Abstract
Plastics are usually made up of low-density polyethylene (LDPE) that serve as the environmental nuisance. The recalcitrant nature of plastics is a huge concern, whereas the increasing demand has made it difficult to handle the plastic waste that eventually leads to plastic pollution. In recent years, due to increasing demand and high pressure for its safe disposal, plastic biodegradation has gained a lot of attention. In the current study, four bacterial strains were isolated from the solid-waste dumpsites of Faisalabad, Pakistan, using enrichment culture technique. The isolated bacterial strains were capable of growing on media having polystyrene as the sole carbon source. Based on 16S rRNA gene sequencing and phylogenetic analysis of the isolated strains Serratia sp., Stenotrophomonas sp. and Pseudomonas sp. were identified as the potential strains for the biodegradation of LDPE. Serratia sp. resulted in 40% weight loss of the LDPE plastic pieces after 150 days of treatment. Stenotrophomonas sp. and Pseudomonas species resulted in 32 and 21% weight loss of the treated piece of plastics (LDPE), respectively. Polyethylene pieces were characterized by Fourier-transform infrared spectroscopy (FTIR) analysis before and after biodegradation. The FTIR spectra indicated that the isolated bacterial strains have a good potential to degrade LDPE. Future studies are required to investigate the bacterial genetic makeup, mechanisms of LDPE biodegradation and the factors that can enhance the biodegradable characteristics of these indigenously isolated bacterial strains.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Plastics are petrochemical-based polymers that are integral part of our society because of their indispensable use in electronics, manufacturing of various equipment and in packaging material (Koller 2019). Due to their chemical inertness and resistance, plastics are highly employed in industrial manufacturing for a wide range of commodities and domestic products (Godfrey 2019; Muneer et al. 2020). Plastics are categorized into macro, micro and nano plastics depending upon the size of their constituent particles, all of these types have polluted the water reservoirs, resulting in serious threats to the aquatic and marine life (Yoshida et al. 2016; Allen et al. 2019). In environment, plastic products are persistent due to the complete absence or low activity of the catabolic enzymes that have the potential to degrade them (Alshehrei 2017). Various studies have concluded that the high ratio of aromatic components in the plastics make them resistant to degradation (Yoshida et al. 2016).
Currently, the rate of plastic production in the world is more than 410 million tons per annum, of which polyethylene terephthalate (PET) alone constitutes more than 70 million tons (Hahladakis et al. 2018; Palm et al. 2019). According to an estimation, more than 13 million tons of plastics entered the ocean from land-based sources in 2010 (Beaumont et al. 2019). Due to serious environmental effects of the plastic pollution, international forums such as the United Nations environmental program (UNEP) has included it in the list of global problems (Ivanova et al. 2018). Human activities can increase the declining threshold of biological diversity and extinction rates of biological species of fauna and flora (Villarrubia-Gómez et al. 2018). Because of poor waste management, major share of plastics enter into the marine ecosystem and have adverse effects to marine and human life. Polyethylene bags in oceans and rivers have life threatening effects on fish, turtles, jellyfish and various other aquatic species (Beaumont et al. 2019). Mismanagement of the plastic waste leads to the accumulation of plastics into the oceans and the fragmentation of such plastics results into production of micro-plastics (Wang et al. 2019). Recently, the microscopic remnants of plastic objects have been found in the intestinal tracts of various invertebrates, fish and birds that have a lot of impacts on their physiological activities, such as growth and development (Yu et al. 2018; Barboza et al. 2018). Plastic items also cause aesthetic problems, which is a nuisance rather than a hazard. Studies have shown that the used fishing nets and plastic bags thrown in the oceans and water bodies result in chocking and clogging of the respiratory tracts of aquatic animals such as fish, turtles and jellyfish. Micro and nano plastic particles are ingested by marine animals and remain in their digestive tract, which results in their retarded growth and improper physiological functions (Rhodes 2018; Muneer et al. 2021a).
Degradation of plastics by physical and chemical methods is expensive and is not safe for the environment because of the production of persistent organic pollutants (POPs) known as dioxins and furans (Alshemmari 2021). The byproducts of plastic degradation by physical and chemical methods are toxic irritants and harmful gaseous foams that not only damage the ozone layer in the upper atmosphere but also destroy the underground water resources, cause infertility of soil, prevent degradation of normal substances and are proved dangerous to human, animals and the ecosystem (Windsor et al. 2019). The ecofriendly methods to resolve plastic pollution is either biodegradation of plastics using microbes or bioplastic production using biomass that can be obtained from lignocellulosic plant materials or from polyesters produced by different microbes (Muneer et al. 2021b, c). Recent research suggests that there are a lot of microbial species that can degrade synthetic polymers at faster rates due to the presence of natural catalytic enzymes that break apart the strong hydrocarbon bonds in the plastic materials (Charnock 2021). The enzymes like lipases excreted by Rhizopus arrhizus, Achromobacter sp., Rhizopus delemar and Candida cylindracea and esterases from hog liver have potential to degrade poly(ε-caprolactone) and polyethylene adipate (Alshehrei 2017). It has been reported earlier that certain bacteria like Rhodococcus ruber, Brevibacillus borstelensis and some fungi such as Fusarium solani, Penicillium simplicissimum YK, Penicillium oxalicum NS4 (KU559906) and Penicillium chrysogenum NS10 (KU559907) have the potential to degrade natural and synthetic polyethylene (Ojha et al. 2017). Bioplastic research has also emerged as an alternative and ecofriendly solution to counter plastic pollution (Khan et al. 2019). Biobased polymers and polyesters such as polyhydroxyalkanoates (PHAs) and polyhydroxybutyrate (PHB) derived from microorganisms have shown a great potential to replace the conventional synthetic plastics. The search for new microbes, enzymes and catalytic systems is of global interest (Yuan et al. 2020). Due to the sophisticated techniques and expensiveness of the methods employed for the production of such biopolymers and polyesters, highly advanced research is required (Muneer et al. 2021b; Karan et al. 2019).
The aim of this study is to isolate and identify some novel bacterial strains that are capable of plastic degradation. The present study also aims to set up a biological method for the biodegradation of synthetic plastics using the indigenously isolated bacteria. Phylogenetic analysis of these isolated bacterial strains is also included as a major part of this study. The current study will pave the way for developing innovative strategies for plastic degradation and will moderate the environmental impacts of the plastics with an economically feasible approach. In the future, the isolated bacterial strains can be further investigated for the purification of the catalytic enzymes that have the potential to degrade synthetic plastics.
Materials and methods
Isolation of plastic-degrading microorganisms
To obtain plastic-degrading microbes samples of decomposing plastic bags from solid waste-dumping sites of Faisalabad, Pakistan, were collected. The collected plastic samples were dipped in 0.9% NaCl (saline) solution and gently shaken for 24 h to remove attached soil and dust particles. Under aseptic conditions, 10 ml of the saline solution was inoculated into 40 ml liquid carbon-free basal medium with plastic pieces as the sole carbon source for microorganisms. The cultures were incubated at 37 °C for 14 days. The microbes obtained from previous inoculation were streaked on Luria Broth (LB) medium solidified with 1.5% agar. These agar plates were incubated at 37 °C for 24 h. Colonies of microorganisms obtained from this step were streaked on separate LB-agar plates. The liquid carbon-free basal medium had KH2PO4 (0.7 g/L), K2HPO4 (0.7 g/L), MgSO4·7H2O (0.7 g/L), NH4NO3 (1.0 g/L), NaCl (0.005 g/L), FeSO4·7H2O (0.002 g/L), ZnSO4·7H2O (0.002 g/L), and MnSO4·H2O (0.001 g/L). The microbial colonies obtained were separately streaked on LB-agar plates to maintain the inoculum of pure cultures.
Screening of microbes having plastic degradation potential
Microorganisms from the inoculum were grown on growth media having 0.016% K2HPO4, 0.02% KH2PO4, 0.002% CaCl2.2H2O, 0.01% (NH4)2SO4, 0.02% MgSO4.7H2O, 0.005% ZnSO4.7H2O, 0.005% MnSO4.H2O, and 0.001% FeSO4.7H2O, which was solidified with 1.5% agar, having polystyrene powder as carbon source at final concentration of 0.1% (w/v).
Biodegradation assay
Pure cultures of microbes obtained by above screening (Sect. 2.2) were inoculated in mineral salt medium (MSM) along with plastic pieces as carbon source. The composition of MSM used was; 0.01% yeast extract, 0.016% K2HPO4, 0.02% KH2PO4, 0.002% CaCl2.2H2O, 0.01% (NH4)2SO4, 0.02% MgSO4.7H2O, 0.005% ZnSO4.7H2O, 0.005% MnSO4.H2O, 0.001% FeSO4.7H2O. Each reaction contains 45 mL of growth medium along with 5 mL of microbial culture and 50 mg (5 plastic pieces of 10 mg) of polyethylene pieces. The biodegradation assay was established for polyethylene. The microorganisms were cultured in MSM along with plastic pieces for 5 months at 30 °C and 120 rpm in a shaking incubator. During incubation, one plastic piece was separated from the reaction mixture (each flask) after interval of 30 days to determine the weight loss. The media was also refreshed after fifteen days. The percentage weight loss was calculated by following formula
FTIR analysis
After incubation of 150 days, the plastic pieces were subjected to FTIR analysis to determine the change in polymer bonds. For FTIR analysis samples were submitted to Central High Tech Laboratory, Government College University Faisalabad. Spectrum Two™ Perkin Elmer FTIR was used to carry out the FTIR analysis.
Identification of isolated microorganisms
Polymerase chain reaction and gel electrophoresis
The genomic DNA was extracted from the microbial cultures and were amplified separately for 16S rRNA gene (Mohan et al. 2016). For PCR followed the method as adopted by (Dey et al. 2020). DNA extracted from four different types of microbial cultures were subjected to 16S rRNA gene amplification and the PCR results were confirmed by gel electrophoresis. The gel electrophoresis of these strains confirmed the amplification of 16S rRNA. The sequencing and amplification was done using universal primers 785F (5’-GGATTAGATACCCTGGTA-3’), 907R (5’-CCGTCAATTCMTTTRAGTTT-3’) and 27F (5’-AGAGTTTGATCMTGGCTCAG-3’) and 1492R (5’-TACGGYTACCTTGTTACGACTT-3’), respectively (Hussein et al. 2015; Dey et al. 2020). Each illuminated band on gel represented the amplified 16S ribosomal gene of each strain. These samples were then delivered to a Korean company “Macrogen” to get sequences of 16S rRNA of AK-1, AK-2, AK-3 and AK-4.
Phylogenetic analysis
PCR products of four different strains obtained in the study, named as AK-1, AK-2, AK-3 and AK-4 were delivered to Macrogen Korea for 16S rRNA gene sequencing. After sequencing, results were visualized by chromas (https://technelysium.com.au/wp/chromaspro/) (Treves 2010). DNA Baser Sequence Assembler was used to assemble both the forward and reverse sequence files and the obtained sequences were subjected to NCBI BLAST separately to search the most similar sequences. Sequences that showed the highest similarity were selected for further analysis. The phylogenetic analysis was performed by a computational tool, MEGA7 (Kumar et al. 2016). ClustalW aligned the data sets for each strain, then the phylogenetic tree for each strain was constructed by Neighbor-Joining method (Li 2003). The evolutionary distances among the bacterial strains were computed using the Maximum Composite Likelihood method and were in the units of the number of base substitutions per site.
Results and discussions
Isolation of plastic-degrading microorganisms
Eight different types of colonies were observed (Supplementary Fig. A1) when microbial strains from enrichment culture were streaked on agar plates and allowed to grow for 24 h at 37 °C. Each obtained colony was separately streaked on agar plate to get pure cultures of microorganisms. When these eight strains were grown on media having polystyrene powder as sole carbon source, only four of these strains showed the ability to grow in the presence of polystyrene as sole carbon source in MSM (Supplementary Fig. A2).
The technique of utilizing plastic-degrading bags for isolation of microorganisms has been previously reported on several times. Kowalczyk et al. (2016) isolated a polyethylene degrading bacterium, Achromobacter xylosoxidans, from polyethylene bags collected from waste sites. The current study first utilized the enrichment culture technique to isolate the microorganisms having potential to grow in the presence of plastic pieces as sole carbon source (Kowalczyk et al., 2016). According to the hypothesis, only those microbes will survive in the given environment that can utilize plastics. Later, the potential of microorganisms obtained from the enrichment culture was checked by growing microbes on medium containing polystyrene powder as sole carbon source. Microbes that have the potential to utilize polystyrene actively grow on the provided medium. Hussein et al. (2015) utilized this method to screen plastic-degrading microbes. Mehmood et al. (2016) isolated the low-density polyethylene degrading strains of various microbes from the solid-waste dumpsite using enrichment culture technique.
Biodegradation assay
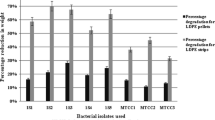
In biodegradation assay, five pieces of polyethylene sheets having weight of 10 mg each were added in the mineral salt medium along with the microorganisms. Four different strains were isolated in the study therefore, biodegradation assay for each strain was established separately. After 150 days of incubation, the percentage weight loss of 10 mg plastic piece noted for AK-1 (Stenotrophomonas sp.) was 32%, AK-2 (Serratia sp.) 40%, AK-3 (Pseudomonas sp.) and AK-4 (Pseudomonas sp.) was 21% each.
Phylogenetic analysis
A computational tool, MEGA7, was used to perform the phylogenetic analysis for each strain. The phylogenetic analysis of AK-1 (MW898426), AK-2 (MW898428), AK-3 (MW898430) and AK-4 (MW898432) showed that these strains belong to Stenotrophomonas sp., Serratia sp., and Pseudomonas sp. respectively (Fig. 1). Two of the bacterial strains that had the potential to degrade plastic i.e. Stenotrophomonas sp. and strain Serratia sp. AK-2 has been previously reported in bacterial-mediated biodegradation of linear low-density polyethylene (Azeko et al. 2015). Sekhar et al. (2016), identified polystyrene degrading microorganisms by sequencing 16S rRNA gene and have reported Stenotrophomonas sp. for the degradation of low-density polyethylene. Closely related type strains of P. stutzeri and S. marcescens are on the same line because these share almost 100% similarity.
FTIR analysis
FTIR was employed for the detection of change in functional group or in the chemical structure of plastic pieces because of incubation with isolated strains. As plastic pieces were exposed to four different strains AK-1 (Stenotrophomonas sp.), AK-2 (Serratia sp.) AK-3 (Pseudomonas sp.) and AK-4 (Pseudomonas sp.) separately and results were presented in various FTIR graphs (Fig. 2b–e). The change was observed in the structure of all plastic pieces that were exposed to all isolates. It was noted that AK-1 (Stenotrophomonas sp.) and AK-2 represented the increase in transmittance at 1459.7 cm−1 and 872.96 cm−1 that indicated the reduction in bonds. These peaks represented reduction in CH2 bends and skeletal C–C vibrations respectively. The change was also observed at 1107.66 cm−1. The change at 1651.47–1670 cm−1, at this point reduction in transmittance was observed. When plastic pieces were exposed to strain AK-2 (Serratia sp.), the increase in transmittance was observed at wavenumbers 2915.12, 2846.42, 1456.84, 874.39 cm−1. These peaks represented –C-H stretch, –C-H bend, CH2 bends and skeletal C–C vibrations respectively. Reduction in transmittance was observed at 1651.47–1670 cm−1, when plastic pieces were degraded by Serratia sp. the decrease in transmittance observed at 1558.45, 1398.17 cm−1, and 1103.36 cm−1 represented the increase in C = O, C–F, C–OH stretch. FTIR spectra of the plastic pieces treated with AK-3 and AK-4 strains showed less importance in terms of change in the plastic degradation when compared with the control (Fig. 2a–e.) of the same size and weight. By comparing all the spectra obtained from plastic pieces that were exposed to four different strains separately, it was concluded that the identified strains successfully reduced the C-H stretches and skeletal C–C vibrations. The range of approximately 1730–1650 cm−1 represented the presence of carbonyl group -C = O. The carbonyl group is formed by embodying an oxygen atom in the damaged structure of the polymer chains (Kowalczyk et al., 2016). In the present study, the decrease in transmittance was observed, which indicated the presence of carbonyl group that was a clear indication of the damaged structure of the polymer chains. The decrease in transmittance at 1100–1150 cm−1 was observed, that indicated the formation of new bonds (–C–O–C) due to weakness of C–C bonds. At 900–730 cm−1, the increase in transmittance was observed. The peaks at 1150–1075 cm−1 represented the presence of ether bonds –C -O –C. This bond was formed because of oxygen atom embodying between the C–C bonds weakened due to the high electronegativity of oxygen atom. The peaks at 900–735 cm−1 and less than 700 cm−1 represented the skeletal –C–C vibrations. The higher absorbance indicated the weakness in structure of the test compound. This is because of cleavage and formation of new bonds, which indicated the change in structure of the low-density polyethylene. The structure becomes looser that reduces the energy level and enhances atomic vibrations (Kowalczyk et al., 2016).
Conclusion
Plastics are synthetic polymers having large spectrum of applications. Degradation of plastics by microorganisms, i.e. biodegradation, is known for several years. The current study was performed for the addition of plastic-degrading microorganisms by isolating from waste plastic bags taken from the soil waste-dumping sites of Faisalabad, Pakistan. Indigenously isolated bacterial strains can be effectively investigated for plastic degradation. Nature has a huge reserve for microbial organisms it is therefore of vital importance that these shall be studied for their plastic biodegradation characteristics. Efficient and non-pathogenic microbial strains that have the catalytic power to degrade synthetic polymers can be released in plastic dumpsites to effectively degrade the waste and rescue the environment from the environmental problems generated because of the plastics. To use bacteria for biodegradation purposes, it is highly required that novel bacterial strains must be employed in plastic degradation research and investigated for their degradation efficacy.
References
Allen B, Coumoul X, Lacorte S (2019) Microplastic freshwater contamination: an issue advanced by science with public engagement. Environ Sci Pollut Res 26:16904–16905. https://doi.org/10.1007/s11356-019-05300-0
Alshehrei F (2017) Biodegradation of synthetic and natural plastic by microorganisms. J Appl Environ Microbiol 5:8–19. https://doi.org/10.12691/JAEM-5-1-2
Alshemmari H (2021) An overview of persistent organic pollutants along the coastal environment of Kuwait. Open Chem 19:149–156. https://doi.org/10.1515/chem-2021-0198
Azeko ST, Etuk-Udo GA, Odusanya OS, Malatesta K, Anuku N, Soboyejo WO (2015) Biodegradation of linear low density polyethylene by Serratia marcescens subsp. Marcescens and its cell free extracts. Waste Biomass Valoriz 6:1047–1057. https://doi.org/10.1007/s12649-015-9421-0
Barboza LGA, Vieira LR, Branco V, Figueiredo N, Carvalho F, Carvalho C, Guilhermino L (2018) Microplastics cause neurotoxicity, oxidative damage and energy-related changes and interact with the bioaccumulation of mercury in the European seabass, Dicentrarchus labrax (Linnaeus, 1758). Aquat Toxicol 195:49–57. https://doi.org/10.1016/j.aquatox.2017.12.008
Beaumont NJ, Aanesen M, Austen MC, Börger T, Clark JR, Cole M, Hooper T, Lindeque PK, Pascoe C, Wyles KJ (2019) Global ecological, social and economic impacts of marine plastic. Mar Pollut Bull 142:189–195. https://doi.org/10.1016/j.marpolbul.2019.03.022
Charnock C (2021) Norwegian soils and waters contain mesophilic, plastic-degrading bacteria. Microorganisms 9:1–18. https://doi.org/10.3390/microorganisms9010094
Dey AS, Bose H, Mohapatra B, Sar P (2020) Biodegradation of unpretreated low-density polyethylene (LDPE) by Stenotrophomonas sp. and Achromobacter sp., isolated from waste dumpsite and drilling fluid. Front Microbiol 11:1–15. https://doi.org/10.3389/fmicb.2020.603210
Godfrey L (2019) Waste plastic, the challenge facing developing countries—ban it, change It, collect It? Recycling 4:3. https://doi.org/10.3390/recycling4010003
Hahladakis JN, Velis CA, Weber R, Iacovidou E, Purnell P (2018) An overview of chemical additives present in plastics: migration, release, fate and environmental impact during their use, disposal and recycling. J Hazard Mater 344:179–199. https://doi.org/10.1016/j.jhazmat.2017.10.014
Hussein AA, Al-Mayaly IK, Kudier SH (2015) Screening and identification of low density polyethylene (LDPE) degrading bacteria from contaminated soil with plastic wastes. Mesop Environ J 1:1–14
Ivanova L, Sokolov K, Kharitonova G (2018) Plastic pollution tendencies of the Barents Sea and adjacent waters under the climate change. Arct North 32:121–145. https://doi.org/10.17238/issn2221-2698.2018.32.121
Karan H, Funk C, Grabert M, Oey M, Hankamer B (2019) Green bioplastics as part of a circular bioeconomy. Trends Plant Sci 24:237–249. https://doi.org/10.1016/j.tplants.2018.11.010
Khan FI, Aktar L, Islam T, Saha ML (2019) Isolation and identification of indigenous poly-β-hydroxybutyrate (PHB) producing bacteria from different waste materials. Plant Tissue Cult Biotechnol 29:15–24. https://doi.org/10.3329/ptcb.v29i1.41975
Koller M (2019) Switching from petro-plastics to microbial polyhydroxyalkanoates (PHA): the biotechnological escape route of choice out of the plastic predicament? EuroBiotech J 3:32–44. https://doi.org/10.2478/ebtj-2019-0004
Kowalczyk A, Chyc M, Ryszka P, Latowski D (2016) Achromobacter xylosoxidans as a new microorganism strain colonizing high-density polyethylene as a key step to its biodegradation. Environ Sci Pollut Res 23:11349–11356. https://doi.org/10.1007/s11356-016-6563-y
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Li KB (2003) ClustalW-MPI: clustalW analysis using distributed and parallel computing. Bioinformatics 19:1585–1586. https://doi.org/10.1093/bioinformatics/btg192
Mehmood CT, Qazi IA, Hashmi I, Bhargava S, Deepa S (2016) Biodegradation of low density polyethylene (LDPE) modified with dye sensitized titania and starch blend using Stenotrophomonas pavanii. Int Biodeterior Biodegrad 113:276–286. https://doi.org/10.1016/j.ibiod.2016.01.025
Mohan AJ, Sekhar VC, Bhaskar T, Nampoothiri KM (2016) Microbial assisted high impact polystyrene (HIPS) degradation. Bioresour Technol 213:204–207. https://doi.org/10.1016/j.biortech.2016.03.021
Muneer F, Rasul I, Azeem F, Siddique MH, Zubair M, Nadeem H (2020) Microbial polyhydroxyalkanoates (PHAs): efficient replacement of synthetic polymers. J Polym Environ 28:2301–2323. https://doi.org/10.1007/s10924-020-01772-1
Muneer F, Azam MH, Zubair M, Farooq T, Ibrahim M, Rasul I, Afzal M, Ahmad A, Nadeem H (2021a) Remediation of water pollution by plastics. Environ Chem Sustain World 54:89–117. https://doi.org/10.1007/978-3-030-52395-4_3
Muneer F, Hussain S, Sidra-tul-Muntaha MR, Nadeem H (2021b) Plastics versus bioplastics. Mater Res Forum LLC. https://doi.org/10.21741/9781644901335-9
Muneer F, Nadeem H, Arif A, Zaheer W (2021c) Bioplastics from biopolymers : an eco-friendly and sustainable solution of plastic pollution. Polym Sci Ser C 63:47–63. https://doi.org/10.1134/S1811238221010057
Ojha N, Pradhan N, Singh S, Barla A, Shrivastava A, Khatua P, Rai V, Bose S (2017) Evaluation of HDPE and LDPE degradation by fungus, implemented by statistical optimization. Sci Rep 7:1–13. https://doi.org/10.1038/srep39515
Palm GJ, Reisky L, Böttcher D, Müller H, Michels EAP, Walczak MC, Berndt L, Weiss MS, Bornscheuer UT, Weber G (2019) Structure of the plastic-degrading Ideonella sakaiensis MHETase bound to a substrate. Nat Commun 10:1–10. https://doi.org/10.1038/s41467-019-09326-3
Rhodes CJ (2018) Plastic pollution and potential solutions. Sci Prog 101:207–260. https://doi.org/10.3184/003685018x15294876706211
Sekhar VC, Nampoothiri KM, Mohan AJ, Nair NR, Bhaskar T, Pandey A (2016) Microbial degradation of high impact polystyrene (HIPS), an e-plastic with decabromodiphenyl oxide and antimony trioxide. J Hazard Mater 318:347–354. https://doi.org/10.1016/j.jhazmat.2016.07.008
Treves DS (2010) Review of three DNA analysis applications for use in the microbiology or genetics classroom. J Microbiol Biol Educ 11:186–186. https://doi.org/10.1128/jmbe.v11i2.205
Villarrubia-Gómez P, Cornell SE, Fabres J (2018) Marine plastic pollution as a planetary boundary threat–the drifting piece in the sustainability puzzle. Mar Policy 96:213–220. https://doi.org/10.1016/j.marpol.2017.11.035
Wang MH, He Y, Sen B (2019) Research and management of plastic pollution in coastal environments of China. Environ Pollut 248:898–905. https://doi.org/10.1016/j.envpol.2019.02.098
Windsor FM, Durance I, Horton AA, Thompson RC, Tyler CR, Ormerod SJ (2019) A catchment-scale perspective of plastic pollution. Glob Chang Biol 25:1207–1221. https://doi.org/10.1111/gcb.14572
Yoshida S, Hiraga K, Takehana T, Taniguchi I, Yamaji H, Maeda Y, Toyohara K, Miyamoto K, Kimura Y, Oda K (2016) A bacterium that degrades and assimilates poly(ethylene terephthalate) material complementario. Science 351:1196–1199. https://doi.org/10.1126/science.aad6359
Yu Y, Zhou D, Li Z, Zhu C (2018) Advancement and challenges of microplastic pollution in the aquatic environment: a review. Water Air Soil Pollut 229:2–18. https://doi.org/10.1007/s11270-018-3788-z
Yuan J, Ma J, Sun Y, Zhou T, Zhao Y, Yu F (2020) Microbial degradation and other environmental aspects of microplastics/plastics. Sci Total Environ 715:136968. https://doi.org/10.1016/j.scitotenv.2020.136968
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nadeem, H., Alia, K.B., Muneer, F. et al. Isolation and identification of low-density polyethylene degrading novel bacterial strains. Arch Microbiol 203, 5417–5423 (2021). https://doi.org/10.1007/s00203-021-02521-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02521-1