Abstract
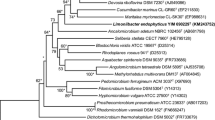
A taxonomic study was carried out on Gsoil 142T, a bacterial strain isolated from the soil collected in a ginseng field in Pocheon province, South Korea. Comparative 16S rRNA gene sequence studies showed a clear affiliation of this bacterium to the Gammaproteobacteria, and it was most closely related to Hydrocarboniphaga effusa ATCC BAA 332T (94.4%, 16S rRNA gene sequence similarity), Nevskia ramosa DSM 11499T (94.1%) and Alkanibacter difficilis MN154.3T (92.0%). Strain Gsoil 142T was a Gram-negative, strictly aerobic, motile, and rod-shaped bacterium. The G+C content of the genomic DNA was 69.9% and predominant ubiquinone was Q-8. Major fatty acids were summed feature 8 (C18:1 ω7c and/or ω6c, 36.3%), summed feature 3 (iso-C15:0 2-OH and/or C16:1 ω7c, 20.6%) and C16:0 (17.4%). The major polar lipids detected in strain Gsoil 142T were phosphatidylethanolamine, phosphatidylglycerol, diphosphatidylglycerol, and an unknown glycolipid. On the basis of polyphasic evidence, it is proposed that strain Gsoil 142T should be placed in a novel genus and species, for which the name Panacagrimonas perspica gen. nov., sp. nov. is proposed. The type strain is Gsoil 142T (= KCTC 12982T = LMG 23239T).
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Atlas, R.M. 1993. Handbook of Microbiological Media. CRC Press, Boca Raton, Florida, USA.
Buck, J.D. 1982. Nonstaining (KOH) method for determination of Gram reactions of marine bacteria. Appl. Environ. Microbiol. 44, 992–993.
Chun, J., J.-H. Lee, Y. Jung, M. Kim, S. Kim, B.K. Kim, and Y.W. Lim. 2007. EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int. J. Syst. Evol. Microbiol. 57, 2259–2261.
Felsenstein, J. 1985. Confidence limit on phylogenies: an approach using the bootstrap. Evolution 39, 783–791.
Fitch, W.M. 1971. Toward defining the course of evolution: minimum change for a specific tree topology. Syst. Zool. 20, 406–416.
Hall, T.A. 1999. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98.
Hiraishi, A., Y. Ueda, J. Ishihara, and T. Mori. 1996. Comparative lipoquinone analysis of influent sewage and activated sludge by high-performance liquid chromatography and photodiode array detection. J. Gen. Appl. Microbiol. 42, 457–469.
Im, W.-T., H.-M. Jung, Y.-S. Cui, Q.-M. Liu, S.-L. Zhang, and S.-T. Lee. 2005. Cultivation of the three hundreds of bacterial species from soil of a ginseng field and mining the novel lineage bacteria. In Proceedings of the International Meeting of the Federation of Korean Microbiological Societies, abstract A035, p. 169. Seoul: Federation of Korean Microbiological Societies.
Kimura, M. 1983. The Neutral Theory of Molecular Evolution. Cambridge: Cambridge University Press, Cambridge, New York, N.Y., USA.
Kouker, G. and K.-E. Jaeger. 1987. Specific and sensitive plate assay for bacterial lipases. Appl. Environ. Microbiol. 53, 211–213.
Kumar, S., J. Dudley, M. Nei, and K. Tamura. 2008. MEGA: A biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 9, 299–306.
Mesbah, M., U. Premachandran, and W. Whitman. 1989. Precise measurement of the G+C content of deoxyribonucleic acid by high performance liquid chromatography. Int. J. Syst. Bacteriol. 39, 159–167.
Minnikin, D.E., P.V. Patel, L. Alshamaony, and M. Goodfellow. 1977. Polar lipid composition in the classification of Nocardia and related bacteria. Int. J. Syst. Bacteriol. 27, 104–117.
Moore, D.D. and D. Dowhan. 1995. Preparation and analysis of DNA, pp. 2–11. In F.W. Ausubel, R. Brent, R.E. Kingston, D.D. Moore, J.G. Seidman, J.A. Smith, and K. Struhl (eds.), Current Protocols in Molecular Biology. Wiley, New York, N.Y., USA.
Palleroni, N.J., A.M. Port, H.-K. Chang, and G.J. Zylstra. 2004. Hydrocarboniphaga effusa gen. nov., sp. nov., a novel member of the γ-Proteobacteria active in alkane and aromatic hydrocarbon degradation. Int. J. Syst. Evol. Microbiol. 54, 1203–1207.
Saitou, N. and M. Nei. 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425.
Sasser, M. 1990. Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical Note 101. MIDI Inc., Newark, DE, USA.
Stürmeyer, H., J. Overmann, H.D. Babenzien, and H. Cypionka. 1998. Ecophysiological and phylogenetic studies of Nevskia ramosa in pure culture. Appl. Environ. Microbiol. 64, 1890–1894.
Ten, L.N., W.-T. Im, M.-K. Kim, M.-S. Kang, and S.-T. Lee. 2004. Development of a plate technique for screening of polysaccharide-degrading microorganisms by using a mixture of insoluble chromogenic substrates. J. Microbiol. Methods 56, 375–382.
Ten, L.N., H.-M. Jung, S.-A. Yoo, W.-T. Im, and S.-T. Lee. 2008. Lysobacter daecheongensis sp. nov., isolated from sediment of stream near the Daechung dam in South Korea. J. Microbiol. 46, 519–524.
Thompson, J.D., T.J. Gibson, F. Plewniak, F. Jeanmougin, and D.G. Higgins. 1997. The Clustal_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 24, 4876–4882.
Tschech, A. and N. Pfennig. 1984. Growth yield increase linked to caffeate reduction in Acetobacterium woodii. Arch. Microbiol. 137, 163–167.
Widdel, F. and F. Bak. 1992. Gram-negative mesophilic sulfatereducing bacteria, The Prokaryotes, 2nd ed., pp. 3352–3378. In A. Balows, H.G. Trüper, M. Dworkin, W. Harder, and K.H. Schleifer (eds.). Springer, New York, N.Y., USA.
Widdel, F., G. Kohring, and F. Mayer. 1983. Studies in dissimilatory sulfate-reducing bacteria that decompose fatty acids. III. Characterization of the filamentous gliding Desulfonema limicola gen. nov. sp. nov., and Desulfonema magnum sp. nov. Arch. Microbiol. 134, 286–294.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplemental material for this article may be found at http://www.springerlink.com/content/120956
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Im, WT., Liu, QM., Yang, JE. et al. Panacagrimonas perspica gen. nov., sp. nov., a novel member of Gammaproteobacteria isolated from soil of a ginseng field. J Microbiol. 48, 262–266 (2010). https://doi.org/10.1007/s12275-010-0067-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-010-0067-0