Abstract
A Gram-staining-negative, non-spore-forming, non-flagellated, rod-shaped, catalase- and oxidase-negative bacterium, designated as Gsoil 193T, was isolated from the soil of ginseng field in Pocheon province, South Korea. 16S rRNA gene sequence analysis showed that strain Gsoil 193T belonged to family Xanthomonadaceae and was most closely related to Lysobacter daecheongensis KCTC 12,600T (96.4 %), Lysobacter panaciterrae KCTC 12601T (96.3 %), Lysobacter dokdonensis DSM 17958T (96.3 %) and Lysobacter oligotrophicus JCM 18257T (95.6 %). Strain Gsoil 193T grew at temperatures between 20 and 30 °C with an optimum of 30 °C. The pH range for growth was 5–9 pH (optimum 6–7 pH). The predominant respiratory quinone was ubiquinone Q-8 and a fatty acid profile with iso-C15:0, iso-C16:0 and summed feature 9 (iso-C17:1 ω9c/C16:0 10-methyl) as the major fatty acids supported the affiliation of strain Gsoil 193T to the genus Lysobacter. The genomic DNA G+C content was 64.8 mol %. On the basis of the genotypic analysis, physiological and chemotaxonomic results indicate that strain Gsoil 193T represents a novel species of the genus Lysobacter, for which the name Lysobacter pocheonensis sp. nov. is proposed. The type strain is Gsoil 193T (= DSM 18338T = KCTC 12624T).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Lysobacter was first described by Christensen and Cook (1978), and the description was emended by Park et al. (2008). Members of the genus Lysobacter, in the family Xanthomonadaceae, contain ubiquinone Q-8 as the major respiratory quinone and have a high DNA G+C content. (Park et al. 2008; Wang et al. 2009). Members of the genus Lysobacter are Gram-negative, aerobic, non-fruiting, gliding organisms and with colonies that were very mucoid and creamy, pink or yellow-brownish in color. Members of this genus are strongly proteolytic and were found to have great potential antibiotic compounds against human pathogens (Ahmed et al. 2003; Hashizume et al. 2004) and may be used as biocontrol agents for plant fungal pathogens (Islam et al. 2005; Park et al. 2008). There are 33 recognized species in the genus Lysobacter (http://www.bacterio.net), and most of the species of this genus were isolated from soil, Lysobacter thermophilus (Wei et al. 2012), Lysobacter arseniciresistens (Luo et al. 2012), Lysobacter panacisoli (Choi et al. 2014), Lysobacter terrae (Ngo et al. 2014) and Lysobacter novalis [(Singh et al. 2015) IJSEM in press]. Some of them are also isolated from sludge: Lysobacter caeni (Ye et al. 2015) and Lysobacter mobilis (Yang et al. 2015). Until now 19 species were isolated from variety of soil samples.
During the course of a study on the culturable aerobic and facultative anaerobic bacterial community living in the soil of a ginseng field in Pocheon province [(37°91′96″N, 127°22.4′59.1″E) May 2005, South Korea], a large number of bacterial strains were isolated on modified R2A agar plates (Ten et al. 2009). Here, we report on the taxonomic characterization of one of these strains, designated Gsoil 193T, which appeared to be a member of the genus Lysobacter.
Materials and methods
Isolation of bacterial strain
Strain Gsoil 193T was cultured routinely on R2A agar at 30 °C and preserved, as a suspension in R2A broth with 20 % (v/v) glycerol, at −70 °C. The type strains, Lysobacter daecheongensis KCTC 12600T, Lysobacter panaciterrae KCTC 12601T, Lysobacter dokdonensis DSM 17958T and Lysobacter oligotrophicus JCM 18257T, respectively, were obtained from our laboratory’s storage box or culture collections (KCTC and JCM), grown under the same conditions and used as reference strains with Gsoil 193T.
Phylogenetic tree construction and DNA G+C content analysis
The genomic DNA of strain Gsoil 193T was extracted using commercial DNA extraction kit (Solgent) and a PCR assay with the universal bacterial primer pair 9F and 1512R to amplify the 16S rRNA gene (Weisburg et al. 1991). The purified PCR products were sequenced by Solgent Co. Ltd (Im et al. 2010). The almost complete (1501 nt) sequence of the 16S rRNA gene was assembled using SeqMan software (DNASTAR) and compared with the 16S rRNA gene sequences of related taxa, which were obtained from the GenBank database or [http://www.ezbiocloud.net/eztaxon; Kim et al. (2012)]. Multiple alignments were made using CLUSTAL_X program (Thompson et al. 1997) with gaps edited using the BioEdit program (Hall 1999). Evolutionary distances were calculated using the Kimura two-parameter model (Kimura 1983), neighbor-joining (Saitou and Nei 1987), maximum-parsimony (Fitch 1971) and maximum-likelihood method using the MEGA6 program (Tamura et al. 2013) with bootstrap values based on 1000 replications (Felsenstein 1985).
For the measurement of the DNA G+C content, the genomic DNA of the novel strain was extracted and purified as described by Moore and Dowhan (1995) and enzymatically degraded into nucleosides, and the base composition was determined as described by Mesbah et al. (1989) using a reverse-phase HPLC.
Physiological and biochemical characteristics
The Gram reaction was determined using the non-staining method, as described by Buck (1982). Cell morphology was observed under a Nikon light microscope at ×1000, with cells grown on R2A agar for 2 days at 30 °C. Cell morphology was examined with the scanning electron microscope (Hitachi SU-3500), using cells grown for 2 days at 30 °C on R2A agar (BD). Gliding motility was investigated using the hanging drop method described by Bernardet et al. (2002). Catalase and oxidase tests were performed as outlined by Cappuccino and Sherman (2002). Biochemical tests were carried out by using API 20NE, API ID 32GN and API ZYM kits according to the instructions of the manufacturer (bioMérieux). Tests for degradation of DNA (using DNase agar from Scharlau, with DNase activity by flooding plates with 1 N HCl), casein, starch (Atlas 1993), olive oil (Kouker and Jaeger 1987), xylan and carboxyl methyl cellulose (Ten et al. 2004) were performed and evaluated after 7 days. Growth at different temperatures (4, 10, 15, 25, 30, 37 and 42 °C) and various pH values (pH 4–10 at intervals of 0.5 pH units) was assessed after 5-day incubation at 30 °C. Three different buffers (final concentration, 50 mM) were used to adjust the pH of R2A broth. Acetate buffer was used for pH 4.0–5.5, phosphate buffer was used for pH 6.0–8.0, and Tris buffer was used for pH 8.5–10.0. Salt tolerance was tested on R2A medium supplemented with 1–10 % (w/v at intervals of 1 % unit) NaCl, and growth was assessed after 7 days of incubation. Growth on nutrient agar (NA, BD), trypticase soy agar (TSA, BD), LB agar (BD) and MacConkey agar (BD) was also evaluated at 30 °C.
Chemotaxonomic analysis
Analysis of respiratory quinone
Cells were grown in R2A broth at 30 °C, shaken at 160 rpm for 2 days and then centrifuged. The pallets were dissolved in chloroform/methanol (2/1, v/v). Isoprenoid quinone was extracted, and afterward concentrated at 40 °C using vacuum rotary evaporator, then the residue was subsequently extracted with n-hexane/water (1/1, v/v). The crude n-hexane–quinone solution was purified using Sep-Pak Vac Cartridges Silica (Waters) and analyzed by HPLC as previously described (Hiraishi et al. 1996).
Fatty acids analysis
Cellular fatty acid profile was determined for strain grown on R2A agar (BD) for 48 h at 30 °C. The fatty acids were extracted, methylated and saponified by the described method of Sherlock Microbial Identification System (MIDI). Then, it was analyzed by capillary GLC (Hewlett Packard 6890) using the TSBA library (version 6.1) Sasser (1990).
Results and discussion
Phylogenetic tree and DNA G+C content
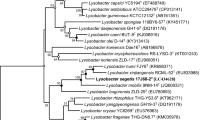
The 16S rRNA gene sequence of strain Gsoil 193T determined in this study is a continuous stretch of 1,501 bp, which has been deposited in the GenBank database (accession number EU273938). A sequence similarity calculation from using the EzTaxon-e server indicated that strain Gsoil 193T shows highest sequence similarity Lysobacter daecheongensis KCTC 12600T (96.4 %), Lysobacter panaciterrae KCTC 12601T (96.3 %), Lysobacter dokdonensis DSM 17958T (96.3 %) and Lysobacter oligotrophicus JCM 18257T (95.6 %). However, in the phylogenetic tree constructed by the neighbor-joining algorithm (Fig. 1), strain Gsoil 193T clustered with Lysobacter dokdonensis DSM 17958T. This phylogenetic relationship was also confirmed in the trees generated with the maximum-likelihood (Supplementary Fig. S2) and maximum-parsimony algorithms. Strain Gsoil 193T and Lysobacter dokdonensis DSM 17958T formed a monophyletic group with a bootstrap high value (80 %), which was supported by all three trees making methods used in this study.
Phylogenetic relationship of strain Gsoil 193T with recognized Lysobacter species. The tree was constructed by using the neighbor-joining method based on 16S rRNA gene sequences. Bootstrap values (expressed as percentages of 1000 replications) >60 % are shown at branch points. Nevskia ramosa Soe1T (AJ001010) was used as an out group. Filled circles indicate that the corresponding nodes were also recovered in the tree generated with maximum-parsimony algorithms and maximum likelihood. Bar 0.02 substitutions per nucleotide position
On the basis of these phylogenetic results, L. dokdonensis DSM 17958T, L. daecheongensis KCTC 12600T, L. panaciterrae KCTC 12601T and L. oligotrophicus JCM 18257T were selected as the closest recognized neighbors of strain Gsoil 193T.
The genomic DNA G+C content of strain Gsoil 193T was 64.8 mol %, which lies within the range observed for members of the genus Lysobacter (61.7–69.2 mol %, respectively).
Physiological biochemical tests
Cells of strain Gsoil 193T were Gram-reaction-negative, aerobic, non-spore-forming and non-motile. Cells were rod shaped (measuring 0.3–0.4 and 2.5–5 μm) shown in Supplementary Fig. S1. Colonies grown on the R2A agar plates for 2 days at 30 °C were convex, translucent, circular, light yellowish in color and 0.3–1.0 mm in diameter after incubation at 30 °C for 2 days. The morphological, physiological and biochemical characteristics of strain Gsoil 193T are given in the species description and in Table 1. Some physiological characteristics of strain Gsoil 193T compared with those of the three reference type strains are given in Table 1.
Quinone and fatty acids
The fatty acid profile of strain Gsoil 193T (Table 2) was compared with those of the type strains of recognized Lysobacter species. The major components were branched fatty acids iso-C15:0 (28.9 %), iso-C16:0 (25.2 %) and summed feature 9 [(iso-C17:1 ω9c/C16:0 10-methyl), 12.4 %], as the major, which is a profile typical of members of the genus Lysobacter (Bae et al. 2005; Weon et al. 2006, 2007; Romanenko et al. 2007). However, some minor qualitative and quantitative differences in fatty acid content could be observed between strain Gsoil 193T and its phylogenetically closest relatives. Strain Gsoil 193T contained ubiquinone Q-8 as the major respiratory quinone. These data are in good agreement with those of other members of the genus Lysobacter (Bae et al. 2005; Weon et al. 2006, 2007; Park et al. 2008; Romanenko et al. 2007; Yassin et al. 2007).
The results obtained from the phenotypic and phylogenetic characterizations indicated that strain Gsoil 193T belongs to the genus Lysobacter. The phylogenetic distinctiveness confirmed that this isolate represents a species that is distinct from recognized Lysobacter species. There are some phenotypic differences between strain Gsoil 193T and phylogenetically related Lysobacter species (Table 1). Therefore, on the basis of the data presented, strain Gsoil 193T should be classified within the genus Lysobacter as representing a novel species, for which the name Lysobacter pocheonensis sp. nov. is proposed.
Description of Lysobacter pocheonensis sp. nov
Lysobacter pocheonensis (po.che.on.en´sis. N.L. mac. adj. pocheonensis pertaining to Pocheon province in South Korea, the location of the soil sample from which the type strain was isolated).
Cells are Gram-reaction-negative, aerobic, non-motile, non-spore-forming, rod-shaped (0.3–0.4 μm in diameter and 2.5–5 μm in length), catalase- and oxidase-negative. Colonies grown on the R2A agar plates for 2 days at 30 °C were convex, translucent, circular, light yellowish in color and 0.3–1.0 mm in diameter. Growth occurs at 20–30 °C with pH 5–9 (optimum at 30 °C with pH 6–7.0) without additional NaCl supplement. Growth is inhibited in the presence of 0.5 % (w/v) NaCl. Hydrolyze gelatin and casein, but not chitin, starch, CM-cellulose, xylan, olive oil and DNase. Using the API kit (API 32 GN, API ZYM and API 20 NE), the following substrates are utilized: urea, alkaline phosphatase, esterase, esterase, lipase, leucine arylamidase, valine arylamidase, cysteine arylamidase, trypsin, α-chymotrypsin, acid phosphatase, gelatin and naphthol-AS-BI-phosphohydrolase. List of all negative traits of commercial kits is given in Table S1. Ubiquinone 8 (Q-8) is the predominant quinone. The major fatty acids are iso-C15:0, iso-C16:0 and summed feature 9 (iso-C17:1 ω9c). The G+C content of genomic DNA is 64.8 mol %.
The type strain Gsoil 193T (= DSM 18338T = KCTC 12624T) was isolated from soil from a ginseng field in Pocheon Province, South Korea.
References
Ahmed K, Chohnan S, Ohashi H, Hirata T, Masaki T, Sakiyama F (2003) Purification, bacteriolytic activity, and specificity of β-lytic protease from Lysobacter sp. IB-9374. J Biosci Bioeng 95:27–34
Atlas RM (1993) Handbook of microbiological media, 3rd edn. CRC Press, Boca Raton, FL
Bae HS, Im WT, Lee ST (2005) Lysobacter concretionis sp. nov., isolated from anaerobic granules in an upflow anaerobic sludge blanket reactor. Int J Syst Evol Microbiol 55:1155–1161
Bernardet JF, Nakagawa Y, Holmes B (2002) Proposed minimal standards for describing new taxa of the family Flavobacteriaceae and emended description of the family. Int J Syst Evol Microbiol 52:1049–1070
Buck JD (1982) Nonstaining (KOH) method for determination of Gram reactions of marine bacteria. Appl Environ Microbiol 44:992–993
Cappuccino JG, Sherman N (2002) Microbiology: a laboratory manual, 6th edn. Benjamin Cummings, San Francisco
Choi JH, Seok JH, Cha JH, Cha CJ (2014) Lysobacter panacisoli sp. nov., isolated from ginseng soil. Int J Syst Evol Microbiol 64:2193–2197
Christensen P, Cook FD (1978) Lysobacter, a new genus of nonfruiting, gliding bacteria with a high base ratio. Int J Syst Bacteriol 28:367–393
Felsenstein J (1985) Confidence limit on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Fukuda W, Kimura T, Araki S, Miyoshi Y, Atomi H, Imanaka T (2013) Lysobacter oligotrophicus sp. nov., isolated from an Antarctic freshwater lake in Antarctica. Int J Syst Evol Microbiol 63:3313–3318
Hall TA (1999) BioEdit: a user–friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hashizume H, Hattori S, Igarashi M, Akamatsu Y (2004) Tripropeptin E, a new tripropeptin group antibiotic produced by Lysobacter sp. BMK333-48F3. J Antibiot (Tokyo) 57:394–399
Hiraishi A, Ueda Y, Ishihara J, Mori T (1996) Comparative lipoquinone analysis of influent sewage and activated sludge by high-performance liquid chromatography and photodiode array detection. J Gen Appl Microbiol 42:457–469
Im WT, Liu QM, Yang JE, Kim MS, Kim SY, Lee ST, Yi TH (2010) Panacagrimonas perspica gen. nov., sp. nov., a novel member of Gammaproteobacteria isolated from soil of a ginseng field. J Microbiol 48:262–266
Islam MT, Hashidoko Y, Deora A, Ito T, Tahara S (2005) Suppression of damping-off disease in host plants by the rhizoplane bacterium Lysobacter sp. strain SB-K88 is linked to plant colonization and antibiosis against soilborne peronosporomycetes. Appl Environ Microbiol 71:3786–3796
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
Kouker G, Jaeger KE (1987) Specific and sensitive plate assay for bacterial lipases. Appl Environ Microbiol 53:211–213
Luo G, Shi Z, Wang G (2012) Lysobacter arseniciresistens sp. nov., an arsenite-resistant bacterium isolated from iron-mined soil. Int J Syst Evol Microbiol 62:1659–1665
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high–performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Moore DD, Dowhan D (1995) Preparation and analysis of DNA. In: Ausubel FW, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (eds) Current protocols in molecular biology. Wiley, New York, pp 2–11
Ngo HTT, Won K, Du J, Son HM, Park Y, MooChang K, Yi TH (2014) Lysobacter terrae sp. nov. isolated from Aglaia odorata rhizosphere soil. Int J Syst Evol Microbiol 65:587–592
Oh KH, Kang SJ, Jung YT, Oh TK, Yoon JH (2011) Lysobacter dokdonensis sp. nov., isolated from soil. Int J Syst Evol Microbiol 61:1089–1093
Park JH, Kim R, Aslam Z, Jeon CO, Chung YR (2008) Lysobacter capsici sp. nov., with antimicrobial activity, isolated from the rhizosphere of pepper, and emended description of the genus Lysobacter. Int J Syst Evol Microbiol 58:387–392
Romanenko LA, Uchino M, Tanaka N, Frolova GM, Mikhailov VV (2007) Lysobacter spongiicola sp. nov., isolated from a deep-sea sponge. Int J Syst Evol Microbiol 58:370–374
Saitou N, Nei M (1987) The neighbor–joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical Note 101, MIDI Inc., Newark
Singh H, Du J, Won KH, Yang JE, Akter S, Kim KY, Yi TH (2015) Lysobacter novalis sp. nov. isolated from fallow farmland soil. Int J Syst Evol Microbiol 65:313–3136 (Epub ahead of print)
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Ten LN, Im WT, Kim MK, Kang MS, Lee ST (2004) Development of a plate technique for screening of polysaccharide-degrading microorganisms by using a mixture of insoluble chromogenic substrates. J Microbiol Methods 56:375–382
Ten LN, Jung HM, Im WT, Yoo SA, Lee ST (2008) Lysobacter daecheongensis sp. nov., isolated from sediment of stream near the Daechung dam in South Korea. J Microbiol 5:519–524
Ten LN, Jung HM, Im WT, Yoo SA, Oh HM, Lee ST (2009) Lysobacter panaciterrae sp. nov., isolated from soil of a ginseng field. Int J Syst Evol Microbiol 59:958–963
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wang Y, Dai J, Zhang L, Luo X, Li Y, Chen G, Tang Y, Meng Y, Fang C (2009) Lysobacter ximonensis sp. nov., isolated from soil. Int J Syst Evol Microbiol 59:786–789
Wei DQ, Yu TT, Yao JC, Zhou EM, Song ZQ, Yin YR, Ming H, Tang SK, Li WJ (2012) Lysobacter thermophilus sp. nov., isolated from a geothermal soil sample in Tengchong, south-west China. Antonie Van Leeuwenhoek 102:643–651
Weon HY, Kim BY, Baek YK, Yoo SH, Kwon SW, Stackebrandt E, Go SJ (2006) Two novel species, Lysobacter daejeonensis sp. nov. and Lysobacter yangpyeongensis sp. nov., isolated from Korean greenhouse soils. Int J Syst Evol Microbiol 56:947–951
Weon HY, Kim BY, Kim MK, Yoo SH, Kwon SW, Go SJ, Stackebrandt E (2007) Lysobacter niabensis sp. nov. and Lysobacter niastensis sp. nov., isolated from greenhouse soils in Korea. Int J Syst Evol Microbiol 57:548–551
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Yang SZ, Feng GD, Zhu HH, Wang YH (2015) Lysobacter mobilis sp. nov., isolated from abandoned lead-zinc ore. Int J Syst Evol Microbiol 65:833–837
Yassin AF, Chen WM, Hupfer H, Siering C, Kroppenstedt RM, Arun AB, Lai WA, Shen FT, Rekha PD, Young CC (2007) Lysobacter defluvii sp. nov., isolated from municipal solid waste. Int J Syst Evol Microbiol 57:1131–1136
Ye XM, Chu CW, Shi C, Zhu JC, He Q, He J (2015) Lysobacter caeni sp. nov., isolated from the sludge of a pesticide manufacturing factory. Int J Syst Evol Microbiol 65:845–850
Acknowledgments
This work was supported by the project on survey and excavation of Korean indigenous species of the National Institute of Biological Resources (NIBR) under the Ministry of Environment.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
The NCBI GenBank accession number for 16S rRNA gene sequence of strain Gsoil 193T is EU273938.
Electronic supplementary material
Below is the link to the electronic supplementary material.
203_2016_1214_MOESM2_ESM.pptx
Maximum-likelihood tree based on 16S rRNA gene sequences showing the relationships between strain Gsoil 193T and other related type species. Numbers at nodes (over 60 % are shown) represent percentages of bootstrap support based on a maximum–likelihood analysis of 1000 resampled datasets. Nevskia ramosa Soe1T (AJ001010) was used as outgroup. (PPTX 105 kb)
203_2016_1214_MOESM3_ESM.pptx
All negative traits of Gsoil 193T for the utilization of following substrates in API kits (API 20NE, 32GN and ZYM) system. (PPTX 33 kb)
Rights and permissions
About this article
Cite this article
Siddiqi, M.Z., Im, WT. Lysobacter pocheonensis sp. nov., isolated from soil of a ginseng field. Arch Microbiol 198, 551–557 (2016). https://doi.org/10.1007/s00203-016-1214-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-016-1214-8