Abstract
MicroRNAs (miRNAs) are endogenously expressed small noncoding RNAs that act as posttranscriptional regulators of gene expression. Dysregulation of these molecules has been observed in many types of cancers. Altered expression levels of several miRNAs were identified also in gliomas. It was been frequently shown that miRNAs are involved in core signaling pathways, which play key roles in cellular processes, such as proliferation, apoptosis, cell cycle regulation, invasion, angiogenesis, and stem cell behavior. Therefore, miRNAs have a great potential to act as new class of diagnostic, prognostic, and predictive biomarkers as well as promising therapeutic targets in gliomas. Here, we summarize the current knowledge about miRNAs significance in glioma molecular pathology, with special focus on their involvement in core signaling pathways, their roles in drug resistance, and their potential clinical implications.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Glial tumors of adults are the most frequently occurring primary tumors of central nervous system with a tendency to invade the surrounding brain tissue, originating in astrocytic glial cells. They are traditionally divided into histopathological subtypes defined by World Health Organization (WHO) classification. Numerous investigations have indicated that the improved understanding of the biology and molecular factors involved in the development, progression, and drug resistance of gliomas will be necessary for prediction of clinical outcome and therapy response, and mainly for development of new therapeutic strategies. Only a few molecular biomarkers have been validated and widely accepted in clinical practice till now and despite introduction of novel therapeutic approaches (bevacizumab, cilengitide) this cancer remains associated with very poor prognosis.
MicroRNAs (miRNAs) are endogenously expressed small noncoding RNAs that act as posttranscriptional regulators of gene expression. Deregulation of miRNAs has been observed in various types of cancers, including gliomas. There is continuously increasing evidence showing miRNAs involvement in core signaling pathways of glioma pathogenesis and their great potential as biomarkers and therapeutic targets.
2 MicroRNAs Function and Biogenesis
MiRNAs are a class of small 18–25 nucleotides long noncoding RNA molecules, whose function is to regulate gene expression on the posttranscriptional level. This regulation is mediated by the binding of miRNA to the 3′-untranslated region (3′UTR) of its target mRNA that induces either direct mRNA chain degradation or translational inhibition followed by deadenylation and mRNA decay. The decision on the fate of the target molecule is dependent on the degree of complementarity of both molecules. Perfect base pairing leads to the degradation in RISC, whereas imperfect bond leads first to the inhibition of mRNA translation subsequently followed by the degradation of target molecule due to its instability. Being able to regulate the translation even in case of imperfect complementarity causes that a single miRNA can regulate multiple mRNAs while a single mRNA can be targeted by different miRNAs. To date, over 1,400 human miRNAs have been identified (Griffiths-Jones et al. 2008). MiRNA expression is largely tissue and cell type specific and, moreover, these molecules play important role in a wide range of biological processes, such as development, differentiation, proliferation, and apoptosis (Niyazi et al. 2011). It is not surprising that deregulation of miRNAs was described as an important step in initiation as well as in progression and metastasis of many cancers including gliomas. On the basis of the target molecules, miRNAs can be classified as onco-miRs or tumor-suppressive miRNAs; some miRNAs may exhibit both features dependently on cellular context in various cancers (Chen 2005).
Most miRNA genes have their own promoters and are transcribed as autonomous transcription units (Carthew and Sontheimer 2009). Primary transcripts (pri-miRNAs) are generally transcribed by RNA polymerase II. Pri-miRNA can be up to 10 kb long and is characterized by the formation of stem loop structures. Single pri-miRNA frequently contains polycistronic cluster of miRNAs that are co-transcribed (Lawler and Chiocca 2009). In the nucleus, the pri-miRNA precursor is further cleaved through the activity of an enzyme called Drosha and cofactor DGCR8/Pasha. This processing results in a precursor miRNA (pre-miRNA) that is usually between 60 and 100 nucleotides in length. The pre-miRNA is exported from the nucleus in a RanGTP-dependent process after binding to transporter Exportin-5. In the cytoplasm, pre-miRNA is processed by the enzyme Dicer and, subsequently, one (passenger) of two chains of mature duplex miRNA is destroyed while guide chain is stabilized by the Argonaute proteins and incorporated into RISC (RNA-induced silencing complex). Recent findings show that guide chain can be derived from both the 5′ and 3′ ends of pre-miRNA hairpin depending on the thermodynamic stability on the 5′ end of the mature miRNA (Schwarz et al. 2003; Ladomery et al. 2011). The RISC complex with a miRNA guide strand is known as a miRISC complex. MiRNA involved in this macromolecular complex is prepared to perform its regulatory effects.
3 MicroRNAs in Glioma Pathogenesis
Tumor behavior is largely dependent on the abilities of the individual tumor cells to have the capacity of uncontrolled proliferation, to regulate own cell cycle, to escape an apoptosis, to regulate angiogenesis, and to migrate and invade in the surrounding tissues. The other specific properties such as the capability to self-renewal, unlimited proliferation potential, and differentiation have been observed in glioma stem cells. It has been described many times that miRNAs are involved in the regulation of these processes in many cancers including gliomas and, thus, these molecules can significantly influence glioma pathogenesis.
3.1 MicroRNAs in Proliferation, Migration, and Invasiveness
Among the key signaling pathways regulating cell proliferation, migration, and invasiveness belong EGFR (epidermal growth factor receptor), PI3K/AKT (phosphatidylinositol-4,5-bisphosphate 3-kinase/v-akt murine thymoma viral oncogene homolog), and NF-κB (nuclear factor of kappa light polypeptide gene enhancer in B-cells) (Nikaki et al. 2012). These signalizations have been many times described as deregulated in gliomas and, thus, deeper understanding of them could help discover new therapeutic approaches in initial as well as recurrent diseases of glial origin. The EGFR signaling network contributes to promotion, progression, and metastasis of the wide range of solid cancers, including gliomas. It is not surprising that this pathway is regulated by many miRNAs, which are therefore promising therapeutic targets (Fig. 4.1).
MicroRNAs involved in regulation of glioma proliferation, migration, and invasion. Bmi-1 BMI1 polycomb ring finger oncogene; Cdk6 cyclin-dependent kinase 6; EGFR epidermal growth factor receptor; IKKα/β/γ conserved helix-loop-helix ubiquitous kinase α/β/γ; IRS-1/2 insulin receptor substrate 1/2; MMP2/9 matrix metalloproteinase 2/9; NF-κB nuclear factor of kappa light polypeptide gene enhancer in B-cells; p21CIP1 cyclin-dependent kinase inhibitor 1A; p27 p27 protein; PI3K phosphatidylinositol-4,5-bisphosphate 3-kinase; PLD2 phospholipase D2; PTEN phosphatase and tensin homolog; Raf zinc fingers and homeoboxes 2; Ras rat sarcoma viral oncogene homolog. —| direct suppression; —> direct activation; ---| indirect suppression; --> indirect activation
Mechanistic studies identified gene targets of miR-21 among important components of the EGFR signaling pathway. GBM (glioblastoma multiforme) cell lines U251 and LN229 characterized by mutated and wild-type PTEN (phosphatase and tensin homolog), respectively, showed a decreased expression of EGFR, activated AKT, Cyclin D, and Bcl-2 (B-cell CLL/lymphoma 2) after silencing of miR-21 expression (Zhou et al. 2010a; Sana et al. 2011). Although miR-21 is known to regulate PTEN and downregulation of miR-21 led to increased PTEN expression, the GBM suppressor effect of antisense miR-21 is most likely independent of the PTEN status because U251 has mutated PTEN (Ren et al. 2010b; Zhou et al. 2010a; Sana et al. 2011). In addition to the PTEN, miR-21 regulates proliferation, migration, invasiveness, and other cell processes through direct or indirect repression of many other cancer genes (Gabriely et al. 2008; Gaur et al. 2011a). Interestingly, significant synergistic inhibition of proliferation and invasiveness was observed after co-inhibition of miR-21 and miR-10b. It was hypothesized that miR-21 inhibitor may interrupt the activity of EGFR pathways, increasing PDCD4 [programmed cell death 4 (neoplastic transformation inhibitor)] and TPM1 [tropomyosin 1 (alpha)] expression and reducing MMP (matrix metalloproteinase) activities, independently of PTEN status. Meanwhile, miR-10b inhibitor induces translational suppression of the mRNA encoding HOXD10 (homeobox D10) leading to the increase of the expression of the well-characterized pro-metastatic gene RHOC (ras homolog family member C). Taken together, combination of miR-21 inhibitor and miR-10b inhibitor could be an effective GBM therapeutic strategy (Dong et al. 2012).
PTEN downregulation followed by AKT activation was described after transfection of GBM cells with the miR-26a-2 primary transcript. Similarly, the miR-26a mimics decreased PTEN protein levels and increased AKT phosphorylation (Huse et al. 2009; Kim et al. 2010; Sana et al. 2011). Another research group also showed that ectopic expression of miR-26a influenced cell proliferation by targeting PTEN and suggested that miR-26a is regulated by transcription factor c-MYC (v-myc avian myelocytomatosis viral oncogene homolog) (Guo et al. 2013).
Modulation of AKT signaling cascade using miRNAs in GBM cell lines was described also by Nan et al. In this study, transfection of miR-451 mimic reduced expression levels of many molecules including AKT1, MMP-2, and MMP-9. According to phenotypic experiments, miR-451 inhibited invasive ability and cell proliferation in GBM cells in vitro (Nan et al. 2010; Sana et al. 2011). Another miRNA involved in the EGFR signaling pathway is miR-7. Kefas et al. showed that miR-7 directly inhibits EGFR expression and independently suppressed the AKT pathway via targeting upstream regulators IRS-1 (insulin receptor substrate) and IRS-2. Moreover, transfection with miR-7 oligonucleotides decreased viability and invasiveness of primary GBM cell lines (Kefas et al. 2008; Webster et al. 2009). Webster et al. also described Raf1, another member of the EGFR signaling pathway, as a direct target of miR-7 in cancer cells (Webster et al. 2009).
Godlewski et al. published that miR-128 expression significantly reduced glioma cell proliferation in vitro and glioma growth in vivo. This effect was explained by direct regulation of the BMI1 (BMI1 polycomb ring finger oncogene), which is significantly upregulated in GBM compared to normal brain while miR-128 showed an opposite regulation. Moreover, miR-128 expression caused a decrease in histone methylation and AKT phosphorylation, and upregulation of p21(CIP1) (cyclin-dependent kinase inhibitor 1A) levels, consistent with BMI1 downregulation (Godlewski et al. 2008).
Finally, miR-221 and miR-222 were revealed as potential regulators of many target genes involved in AKT signaling pathway. Upregulation of miR-221/222 resulted in remarkable increase of p-AKT (phosphorylated AKT) and significant changes in the expression of AKT-related genes in glioma cells. Consequently, miR-221/222 overexpression increased glioma cell proliferation and invasion in vitro and induced glioma growth in mouse model. These results suggest that miR-221/222 enhance glioma malignant phenotype via activation of the AKT signaling pathway (Zhang et al. 2010b; Sana et al. 2011). Another study revealed that phenotypic effect of miR-221/222 is caused at least in part by targeting Cx43 (gap junction protein, alpha 1, 43 kDa), which has been identified as a tumor suppressor and major component for the establishment of GJIC (gap junction intercellular communication) in glial cells (Hao et al. 2012).
Another signaling pathway mentioned at the beginning of this chapter is the NF-κB pathway. NF-κB is the transcription factor with pleiotropic activity owing to its central roles in various biological processes. Aberrant activation of NF-κB signaling pathway has been proved to be important for invasiveness and metastatic capacity of tumors through upregulation of MMPs and transcription factors regulating E-cadherin, such as Snail (snail family zinc finger 1), Twist (twist family bHLH transcription factor), or Slug (snail family zinc finger 2). A critical component in NF-κB regulation is the IKK-β (conserved helix-loop-helix ubiquitous kinase) complex (Ghosh and Karin 2002; Song et al. 2010). Song et al. identified miR-218 expression in glioma cell lines and in human primary glioma tissues was substantially downregulated, when compared to miR-218 expression in normal human astrocytes and normal brain tissues. Forced upregulation of miR-218 dramatically reduced the migratory speed and invasive ability of analyzed cells. Ectopic expression of miR-218 downregulated MMP-9 and reduced NF-κB transactivity at transcriptional level, whereas inhibition of miR-218 has the opposite effect. Authors also demonstrated that miR-218 could inactivate NF-κB/MMP-9 signaling by directly targeting IKK-β (Song et al. 2010; Sana et al. 2011).
High-throughput analysis of the effects of 319 miRNA precursor molecules on cell proliferation in three (A172, LN405, and U87MG) GBM cell lines revealed nine miRNAs (miR-129, miR-136, miR-145, miR-155, miR-181b, miR-342-5p, miR-342-3p, miR-376a, and miR-376b), which had an antiproliferative effect in glioma cells. Moreover, six miRNA target genes [ROCK1 (Rho-associated, coiled-coil containing protein kinase 1), RHOA (ras homolog family member A), MET (met proto-oncogene), CSF1R (colony-stimulating factor 1 receptor), EIF2AK1 (eukaryotic translation initiation factor 2-alpha kinase 1), and FGF7 (fibroblast growth factor 7)] were subsequently validated for the similar effects (Haapa-Paananen et al. 2013).
MiR-124 is another miRNA that plays an important role in regulating proliferation and invasiveness of GBM cells. As one of the possible mechanisms of these effects was revealed signalization through PPP1R3L, an inhibitory member of the apoptosis-stimulating protein of p53 family (IASPP), which is also able to affect growth, cell cycle progression, apoptosis, and metastasis of various types of cancer (Zhao et al. 2013b). Similarly, miR-203 inhibits the proliferation and invasion of U251 GBM cells partially via direct targeting of PLD2 (phospholipase D2) and/or ROBO1 [roundabout, axon guidance receptor, homolog 1 (Drosophila)]/ERK (mitogen-activated protein kinase 1)/MMP-9 signaling pathway (Dontula et al. 2013; Chen et al. 2013d). The ability of miR-100 to reduce proliferation in vitro was subsequently demonstrated in vivo where this miRNA treatment improved overall survival of treated animals. As a direct target of miR-100 was later confirmed SMRT/NCOR2 (silencing mediator of retinoid or thyroid hormone receptor-2/nuclear receptor corepressor 2) (Alrfaei et al. 2013).
Recently, many other miRNAs and their targets were described as involved in the regulation of proliferation, migration, and invasiveness in gliomas. Among them belong miR-326/NOB1 (Nin one binding protein) (Zhou et al. 2013); miR-125/PIAS3 (protein inhibitor of activated STAT, 3), which contributed to reduced STAT3 [signal transducer and activator of transcription 3 (acute-phase response factor)] transcriptional activity and subsequent decreased expression of MMP-2/9 (Shi et al. 2013); miR-138/ETH2 (methionine adenosyltransferase SAM2)–CDK4/6 (Cyclin-dependent kinase 4/6)–pRb (protein retinoblastoma 1)–E2F1 (E2F transcription factor 1) signal loop (Qiu et al. 2013); miR-495/CDK6 (Chen et al. 2013c); miR-149/AKT1 signaling (Pan et al. 2012); miR-330/SH3GL2 (SH3-domain GRB2-like 2) (Qu et al. 2012); miR-195/CCND3 (cyclin D3) (Zhang et al. 2012b); miR-34a/NOTCH1 (Li et al. 2011); miR-128/BMI1 (Godlewski et al. 2008); and miR-124, miR-137, miR-143, miR-145, and let-7 (Silber et al. 2008; Lee et al. 2011b; Koo et al. 2012).
3.2 MicroRNAs in Cell Cycle
Many miRNAs directly or indirectly target molecules such as cyclins, cyclin-dependent kinases, CDK regulators, and others, which are crucial for a progression of the cell through individual cell cycle checkpoints (Fig. 4.2).
MicroRNAs involved in regulation of glioma cell cycle. CCNE1 cyclin E1; CDK6 cyclin-dependent kinase 6; NOB1 NIN1/RPN12 binding protein 1 homolog (S. cerevisiae); p53 protein 53; PI3K/AKT phosphatidylinositol-4,5-bisphosphate 3-kinase/v-akt murine thymoma viral oncogene homolog; pRB1 protein retinoblastoma 1. —| direct suppression; —> direct activation; ---> indirect activation
The best-described miRNAs in gliomas with regard to the cell cycle are undoubtedly members of miR-34 family. Initially, overexpression of miR-34a in U251 GBM cells was described resulting in inhibition of cell growth and arrest in G0/G1 phase, which is inter alia accompanied by increased apoptosis. Moreover, authors supposed that this effect is due to the regulation of SIRT1 (sirtuin 1), a direct target of miR-34a (Luan et al. 2010). Two other studies confirmed c-Met (met proto-oncogene), NOTCH1, NOTCH2, and CDK6 also as direct targets of miR-34a and observed inhibition of G1/S progression after miR-34a artificial upregulation. It seems that miR-34a regulates cell cycle via more important targets and, thus, presents a promising therapeutic target for brain tumors (Li et al. 2009a; Guessous et al. 2010b). CDK6 is confirmed as target molecule also for miR-495 and miR-107. Similar to miR-34a, cell cycle analysis revealed that upregulation of both miRNAs resulted in cell cycle arrest at the G1/S checkpoint. Interestingly, miR-107 inhibits also translation of NOTCH2 and its upstream regulator is p53 (protein 53) (Chen et al. 2013b). MiR-495 inhibits another crucial molecule of G1/S transition, tumor suppressor pRB, highlighting its potential in regulation of cell cycle (Chen et al. 2013c). Further, G1 cell cycle arrest was associated also with overexpression of miR-326, miR-15b, and miR-124. Nevertheless, the mechanisms seem to be different. While miR-326 targets NOB1, miR-15b decreases expression of downstream CCNE1 gene encoding cyclin E1 (Xia et al. 2009; Zhao et al. 2013b; Zhou et al. 2013).
But miRNAs are not involved just in the disruption of G1/S transition; it seems that miR-21 decreases G2/M transition. Gwak et al. showed that transfection of malignant glioma cells by anti-miR-21 resulted in the increased transition at this phase. The mechanism of miR-21 modulation of G2/M transition is probably similar to its regulation of proliferation, through the activation of the PI3K/AKT pathway (Gwak et al. 2012).
Finally, the ability to arrest a cell cycle progression in gliomas is linked also with miR-524-5p, miR-628-5p, let-7a, miR-23b, miR-10b, and miR-128 (Zhang et al. 2009; Gabriely et al. 2011; Jiang et al. 2013; Li et al. 2013c; Wang et al. 2013c).
3.3 MicroRNAs in Apoptosis
MiRNAs are important regulators of apoptotic pathways. Alterations in functioning of these apoptotic miRNAs are frequently observed also in the glial tumors (Fig. 4.3). One of the most important regulatory mechanisms of apoptosis in glioma cells is cross talk between TGF-β (transforming growth factor, beta) and p53 signaling, which are regulated by the well-known onco-miR, miR-21. Papagiannakopoulos et al. reported that p53, TGF-β, and mitochondrial apoptotic networks are derepressed in response to miR-21 knockdown. They published a panel of genes, which are involved in particular pathways and are simultaneously modulated by miR-21 treatment. Some of these genes were subsequently predicted to be direct targets of miR-21 that can stabilize p53 protein levels by interfering with MDM2 (MDM2 oncogene, E3 ubiquitin protein ligase) and/or act as p53 transcriptional cofactors (Papagiannakopoulos et al. 2008; Sana et al. 2011). Inhibition of miR-21 increased also endogenous levels of PDCD4 in human glioma cell lines and activated caspases 9/3, which may be mediated by modulating multiple potential target genes, such as TIMP3 (tissue inhibitor of matrix metalloproteinase 3) (Chen et al. 2008; Zhou et al. 2010b; Sana et al. 2011). The upregulation of PDCD4 and caspase 3 is in contrast to the low level of miR-21 and inactivation of TGF-β1/SMAD signaling, which are the critical upstream regulators of miR-21 in dose- and time-dependent manner in human GBM cell line U251 (Wang et al. 2012a).
MicroRNAs involved in regulation of glioma mitochondrial associated apoptosis. APAF apoptotic peptidase activating factor; CASP3/7/9 caspase 3/7/9, apoptosis-related cysteine peptidase; DAXX death-domain associated protein; HNRNPK heterogeneous nuclear ribonucleoprotein K; JMY unction mediating and regulatory protein, p53 cofactor; p53 protein 53; p63 protein 63; PKM2 pyruvate kinase 2; PPP1R3L an inhibitory member of the apoptosis-stimulating protein of p53 family (IASPP); SH3GL2 SH3-domain GRB2-like 2; TGFB1/2 transforming growth factor, beta 1; TGFBR2/3 transforming growth factor, beta receptor 2/3; TOPORS topoisomerase I binding, arginine/serine-rich, E3 ubiquitin protein ligase; TP53BP2 tumor protein p53 binding protein, 2. —| direct suppression; —> direct activation; ---> indirect activation
Other direct targets of miR-21 involved in apoptosis were confirmed molecules ANP32A (acidic (leucine-rich) nuclear phosphoprotein 32 family, member A) and SMARCA4 (SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 4). In A172 GBM cells, enhanced ANP32A expression compensated for the positive effects of anti-miR-21 treatment on cell viability and apoptosis (Schramedei et al. 2011). MiR-21 silencing also enhances apoptosis of GBM cells also after treatment by the antiangiogenic drug sunitinib and alkylation agent temozolomide (TMZ) (Zhang et al. 2012c; Costa et al. 2013). In relation to the enhancement of chemosensitivity was described also miR-155. This miRNA indicated oncogenic character and its silencing led to the elevation of apoptosis after taxol treatment via EAG1 (potassium voltage-gated channel, subfamily H (eag-related), member 1) expression (Meng et al. 2012).
Interestingly, miR-211 overexpression and MMP-9 treatments led to the activation of the intrinsic mitochondrial/caspase-9/3-mediated apoptotic pathway in both glioma cells and GSCs (Asuthkar et al. 2012). MiR-34a was identified also as regulator of TGF-β signaling in GBM (Genovese et al. 2012). This miRNA was suggested to be promoter of mitochondrial apoptosis in several cancers including gliomas (Li et al. 2011; Sasaki et al. 2012; Bienertova-Vasku et al. 2013). Therefore, it is probable that miR-34a regulates apoptosis in glioma also through the TGF-β signaling pathway.
There are also other miRNAs described as regulators of apoptosis in glioma cells. Overexpression of miR-326 in human glioma cell lines A172 and U373 caused cell cycle arrest at the G1 phase, delayed cell proliferation, and mainly enhanced apoptosis (Zhou et al. 2013). Interestingly, miR-326 targets PKM2 (pyruvate kinase M2), which plays a key role in cancer cell metabolism. This could be a mechanism how miR-326 regulates glioma cell survival (Kefas et al. 2010). MiR-10b suppresses many tumor suppressors including p53, FOXO3 (forkhead box O3), CYLD [cylindromatosis (turban tumor syndrome)], PAX6 (paired box 6), PTCH1 (patched 1), HOXD10, and NOTCH1. Hence, this miRNA shows pleiotropic feature and regulates many key cellular processes (Lin et al. 2012) and its downregulation; it is not expressed in human brain and strongly upregulated in both low-grade and high-grade gliomas and reduces glioma cell growth through cell cycle arrest and activation of apoptosis (Gabriely et al. 2011). MiR-124 regulates apoptosis and other cell processes via PPP1R3L (Zhao et al. 2013b). Similar pleotropic nature has also onco-miR miR-330 targeting SH3GL2 (Qu et al. 2012).
3.4 MicroRNAs in Angiogenesis
Angiogenesis is essential for many physiological processes as well as for tumor development. It can be triggered by extracellular signals such as vascular endothelial growth factors and by genetic alterations such as activation of oncogenes (Sternlicht et al. 1999; Folkman 2006; Zheng et al. 2013). In gliomas, aberrant activation of angiogenic signaling pathways EGFR/PI3K/AKT, NF-κB, VEGF (vascular endothelial growth factor), and PDGF (platelet-derived growth factor) was repeatedly described (Kozomara and Griffiths-Jones 2011; Mizoguchi et al. 2012).
Glioma is one of the first cancers in which angiogenesis was found to be a key phenotypic feature to disease progression. Indeed, GBM is one of the most highly vascularized human cancers. It is well recognized that the progression of human glioma is angiogenesis dependent. Accumulated evidence has shown that high vascular density and overexpression of angiogenic factors correlate with poor prognosis of glioma patients (Mizoguchi et al. 2012).
Recently, several miRNAs were described to be involved in glioma angiogenesis regulation (Fig. 4.4). In addition to miR-30e* and miR-182 activating NF-κB signaling and subsequent expression of downstream angiogenic factors such as VEGF-C and MMPs (Mizoguchi et al. 2012), tumor-suppressive miR-205 can specifically suppress expression of the key angiogenic factor, VEGF-A. The effect of the miR-205 ectopic expression has not only anti-angiogenic impact, but it is accompanied by the induction of apoptosis, cell cycle arrest, impairing of cell viability, clonability, and invasiveness (Yue et al. 2012). Angiogenic function was supposed also for miR-15b and miR-152, but subsequent evaluation of tube formation in cultured endothelial cells with culture supernatant from 9L cells, rat glial cell line derived from gliosarcoma, treated with these miRNAs revealed that only miR-15b significantly reduced capillary-like tube formation. Authors have suggested that this effect is caused by NRP2 (neuropilin 2), which was confirmed to be a direct target of miR-15b. NRPs are receptors for the SEMA (class-3 semaphorin) family of axon guidance molecules and also for VEGF family of angiogenic factors. VEGF–NRP interactions promote developmental angiogenesis and also metastases (Geretti and Klagsbrun 2007; Zheng et al. 2013).
MicroRNAs involved in regulation of glioma angiogenesis. HGS hepatocyte growth factor-regulated tyrosine kinase substrate; MMPs matrix metalloproteinases; NF-κB nuclear factor of kappa light polypeptide gene enhancer in B-cells; NRP2 neuropilin 2; PDGFRB platelet-derived growth factor receptor, beta polypeptide; VEGF-A vascular endothelial cell growth factor A; VEGFR2 VEGF kinase insert domain receptor (a type III receptor tyrosine kinase). —| direct suppression; —> direct activation; ---> indirect activation
Glioma cells and angiogenic growth factors elevate the level of miR-296 in primary human brain microvascular endothelial cells in culture. Subsequently, growth factor-induced miR-296 significantly contributes to angiogenesis by directly targeting the HGS (hepatocyte growth factor-regulated tyrosine kinase substrate) leading to reduced HGS-mediated degradation of the VEGFR2 (vascular endothelial growth factor receptor 2) and PDGFRβ (platelet-derived growth factor receptor, beta). Furthermore, inhibition of miR-296 reduces angiogenesis in tumor xenografts in vivo (Wurdinger et al. 2008). Similarly, when miR-93-overexpressing U87 glioma cells were cocultured with endothelial cells, they supported endothelial cell spreading, growth, migration, and tube formation. In vivo studies revealed that miR-93-expressing cells induced blood vessel formation allowing blood vessels to extend to tumor tissues in high densities. This effect is explained by suppressing, at least in part, integrin-β8 expression (Fang et al. 2011).
MiR-26a has been described above as a regulator of proliferation. In addition, forced expression of miR-26a in glioma significantly increased tumor growth and angiogenesis in vivo, while reduced expression of this miRNA played opposite role (Qian et al. 2013). Functionally pleiotropic miR-10b is able to regulate angiogenicity in tumor cells resembling mesenchymal subtype of GBM (Lin et al. 2012).
3.5 MicroRNAs in Immune Response
Interferons (IFNs) are cytokines released by lymphocytes that have antiviral, antiproliferative, and immunomodulatory effects. They are connected with the JAK-STAT (Janus kinase-Signal Transducer and Activator of Transcription) signaling pathway and allow communication between cells to trigger protective defenses of the immune system leading to eradication of affected cells. Therefore, deregulation of JAK-STAT cascade is the key step to mediating immunosuppression in the tumor microenvironment (Platanias 2005; Sana et al. 2011). Recently, the possibility was investigated if IFN-β may induce or reduce cellular miRNAs in human gliomas. Analysis of IFN-β treatment on miR-21 expression in glioma cells and intracranial glioma xenografts revealed that systemic delivery of this cytokine markedly reduced the level of miR-21 in glioma cells. In contrast, the addition of the STAT3-specific inhibitor increased the level of miR-21 (Ohno et al. 2009; Sana et al. 2011). Another study revealed miR-221 and miR-222 as possible regulators of IFN pathways. Zhang et al. found that the IFN-α signaling pathway is the most significant pathway modulated by genes with the different expression after knockdown of miR-221 and miR-222. The authors showed that STAT1 and STAT2 expression and phosphorylation were upregulated in U251 cells with silenced miR-221/222. Tyrosine phosphorylation of STAT1 and STAT2 was present in the nucleus after repression of the same miRNAs. These data illustrate a mechanism of STAT1/2 upregulation under the transcriptional control of IFN-α signaling after knockdown of miR-221/222 cluster in U251 glioma cells (Zhang et al. 2010a; Sana et al. 2011).
Interestingly, on the basis of miRNA expression in gliomas using tissue microarrays, in situ hybridization, and molecular modeling, miR-124 was identified as another candidate for modulating STAT3 signaling pathway. MiR-124 is absent in all grades and pathologic types of gliomas. Upon replacement of miR-124 in GSCs (glioma stem cells), the STAT3 pathway was inhibited, and miR-124 reversed GSC-mediated immunosuppression of T-cell proliferation and induction of Foxp3 (forkhead box P3) regulatory T cells (Treg). Treatment of T cells from immunosuppressed GBM patients with miR-124 induced remarkable effector response including upregulation of IL-2 (interleukin 2), IFN-γ, and TNF-α (tumor necrosis factor, alpha). Both systemic administration of miR-124 and adoptive miR-124-transfected T-cell transfers caused strong anti-glioma therapeutic effects and enhanced effector responses in the local tumor microenvironment in vivo. These findings highlight the potential application of miR-124 as a novel immunotherapeutic agent for gliomas (Shi et al. 2013).
3.6 MicroRNAs in Glioma Stem Cells
During the last years there is an increasing interest in the small population of glioma cells, later termed as glioma stem cells (GSCs) or glioma initiating cells (GICs), which have the ability to self-renew (Koshkin et al. 2013). Conventional therapies target the cells that rapidly divide, while the benefit of GICs is to divide slowly but infinitely. These properties enable them to initiate new tumors and also to serve as a source of recurrence (Martin et al. 2013). These cells, which maintain stem-like phenotype, show generally much more resistance to radiation or chemotherapy in comparison to “normal” glioma cells (Wang et al. 2010; Yamada and Nakano 2012). Similar to normal stem cells, GSCs express CD133 (prominin 1), NESTIN, OCT4 (POU class 5 homeobox 1), SOX2 (SRY (sex determining region Y)-box 2), and NANOG (Nanog homeobox) markers and are capable of self-renewal as well as they have the ability to initiate neurosphere growth in vitro and generation of highly malignant tumor in NOD/SCID (nonobese diabetic/severe combined immunodeficient) mice (Altaner 2008; Chen et al. 2010b). There is increasing amount of evidence suggesting involvement of miRNAs in the regulation of stem-like phenotype of GSCs (Fig. 4.5).
MicroRNAs involved in regulation of glioma stem cells. BMI1 BMI1 polycomb ring finger oncogene; c-MET met proto-oncogene; CAMTA1 calmodulin binding transcription activator 1; CDK6 cyclin-dependent kinase 6; CTGF connective tissue growth factor; CXCR4 chemokine (C-X-C motif) receptor 4; E2F2 E2F transcription factor 2; NPPA natriuretic peptide A; p14ARF cyclin-dependent kinase inhibitor 2A; p16INK4A cyclin-dependent kinase inhibitor 2A; PTEN phosphatase and tensin homolog; SCC1 protein tyrosine phosphatase; SLUG snail family zinc finger 2; SMAD SMAD family member; SNAIL snail family zinc finger 1; STAT3 signal transducer and activator of transcription 3 (acute-phase response factor); ZEB1 zinc finger E-box binding homeobox 1. —| direct suppression; —> direct activation; ---> indirect activation
Two independent research groups found out that miR-128 inhibits self-renewal and proliferation capacities of GSCs through the direct targeting of BMI1 (Godlewski et al. 2008; Fu et al. 2013), which negatively regulates expression of tumor-suppressor proteins p16INK4a (cyclin-dependent kinase inhibitor 2A) and p14ARF (CDKN2A; cyclin-dependent kinase inhibitor 2A). These two molecules subsequently regulate cyclin D and p53 leading to the cell cycle arrest (Siddique and Saleem 2012). Fu et al. described capability of miR-200 family to suppress an epithelial–mesenchymal transition (EMT) in GSCs via E-cadherin, N-cadherin, SNAIL, SLUG, and ZEB1 (zinc finger E-box binding homeobox 1) (Fu et al. 2013). It was also discovered that miR-200c is involved in the regulation of BMI1 in breast cancer and, thus, this miRNA could function analogically to miR-128 in GSCs (Siddique and Saleem 2012).
Chan et al. published data showing miR-138 being an important regulator of self-renewal, proliferation, and apoptosis in GSCs. Interestingly, this miRNA did not affect healthy neural stem cells. This fact is remarkable accordingly to potential use of miR-138 in therapy (Chan et al. 2012). The similar effects indicated also miR-10b, the inhibition of which decreased proliferation, migration, and invasiveness in vitro and, moreover, the ability of GSCs to initiate tumorigenesis in vivo (Guessous et al. 2013). Oncogenic properties have also miR-9/9* that inhibit expression tumor suppressor CAMTA1 (calmodulin-binding transcription activator 1) and thus support tumorigenic potential of CD133 positive cells. CAMTA1 activates transcription of NPPA (antiproliferative peptide), protein with proven antiproliferative effects in GBM. Furthermore, expression levels of both CAMTA1 and NPPA correlate with prognosis and overall survival of GBM patients (Schraivogel et al. 2011).
On the contrary, miR-34a was observed as tumor-suppressive miRNA in GBM. This molecule targets genes involved in tumorigenesis as well as in regulation of GSCs (c-MET, NOTCH1, NOTCH2). It was demonstrated that miR-34a has significantly lower expression in glioma cells and, further, decreases cell proliferation, migration, invasiveness, and viability when replaced in vitro (Li et al. 2009a). This replacement approach led in GSCs to the induction of apoptosis and differentiation, which was connected with significant reduction of stem cell markers CD133 and nestin (Guessous et al. 2010a). Another regulator of NOTCH signaling is miR-326 that also decreases viability, invasiveness, and tumorigenic potential in GBM when its levels are upregulated (Kefas et al. 2009). MiR-125 is another molecule involved in the regulation of neural differentiation (Le et al. 2009), which is downregulated in GBM tissue and CD133 positive GBM cells. Ectopic expression of miR-125 led to the inhibition of proliferation and sphere formation, to cell cycle arrest in G0–G1 phase, and to the decreased expression of stemness markers indicating cell differentiation. Further, E2F2 (E2F transcription factor 2), critical regulator of cell cycle, is the direct target of miR-125b (Wu et al. 2012). Moreover, miR-124 and miR-137 are involved in a regulation of the cell cycle progression through G1 phase into early S phase in GSCs via inhibition of CDK6. This results in decreased proliferation and enhanced differentiation (Silber et al. 2008). In another study was observed that miR-125b-2 is overexpressed in initial GBMs as well in relapsed tumors after concomitant RT/TMZ (radiochemotherapy with temozolomide) therapy and, thus, this miRNA indicates oncogenic character. Increased levels of miR-125b-2 were also detected in CD133 positive GBM cells. Inhibition of this miRNA leads to the induction of mitochondrial apoptosis after TMZ treatment in CD133 positive cells Shi et al. 2012.
As an important regulators of glioblastoma tumor initiating cells (GICs; this term is sometimes used by authors, who decline to use the term glioma stem cell) have been described also members of cluster miR-302-367. This cluster is able to suppress the self-renewal, infiltration, and proliferation potential of GICs through a repression of CXCR4 (chemokine C-X-C motif, receptor 4) leading to the suppression of SHH-GLI-NANOG signaling pathway (Card et al. 2008; Zbinden et al. 2010; Fareh et al. 2012). Similarly, another cluster miR-17-92 regulates neurosphere formation, differentiation, proliferation, and apoptosis in vitro in GICs. Supposed mediator of these effects is the gene CTFG (connective tissue growth factor), a direct target of this cluster (Ernst et al. 2010). Global expression analysis of miRNAs revealed changes during differentiation of GICs, including overexpression of miR-21, miR-29a, miR-29b, miR-221, and miR-222 and downregulation of miR-93 and miR-106a. Functional studies showed that miR-21 overexpression in GICs induced comparable cell differentiation features and targeted SPRY1 [sprouty homolog 1, antagonist of FGF signaling (Drosophila)] mRNA, which encodes for a negative regulator of neural stem cell differentiation. In addition, miR-221 and miR-222 inhibition in differentiated cells restored the expression of stem cell markers while reducing differentiation markers. Finally, miR-29a and miR-29b targeted MCL1 [myeloid cell leukemia sequence 1 (BCL2-related)] mRNA in GICs and increased apoptosis (Aldaz et al. 2013).
Analysis of the expression profiles of both CD133 positive and negative GBM cells revealed several differentially expressed miRNAs including miR-451, miR-486, and miR-425. Ectopic expression of these miRNAs led to the inhibition of cell growth and neurosphere formation in GBM cells. As positive regulator of miR-451 was subsequently revealed SMAD (Gal et al. 2008). Another analysis showed that miR-153 expression was downregulated in GBM tissues relative to normal brain tissues and in CD133 positive cells relative to CD133 negative cells (Zhao et al. 2013a).
4 MicroRNAs as Biomarkers in Gliomas
Recent research suggests that deregulation of miRNAs is involved in initiation and progression of many cancers, including gliomas, and that miRNAs hold great potential as future diagnostic, prognostic, and predictive biomarkers in cancer. The rationale of this potential is mainly based on the miRNAs’ increased stability when compared to mRNAs and the ability of a single miRNA molecule to affect numerous targets, oncogenes and tumor-suppressor genes including (Hermansen and Kristensen 2013).
4.1 Diagnostic Biomarkers
Study of miRNA expression profiles in tumor tissue of gliomas of different grades and their comparison to miRNA profiles in non-tumoral brain tissue is not only the first step in the determination of miRNAs involved in glioma pathogenesis, but also important approach enabling new level of glioma molecular taxonomy, which is not based on morphology but on concrete molecular alterations which are present in individual tumors. This molecular classification has potential to be more accurate in stratification of the patients accordingly to their prognosis and therapeutic response in comparison to conventional histopathological classification and open new diagnostic possibilities for glioma patients. MiRNAs which were repeatedly found to be differentially expressed between glioma tumor tissue and non-tumoral brain tissue are summarized in Table 4.1.
The first work aiming miRNA profiling in glioblastoma tissue was published by Ciafre et al. (2005). These authors analyzed 254 miRNAs in nine paired GBM and non-tumor brain tissue samples. The findings showed that nine miRNAs (miR-10b, miR-130a, miR-221, miR-125b-1, miR-125b-2, miR-9-2, miR-21, miR-25, and miR-123) were upregulated and four miRNAs (miR-128a and miR-181a/b/c) downregulated in GBM. The greatest expression changes were observed in miR-221, miR-128a, and miR-181a/b/c (Ciafre et al. 2005). A similar analysis was performed by Chan et al., which compared three primary high-grade gliomas with eight samples of fetal and adult brain tissues. In this study, five miRNAs (miR-21, miR-138, miR-347, miR-135, and miR-291-5) were significantly overexpressed and three miRNAs (miR-198, miR-188, and miR-202) showed lower expression levels in tumor tissue. MiR-21 was almost nine times more expressed in GBM compared to control samples (Chan et al. 2005). Silber et al. analyzed 192 matured miRNAs in four anaplastic astrocytomas (grade III), four GBMs (grade IV), and four non-tumor tissues obtained from surgically resected tissue of temporal lobes of epileptic patients. Statistical analysis revealed 13 downregulated and three upregulated miRNAs in GBM compared with the non-tumor tissue. Furthermore, only six deregulated miRNAs were observed between grade III and IV gliomas (Silber et al. 2008).
Similar study was published comparing global expression profiles of miRNAs in 39 gliomas (13 primary and 13 secondary GBMs, 13 anaplastic astrocytomas) and seven normal brain tissues and revealed 55 upregulated and 29 downregulated miRNAs in glioma tissue. Moreover, 67 differentially expressed miRNAs were identified by a comparison of anaplastic astrocytomas and GBMs. The comparison of primary and secondary GBMs revealed seven differentially expressed miRNAs. Several miRNAs significantly differed also between sequentially progressive astrocytomas (anaplastic astrocytomas and secondary GBM) and primary GBMs (21 miRNAs), anaplastic astrocytomas and primary GBMs (76 miRNAs), as well as anaplastic astrocytomas and secondary GBMs (68 miRNAs) (Rao et al. 2010). Indeed, D’Urso et al. found that a set of 10 miRNAs is able to classify primary and secondary GBMs (D’Urso et al. 2012).
Skalsky and Cullen investigated miRNA expression profiles in six GBMs and three samples of non-tumor brain tissues using high-throughput sequencing technology. They published 68 differentially expressed miRNAs between both examined sets (p < 0.05, Student’s t-test). Nevertheless, when they applied the standard FDR (false discovery rate)-adjusted p-value < 0.05 cutoff, only nine miRNAs (miR-124, miR-10b*, miR-139-5p, miR-7, miR-10b, miR-132, miR-95, miR-543, miR-7d) were identified that met the criteria. Given the much broader dynamic range of deep sequencing compared to finite platforms such as miRNA microarrays or real-time PCR (Polymerase Chain Reaction) arrays, they therefore used an FDR cutoff of p < 0.1. Under these conditions 12 more miRNAs (miR-323-3p, miR-128, miR-139-3p, miR-598, miR-103a, miR-103b, miR-873, miR-891a, miR-487b, miR-323b-3p, miR-138-1*, miR-93) met the criteria (Skalsky and Cullen 2011).
Sasayama et al. analyzed three paired samples of GBM and non-tumor brain tissues. They found that five miRNAs (miR-10b, miR-21, miR-183, miR-92b, and miR-106b) were upregulated and five miRNAs (miR-302c*, miR-379, miR-329, miR-134, and miR-369-3p) were downregulated in GBMs (Sasayama et al. 2009). Wang et al. also analyzed miRNA expression profiles in three GBMs and three paired non-tumor adjacent tissues. They identified 91 miRNAs, whose expressions were at least twofold changed between both sets of samples. Interestingly, an expression of miR-483-5p was almost 100-fold decreased in tumors (Wang et al. 2012b). Another study validated expression of eight selected miRNAs in 10 GBM tissues in comparison to the four non-tumor samples of brain tissues obtained from epileptic patients. MiR-21 and miR-221 showed higher and miR-181b lower expression levels in tumor samples (Conti et al. 2009). Slaby et al. compared expressions of eight selected miRNAs in GBMs and non-tumor brain tissues obtained from surgeries of arteriovenous malformations (AVM). Results showed that while miR-21 and miR-125b were overexpressed in GBMs, miR-128a, miR-181a/b/c, and miR-221/222 were downregulated in tumors (Slaby et al. 2010). Other studies show that miR-31 (Hua et al. 2012), miR-205 (Yue et al. 2012), miR-124a (Fowler et al. 2011), and miR-34a (Li et al. 2011) have lower expression levels in GBM in comparison with normal brain tissue.
Malzkorn et al. analyzed miRNA expression profiles among different grades of gliomas (low-grade astrocytomas vs. anaplastic astrocytomas vs. secondary GBMs). Authors studied expression profiles of 157 miRNAs in patients affected by low-grade astrocytomas (grade II) that gradually progressed to secondary GBMs (grade IV). It was found that 12 miRNAs (miR-9, miR-15a, miR-16, miR-17, miR-19a, miR-20a, miR-21, miR-25, miR-28, miR-130b, miR-140, and miR-210) are upregulated and two miRNAs (miR-184 and miR-328) are downregulated dependently on the grade of glioma (Malzkorn et al. 2010).
There are also many studies characterizing expression pattern of only one miRNA in gliomas and their different clinicopathological features. From these studies it implies that miR-34a, miR-203, miR-326, and miR-375 are reduced in tumors (Chang et al. 2012; Gao et al. 2013; He et al. 2013; Wang et al. 2013b) and miR-17, miR-19a/b, miR-224, miR-335, and miR-372 are overexpressed in gliomas (Jiang et al. 2012; Lu et al. 2012, 2013; Jia et al. 2013; Li et al. 2013a) compared to non-tumor brain tissues. Moreover, most of these miRNAs were significantly correlated with grading of gliomas.
4.2 Prognostic and Predictive Biomarkers
One of the main aims in glioma research is the discovery of highly sensitive prognostic and predictive biomarkers enabling stratification of the patients accordingly to their risk of progression and predicted therapeutic response. The importance of this challenge increases with the advent of new therapeutic possibilities in the treatment of gliomas, e.g., TMZ, bevacizumab, and cilengitide (Chamberlain 2011). MiRNAs with potential as prognostic and/or predictive biomarkers are summarized in Table 4.2.
Srinivasan et al. analyzed 10 selected miRNAs in 222 GBM patients. The aim of this study was to find differently expressed miRNAs between short-time and long-time surviving GBM patients. Tumor samples of the patients with long-time OS (overall survival) showed higher expression levels of protective miRNAs (miR-20a, miR-106a, miR-17-5p). Conversely, overexpression of oncogenic miRNAs (miR-31, miR-221, miR-222, miR-148a, miR-146b, miR-200b, miR-193a) was observed in tumors of patients with short-time OS (Srinivasan et al. 2011). Further, Niyazi et al. performed retrospective study analyzing 1,100 miRNAs in 35 GBM patients, who underwent adjuvant concomitant chemoradiotherapy. Subsequently, 30 miRNAs, which mostly differ between short- and long-time surviving patients, were selected for the following analysis. However, only five miRNAs were significantly deregulated: miR-3163 (fold change (FC) = 2.0, p = 0.05), miR-539 (FC = 0.5, p = 0.001), miR-1305 (FC = 0.5, p = 0.05), miR-1260 (FC = 0.5, p = 0.03), and let-7a (FC = 0.3, p = 0.02). Nevertheless, an application of all 30 miRNAs classified tested patients into two groups according to overall survival. The miRNA pattern and the prognostic power were both independent of the MGMT (O-6-methylguanine-DNA methyltransferase) methylation status. When performing a multivariate analysis for all patients including the factors MGMT status, miRNA pattern, adjuvant TMZ, age category, and RPA (recursive partitioning analysis) class, it turns out that only adjuvant TMZ remains a prognostic factor (p = 0.01) and MGMT status and miRNA pattern lose their prognostic significance (p = 0.17 and p = 0.22) (Niyazi et al. 2011).
Recent study published by Slaby et al. described significant negative impact of miR-181c and miR-181b on the response to the concomitant chemoradiotherapy with TMZ (Slaby et al. 2010). Lakomy et al. analyzed expression of eight miRNAs (miR-21, miR-128a, miR-181c, miR-195, miR-196a, miR-196b, miR-221, and miR-222) in GBM patients treated with concomitant chemoradiotherapy with TMZ and results subsequently correlated with MGMT status and clinical data. It was found that miR-195 (p = 0.0124) and miR-196b (p = 0.0492) had a positive impact on overall survival of the patients. Moreover, combination of miR-181c and miR-21 enabled to identify patients, which progressed in the 6 months after diagnosis (sensitivity 92 %, specificity 81 %, p < 0.0001) (Lakomy et al. 2011). These results are partially conflicting with conclusions of Ujifuku at al., who identified miR-195, miR-455-3p, and miR-10a* to be three most increased miRNAs in the TMZ-resistant cell lines. Moreover, miR-195 silencing led to the increased sensitivity to TMZ in these cells (Ujifuku et al. 2010). Similarly, Guan et al. described significant relationship between high expression of miR-196a/b and a worse prognosis of GBM and anaplastic astrocytoma patients (Guan et al. 2010). However, it is interesting that miR-195 and miR-196b have the same impact on prognosis in colorectal, hepatocellular, and adrenocortical carcinomas as described by Lakomy et al. (Soon et al. 2009; Xu et al. 2009; Liu et al. 2010; Wang et al. 2012c).
Wang et al. evaluated the prognostic value of miR-214 in overall survival of 108 glioma patients (WHO I—18, WHO II—12, WHO III—32, WHO IV—46). The overall survivals of patients, whose tumors expressed low level of miR-214, were significantly shorter than those with high level of miR-214 (p < 0.001). Moreover miR-214 was significantly downregulated compared to 20 normal brain tissues (p = 0.001) (Wang et al. 2014). Similarly, miR-34a, miR-203, miR-326, and miR-375 were reduced in tumors and their lower expression levels were significantly associated with worse progression-free survival and overall survival of glioma patients. Furthermore, multivariate Cox regression analyses indicated that all these miRNAs have been independent prognostic factors (Chang et al. 2012; Gao et al. 2013; He et al. 2013; Wang et al. 2013b).
On the contrary, miR-9 expression in 128 glioma tissues (WHO I—18, WHO II—14, WHO III—38, WHO IV—58) was significantly higher in comparison to 10 corresponding non-neoplastic brain tissues (p < 0.001). The increased expression of miR-9 was more frequently observed in the tissue of high-grade gliomas (p = 0.001). The expression levels of miR-9 in glioma tissues with low Karnofsky performance score (KPS) were also significantly higher than those with high KPS (p = 0.008). Moreover, the overall survival of glioma patients with high miR-9 expression was obviously lower than that with low miR-9 expression (p < 0.001). Multivariate analysis further showed that high miR-9 expression was an independent prognostic factor for overall survival in glioma patients (p = 0.01). More importantly, the subgroup analyses indicated that the overall survival of patients with high-grade gliomas was significantly worse for high miR-9 expression group than for low miR-9 expression group (p < 0.001), but no significant difference was found for patients with low-grade (I–II) gliomas (Wu et al. 2013). Furthermore, miR-650 expression was increased in 168 gliomas (WHO I—35, WHO II—40, WHO III—41, WHO IV—52) compared with 21 normal control specimens (p < 0.001). It was also found that miR-650 expression was related to grade and KPS, whereas high expression was more frequently detected in gliomas of high grade or low KPS score (p < 0.001). The prognosis of glioma with high miR-650 expression was significantly worse compared with gliomas with low miR-650 expression (Sun et al. 2013).
MiR-21 is the most consistently overexpressed miRNA in many cancers including gliomas. To better understand the role of miR-21 in gliomas, paraffin-embedded glioma tissue samples from 193 patients with WHO I–IV tumors were analyzed by in situ hybridization. It was found that miR-21 expression is localized in tumor cells and tumor-associated blood vessels, whereas no expression was seen in adjacent normal brain parenchyma. Only tumor cell miR-21 was associated with poor prognosis when adjusting for known clinical parameters in a multivariate analysis (Hermansen et al. 2013). Similarly, miR-335 expression level was also described to be significantly higher than that in corresponding non-neoplastic brain tissues (p < 0.001) and, in addition, survival analysis demonstrated that patients with high miR-335 expression tumors had significantly shorter survival times (p = 0.01) and that miR-335 is an independent prognostic factor (p = 0.02) (Jiang et al. 2012). Similarly, miR-17, miR-224, and miR-372 are overexpressed in gliomas and their higher expression levels are significantly associated with worse progression-free survival and overall survival of glioma patients. Multivariate Cox regression analyses indicated that all these miRNAs are independent prognostic factors (Lu et al. 2012, 2013; Li et al. 2013a).
5 MicroRNAs Involved in Chemoradioresistance of Gliomas
DNA damage, caused by radiation, activates pathways that protect cells from apoptosis and give them the ability to grow and proliferate. There are two well-known pathways: PI3K/AKT and ATM (ataxia telangiectasia mutated)/Chk2 (checkpoint kinase 2)/p53 that are activated as a response to the treatment. These pathways are regulated by several miRNAs. As for radiation, DNA damage is the main reason for cell death in normal cells, where AKT is important activator of multiple proteins, such as DNA-PK (protein kinase, DNA-activated, catalytic polypeptide), which subsequently leads to DNA repair, BCL-2, or mTOR [mechanistic target of rapamycin (serine/threonine kinase)], which ensures cell growth and proliferation. Next possibility of cells to recognize DSB (double strand breaks) or DNA damage is via BRCA1 (breast cancer 1, early onset) and activation of ATM/CHK2/p53 signaling pathway. The survival and cell apoptosis is dependent on activation of downstream proteins of this pathway.
Among treatment approaches that modulate and target DNA, TMZ is one of the currently frequently used. TMZ is an alkylating agent, which methylates nucleotides and so causes inhibition of DNA replication. Resistant cancer cells have evolved ways how to overcome this process. First mechanism is based on the MGMT enzyme that is able to excise nucleotides in DNA base-pair mismatch, and so neutralize the effect of TMZ. Other mechanism is efflux of TMZ by ATP-binding cassette transporters, which decrease the concentration of the drug within the cell. ATP-binding cassettes are probably the main reason for multidrug resistance. These processes were experimentally proved to be re-regulated by miRNAs (Fig. 4.6).
MicroRNAs involved in radiotherapy and chemotherapy resistance. AKT v-akt murine thymoma viral oncogene homolog; ATM ataxia telangiectasia mutated; BAD BCL2-associated agonist of cell death; BCL2 B-cell CLL/lymphoma 2; BRCA1 breast cancer 1; CCND3 cyclin D3; CDK4/6 cyclin-dependent kinase 4/6; DNA-PK protein kinase, DNA-activated, catalytic polypeptide; E2F3 E2F transcription factor 3; EGFR epidermal growth factor receptor; KU70/80 X-ray repair complementing defective repair in Chinese hamster cells 5/6; MGMT O-6-methylguanine-DNA methyltransferase; mTOR mechanistic target of rapamycin (serine/threonine kinase); p27 protein 27; p53 protein 53; PI3K phosphatidylinositol-4,5-bisphosphate 3-kinase; PIP2 phosphatidylinositol 4,5-bisphosphate; PIP3 phosphatidylinositol (3,4,5)-triphosphate; PTEN phosphatase and tensin homolog; RB retinoblastoma; SMARCA5 (SNF2H) SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 5; TMZ temozolomide. —| direct suppression; —> direct activation; ---| indirect suppression; ---> indirect activation
5.1 MicroRNAs Involved in Radioresistance
PI3K/AKT (Lee et al. 2011a) and ATM/CHk2/p53 (Squatrito et al. 2010) are two main pathways involved in radioresistance. These pathways are responsible for either activation of downstream proteins involved in repair of damaged DNA caused by radiation or regulation of apoptosis and survival rate (Li et al. 2009a). EGFR which triggers PI3K/AKT signaling pathways is important for proliferation activity of cells and is highly expressed in various types of cancer including gliomas. Furthermore, there is substantial experimental evidence supporting a causal role for aberrant EGFR signaling in cancer pathogenesis and resistance to treatment (Huang et al. 2009; Hatanpaa et al. 2010). PI3K is crucial for normal brain development and function. It was found to be hyperactivated in brain tumors (Guillamo et al. 2009; Kwiatkowska and Symons 2013). Studies concerning miRNAs and resistance revealed a connection between miRNAs and downstream proteins of EGFR/PI3K/AKT signaling.
PI3K/AKT signaling pathway is usually activated in GBM. Inactivation of this pathway impairs mechanisms of DNA repair, which can consequently enhance radiosensitivity. MiR-21 was shown to activate this pathway through suppression of PI3K/AKT inhibitor PTEN (Chakravarti et al. 2004; Kao et al. 2007; Gwak et al. 2012). MiR-21 belongs to one of the most studied miRNAs across many tumors including gliomas and is classified as an onco-miR. (Hong et al. 2013). It seems than miR-21 act as a major player in radioresistance, as decrease of this miRNA sensitizes GBM cell lines to irradiation. Sensitization of cells is mediated via γ-H2AX (H2A histone family, member X, gamma) blocking, which serves as an indicator of DSB and subsequently leads to prolongation of DNA repair (Gwak et al. 2012).
Further studies involved miR-7 in the regulation of PI3K/AKT signaling (Kefas et al. 2008). The involvement of miR-7 in this pathway was evaluated on two glioma cell lines U251 and U87, where increased level or miR-7 induced attenuation of EGFR and AKT, and caused a radio-sensitization (Lee et al. 2011a). AKT is also important player in DNA damage repair as it controls expression of DNA-PK. DNA-PK serves as an activator of heterodimers Ku70/Ku80 (X-ray repair complementing defective repair in Chinese hamster cells 5/6), which are able to recognize DNA DSB during NHEJ (non-homologous end joining), and so it enables to repair DNA after DNA damage (Fell and Schild-Poulter 2012). As by decrease of miR-21, also miR-7 was able to regulate response of cell to DNA damage, through regulation of DNA-PK. This indicates the relationship between delayed DNA damage repair and increased radiation-induced cell killing by miR-7 overexpression. It seems that miR-7 could be a useful therapeutic target for overcoming the radioresistance of GBM and other tumors with activated EGFR-PI3K-AKT signaling, as it had an effect in different cancer types treated with radiation (Lee et al. 2011a).
MiR-181a is next miRNA which has been described in connection with GBM and radiation treatment. It affects anti-apoptotic gene BCL2 and it was shown that irradiation exhibited and induced expression of miR-181a, while overexpression of miR-181a caused sensitization to radiation (Chen et al. 2010a).
ATM/Chk2/p53 pathway is closely connected to cellular processes such as regulation of apoptosis, cell cycle, or senescence (Cao et al. 2006). The importance of ATM level in radioresistance was shown on two GBM cell lines, which differ in ATM expression. An M059J radiosensitive cell line is characterized by lower level of ATM and M059K radioresistant cell line, which show higher expression of ATM due to deficiency in DNA-PK. There is an evidence that miR-100 could be involved in this process, as it is differently expressed in these cell lines and accordingly to the computational analyses is a direct regulator of ATM (Ng et al. 2010). Irradiation of M059K and M059J cell lines caused upregulation of several miRNAs: miR-17-3p, miR-17-5p, miR-19a, miR-19b miR-142-3p, and miR142-5p. However, cell line with lower level of ATM and normal DNA-PK activity exhibited induction of miR-15a, miR-16, miR-21, miR-143, and miR-155 level (Chaudhry et al. 2010b). Establishment of these miRNAs in radiation response was confirmed by other studies. Study focused on lymphoblastoid cell revealed that low dosage of IR elevates expression level of miR-15a, miR-16, and miR-21 (Chaudhry et al. 2010a). Mir-143 was found to directly target gene FHIT (fragile histidine triad) the deregulation of which is often seen in epithelial tumors and homozygous deletion contributes to higher radioresistance. Regulation of FHIT was shown via miR-143 overexpression which consequently leads to cell cycle arrest in G2 phase (Lin et al. 2011). It was also shown that elevated level of miR-155 serves as a radioprotectant in lung cancer cells. Inhibition of miR-155 led to radio-sensitization (Babar et al. 2011). ATM and DNA-PK were confirmed as targets of miR-101. These miRNAs sensitized U87MG glioma cells to radiation after lentiviral transduction in vitro and in vivo (Yan et al. 2010).
For radioresistance of cells it is very important to recognize and repair DSB or DNA damage, chromatin remodeling protein complex. A positive correlation between miR-99 and SWI/SNF (SWItch/Sucrose NonFermentable) chromatin remodeling factor SNF2H/SMARCA5 (SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 5), a component of the ACF1 (bromodomain adjacent to zinc finger domain, 1A) complex, was found. Moreover, it has been elucidated that reduction of BRCA1 level on DNA damage site was arranged due to the downregulation of SNF2H, miR-99a, and miR-100. Ectopic expression of the miR-99 family in cells reduced the rate and overall efficiency of repair by both homologous recombination and non-homologous end joining, which led to sensitization of cells to radiation (Nakano and Kornblum 2006; Mueller et al. 2013; Besse et al. 2013). Effects of miRNAs and their targets are summarized in Table 4.3 and Fig. 4.6.
5.2 MiRNAs Involved in Chemoresistance
Temozolomide is a cytotoxic prodrug agent, which after hydrolyzation methylates nucleotides and causes inhibition of DNA replication. The resistance after long-time exposure is nearly universal. This is caused by three main mechanisms involved in DNA repair: increased levels of MGMT; a deficient mismatch repair process (MMR); and activation of the PARP (poly(ADP)-ribose polymerase) pathway (Darkes et al. 2002; Carmo et al. 2011).
MGMT repairs mutagenic DNA lesions, caused by TMZ guanines methylation on O6 position that subsequently leads to base-pair mismatch with thymine. Further, MGMT prevents mismatches and errors during DNA replication and transcription. Cells with increased level of MGMT become resistant to the exposure of TMZ, while in nonresistant glioma cells the pathway induced by TMZ is active and is able to induce cell death (Darkes et al. 2002; Sharma et al. 2009).
As described previously, miRNAs have the possibility to regulate multiple genes, as was observed in miR-181, which is involved in the regulation of BCL2 and MGMT. Silencing of miR-181 family led to increased level of MGMT. Therefore, miR-181 could be used as a predictive factor to TMZ response (Fig. 4.6) (Chen et al. 2010a; Zhang et al. 2012c).
Correlation of TMZ chemoresistance and survival pathways PI3K/AKT and ERK1/2/MAPK (mitogen-activated protein kinase) was confirmed on U118 cell line, which evinces chemoresistance, while after inhibition of these pathways chemoresistance was partially eradicated (Carmo et al. 2011).
Regulation of PI3K/AKT by miR-21 was described also by TMZ-treated glioma cell lines. Decreased level of this miRNAs caused inhibition of proliferation, increased apoptosis, and cell cycle arrest. This was particularly caused by the decrease of BAX (BCL2-associated X protein)/BCL-2 ratio and decreased caspase-3 activity (Ren et al. 2010b; Shi et al. 2010). Among miRNAs targeting apoptotic factors such as BAX, cytochrome c, and cleaved caspase-3 belongs miR-221/222 as well. Their decreased levels enhanced sensitivity to TMZ, but independently of the p53 status (Chen et al. 2012).
To evaluate differences in cell resistant to TMZ, two cell lines U251R, U251 were compared on their miRNA profile. This revealed an elevated level of miR-195, miR-455-3p, and miR-10a* in U251R (TMZ-resistant) cells, which could be important in TMZ resistance. Inhibition of miR-455-3p or miR-10a(*) showed modest cell killing effect in the presence of TMZ. However, miR-195 with TMZ in combination strongly increased cell death (Ujifuku et al. 2010). This was caused by inverse correlation of E2F3 (E2F transcription factor 3) and CCND3 (cyclin D3) and expression level of miR-195. These results indicate that E2F3 and CCND3 are targets of this miRNA. E2F3 is able to suppress transcription of genes related to cell cycle progression and so causes G1 phase arrest. On the other hand CCND3 suppression mediated by miR-195 allows an increase of p27Kip1 (cyclin-dependent kinase inhibitor 1B) expression level, which consequently influences proteins involved in cell migration, resulting in repression of GBM cell invasion (Zhang et al. 2012a).
Although TMZ is the most often used chemotherapeutical agent, there are other chemotherapeutics used in GBM treatment. VM-26 (Teniposide) was tested on U373MG cell line in combination with miR-21 inhibitor, which led to enhancement of cytotoxicity. As a direct target LRRFIP1 (leucine-rich repeat (in FLII) interacting protein 1) was validated, whose product is an inhibitor of NF-kB, a downstream effector of PI3K/AKT signaling (Li et al. 2009b). MiR-21 effect was also examined in cells treated by 5-FU (5-fluorouracil) and showed increased apoptosis and decreased ability to migrate (Ren et al. 2010a). Not only deregulation of PI3K pathway and its downstream factor is responsible for resistance, but also family of proteins called ABCG2 (ATP-binding cassettes, a subfamily of G member 2 protein family), which contribute to multidrug resistance. These transporters cause selective survival of cells through the efflux of drugs. MiR-328 contributes to the regulation of ABCG2 expression, while increased level of this miRNA led to decrease of ABCG2 expression (Li et al. 2010). Regulatory effects of miRNAs involved in chemoresistance of gliomas are summarized in Table 4.4 and Fig. 4.6.
5.3 Glioma Initiating Cells/Stem Cells and Their Role in Tumor Resistance
GSCs, which maintain stem-like phenotype, show generally much more resistance to radiation or chemotherapy in comparison to “normal” glioma cells (Wang et al. 2010; Yamada and Nakano 2012). Therefore, it seems that the best way how to decrease the number of recurrences which is connected with high treatment resistance is to target these cells and their signaling pathways. This population of CD133 positive (CD133+) cancer stem cells contributes to the radio- and chemoresistance and bears the activation of NOTCH and SHH (Sonic hedgehog) pathways. Inhibition of these pathways led to enhancement of CD133+ glioma cells to TMZ treatment (Ulasov et al. 2011). Further, SHH has been found to play a role in maintaining undifferentiated stem cell state in GSCs and Notch signaling in active GSCs as well as in GBM pathogenesis (Yamada and Nakano 2012). In fact, NOTCH and SHH were shown to be upregulated in GSCs and their inhibition led to increased susceptibility of these cells to TMZ. Involvement of miRNAs in regulation of stem-like phenotype of GSCs is described in detail in Sect. 4.3.6.
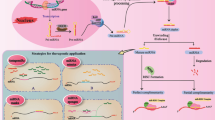
6 MicroRNA as Therapeutic Targets
The role of miRNA in carcinogenesis depends on the functions of their targets (oncogenes or tumor suppressors). Downregulated miRNAs, which regulate oncogenes, are tumor-suppressive miRNAs (TS-miRNAs), while miRNAs overexpressed in tumors and inhibiting tumor suppressors are classified as an onco-miRs. Therefore, there are two possible miRNA-based strategies: therapeutic inhibition (in case of onco-miRs) or substitution (in case of TS-miRNAs) (Fig. 4.7).
Specific inhibitory effect of miRNAs can be achieved by use of their complementary antagonists (anti-miRs). Whereas in the most cancers tumor tissues indicate global decrease in miRNA expression levels, therapeutic substitution presents the most promising approach. So far, many studies have described oncogenic/tumor-suppressive miRNAs and their contribution to development and progression of gliomas. Most of these studies were performed under in vitro conditions (Li et al. 2013b). In vitro models are suitable for studies on specific targets and regulatory pathways; however, to evaluate the real therapeutic potential, it is necessary to study miRNAs on animal models. MiRNAs in vivo studies are mainly focused on the evaluation of particular miRNA effects on the tumor growth and confirmation of phenotypic observations from in vitro studies.
It is presumed that GSCs may stand for the cells responsible for tumor formation and treatment failure. Notch signaling is important for maintaining this stem-like phenotype and has been found to be aberrantly expressed in GSCs. Inhibition of this signaling pathway decreases proliferation, increases neuronal differentiation, and reduces CD133+ cell fraction in vitro; moreover, in vivo such inhibition attenuates tumorigenicity (Stockhausen et al. 2010). MiR-34a is often described as direct regulator of Notch signaling and could serve as potential tumor suppressor in brain tumors. This was demonstrated by transfection of pre-miR-34a to cells, which were consequently implanted to immunocompromised mice. Four weeks after implantation, xenografts of pre-miR-34a-transfected cells developed tumor of significantly smaller volume compared to control cells (Li et al. 2009a, 2011). Moreover, other miRNAs were confirmed as direct regulators of Notch signaling, such as miR-326 and miR-107. Similarly to miR-34a, exogenous expression of these miRNAs in glioma cells caused decrease of the tumor bearing in mice (Kefas et al. 2009; Chen et al. 2013a). Inhibition of NOTCH can be also achieved by miR-92b. It was shown that this miRNA regulates NLK (Nemo-like kinase), which negatively regulates NOTCH--dependent transcriptional activation. Furthermore, expression of NLK was inversely correlated with miR-92b in clinical glioma samples and enables prediction of clinical outcome. An effect of miR-92b on glioma tumors was evaluated in vivo by using nude mice. Glioma cell lines were subcutaneously injected to mice and after reaching 100 mm3 tumor volumes were animals treated with anti-miR-92b oligonucleotide. Anti-miR-92b triggered growth inhibition, induced apoptosis, and suppressed invasion of glioma (Ishitani et al. 2010; Wang et al. 2013a).
Consistently, miR-204 was found to be downregulated in glioma cells and also in neural stem cells. Inhibition of miR-204 promoted not only cellular stemness, but also the biological properties essential to lethality of the disease; however, restoration of miR-204 greatly abrogates the aggressiveness of glioma cells. This observation is supported by Kaplan–Meier survival analysis of mice transduced by miR-204 compared to control. MiR-204 prolonged the survival of mice by approx. 15 %. This is probably caused by miR-204 targeting of SOX4 gene, which is the core regulator governing the stemness of both glioma and neural stem cells (Ying et al. 2013).
Aggressiveness of stem-like cells and ability to form new in situ tumors is closely related to proliferation and invasive ability. Receptor tyrosine kinases such as EGFR and PDGFR play a major role in glioma proliferation and invasion (Nakada et al. 2013). In gliomas, regulation of these receptors by miRNAs was observed in many in vitro studies, which were consequently confirmed in animal models. It was shown that miR-128 represses GSC growth by enhancing neuronal differentiation and mediates differentiation by targeting EGFR and PDGFRα. Authors of this study proved this by utilization and autochthonous glioma mouse model, which arises from activation of oncogenic H-RasV12 (Harvey-RasV12) and loss of p53. Developed glioma tumors in mice showed significant decrease of miR-128 and high levels of PDGFRα compared to normal brain tissue. The introduction of miR-128 led to remarkable improvement of survival in treated animals. This study validated miR-128 as effective suppressor of glioma growth (Papagiannakopoulos et al. 2012).
Another interesting example of onco-miR is miR-93. It enhances tumor cell survival, blood vessel expansion, and tumor growth, by targeting, at least in part, integrin-β8 expression. Consistent with this was the decreased survival rate of mice bearing the miR-93 positive tumors compared with the controls and Kaplan–Maier survival curves which indicated that miR-93 expression decreased rates of mouse survival. Further, higher levels of integrin-β8 are associated with cell death in tumor mass and in human glioblastoma (Fang et al. 2011). Accelerated proliferation is a characteristic for tumor cells and contributes to excessive growth and tumor progression. PDCD4 is a well-known tumor-suppressor gene. Overexpression of PDCD4 caused by downregulation of miR-21 led to decreased proliferation and apoptotic rate and in the xenograft model decreased tumor formation and growth (Gaur et al. 2011b). Clinical relevance of miR-21 was further studied by Xuan Zhou et al. In the in vivo pharmacological study 30 mice were included and screened for tumor volume of xenografts treated with PBS, negative control, and anti-sense-miR-21 (as-miR-21). Significant decrease in tumor volume was observed only in as-miR-21-treated group. These data suggest miR-21 as a promising therapeutic target for malignant gliomas (Zhou et al. 2010a).
Even if miRNA-based therapies were not clinically implemented to the therapy, they have several important advantages to conventional drugs and targeted therapy in oncology. MiRNAs have ability to target multiple genes, which give them a priority in cancer treatment, considering cancer as a heterogeneous disease with multiple molecular alterations. In addition, miRNAs are not detected by immune system and so they could be an attractive option for clinical drug development.
Conclusions and Future Perspectives
The discovery of miRNA function has markedly spread the view on regulation of gene expression. Its remarkable ability to regulate large number of genes, including oncogenes and tumor-suppressor genes, has catapulted miRNAs into the center of cancer molecular biology over the past 5 years. It is now evident that dysregulation of miRNAs is an important step in the development of many cancers, including gliomas. Several studies based on expression profiling have proved that there are significant changes of miRNA expression levels in gliomas compared to non-tumor brain tissue. These studies also identified groups of miRNAs with potential for prognostic stratification and prediction of responses to chemoradiotherapy in glioma patients. But much more studies have been focused on the improvement of our knowledge about miRNAs involvement in glioma core signaling pathways. The results of these studies suggest a great potential and relevance of miRNAs as a novel class of therapeutic targets and possibly powerful intervention tools in glioma.
Abbreviations
- BMI1:
-
BMI1 polycomb ring finger oncogene
- Cdk:
-
Cyclin-dependent kinase
- c-MYC:
-
v-myc avian myelocytomatosis viral oncogene homolog
- EGFR:
-
Epidermal growth factor receptor
- GBM:
-
Glioblastoma multiforme
- GSC:
-
Glioma stem cell
- IFN:
-
Interferon
- IKKα/β/γ:
-
Conserved helix-loop-helix ubiquitous kinase α/β/γ
- IRS:
-
Insulin receptor substrate
- JAK-STAT:
-
Janus kinase-signal transducer and activator of transcription
- miRNA:
-
MicroRNA
- MMP:
-
Matrix metalloproteinase
- NF-κB:
-
Nuclear factor of kappa light polypeptide gene enhancer in B-cells
- p21CIP1:
-
Cyclin-dependent kinase inhibitor 1A
- p27:
-
p27 protein
- PDGFR:
-
Platelet-derived growth factor receptor
- PI3K/AKT:
-
Phosphatidylinositol-4,5-bisphosphate 3-kinase/v-akt murine thymoma viral oncogene homolog)
- PLD2:
-
Phospholipase D2
- PTEN:
-
Phosphatase and tensin homolog
- RAS:
-
Rat sarcoma viral oncogene homolog
- TMZ:
-
Temozolomide
- VEGF:
-
Vascular endothelial cell growth factor
- VEGFR:
-
VEGF receptor
- WHO:
-
World Health Organization
References
Aldaz B, Sagardoy A, Nogueira L, Guruceaga E, Grande L, Huse JT et al (2013) Involvement of miRNAs in the differentiation of human glioblastoma multiforme stem-like cells. PLoS One 8:e77098
Alrfaei BM, Vemuganti R, Kuo JS (2013) microRNA-100 targets SMRT/NCOR2, reduces proliferation, and improves survival in glioblastoma animal models. PLoS One 8:e80865
Altaner C (2008) Glioblastoma and stem cells. Neoplasma 55:369–374
Asuthkar S, Velpula KK, Chetty C, Gorantla B, Rao JS (2012) Epigenetic regulation of miRNA-211 by MMP-9 governs glioma cell apoptosis, chemosensitivity and radiosensitivity. Oncotarget 3:1439–1454
Babar IA, Czochor J, Steinmetz A, Weidhaas JB, Glazer PM, Slack FJ (2011) Inhibition of hypoxia-induced miR-155 radiosensitizes hypoxic lung cancer cells. Cancer Biol Ther 12:908–914
Besse A, Sana J, Fadrus P, Slaby O (2013) MicroRNAs involved in chemo- and radioresistance of high-grade gliomas. Tumour Biol 34(4):1969–1978
Bienertova-Vasku J, Sana J, Slaby O (2013) The role of microRNAs in mitochondria in cancer. Cancer Lett 336:1–7
Cao L, Kim S, Xiao C, Wang RH, Coumoul X, Wang X et al (2006) ATM-Chk2-p53 activation prevents tumorigenesis at an expense of organ homeostasis upon Brca1 deficiency. EMBO J 25:2167–2177
Card DA, Hebbar PB, Li L, Trotter KW, Komatsu Y, Mishina Y et al (2008) Oct4/Sox2-regulated miR-302 targets cyclin D1 in human embryonic stem cells. Mol Cell Biol 28:6426–6438
Carmo A, Carvalheiro H, Crespo I, Nunes I, Lopes MC (2011) Effect of temozolomide on the U-118 glioma cell line. Oncol Lett 2:1165–1170
Carthew RW, Sontheimer EJ (2009) Origins and mechanisms of miRNAs and siRNAs. Cell 136:642–655
Ciafre SA, Galardi S, Mangiola A, Ferracin M, Liu CG, Sabatino G et al (2005) Extensive modulation of a set of microRNAs in primary glioblastoma. Biochem Biophys Res Commun 334:1351–1358
Conti A, Aguennouz M, La Torre D, Tomasello C, Cardali S, Angileri FF et al (2009) miR-21 and 221 upregulation and miR-181b downregulation in human grade II-IV astrocytic tumors. J Neurooncol 93:325–332
Costa PM, Cardoso AL, Nobrega C, Pereira de Almeida LF, Bruce JN, Canoll P et al (2013) MicroRNA-21 silencing enhances the cytotoxic effect of the antiangiogenic drug sunitinib in glioblastoma. Hum Mol Genet 22:904–918
D’Urso PI, D’Urso OF, Storelli C, Mallardo M, Gianfreda CD, Montinaro A et al (2012) miR-155 is up-regulated in primary and secondary glioblastoma and promotes tumour growth by inhibiting GABA receptors. Int J Oncol 41:228–234
Darkes MJM, Plosker GL, Jarvis B (2002) Temozolomide: a review of its use in the treatment of malignant gliomas, malignant melanoma and other advanced cancers. Am J Cancer 1:55–80
Dong CG, Wu WK, Feng SY, Wang XJ, Shao JF, Qiao J (2012) Co-inhibition of microRNA-10b and microRNA-21 exerts synergistic inhibition on the proliferation and invasion of human glioma cells. Int J Oncol 41:1005–1012
Dontula R, Dinasarapu A, Chetty C, Pannuru P, Herbert E, Ozer H et al (2013) MicroRNA 203 modulates glioma cell migration via Robo1/ERK/MMP-9 signaling. Genes Cancer 4:285–296
Ernst A, Campos B, Meier J, Devens F, Liesenberg F, Wolter M et al (2010) De-repression of CTGF via the miR-17-92 cluster upon differentiation of human glioblastoma spheroid cultures. Oncogene 29:3411–3422
Fang L, Deng Z, Shatseva T, Yang J, Peng C, Du WW et al (2011) MicroRNA miR-93 promotes tumor growth and angiogenesis by targeting integrin-beta8. Oncogene 30:806–821
Fareh M, Turchi L, Virolle V, Debruyne D, Almairac F, de-la-Forest Divonne S et al (2012) The miR 302-367 cluster drastically affects self-renewal and infiltration properties of glioma-initiating cells through CXCR4 repression and consequent disruption of the SHH-GLI-NANOG network. Cell Death Differ 19:232–244
Fell VL, Schild-Poulter C (2012) Ku regulates signaling to DNA damage response pathways through the Ku70 von Willebrand A domain. Mol Cell Biol 32:76–87
Folkman J (2006) Angiogenesis. Annu Rev Med 57:1–18
Fowler A, Thomson D, Giles K, Maleki S, Mreich E, Wheeler H et al (2011) miR-124a is frequently down-regulated in glioblastoma and is involved in migration and invasion. Eur J Cancer 47:953–963
Fu J, Rodova M, Nanta R, Meeker D, Van Veldhuizen PJ, Srivastava RK et al (2013) NPV-LDE-225 (Erismodegib) inhibits epithelial mesenchymal transition and self-renewal of glioblastoma initiating cells by regulating miR-21, miR-128, and miR-200. Neuro Oncol 15:691–706
Gabriely G, Wurdinger T, Kesari S, Esau CC, Burchard J, Linsley PS et al (2008) MicroRNA 21 promotes glioma invasion by targeting matrix metalloproteinase regulators. Mol Cell Biol 28:5369–5380
Gabriely G, Yi M, Narayan RS, Niers JM, Wurdinger T, Imitola J et al (2011) Human glioma growth is controlled by microRNA-10b. Cancer Res 71:3563–3572
Gal H, Pandi G, Kanner AA, Ram Z, Lithwick-Yanai G, Amariglio N et al (2008) MIR-451 and Imatinib mesylate inhibit tumor growth of Glioblastoma stem cells. Biochem Biophys Res Commun 376:86–90
Gao H, Zhao H, Xiang W (2013) Expression level of human miR-34a correlates with glioma grade and prognosis. J Neurooncol 113:221–228
Gaur AB, Holbeck SL, Colburn NH, Israel MA (2011) Downregulation of Pdcd4 by mir-21 facilitates glioblastoma proliferation in vivo. Neuro Oncol 13:580–590
Genovese G, Ergun A, Shukla SA, Campos B, Hanna J, Ghosh P et al (2012) microRNA regulatory network inference identifies miR-34a as a novel regulator of TGF-beta signaling in glioblastoma. Cancer Discov 2:736–749
Geretti E, Klagsbrun M (2007) Neuropilins: novel targets for anti-angiogenesis therapies. Cell Adh Migr 1:56–61
Ghosh S, Karin M (2002) Missing pieces in the NF-kappaB puzzle. Cell 109(Suppl):S81–S96
Godlewski J, Nowicki MO, Bronisz A, Williams S, Otsuki A, Nuovo G et al (2008) Targeting of the Bmi-1 oncogene/stem cell renewal factor by microRNA-128 inhibits glioma proliferation and self-renewal. Cancer Res 68:9125–9130
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res 36:D154–D158
Guan Y, Mizoguchi M, Yoshimoto K, Hata N, Shono T, Suzuki SO et al (2010) MiRNA-196 is upregulated in glioblastoma but not in anaplastic astrocytoma and has prognostic significance. Clin Cancer Res 16:4289–4297
Guessous F, Alvarado-Velez M, Marcinkiewicz L, Zhang Y, Kim J, Heister S et al (2013) Oncogenic effects of miR-10b in glioblastoma stem cells. J Neurooncol 112:153–163
Guessous F, Zhang Y, Kofman A, Catania A, Li Y, Schiff D et al (2010) microRNA-34a is tumor suppressive in brain tumors and glioma stem cells. Cell Cycle (Georgetown, TX) 9:1031–1036
Guillamo JS, de Bouard S, Valable S, Marteau L, Leuraud P, Marie Y et al (2009) Molecular mechanisms underlying effects of epidermal growth factor receptor inhibition on invasion, proliferation, and angiogenesis in experimental glioma. Clin Cancer Res 15:3697–3704
Guo P, Nie Q, Lan J, Ge J, Qiu Y, Mao Q (2013) C-Myc negatively controls the tumor suppressor PTEN by upregulating miR-26a in glioblastoma multiforme cells. Biochem Biophys Res Commun 441:186–190
Gwak HS, Kim TH, Jo GH, Kim YJ, Kwak HJ, Kim JH et al (2012) Silencing of microRNA-21 confers radio-sensitivity through inhibition of the PI3K/AKT pathway and enhancing autophagy in malignant glioma cell lines. PLoS One 7:e47449
Haapa-Paananen S, Chen P, Hellstrom K, Kohonen P, Hautaniemi S, Kallioniemi O et al (2013) Functional profiling of precursor MicroRNAs identifies MicroRNAs essential for glioma proliferation. PLoS One 8:e60930
Hao J, Zhang C, Zhang A, Wang K, Jia Z, Wang G et al (2012) miR-221/222 is the regulator of Cx43 expression in human glioblastoma cells. Oncol Rep 27:1504–1510
Hatanpaa KJ, Burma S, Zhao D, Habib AA (2010) Epidermal growth factor receptor in glioma: signal transduction, neuropathology, imaging, and radioresistance. Neoplasia 12:675–684
He J, Deng Y, Yang G, Xie W (2013) MicroRNA-203 down-regulation is associated with unfavorable prognosis in human glioma. J Surg Oncol 108:121–125
Hermansen SK, Dahlrot RH, Nielsen BS, Hansen S, Kristensen BW (2013) MiR-21 expression in the tumor cell compartment holds unfavorable prognostic value in gliomas. J Neurooncol 111:71–81
Hermansen SK, Kristensen BW (2013) MicroRNA biomarkers in glioblastoma. J Neurooncol 114:13–23
Hong L, Han Y, Zhang Y, Zhang H, Zhao Q, Wu K et al (2013) MicroRNA-21: a therapeutic target for reversing drug resistance in cancer. Expert Opin Ther Targets 17:1073–1080
Hua D, Ding D, Han X, Zhang W, Zhao N, Foltz G et al (2012) Human miR-31 targets radixin and inhibits migration and invasion of glioma cells. Oncol Rep 27:700–706
Huang PH, Xu AM, White FM (2009) Oncogenic EGFR signaling networks in glioma. Sci Signal 2:re6
Huse JT, Brennan C, Hambardzumyan D, Wee B, Pena J, Rouhanifard SH et al (2009) The PTEN-regulating microRNA miR-26a is amplified in high-grade glioma and facilitates gliomagenesis in vivo. Genes Dev 23:1327–1337
Chakravarti A, Zhai G, Suzuki Y, Sarkesh S, Black PM, Muzikansky A et al (2004) The prognostic significance of phosphatidylinositol 3-kinase pathway activation in human gliomas. J Clin Oncol 22:1926–1933
Chamberlain MC (2011) Bevacizumab for the treatment of recurrent glioblastoma. Clin Med Insights Oncol 5:117–129
Chan JA, Krichevsky AM, Kosik KS (2005) MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 65:6029–6033
Chan XH, Nama S, Gopal F, Rizk P, Ramasamy S, Sundaram G et al (2012) Targeting glioma stem cells by functional inhibition of a prosurvival oncomiR-138 in malignant gliomas. Cell Rep 2:591–602
Chang C, Shi H, Wang C, Wang J, Geng N, Jiang X et al (2012) Correlation of microRNA-375 downregulation with unfavorable clinical outcome of patients with glioma. Neurosci Lett 531:204–208
Chaudhry MA, Kreger B, Omaruddin RA (2010a) Transcriptional modulation of micro-RNA in human cells differing in radiation sensitivity. Int J Radiat Biol 86:569–583
Chaudhry MA, Sachdeva H, Omaruddin RA (2010b) Radiation-induced micro-RNA modulation in glioblastoma cells differing in DNA-repair pathways. DNA Cell Biol 29:553–561
Chen C-Z (2005) MicroRNAs as oncogenes and tumor suppressors. New Engl J Med 353:1768–1771
Chen G, Zhu W, Shi D, Lv L, Zhang C, Liu P et al (2010a) MicroRNA-181a sensitizes human malignant glioma U87MG cells to radiation by targeting Bcl-2. Oncol Rep 23:997–1003
Chen L, Chen XR, Chen FF, Liu Y, Li P, Zhang R et al (2013a) MicroRNA-107 inhibits U87 glioma stem cells growth and invasion. Cell Mol Neurobiol 33:651–657
Chen L, Zhang J, Han L, Zhang A, Zhang C, Zheng Y et al (2012) Downregulation of miR-221/222 sensitizes glioma cells to temozolomide by regulating apoptosis independently of p53 status. Oncol Rep 27:854–860
Chen L, Zhang R, Li P, Liu Y, Qin K, Fa ZQ et al (2013b) P53-induced microRNA-107 inhibits proliferation of glioma cells and down-regulates the expression of CDK6 and Notch-2. Neurosci Lett 534:327–332
Chen R, Nishimura MC, Bumbaca SM, Kharbanda S, Forrest WF, Kasman IM et al (2010b) A hierarchy of self-renewing tumor-initiating cell types in glioblastoma. Cancer Cell 17:362–375
Chen SM, Chen HC, Chen SJ, Huang CY, Chen PY, Wu TW et al (2013c) MicroRNA-495 inhibits proliferation of glioblastoma multiforme cells by downregulating cyclin-dependent kinase 6. World J Surg Oncol 11:87
Chen Y, Liu W, Chao T, Zhang Y, Yan X, Gong Y et al (2008) MicroRNA-21 down-regulates the expression of tumor suppressor PDCD4 in human glioblastoma cell T98G. Cancer Lett 272:197–205
Chen Z, Li D, Cheng Q, Ma Z, Jiang B, Peng R et al (2013d) MicroRNA-203 inhibits the proliferation and invasion of U251 glioblastoma cells by directly targeting PLD2. Mol Med Rep 9(2):503–508
Ishitani T, Hirao T, Suzuki M, Isoda M, Ishitani S, Harigaya K et al (2010) Nemo-like kinase suppresses Notch signalling by interfering with formation of the Notch active transcriptional complex. Nat Cell Biol 12:278–285
Jia Z, Wang K, Zhang A, Wang G, Kang C, Han L et al (2013) miR-19a and miR-19b overexpression in gliomas. Pathol Oncol Res 19:847–853
Jiang J, Sun X, Wang W, Jin X, Bo X, Li Z et al (2012) Tumor microRNA-335 expression is associated with poor prognosis in human glioma. Med Oncol 29:3472–3477
Jiang J, Yang J, Wang Z, Wu G, Liu F (2013) TFAM is directly regulated by miR-23b in glioma. Oncol Rep 30:2105–2110
Kao GD, Jiang Z, Fernandes AM, Gupta AK, Maity A (2007) Inhibition of phosphatidylinositol-3-OH kinase/Akt signaling impairs DNA repair in glioblastoma cells following ionizing radiation. J Biol Chem 282:21206–21212
Kefas B, Comeau L, Erdle N, Montgomery E, Amos S, Purow B (2010) Pyruvate kinase M2 is a target of the tumor-suppressive microRNA-326 and regulates the survival of glioma cells. Neuro Oncol 12:1102–1112
Kefas B, Comeau L, Floyd DH, Seleverstov O, Godlewski J, Schmittgen T et al (2009) The neuronal microRNA miR-326 acts in a feedback loop with notch and has therapeutic potential against brain tumors. J Neurosci 29:15161–15168
Kefas B, Godlewski J, Comeau L, Li Y, Abounader R, Hawkinson M et al (2008) microRNA-7 inhibits the epidermal growth factor receptor and the Akt pathway and is down-regulated in glioblastoma. Cancer Res 68:3566–3572
Kim H, Huang W, Jiang X, Pennicooke B, Park PJ, Johnson MD (2010) Integrative genome analysis reveals an oncomir/oncogene cluster regulating glioblastoma survivorship. Proc Natl Acad Sci U S A 107:2183–2188
Koo S, Martin GS, Schulz KJ, Ronck M, Toussaint LG (2012) Serial selection for invasiveness increases expression of miR-143/miR-145 in glioblastoma cell lines. BMC Cancer 12:143
Koshkin PA, Chistiakov DA, Chekhonin VP (2013) Role of microRNAs in mechanisms of glioblastoma resistance to radio- and chemotherapy. Biochemistry (Mosc) 78:325–334
Kozomara A, Griffiths-Jones S (2011) miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 39:D152–D157
Kwiatkowska A, Symons M (2013) Signaling determinants of glioma cell invasion. Adv Exp Med Biol 986:121–141
Ladomery MR, Maddocks DG, Wilson ID (2011) MicroRNAs: their discovery, biogenesis, function and potential use as biomarkers in non-invasive prenatal diagnostics. Int J Mol Epidemiol Genet 2:253–260
Lakomy R, Sana J, Hankeova S, Fadrus P, Kren L, Lzicarova E et al (2011) MiR-195, miR-196b, miR-181c, miR-21 expression levels and O-6-methylguanine-DNA methyltransferase methylation status are associated with clinical outcome in glioblastoma patients. Cancer Sci 102:2186–2190
Lawler S, Chiocca EA (2009) Emerging functions of microRNAs in glioblastoma. J Neurooncol 92:297–306
Le MT, Xie H, Zhou B, Chia PH, Rizk P, Um M et al (2009) MicroRNA-125b promotes neuronal differentiation in human cells by repressing multiple targets. Mol Cell Biol 29:5290–5305
Lee KM, Choi EJ, Kim IA (2011a) microRNA-7 increases radiosensitivity of human cancer cells with activated EGFR-associated signaling. Radiother Oncol 101:171–176
Lee ST, Chu K, Oh HJ, Im WS, Lim JY, Kim SK et al (2011b) Let-7 microRNA inhibits the proliferation of human glioblastoma cells. J Neurooncol 102:19–24
Li G, Zhang Z, Tu Y, Jin T, Liang H, Cui G et al (2013a) Correlation of microRNA-372 upregulation with poor prognosis in human glioma. Diagn Pathol 8:1
Li M, Li J, Liu L, Li W, Yang Y, Yuan J (2013b) MicroRNA in human glioma. Cancers (Basel) 5:1306–1331
Li W-Q, Li Y-M, Tao B-B, Lu Y-C, Hu G-H, Liu H-M et al (2010) Downregulation of ABCG2 expression in glioblastoma cancer stem cells with miRNA-328 may decrease their chemoresistance. Med Sci Monit 16:HY27–HY30
Li WB, Ma MW, Dong LJ, Wang F, Chen LX, Li XR (2011) MicroRNA-34a targets notch1 and inhibits cell proliferation in glioblastoma multiforme. Cancer Biol Ther 12:477–483
Li Y, Guessous F, Zhang Y, Dipierro C, Kefas B, Johnson E et al (2009a) MicroRNA-34a inhibits glioblastoma growth by targeting multiple oncogenes. Cancer Res 69:7569–7576
Li Y, Li W, Yang Y, Lu Y, He C, Hu G et al (2009b) MicroRNA-21 targets LRRFIP1 and contributes to VM-26 resistance in glioblastoma multiforme. Brain Res 1286:13–18
Li Y, Xu J, Chen H, Bai J, Li S, Zhao Z et al (2013c) Comprehensive analysis of the functional microRNA-mRNA regulatory network identifies miRNA signatures associated with glioma malignant progression. Nucleic Acids Res 41(22):e203
Lin J, Teo S, Lam DH, Jeyaseelan K, Wang S (2012) MicroRNA-10b pleiotropically regulates invasion, angiogenicity and apoptosis of tumor cells resembling mesenchymal subtype of glioblastoma multiforme. Cell Death Dis 3:e398
Lin YX, Yu F, Gao N, Sheng JP, Qiu JZ, Hu BC (2011) microRNA-143 protects cells from DNA damage-induced killing by downregulating FHIT expression. Cancer Biother Radiopharm 26:365–372
Ling N, Gu J, Lei Z, Li M, Zhao J, Zhang HT et al (2013) microRNA-155 regulates cell proliferation and invasion by targeting FOXO3a in glioma. Oncol Rep 30:2111–2118
Liu L, Chen L, Xu Y, Li R, Du X (2010) microRNA-195 promotes apoptosis and suppresses tumorigenicity of human colorectal cancer cells. Biochem Biophys Res Commun 400:236–240
Lu S, Wang S, Geng S, Ma S, Liang Z, Jiao B (2012) Increased expression of microRNA-17 predicts poor prognosis in human glioma. J Biomed Biotechnol 2012:970761
Lu S, Wang S, Geng S, Ma S, Liang Z, Jiao B (2013) Upregulation of microRNA-224 confers a poor prognosis in glioma patients. Clin Transl Oncol 15:569–574
Luan S, Sun L, Huang F (2010) MicroRNA-34a: a novel tumor suppressor in p53-mutant glioma cell line U251. Arch Med Res 41:67–74
Malzkorn B, Wolter M, Liesenberg F, Grzendowski M, Stuhler K, Meyer HE et al (2010) Identification and functional characterization of microRNAs involved in the malignant progression of gliomas. Brain Pathol 20:539–550
Martin V, Sanchez-Sanchez AM, Herrera F, Gomez-Manzano C, Fueyo J, Alvarez-Vega MA et al (2013) Melatonin-induced methylation of the ABCG2/BCRP promoter as a novel mechanism to overcome multidrug resistance in brain tumour stem cells. Br J Cancer 108:2005–2012
Meng W, Jiang L, Lu L, Hu H, Yu H, Ding D et al (2012) Anti-miR-155 oligonucleotide enhances chemosensitivity of U251 cell to taxol by inducing apoptosis. Cell Biol Int 36:653–659
Mizoguchi M, Guan Y, Yoshimoto K, Hata N, Amano T, Nakamizo A et al (2012) MicroRNAs in human malignant gliomas. J Oncol 2012:732874
Mueller AC, Sun D, Dutta A (2013) The miR-99 family regulates the DNA damage response through its target SNF2H. Oncogene 32(9):1164–1172
Nakada M, Kita D, Teng L, Pyko IV, Watanabe T, Hayashi Y et al (2013) Receptor tyrosine kinases: principles and functions in glioma invasion. Adv Exp Med Biol 986:143–170
Nakano I, Kornblum HI (2006) Brain tumor stem cells. Pediatr Res 59:54R–58R
Nan Y, Han L, Zhang A, Wang G, Jia Z, Yang Y et al (2010) MiRNA-451 plays a role as tumor suppressor in human glioma cells. Brain Res 1359:14–21
Ng WL, Yan D, Zhang X, Mo YY, Wang Y (2010) Over-expression of miR-100 is responsible for the low-expression of ATM in the human glioma cell line: M059J. DNA Repair (Amst) 9:1170–1175
Nikaki A, Piperi C, Papavassiliou AG (2012) Role of microRNAs in gliomagenesis: targeting miRNAs in glioblastoma multiforme therapy. Expert Opin Investig Drugs 21:1475–1488
Niyazi M, Zehentmayr F, Niemoller OM, Eigenbrod S, Kretzschmar H, Schulze-Osthoff K et al (2011) MiRNA expression patterns predict survival in glioblastoma. Radiat Oncol 6:153
Ohno M, Natsume A, Kondo Y, Iwamizu H, Motomura K, Toda H et al (2009) The modulation of microRNAs by type I IFN through the activation of signal transducers and activators of transcription 3 in human glioma. Mol Cancer Res 7:2022–2030
Pan SJ, Zhan SK, Pei BG, Sun QF, Bian LG, Sun BM (2012) MicroRNA-149 inhibits proliferation and invasion of glioma cells via blockade of AKT1 signaling. Int J Immunopathol Pharmacol 25:871–881
Papagiannakopoulos T, Friedmann-Morvinski D, Neveu P, Dugas JC, Gill RM, Huillard E et al (2012) Pro-neural miR-128 is a glioma tumor suppressor that targets mitogenic kinases. Oncogene 31:1884–1895
Papagiannakopoulos T, Shapiro A, Kosik KS (2008) MicroRNA-21 targets a network of key tumor-suppressive pathways in glioblastoma cells. Cancer Res 68:8164–8172
Platanias LC (2005) Mechanisms of type-I- and type-II-interferon-mediated signalling. Nat Rev Immunol 5:375–386
Qian X, Zhao P, Li W, Shi ZM, Wang L, Xu Q et al (2013) MicroRNA-26a promotes tumor growth and angiogenesis in glioma by directly targeting prohibitin. CNS Neurosci Ther 19:804–812
Qiu S, Huang D, Yin D, Li F, Li X, Kung HF et al (2013) Suppression of tumorigenicity by microRNA-138 through inhibition of EZH2-CDK4/6-pRb-E2F1 signal loop in glioblastoma multiforme. Biochim Biophys Acta 1832:1697–1707
Qu S, Yao Y, Shang C, Xue Y, Ma J, Li Z et al (2012) MicroRNA-330 is an oncogenic factor in glioblastoma cells by regulating SH3GL2 gene. PLoS One 7:e46010
Rao SA, Santosh V, Somasundaram K (2010) Genome-wide expression profiling identifies deregulated miRNAs in malignant astrocytoma. Mod Pathol 23:1404–1417
Ren Y, Kang CS, Yuan XB, Zhou X, Xu P, Han L et al (2010a) Co-delivery of as-miR-21 and 5-FU by poly(amidoamine) dendrimer attenuates human glioma cell growth in vitro. J Biomater Sci Polym Ed 21:303–314
Ren Y, Zhou X, Mei M, Yuan XB, Han L, Wang GX et al (2010b) MicroRNA-21 inhibitor sensitizes human glioblastoma cells U251 (PTEN-mutant) and LN229 (PTEN-wild type) to taxol. BMC Cancer 10:27
Sana J, Hajduch M, Michalek J, Vyzula R, Slaby O (2011) MicroRNAs and glioblastoma: roles in core signalling pathways and potential clinical implications. J Cell Mol Med 15:1636–1644
Sasaki A, Udaka Y, Tsunoda Y, Yamamoto G, Tsuji M, Oyamada H et al (2012) Analysis of p53 and miRNA expression after irradiation of glioblastoma cell lines. Anticancer Res 32:4709–4713
Sasayama T, Nishihara M, Kondoh T, Hosoda K, Kohmura E (2009) MicroRNA-10b is overexpressed in malignant glioma and associated with tumor invasive factors, uPAR and RhoC. Int J Cancer 125:1407–1413
Sharma S, Salehi F, Scheithauer BW, Rotondo F, Syro LV, Kovacs K (2009) Role of MGMT in tumor development, progression, diagnosis, treatment and prognosis. Anticancer Res 29:3759–3768
Shi L, Chen J, Yang J, Pan T, Zhang S, Wang Z (2010) MiR-21 protected human glioblastoma U87MG cells from chemotherapeutic drug temozolomide induced apoptosis by decreasing Bax/Bcl-2 ratio and caspase-3 activity. Brain Res 1352:255–264
Shi L, Zhang S, Feng K, Wu F, Wan Y, Wang Z, Zhang J, Wang Y, Yan W, Fu Z, You Y (2012) MicroRNA-125b-2 confers human glioblastoma stem cells resistance to temozolomide through the mitochondrial pathway of apoptosis. Int J Oncol 40:119–129
Shi L, Wan Y, Sun G, Zhang S, Wang Z, Zeng Y (2013) miR-125b inhibitor may enhance the invasion-prevention activity of temozolomide in glioblastoma stem cells by targeting PIAS3. BioDrugs 28(1):41–54
Schraivogel D, Weinmann L, Beier D, Tabatabai G, Eichner A, Zhu JY et al (2011) CAMTA1 is a novel tumour suppressor regulated by miR-9/9* in glioblastoma stem cells. EMBO J 30:4309–4322
Schramedei K, Morbt N, Pfeifer G, Lauter J, Rosolowski M, Tomm JM et al (2011) MicroRNA-21 targets tumor suppressor genes ANP32A and SMARCA4. Oncogene 30:2975–2985
Schwarz DS, Hutvagner G, Du T, Xu Z, Aronin N, Zamore PD (2003) Asymmetry in the assembly of the RNAi enzyme complex. Cell 115:199–208
Siddique HR, Saleem M (2012) Role of BMI1, a stem cell factor, in cancer recurrence and chemoresistance: preclinical and clinical evidences. Stem Cells 30:372–378
Silber J, Lim DA, Petritsch C, Persson AI, Maunakea AK, Yu M et al (2008) miR-124 and miR-137 inhibit proliferation of glioblastoma multiforme cells and induce differentiation of brain tumor stem cells. BMC Med 6:14
Skalsky RL, Cullen BR (2011) Reduced expression of brain-enriched microRNAs in glioblastomas permits targeted regulation of a cell death gene. PLoS One 6:e24248
Slaby O, Lakomy R, Fadrus P, Hrstka R, Kren L, Lzicarova E et al (2010) MicroRNA-181 family predicts response to concomitant chemoradiotherapy with temozolomide in glioblastoma patients. Neoplasma 57:264–269
Song L, Huang Q, Chen K, Liu L, Lin C, Dai T et al (2010) miR-218 inhibits the invasive ability of glioma cells by direct downregulation of IKK-beta. Biochem Biophys Res Commun 402:135–140
Soon PS, Tacon LJ, Gill AJ, Bambach CP, Sywak MS, Campbell PR et al (2009) miR-195 and miR-483-5p Identified as Predictors of Poor Prognosis in Adrenocortical Cancer. Clin Cancer Res 15:7684–7692
Squatrito M, Brennan CW, Helmy K, Huse JT, Petrini JH, Holland EC (2010) Loss of ATM/Chk2/p53 pathway components accelerates tumor development and contributes to radiation resistance in gliomas. Cancer Cell 18:619–629
Srinivasan S, Patric IR, Somasundaram K (2011) A ten-microRNA expression signature predicts survival in glioblastoma. PLoS One 6:e17438
Sternlicht MD, Lochter A, Sympson CJ, Huey B, Rougier JP, Gray JW et al (1999) The stromal proteinase MMP3/stromelysin-1 promotes mammary carcinogenesis. Cell 98:137–146
Stockhausen MT, Kristoffersen K, Poulsen HS (2010) The functional role of Notch signaling in human gliomas. Neuro Oncol 12:199–211
Sun B, Pu B, Chu D, Chu X, Li W, Wei D (2013) MicroRNA-650 expression in glioma is associated with prognosis of patients. J Neurooncol 115:375–380
Ujifuku K, Mitsutake N, Takakura S, Matsuse M, Saenko V, Suzuki K et al (2010) miR-195, miR-455-3p and miR-10a( *) are implicated in acquired temozolomide resistance in glioblastoma multiforme cells. Cancer Lett 296:241–248
Ulasov IV, Nandi S, Dey M, Sonabend AM, Lesniak MS (2011) Inhibition of Sonic hedgehog and Notch pathways enhances sensitivity of CD133(+) glioma stem cells to temozolomide therapy. Mol Med 17:103–112
Wang J, Li Y, Wang X, Jiang C (2012a) Ursolic acid inhibits proliferation and induces apoptosis in human glioblastoma cell lines U251 by suppressing TGF-beta1/miR-21/PDCD4 pathway. Basic Clin Pharmacol Toxicol 111:106–112
Wang J, Wakeman TP, Lathia JD, Hjelmeland AB, Wang XF, White RR et al (2010) Notch promotes radioresistance of glioma stem cells. Stem Cells 28:17–28
Wang K, Wang X, Zou J, Zhang A, Wan Y, Pu P et al (2013a) miR-92b controls glioma proliferation and invasion through regulating Wnt/beta-catenin signaling via Nemo-like kinase. Neuro Oncol 15:578–588
Wang L, Shi M, Hou S, Ding B, Liu L, Ji X et al (2012b) MiR-483-5p suppresses the proliferation of glioma cells via directly targeting ERK1. FEBS Lett 586:1312–1317
Wang S, Jiao B, Geng S, Ma S, Liang Z, Lu S (2014) Combined aberrant expression of microRNA-214 and UBC9 is an independent unfavorable prognostic factor for patients with gliomas. Med Oncol 31:767
Wang S, Lu S, Geng S, Ma S, Liang Z, Jiao B (2013b) Expression and clinical significance of microRNA-326 in human glioma miR-326 expression in glioma. Med Oncol 30:373
Wang X, Wang J, Ma H, Zhang J, Zhou X (2012c) Downregulation of miR-195 correlates with lymph node metastasis and poor prognosis in colorectal cancer. Med Oncol 29:919–927
Wang XR, Luo H, Li HL, Cao L, Wang XF, Yan W et al (2013c) Overexpressed let-7a inhibits glioma cell malignancy by directly targeting K-ras, independently of PTEN. Neuro Oncol 15:1491–1501
Webster RJ, Giles KM, Price KJ, Zhang PM, Mattick JS, Leedman PJ (2009) Regulation of epidermal growth factor receptor signaling in human cancer cells by microRNA-7. J Biol Chem 284:5731–5741
Wu N, Xiao L, Zhao X, Zhao J, Wang J, Wang F et al (2012) miR-125b regulates the proliferation of glioblastoma stem cells by targeting E2F2. FEBS Lett 586:3831–3839
Wu Z, Wang L, Li G, Liu H, Fan F, Li Z et al (2013) Increased expression of microRNA-9 predicts an unfavorable prognosis in human glioma. Mol Cell Biochem 384(1–2):263–268
Wurdinger T, Tannous BA, Saydam O, Skog J, Grau S, Soutschek J et al (2008) miR-296 regulates growth factor receptor overexpression in angiogenic endothelial cells. Cancer Cell 14:382–393
Xia H, Qi Y, Ng SS, Chen X, Chen S, Fang M et al (2009) MicroRNA-15b regulates cell cycle progression by targeting cyclins in glioma cells. Biochem Biophys Res Commun 380:205–210
Xu T, Zhu Y, Xiong Y, Ge YY, Yun JP, Zhuang SM (2009) MicroRNA-195 suppresses tumorigenicity and regulates G1/S transition of human hepatocellular carcinoma cells. Hepatology 50:113–121
Yamada R, Nakano I (2012) Glioma stem cells: their role in chemoresistance. World Neurosurg 77:237–240
Yan D, Ng WL, Zhang X, Wang P, Zhang Z, Mo YY et al (2010) Targeting DNA-PKcs and ATM with miR-101 sensitizes tumors to radiation. PLoS One 5:e11397
Ying Z, Li Y, Wu J, Zhu X, Yang Y, Tian H et al (2013) Loss of miR-204 expression enhances glioma migration and stem cell-like phenotype. Cancer Res 73:990–999
Yue X, Wang P, Xu J, Zhu Y, Sun G, Pang Q et al (2012) MicroRNA-205 functions as a tumor suppressor in human glioblastoma cells by targeting VEGF-A. Oncol Rep 27:1200–1206
Zbinden M, Duquet A, Lorente-Trigos A, Ngwabyt SN, Borges I, Ruiz i Altaba A (2010) NANOG regulates glioma stem cells and is essential in vivo acting in a cross-functional network with GLI1 and p53. EMBO J 29:2659–2674
Zhang C, Han L, Zhang A, Yang W, Zhou X, Pu P et al (2010a) Global changes of mRNA expression reveals an increased activity of the interferon-induced signal transducer and activator of transcription (STAT) pathway by repression of miR-221/222 in glioblastoma U251 cells. Int J Oncol 36:1503–1512
Zhang J, Han L, Ge Y, Zhou X, Zhang A, Zhang C et al (2010b) miR-221/222 promote malignant progression of glioma through activation of the Akt pathway. Int J Oncol 36:913–920
Zhang QQ, Xu H, Huang MB, Ma LM, Huang QJ, Yao Q et al (2012a) MicroRNA-195 plays a tumor-suppressor role in human glioblastoma cells by targeting signaling pathways involved in cellular proliferation and invasion. Neuro Oncol 14:278–287
Zhang S, Wan Y, Pan T, Gu X, Qian C, Sun G et al (2012b) MicroRNA-21 inhibitor sensitizes human glioblastoma U251 stem cells to chemotherapeutic drug temozolomide. J Mol Neurosci 47:346–356
Zhang W, Zhang J, Hoadley K, Kushwaha D, Ramakrishnan V, Li S et al (2012c) miR-181d: a predictive glioblastoma biomarker that downregulates MGMT expression. Neuro Oncol 14:712–719
Zhang Y, Chao T, Li R, Liu W, Chen Y, Yan X et al (2009) MicroRNA-128 inhibits glioma cells proliferation by targeting transcription factor E2F3a. J Mol Med (Berl) 87:43–51
Zhao S, Deng Y, Liu Y, Chen X, Yang G, Mu Y et al (2013a) MicroRNA-153 is tumor suppressive in glioblastoma stem cells. Mol Biol Rep 40:2789–2798
Zhao WH, Wu SQ, Zhang YD (2013b) Downregulation of miR-124 promotes the growth and invasiveness of glioblastoma cells involving upregulation of PPP1R13L. Int J Mol Med 32:101–107
Zheng X, Chopp M, Lu Y, Buller B, Jiang F (2013) MiR-15b and miR-152 reduce glioma cell invasion and angiogenesis via NRP-2 and MMP-3. Cancer Lett 329:146–154
Zhou J, Xu T, Yan Y, Qin R, Wang H, Zhang X et al (2013) MicroRNA-326 functions as a tumor suppressor in glioma by targeting the Nin one binding protein (NOB1). PLoS One 8:e68469
Zhou X, Ren Y, Moore L, Mei M, You Y, Xu P et al (2010a) Downregulation of miR-21 inhibits EGFR pathway and suppresses the growth of human glioblastoma cells independent of PTEN status. Lab Invest 90:144–155
Zhou X, Zhang J, Jia Q, Ren Y, Wang Y, Shi L et al (2010b) Reduction of miR-21 induces glioma cell apoptosis via activating caspase 9 and 3. Oncol Rep 24:195–201
Acknowledgment
This work was supported by grants of Internal Grant Agency of the Czech Ministry of Health no. NT13514-4/2012 and NT11214-4/2010; project “CEITEC—Central European Institute of Technology” (CZ.1.05/1.1.00/02.0068).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Wien
About this chapter
Cite this chapter
Sana, J., Besse, A., Slaby, O. (2014). MicroRNAs in the Molecular Pathology of Gliomas. In: Sedo, A., Mentlein, R. (eds) Glioma Cell Biology. Springer, Vienna. https://doi.org/10.1007/978-3-7091-1431-5_4
Download citation
DOI: https://doi.org/10.1007/978-3-7091-1431-5_4
Published:
Publisher Name: Springer, Vienna
Print ISBN: 978-3-7091-1430-8
Online ISBN: 978-3-7091-1431-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)