR-93, part of the miR-17 family, promotes cell viability and blood vessel formation by silencing integrin-β8 expression (Buruiană et al. 2020). Enhancing the expression of miR-93 in GBM cells induces the proliferation of endothelial cells and accelerates the formation of blood vessels in mouse xenografts.
Long noncoding RNAs
Long noncoding RNA production
In the realm of cancer research, lncRNAs have emerged as a subject of intense scrutiny. These RNA molecules, surpassing 200 nucleotides in length, exhibit gene-like functions despite lacking protein-coding potential. LncRNAs have a 5′ cap and a 3′ polyadenylated tail (Liu et al. 2016). The generation of these RNAs is guided by RNA polymerase II. Generally, lncRNAs have more than two exons, with more than 60% of them having poly(A) tails. lncRNAs can be categorized according to their genomic origin and distribution, including promoter, antisense, enhancer, bidirectional, intergenic, intron, sense, or bidirectional genes. Although current research has extensively examined the methods by which lncRNAs control GBM, given their complicated and diverse mechanisms, more research is necessary to understand their biogenesis and pathogenesis fully (Statello et al. 2021).
Long noncoding RNAs’ functions
To elucidate the regulatory mechanisms of lncRNAs in GBM, an aggressive and invasive brain cancer, researchers aim to uncover how these long non-coding RNAs contribute to disease progression and potential therapeutic targets. By identifying specific lncRNAs implicated in GBM, this study aims to pinpoint critical molecular pathways and therapeutic targets crucial to the disease’s progression. These insights will facilitate the development of innovative therapeutic approaches, including lncRNA-targeted therapies, to advance GBM treatment strategies effectively. lncRNAs play diverse roles in gene regulation, encompassing chromatin remodeling, transcriptional regulation, and post-transcriptional processes (Krysler et al. 2022). They influence gene expression by interacting with transcription factors, forming R-loops on gene promoters, and modulating the translation of effector proteins. This functional versatility highlights the multifaceted nature of lncRNAs and their profound impact on cellular processes. Notably, lncRNAs MEG3 suppresses cellular-myelocytomatosis (c-Myc), inhibiting bladder cancer cell invasion by enhancing PHLPP2 gene translation. LncRNAs also participate in chromatin modification, impacting the structure and catalyzing covalent modifications (Mattick et al. 2023). LINC01123 has recently been identified as a miRNA sponge, capable of sequestering specific miRNA sites and competing with endogenous RNAs for binding. In nonsmall cell lung cancer, LINC01123 plays a central role in augmenting c-Myc expression by acting as a decoy for miR-199a-5p. This effectively sequesters miR-199a-5p, preventing it from binding to and suppressing c-Myc activity. LncRNAs function at both the transcriptional and translational levels to regulate various aspects of tumor cell growth, including differentiation, proliferation, epigenetic inheritance, and genomic imprinting. This emphasizes how urgent it is to conduct additional studies to clarify their biological roles and regulatory mechanisms (Zhang et al. 2020a, 2020b).
Long noncoding RNAs in GBM
Recent findings highlight the ability of lncRNAs to influence both GBM cell behavior and patient outcomes, suggesting their potential as therapeutic targets or prognostic markers. It has been discovered that lncRNAs significantly influence the growth, invasion, apoptosis, metastasis, and GBM cells’ resilience to chemotherapeutic drugs. GBM cells’ resilience to chemotherapeutic drugs. Expanding our understanding of lncRNAs in GBM pathogenesis may lead to the development of novel biomarkers for early detection, and improved patient outcomes are listed in Table 2. lncRNA expressions in low-grade to high-grade GBM are depicted in Fig. 5. The outcomes presented in these figures and tables provide insight into potential biomarkers in lncRNAs for predicting GBM.
Heatmap of the differentially expressed lncRNAs between low-grade GBM (n = 3) and high-grade GBM (n = 3). Each row represents the expression levels of a specific lncRNA, and each column represents a single sample: Blue color denotes low lncRNA expression, whereas red denotes high lncRNA expression. H, high-grade GBM; L, low-grade GBM
Long noncoding RNAs revealing mysteries in GBM
LncRNAs are essential for controlling several important GBM activities, such as cellular invasion, migration, proliferation, and GSC behavior. These pathways function as bridges, connecting the intricate network of signaling cascades that govern GBM progression, including PI3K/Akt/mTOR, Wnt-β-catenin, MAPK, and Notch pathways (Chaudhary 2021). Studies by Zhang and Han, employing microarray mining and high-throughput screening, identified differentially expressed lncRNAs in GBM compared to normal brain regions (Zhang et al. 2020a, 2020b). Exploration of RNA-sequencing data from the Cancer Genome Atlas (TCGA) unveiled 1288 lncRNAs with altered expression levels in all GBM. Within this set, 282 were associated with enhanced survival rates, while 584 were correlated with an unfavorable prognosis specifically for GBM. Noteworthy lncRNAs like MEG3, HOTAIRM1, and CRNDE consistently exhibited abnormal expression in GBM, indicating their potential significance in gliomagenesis (Ding et al. 2020).
Cell proliferation and apoptosis
Colorectal neoplasia differentially expressed (CRNDE), initially recognized for its elevated expression in colorectal cancer, demonstrates increased levels in diverse malignancies, particularly in GBM. Experimental suppression of CRNDE demonstrated its oncogenic function, resulting in diminished cell proliferation and migration in vitro. Furthermore, it hampered GBM tumor growth in vivo. The ability of CRNDE to sequester miR-384, a miRNA responsible for targeting and suppressing PIWIL4, contributes to GBM progression by hindering the silencing of PIWIL4 and its subsequent effector, STAT3. Furthermore, CRNDE modulates the growth, migration, and invasion of GSCs by capturing miR-186 (Xie et al. 2020). Low expression of the lncRNA maternally expressed gene 3 (MEG3) in GBM is linked to a less favorable prognosis, characterized by decreased overall survival, increased risk of recurrence, higher tumor grades, and wild-type isocitrate dehydrogenase (IDH) gene status. Overexpression of MEG3 inhibits GBM, a cell growth, and increases apoptosis and autophagy in vitro, suggesting a potential role as a ceRNAs due to its direct binding sites for several miRNAs (Huang et al. 2022). In GBM, the ncRNA PLAC2 plays a role in tumor suppression by initiating cell cycle arrest and hindering the proliferation of cancer cells. Increased PLAC2 levels control the expression of ribosomal protein L36 (RPL36), which results in G1/S arrest and successfully prevents the growth of GBM in vivo (Kovalchuk 2021). PLAC2’s regulatory actions are facilitated by its interactions with RPL36 and signal transducer and activator of transcription 1 (STAT1) (Momtazmanesh and Rezaei 2021).
Migration and invasion
It has been demonstrated that the recently discovered lncRNA-ATB (an RNA gene affiliated with the lncRNAs class) stimulates GBM cell invasion when transforming growth factor-beta (TGF-β) is present. Elevated ATB levels initiate the nuclear factor kappa B (NF-Κb) signaling cascade, resulting in p65’s nuclear translocation. Stimulation of the NF-κB signaling pathway amplifies TGF-β-mediated invasion of GBM cells. The upregulation of lncRNA-ATB stimulates the activation of the NF-κB signaling pathway, which leads to the transport of p65 into the cell nucleus. This process facilitates the TGF-penetration β’s of GBM cells. Furthermore, lncRNA-ATB activates the p38/MAPK pathway in GBM cells to improve TGF-β-mediated invasion. Epigenetically induced MYC interacting lncRNA-1 (EPIC1), a lncRNAs, has emerged as a critical regulator of cellular processes and exhibits a striking increase in GBM, a malignant brain tumor. By focusing on Cdc20, its inhibition causes apoptosis, lowers cell viability, enhances sensitivity to temozolomide (TMZ), and considerably lessens cell invasion (Tang et al. 2019).
Angiogenesis
One of the first lncRNAs to be found was H19, which is expressed by the mother and is also higher in GBM. A well-known oncogene called c-Myc has been linked to the control of H19, a ncRNA, in GBM. H19, in turn, has been shown to interact with miR-138, a miRNA, acting as a competitive endogenous RNA (ceRNA) to modulate the expression of HIF-1α, a hypoxia-inducible factor. HIF-1α plays a crucial role in angiogenesis, the formation of new blood vessels. Therefore, the c-Myc-H19-miR-138-HIF-1α axis may contribute to GBM angiogenesis. Furthermore, GBM exhibits an increased expression of CCAT-1, which also functions as a ceRNA (Liao et al. 2023). CCAT-1 plays a role in advancing GBM tumor progression by capturing miR-181b and reinstating the activity of its original targets, PDGFR-α and fibroblast growth factor receptor 3 (FGFR3). Furthermore, CCAT-1 knockdown impedes the proliferation, migration, and epithelial-mesenchymal transition (EMT) of GBM cells, while simultaneously inducing apoptosis (Wang and Li 2020). Notably, lncRNAs are increasingly recognized as crucial players in cancer progression and metastasis, modulating various physiological and cellular regulatory pathways. Unlocking the diagnostic and therapeutic potential of lncRNAs in GBM and other cancers necessitates further research. The exploration of lncRNAs as potential molecular targets in GBM is still in its early stages (Yin et al. 2020).
Biogenesis of circular RNAs
This study seeks to investigate the regulatory mechanisms of circRNAs in GBM, a highly aggressive and invasive brain cancer. By identifying circRNAs specifically involved in GBM, the research aims to delineate pivotal molecular pathways and therapeutic targets essential for understanding disease progression. These findings are anticipated to drive the development of novel therapeutic strategies, including circRNA-targeted therapies, to enhance the efficacy of GBM treatment approaches. The discovery of circRNAs in RNA viruses was made by using electron microscopy. The structure of the circRNAs was confirmed in 1993. RNA polymerase II-transcribed precursor RNA and RNA-binding protein are required to produce circRNAs. Intronic, exonic, and eciRNAs are the three different types of circRNAs (Patop et al. 2019).
Functions of circular RNAs
CircRNAs primarily act as miRNA sponges, hindering the interaction between miRNAs and their target genes. This mechanism safeguards target gene stability by protecting mRNA from miRNA-mediated degradation. CircENTPD7 promotes the survival and proliferation of GBM cells by sequestering miR-101-3p, leading to increased expression of ROS1. Similarly, circFOXO3, discovered by Zhang et al. (2019a, 2019b) acts as competitive endogenous RNAs (ceRNAs), sequestering miR-138-5p and miR-432-5p, thereby promoting the overexpression of nuclear factor of activated T cells 5 (NFAT5) (Zhang et al. 2019a, 2019b). This mechanism facilitates cancer cell spread and dissemination (Xiao et al. 2022). CircRNAs can directly modulate the expression of target genes by interacting with RNA-binding proteins. They can also disrupt the interactions between miRNAs and RNA-binding proteins, indirectly affecting the function of RNA-binding proteins. For example, circ-AMOTL1 interacts with c-Myc, enhancing c-Myc expression and stability in the nucleus. Furthermore, circ-AMOTL1 expression augments c-Myc’s affinity for various promoters. Moreover, circRNAs impact the pace of processes such as phosphorylation and ubiquitination by facilitating the binding of specific enzymes to substrates. Residing predominantly within the nucleus, circRNAs possess the remarkable ability to recruit proteins to specific locations, such as promoters, during transcription (Ali Syeda et al. 2020). This unique capability empowers the modulation of gene expression and the amplification of ribonucleoprotein function. Some circRNAs are transported by exosomes, participating in intercellular communication (Hanif et al. 2017). CircRNAs can be translated into proteins by utilizing m6A modifications and internal ribosome entry sites (IRES) to interact with ribosomes at the 5′ untranslated region (UTR) of mRNA. During translation initiation, m6A modifications interact with eukaryotic initiation factor 3 (EIF3), while IRES promotes circRNA translation by facilitating ribosome recruitment (Huang et al. 2020). In summary, circRNAs serve diverse roles, with their well-studied function as a miRNA sponge, particularly in diseases like GBM. Future research endeavors are expected to uncover new avenues for identifying and managing GBM using circRNA (Han et al. 2022).
Circular RNA-redefining the genetic landscape
CircRNAs are a distinct class of ncRNAs that form a closed loop due to the covalent linkage of their ends, lacking both 3′ poly(A) tails and 5′ caps. Formed through a distinctive splicing process known as back-splicing, circRNAs display tissue-specific roles. Despite generally having low expression levels, their closed-loop structure imparts remarkable stability, making them resistant to degradation by ribonucleases. Discovered in 1976 in association with potato spindle tuber disease, circRNAs were initially considered “splicing noise” but have since been recognized for their abundance and crucial roles, especially in cancer (Liu et al. 2022a, 2022b, 2022c).
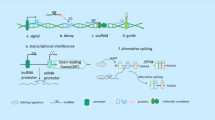
Three main types of circular RNAs comprise exon–intron circRNAs (EIciRNAs), exon circRNAs (ecircRNAs), and circular intron RNAs (ciRNAs). CircRNAs arise from various mechanisms, including lariat-driven circularization, RNA-binding protein (RBP) mediated circularization, and intron-pairing-driven cyclization. EIciRNAs, defined by stable intron retention between exons, are considered intermediates in circRNA biosynthesis. The canonical splicing machinery creates circular RNAs by excising lariat introns, and their resistance to debranching enzymes is attributed to specific motifs. CircRNAs serve diverse functions, such as acting as decoys for proteins, sponging miRNAs, and influencing transcriptional control by competing with linear pre-mRNA splicing to modulate gene expression. Some developmental models also propose the potential for circRNAs to undergo translation. Figure 6 illustrates the biogenesis or mechanism of circular RNA which becomes dysregulated, potentially triggering GBM-like miRNA sponges, proliferation, and metastasis (Zhao et al. 2019).
CircRNAs are crucial players in the development and growth of GBM. Analyzing RNA-seq data from GBM and normal brain tissues in this study revealed 476 circRNAs with unique expression patterns, highlighting their potential as diagnostic tools or therapeutic targets for GBM. One such circRNA is cir-ITCH, a downregulated circRNA serving as a predictive biomarker for GBM. Cir-ITCH enhances the activity of the tumor suppressor gene ITCH by suppressing the Wnt/β-catenin signaling pathway. This suppression prevents cell division and enhances ITCH expression by sequestering miR-214. In contrast, circ-TTBK2, discovered by Zheng et al., is elevated in GBM, promoting cell invasion, migration, proliferation, and suppressing apoptosis (Zheng et al. 2017). Its mechanism involves control of the HNF1β/Derlin-1 pathway and acting as a miR-217 sponge. Another notable circRNA, cZNF292, is expressed under hypoxic conditions and modifies the Wnt/β-catenin pathway, leading to the inhibition of GBM cell growth, tube formation, and induction of S/G2/M phase cell cycle arrest (Zhang et al. 2017). This version of the sentence is more concise and emphasizes the multifaceted nature of circRNAs’ involvement in GBM biology. It also uses the word “roles” instead of “functions” to convey a more specific and active involvement. The prevalence of circRNAs in the human brain underscores their growing importance in GBM development and spread. Harnessing circRNAs for GBM diagnosis and therapy necessitates a thorough understanding of these biological entities and their involvement in GBM development (Tang et al. 2024).
Circular RNAs in GBM
Growing data points to circRNAs as a new focus for examining the relationship between different types of cancer and noncoding RNAs. It is currently unknown how circRNAs function in cancer, particularly in GBM. The prevailing focus of recent circRNA research in GBM has been on their miRNA sponge function. This entails circRNAs contributing to the advancement of GBM by attaching to miRNA’s downstream target genes and preventing miRNA from binding to target mRNA. As a result, mRNAs protected from miRNA degradation by this protective effect (Tang et al. 2021). Furthermore, circRNAs have been linked in several studies to possible implications for GBM prognosis and diagnosis. This paper sheds light on the molecular mechanisms underlying circRNA function and synthesizes their oncogenic and tumor-suppressive roles, particularly their miRNA sponge activity in GBM is detailed in Table 3. CircRNA expression in low-grade and high-grade GBM is explained in the heat map of Fig. 7. The discovery of alterations in circRNA oncogenes and tumor suppressor genes sheds light on potential therapeutic targets for GBM.
Heatmap of the differentially expressed circRNAs between low-grade GBM (n = 3) and high-grade GBM (n = 3). Each row represents the expression levels of a specific circRNA, and each column represents a single sample: Blue color denotes low circRNA expression, whereas red denotes high circRNA expression. H, high-grade GBM; L, low-grade GBM
Circular RNAs as tumor promoters
New evidence suggests that H19 can interact with miR-138 and function as ceRNA to modulate HIF-1α expression, thereby promoting angiogenesis in GBM. Interestingly, it was found that GBM had elevated expression of circ-MELK and that this higher expression was linked to enhanced viability and proliferation of GBM cells. In stark contrast, a substantial reduction in circ-MELK levels significantly suppressed GBM proliferation in both in vitro and in vivo settings. A prior investigation demonstrated the function of circ-MELK in supporting GBM cells’ mesenchymal transformation and maintaining their tumor stem cell qualities. Sponging miR-593 was used to target EPH receptor B2 (EphB2) to produce this result. The RNA-binding protein adenosine deaminase RNA-specific B2 (ADARB2) interacts with the flanking region of the circRNA’s intron, promoting the formation of CircNT5E, also known as circular 50-nucleotidase ecto (NT5E). In GBM, heightened CircNT5E expression is frequently linked to enhanced proliferation and distant metastasis of GBM cells (Liu et al. 2020). CircNT5E inhibits the proliferation, invasion, and migration of GBM cells by interacting with miR-422a, thereby counteracting the oncogenic effects of PIK3CA and NT5E (Zhang et al. 2020a, 2020b). Conversely, circ-LGMN promotes GBM development by modulating the miR127-3p/Legumain LGMN axis, enhancing tumor cell growth and infiltration. EIF4A3-induced circASAP1 is elevated in temozolomide-resistant GBM, contributing to drug resistance by upregulating NRAS through miR-502-5p sequestration. CircFOXO3 (Zhang et al. 2019a, 2019b), highly expressed in GBM tissues, promotes invasion and migration by sequestering miR (432-5p and 138-5p) and upregulating NFAT5. E-cadherin-RNA activates EGFR signaling in GBM, crucial for tumor initiation and progression (Chen et al. 2021). CircMMP9 increases CDK4 and AURKA production, facilitating GBM cell proliferation, invasion, and migration. By altering proteins involved in DNA damage repair, low-dose radiation causes the release of ldrEXOs and circ-METRN, which influences the course of GBM and the response to radiotherapy. Through its binding to GRB14/PDGFRα and inhibition of its expression, miR-4709-3p facilitates the growth, invasion, migration, and resistance to radiation of GBM cells. Overall, these findings underscore the significance of circRNAs in GBM progression, invasion, metastasis, and therapeutic resistance (Gao et al. 2021).
Circular RNAs as tumor suppressors
circRNAs, a relatively new area of research, are showing promise as tumor suppressors. These unique RNA molecules with a closed-loop structure act like sponges, soaking up miRNAs that would normally target and inhibit genes (Chen et al. 2021a, 2021b). By binding these miRNAs, circRNAs allow genes with tumor-suppressing functions to operate freely. This “miRNA sponge” effect is just one way circRNAs can suppress tumors. They can also directly interact with proteins involved in cell growth and division, or even produce functional proteins themselves. Researchers are finding that abnormal circRNA expression is linked to various cancers, suggesting their potential as biomarkers for early cancer detection. While some circRNAs promote cancer, the tumor-suppressive ones hold promise for future therapies. By manipulating circRNA activity, scientists hope to develop treatments that can boost the body’s natural anti-tumor defenses. Understanding circRNAs is a growing field of cancer research with exciting possibilities. Despite their widespread involvement in gene regulation and their roles in the pathophysiological processes and carcinogenesis of tissues and cells within GBM, specific circRNAs have demonstrated tumor-suppressive properties in GBM. The finding that GBM patients have higher expression levels of these circRNAs lends additional credence to this. In healthy brain tissue, SNF2 histone linker PHD RING helicase (circ-SHPRH) is robustly expressed, but its expression is markedly diminished in GBM. The SHPRH-146AA variation protects SHPRH from ubiquitin–proteasome degradation, which improves SHPRH’s capacity to ubiquitinate PCNA. Consequently, there is a decrease in both tumorigenicity and tumor growth. A further illustration is the AKT transcriptional variation circAKT3, which has significantly reduced expression levels in GBM tissues as compared to nearby typical safeguards (Ahmed et al. 2022). The circAKT3 gene interacts with phosphorylated pyruvate dehydrogenase kinase 1 to produce the AKT3-174aa protein, which is present in GBM and inhibits the PI3K/AKT signaling pathway (PDK1). Radiation resistance, tumorigenic potential, and GBM cell proliferation are all inhibited as a result of this relationship. Patients’ long-term prognosis may benefit from such inhibitory effects (Khan et al. 2022). Cerebellar degeneration-related protein 1 (circCDR1) showed significantly lower protein expression in GBM, suggesting its potential as a predictor for overall survival (OS) in GBM patients. By removing ubiquitin modifications from p53, CircCDR1 stabilizes the protein and prevents the p53/MDM2 complex from forming. This disruption shields GBM cells from DNA damage, acting as a tumor suppressor (Dai et al. 2023). In GBM, Yang et al. investigation showed that circ-FBXW7 and its functional protein FBXW7-185aa were less expressed. Increasing their expression improved OS by reducing cell proliferation, accelerating the cell cycle, and highlighting their potential as prognostic markers. Moreover, circCD44 was demonstrated to act as a molecular sponge for miR-326 and miR-330-5p, thereby regulating SMAD6 expression and impeding the growth, invasion, and migration of GBM cells (Yang et al. 2018). Despite these findings, the exact mechanism by which circular RNAs suppress tumor growth in GBM remains unknown, emphasizing the need for further research into their unique roles and mechanisms.
Therapeutic targeting of noncoding RNAs
For decades, the focus of our understanding of RNA has been on mRNA, the molecule that carries genetic instructions from DNA to ribosomes for protein production. However, recent research has revealed a vast and exciting world of ncRNAs that play crucial roles in regulating gene expression and various cellular processes. This newfound knowledge has opened a new frontier in therapeutic development: targeting ncRNAs for disease treatment. ncRNAs encompass a diverse group of RNA molecules that lack the protein-coding capacity of mRNA. These include miRNAs, lncRNAs, and circRNAs. Each type has its unique structure and function, influencing gene expression through various mechanisms. miRNAs, for example, act as post-transcriptional regulators, binding to specific mRNA sequences and promoting their degradation or inhibiting translation. LncRNAs, on the other hand, can function as either activators or repressors of gene expression by interacting with DNA, RNA, or proteins. The exciting potential of ncRNAs as therapeutic targets lies in their involvement in various diseases. Many ncRNAs are dysregulated in diseases like cancer, neurological disorders, and cardiovascular diseases. For instance, specific miRNAs might be overexpressed in cancer cells, promoting uncontrolled proliferation. Conversely, other miRNAs might be underexpressed, hindering tumor suppressor genes. By targeting these dysregulated ncRNAs, we can potentially restore a healthy balance in gene expression and influence disease progression. Several strategies are being explored to therapeutically target ncRNAs. Antisense oligonucleotides (ASOs) are short, single-stranded DNA or RNA molecules designed to bind complementary sequences on target ncRNAs. This binding can either promote RNase H-mediated degradation of the target ncRNA or block its interaction with other molecules, altering its regulatory function. Another approach utilizes miRNA mimics or antagonists. Mimics are synthetic versions of miRNAs that can replace those deficient in a disease state, while antagonists are molecules that block the activity of overexpressed miRNAs.
Furthermore, researchers are developing small interfering RNA (siRNA) therapies specifically designed to target disease-associated ncRNAs. siRNAs are double-stranded RNA molecules that induce the silencing of a specific target RNA through a cellular machinery called RNA interference (RNAi). This approach holds promise for targeting not only ncRNAs but also mRNAs associated with disease (Teplyuk et al. 2016). Despite the immense potential, therapeutic targeting of ncRNAs faces several challenges. One major hurdle is ensuring the specificity of the therapeutic molecule. Off-target effects, where the molecule binds to unintended RNA sequences, can lead to unintended consequences. Additionally, delivering these therapies to the desired tissues within the body can be challenging (Nappi 2024). ncRNAs often reside within complex cellular environments, and efficient delivery systems are crucial for maximizing therapeutic efficacy. Another challenge lies in fully understanding the complex roles of ncRNAs. While we are rapidly unraveling their functions, many ncRNAs remain poorly characterized (Carrano et al. 2021). Continued research is essential to identify ncRNAs with therapeutic potential and elucidate their precise mechanisms of action (Yadav et al. 2024).
ncRNAs are found in large quantities in GBM and have been shown to have important functions in controlling how these tumors grow and behave. They may also be useful as biomarkers or targets for therapy. The use of morpholino oligonucleotides (MOs), ASOs, locked nucleic acids (LNAs), and peptide nucleic acids (PNAs) are emerging methods for precisely addressing noncoding RNAs. Single-stranded oligonucleotides called ASOs are created with sequences that match target RNAs. They attach to the target directly and encourage RNase H to break them down (Desgraves et al. 2024). This is how they exert their inhibitory effect and can be utilized to target various RNA types, including mRNAs, circRNAs, lncRNAs, miRNAs, and piRNAs. Amazingly, they can be transported without a carrier. Interestingly, ASOs are successful in suppressing the expression of lncRNA MALAT1, which reduces tumor burden and cancer cell metastasis in mice models (Dhuri et al. 2020). Teplyuk et al. also employed ASOs to suppress miR-10b in murine GL261 allograft models and xenografts produced from human GBM progenitor cells. The administration of miR-10b ASO via three different routes intravenous injections, osmotic delivery pumps, and direct tumor injections successfully suppressed miR-10b activity with minimal adverse effects. Inhibiting the activity of miR-21 led to the liberation of target RNAs, causing a notable decrease in tumor growth and progression. LNAs, single-stranded oligonucleotides with exceptional resistance to nuclease degradation, effectively counteracted the anti-apoptotic effects of miR-21 in the GBM cell line (Teplyuk et al. 2016). LNCs linked to LNAs were created by Griveau et al. to successfully sensitize silencing miR-21 sensitized U87MG GBM cells to radiation-induced cell death (Griveau et al. 2013). In another study, Wang et al. demonstrated that silencing miR-381 using an LNA enhanced the sensitivity of U251 GBM cells to TMZ (Wang et al. 2015).
Peptide nucleic acids (PNAs), modified with four lysine residues, effectively blocked the function of miR-155 in vivo in mice. Employing GDC-0084, a modified morpholino engineered for enhanced blood–brain barrier (BBB) permeability (Salphati et al. 2016), effectively inhibited the PI3K pathway, thereby impeding tumor growth in an orthotopic xenograft model of GBM (Salphati et al. 2016). Apart from MOs, miRNA-based therapies include miRNA mimics and antagomirs. Antagomirs, 22–23 nucleotide RNA analogs, suppress miRNA expression and can be directly administered to cells without delivery vehicles. MiRNA mimics, unlike miRNA inhibitors, function by replicating the sequence of mature endogenous miRNAs, leading to an increase in target miRNA expression. Sun et al. study demonstrated that miR-137 mimics inhibited the proliferation of brain stem cells (Sun et al. 2011). In addition to nucleotide-based therapies, sncRNA molecules, such as siRNAs and shRNAs, have emerged as promising therapeutic agents. SiRNAs interact with target RNAs via complementary binding, effectively silencing them through the RISC. Similar to siRNAs, shRNAs find widespread use for targeting and silencing various noncoding RNAs, both in vivo and in vitro. The CRISPR-Cas9 system enables precise gene editing, offering a medicinal approach to noncoding RNAs. Peng et al. effectively knocked down linc-RoR in breast cancer cell lines using CRISPR-Cas9, attenuating ERK activation induced by estrogen deprivation (Peng et al. 2017). Overall, these diverse therapeutic approaches hold promise for targeting noncoding RNAs in various diseases. In addition to nucleotide- and RNAi-based methods, small compounds may potentially be used, depending on their structural characteristics, to target noncoding RNAs therapeutically (Chen et al. 2018). Certain high-throughput screening methods have been utilized to identify small compounds capable of binding to various structural configurations of RNA, and this includes the use of small-molecule microarrays.
Bose et al. targeted screening of MCF-7 cells revealed streptomycin as a potent inhibitor of miR-21, with its inhibitory efficacy matching that of a miR-21-specific antisense oligonucleotide (ASO) (Bose et al. 2012). Informa, a chemoinformatics method created by Disney and associates, has been applied more recently to find multiple lead drugs for RNA targeting. This approach involves utilizing two-dimensional combinatorial screening (2DCS) to identify and validate interactions between small molecules and RNA motifs. Subsequently, sequencing structure–activity relationship through sequencing (StARTS) is employed to determine binding fitness scores based on structure–activity relationships. This strategy offers a promising avenue for the development of novel therapeutic agents by targeting specific RNA motifs with small molecules. As a result, Inforna can forecast the targetable structural patterns of any RNA, including ncRNA, as well as the fitness of their interactions and the matching small molecule matches. Utilizing this approach unveiled G Neomycin B (G Neo B) as a prospective small-molecule therapeutics. G Neo B functions by directly binding to the Drosha binding site, effectively hindering the maturation process of miR-10b. This disruption of miR-10b maturation holds promise for therapeutic intervention in diseases associated with miR-10b dysregulation (Grillone et al. 2020).
Emerging paradigms of miRNAs, lncRNAs, and circRNAs in angiogenesis
Emerging evidence states that miRNAs, lncRNAs, and circRNAs are important for controlling many aspects of tumor biology, such as invasion, metastasis, proliferation, migration, and changes in the local tumor microenvironment. Figure 8 depicts the role of ncRNAs in GBM development, progression, and metastasis. The main topic of this article is research on targeting the ncRNAs in GBM that explores their role in immune evasion, migration, intravasation, extravasation, and the creation of pre-metastatic niches. Immune escape, a crucial stage in the genesis and spread of tumors, involves the recruitment of immune cells and the polarization of macrophages into M1 (classically activated) or M2 (alternatively activated) states. lncRNAs are essential for controlling the activity of macrophages and innate immunity. For instance, studies have shown that lncRNA-MM2P and SNHG15 trigger M2 polarization, while CamK-A encourages macrophage recruitment (Zhao et al. 2021). Tumor cells change from epithelial to mesenchymal through a biological process called epithelial-mesenchymal transition (EMT) (Lu et al. 2023b). This makes it easier for them to spread and get into nearby tissues. In mouse models, GATA6-AS amplifies TGF-β2-induced EMT and encourages angiogenesis, while MALAT1 increases the expression of VEGFA, SLUG, and TWIST to support angiogenesis and EMT in colorectal cancer (Duan et al. 2023). Similarly, circRNA-MYLK and circ-CSPP1 aid in EMT and cancer advancement by initiating relevant signaling pathways. Critical mechanisms called intravasation and extravasation allow cancer cells to enter and exit the bloodstream, which promotes metastasis (Singh et al. 2017). Researchers have found that lncRNA-ATB controls TGF-β signaling to help tumor cells invade, intravasate, and colonize, while circ-CCAC1 damages the integrity of the endothelial barrier and encourages the growth of new blood vessels (Zepeda et al. 2023). The term “pre-metastatic niche” describes the altered microenvironment of distant organs before the arrival of metastatic tumor cells (Gupta et al. 2024). These alterations include those in the extracellular matrix (ECM), blood supply, immunological response, and cellular makeup (Thakur et al. 2021). H19 and CDR1as play a role in angiogenesis and ECM remodeling, while XIST and MALAT1 affect fibroblast growth factor production and release (Xia et al. 2023). Recent studies demonstrate how circRNAs and lncRNAs alter the tumor microenvironment to influence several aspects of tumor development. Finding out how Angio-LncRNAs and Angio-CircRNAs work in angiogenesis could help scientists make drugs that stop tumors from growing by blocking angiogenic processes.
LncRNAs and circRNAs play pivotal roles in the multistep progression of tumors, including ECM remodeling, pre-metastasis events, angiogenesis, and macrophage recruitment. Abbreviations: CD31, cluster of differentiation 31; SNHG15, small nucleolar RNA host gene 15; MP, macrophage polarization; TGF-β2, transforming growth factor beta 2; CDK6, cyclin-dependent kinase 6; CD206, cluster of differentiation 206 (also known as MRC1); lncRNA, long noncoding RNA; M1, classically activated macrophage; M2, alternatively activated macrophage; EMT, epithelial-mesenchymal transition; FGF, fibroblast growth factor; VEGF, vascular endothelial growth factor; FB, fibroblast; MALAT1, metastasis-associated lung adenocarcinoma transcript 1; H19, long noncoding RNA H19; CDR1as, cerebellar degeneration-related protein 1 antisense; ECM, extracellular matrix; HMGA2, high mobility group AT-Hook 2; MED12, mediator complex subunit 12; DC, dendritic cell; TAP, tumor antigen cross-presentation; MDSC, myeloid-derived suppressor cell; T cell, T lymphocyte; NK cell, natural killer cell; ITL, inhibition of tumor cell lysis; CC, cancer cell; CCP, cancer cell proliferation; MT, metastatic tumor; circRNA MYLK, circular RNA myosin light chain kinase. Created with Biorender.com
Clinical applications of miRNAs, lncRNAs and circRNAs
Angiogenesis formation by miRNAs, lncRNAs, and circRNAs, as previously mentioned, have a major impact on a variety of phenotypes that include tumor angiogenesis, cell proliferation, EMT, apoptosis, and metastasis (Zhou et al. 2024). As a result, these compounds have the potential to serve as useful indicators and potent targets and biomarkers for cancer treatment. Despite their potential, clinical settings have proven the usefulness of a small number of circRNAs and lncRNAs as therapeutic, prognostic, and diagnostic biomarkers. A variety of cancer types have found tumor-associated Angio-LncRs, such as H19, MALAT1, and HULC, to be plasma biomarkers. Similarly, angio-CircRs such as circHIPK3 serve as prognostic markers in various malignancies (Badowski et al. 2022). However, further research is necessary to comprehensively assess their functional roles both in vivo and in vitro, intending to validate LncRNAs and circRNAs as innovative and effective therapeutic alternatives in clinical settings.
Targeting angiogenesis by miRNAs, lncRNA, and circRNA provides a viable method of preventing tumor growth and spread. Current RNA-based treatment strategies, such as RNA interference (RNAi) and antisense oligonucleotides (ASOs), target different RNAs and specific areas. The role of noncoding RNAs (ncRNAs) in angiogenesis is illustrated in Fig. 9, while RNA interaction-based therapies are also depicted. (Di Martino et al. 2022). Notably, the US FDA has approved patisiran, the first medication based on RNA interference, to treat inherited transthyretin amyloidosis. Studies on people with Angelman syndrome have indicated that ASOs can target lncRNAs. Additionally, intravenous ASOs that target the lncRNAs TUG1 and PVT1 have been shown to stop tumor growth and differentiation when used with the right drug delivery systems (Urits et al. 2020). The successful suppression of tumor angiogenesis and metastasis in cancer models has demonstrated the therapeutic promise of in vivo RNA interference (RNAi) and pharmaceutical therapies targeting FLANC (Tian et al. 2021). Furthermore, certain small interfering RNAs (siRNAs) have proven to be quite effective in inhibiting metastasis. Additionally, peptides encoded by Angio-CircRs may be useful in cancer treatment, according to recent research (Zhang et al. 2023). In the future, combining traditional treatments with focused strategies against Angiogenesis by targeting MiRNAs, LncRNAs, and CircRNAs may synergistically improve anti-angiogenic cancer treatments, improving our knowledge of and ability to treat cancer.
circRNAs, lncRNAs, and miRNAs play crucial roles in angiogenesis development in GBM, facilitated by growth factors such as VEGF and FGF. Their dysregulation can be targeted through RNA interaction–based therapies, including antisense oligonucleotides (ASOs), morpholinos (MOs), locked nucleic acids (LNAs), and peptide nucleic acids (PNAs) which have been shown to halt tumor growth and differentiation when administered with appropriate drug delivery systems
Recent studies have found miRNA, lncRNAs, and circRNAs to be important regulators influencing immunotherapy tolerance, radiation sensitivity, and chemotherapy resistance in cancer treatment for GBM (Ma et al. 2024). For instance, various cancer types have linked resistance to temozolomide to LncRNAs like MALAT1, MEG3, and CRNDE. Similarly, circRNAs such as circ-SMARCA5 and hsa_circ_0023404 have been associated with chemotherapy resistance in cancers (Qin et al. 2023). Also, when GBM is treated with radiation, circRNAs like cZNF292 affect how sensitive hypoxic tumor cells are to radiation. has-circRNA-002178 can change the expression of immune checkpoint molecules PD-1 and PD-L1, which can change how well immunotherapy works (Salami et al. 2022). Despite the approval of anti-angiogenic medicines like bevacizumab that targets angiogenesis in cancer therapy, there are still limited clinical applications that specifically target angiogenesis by miRNAs, lncRNAs, and circRNAs. Finding these patients’ levels of miRNAs, lncRNAs, and circRNAs may help predict the safety and effectiveness of medications, allowing for more individualized treatment plans with specific dosages or administration techniques. Overall, it looks like better clinical outcomes in cancer treatment are possible with the help of new biomarkers and treatment methods for targeting angiogenesis by miRNAs, lncRNAs, and circRNAs.
Conclusions and future directions
The successful outcomes achieved in recent years have spurred the scientific community to delve deeper into RNA research. Reflecting on the past, we now find ourselves at the threshold of the RNA therapeutics era, having achieved several milestones. This review highlights the potential for diagnostic and therapeutic uses of angiogenic miRNAs, lncRNAs, and circRNAs, while also highlighting their increasing roles and mechanisms in tumor progression. Angiogenic molecules, responsible for a variety of malignancies, impact angiogenesis, cell proliferation, EMT, apoptosis, and metastasis through a variety of molecular processes. Frequently occurring dysregulations in miRNAs, lncRNAs, and circRNAs affect several signaling pathways that are critical to tumor development. Examining these functional overlaps shows that ncRNAs may have therapeutic value in tumor angiogenesis and shed light on their biological functions. The known roles of numerous lncRNAs and circRNAs in cancer biology are only a small portion of their potential activities, even though their exact methods of action are still unknown. Several LncRNAs, circRNAs, and angiogenic miRNAs play diverse roles in tumor angiogenesis. For example, PVT1 acts as a sponge for various miRNAs that control genes further down the line. It also manages chromatin remodeling, transcriptional activation, and protein modification. In addition to angiogenesis, angio-lncRNAs and angio-circRNAss have an impact on a variety of functional characteristics. MALAT1, for example, controls EMT, invasion, migration, metastasis, and ECM remodeling. This review does not go into great detail on alternative functional pathways, instead concentrating on their roles in tumor angiogenesis. Despite encouraging developments, lncRNAs and circRNAs still have a lot of obstacles to overcome. To get around these problems and make preclinical research on anticancer therapies better, we need new technologies like genome-wide chromatin analysis and CRISPR-mediated gene editing. Methods such as quantitative RT-PCR (qRT-PCR), global run-on sequencing (GRO-seq), and RNA sequencing (RNA-seq) have improved our understanding of gene transcription and ncRNA activity. Additionally, RNA interference techniques like ASOs, MOs, LNAs, PNAs, shRNA, and siRNA, along with RNA pull-down experiments and immunoprecipitation (RIP), have simplified the study of ncRNA interactions with proteins. Despite these developments, problems, including low cellular absorption and stability, make it difficult to translate ncRNA research into practical in vivo therapeutics. These days, liposomes, lentiviruses, adenoviruses, exosomes, and nanoparticles are used to deliver miRNAs, lncRNAs, and circRNAs inside living cells.
However, there is an urgent need to harness the diverse biological properties of noncoding RNAs to devise strategic alternatives for combating pathological processes. RNA-based therapies offer significant advantages, including their specificity, low toxicity, and potential for synergistic action in targeting complex pathways. Additionally, certain ncRNAs exhibit remarkable safety and stability, whether in their naked or encapsulated forms, enhancing their applicability and value in therapy. These attributes hold promise for more efficacious treatments and personalized medicine. Further exploration into the realm of ncRNAs is essential for elucidating their mechanisms of action, identifying novel ncRNA classes, and devising innovative therapeutic approaches. Future developments in RNA-based therapies must prioritize minimizing off-target effects, optimizing immunogenicity, safety, stability, and tailoring treatments to individual patients. In essence, the key areas of focus revolve around conceiving novel delivery methods to enhance the efficiency, specificity, and clinical applicability of RNA molecules. To achieve these goals, interdisciplinary collaboration across molecular biology, cell biology, chemistry, nanotechnology, and immunology is imperative. Prospective research ought to concentrate on creating customized delivery strategies for particular tissues and cells. Despite persistent obstacles, notable advancements in ncRNA research continue to expand our knowledge of miRNAs, circRNAs, and lncRNAs, providing fresh perspectives on cancer detection and treatment.
Data availability
No datasets were generated or analysed during the current study.
Abbreviations
- GBM:
-
Glioblastoma
- mRNAs:
-
Messenger RNAs
- ncRNAs:
-
Noncoding RNAs
- sncRNAs:
-
Small noncoding RNAs
- snRNAs:
-
Including small nuclear RNAs
- tRNAs:
-
Transfer RNAs
- snoRNAs:
-
Small nucleolar RNAs
- miRNAs:
-
MicroRNAs
- CNS:
-
Central nervous system
- lncRNAs:
-
Long noncoding RNAs
- GSCs:
-
Glioma stem cells
- siRNA:
-
Small interfering RNA
- UTR:
-
3′ Untranslated regions
- MiRISC:
-
MiRNA-induced silencing complex
- pre-miRNA:
-
Precursor miRNA
- RISC:
-
RNA-induced silencing complex
- shRNA:
-
Small hairpin RNA
- MREs:
-
MiRNA response elements
- DNA:
-
Deoxyribonucleic acid
- RNA:
-
Ribonucleic acid
- XRN1:
-
Enzyme 5′-3′ exoribonuclease-1
- RER:
-
Rough endoplasmic reticulum
- PDCD4:
-
Programmed cell death protein 4
- ANP32A :
-
Acidic leucine-rich nuclear phosphoprotein 32
- PTEN:
-
Phosphatase and tensin homolog deleted on chromosome 10
- SMARCA4:
-
SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 4
- SPRY2:
-
Sprouty homolog 2
- TIMP3:
-
Tissue inhibitor of metalloproteinase 3
- MMP16:
-
Matrix metalloproteinase 16
- MMP14:
-
Matrix metalloproteinase 14
- c-Myc:
-
Cellular myelocytomatosis
- PI3K/Akt/Mtor:
-
Phosphoinositide 3 kinase/Akt/mammalian (or mechanistic) target of rapamycin
- Wnt-β-catenin:
-
Wingless/integrated β-catenin
- MAPK:
-
Mitogen-activated protein kinase
- TCGA:
-
Cancer genome atlas
- CRNDE:
-
Colorectal neoplasia differentially expressed
- MEG3:
-
Maternally expressed gene 3
- IDH:
-
Isocitrate dehydrogenase
- STAT1:
-
Signal transducer and activator of transcription 1
- RPL36:
-
Ribosomal protein L36
- TGF-β:
-
Transforming growth factor-beta
- NF-Κb:
-
Nuclear factor kappa B
- EPIC1:
-
Epigenetically induced MYC interacting lncRNA-1
- CeRNA:
-
Competitive endogenous RNA
- FGFR3:
-
Fibroblast growth factor receptor 3
- EMT:
-
Epithelial-mesenchymal transition
- NFAT5:
-
Nuclear factor of activated T cells 5
- IRES:
-
Internal ribosome entry sites
- UTR :
-
5′ Untranslated region
- EIF3:
-
Eukaryotic initiation factor 3
- ecircRNAs :
-
Exon circRNAs
- ciRNAs:
-
Circular intron RNAs and exon–intron circRNAs (EIciRNAs).
- CeRNA:
-
Competitive endogenous RNA
- ADARB2:
-
Adenosine deaminase RNA-specific B2
- NT5E :
-
Nucleotidase ecto
- circ-SHPRH:
-
SNF2 histone linker PHD RING helicase
- OS:
-
Overall survival
- circCDR1:
-
Cerebellar degeneration-related protein 1
- MOs :
-
Morpholino oligonucleotides
- PNAs:
-
Peptide nucleic acids
- LNAs:
-
Locked nucleic acids
- ASOs:
-
Antisense oligonucleotides
- LNCs:
-
Lipid nanocapsules
- PNAs:
-
Peptide nucleic acids
- PI3K:
-
Phosphatidylinositol 3-kinase
- BBB:
-
Blood-brain barrier
- ASO:
-
Antisense oligonucleotide
- 2DCS:
-
2-Dimensional combinatorial screening
- G Neo B:
-
G neomycin B
- StARTS:
-
Structure–activity relationship
References
Ahmadi A, Moqadami A, Khalaj-Kondori M, Ghiasvand S (2024) Non-coding RNAs affect breast cancer development through the notch signaling pathway: An overview. Gene Expr 23(1):44–56. https://doi.org/10.14218/GE.2023.00084
Ahmed SP, Castresana JS, Shahi MH (2022) Role of circular RNA in brain tumor development. Cells 11(14). https://doi.org/10.3390/cells11142130
Akhlaghdoust M, Bordbar S, Nikoohemmat M, Meftah E, Rahimzadegan M, Akbari S, Zali A (2024) Precision medicine in brain tumors: New approaches. https://doi.org/10.1007/16833_2024_274
Ali Syeda Z, Langden SSS, Munkhzul C, Lee M, Song J (2020) Regulatory mechanism of microrna expression in cancer. Int J Mol Sci 21(5):1723. https://doi.org/10.3390/ijms21051723
Aloizou A-M, Pateraki G, Siokas V, Mentis A-F, Liampas I, Lazopoulos G, Kovatsi L, Mitsias PD, Bogdanos DP, Paterakis K, Dardiotis E (2020) The role of MiRNA-21 in gliomas: hope for a novel therapeutic intervention? Toxicol Rep 7:1514–1530. https://doi.org/10.1016/j.toxrep.2020.11.001
Annese T, Tamma R, De Giorgis M, Ribatti D (2020) MicroRNAs biogenesis, functions and role in tumor angiogenesis. Front Oncol 10:581007. https://doi.org/10.3389/fonc.2020.581007
Badowski C, He B, Garmire L (2022) Blood-derived lncRNAs as biomarkers for cancer diagnosis: the Good, the Bad and the Beauty. NPJ Precision Oncology 6(1):40. https://doi.org/10.1038/s41698-022-00283-7
Bao Q, Yu X, Qi X (2024) Integrated analysis of single‐cell sequencing and weighted co‐expression network identifies a novel signature based on cellular senescence‐related genes to predict prognosis in glioblastoma. Environ Toxicol 39(2):643–656. https://doi.org/10.1002/tox.23921
Beňačka R, Szabóová D, Guľašová Z, Hertelyová Z, Radoňak J (2023) Non-coding RNAs in human cancer and other diseases: Overview of the diagnostic potential. Int J Mol Sci 24(22). https://doi.org/10.3390/ijms242216213
Bose D, Jayaraj G, Suryawanshi H, Agarwala P, Pore SK, Banerjee R, Maiti S (2012) The tuberculosis drug streptomycin as a potential cancer therapeutic: inhibition of MiR-21 function by directly targeting its precursor. Angew Chem 124(4):1043–1047. https://doi.org/10.1002/ange.201106455
Budi HS, Younus LA, Lafta MH, Parveen S, Mohammad HJ, Al-Qaim ZH, Jawad MA, Parra RMR, Mustafa YF, Alhachami FR, Karampoor S, Mirzaei R (2022) The role of MiR-128 in cancer development, prevention, drug resistance, and immunotherapy. Front Oncol 12:1067974. https://doi.org/10.3389/fonc.2022.1067974
Buruiană A, Florian ȘI, Florian AI, Timiș T-L, Mihu CM, Miclăuș M, Oșan S, Hrapșa I, Cataniciu RC, Farcaș M, Șușman S (2020) The roles of MiRNA in glioblastoma tumor cell communication: diplomatic and aggressive negotiations. Int J Mol Sci 21(6). https://doi.org/10.3390/ijms21061950
Byron SA, Min E, Thal TS, Hostetter G, Watanabe AT, Azorsa DO, Little TH, Tapia C, Kim S (2012) Negative regulation of NF-ΚB by the ING4 tumor suppressor in breast cancer. PLoS ONE 7(10):e46823. https://doi.org/10.1371/journal.pone.0046823
Cai H, Yanjiao Yu, Ni X, Li C, Yuanjun Hu, Wang J, Chen F, Xi S, Chen Z (2020) LncRNA LINC00998 inhibits the malignant glioma phenotype via the CBX3-mediated c-Met/Akt/MTOR axis. Cell Death Dis 11(12):1032. https://doi.org/10.1038/s41419-020-03247-6
Carrano A, Juarez JJ, Incontri D, Ibarra A, Cazares HG (2021) Sex-specific differences in glioblastoma. Cells 10(7). https://doi.org/10.3390/cells10071783
Chaudhary R (2021) Potential of long non-coding RNAs as a therapeutic target and molecular markers in glioblastoma pathogenesis. Heliyon 7(3):e06502. https://doi.org/10.1016/j.heliyon.2021.e06502
Chaudhary U, Banerjee S (2024) Decoding the non-coding: tools and databases unveiling the hidden world of “Junk” RNAs for innovative therapeutic exploration. ACS Pharmacol Transl Sci. https://doi.org/10.1021/acsptsci.3c00388
Chen GR, Sive H, Bartel DP (2017) A seed mismatch enhances argonaute2-catalyzed cleavage and partially rescues severely impaired cleavage found in fish. Mol Cell 68(6):1095-1107.e5. https://doi.org/10.1016/j.molcel.2017.11.032
Chen X, Mangala LS, Rodriguez-Aguayo C, Kong X, Lopez-Berestein G, Sood AK (2018) RNA interference-based therapy and its delivery systems. Cancer Metastasis Rev 37(1):107–124. https://doi.org/10.1007/s10555-017-9717-6
Chen B, Wang M, Huang R, Liao K, Wang T, Yang R, Zhang W, Shi Z, Ren Li, Lv Qi, Ma C, Lin Y, Qiu Y (2021a) Circular RNA CircLGMN facilitates glioblastoma progression by targeting MiR-127-3p/LGMN axis. Cancer Lett 522:225–237. https://doi.org/10.1016/j.canlet.2021.09.030
Chen B, Chen C, Zhang Y, Jianguo Xu (2021b) Recent incidence trend of elderly patients with glioblastoma in the United States, 2000–2017. BMC Cancer 21(1):54. https://doi.org/10.1186/s12885-020-07778-1
Chen B, Wang M, Huang R, Liao K, Wang T, Yang R, Zhang W, Shi Z, Ren L, Lv Q, Ma C (2021) Circular RNA circLGMN facilitates glioblastoma progression by targeting miR-127-3p/LGMN axis. Cancer Lett 522:225–237. https://doi.org/10.1016/j.canlet.2021.09.030
Chen L, Li Z, Shuaibing Hu, Deng Q, Hao P, Guo S (2022) Extracellular vesicles carry MiR-27a-3p to promote drug resistance of glioblastoma to temozolomide by targeting BTG2. Cancer Chemother Pharmacol 89(2):217–229. https://doi.org/10.1007/s00280-021-04392-1
Dai L, Liang W, Shi Z, Li X, Zhou S, Weihua Hu, Yang Z, Wang X (2023) Systematic characterization and biological functions of non-coding RNAs in glioblastoma. Cell Prolif 56(3):e13375. https://doi.org/10.1111/cpr.13375
DeOcesano-Pereira C, Machado C, Chudzinski-Tavassi A, Sogayar C (2020) Emerging roles and potential applications of non-coding RNAS in glioblastoma. Int J Mol Sci 21(7). https://doi.org/10.3390/ijms21072611
Desgraves F, Mendez Valdez J, Chandar J, Gurses E, Henderson L, Castro J, Seetheram D, Ivan E, Komotar J, Shah H (2024) Antisense oligonucleotides for rapid translation of gene therapy in glioblastoma. Cancers 16(10):1944. https://doi.org/10.3390/cancers16101944
Dhuri K, Bechtold C, Quijano E, Pham H, Gupta A, Vikram A, Bahal R (2020) Antisense oligonucleotides: an emerging area in drug discovery and development. J Clin Med 9(6). https://doi.org/10.3390/jcm9062004
Di Martino MT, Arbitrio M, Caracciolo D, Cordua A, Cuomo O, Grillone K, Riillo C, Caridà G, Scionti F, Labanca C, Romeo C, Siciliano MA, D'Apolito M, Napoli C, Montesano M, Farenza V, Uppolo V, Tafuni M, Falcone F, D'Aquino G, Calandruccio ND, Luciano F, Pensabene L, Tagliaferri P, Tassone P (2022) miR-221/222 as biomarkers and targets for therapeutic intervention on cancer and other diseases: A systematic review. Mol Ther Nucleic Acids 27:1191–1224. https://doi.org/10.1016/j.omtn.2022.02.005
Ding Yi, Wang X, Pan J, Ji M, Luo Z, Zhao P, Zhang Y, Wang G (2020) Aberrant expression of long non-coding RNAs (LncRNAs) is involved in brain glioma development. Arch Med Sci 16(1):177–188. https://doi.org/10.5114/aoms.2020.91290
Duan Y, Yue K, Ye B, Chen P, Zhang J, He Q, Wu Y, Lai Q, Li H, Wu Y, Jing C, Wang X (2023) LncRNA MALAT1 promotes growth and metastasis of head and neck squamous cell carcinoma by repressing VHL through a non-canonical function of EZH2. Cell Death Dis 14(2):149. https://doi.org/10.1038/s41419-023-05667-6
Fan X, Liu M, Fei Li, Huang Z, Yan Y (2022a) CircFOXM1 promotes the proliferation, migration, invasion, and glutaminolysis of glioblastoma by regulating the MiR-577/E2F5 axis. Bosn J Basic Med Sci 22(2):205–216. https://doi.org/10.17305/bjbms.2021.6028
Fan Y, Zhang X, Gao C, Jiang S, Haoze Wu, Liu Z, Dou T (2022b) Burden and trends of brain and central nervous system cancer from 1990 to 2019 at the global, regional, and country levels. Arch Public Health 80(1):209. https://doi.org/10.1186/s13690-022-00965-5
Feng Y-H, Tsao C-J (2016) Emerging role of microRNA-21 in cancer. Biomed Rep 5(4):395–402. https://doi.org/10.3892/br.2016.747
Foster C, Couey A, Kochanny S, Khattri A, Acharya K, Tan C, Brisson J, Leidner S, Seiwert Y (2021) Immune‐related adverse events are associated with improved response, progression‐free survival, and overall survival for patients with head and neck cancer receiving immune checkpoint inhibitors. Cancer 127(24):4565–4573. https://doi.org/10.1002/cncr.33780
Gao X, Xia X, Li F, Zhang M, Zhou H, Xujia Wu, Zhong J, Zhao Z, Zhao K, Liu D, Xiao F, Qiang Xu, Jiang T, Li Bo, Cheng S-Y, Zhang Nu (2021) Circular RNA-encoded oncogenic E-cadherin variant promotes glioblastoma tumorigenicity through activation of EGFR–STAT3 signalling. Nat Cell Biol 23(3):278–291. https://doi.org/10.1038/s41556-021-00639-4
Gao P, Wang H, Li H, Shu L, Han Z, Li S, Cheng H, Dai X (2023) MiR-21-5p inhibits the proliferation, migration, and invasion of glioma by targeting S100A10. J Cancer 14(10):1781–1793. https://doi.org/10.7150/jca.84030
Garbo S, Maione R, Tripodi M, Battistelli C (2022) Next RNA therapeutics: The mine of non-coding. Int J Mol Sci 23(13). https://doi.org/10.3390/ijms23137471
Gheidari F, Arefian E, Adegani FJ, Kalhori MR, Seyedjafari E, Kabiri M, Teimoori-Toolabi L, Soleimani M (2021) MiR-424 induces apoptosis in glioblastoma cells and targets AKT1 and RAF1 oncogenes from the ERBB signaling pathway. Eur J Pharmacol 906:174273. https://doi.org/10.1016/j.ejphar.2021.174273
Goel B, Tiwari A, Pandey K, Singh A, Kumar S, Sinha A, Jain S, Khattri A (2022) Therapeutic approaches for the treatment of head and neck squamous cell carcinoma–An update on clinical trials. Transl Oncol 21:101426. https://doi.org/10.1016/j.tranon.2022.101426
Grillone K, Riillo C, Scionti F, Rocca R, Tradigo G, Guzzi PH, Alcaro S, Martino MTD, Tagliaferri P, Tassone P (2020) Non-coding RNAs in cancer: platforms and strategies for investigating the genomic ‘dark matter.’ J Exp Clin Cancer Res 39(1):117. https://doi.org/10.1186/s13046-020-01622-x
Griveau A, Bejaud J, Anthiya S, Avril S, Autret D, Garcion E (2013) Silencing of miR-21 by locked nucleic acid–lipid nanocapsule complexes sensitize human glioblastoma cells to radiation-induced cell death. Int J Pharm 454(2):765–774. https://doi.org/10.1016/j.ijpharm.2013.05.049
Grochans S, Cybulska A, Simińska D, Korbecki J, Kojder K, Chlubek D, Baranowska-Bosiacka I (2022) Epidemiology of glioblastoma multiforme-literature review. Cancers 14(10). https://doi.org/10.3390/cancers14102412
Gu K, Mok L, Chong MMW (2018) Regulating gene expression in animals through RNA endonucleolytic cleavage. Heliyon 4(11):e00908. https://doi.org/10.1016/j.heliyon.2018.e00908
Gu J, Zhang Bo, An R, Qian W, Han L, Duan W, Wang Z, Ma Q (2022) Molecular interactions of the long noncoding RNA NEAT1 in cancer. Cancers 14(16):4009. https://doi.org/10.3390/cancers14164009
Gu P, Ding Y, Zheng G, Xu P, Xia X (2023) Extracranial metastasis of glioblastoma: A case report and literature review. Int J Surg Case Rep 111:108895. https://doi.org/10.1016/j.ijscr.2023.108895
Gupta A, Singh MS, Singh B (2024) Deciphering the functional role of clinical mutations in ABCB1, ABCC1, and ABCG2 ABC transporters in endometrial cancer. Front Pharmacol 15. https://doi.org/10.3389/fphar.2024.1380371
Han Z, Chen H, Guo Z, Shen J, Luo W, Xie F, Wan Y, Wang S, Li J, He J (2022) Circular RNAs and Their Role in Exosomes. Front Oncol 12:848341. https://doi.org/10.3389/fonc.2022.848341
Hanif F, Muzaffar K, Perveen K, Malhi SM, Simjee SU (2017) Glioblastoma multiforme: a review of its epidemiology and pathogenesis through clinical presentation and treatment. Asian Pac J Cancer Prev : APJCP 18(1):3–9. https://doi.org/10.22034/APJCP.2017.18.1.3
Ho K-H, Shih C-M, Liu A-J, Chen K-C (2022) Hypoxia-inducible LncRNA MIR210HG interacting with OCT1 is involved in glioblastoma multiforme malignancy. Cancer Sci 113(2):540–552. https://doi.org/10.1111/cas.15240
Hua Y, Qin Z, Gao L, Zhou M, Xue Y, Li Y, Xie J (2024) Protein nanoparticles as drug delivery systems for cancer theranostics. J Control Release 371:429–444. https://doi.org/10.1016/j.jconrel.2024.06.004
Huang A, Zheng H, Zhiye Wu, Chen M, Huang Y (2020) Circular RNA-protein interactions: functions, mechanisms, and identification. Theranostics 10(8):3503–3517. https://doi.org/10.7150/thno.42174
Huang Y, Chiao M T, Hsiao T, Zhan X, Chen Y, Lee H, Liu Y, Liao H, Cheng Y, Yen M, Lai M, Chen P, Shen C, Yang Y (2024) Genetic mutation patterns among glioblastoma patients in the Taiwanese population – insights from a single institution retrospective study. Cancer Gene Ther. https://doi.org/10.1038/s41417-024-00746-y
Huang Z-H, Yu-ping Du, Wen J-T, Bing-feng Lu, Zhao Y (2022) SnoRNAs: functions and mechanisms in biological processes, and roles in tumor pathophysiology. Cell Death Disc 8(1):259. https://doi.org/10.1038/s41420-022-01056-8
Ismail NH, Mussa A, Al-Khreisat M, Mohamed Yusoff S, Husin A, Al-Jamal HAN, Johan M, Islam M (2023) Dysregulation of non-coding RNAs: Roles of miRNAs and lncRNAs in the pathogenesis of multiple myeloma. Non-Coding RNA 9(6). https://doi.org/10.3390/ncrna9060068
Iwakawa H-O, Tomari Y (2022) Life of RISC: formation, action, and degradation of RNA-induced silencing complex. Mol Cell 82(1):30–43. https://doi.org/10.1016/j.molcel.2021.11.026
Kciuk M, Yahya E, Mohamed M, Abdulsamad M, Allaq A, Gielecińska A, Kontek R (2023) Insights into the Role of LncRNAs and miRNAs in glioma progression and their potential as novel therapeutic targets. Cancers 15(13):3298. https://doi.org/10.3390/cancers15133298
Khan FA, Nsengimana B, Khan NH, Song Z, Ngowi EE, Wang Y, Zhang W, Ji S (2022) Chimeric peptides/proteins encoded by CircRNA: an update on mechanisms and functions in human cancers. Front Oncol 12:781270. https://doi.org/10.3389/fonc.2022.781270
Khattri A, Zuo Z, Brägelmann J, Keck MK, El Dinali M, Brown CD, Stricker T, Munagala A, Cohen E, Lingen W, White P, Vokes E, Seiwert Y (2015) Rare occurrence of EGFRvIII deletion in head and neck squamous cell carcinoma. Oral Oncol 51(1):53–58. https://doi.org/10.1016/j.oraloncology.2014.08.014
Kim J, Park MW, Park YJ, Ahn JW, Sim JM, Kim S, Heo J, Jeong JH, Lee M, Lim J, Moon J-S (2021) MiR-542–3p contributes to the HK2-mediated high glycolytic phenotype in human glioma cells. Genes 12(5). https://doi.org/10.3390/genes12050633
Koffler-Brill T, Noy Y, Avraham KB (2023) The long and short: Non-coding RNAs in the mammalian inner ear. Hear Res 428:108666. https://doi.org/10.1016/j.heares.2022.108666
Komatsu S, Kitai H, Suzuki HI (2023) Network regulation of microRNA biogenesis and target interaction. Cells 12(2). https://doi.org/10.3390/cells12020306
Kovalchuk I (2021) Non-coding RNAs in genome integrity. Genome Stability 453–475. https://doi.org/10.1016/B978-0-323-85679-9.00024-6
Krysler AR, Cromwell CR, Tommy Tu, Jovel J, Hubbard BP (2022) Guide RNAs containing universal bases enable Cas9/Cas12a recognition of polymorphic sequences. Nat Commun 13(1):1617. https://doi.org/10.1038/s41467-022-29202-x
Kwak S, Park S-H, Kim S-H, Sung G-J, Song J-H, Jeong J-H, Kim H, Ha CH, Kim SW, Choi K-C (2022) MiR-3189-targeted GLUT3 repression by HDAC2 knockdown inhibits glioblastoma tumorigenesis through regulating glucose metabolism and proliferation. J Exp Clin Cancer Res 41(1):87. https://doi.org/10.1186/s13046-022-02305-5
Lee SM, Kaye KM, Slack FJ (2021) Cellular microRNA-127–3p suppresses oncogenic herpesvirus-induced transformation and tumorigenesis via down-regulation of SKP2. Proc Natl Acad Sci USA 118(45). https://doi.org/10.1073/pnas.2105428118
Liang J, Liu C, Dezhi Xu, Xie K, Li A (2022) LncRNA NEAT1 facilitates glioma progression via stabilizing PGK1. J Transl Med 20(1):80. https://doi.org/10.1186/s12967-022-03273-2
Liao J, Chen B, Zhu Z, Chengcheng Du, Gao S, Zhao G, Zhao P, Wang Y, Wang A, Schwartz Z, Song L, Hong J, Wagstaff W, Haydon RC, Luu HH, Fan J, Reid RR, He T-C, Shi L, Ning Hu, Huang W (2023) Long noncoding RNA (LncRNA) H19: an essential developmental regulator with expanding roles in cancer, stem cell differentiation, and metabolic diseases. Genes Dis 10(4):1351–1366. https://doi.org/10.1016/j.gendis.2023.02.008
Lin P, He L, Tian N, Qi X (2023) The evaluation of six genes combined value in glioma diagnosis and prognosis. J Cancer Res Clin Oncol 149(13):12413–12433. https://doi.org/10.1007/s00432-023-05082-6
Ling C, Wang X, Zhu J, Tang H, Wenwen Du, Zeng Y, Sun L, Huang J-A, Liu Z (2019) MicroRNA-4286 promotes cell proliferation, migration, and invasion via PTEN regulation of the PI3K/Akt pathway in non-small cell lung cancer. Cancer Med 8(7):3520–3531. https://doi.org/10.1002/cam4.2220
Liu Y, Li S, Chen Y, Kimberlin AN, Cahoon EB, Bin Yu (2016) SnRNA 3′ end processing by a CPSF73-containing complex essential for development in Arabidopsis. PLoS Biol 14(10):e1002571. https://doi.org/10.1371/journal.pbio.1002571
Liu W, Zhang B, Xu N, Wang MJ, Liu Q (2017) MiR-326 regulates EMT and metastasis of endometrial cancer through targeting TWIST1. Eur Rev Med Pharmacol Sci 21(17):3787–3793
Liu H, Lei C, He Q, Pan Z, Xiao D, Tao Y (2018) Nuclear functions of mammalian microRNAs in gene regulation, immunity and cancer. Mol Cancer 17(1):64. https://doi.org/10.1186/s12943-018-0765-5
Liu ZZ, Tian YF, Wu H, Ouyang SY, Kuang WL (2020) LncRNA H19 promotes glioma angiogenesis through MiR-138/HIF-1α/VEGF axis. Neoplasma 67(01):111–118. https://doi.org/10.4149/neo_2019_190121N61
Liu C, Zhu B, Zhong M, Bao J (2022a) MiRNA-448 regulates the development of glioblastoma (GBM) by regulating Rho-associated protein kinase 1. Comput Math Methods Med 2022:2502010. https://doi.org/10.1155/2022/2502010
Liu K, Deng Y, Yang Y, Wang H, Zhou P (2022b) MicorRNA-195 links long non-coding RNA SEMA3B antisense RNA 1 (head to head) and cyclin D1 to regulate the proliferation of glioblastoma cells. Bioengineered 13(4):8798–8805. https://doi.org/10.1080/21655979.2022.2052646
Liu X, Zhang Yu, Zhou S, Dain L, Mei L, Zhu G (2022c) Circular RNA: an emerging frontier in RNA therapeutic targets, RNA therapeutics, and MRNA vaccines. J Control Release 348:84–94. https://doi.org/10.1016/j.jconrel.2022.05.043
Loubalova Z, Konstantinidou P, Haase AD (2023) Themes and variations on PiRNA-guided transposon control. Mob DNA 14(1):10. https://doi.org/10.1186/s13100-023-00298-2
Lu J, Kornmann M, Traub B (2023a) Role of epithelial to mesenchymal transition in colorectal cancer. Int J Mol Sci 24(19):14815. https://doi.org/10.3390/ijms241914815
Lu Q, Xie Y, Qi X, Yang S (2023b) TREM1 as a novel prognostic biomarker and tumor immune microenvironment evaluator in glioma. Medicine 102(48):e36410. https://doi.org/10.1097/MD.0000000000036410
Lu Y, Tian M, Liu J, Wang K (2021) LINC00511 facilitates temozolomide resistance of glioblastoma cells via sponging MiR‐126‐5p and activating Wnt/Β‐catenin signaling. J Biochem Mol Toxicol 35(9). https://doi.org/10.1002/jbt.22848
Lui A, Do T, Alzayat O, Yu N, Phyu S, Santuya H, Liang B, Kailash V, Liu D, Inslicht S, Shahlaie K, Liu D (2024) Tumor suppressor MicroRNAs in clinical and preclinical trials for neurological disorders. Pharmaceuticals 17(4):426. https://doi.org/10.3390/ph17040426
Lulli V, Buccarelli M, Ilari R, Castellani G, De Dominicis C, Di Giamberardino A, Alessandris QGD, Giannetti S, Martini M, Stumpo V, Boe A, De Luca G, Biffoni M, Marziali G, Pallini R, Ricci-Vitiani L (2020) “Mir-370–3p impairs glioblastoma stem-like cell malignancy regulating a complex interplay between HMGA2/HIF1A and the oncogenic long non-coding RNA (LncRNA) NEAT1. Int J Mol Sci 21(10). https://doi.org/10.3390/ijms21103610.
Luo Y, Na Z, Slavoff SA (2018) P-bodies: composition, properties, and functions. Biochemistry 57(17):2424–2431. https://doi.org/10.1021/acs.biochem.7b01162
Lv T, Jin Y, Miao Y, Tianqi Xu, Jia F, Feng H, Zhang X (2022) LncRNA PVT1 promotes tumorigenesis of glioblastoma by recruiting COPS5 to deubiquitinate and stabilize TRIM24. Mol Ther - Nucleic Acids 27:109–121. https://doi.org/10.1016/j.omtn.2021.11.012
Ma C, Wang H, Zong G, He J, Wang Y, Yang F, Yang Z, Bian E, Zhao B (2021) EGR1 modulated LncRNA HNF1A-AS1 drives glioblastoma progression via MiR-22-3p/ENO1 axis. Cell Death Disc 7(1):350. https://doi.org/10.1038/s41420-021-00734-3
Ma Y, Wang T, Zhang X, Wang P, Long F (2024) The role of circular RNAs in regulating resistance to cancer immunotherapy: mechanisms and implications. Cell Death Dis 15(5):312. https://doi.org/10.1038/s41419-024-06698-3
Mafi A, Rahmati A, Aghdam ZB, Salami R, Salami M, Vakili O, Aghadavod E (2022) Recent insights into the microRNA-dependent modulation of gliomas from pathogenesis to diagnosis and treatment. Cell Mol Biol Lett 27(1):65. https://doi.org/10.1186/s11658-022-00354-4
Manzoni C, Kia DA, Vandrovcova J, Hardy J, Wood NW, Lewis PA, Ferrari R (2018) Genome, transcriptome and proteome: the rise of omics data and their integration in biomedical sciences. Brief Bioinform 19(2):286–302. https://doi.org/10.1093/bib/bbw114
Martinez I, Hayes KE, Barr JA, Harold AD, Xie M, Bukhari SIA, Vasudevan S, Steitz JA, DiMaio D (2017) An exportin-1–dependent microRNA biogenesis pathway during human cell quiescence. Proceedings of the National Academy of Sciences 114(25). https://doi.org/10.1073/pnas.1618732114
Matsui M, Chu Y, Zhang H, Gagnon KT, Shaikh S, Kuchimanchi S, Manoharan M, Corey DR, Janowski BA (2013) Promoter RNA links transcriptional regulation of inflammatory pathway genes. Nucleic Acids Res 41(22):10086–10109. https://doi.org/10.1093/nar/gkt777
Mattei V, Santilli F, Martellucci S, Delle Monache S, Fabrizi J, Colapietro A, Angelucci A, Festuccia C (2021) The importance of tumor stem cells in glioblastoma resistance to therapy. Int J Mol Sci 22(8). https://doi.org/10.3390/ijms22083863
Mattick JS, Amaral PP, Carninci P, Carpenter S, Chang HY, Chen L-L, Chen R, Dean C, Dinger ME, Fitzgerald KA, Gingeras TR, Guttman M, Hirose T, Huarte M, Johnson R, Kanduri C, Kapranov P, Lawrence JB, Lee JT, Mendell JT, Mercer TR, Moore KJ, Nakagawa S, Rinn JL, Spector DL, Ulitsky I, Wan Y, Wilusz JE, Mian Wu (2023) Long non-coding RNAs: definitions, functions, challenges and recommendations. Nat Rev Mol Cell Biol 24(6):430–447. https://doi.org/10.1038/s41580-022-00566-8
Mazurek M, Grochowski C, Litak J, Osuchowska I, Maciejewski R, Kamieniak P (2020) Recent trends of microRNA significance in pediatric population glioblastoma and current knowledge of micro RNA function in glioblastoma multiforme. Int J Mol Sci 21(9):3046. https://doi.org/10.3390/ijms21093046
McGeary SE, Lin KS, Shi CY, Pham TM, Bisaria N, Kelley GM, Bartel DP (2019) The biochemical basis of microRNA targeting efficacy. Science (New York, N.Y.) 366(6472). https://doi.org/10.1126/science.aav1741
Medley JC, Panzade G, Zinovyeva AY (2021) MicroRNA strand selection: unwinding the rules. Wiley Interdiscip Rev RNA 12(3):e1627. https://doi.org/10.1002/wrna.1627
Møller HG, Rasmussen AP, Andersen HH, Johnsen KB, Henriksen M, Duroux M (2013) A systematic review of microRNA in glioblastoma multiforme: micro-modulators in the mesenchymal mode of migration and invasion. Mol Neurobiol 47(1):131–144. https://doi.org/10.1007/s12035-012-8349-7
Momtazmanesh S, Rezaei N (2021) Long non-coding RNAs in diagnosis, treatment, prognosis, and progression of glioma: a state-of-the-art review. Front Oncol 11:712786. https://doi.org/10.3389/fonc.2021.712786
Mulcahy EQX, Zhang Y, Colόn RR, Cain SR, Gibert MK, Dube CJ, Hafner M, Abounader R (2022) MicroRNA 3928 suppresses glioblastoma through downregulation of several oncogenes and upregulation of P53. Int J Mol Sci 23(7). https://doi.org/10.3390/ijms23073930
Nabariya DK, Heinz A, Derksen S, Krauß S (2022) Intracellular and intercellular transport of RNA organelles in CXG repeat disorders: the strength of weak ties. Front Mol Biosci 9. https://doi.org/10.3389/fmolb.2022.1000932
Nappi F (2024) Non-coding RNA-targeted therapy: A state-of-the-art review. Int J Mol Sci 25(7). https://doi.org/10.3390/ijms25073630
O’Brien J, Hayder H, Zayed Y, Peng C (2018a) Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol 9:402. https://doi.org/10.3389/fendo.2018.00402
O’Brien J, Hayder H, Zayed Y, Peng C (2018b) Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol 9. https://doi.org/10.3389/fendo.2018.00402
Park J, Cho M, Cho J, Kim EE, Song EJ (2022) MicroRNA-101–3p suppresses cancer cell growth by inhibiting the USP47-induced deubiquitination of RPL11. Cancers 14(4). https://doi.org/10.3390/cancers14040964
Patop IL, Wüst S, Kadener S (2019) Past, present, and future of CircRNAs. EMBO J 38(16):e100836. https://doi.org/10.15252/embj.2018100836
Peng W, Qian Y, Qi X (2023) Efficacy of a novel glioma therapy based on ferroptosis induced by layered double hydroxide loaded with simvastatin. Environ Res 238:117112. https://doi.org/10.1016/j.envres.2023.117112
Peng W-X, Huang J-G, Yang L, Gong A-H, Mo Y-Y (2017) Linc-RoR promotes MAPK/ERK signaling and confers estrogen-independent growth of breast cancer. Mol Cancer 16(1):161. https://doi.org/10.1186/s12943-017-0727-3
Peng X, Zhang Y, Gao J, Cai C (2020) MiR-1258 promotes the apoptosis of cervical cancer cells by regulating the E2F1/P53 signaling pathway. Exp Mol Pathol 114:104368. https://doi.org/10.1016/j.yexmp.2020.104368
Pong SK, Gullerova M (2018) Noncanonical functions of micro RNA pathway enzymes–Drosha, DGCR 8, Dicer and Ago proteins. FEBS letters 592(17):2973–2986. https://doi.org/10.1002/1873-3468.13196
Pradeep P, Malik S, Slack J, Bahal R (2023) Unlocking the potential of chemically modified peptide nucleic acids for RNA-based therapeutics. RNA (New York, N.Y.) 29(4):434–445. https://doi.org/10.1261/rna.079498.122
Qin M, Zhang C, Li Y (2023) Circular RNAs in gynecologic cancers: mechanisms and implications for chemotherapy resistance. Front Pharmacol 14. https://doi.org/10.3389/fphar.2023.1194719
Rhim J, Baek W, Seo Y, Kim JH (2022) From molecular mechanisms to therapeutics: understanding microRNA-21 in cancer. Cells 11(18):2791. https://doi.org/10.3390/cells11182791
Rios S, Oyervides S, Uribe D, Reyes A, Fanniel V, Vazquez J, Keniry M (2024) Emerging therapies for glioblastoma. Cancers 16(8):1485. https://doi.org/10.3390/cancers16081485
Sadakierska-Chudy A (2020) MicroRNAs: diverse mechanisms of action and their potential applications as cancer epi-therapeutics. Biomolecules 10(9):1285. https://doi.org/10.3390/biom10091285
Salami R, Salami M, Mafi A, Vakili O, Asemi Z (2022) Circular RNAs and glioblastoma multiforme: focus on molecular mechanisms. Cell Commun Signaling 20(1):13. https://doi.org/10.1186/s12964-021-00809-9
Saloura V, Fatima A, Zewde M, Kiyotani K, Brisson R, Park J, Ikeda Y, Vougiouklakis T, Bao R, Khattri A, Seiwert T, Cipriani N, Lingen M, Vokes E, Nakamura Y (2017) Characterization of the T-Cell receptor repertoire and immune microenvironment in patients with locoregionally advanced squamous cell carcinoma of the head and neck. Clin Cancer Res 23(16):4897–4907. https://doi.org/10.1158/1078-0432.CCR-17-0103
Saloura V, Izumchenko E, Zuo Z, Bao R, Korzinkin M, Ozerov I, Zhavoronkov A, Sidransky D, Bedi A, Hoque O, Koeppen H, Keck K, Khattri A, London N, Kotlov N, Fatima A, Vougiouklakis T, Nakamura Y, Lingen M, … Seiwert Y (2019) Immune profiles in primary squamous cell carcinoma of the head and neck. Oral Oncol 96:77–88. https://doi.org/10.1016/j.oraloncology.2019.06.032
Salphati L, Alicke B, Heffron TP, Shahidi-Latham S, Nishimura M, Cao T, Carano RA, Cheong J, Greve J, Koeppen H, Lau S, Lee LB, Nannini-Pepe M, Pang J, Plise EG, Quiason C, Rangell L, Zhang X, Gould SE, Phillips HS, Olivero AG (2016) Brain distribution and efficacy of the brain penetrant PI3K inhibitor GDC-0084 in orthotopic mouse models of human glioblastoma. Drug Metab Dispos 44(12):1881–1889. https://doi.org/10.1124/dmd.116.071423
Schäfer A, Evers L, Meier L, Schlomann U, Bopp MHA, Dreizner G-L, Lassmann O, Bacha AB, Benescu A-C, Pojskic M, Preußer C, von Strandmann EP, Carl B, Nimsky C, Bartsch JW (2022) The metalloprotease-disintegrin ADAM8 alters the tumor suppressor MiR-181a-5p expression profile in glioblastoma thereby contributing to its aggressiveness. Front Oncol 12. https://doi.org/10.3389/fonc.2022.826273
Shahzad U, Krumholtz S, Rutka JT, Das S (2021) Noncoding RNAs in glioblastoma: emerging biological concepts and potential therapeutic implications. Cancers 13(7). https://doi.org/10.3390/cancers13071555
Shi P, Jie Xu, Cui H (2023) The recent research progress of NF-ΚB signaling on the proliferation, migration, invasion, immune escape, and drug resistance of glioblastoma. Int J Mol Sci 24(12):10337. https://doi.org/10.3390/ijms241210337
Shivakumar K, Mahendran G, Brown J (2024) Locked nucleic acid oligonucleotides facilitate RNA LNA-RNA triple-helix formation and reduce malat1 levels. Int J Mol Sci 25(3):1630. https://doi.org/10.3390/ijms25031630
Singh CP (2020) Role of microRNAs in insect-baculovirus interactions. Insect Biochem Mol Biol 127:103459. https://doi.org/10.1016/j.ibmb.2020.103459
Singh MS, Tammam N, Shetab Boushehri A, Lamprecht A (2017) MDR in cancer: Addressing the underlying cellular alterations with the use of nanocarriers. Pharmacol Res 126:2–30. https://doi.org/10.1016/j.phrs.2017.07.023
Širvinskas D, Steponaitis G, Stakaitis R, Tamašauskas A, Vaitkienė P, Skiriutė D (2023) Antisense lncRNA CHROMR is linked to glioma patient survival. Front Mol Biosci 10. https://doi.org/10.3389/fmolb.2023.1101953
Song Z, Xue Z, Wang Y, Imran M, Assiri M, Fahad S (2024) Insights into the roles of non-coding RNAs and angiogenesis in glioblastoma: An overview of current research and future perspectives. Biochim Biophys Acta Gen Subj 1868(4):130567. https://doi.org/10.1016/j.bbagen.2024.130567
de Souza A, Davis W, Zhang Y, Khattri A, Seiwert Y, Aktolga S, Wong J, Kozloff F, Nattam S, Lingen M W, Kunnavakkam R, Stenson M, Blair A, Bozeman J, Dancey E, Vokes E, Cohen W (2012) A phase II study of lapatinib in recurrent/metastatic squamous cell carcinoma of the head and neck. Clin Cancer Res 18(8):2336–2343. https://doi.org/10.1158/1078-0432.CCR-11-2825
Statello L, Guo C-J, Chen L-L, Huarte M (2021) Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol 22(2):96–118. https://doi.org/10.1038/s41580-020-00315-9
Sun G, Ye P, Murai K, Lang M-F, Li S, Zhang H, Li W, Chelsea Fu, Yin J, Wang A, Ma X, Shi Y (2011) MiR-137 forms a regulatory loop with nuclear receptor TLX and LSD1 in neural stem cells. Nat Commun 2:529. https://doi.org/10.1038/ncomms1532
Svoboda P (2020) Introduction to RNAi and MiRNA Pathways. Karolinum press Prague, pp 5–5. https://doi.org/10.14712/9788024643724.1
Tabnak P, Bashkandi AH, Ebrahimnezhad M, Soleimani M (2023) Forkhead box transcription factors (FOXOs and FOXM1) in glioma: from molecular mechanisms to therapeutics. Cancer Cell Int 23(1):238. https://doi.org/10.1186/s12935-023-03090-7
Tang F, Wang H, Chen E, Bian E, Yadi Xu, Ji X, Yang Z, Hua X, Zhang Y, Zhao B (2019) LncRNA-ATB promotes TGF-β-induced glioma cells invasion through NF-κB and P38/MAPK pathway. J Cell Physiol 234(12):23302–23314. https://doi.org/10.1002/jcp.28898
Tang J, Zhang J, Lu Y, He J, Wang H, Liu B, Tu C, Li Z (2023) Novel insights into the multifaceted roles of m6A-modified LncRNAs in cancers: biological functions and therapeutic applications. Biomarker Research, 11(1):42. https://doi.org/10.1186/s40364-023-00484-7
Tang X, Ren H, Guo M, Qian J, Yang Ye, Chunyan Gu (2021) Review on circular RNAs and new insights into their roles in cancer. Comput Struct Biotechnol J 19:910–928. https://doi.org/10.1016/j.csbj.2021.01.018
Tang C, He X, Jia L, Zhang X (2024) Circular RNAs in glioma: molecular functions and pathological implications. Non-Coding RNA Res 9(1):105–115. https://doi.org/10.1016/j.ncrna.2023.10.007
Teplyuk NM, Uhlmann EJ, Gabriely G, Volfovsky N, Wang Y, Teng J, Karmali P, Marcusson E, Peter M, Mohan A, Kraytsberg Y (2016) Therapeutic potential of targeting micro RNA‐10b in established intracranial glioblastoma: first steps toward the clinic. EMBO Mol Med 8(3):268–287. https://doi.org/10.15252/emmm.201505495
Tepper S, Zuo Z, Khattri A, Heß J, Seiwert Y (2016) Growth factor expression mediates resistance to EGFR inhibitors in head and neck squamous cell carcinomas. Oral Oncol 56:62–70. https://doi.org/10.1016/j.oraloncology.2016.03.008
Thakur K, Singh MS, Feldstein-Davydova S, Hannes V, Hershkovitz D, Tsuriel S (2021) Extracellular vesicle-derived DNA vs. CfDNA as a biomarker for the detection of colon cancer. Genes 12(8):1171. https://doi.org/10.3390/genes12081171
Tian Z, Liang G, Cui K, Liang Y, Wang Q, Lv S, Cheng X, Zhang L (2021) Insight into the prospects for RNAi therapy of cancer. Front Pharmacol 12. https://doi.org/10.3389/fphar.2021.644718
Urits I, Swanson D, Swett M, Patel A, Berardino K, Amgalan A, Berger A A, Kassem H, Kaye AD, Viswanath O (2020) A review of patisiran (ONPATTRO®) for the treatment of polyneuropathy in people with hereditary transthyretin amyloidosis. Neurol Ther 9(2):301–315. https://doi.org/10.1007/s40120-020-00208-1
Veys C, Jammes M, Rédini F, Poulain L, Denoyelle C, Legendre F, Galera P (2022) Tumor suppressive role of MiR-342–5p and MiR-491–5p in human osteosarcoma cells. Pharmaceuticals (Basel, Switzerland) 15(3). https://doi.org/10.3390/ph15030362
Wang Z, Yang J, Gang Xu, Wang W, Liu C, Yang H, Zhibin Yu, Lei Q, Xiao L, Xiong J, Zeng L, Xiang J, Ma J, Li G, Minghua Wu (2015) Targeting MiR-381-NEFL axis sensitizes glioblastoma cells to temozolomide by regulating stemness factors and multidrug resistance factors. Oncotarget 6(5):3147–3164. https://doi.org/10.18632/oncotarget.3061
Wang X, Xiaojun Yu, Haoran Xu, Wei K, Wang S, Wang Y, Han J (2022) Serum-derived extracellular vesicles facilitate temozolomide resistance in glioblastoma through a HOTAIR-dependent mechanism. Cell Death Dis 13(4):344. https://doi.org/10.1038/s41419-022-04699-8
Wang L, Li H (2020) MiR-770-5p facilitates podocyte apoptosis and inflammation in diabetic nephropathy by targeting TIMP3. Biosci Rep 40(4). https://doi.org/10.1042/BSR20193653
Wang Q, Shao X, Zhang Y, Zhu M, Wang F, Mu J, Li J, Yao H, Chen K (2023) Role of tumor microenvironment in cancer progression and therapeutic strategy. Cancer Med 12(10):11149–11165. https://doi.org/10.1002/cam4.5698
Wu S, Huang S, Ding J, Zhao Y, Liang L, Liu T, Zhan R, He X (2010) Multiple microRNAs modulate P21Cip1/Waf1 expression by directly targeting its 3′ untranslated region. Oncogene 29(15):2302–2308. https://doi.org/10.1038/onc.2010.34
Wu Y, Qian Y, Peng W, Qi X (2023) Functionalized nanoparticles crossing the brain–blood barrier to target glioma cells. PeerJ 11:e15571. https://doi.org/10.7717/peerj.15571
Xia B, Liu Y, Wang J, Lu Q, Lv X, Deng K, Yang J (2023) Emerging role of exosome-shuttled noncoding RNAs in gastrointestinal cancers: From intercellular crosstalk to clinical utility. Pharmacol Res 195:106880. https://doi.org/10.1016/j.phrs.2023.106880
Xiao J, Joseph S, Xia M, Teng F, Chen X, Huang R, Zhai L, Deng W (2022) Circular RNAs acting as MiRNAs’ sponges and their roles in stem cells. J Clin Med 11(10). https://doi.org/10.3390/jcm11102909
Xie S-C, Zhang J-Q, Jiang X-L, Hua Y-Y, Xie S-W, Qin Y-A, Yang Y-J (2020) LncRNA CRNDE facilitates epigenetic suppression of CELF2 and LATS2 to promote proliferation, migration and chemoresistance in hepatocellular carcinoma. Cell Death Dis 11(8):676. https://doi.org/10.1038/s41419-020-02853-8
Xue C, Gu X, Bao Z, Su Y, Lu J, Li L (2022) The mechanism underlying the ncRNA dysregulation pattern in hepatocellular carcinoma and its tumor microenvironment. Front Immunol 13. https://doi.org/10.3389/fimmu.2022.847728
Yadav A, Mathan J, Dubey A, Singh A (2024) The emerging role of non-coding RNAs (ncRNAs) in plant growth, development, and stress response signaling. Non-Coding RNA 10(1):13. https://doi.org/10.3390/ncrna10010013
Yang T, Li Y, Zhao F, Zhou L, Jia R (2021) Circular RNA Foxo3: a promising cancer-associated biomarker. Front Genet 12:652995. https://doi.org/10.3389/fgene.2021.652995
Yang Y, Gao X, Zhang M, Yan S, Sun C, Xiao F, Huang N, Yang X, Zhao K, Zhou H, Huang S, Xie B, Zhang N (2018). Novel role of FBXW7 circular RNA in repressing glioma tumorigenesis. JNCI: J Natl Cancer Inst 110(3):304–15. https://doi.org/10.1093/jnci/djx166
Yeh M, Wang Y-Y, Yoo JY, Christina Oh, Otani Y, Kang JM, Park ES, Kim E, Chung S, Jeon Y-J, Calin GA, Kaur B, Zhao Z, Lee TJ (2021) MicroRNA-138 suppresses glioblastoma proliferation through downregulation of CD44. Sci Rep 11(1):9219. https://doi.org/10.1038/s41598-021-88615-8
Yin J, Shi Z, Wei W, Lu C, Wei Y, Yan W, Li R, Zhang J, You Y, Wang X (2020) MiR-181b suppress glioblastoma multiforme growth through inhibition of SP1-mediated glucose metabolism. Cancer Cell Int 20:69. https://doi.org/10.1186/s12935-020-1149-7
Yoo J-Y, Yeh M, Wang Y-Y, Oh C, Zhao Z-M, Kaur B, Lee T-J (2021) MicroRNA-138 increases chemo-sensitivity of glioblastoma through downregulation of survivin. Biomedicines 9(7). https://doi.org/10.3390/biomedicines9070780
Yu B, Pang J, You J (2022) Effects and mechanism of MiR-133a on invasion and migration of lung cancer cells. Am J Transl Res 14(2):728–739
Zepeda-Enríquez P, Silva-Cázares B, López-Camarillo C (2023) Novel insights into circular RNAs in metastasis in breast cancer: An update. Non-Coding RNA 9(5):55. https://doi.org/10.3390/ncrna9050055
Zhang Y, Liang W, Zhang P, Chen J, Qian H, Zhang X, Xu W (2017) Circular RNAs: emerging cancer biomarkers and targets. J Exp Clin Cancer Res 36(1):152. https://doi.org/10.1186/s13046-017-0624-z
Zhang P, Wu W, Chen Q, Chen M (2019a) Non-coding RNAs and their integrated networks. J Integr Bioinforma 16(3). https://doi.org/10.1515/jib-2019-0027
Zhang S, Liao K, Miao Z, Wang Q, Miao Y, Guo Z, Qiu Y, Chen B, Ren Li, Wei Z, Lin Y, Xiaojie Lu, Qiu Y (2019b) CircFOXO3 promotes glioblastoma progression by acting as a competing endogenous RNA for NFAT5. Neuro Oncol 21(10):1284–1296. https://doi.org/10.1093/neuonc/noz128
Zhang P, Long Q, Zeng S, Wen M, Qing Lu (2020a) FOXC1-induced LINC01123 acts as a mediator in triple negative breast cancer. Cancer Cell Int 20:199. https://doi.org/10.1186/s12935-020-01258-z
Zhang X, Niu W, Maolin Mu, Shanshan Hu, Niu C (2020b) Long non-coding RNA LPP-AS2 promotes glioma tumorigenesis via MiR-7-5p/EGFR/PI3K/AKT/c-MYC feedback loop. J Exp Clin Cancer Res 39(1):196. https://doi.org/10.1186/s13046-020-01695-8
Zhang D, Ma S, Zhang C, Li P, Mao B, Guan X, Zhou W, Peng J, Wang Xi, Li S, Jia W (2021) MicroRNA-935 directly targets FZD6 to inhibit the proliferation of human glioblastoma and correlate to glioma malignancy and prognosis. Front Oncol 11:566492. https://doi.org/10.3389/fonc.2021.566492
Zhang J, Chen B, Gan C, Sun H, Zhang J, Feng L (2023) A comprehensive review of small interfering RNAs (siRNAs): mechanism, therapeutic targets, and delivery strategies for cancer therapy. Int J Nanomedicine 18:7605–7635. https://doi.org/10.2147/IJN.S436038
Zhao Y, Cui S, Wang Y, Ruirong Xu (2022) The extensive regulation of microRNA in immune thrombocytopenia. Clin Appl Thromb Hemost 28:107602962210935. https://doi.org/10.1177/10760296221093595
Zhao Q, Pang G, Yang L, Chen S, Xu R, Shao W (2021) Long noncoding RNAs regulate the inflammatory responses of macrophages. Cells 11(1):5. https://doi.org/10.3390/cells11010005
Zhao X, Cai Y, Xu J (2019) Circular RNAs: biogenesis, mechanism, and function in human cancers. Int J Mol Sci 20(16). https://doi.org/10.3390/ijms20163926
Zheng J, Liu X, Xue Y, Gong W, Ma J, Xi Z, Que Z, Liu Y (2017) TTBK2 circular RNA promotes glioma malignancy by regulating MiR-217/HNF1β/Derlin-1 pathway. J Hematol Oncol 10(1):52. https://doi.org/10.1186/s13045-017-0422-2
Zhong X, Cai Y (2021) Long non-coding RNA (LncRNA) HOXD-AS2 promotes glioblastoma cell proliferation, migration and invasion by regulating the MiR-3681-5p/MALT1 signaling pathway. Bioengineered 12(2):9113–9127. https://doi.org/10.1080/21655979.2021.1977104
Zhou T, Li Z, Jiang Y. Su K, Xu C, Yi H (2024) Emerging roles of circular RNAs in regulating the hallmarks of thyroid cancer. Cancer Gene Ther 31(4):507–516. https://doi.org/10.1038/s41417-024-00736-0
Acknowledgements
We would like to acknowledge the support and resources provided by the SRM Institute of Science and Technology for this review article. I acknowledge Biorender (www.biorender.com) for providing templates and tools used in the creation of figures in this manuscript.
Author information
Authors and Affiliations
Contributions
S.K. Resources, Conceptualization, data curation, methodology, Formal analysis, writing (original draft), and writing (review). T.T. Conceptualization, Supervision, Formal analysis, validation, and editing. All authors have read and agreed to the final version of the manuscript. The authors confirm that no paper mill and artificial intelligence was used.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sandhanam, K., Tamilanban, T. Unraveling the noncoding RNA landscape in glioblastoma: from pathogenesis to precision therapeutics. Naunyn-Schmiedeberg's Arch Pharmacol (2024). https://doi.org/10.1007/s00210-024-03265-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00210-024-03265-7