Abstract
Kriging metamodeling (also called Gaussian Process regression) is a popular approach to predict the output of a function based on few observations. The Kriging method involves length-scale hyperparameters whose optimization is essential to obtain an accurate model and is typically performed using maximum likelihood estimation (MLE). However, for high-dimensional problems, the hyperparameter optimization is problematic and often fails to provide correct values. This is especially true for Kriging-based design optimization where the dimension is often quite high. In this article, we propose a method for building high-dimensional surrogate models which avoids the hyperparameter optimization by combining Kriging sub-models with randomly chosen length-scales. Contrarily to other approaches, it does not rely on dimension reduction techniques and it provides a closed-form expression for the model. We present a recipe to determine a suitable range for the sub-models length-scales. We also compare different approaches to compute the weights in the combination. We show for a high-dimensional test problem and a real-world application that our combination is more accurate than the classical Kriging approach using MLE.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Kriging models (Cressie 1993; Stein 1999) are non-parametric statistical models which have been used in many different fields to infer the output of a function y based on a few observations. Applications include geostatistics (Krige 1951; Matheron 1963), the approximation of numerical experiments (Sacks et al 1989; Santner et al 2003), machine learning where the method is known as Gaussian process (GP) regression (Rasmussen and Williams 2006).
One of the main drawbacks of the Kriging method is that it scales poorly for large-scale problems: it suffers from the curse of dimensionality (Bellman 1966) when the dimension of the input is large. This issue is especially prevalent in engineering design optimization (Sobester et al 2008) as industrial designs are commonly parametrized by more than 50 shape parameters (Shan and Wang 2010; Gaudrie et al 2020). In this context, Kriging surrogate models are used to approximate the response of a computationally expensive numerical experiment based on a limited number of observations. The sample plan is usually built using a sequential strategy where an initial design of experiments is completed with new samples obtained by maximizing an acquisition criterion on the surrogate at each iteration (Jones et al 1998). It is therefore important that the surrogate is accurate, even with few observations as in the first iterations, since it will directly impact the number of additional samples needed for the optimization to converge (and thus the convergence speed).
The main challenge for building high-dimensional Kriging models resides in the hyperparameter optimization. Most Kriging models consider one length-scale hyperparameter per dimension which all need to be optimized simultaneously and this multidimensional optimization problem can be difficult to solve. Typically, the optimization is either performed by maximizing the likelihood of the model (Jones 2001) or by minimizing the leave-one-out cross-validation (LOOCV) error (Bachoc 2013). However, both methods involve the inversion of the covariance matrix with cost in \(O(n^3)\) (where \(n\) is the number of sample points). In design optimization, the number of samples is usually limited due to the cost of obtaining each of them, and thus the inversion is manageable. Yet, the optimization requires many of these inversions, especially in high-dimension as the size of the search space grows exponentially with the number of hyperparameters. As such, due to the large number of iterations needed to converge, the hyperparameter optimization can be prohibitively expensive even for a limited number of samples. One way to reduce the cost of the optimization is to use an approximation of the covariance matrix inverse such as those developed for Kriging models with large number of observations where the cost of an inversion is prohibitive (see Liu et al (2020) for a review). For example, in Quinonero-Candela and Rasmussen (2005), Titsias (2009) and Hensman et al (2013), the authors use a low-rank approximation of the covariance matrix to reduce the computational cost of the inversion. However, most of those methods are only designed for a large number of samples.
Besides the cost of the hyperparameter optimization, in high-dimension the input space training data is often sparse since the design space grows exponentially with the dimension. This, along with the large number of hyperparameters, can cause the usual criterion for the optimization to over-fit the training data leading to a poor estimation of the hyperparameters even when the optimization has converged (Ginsbourger et al 2009; Mohammed and Cawley 2017). Reducing the dimension of the problem is a way to solve these issues (see Binois and Wycoff (2021) for a review), but, because y is computationally expensive, classical sensibility analysis (Saltelli et al 2008) cannot be performed beforehand for variable selection. Some methods reduce the dimension by embedding the design space into a lower-dimension space (Constantine 2015; Bouhlel et al 2016). Additive Kriging (Durrande et al 2012) is another approach where y is decomposed into a sum of one dimensional components, enabling a sequential optimization of the length-scale hyperparameters.
In this paper, we propose a new method to tackle the challenging hyperparameter optimization for high-dimensional problems. Our approach avoids this optimization by combining Kriging sub-models with random length-scales. It replaces the challenging inner optimization of the length-scales by an optimization of the combination weights which is much simpler and whose solution can be obtained in closed-form. It also avoids reducing the dimension of the design space and preserves the correlation between all the input variables. This article starts by briefly recalling the main concepts in Kriging and introduces the employed notations in Sect. 2. Our combined Kriging method is detailed in Sect. 3. Finally, results of our method on numerical test problems are presented and discussed in Sect. 4.
2 Kriging model
2.1 Kriging predictions
This section briefly recalls the Kriging method and introduces the notations used throughout this paper. We denote by \(y: \varvec{x}\in {\mathcal {X}} \subset {\mathbb {R}}^d\rightarrow y(\varvec{x}) \in {\mathbb {R}}\) the \(d\)-dimensional black-box function that we want to approximate. We suppose y is known on an ensemble of \(n\) sample points \(\varvec{X}= \left( \varvec{x}_1,\dots ,\varvec{x}_n\right) ^T\) and we denote \(\varvec{Y}= \left( y(\varvec{x}_1),\dots ,y(\varvec{x}_n) \right) ^T\) the observed values at these locations. The Kriging method approximates y as the realization of a Gaussian process on \({\mathcal {X}}\):
Without loss of generality, we can assume that the GP is centered (\(\mu =0\)). \(k_{\varvec{\theta }}: {\mathcal {X}} \times {\mathcal {X}} \rightarrow [-1,1]\) is the positive definite correlation function indexed by the hyperparameters \(\varvec{\theta }\in {\mathbb {R}}^d\), the correlation length-scales vector (also called range or scale parameters) with one length-scale value per dimension of the input space. Finally, \(\sigma ^2k_{\varvec{\theta }}\) is the covariance function (also called kernel) with \(\sigma ^2 \in {\mathbb {R}}^+\) the variance of the GP. A stationary GP with a Matérn-class covariance function is often recommended (Stein 1999; Rasmussen and Williams 2006). Throughout this paper, we use the radial Matérn 5/2 correlation defined as:
where \(\left\| \frac{\varvec{x}-\varvec{x}'}{\varvec{\theta }}\right\| \) is the scaled distance between two points \(\varvec{x}, \varvec{x}'\in {\mathcal {X}}\) using component-wise division: \(\left\| \frac{\varvec{x}-\varvec{x}'}{\varvec{\theta }}\right\| ^2 :=\sum _{i=1}^{d} \left( \frac{{x}^{(i)}-{x'}^{(i)}}{\theta ^{(i)}}\right) ^2.\) This is a typical choice for design optimization (Roustant et al 2012), and even when the covariance is misspecified, a proper estimation of the hyperparameters can still yield a model with good predictive capacities (Bachoc 2013). Other covariances could be used if a priori knowledge about the unknown function is available.
The Simple Kriging (SK) predictor is a linear combination of the observations which is obtained by conditioning the Gaussian process Y over \({\mathcal {D}}=(\varvec{X},\varvec{Y})\):
where \(k_{\varvec{\theta }}(\varvec{x},\varvec{X})\) is the vector of correlations between the prediction point \(\varvec{x}\) and the sample points \(\varvec{X}\), and \(k_{\varvec{\theta }}(\varvec{X},\varvec{X})\) is the correlation matrix of the model, i.e. the \(n\times n\) matrix of correlations between the components of \(\varvec{X}\). Note that this predictor does not depend on \(\sigma ^2\). The predictive variance of the model can also be obtained as:
In the following, we will simply denote the correlation matrix as \(\varvec{K}_{\varvec{\theta }} :=k_{\varvec{\theta }}(\varvec{X},\varvec{X})\).
2.2 Hyperparameter estimation
The covariance hyperparameters must be chosen appropriately to obtain an accurate model. Usually, they are set using the maximum likelihood estimation (MLE) (Jones 2001), which consists on maximizing the marginal likelihood of the model:
This is equivalent to minimizing \(-log({\mathcal {L}}(\sigma ,\varvec{\theta }))\). For a fixed \(\varvec{\theta }\), the MLE estimator for \(\sigma ^2\) is:
After substituting (5) into the log-likelihood, we obtain the length-scales \(\varvec{\theta }\) by solving:
An alternative to MLE is to minimize the leave-one-out cross-validation (LOOCV) error (Bachoc 2013) of the model:
where \({M_{\varvec{\theta }}}_{-k}\) is the simple Kriging model built by removing the \(k\)th sample point \(\varvec{x}_k\). For Kriging models, the LOOCV error can be computed easily (Ginsbourger and Schärer 2021) without having to build \(n\) models using the formula:
Finally, the LOOCV estimation of the length-scales is obtained as:
In practice, both optimization problems (6) and (9) can be difficult to solve numerically due to their multi-modality, to flat areas of the objectives, and to the fact that the objective evaluations can be expensive (cost in \(O(n^3)\) for both objectives and their gradients). This is particularly true for high-dimensional problems since \(\varvec{\theta }\) has dimension \(d\). Equations (6) and (9) are typically solved using gradient-based method (e.g BFGS) with multi-start, or using evolutionary algorithms (Roustant et al 2012). However, as we will show in Sect. 4, these methods can fail to produce suitable values of the hyperparameters in high-dimensional problems, when the data is relatively sparse.
In the next section, we propose an alternative method for building a Kriging-based surrogate model which avoids this challenging optimization of the length-scale hyperparameters.
3 Combined Kriging with fixed length-scales
3.1 Description of the method
Combining different surrogate models using weights has been explored by many authors in the past years. The proposed methods differ in the purpose of the combination, in the type of the surrogate models employed, and in the way the weights are computed. For example, Bayesian model averaging (Gelman et al 1995; Hoeting et al 1999; Burnham et al 2011) combines different models using different parameters to perform a multimodel inference while accounting for the uncertainty in the choice of the model. In Goel et al (2007), Acar and Rais-Rohani (2009) and Viana et al (2009), different metamodels build on the same data set are combined to obtain an ensemble of surrogates whose accuracy is better than the one of the best metamodel. To circumvent the difficulties of Kriging metamodels in the presence of large datasets, several methods combining local Kriging sub-models optimized on subset of points have also been proposed with different weighting schemes (Rasmussen and Ghahramani 2001; Cao and Fleet 2014; Deisenroth and Ng 2015; Rullière et al 2018). In the context of Bayesian optimization, Ginsbourger et al (2008) present a method to combine Kriging sub-models with various covariance functions, or with different hyperparameter optimization criteria. The combination of Kriging sub-models for selecting the covariance function is further explored in Palar and Shimoyama (2018), and Pronzato and Rendas (2017) combine several local Kriging sub-models with different covariance functions in a fully Bayesian manner to build a non-stationary model.
Contrarily to the combinations of Kriging sub-models presented above, in the method we propose, the length-scale hyperparameters are not optimized but randomly chosen. It avoids the expensive and difficult optimization of these hyperparameters for high dimensional problems by emphasizing the appropriate random length-scales through their weights in the combination, which are obtained in closed-form.
The combined model writes as:
where \(M_{tot}\) is the combined model, \(p\) is the number of sub-models, and \(w_i\), \(i=1,\dots ,p,\) are the weights of each sub-model. The sub-models \(M_i\) are simple Kriging models with random length-scales, hence:
where \(\varvec{\theta }_i\) is the random length-scale vector and \({\mathcal {D}}_i=(\varvec{X}_i,\varvec{Y}_i)\) the training data set of the \(i\)th sub-model. We also have access to the variance of each sub-model:
The proposed method enables the construction of a Kriging model for high dimensional problems without reducing the dimension. It both preserves the correlation between all input variables and avoids a loss of information due to a truncated design space. Moreover, this method is very flexible since each sub-model can for instance be constructed on different subsets of points, can take into account different design variables, or can have different covariance functions. Sub-models with very different behaviors sweeping through a wide range of length-scales can therefore be combined. The interest in this paper is for high-dimensional problems with typical dimension \(d> 20\). The number of sub-models is limited to \(p< d\ll n\), and we will show empirically in Sect. 4 that for our test problem with dimension \(d=50\), a small number of sub-models is sufficient as adding more no longer improves the combinations. Finally, we consider a moderate number of, at most, few thousands samples so that, albeit slightly expensive, the inverse of the covariance matrix can be computed for the \(p\) sub-models. Thus, the complexity of the combination is \(O(pn^3)\) which is generally less than the cost of an ordinary Kriging model in \(O(\alpha _{iter}n^3)\) where \(\alpha _{iter}\) is the number of matrix inversions (i.e. the number of iterations, typically of the order of 100) in the optimization of the \(d{}\) hyperparameters. To fully define the combination, the first step is to define the sub-models which is detailed in Sect. 3.2. Then, the choice of the weights for the combination is discussed in Sect. 3.3.
3.2 Choice of the sub-models
In this paper, all Kriging sub-models are constructed with all sample points and all design variables: \({\mathcal {D}}_i={\mathcal {D}}=(\varvec{X},\varvec{Y})\), \(i=1,\dots ,p\) so that the length-scales are the unique source of difference between the \(M_i\)’s. An appropriate choice of the length-scales is essential to obtain a combined Kriging model with a good accuracy. In particular, variety among the sub-models is crucial so that the combined model can select the most well-suited behaviors through the weights in the combination. Since no prior knowledge is available for the length-scales, we choose them randomly in a bounded interval:
where \(\theta _{min}^{(\ell )}\) and \(\theta _{max}^{(\ell )}\) are lower and upper bounds for the \(\ell \)th component \(\theta ^{(\ell )}\) of \(\varvec{\theta }\). To the best of our knowledge, only few works in the literature deal with length-scale bounds. Mohammadi et al (2016, 2018) deal with these bounds in an optimization context, but it is common practice to assume pre-specified bounds. In the DiceKriging R package (Roustant et al 2012), by default, \(\theta ^{(\ell )}_{min}=10^{-10}\) and \(\theta ^{(\ell )}_{max}\) is twice the observed range in the \(\ell \)th dimension. Obrezanova et al (2007) fix the length-scales based on the standard deviation of the data. Issues related to flat likelihood landscapes may occur for too small length-scales and a suitable lower bound for maximum likelihood estimation is proposed in Richet (2018).
Intuitively, the choice of the length-scales range depends both on the design and on the covariance function family. We study hereafter both factors separately, although it would be possible to study them jointly.
3.2.1 Design impact
If the length-scales are large compared to most observed pairwise distances, then the correlations will tend to one. If they are smaller than most distances, trajectories with higher frequencies than observed in the given samples are implicitly considered. Therefore, length-scales should be of the order of most of the observed pairwise distances.
Let us investigate the distribution of observed distances between design points. Assume that design points are distributed as a random vector \({\textbf{X}}=(X^{(1)}, \ldots , X^{(d)})\), with respective standard deviations \(\sigma ^{(1)}, \ldots , \sigma ^{(d)}\). As we do not consider here the cross influence of joint length-scales, we investigate the impact of length-scales variations along the curve:
Now denote by \(\left\| \frac{\textbf{R}}{\varvec{\theta }}\right\| \) the scaled random distance between two distinct independent points \({\textbf{X}}\) and \(\mathbf {X'}\) of the design, using component-wise division. When \(\varvec{\theta }\in {\mathcal {C}}\), this distance can be expressed as a function of \(\theta ^{(\ell )}\):
Assuming the finiteness of the first four moments, when all components of \({\textbf{X}}\) and \(\mathbf {X'}\) are mutually independent with common kurtosis \(\kappa \),
Along the direction \(\ell \), using a simplified model, for \(d\) large enough, typical values of the unscaled distance \(\Vert \theta ^{(\ell )}\frac{\textbf{R}}{\varvec{\theta }}\Vert \) are given by the root of a Gaussian \(95\%\) confidence interval (for a Gaussian design, one should use the confidence interval of a \(\chi \) distribution):
This interval corresponds to typically observed unscaled distances in the dimension \(\ell \). Notice that it grows as \(\sqrt{d}\) and that it is built around an average distance \(\sigma ^{(\ell )}\sqrt{2d}\) along this axis. For uniform random variables, the kurtosis is \(\kappa = 9/5\), for Gaussian ones, it is \(\kappa = 3\).
3.2.2 Covariance family impact
The impact of a change in the length-scales depends on the covariance function: for instance, the covariance varies slowly at short distances for Gaussian kernels, whereas it varies rapidly for exponential ones. This has to be taken into account when choosing length-scales bounds.
Let \(k(\left\| \frac{\textbf{r}}{\varvec{\theta }}\right\| )\) be the covariance between two design points \({\textbf{x}}\) and \(\mathbf {x'}\), where \(\left\| \frac{\textbf{r}}{\varvec{\theta }}\right\| \) is the scaled distance between the points and k(.) is the covariance function of an isotropic stationary Gaussian Process. When \(\varvec{\theta }\in {\mathcal {C}}\), \(\left\| \frac{\textbf{r}}{\varvec{\theta }}\right\| \) can be expressed using only \(\theta ^{(\ell )}\). The influence of \(\theta ^{(\ell )}\) on the covariance can be measured by the following normalized derivative:
The derivative with respect to the length-scale can be obtained easily by direct calculation for the usual covariance functions. Along the axis \(\ell \), at a scaled distance \(\left\| \frac{\textbf{r}}{\varvec{\theta }}\right\| =\frac{r}{\theta ^{(\ell )}}\), a length-scale \(\theta ^{(\ell )}\) is considered influential enough if it belongs to:
where \(\delta \in (0,1)\) is a user-defined threshold that we set to \(\delta =1/10\) in the following.
For \(r \in \left[ r_{min}^{(\ell )}, r_{max}^{(\ell )}\right] :=\sigma ^{(\ell )}[r_{min}, r_{max}]\), length-scales bounds are chosen as:
Note that multiplying distances by a scale factor \(\alpha >0\) changes the set of admissible length-scales by the same factor, \(\Theta ^{(\ell )}_{adm}(\alpha r)= \alpha \Theta ^{(\ell )}_{adm}(r)\). Therefore, one only has to solve for \(r=1\) in \(\theta \):
We denote respectively as \(\theta ^{-}(k)\) and \(\theta ^{+}(k)\) the smallest and largest roots of (15), which depend only on the chosen covariance function k(.). The influence index and its roots are illustrated in Fig. 1 for the exponential and Gaussian kernels. The roots do not depend on the component number \(\ell \), and we finally get:
Notice that \(r_{min}\), \(r_{max}\) depend only on the design kurtosis \(\kappa \) and on the dimension \(d\). Examples of obtained bounds for uniformly sampled designs are given in Table 1. When d tends to infinity, the length-scale range becomes equivalent to \(\sigma ^{(\ell )}\sqrt{2d} [\theta ^{-}(k), \theta ^{+}(k)]\), and only depends on the design distribution through \(\sigma ^{(\ell )}\). The surprising result of distance concentration in high dimension (\(r_{min}\) and \(r_{max}\) both equivalent to \(\sqrt{2d}\) as d increases, see last lines in Table 1) is also discussed in the literature, see e.g. Aggarwal et al (2001).
3.2.3 Sampled bounds
The length-scales of the sub-models are sampled randomly in their corresponding interval. Different sampling strategies can be considered (i.e. space-filling designs, sample plans biased towards the center of the length-scale space). In this paper we use a uniform sampling scheme: \(\theta ^{(\ell )}\thicksim {\mathcal {U}}(\theta _{min}^{(\ell )},\theta _{max}^{(\ell )}), \quad \ell =1,\dots ,d\,.\)
3.3 Choice of the weighting method
As detailed in Sect. 3.1, the literature on model combination is vast and many weighting methods have been developed. Five of those are investigated in this paper in order to compute the weights of the sub-models in (10). This Section briefly describes the resulting five weighting schemes, more details on each method can be found in Appendix A. Their performances are then compared on numerical experiments in Sect. 4.
3.3.1 PoE approach
The first approach to obtain the weights is based on Product of Experts (PoE) (Hinton 2002). The PoE weights are given by:
where \(\hat{s}^{2}_i(\varvec{x})\) is the variance of the \(i\)th sub-model given in Eq. (12). Note that these weights depend only on the position of the sample points and not on the observed values. When the Kriging sub-models are all built with the same sample points, this method will emphasize the sub-models with large length-scales because these are the ones with smallest predicted variance. Thus, we expect this method to favor large length-scales and to fail in selecting the correct sub-models.
3.3.2 gPoE approach
The second approach is based on generalized Product of Experts (gPoE) (Cao and Fleet 2014; Deisenroth and Ng 2015). The gPoE weights are:
In this paper, we use the gPoE approach to adjust the PoE weights in order to account for the observed values at the sample points. To this aim, the internal weights \(\varvec{\beta }^*\) are optimized numerically to minimize the LOOCV error of the combined model given by Eq. (7). However, a closed-form expression of the weights is no longer available because of this inner optimization.
3.3.3 LOOCV and LOOCV diag approaches
The third approach is to directly minimize the LOOCV error of the combination in Eq. (7) (Viana et al 2009) giving the LOOCV weights:
where the components of the matrix \(\varvec{C} \in {\mathbb {R}}^{p\times p}\) are \(c_{ij} = \frac{1}{n} \varvec{e}_i^T \varvec{e}_j, \quad i=1,\dots ,p, \quad j=1,\dots ,p,\) with \(\varvec{e}_i\) the LOOCV vector for the \(i\)th sub-model: \(\varvec{e}_i=(\varvec{e}_i^{(1)},\dots ,\varvec{e}_i^{(n)})\). Using (8), these elements can be expressed easily as: \(\varvec{e}_i^{(k)} = [\varvec{K}_{\varvec{\theta }_i} \varvec{Y}]_k/ [\varvec{K}_{\varvec{\theta }_i}]_{k,k}\). Contrarily to the two previous approaches, these weights are constant and do not depend on \(\varvec{x}\). We also note that this method might lead to negative or greater than one weights. As we will discuss in Sect. 4, negative weights can raise some issues for the combination. Thus, following the suggestion of Viana et al , we propose the fourth weight definition enforcing \(w_i\in [0,1]\) by keeping only the diagonal elements of the matrix \(\varvec{C}\) in Eq. (19):
3.3.4 MoE approach
The fifth approach based on Mixture of Experts (MoE) (Yuksel et al 2012) gives the MoE weights:
Here, \({\mathcal {L}}(\varvec{\theta }_i)\) is the marginal likelihood of the \(i\)th sub-model. One drawback of MoE is that the likelihood of different sub-models can vary by several orders of magnitude. Thus this method may emphasize one single sub-model with the best likelihood instead of combining different sub-models.
4 Numerical results
4.1 Experiment setup
We compare the performances of the different combined models described in Sect. 3 with the simple Kriging method on simulated data and on a real-world application. To build the sub-models and the simple Kriging model, \(n_{train}=500\) random training points \(\varvec{x}_1,\dots ,\varvec{x}_{n_{train}} \in [0,1]^d\) are uniformly sampled. We use the Matérn 5/2 covariance function defined in Eq. (1). For the random sub-models, we follow the methodology detailed in Sect. 3.2, where we take a threshold \(\delta =1/10\), a kurtosis corresponding to a uniform distribution \(\kappa =9/5\), and the empirical standard deviation of the design along each direction. The weights of the combinations are computed according to Eqs. (17, 18, 19, 20) and (21). The performances of the five combined models are compared with the accuracy of a simple Kriging model with hyperparameters optimized by MLE (the optimization is performed using the package DiceKriging in the R language Roustant et al (2012) with 300 maximum iterations). The experiments are repeated for 10 different random seeds, with different random length-scales for the sub-models as well.
4.2 Simulated data
For the simulated data, the functions to surrogate are random samples of a high dimensional (\(d=50\)) centered Gaussian process Y using a Matérn 5/2 covariance function with known isotropic length-scales \(\theta _{true}=2\). Since in this case the true length-scales are known, we also compare the combinations and the simple Kriging to a Kriging model with the true hyperparameters \(\varvec{\theta }_{true}\) as a reference.
In a first experiment, we consider a combination of \(p=\)10 sub-models where, only this time, the length-scales are fixed to isotropic values ranging from 1 to 10. The purpose of this experiment is to observe how the different weights behave in a case where we know how close to the true function each sub-model is. In a second experiment, we build a set of \(p=40\) Kriging sub-models \({\mathcal {M}} = \{M_1,\dots ,M_{p}\}\), this time with random length-scales. The combined models are then constructed by aggregating a gradually increasing number of these sub-models (from 5 to all 40), using the 5 different methods for computing the weights. Additionally, to investigate the robustness of the combinations to “wrong” sub-models, we also add 5 sub-models with fixed isotropic large length-scales \(\theta =10\). The quality of each prediction \(M_{tot}\) is assessed by the mean-square error (MSE) computed on a test set of \(n_{test}\) = 5000 random test point \(\varvec{x}_1^{(t)},\dots ,\varvec{x}_{n_{test}}^{(t)} \in [0,1]^d\):
In order to further interpret the results, the LOOCV error of each model is also computed:
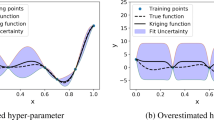
The results of the first experiment are presented in the Figs. 2 and 3. The first result to note is the weak performance of the SK model with estimated hyperparameters (KrgMLE). Since the model is well specified (the underlying function we try to approximate is a GP sample with the same covariance structure), we would expect the MLE method to recover the true hyperparameters (see Bachoc (2013)). Moreover, as the high-dimensional optimization can be difficult, we use multi-start along with a large number of iterations (300 iterations) to ensure convergence. However, the maximum likelihood optimization still results in a wrong estimation of the length-scales \(\varvec{\theta }_{MLE}\), far from the truth \(\varvec{\theta }_{true}\). This is because with the small number of observations available, the maximum likelihood criterion over-fits the training data as highlighted in Fig. 2b by the LOOCV error of the model with estimated hyperparameters which is much smaller than that of the reference model. Because of the poor estimation of the length-scales, the MSE error of this model is also much worse than the MSE of the model with the true hyperparameters as shown in Fig. 2a. The PoE method clearly performs the worst among the combined models because it gives almost all the weight to the subs-models with large length-scales. The gPoE method avoids this issue thanks its internal weights as shown in Fig. 3a, and this method performs similarly to the three others. The different weighting strategies of each method are shown in Fig. 3. For the LOOCV method, the weights are fluctuating and hard to interpret, since their values are not in the [0,1] interval. The LOOCV diag method gives weights which are distributed quite uniformly among all sub-models though sub-models with \(\theta _{i}\approx \theta _{true}\) are highlighted, while the gPoE and MoE methods focus on the two more accurate sub-models.
MSE and LOOCV error (the lower the better) for the combinations of isotropic sub-models for the approximation of an isotropic Gaussian process sample. The 5 five first boxes (blue) correspond to the 5 weighting methods, the second to last box (red) to the simple Kriging model with hyperparameters estimated by MLE, the last box (green) to a simple Kriging model with \(\theta =\theta _{true}\) (colour figure online)
Weights of the isotropic sub-models for the first experiment. The x-axis values represent the isotropic length-scale of each sub-model, \(\theta \). For the gPoE method, the weights are the \(\varvec{\beta }\) internal weights in Eq. 18, for the 3 other methods the weights are the constant weights used for the combination
The results of the second experiment are given in Fig. 4. First, we can note that the SK model with estimated hyperparameters still overfits the data which results in a high MSE. Contrarily to the first experiment, the PoE method performs well as seen in Fig. 4a and f. This is because, in this experiment, the sub-models are no longer isotropic and are all composed of both small and large length-scales. As such, the PoE which discriminates against small length-scales leads to a good MSE since small length-scales often result in a Kriging model with large variations, and thus potentially large MSE values, while large length-scales give flatter models with moderate MSEs. However, for the same reason, PoE is not robust to the addition of “wrong” sub-models with large length-scales. Figure 4c shows that the accuracy of the combined model using the LOOCV method steadily decreases when too many sub-models are aggregated (more than 10). This is in contrast to the Fig. 4h where the LOOCV error of this method keeps decreasing when more sub-models are added, which is to be expected as the weights in this method are designed to minimize this very error. This, again, can be explained by the fact that this combination starts to overfit the data with too many sub-models. However, this issue does not occur for the LOOCV diag method in Fig. 4d and i where the MSE always decreases with \(p\) and converges to a threshold at about 15 sub-models in the combination. Figure 4e and j show that in this experiment the MoE method produces poor results. This is because, in this 50-dimensional example, the likelihoods of the sub-models are very small and differ by several order of magnitudes. This results in an MoE weight of almost one for the sub-model with the best likelihood, hence the method is almost equivalent to choosing only the best sub-model, thus using only one single pre-specified covariance. We also note that, for the methods which give the best accuracy (PoE, gPoE and LOOCV diag), the combined model is generally better than the best sub-model with as few as five sub-models in the combination. This shows well that the combination strategy is more effective compared to choosing the best sub-model among random samples.
Results of the second experiment for an initial GP sample with isotropic length-scales \(\theta = 2\). The top row shows the MSE results for the 5 combination methods, and the bottom row the LOOCV error results (the lower the better). In each boxplot, the first box gives the accuracy of the best sub-model Sub (yellow), the next 8 boxes (blue) give the accuracy of the combined model with an increasing number of sub-models (from 5 to 40), the third to last +Bad (purple) gives the accuracy when the combination is perturbed by the addition of 5 bad sub-models with large length-scales (\(\theta =10\)), the second to last box MLE (red) gives the performance of a simple Kriging model with hyperparameters estimated by MLE, the last box True (green) gives the precision of a simple Kriging model using the same length-scale as the initial GP sample (colour figure online)
Table 2 gives a summary of the different properties observed for the 5 weighting methods in these numerical experiments. The only method without a closed-form expression is gPoE because of the inner weights optimization (Eq. (A3)). As seen in the third experiment (Fig. 4b) PoE is not robust to “wrong” sub-models.Only PoE is not robust to “wrong” sub-models as seen in Fig. 4b. Figure 4c shows that LOOCV overfits when there are too many sub-models. Finally, both experiments (Figs. 2a and 4e) show that MoE does not suitably balance the weight between all sub-models.
4.3 Real-world application
To validate the method on a more realistic problem, we study a real-world application corresponding to the design of an electrical machine. The shape of the machine is parameterized with \(d=37\) design variables. These variables represent the position and size of air holes and magnets, as well as the radius of the machine. The layout of the machine is illustrated in Fig. 5. We are interested in the performances measured by two objectives to minimize: the consumption and cost of the machine; subject to ten constraints which characterize the dynamic of the car (maximum speed, acceleration,...), the reducer dimensioning, and the dynamic of the machine (oscillations,...). These two objectives and ten constraints are obtained via numerical simulation. Thus, we build 12 surrogates (one for each objective and constraint).
For a fixed number of \(p=20\) random sub-models, we compare the accuracy of the combinations with that of simple Kriging. To measure the accuracy, as the scales and units of each objectives and constraints cannot be compared, instead of the MSE and LOOCV we use the \(Q^2\) coefficient computed on a test set of \(n_{test}\) = 4500 random test point \(\varvec{x}_1^{(t)},\dots ,\varvec{x}_{n_{test}}^{(t)} \in [0,1]^d\):
The results for the 2 objective and 10 constraints of the electrical machine is summarized in Fig. 6. Note that here the boxplots represent the result over the 12 surrogates (averaged over 10 different random seeds).
Results of the real-world application. The boxplots represent the \(Q^2\) (the higher the better) over the 12 objectives and constraints. The leftmost box gives the accuracy of the best sub-model Best sub (yellow), the next 5 boxes (blue) give the accuracy of the 5 methods for the combination, the last box Krg MLE (red) gives the performance of a Kriging model with hyperparameters estimated by MLE (colour figure online)
The results obtained confirm those for the simulated data. The accuracy using a combination of random sub-models is better than Kriging with hyperparameters optimized via maximum likelihood. Among the 5 methods for the weights, gPoE and LOOCV diag are to be preferred as the conclusion from the simulated data still applies here.
5 Conclusion
In this paper, we have proposed a new method to construct a surrogate model as a combination of Kriging sub-models which avoids the cumbersome optimization of the length-scales hyperparameters. The length-scales of the sub-models are pre-specified, for instance randomly, and the combined model emphasizes the important ones. We also provided a recipe for the choice of the length-scale bounds, as well as a comparison of different methods for weighting the sub-models.
Compared to other approaches, our method provides a novel way to build a Kriging-based surrogate model for high dimensional problems without employing dimension reduction techniques. The accuracy of our surrogate model is improved in comparison to simple Kriging models where the length-scales are optimized by MLE and which performs poorly in high-dimension, especially when the number of observations is limited. Moreover, the computational cost of the model is reduced as only \(p\) matrix inversions are needed to build the \(p\) sub-models, which, for a reasonable number of sub-models, is less expensive than the standard length-scale optimization which requires iterative covariance matrix inversions.
The numerical results for the 50 dimensional test problem and for the real-world application show that our method performs significantly better than simple Kriging with hyperparameters optimized by MLE for this type of problems. In particular, both the gPoE and LOOCV diag stand out as the best approaches to combine the sub-models and give an accuracy close to that of the reference model with as few as 15 sub-models.
Several aspects still need to be explored in further research. First, we can think of combining different kinds of sub-models, for example for problems where the design variables can naturally be separated into different groups, or by varying the covariance function, to further diversify the sub-models instead of considering identical sub-models sharing the same points and design variables. We could also consider sub-models built with subsets of samples in order to handle cases where in addition to the high-dimension, the number of observations is also large enough that the cost of the covariance matrix inversion becomes prohibitive. Second, we could try to induce sparsity in the weights in order to improve the interpretability of the combination. Finally, the variance of the aggregated model, which is mandatory to apply our method to the EGO framework for Bayesian optimization, is currently available only for the MoE weighting. Extending the current method to obtain variance estimates for other weighting approaches and applying it to Bayesian optimization constitutes an interesting research direction.
References
Acar E, Rais-Rohani M (2009) Ensemble of metamodels with optimized weight factors. Struct Multidiscip Optim 37(3):279–294
Aggarwal CC, Hinneburg A, Keim DA (2001) On the surprising behavior of distance metrics in high dimensional space. In: International conference on database theory, Springer, pp 420–434
Bachoc F (2013) Cross validation and maximum likelihood estimations of hyper-parameters of gaussian processes with model misspecification. Comput Stat Data Anal 66:55–69
Bellman R (1966) Dynamic programming. Science 153(3731):34–37
Binois M, Wycoff N (2021) A survey on high-dimensional gaussian process modeling with application to bayesian optimization. arXiv preprint arXiv:2111.05040
Bouhlel MA, Bartoli N, Otsmane A et al (2016) Improving kriging surrogates of high-dimensional design models by partial least squares dimension reduction. Struct Multidiscip Optim 53(5):935–952
Burnham KP, Anderson DR, Huyvaert KP (2011) Aic model selection and multimodel inference in behavioral ecology: some background, observations, and comparisons. Behav Ecol Sociobiol 65(1):23–35
Cao Y, Fleet DJ (2014) Generalized product of experts for automatic and principled fusion of gaussian process predictions. arXiv preprint arXiv:1410.7827
Constantine PG (2015) Active subspaces: Emerging ideas for dimension reduction in parameter studies. SIAM
Cressie N (1993) Statistics for spatial data. Wiley, Amsterdam
Deisenroth M, Ng JW (2015) Distributed gaussian processes. In: International Conference on Machine Learning, PMLR, pp 1481–1490
Durrande N, Ginsbourger D, Roustant O (2012) Additive Covariance kernels for high-dimensional Gaussian Process modeling. Annales de la Faculté des sciences de Toulouse : Mathématiques Ser. 6, 21(3):481–499
Gaudrie D, Le Riche R, Picheny V et al (2020) Modeling and optimization with gaussian processes in reduced eigenbases. Struct Multidiscip Optim 61(6):2343–2361
Gelman A, Carlin JB, Stern HS, et al (1995) Bayesian data analysis. Chapman and Hall/CRC
Ginsbourger D, Schärer C (2021) Fast calculation of gaussian process multiple-fold cross-validation residuals and their covariances. arXiv preprint arXiv:2101.03108
Ginsbourger D, Helbert C, Carraro L (2008) Discrete mixtures of kernels for kriging-based optimization. Qual Reliab Eng Int 24(6):681–691
Ginsbourger D, Dupuy D, Badea A et al (2009) A note on the choice and the estimation of kriging models for the analysis of deterministic computer experiments. Appl Stoch Models Bus Ind 25(2):115–131
Goel T, Haftka RT, Shyy W et al (2007) Ensemble of surrogates. Struct Multidiscip Optim 33(3):199–216
Hensman J, Fusi N, Lawrence ND (2013) Gaussian processes for big data. arXiv preprint arXiv:1309.6835
Hinton GE (2002) Training products of experts by minimizing contrastive divergence. Neural comput 14(8):1771–1800
Hoeting JA, Madigan D, Raftery AE et al (1999) Bayesian model averaging: a tutorial (with comments by m. clyde, david draper and ei george, and a rejoinder by the authors. Stat Sci 14(4):382–417
Jones DR (2001) A taxonomy of global optimization methods based on response surfaces. J Global Optim 21(4):345–383
Jones DR, Schonlau M, Welch WJ (1998) Efficient global optimization of expensive black-box functions. J Global Optim 13(4):455–492
Krige DG (1951) A statistical approach to some basic mine valuation problems on the witwatersrand. J Southern Afr Instit Min Metall 52(6):119–139
Liu H, Ong YS, Shen X et al (2020) When gaussian process meets big data: a review of scalable gps. IEEE Trans Neural Netw Learn Syst 31(11):4405–4423
Matheron G (1963) Principles of geostatistics. Econ Geol 58(8):1246–1266
Mohammadi H, Riche RL, Touboul E (2016) Small ensembles of kriging models for optimization. arXiv preprint arXiv:1603.02638
Mohammadi H, Le Riche R, Bay X et al (2018) An analysis of covariance parameters in gaussian process-based optimization. Croat Operat Res Rev 9:1–10
Mohammed RO, Cawley GC (2017) Over-fitting in model selection with gaussian process regression. In: International conference on machine learning and data mining in pattern recognition, Springer, pp 192–205
Obrezanova O, Csányi G, Gola JM et al (2007) Gaussian processes: a method for automatic qsar modeling of adme properties. J Chem Inf Model 47(5):1847–1857
Palar PS, Shimoyama K (2018) On efficient global optimization via universal kriging surrogate models. Struct Multidiscip Optim 57(6):2377–2397
Pronzato L, Rendas MJ (2017) Bayesian local kriging. Technometrics 59(3):293–304
Quinonero-Candela J, Rasmussen CE (2005) A unifying view of sparse approximate gaussian process regression. J Mach Learn Res 6:1939–1959
Rasmussen C, Ghahramani Z (2001) Infinite mixtures of gaussian process experts. Adv Neural Inf Process Syst 14
Rasmussen CE, Williams CK (2006) Gaussian processes for machine learning. MIT press Cambridge, MA
Richet Y (2018) Cookbook: lower bounds for kriging maximum likelihood estimation (mle). https://dicekrigingclub.github.io/www/r/jekyll/2018/05/21/KrigingMLELowerBound.html, accessed: 2022-09-21
Roustant O, Ginsbourger D, Deville Y (2012) Dicekriging, diceoptim: two r packages for the analysis of computer experiments by kriging-based metamodeling and optimization. J Stat Softw 51:1–55
Rullière D, Durrande N, Bachoc F et al (2018) Nested kriging predictions for datasets with a large number of observations. Stat Comput 28(4):849–867
Sacks J, Welch WJ, Mitchell TJ et al (1989) Design and analysis of computer experiments. Stat Sci 4(4):409–423
Saltelli A, Ratto M, Andres T et al (2008) Global sensitivity analysis: the primer. Wiley, Amsterdam
Santner TJ, Williams BJ, Notz WI et al (2003) The design and analysis of computer experiments, vol 1. Springer, Berlin
Shan S, Wang GG (2010) Survey of modeling and optimization strategies to solve high-dimensional design problems with computationally-expensive black-box functions. Struct Multidiscip Optim 41(2):219–241
Sobester A, Forrester A, Keane A (2008) Engineering design via surrogate modelling: a practical guide. Wiley, Amsterdam
Stein ML (1999) Interpolation of spatial data: some theory for kriging. Springer, Berlin
Titsias M (2009) Variational learning of inducing variables in sparse gaussian processes. In: Artificial intelligence and statistics, PMLR, pp 567–574
Viana FA, Haftka RT, Steffen V (2009) Multiple surrogates: how cross-validation errors can help us to obtain the best predictor. Struct Multidiscip Optim 39(4):439–457
Yuksel SE, Wilson JN, Gader PD (2012) Twenty years of mixture of experts. IEEE Trans Neural Netw Learn Syst 23(8):1177–1193
Acknowledgements
This research was conducted with the support of the consortium in Applied Mathematics CIROQUO, gathering partners in technological and academia in the development of advanced methods for Computer Experiments. This research was partly funded by a CIFRE grant (convention #2021/1284) established between the ANRT and Stellantis for the doctoral work of Tanguy Appriou. The author also thank the editor and all the reviewers for their valuable comments on this paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix A more details on the weighting methods
Appendix A more details on the weighting methods
1.1 PoE approach
Product of Experts (PoE) (Hinton 2002) arises from the hypothesis that the posterior probability distribution of the combined model can factorize as a product of the posterior distribution of each sub-model (experts). Thus, it assumes independence between each sub-model, which is not the case for the proposed method where the sub-models are correlated. The PoE can also be seen as the best convex combination in the case of independent sub-models, which minimizes the variance of the combined model. The PoE weights are given by:
where \(\hat{s}^{2}_i(\varvec{x})\) is the variance of the \(i\)th sub-model given in Eq. (12).
1.2 gPoE approach
The generalized Product of Experts (gPoE) approach (Cao and Fleet 2014; Deisenroth and Ng 2015) was originally developed in the context of aggregating Kriging sub-models to alleviate some shortcomings of the PoE method, namely the fact that a single poor sub-model can cause a biased mean prediction and an overconfident variance. In gPoE, flexibility in the model is added by introducing internal weights \(\varvec{\beta }^*\) to the contribution of each sub-model. This results in the following expression for the gPoE weights:
Cao and Fleet (2014) suggest to compute \(\varvec{\beta }^*\) as the difference in entropy between the prior and posterior of each sub-model. In Deisenroth and Ng (2015), in order to recover the prior outside the data, the authors imposed the constraint \(\sum _{i=1}^{p}\beta _i^* = 1\) and proposed uniform weights \(\beta _i^*=1/p\).
In this paper, we use the gPoE approach to adjust the PoE weights in order to account for the observed values at the sample points. To this aim, the internal weights are optimized to minimize the LOOCV error of the combined model, with the constraint that their sum must be equal to one.
This is equivalent to optimizing the LOOCV error of a Kriging model whose precision matrix is the weighted sum of the precision matrices of each sub-models: \(\varvec{K}_{tot}^{-1}(\varvec{\beta })=\sum _{i=1}^{p} \beta _i\varvec{K}_{\varvec{\theta }_i}^{-1} \in {\mathbb {R}}^{n\times n}\). Thus, using the LOOCV formula for Kriging models given in Eq. (8):
This inner optimization can be performed numerically, and since only the inverses of the covariance matrices of each sub-model are required, it is inexpensive to perform as these inverses are already computed to build the sub-models. Although it accounts for the observed values, a closed-form expression of the weights is no longer available because of the inner optimization.
1.3 LOOCV and LOOCV diag approaches
To combine different surrogates, Viana et al (2009) proposed a method which minimizes the global mean-square error (MSE) of the combination given by \(E\left[ \left( M_{tot}(\varvec{X})-y(\varvec{X})\right) ^2 \right] \), where \(\varvec{X}\) is a random variable uniformly distributed over the design space. Since in practice we only dispose of a few observations, the global MSE is approximated using cross-validation. A discrete approximation of the global MSE using the LOOCV error can be obtained as:
where the components of the matrix \(\varvec{C} \in {\mathbb {R}}^{p\times p}\) are \(c_{ij} = \frac{1}{n} \varvec{e}_i^T \varvec{e}_j, \quad i=1,\dots ,p, \quad j=1,\dots ,p,\) with \(\varvec{e}_i\) the LOOCV vector for the \(i\)th sub-model: \(\varvec{e}_i=(\varvec{e}_i^{(1)},\dots ,\varvec{e}_i^{(n)})\). Using (8), these elements can be expressed easily as: \(\varvec{e}_i^{(k)} = [\varvec{K}_{\varvec{\theta }_i} \varvec{Y}]_k/ [\varvec{K}_{\varvec{\theta }_i}]_{k,k}\).
The weights are obtained by minimizing (A4) with respect to \(\varvec{w}\):
Using a Lagrange multiplier and setting the derivatives to zero, this gives the following weights for the LOOCV approach:
We note that this method might lead to negative or greater than one weights. Thus, following the suggestion of Viana et al , we propose a second weight definition enforcing \(w_i\in [0,1]\) by keeping only the diagonal elements of the matrix \(\varvec{C}\) in Eq. (19):
1.4 MoE approach
In Mixture of Experts (MoE) (Yuksel et al 2012) predictions of each expert are weighted by their posterior probability. The posterior predictive distribution of the mixture at \(\varvec{x}^*\) given the data \({\mathcal {D}}\) is:
where \(p(\varvec{\theta }=\varvec{\theta }_i\vert {\mathcal {D}})\) is the posterior probability of the sub-model \(i\) (with length-scales \(\varvec{\theta }_i\)). Using Bayes formula, we can express this posterior probability as:
The prior for the \(i\)th sub-model, \(p(\varvec{\theta }=\varvec{\theta }_i)\) is taken constant and \(p({\mathcal {D}}\vert \varvec{\theta }=\varvec{\theta }_i)\) is the marginal likelihood \({\mathcal {L}}(\varvec{\theta }_i)\) of the \(i\)th sub-model which is Gaussian and whose expression was given in Eq. (4).
From Eq. (A7), we can then obtain the predictive mean and variance of the combination:
where the weights \(w_{{MoE}_i}\) are obtained combining Eqs. (4) and (A8):
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Appriou, T., Rullière, D. & Gaudrie, D. Combination of optimization-free kriging models for high-dimensional problems. Comput Stat 39, 3049–3071 (2024). https://doi.org/10.1007/s00180-023-01424-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00180-023-01424-7