Abstract
Acute myeloid leukemia (AML) is a widely prevalent disease worldwide and poses a large threat to public health. Previous studies have shown that AML is associated with cytogenetic heterogeneity, complex subtypes, and different therapeutic approaches. In this study, we found that miR-486 was upregulated in AML using both The Cancer Genome Atlas (TCGA) database and patient tissues. After knockdown of miR-486 by short hairpin RNA (shRNA), we discovered that miR-486 was required for cell proliferation. Through miRNA profile analysis and a dual-luciferase reporter assay, suppressor of cytokine signaling 2 (SOCS2) was identified as a direct target of miR-486. Therefore, by silencing SOCS2, a negative regulator of the Janus kinase (JAK)–signal transducer and activator of transcription (STAT) pathway, miR-486 enhanced JAK-STAT3 activity and promoted cell proliferation. The miR-486-SOCS2-STAT3 proliferation axis is therefore involved in the pathogenesis of AML, providing a novel molecular mechanism and diagnostic and therapeutic clues for AML.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Acute myeloid leukemia (AML) is characterized as abnormally differentiated hematopoietic cell infiltration into the bone marrow and other tissues (Dohner et al. 2015). Its cytogenetic heterogeneity is widely accepted, as genes, including NPM1, CEBPA, RUNX2, and FLT3-ITD, are highly mutated in AML (Dohner et al. 2015). Although advanced techniques, such as next-generation sequencing, have provided us new insights into its tumorigenesis, the detailed mechanisms and translational medicine have not been fully investigated.
The JAK-STAT pathway is a signaling cascade by which cytokines regulate cell proliferation, immune response, and inflammation (Morris et al. 2018). In particular, STAT3 promotes cell proliferation. On the one hand, phosphorylated STAT3 binds to the transcription factor IRF9 and upregulates the anti-apoptosis genes Bcl-2, Mcl-1, and Bcl-xL (Bournazou and Bromberg 2013). On the other hand, the family of suppressors of cytokine signaling (SOCS) inhibits the activity of JAK-STAT signaling, forming a negative feedback loop (Honda et al. 2005; Ivashkiv and Donlin 2014; Schneider et al. 2014).
MicroRNAs (miRNAs) are approximately 22-nucleotide, small, non-coding RNAs that regulate many biological processes, such as cell proliferation, differentiation, and apoptosis, by binding to the 3′UTR of target mRNAs, resulting in gene silencing (Duan et al. 2017; Peng et al. 2017). Increasing evidence has shown that abnormal expression patterns of miRNAs occur in many diseases (Feng et al. 2017; Zhang et al. 2017). In AML, each subtype displays a specific miRNA expression spectrum (Liu et al. 2019). For instance, the oncogene miR-9 is highly expressed in mixed-lineage leukemia (Chen et al. 2013), and miR-155 is overexpressed in FLT3-ITD-mutated patients (Gerloff et al. 2015). Above all, miRNAs play vital roles and act as promising diagnostic and therapeutic targets in AML.
Recently, evidence showed that miR-486 participates in signaling transduction and tumorigenesis. miR-486 is downstream of PAX3-FOXO1 and is highly expressed in rhabdomyosarcoma (RMS) tissues (Ghamloush et al. 2019). In colorectal neoplasia (CRN) patients, miR-486 was low and may serve as a prognostic biomarker (Yang et al. 2018). miR-486 promoted Capan-2 cell proliferation and targeted PTEN (Xia et al. 2019). Based on the above work, we speculated that miR-486 may also play roles in AML. Herein, we evaluated the expression level of miR-486 from The Cancer Genome Atlas (TCGA) database and AML tissues and investigated potential mechanisms. The results showed that miR-486 is highly expressed in AML and promotes JAK-STAT3 activity and cell proliferation by silencing SOCS2. The miR-486-SOCS2-STAT3-proliferation axis could act as a potential diagnostic and therapeutic target.
Methods and materials
Patient specimens
In the present study, bone marrow was collected from AML patients (n = 45) diagnosed at First People’s Hospital of Lianyungang as well as non-AML patients (n = 43). All specimens derived from patients were approved by patients and the Ethics Committee of First People’s Hospital of Lianyungang. All tissues were stored in liquid nitrogen.
Cell culture, stable cell construction, and transfection
The cancer cell lines SKM-1 and AML-193 were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA) and maintained in Dulbecco’s Modified Eagle’s Medium (DMEM) (Sigma, St. Louis, MO, USA) with 10% fetal bovine serum (FBS) (HyClone, Logan, UT, USA), 1% streptomycin (v/v) (Sigma), and 1% penicillin (v/v) (Sigma) in an incubator at 37 °C with 5% CO2. pLKO.1-TRC (Addgene, #10878) and pLVX-CMV100 (Addgene, no. 110718) backbones were digested by restriction enzymes, and shRNAs and miRNA/NC were ligated by T4 DNA ligase (Cat: M0202S, NEB, MA, USA). For lentivirus production, the lentivirus vectors psPAX2 (Addgene, no. 12260) and pMD2.G (Addgene, no. 12259) were co-transfected into HEK 293T cells by Lipofectamine 2000 (Invitrogen, CA, USA) according to the manufacturer’s instructions. Forty-eight hours later, the supernatants were harvested and filtered through a 0.45-um filter, and 10 μg/mL polybrene was added (Cat: TR-1003, Sigma-Aldrich, MO, USA). Then, 2 μg/mL puromycin (Cat: P8833, Sigma-Aldrich, MO, USA) was used to screen positive cells after 48 h.
For plasmid transfection, Lipofectamine 2000 (Invitrogen, CA) was used according to the manufacturer’s instructions.
The oligo sequences involved are as follows: shSOCS2#1:5′-CCG GTC AGA TTG GAC TCT ATC ATA TCT CGA GAT ATG ATA GAG TCC AAT CTG ATT TTT TG-3′; shSOCS2#2: 5′-CCG GAG CTG AAG CTA ATC TAA TTT ACT CGA GTA AAT TAG ATT AGC TTC AGC TTT TTT TG-3′; shSOCS2#3: 5′-CCG GGT TCA TCT GAT CGA CTA CTA TCT CGA GAT AGT AGT CGA TCA GAT GAA CTT TTTG-3′; sh-miR-486: 5′-CCG GCG GGG CAG CTC AGT ACA GGA TCT CGA GAT CCT GTA CTG AGC TGC CCC GTT TTTG-3′.
TCGA database analysis
The TCGA-LAML expression data were downloaded, and analyses were performed using TCGAbiolinks. The TPM was normalized and log2-transformed before comparison. LAML patients (173) were compared with the normal cohort (70) by t test and yielded differentiated expressed genes. The has-miR-486 survival analysis was performed using the TCGAbiolinks function to analyze survival.
Cell viability assay
Cell viability was evaluated using the cell counting kit-8 (CCK-8) (Dojindo, Mashiki-machi, Japan) according to the manufacturer’s instructions. Briefly, stable pLOK.1-sh-vec and pLOK.1-sh-miR-486#1/2/3 cells (2 × 103 per well) were plated in 96-well plates and treated with the indicated conditions. To determine the percentage viability, the absorbance was measured at 450 nm using a microplate reader (MQX 200, BioTek Instruments, Winooski, VT, USA) followed by treatment with CCK-8 for 2 h at 37 °C. Five repetition wells were carried out.
Luciferase assay
The wild-type and mutant-3′UTR of human SOCS2 were cloned into the pMIR Report luciferase vector (Ambion, Carlsbad, CA). miR-486 or a negative control (NC), pMIR-SOCS2-3′ UTR or pMIR empty vector pRL-TK plasmid (Promega, Madison, WI), was co-transfected into HEK 293T cells using Lipofectamine 2000. The pGL3 vector or STAT3 reporter firefly plasmid and pRL-TK were co-transfected into the indicated cell lines. Twenty-four hours after transfection, cell harvest and luciferase assay were conducted according to the manufacturer’s instructions of the Dual-Luciferase Reporter Assay System (Promega, Beijing, China). Plasmid extraction kits were purchased from TIANGEN (Beijing, China).
The oligos involved in this study were as follows: SOCS2-3′ UTR-WT: 5′-TTT ACT GTA GTT CAA TTA CAG GA-3′; SOCS2-3′ UTR mutant: 5′-TTT ACT ATT ATG CAG TAT GTA AA-3′; miR-486: 5′-CGG GGC AGC TCA GTA CAG GAT-3′; NC: 5′-TCC TGT ACT GAG CTG CCC CGA G-3′.
RNA extraction and RT-PCR
Trizol (Invitrogen, CA, USA) and the First-Strand cDNA Synthesis Kit (Invitrogen, CA, USA) were used for total RNA extraction and cDNA synthesis, respectively. miR-486 and SOCS2, Bcl-2, Mcl-1, and Bcl-xL mRNA levels were detected by Power SYBR Green PCR Master Mix (Invitrogen, Carlsbad, CA, USA) on a 7900HT FAST real-time PCR system (Life Technologies, Carlsbad, CA). GAPDH was used as an internal reference control. Six repetitions were carried out. The relative mRNA levels were calculated with the 2*(−∆∆CT) method.
The following primer sequences were used: SOCS2, forward, 5′-TTA AAA GAG GCA CCA GAA GGA AC-3′, reverse, 5′-AGT CGA TCA GAT GAA CCA CAC T-3′; Bcl-2, forward, 5′-GGT GGG GTC ATG TGT GTG G-3′, reverse, 5′-CGG TTC AGG TAC TCA GTC ATCC-3′; Mcl-1, forward, 5′-TGC TTC GGA AAC TGG ACA TCA-3′, reverse, 5′-TAG CCA CAA AGG CAC CAA AAG-3′; Bcl-xL, forward, 5′-GAG CTG GTG GTT GAC TTT CTC-3′, reverse, 5′-TCC ATC TCC GAT TCA GTC CCT-3′. In particular, miR-486 was quantified using MystiCq® microRNA qPCR Assay Primers (Cat: MIRAP00422, Sigma-Aldrich, MO, USA).
Cell cycle analysis
Cell samples were treated with the FITC BrdU Flow Kit, and the cell cycle was analyzed on a BD ACCURI C6 according to the manufacturer’s instructions. Cells were gated according to their BrdU and 7-AAD levels, e.g., G1 cells have 2n DNA and low BrdU, whereas S phase cells have > 2n DNA and high BrdU.
Western blot
Protein (50 mg) was loaded into every well of a 10% SDS-PAGE gel. After transferring onto polyvinylidene fluoride (PVDF) membranes, the membranes were blocked with 5% BSA for 1 h at room temperature. The primary antibody was incubated overnight at 4 °C, and peroxidase-conjugated secondary antibody was then incubated for 2 h at room temperature. The antibodies for SOCS2 (Cat: no. 2779) and GAPDH (Cat: no. 5174) were purchased from Cell Signaling Technology (CST, Danvers, MA, USA).
Immunofluorescence
After three PBS washes, cells were fixed for 15 min with 4% paraformaldehyde and permeabilized for 10 min with 0.2% Triton X-100. Anti-STAT3 antibody (Cat: no. 12640, CST, Danvers, MA, USA) was incubated overnight at 4 °C, and anti-rabbit-488 secondary antibody (Cat: A32731, Invitrogen, Carlsbad, CA, USA) and DAPI dye (Cat: 62247, Invitrogen, Carlsbad, CA, USA) were then incubated at room temperature for 2 h. A Nikon DXM1200 fluorescence microscope and Image-Pro Plus version 6.0 software (Media Cybernetics, Washington) were used to capture and analyze the images, respectively.
Statistical analysis
The results were expressed as the means ± SEM. Statistical analysis was performed using one-way ANOVA to compare between multiple groups and Student’s t test to compare between two groups. A value of P < 0.05 was considered statistically significant. Statistical analysis was performed using GraphPad Prism 7 software (CA, USA).
Results
miR-486 is upregulated in LAML patients
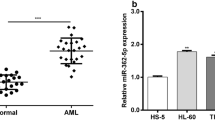
To investigate the role of miR-486 in AML, we first explored the relationship between miR-486 expression levels and patient survival rate from the online TCGA database. We found that has-miR-486 levels and survival rates were negatively correlated (Fig. 1a). Patients expressing lower levels of miR-486 had a longer survival range. Meanwhile, when comparing tissues from normal donors (ND) and AML patients, the real-time polymerase chain reaction (RT-PCR) results showed that the miR-486 expression levels in AML (n = 45) were approximately twofold those of ND (n = 43) (Fig. 1b). Additionally, agarose gel electrophoresis of six randomly selected specimens (Fig. 1c) confirmed the results.
MiR-486 is upregulated in LAML patients. a The TCGA database has-miR-486 expression levels in AML and survival rates. The has-miR-486 levels and survival rates were negatively correlated. b Relative miR-486 levels of normal donors (n = 43) and AML (N = 45) specimens. miR-486 was quantified using a microRNA qPCR assay. c miR-486 agarose gel electrophoresis of six randomly selected specimens of ND and AML in b
Collectively, these data indicate that miR-486 is upregulated in AML patients.
miR-486 is required for LAML cell proliferation
We used three different shRNA sequences to silence miR-486 in the AML cell lines SKM-1 and AML-193. The miR-486 expression levels in stable sh-miR-486#1/2/3 cells were approximately 20% of sh-vec cells (Fig. 2 a and c). Using the 3-(4,5)-dimethylthiahiazo (-z-y1)-3,5-di-phenytetrazoliumromide (MTT) assay, sh-miR-486 stable cells showed lower cell viability than sh-vec cells in a time-dependent manner, both for SKM-1 (Fig. 2b) and AML-193 (Fig. 2d) cells. The proliferation rate of knockdown cells was approximately 75% that of control cells on day 5 (Fig. 2 b and d). Meanwhile, flow cytometry (FCM) analysis indicated that the percentages of sh-vec cells and sh-miR-486 cells in the G0/G1 phase were approximately 30% and 50%, respectively (Fig. 2 e and f). However, the percentage of sh-miR-486 cells in the S phase was significantly lower than that of sh-vec cells (Fig. 2 e and f). Furthermore, both cell lines had a similar number in the G2/M phase (Fig. 2 e and f). The FCM analysis illustrated the sh-miR-486 cells had a lower proliferation rate than sh-vec cells.
miR-486 is required for LAML cell proliferation. a, c miR-486 knockdown efficiency of three shRNA sequences in SKM-1 and AML-193 cells. Vec was the control cell line constructed by the pLKO.1 empty vector. miR-486 was quantified using a microRNA qPCR assay. Six repetitions were carried out. b, d Cell viability of miR-486 knockdown and control cells from day 0 to day 5 in SKM-1 and AML-193 cells. Vec was the control cell line constructed by the pLKO.1 empty vector. For five repetition wells of sh-miR-486 cells, two wells were pLOK.1-sh-miR-486#1 cells, two wells were pLOK.1-sh-miR-486#2 cells, and one well was pLOK.1-sh-miR-486#3 cells. e, f Percentage of cell cycle phase of miR-486 knockdown and control cells in SKM-1 and AML-193 cells. The stable cell lines were the same as those in b and d
These data demonstrate that knockdown of miR-486 results in low cell viability and induces G0/G1 cell cycle arrest, indicating that miR-486 is required for AML cell proliferation.
SOCS2 is a direct target of miR-486
Subsequently, we explored the molecular target of miR-486. Through a target gene prediction database analysis, SOCS2 was screened and identified as a target of miR-486. Oligonucleotide sequences showed that SOCS2 has a conserved miR-486 seed sequence at position 1457–1473 (Fig. 3a), and the luciferase reporter system verified the analysis. miR-486 decreased approximately 90% of the SOCS2-WT-3′UTR-driven firefly luciferase expression, while it had no effect on that of SOCS2-Mut-3′UTR (Fig. 3b). When miR-486 was overexpressed in SKM-1 and AML-193 cells, the SOCS2 mRNA levels decreased in a miR-486 dose-dependent manner (Fig. 3 c and d), indicating that miR-486 represses SOCS2 expression. In the TCGA database, the SOCS2 expression in normal donors was approximately fourfold higher than that of AML patients (Fig. 3e), consistent with the result that miR-486 is high in AML patients. Convincingly, in AML and ND specimens, the mRNA (Fig. 3f) and protein levels (Fig. 3 g and h) further confirmed that SOCS2 was lowly expressed in AML patients. The SOCS2 protein level in ND was approximately 2-fold that of AML patients.
SOCS2 is a direct target of miR-486. a Sequence alignment of miR-486 and SOCS2 3′-UTR sequences. SOCS2 3′-UTR has a conserved miR-486 seed. b Co-transfection of miR-486 or negative control (NC) and SOCS2 3′ UTR-WT/Mut into 293T cells. The luciferase activity was normalized to the (NC + SOCS2 3′UTR-WT) group. c, d SOCS2 mRNA levels in miR-486-transfected SKM-1 and AML-193 cells. The mRNA levels were normalized to the untransfected group. e TCGA database of SOCS2 mRNA expression levels in negative controls (n = 70) and AML (n = 173). f SOCS2 mRNA expression levels in normal donors (ND) and AML patient specimens. The mRNA levels were normalized to the ND group. g SOCS2 protein expression levels in normal donors and AML patient specimens. h Western blot quantification of SOCS2 protein in normal donors and AML patient specimens. The gray intensity of Western blot data was quantified by ImageJ software and normalized to the ND group
To summarize, SOCS2 was first identified as a miR-486 target by oligonucleotide sequence analysis. Then, cellular overexpression, database retrieval, and patient specimens demonstrated that miR-486 represses SOCS2 expression. SOCS2 is therefore a direct target of miR-486.
miR-486 enhances JAK-STAT activity
Because SOCS2 is a negative regulator of JAK-STAT signaling (Honda et al. 2005; Ivashkiv and Donlin 2014; Schneider et al. 2014), we speculated that miR-486 may influence JAK-STAT activity. Both knockdown and stably overexpressing cells were used, and the miR-486 levels were examined by RT-PCR (Fig. 4a). After transfection with the pGL3 vector or STAT3 activity reporter plasmid, knockdown of miR-486 decreased STAT3 luciferase activity, whereas overexpression of miR-486 increased STAT3 luciferase activity (Fig. 4b), indicating that miR-486 enhances JAK-STAT signaling. The subcellular localization of STAT3 was also tested. In miR-486 knockdown cells, nuclear-localized STAT3 was approximately 50% that of sh-vec cells; however, in miR-486-overexpressing cells, nuclear-localized STAT3 was 140% that of negative control (NC) cells (Fig. 4c), showing that miR-486 promotes STAT3 translocation into the nucleus. Furthermore, the RT-PCR results also illustrated that miR-486 knockdown decreased STAT3 target gene expression by nearly 50%, while miR-486 overexpression increased the expression of target genes, including Bcl-2 (Fig. 4d), Mcl-1 (Fig. 4e), and Bcl-xL (Fig. 4f), by 2–3-fold.
miR-486 enhances JAK-STAT activity. a miR-486 mRNA levels in SKM-1 and AML-193 cells. Vec and sh-miR-486, NC, and miR-486 were stable cells constructed by the pLKO.1 and pLVX-CMV100 lentivirus vectors, respectively. Six repetitions were carried out. b STAT3 luciferase reporter activity in SKM-1 and AML-193 cells described in a. The luciferase activity was normalized to the vec group. c Statistical results of nuclear-translocated STAT3 and SOCS2 protein expression levels in SKM-1 and AML-193 cells described in a. The nuclear-translocated STAT3 was normalized to the vec group. d–f Relative mRNA levels of Bcl-2, Mcl-1, and Bcl-XL in SKM-1 and AML-193 cells described in a. The mRNA levels were normalized to the vec group
miR-486 promotes cell proliferation via JAK-STAT signaling
Based on the above results, miR-486 both promotes cell proliferation and enhances JAK-STAT signaling, but whether the effect of miR-486 on cell proliferation is mediated by JAK-STAT signaling has not been demonstrated. To clearly elucidate the miR-486-SOCS2-STAT3 proliferation axis, we used two STAT3 selective inhibitors, S3I-201 and Stattic. S3I-201 inhibits STAT3-DNA binding and STAT3-dependent transcription (Lin et al. 2009; Schust et al. 2006; Siddiquee et al. 2007), while Stattic inhibits STAT3 activation (Schust et al. 2006). miR-486 overexpression promoted cell viability from day 2 to day 3, while the promotion effect was blocked by Stattic and S3I-201 (Fig. 5 a and b), demonstrating that miR-486 promotes cell proliferation by stimulating STAT3 activation and transcription activity. These results are consistent with the immunofluorescence results showing that miR-486 promotes STAT3 nuclear translocation (Fig. 4c) and target gene expression (Fig. 4d–f). The FCM analysis also showed that miR-486 overexpression decreased G0/G1 retention and increased S advancement, but these effects were blocked by Stattic (Fig. 5 c and d).
miR-486 promotes cell proliferation via JAK-STAT signaling. a, b Cell viability of miR-486 overexpressed and negative control cells upon treatment with S3I-201, Stattic, or vehicle in SKM-1 and AML-193 cells. The proliferation rate was determined by the CCK-8 kit. c, d Percentage of cell cycle phase of miR-486 overexpressed and negative control cells upon treatment with Stattic or vehicle in SKM-1 and AML-193 cells. e Schematic diagram of the miR-486-SOCS2-STAT3 proliferation axis. miR-486 promotes cell proliferation by silencing SOCS2 to accelerate STAT3 nuclear translocation and JAK-STAT signaling
In brief, these results demonstrate that miR-486 promotes cell proliferation via JAK-STAT signaling dependent on STAT3 activation and nuclear translocation.
Discussion and conclusion
For AML, great progress has been made by advanced techniques, such as next-generation sequencing (Dohner et al. 2015). Some genes, including MYC, TP53, and BCL2, are vital for hematopoiesis or leukemogenesis (Akhter et al. 2018). The overall survival (OS) of children with AML is up to 70% (Lonetti et al. 2019), and highly expressed MAP4K3/4/5 combined with lowly expressed MAP4K1 may be a tool for OS prediction (Bai et al. 2019). However, the cytogenetic heterogeneity of AML poses a great obstacle for translational practice (Dohner et al. 2015). Thus, precise mechanisms and signaling transduction are far from fully discovered.
JAK-STAT signaling has an important role in hematopoietic development, and selective inhibition of this pathway is effective for AML therapy. JAK-STAT inhibitors, such as ruxolitinib, lestaurtinib, and pacritinib, are currently under clinical evaluation (Venugopal et al. 2019). Given that the SOCS family inhibits the activity of JAK-STAT signaling (Honda et al. 2005; Ivashkiv and Donlin 2014; Schneider et al. 2014), much attention has been given to this family. The knockdown of SOCS3 by siRNA can repress TNF-α-induced apoptosis (Zhao et al. 2012). SOCS2 regulates hematopoietic stemness and proliferation (Vitali et al. 2015). However, one study indicated that SOCS2 accompanied by IL2RA, NPDC1, and PHGDH accelerated the stemness and aggressiveness of leukemia (Nguyen et al. 2019), which is contradictory to other reports. One possible explanation is that different AML subtypes display different signatures. More detailed research is needed to disclose the underlying mechanisms.
miRNAs are related to different kinds of biological processes, and their dysregulation may cause some disease lesions (Di Leva et al. 2014). Onco-miRNAs were reported to inhibit the expression of tumor suppressors, and TS-miRNAs are considered repressors of oncogenes. TS-miRNAs are usually expressed in tumors; thus, they might play an important role in the progression of tumors (Lujambio and Lowe 2012). Previous studies have reported that miR-486 participates in the regulation of cell proliferation in many diseases. It was found to be downregulated in the blood of sepsis patients and may be a prognostic predictor (Ojha et al. 2019), and it can also downregulate Olfactomedin 4 (OLFM4) in ovarian adenocarcinoma cells (Liu and Rodgers 2016). In gastric cancer, miR-486 is a tumor suppressor and is strongly downregulated (Ma and Tao 2012). miR-486 induces autophagy in lung cancer (Xu et al. 2017) and suppresses osteosarcoma invasion (He et al. 2017; Li et al. 2017; Liu et al. 2018). It may act as a tumor suppressor in hepatocellular carcinoma (HCC) by regulating CLDN10 and CITRON (Sun et al., 2015). In the cardiovascular system, miR-486 promotes heart ventricular growth in mice (Lange et al. 2019), and it also accelerates fibroblast proliferation after skin injury (Wu et al. 2019). Taken together, miR-486 displays different effects in different organs and tissues.
However, little is known about the role and mechanisms of miR-486 in AML. In this study, we found the miR-486 was upregulated in AML and was required for AML cell proliferation. Upon silencing SOCS2 and enhancing JAK-STAT signaling, miR-486 promoted cell proliferation (Fig. 5e). From this point of view, miR-486 is oncogenic. However, a previous report indicated that miR-486 was a tumor suppressor in gastric cancer (Ma and Tao 2012) and HCC (Sun et al. 2015). This contradiction may result from different cancers and different tissues. Although its elevated expression level was confirmed in patient specimens, the effect of miR-486 on AML cells need to be validated in other conditions or even in vivo.
In summary, our study revealed that miR-486 expression is upregulated in AML and is required for AML cell proliferation and that SOCS2 is a direct target of miR-486. miR-486 promotes cell proliferation by silencing SOCS2 to accelerate STAT3 nuclear translocation and JAK-STAT signaling. Thus, the miR-486-SOCS2-STAT3 proliferation axis is involved in the pathogenesis of AML, which may provide diagnostic and therapeutic clues for AML.
References
Akhter A, Farooq F, Elyamany G, Mughal MK, Rashid-Kolvear F, Shabani-Rad MT, Street L, Mansoor A (2018) Acute myeloid leukemia (AML): upregulation of BAALC/MN1/MLLT11/EVI1 gene cluster relate with poor overall survival and a possible linkage with coexpression of MYC/BCL2 proteins. Appl Immunohistochem Mol Morphol 26:483–488
Bai Z, Yao Q, Sun Z, Xu F, Zhou J (2019) Prognostic value of mRNA expression of MAP4K family in acute myeloid leukemia. Technol Cancer Res Treat 18:1533033819873927
Bournazou E, Bromberg J (2013) Targeting the tumor microenvironment: JAK-STAT3 signaling. Jak-stat 2:e23828
Chen P, Price C, Li Z, Li Y, Cao D, Wiley A, He C, Gurbuxani S, Kunjamma RB, Huang H, Jiang X, Arnovitz S, Xu M, Hong GM, Elkahloun AG, Neilly MB, Wunderlich M, Larson RA, Le Beau MM, Mulloy JC, Liu PP, Rowley JD, Chen J (2013) miR-9 is an essential oncogenic microRNA specifically overexpressed in mixed lineage leukemia-rearranged leukemia. Proc Natl Acad Sci U S A 110:11511–11516
Di Leva G, Garofalo M, Croce CM (2014) MicroRNAs in cancer. Annu Rev Pathol 9:287–314
Dohner H, Weisdorf DJ, Bloomfield CD (2015) Acute myeloid leukemia. N Engl J Med 373:1136–1152
Duan R, Zhang Z, Zheng F, Wang L, Guo J, Zhang T, Dai X, Zhang S, Yang D, Kuang R, Wang G, He C, Hakeem A, Shu C, Yin P, Lou X, Zeng F, Liang H, Xia F (2017) Combining protein and miRNA quantification for bladder cancer analysis. ACS Appl Mater Interfaces 9:23420–23427
Feng C, Sun P, Hu J, Feng H, Li M, Liu G, Pan Y, Feng Y, Xu Y, Feng K, Feng Y (2017) miRNA-556-3p promotes human bladder cancer proliferation, migration and invasion by negatively regulating DAB2IP expression. Int J Oncol 50:2101–2112
Gerloff D, Grundler R, Wurm AA, Brauer-Hartmann D, Katzerke C, Hartmann JU, Madan V, Muller-Tidow C, Duyster J, Tenen DG, Niederwieser D, Behre G (2015) NF-kappaB/STAT5/miR-155 network targets PU.1 in FLT3-ITD-driven acute myeloid leukemia. Leukemia 29:535–547
Ghamloush F, Ghayad SE, Rammal G, Fahs A, Ayoub AJ, Merabi Z, Harajly M, Zalzali H, Saab R (2019) The PAX3-FOXO1 oncogene alters exosome miRNA content and leads to paracrine effects mediated by exosomal miR-486. Sci Rep 9:14242
He M, Wang G, Jiang L, Qiu C, Li B, Wang J, Fu Y (2017) miR-486 suppresses the development of osteosarcoma by regulating PKC-delta pathway. Int J Oncol 50:1590–1600
Honda K, Yanai H, Negishi H, Asagiri M, Sato M, Mizutani T, Shimada N, Ohba Y, Takaoka A, Yoshida N, Taniguchi T (2005) IRF-7 is the master regulator of type-I interferon-dependent immune responses. Nature 434:772–777
Ivashkiv LB, Donlin LT (2014) Regulation of type I interferon responses. Nat Rev Immunol 14:36–49
Lange S, Banerjee I, Carrion K, Serrano R, Habich L, Kameny R, Lengenfelder L, Dalton N, Meili R, Borgeson E, Peterson K, Ricci M, Lincoln J, Ghassemian M, Fineman J, Del Alamo JC, Nigam V (2019) miR-486 is modulated by stretch and increases ventricular growth. JCI Insight 4
Li N, Sun ZH, Fang M, Xin JY, Wan CY (2017) Long non-coding RNA ZFAS1 sponges miR-486 to promote osteosarcoma cells progression and metastasis in vitro and vivo. Oncotarget 8:104160–104170
Lin L, Amin R, Gallicano GI, Glasgow E, Jogunoori W, Jessup JM, Zasloff M, Marshall JL, Shetty K, Johnson L, Mishra L, He AR (2009) The STAT3 inhibitor NSC 74859 is effective in hepatocellular cancers with disrupted TGF-beta signaling. Oncogene 28:961–972
Liu W, Rodgers GP (2016) Olfactomedin 4 expression and functions in innate immunity, inflammation, and cancer. Cancer Metastasis Rev 35:201–212
Liu Y, Zhang J, Xing C, Wei S, Guo N, Wang Y (2018) miR-486 inhibited osteosarcoma cells invasion and epithelial-mesenchymal transition by targeting PIM1. Cancer Biomark Sect A Dis Mark 23:269–277
Liu Y, Cheng Z, Pang Y, Cui L, Qian T, Quan L, Zhao H, Shi J, Ke X, Fu L (2019) Role of microRNAs, circRNAs and long noncoding RNAs in acute myeloid leukemia. J Hematol Oncol 12:51
Lonetti A, Pession A, Masetti R (2019) Targeted therapies for pediatric AML: gaps and perspective. Front Pediatr 7:463
Lujambio A, Lowe SW (2012) The microcosmos of cancer. Nature 482:347–355
Ma YY, Tao HQ (2012) Microribonucleic acids and gastric cancer. Cancer Sci 103:620–625
Morris R, Kershaw NJ, Babon JJ (2018) The molecular details of cytokine signaling via the JAK/STAT pathway. Protein Sci Publ Protein Soc 27:1984–2009
Nguyen CH, Gluxam T, Schlerka A, Bauer K, Grandits AM, Hackl H, Dovey O, Zochbauer-Muller S, Cooper JL, Vassiliou GS, Stoiber D, Wieser R, Heller G (2019) SOCS2 is part of a highly prognostic 4-gene signature in AML and promotes disease aggressiveness. Sci Rep 9:9139
Ojha R, Nandani R, Pandey RK, Mishra A, Prajapati VK (2019) Emerging role of circulating microRNA in the diagnosis of human infectious diseases. J Cell Physiol 234:1030–1043
Peng F, Li H, Xiao H, Li L, Li Y, Wu Y (2017) Identification of a three miRNA signature as a novel potential prognostic biomarker in patients with bladder cancer. Oncotarget 8:105553–105560
Schneider WM, Chevillotte MD, Rice CM (2014) Interferon-stimulated genes: a complex web of host defenses. Annu Rev Immunol 32:513–545
Schust J, Sperl B, Hollis A, Mayer TU, Berg T (2006) Stattic: a small-molecule inhibitor of STAT3 activation and dimerization. Chem Biol 13:1235–1242
Siddiquee K, Zhang S, Guida WC, Blaskovich MA, Greedy B, Lawrence HR, Yip ML, Jove R, McLaughlin MM, Lawrence NJ, Sebti SM, Turkson J (2007) Selective chemical probe inhibitor of Stat3, identified through structure-based virtual screening, induces antitumor activity. Proc Natl Acad Sci U S A 104:7391–7396
Sun H, Cui C, Xiao F, Wang H, Xu J, Shi X, Yang Y, Zhang Q, Zheng X, Yang X, Wu C, Wang L (2015) miR-486 regulates metastasis and chemosensitivity in hepatocellular carcinoma by targeting CLDN10 and CITRON. Hepatol Res 45:1312–1322
Venugopal S, Bar-Natan M, Mascarenhas JO (2019) JAKs to STATs: a tantalizing therapeutic target in acute myeloid leukemia. Blood Rev 100634
Vitali C, Bassani C, Chiodoni C, Fellini E, Guarnotta C, Miotti S, Sangaletti S, Fuligni F, De Cecco L, Piccaluga PP, Colombo MP, Tripodo C (2015) SOCS2 controls proliferation and stemness of hematopoietic cells under stress conditions and its deregulation marks unfavorable acute leukemias. Cancer Res 75:2387–2399
Wu SG, Li HT, Wang LL, Yan L (2019) Lidocaine promotes fibroblast proliferation after thermal injury via up-regulating the expression of miR-663 and miR-486. Kaohsiung J Med Sci
Xia L, Song M, Sun M, Chen W, Yang C (2019) miR-486 promotes Capan-2 pancreatic cancer cell proliferation by targeting phosphatase and tensin homolog deleted on chromosome 10 (PTEN). Front Genet 10:541
Xu Y, Wang Y, Yao A, Xu Z, Dou H, Shen S, Hou Y, Wang T (2017) Low frequency magnetic fields induce autophagy-associated cell death in lung cancer through miR-486-mediated inhibition of Akt/mTOR signaling pathway. Sci Rep 7:11776
Yang Q, Wang S, Huang J, Xia C, Jin H, Fan Y (2018) Serum miR-20a and miR-486 are potential biomarkers for discriminating colorectal neoplasia: a pilot study. J Cancer Res Ther 14:1572–1577
Zhang Z, Gong Q, Li M, Xu J, Zheng Y, Ge P, Chi G (2017) MicroRNA-124 inhibits the proliferation of C6 glioma cells by targeting Smad4. Int J Mol Med 40:1226–1234
Zhao X, Qi R, Sun C, Xie Y (2012) Silencing SOCS3 could inhibit TNF-alpha induced apoptosis in 3T3-L1 and mouse preadipocytes. Mol Biol Rep 39:8853–8860
Author information
Authors and Affiliations
Contributions
Xu Guanghui designed the research and wrote the paper; Cao Sha, Zhao Jingjing, Gao Jia, and Hu Yapeng performed the research and analyzed the data. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Disclaimer
All data were generated in-house, and the authors did not use a paper mill.
Conflict of interest
The authors declare that they have no competing interests.
Ethical standards
Patients were diagnosed according to the criteria of the World Health Organization classification of hematological malignancies. Before the initiation of treatment, clinical features, including Mantle Cell Lymphoma International Prognostic Index (MIPI) score, lactate dehydrogenase (LDH) content in serum, performance status (determined using Eastern Cooperative Oncology Group performance status scores), and white blood cell (WBC) counts, were collected. All experimental protocols and procedures involved in the present study were approved by the Ethics Committee of the First People’s Hospital of Lianyungang. Written informed consent was obtained from all participants prior to publication of this article.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sha, C., Jia, G., Jingjing, Z. et al. miR-486 is involved in the pathogenesis of acute myeloid leukemia by regulating JAK-STAT signaling. Naunyn-Schmiedeberg's Arch Pharmacol 394, 177–187 (2021). https://doi.org/10.1007/s00210-020-01892-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00210-020-01892-4