Abstract
The tight regulation of glucose and lipid metabolism is crucial for maintaining metabolic health. Dysregulation of these processes can lead to the development of metabolic diseases. Secreted factors, or hormones, play an essential role in the regulation of glucose and lipid metabolism, thus also playing an important role in the development of metabolic diseases such as type 2 diabetes and obesity. Given the important roles of secreted factors, there has been significant interest in identifying new secreted factors and new functions for existing secreted factors that control glucose and lipid metabolism. In this review, we evaluate novel secreted factors or novel functions of existing factors that regulate glucose and lipid metabolism discovered in the last decade, including secreted isoform of endoplasmic reticulum membrane complex subunit 10, vimentin, cartilage intermediate layer protein 2, isthmin-1, lipocalin-2, neuregulin-1 and neuregulin-4. We discuss their discovery, tissues of origin, mechanisms of action and sex differences, emphasising their potential to regulate metabolic processes central to diabetes. Additionally, we discuss the translational barriers, particularly the absence of identified receptors, that hamper their functional characterisation and further therapeutic development. Ultimately, the identification of new secreted factors may give insights into previously unidentified pathways of disease progression and mechanisms of glucose and lipid homeostasis.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Maintaining energy homeostasis is a multifaceted process involving many regulatory factors from many tissues. One class of regulatory factors crucial to energy metabolism is secreted factors or hormones. For example, high blood glucose levels promote insulin release from pancreatic beta cells to induce glucose uptake in peripheral tissues, thus lowering blood glucose levels [1,2,3]. Conversely, low blood glucose levels promote glucagon release by pancreatic alpha cells to induce glycogenolysis in the liver, thus raising blood glucose levels [4,5,6]. Postprandially, glucagon-like peptide 1 (GLP-1) is secreted by intestinal L cells to promote insulin secretion, slow down gastric emptying and suppress food intake [7, 8]. When fat mass increases, the adipokine leptin is released to inhibit food intake and increase energy expenditure [9]. Adipocyte secretion of adiponectin reduces blood glucose levels by suppressing hepatic gluconeogenesis and promoting fatty acid oxidation in skeletal muscle [10,11,12,13]. This handful of secreted factors only encompasses a small percentage of the secretome. Computational approaches predict that approximately 9% of human protein-coding genes encode at least one secreted product. This 9% represents 1891 genes and many more putative secreted factors, most of which have unknown functions. We have only scratched the surface of mechanistically understanding the secretome and its relationship with energy metabolism [14, 15]. Recently, we have seen a rapid increase in identification of secreted factors that regulate glucose and/or lipid metabolism, mainly thanks to advances in open-source expression data and proteomic techniques. Large-scale proteomic studies can reveal widespread changes to the secretome in individuals with type 2 diabetes or obesity compared with healthy control individuals [16, 17]. Often, secretome changes are only modestly correlated between studies, depending on the platform used and method of analysis [18]. Additionally, while proteomics provide significant insights into changes in protein levels that correlate with metabolic disease, the information is not sufficient for determining whether the protein is bioactive or for understanding the physiological role of the protein in regulating glucose and/or lipid metabolism. Therefore, a combination of computational approaches, proteomic studies and classical molecular biology studies is necessary for identifying novel secreted factors and gaining mechanistic understanding of how these secreted factors regulate energy metabolism and contribute to the development of metabolic disease.
In this review, we discuss the current understanding of how some of these newly described secreted factors affect energy metabolism in mice and cell lines, as well as correlations between these secreted factors and metabolic disease phenotypes in humans. Secreted factors can largely be divided into two groups: factors that drive the development of phenotypes associated with metabolic disease (e.g. glucose intolerance, insulin resistance and fat mass accumulation); and factors that protect against the development of these phenotypes. Here, we selected seven secreted factors (three driving factors and four protective factors) that are either newly identified or have been attributed with a novel function in the last decade. We discuss the tissues of origin, their roles in fundamental aspects of tissue metabolic homeostasis and their proposed mechanisms of action. We also elaborate on the limitations of our current understanding and the technological challenges in therapeutically harnessing these factors.
Secreted factors that promote the development of metabolic disease
Secreted isoform of endoplasmic reticulum membrane complex subunit 10
Endoplasmic reticulum membrane complex subunit 10 (EMC10) is a 27 kDa protein that is widely expressed and is an integral part of the endoplasmic reticulum (ER) membrane [14, 19]. It has been studied as a regulator of neurodevelopment, sperm motility and angiogenesis [20,21,22,23,24,25,26]. Differential splicing of EMC10 produces two isoforms: a membrane-bound EMC10 (mEMC10); and a secreted isoform [27]. In 2022, novel functions of the secreted isoform of EMC10 (scEMC10) in regulating glucose and lipid metabolism were identified (Fig. 1) [28]. scEMC10 was first implicated in regulating glucose metabolism when a 25 mmol/l dose of glucose to cultured pancreatic beta cells and intact islets led to a threefold increase in mRNA transcript levels for scEMC10 (previously known as INM02) [29]. A decade later, Wang et al used a neutralising antibody to demonstrate a causative role for scEMC10 in the dysregulation of glucose and lipid metabolism [28]. Specifically, they reported that neutralising scEMC10 in male mice on a high-fat diet (HFD) improved lipid metabolism as indicated by decreased fat mass accumulation and circulating NEFA levels, reduced hepatic steatosis and triglyceride levels, decreased circulating leptin levels and increased circulating adiponectin levels. Neutralising the function of scEMC10 in HFD-fed male mice also improved glucose metabolism as demonstrated by improved glucose and insulin tolerance and reduced fasting blood glucose levels. These improvements likely stem from increased thermogenesis and energy expenditure. In response to neutralisation of scEMC10 function, there was a significant increase in protein levels of uncoupled protein 1 (UCP1) and peroxisome proliferator-activated receptor γ coactivator-1α (PGC-1α) in brown adipose tissue (BAT), accompanied by increased energy expenditure [28]. Conversely, adeno-associated virus (AAV)-mediated overexpression of human scEMC10 (hscEMC10) in male mice worsened metabolic functions in both chow- and HFD-fed mice. In terms of glucose metabolism, overexpression of hscEMC10 impaired glucose tolerance and insulin sensitivity, and increased circulating insulin levels [28]. With regards to lipid metabolism, mice overexpressing hscEMC10 displayed increased fat mass accumulation, increased liver mass, increased circulating leptin levels and decreased circulating adiponectin levels [28].
Secreted factors that promote the development of metabolic disease. scEMC10, vimentin and CILP2 are three secreted factors with novel functions that drive the development of phenotypes associated with metabolic disease. Such functions include reducing glucose tolerance and insulin sensitivity, as well as promoting increased fat storage. CREB, cAMP-response element-binding protein. This figure is available as part of a downloadable slideset
While the receptor and tissue of action for scEMC10’s role in regulating energy metabolism is still unknown, scEMC10 is proposed to be transported intracellularly. Recombinant scEMC10 labelled with FITC is found intracellularly in HeLa cells. Co-immunoprecipitation also shows that scEMC10 binds to protein kinase A (PKA) catalytic α (PKA Cα), an intracellular catalytic subunit of PKA [28]. Additionally, scEMC10 signals in adipocytes by inducing phosphorylation of cAMP-response element-binding protein (CREB) at serine 133 [28]. These data together demonstrate the therapeutic potential of inhibiting scEMC10 to improve metabolic function. However, we still lack a full understanding of scEMC10’s endogenous functions. While Wang et al generated a global EMC10-knockout mouse, the results from these in vivo loss-of-function studies cannot differentiate between the effects of scEMC10 and mEMC10 or the whole parent protein itself [28]. Due to technical challenges, we are limited by the inability to create loss-of-function models for a specific peptide that does not affect the parent protein. Nevertheless, scEMC10 is a novel peptide hormone regulating glucose and lipid metabolism (Fig. 1). Additionally, two human studies revealed correlations between serum scEMC10 levels and phenotypes of metabolic disease: scEMC10 serum levels were positively correlated with increasing BMI scores and negatively correlated with decreasing resting metabolic rate [28, 30], two indicators of dysregulated energy metabolism. More studies are needed to determine whether scEMC10 serum levels are linked to dysregulated glucose and lipid metabolism and whether neutralising scEMC10 function in humans can ameliorate dysregulated energy metabolism.
Vimentin
Vimentin, a 57 kDa protein, is one of the most well-studied intermediate filament proteins [31,32,33]. In 2023, it was discovered that vimentin is secreted by adipocytes in response to oxidised LDLs (oxLDLs) to regulate both lipid and glucose metabolism in vitro (Fig. 1) [34]. This oxLDL-dependent secretion of vimentin relies on its PKA-dependent phosphorylation at serine 72 [34]. Vimentin does not contain a signal peptide, thus is not secreted through the conventional secretion pathway. Rather, vimentin may be secreted through the vesicular type III unconventional protein secretion pathway. In tumour endothelial cells, inhibitors of vesicular secretion pathways strongly blocks vimentin secretion into the cell media [35]. Once secreted, vimentin acts in adipocytes to regulate glucose metabolism (Fig. 1). Adipocytes treated with vimentin for 24 h increase their glucose uptake by promoting the translocation of GLUT1 and GLUT4 proteins to the plasma membrane, without an increase in phosphorylated Akt protein levels [34]. Interestingly, administration of vimentin and insulin together have an additive effect on glucose uptake [34], suggesting that vimentin can induce glucose uptake in a parallel pathway to insulin. Surprisingly, this is accompanied by an increase in GLUT1 mRNA levels but a decrease in GLUT4 mRNA levels. The vimentin-dependent regulation of GLUT1 mRNA is mediated by phosphorylation and activation of the IGF-1 receptor (IGF-1R) and ERK [34]. While direct binding between vimentin and IGF-1Rs has not been tested in adipocytes, a previous study demonstrated vimentin–IGF-1R binding in neurons [36]. Thus, it is possible that vimentin mediates its glucose regulatory effects in adipocytes via IGF-1R signalling.
Regarding lipid metabolism, administration of recombinant vimentin to 3T3-L1 adipocytes was found to increase NEFA uptake via increased expression of the NEFA transporter CD36 [34]. Vimentin also increased the proportion of larger adipocytes and larger lipid droplets, consequently increasing lipid stores as measured by Oil-Red-O staining and triglyceride levels [34]. Vimentin treatment of cultured adipocytes simultaneously increased protein levels of sterol regulatory element-binding protein 1 (SREBP1), a key regulator of lipogenesis [34], and reduced protein levels of several lipolytic factors (e.g. peroxisome proliferator-activated receptor γ [PPARγ] and adipose triglyceride lipase [ATGL]) [34]. Additionally, vimentin treatment induced markers of ER stress and impaired autophagy [34]. Together, these in vitro data present a model whereby vimentin promotes energy storage in adipocytes by increasing glucose and NEFA uptake and inducing lipogenesis and ER stress, while inhibiting lipolysis and autophagy. While more in vivo studies are needed to confirm vimentin’s role as a secreted factor in the regulation of energy metabolism in humans, one study found that individuals with type 2 diabetes and increased blood glucose levels also have increased serum vimentin levels [37]. Thus, neutralising vimentin’s effects in vivo may be a viable therapeutic option for improving glucose regulation.
Cartilage intermediate layer protein 2
Cartilage intermediate layer protein 2 (CILP2), a classically secreted 125 kDa glycoprotein, was first identified in 2003 and is largely found in cartilaginous tissues [38]. CILP2 mRNA transcript levels are widely distributed in mice, with the highest levels being found in skeletal and cardiac muscle, adipose tissue and liver [39]. While there is little work on CILP2’s functions and mechanisms related to energy homeostasis, a study in 2019 investigated the causative role of CILP2 in developing phenotypes of metabolic disease (Fig. 1) [40]. Specifically, AAV-mediated global CILP2 overexpression exacerbated HFD-induced increased blood glucose levels, increased serum insulin concentration, impaired glucose tolerance, reduced glucose uptake and reduced insulin sensitivity in male mice [40]. Additionally, CILP2 overexpression further increased serum concentrations of cholesterol, triglycerides and NEFA in HFD-fed mice [40]. Interestingly, AAV-mediated overexpression of CILP2 increased CILP2 transcript levels to the greatest extent in the liver, suggesting a hepatic role for CILP2 in regulating energy metabolism [40]. In vitro studies on cultured hepatocytes revealed that overexpression of CILP2 promotes the expression of PEPCK, a key enzyme in hepatic gluconeogenesis [40]. Simultaneously, overexpression of CILP2 blocked insulin-induced activation of the Akt signalling pathway [40]. Together, this suggests a model whereby CILP2 drives the development of metabolic disease by inhibiting insulin signalling and promoting gluconeogenesis (Fig. 1). However, the field still lacks loss-of-function and mechanistic studies to elucidate the molecular mechanisms underlying CILP2’s action. In particular, the CILP2 receptor remains unknown. Meanwhile, recent studies in humans revealed that high levels of CILP2 in serum are associated with decreased insulin sensitivity, increased blood glucose, serum insulin and serum triglyceride levels, and increased BMI [41, 42]. Notably, serum CILP2 levels are significantly higher in men than in women [41]. As men have decreased insulin sensitivity compared with women [43], it is possible that CILP2 levels are regulated by sex hormones such as testosterone. Together, these lines of evidence demonstrate a clear association between CILP2 and various metabolic phenotypes, although additional studies are needed to determine whether the regulation of CILP2 is a causative driver of metabolic disease.
Secreted factors that protect against the development of metabolic disease
Isthmin-1
Isthmin-1 (ISM1) is a widely expressed, 60 kDa secreted protein that was originally identified in 2002 and studied for its role in neural development [43, 44]. In 2021, it was discovered that ISM1 has dual functions in promoting glucose uptake in adipose tissue and preventing lipid synthesis in the liver (Fig. 2) [45]. AAV-mediated overexpression of ISM1 in male mice was found to significantly improve glucose tolerance and insulin sensitivity and to inhibit triglyceride accumulation in the liver after 10 weeks of HFD [45]. Importantly, administration of recombinant ISM1 significantly improved glucose and insulin tolerance and reduced hepatic lipid accumulation in mice fed a metabolic dysfunction-associated steatotic liver disease (MASLD)-inducing diet. ISM1 administration also blocked insulin-induced lipogenesis, as measured by acetate incorporation in the mouse hepatic cell line AML12 and expression levels of lipogenesis genes in primary mouse hepatocytes [45].
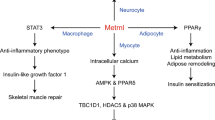
Secreted factors that protect against the development of metabolic disease. ISM1, LCN2, NRG1 and NRG4 are secreted factors with novel functions that protect against the development of phenotypes associated with metabolic disease. These four secreted factors improve glucose tolerance and insulin sensitivity while reducing fat storage via increased fatty acid oxidation and decreased lipogenesis. LCN2 further supports glucose metabolism by promoting beta cell proliferation and thus increased insulin secretion. MC4R, melatonin-4 receptor; STAT5, signal transducer and activator of transcription 5. This figure is available as part of a downloadable slideset
Concurrently, ISM1 promotes the synthesis of hepatic proteins, as measured by increased leucine incorporation and protein levels of phosphorylated S6, a kinase involved in protein synthesis [45]. In adipocytes [45] and myocytes [46], ISM1’s regulatory effects on metabolism are mediated through the phosphoinositide 3-kinase (PI3K)–Akt signalling pathway, independent of insulin. Specifically, in 3T3-F442A cells treated with ISM1, a phosphokinase array revealed that ISM1 phosphorylated Akt most strongly at serine 473 [45]. This was further confirmed via an ELISA and western blot assays to measure phosphorylated Akt levels after ISM1 administration to 3T3-F442A cells [45]. However, while we know that ISM1 activates the PI3K–Akt signalling pathway, the receptor for ISM1 is unknown. Future studies will be needed to determine whether ISM1 is a viable therapeutic target for treating metabolic disease. In adult humans, serum ISM1 levels are decreased in individuals with type 2 diabetes compared with healthy control individuals [47]. ISM1 serum levels were also found to be positively correlated with increasing obesity in a cohort of pubescent boys [48], suggesting an additional function of Ism1 in regulating adiposity. Overall, ISM1 is an interesting secreted factor that may play many roles in the control of energy metabolism.
Lipocalin-2
Lipocalin-2 (LCN2), a 25 kDa osteokine, is widely distributed and is a classically secreted glycoprotein [49]. Originally, LCN2 was studied as an adipokine that decreased insulin sensitivity [50]. However, it was discovered in 2017 that LCN2 expression levels are tenfold higher in bone than in adipose tissue [51], leading to the discovery that osteoblast-derived LCN2 is a potent appetite suppressor and metabolic regulator (Fig. 2). Loss of LCN2 in osteoblasts was found to significantly increase fat pad mass and food intake in male mice [51]. This effect was accompanied by reduced glucose and insulin tolerance as well as impaired glucose-stimulated insulin secretion (GSIS) [51]. Importantly, LCN2 shows therapeutic promise as it acts as an anorexigenic hormone in both lean and obese mice. Daily i.p. injections of 150 ng/g of recombinant LCN2 reduced food intake by 18%, reduced fat mass by 32%, reduced overall body weight by 9.4% and improved glucose and insulin tolerance in wild-type mice [51]. Daily LCN2 injections into leptin-receptor-deficient mice ameliorated their hyperphagia and fat mass gain, improved GSIS and completely reversed glucose intolerance [51]. The appetite-suppressing effects of LCN2 are mediated through melanocortin 4 receptor (MC4R) signalling in the paraventricular hypothalamus [51]. LCN2’s appetite-suppressing effects are also seen in primates where a 0.0375 mg/kg dose of LCN2 reduced food intake by 21% [52]. There may be more to LCN2’s therapeutic potential than just its actions as an anorexigenic factor. Loss of LCN2 in osteoblasts also decreased beta cell mass and proliferation [51]. One major question left to answer regarding LCN2’s anorexigenic capabilities is its mechanism of action. LCN2 has two known receptors, megalin and solute carrier family 22 member 17 (SLC22A17) [49]. However, megalin is not expressed in cultured murine hypothalamic neurons where LCN2 is able to induce cAMP activity and silencing SLC22A17 in these same neurons did not affect LCN2 signalling [51]. There is much left to elucidate regarding osteoblast-derived LCN2’s mechanisms of action as well as LCN2’s role in metabolic disease development in humans. In humans, serum levels of LCN2 are positively correlated with increasing BMI [53]. Notably, serum LCN2 levels are influenced by biological sex, as men have higher concentrations than women [54]. Given the myriad of sex differences in glucose metabolism and beta cell function [55, 56], serum LCN2 levels may be regulated by sex hormones such as oestrogen and testosterone. It will be necessary for future studies investigating LCN2’s function to consider that men and women may regulate and respond to LCN2 differently.
Neuregulins
The neuregulins (NRGs) are a family of four EGF-like secreted proteins (NRG1–4) best known for their roles in development, regulation of the nervous system and inter-organ communication [57].
NRG1
Initial in vitro studies revealed that NRG1 can stimulate glucose uptake by promoting the translocation of GLUT4 to the cell surface via activation of the PI3K–Akt signalling pathway [58,59,60,61]. These findings were not demonstrated in vivo until 2015. In db/db mice, acute or chronic injections of recombinant NRG1 significantly improved glucose tolerance and reduced both blood glucose and serum insulin levels (Fig. 2) [62]. These effects are partially mediated by the liver; an acute injection of NRG1 induced phosphorylation of the known receptor erb-b2 receptor tyrosine kinase 3 (ErbB3) only in the liver [62]. In addition to improved glucose clearance, NRG1 injection into db/db mice inhibited hepatic gluconeogenesis via phosphorylation of Akt and forkhead box protein O1 [62]. Further, in vitro assays in skeletal myocytes previously revealed that NRG1 promotes glucose and fatty acid oxidation via increased mitochondrial mass and activity [63, 64]. In 2017, these findings were replicated in vivo: 8 weeks of treatment with recombinant NRG1 significantly reduced body weight, improved mitochondrial function and increased mitochondrial complex 2 subunit content in male mice [65]. Importantly, serum levels of NRG1 are significantly decreased in individuals with type 2 diabetes compared with healthy control individuals [66], and are significantly higher in women than in men [67]. Given that women have a lower prevalence of type 2 diabetes and increased insulin sensitivity compared with men [55], the sex difference in serum NRG1 levels may be regulated by sex hormones such as oestrogen. Thus, NRG1 may be leveraged as a therapeutic agent to improve glucose regulation and mitochondrial function in humans. However, NRG1 has a very short t½ in circulation [68]. Thus, a fusion protein of human NRG1 and the Fc domain of human IgG1 (NRG1-Fc), with a markedly longer plasma t½ of 40 h, has been generated [69]. A single NRG1-Fc injection significantly improved glucose tolerance and reduced both blood glucose and plasma insulin levels in chow- and HFD-fed mice. Daily injections for 8 days into HFD-fed mice significantly improved glucose regulation and reduced body weight gain and food intake [69]. Together, these studies demonstrate the potential of NRG1 as a therapeutic agent for treating obesity and type 2 diabetes.
NRG4
In addition to NRG1, studies in the last decade have also established novel functions of NRG4 in the regulation of glucose and lipid metabolism (Fig. 2). NRG4 mRNA transcripts were originally detected only in the pancreas and the muscle [70, 71]. However, secretome analysis revealed that NRG4 is more highly expressed in BAT and is moderately expressed in subcutaneous white adipose tissue [72, 73]. In HFD-fed male mice, overexpression of Nrg4 was found to globally attenuate fat mass gain, increase plasma leptin levels and glucose intolerance, and decrease insulin sensitivity [74, 75]. Conversely, whole-body loss of Nrg4 led to impaired glucose metabolism, indicated by worsened glucose and insulin tolerance and increased blood glucose and serum insulin levels [72]. NRG4’s effects on glucose metabolism are mediated by pancreatic beta cells and adipocytes. In cultured rat islets, NRG4 induced the greatest insulin release when compared with the other NRGs [71]. In vitro assays in cultured adipocytes with Nrg4 knockdown revealed decreased protein levels of both insulin receptor and the major GLUT, GLUT4 [76]. Decreased GLUT4 protein levels due to loss of Nrg4 is mediated by reduced mTOR phosphorylation and thus increased autophagy of GLUT4 storage vesicles [76]. Importantly, NRG4 treatment is sufficient to increase both insulin receptor expression and GLUT4 protein levels [76]. Thus, NRG4 may maintain glucose homeostasis by promoting glucose uptake in adipocytes.
In addition to NRG4’s effects on glucose metabolism, NRG4 acts in the liver to suppress hepatic lipogenesis [72]. Whole-body loss of Nrg4 in mice promoted lipid accumulation in the liver, increased circulating triglyceride levels and increased hepatic mRNA transcript levels of de novo lipogenesis genes (e.g. Gck and Fasn), inflammatory markers (e.g. Tnfa, Il1b), and fibrosis markers (e.g. Col1a1, Acta2) [72, 77, 78]. Exogenous NRG4 can block the upregulation of lipogenic genes and incorporation of acetate into lipids induced by pharmaceutically activating SREBP1c [72]. In vitro experiments using primary mouse hepatocytes revealed that NRG4 acts through its receptor ErbB4 to phosphorylate Akt and signal transducer and activator of transcription 5 (STAT5) to modulate its protective functions [72, 77, 78]. While it is clear that adipose-derived NRG4 has important roles in regulating glucose and lipid metabolism, the relationship between serum NRG4 levels and metabolic disease in humans is less clear. Some studies revealed that individuals with gestational diabetes, obesity or MASLD have reduced serum NRG4 levels compared with control individuals [79,80,81,82]. However, one study found no link between serum NRG4 levels and MASLD [83] while another study found a positive correlation between serum NRG4 levels and insulin resistance [84]. Future studies will be needed to determine whether NRG4 can be used as a biomarker of metabolic disease development and whether NRG4 can be leveraged to treat metabolic diseases.
Conclusions and future directions
Significant progress has been made in the last decade in the discovery of novel hormones and in revealing additional functions of known factors that contribute to the regulation of glucose and lipid metabolism. As novel secreted factors and functions are revealed, there is a need to understand the mechanisms underlying their regulatory actions. Specifically, identifying receptors for regulatory hormones is vital. Of the hormones discussed in this review, only vimentin, NRG1 and NRG4 have known receptors that modulate energy homeostasis. Identifying receptors is a major challenge due to technical limitations. To put this into perspective, while insulin was discovered in 1921, its receptor was not discovered until the early 1970s. Similarly, EGF was discovered in 1962 [85] and its receptor was discovered 18 years later [86, 87]. With the advent of computational tools for predicting protein structure, in silico screening for ligand–receptor interactions will speed up this process. However, we are still limited to a candidate approach. The following are examples of recent technologies that have made higher-throughput screening possible: (1) Retrogenix Cell Microarray, used to identify the receptor for growth differentiating factor 15 (GDF15) [88, 89]; (2) a ligand-based receptor capturing technique in live cells and tissues called TRICEPS, used to identify novel receptors for C1q TNF-related protein 3 (CTRP3) [90] and stanniocalcin-1 (STC1) [91,92,93]; and (3) a pooled CRISPR/Cas9-based transcriptional activation screen (CRISPRa), used to deorphanise several receptor protein tyrosine phosphatase ligands and a novel ligand that binds to killer immunoglobulin-like receptors [94, 95]. These newer techniques are now being used alongside classical cDNA cloning methods to identify ligand–receptor pairs but there is still much room for improvement and it will be an important hurdle to cross for the metabolism field in order to design more effective therapeutics.
Altogether, this review highlights some of the most recent discoveries of secreted factors involved in maintaining energy homeostasis. While this review focused on novel secreted peptides and proteins, other classes of secreted factors also contribute to maintaining energy homeostasis such as secreted lipids, metabolites and small open reading frames (sORFs) [15, 96, 97]. These other classes of secreted factors, alongside almost 2000 genes encoding secreted proteins and peptides, reveal the vastness of the mammalian secretome. The secretome is a powerful tool for understanding energy metabolism and studies in this field will be foundational for new and potentially more effective treatments for many metabolic diseases.
Abbreviations
- AAV:
-
Adeno-associated virus
- ATGL:
-
Adipose triglyceride lipase
- BAT:
-
Brown adipose tissue
- CILP2:
-
Cartilage intermediate layer protein 2
- EMC10:
-
Endoplasmic reticulum membrane complex subunit 10
- ER:
-
Endoplasmic reticulum
- ErbB:
-
Erb-b2 receptor tyrosine kinase
- GSIS:
-
Glucose-stimulated insulin secretion
- HFD:
-
High-fat diet
- hscEMC10:
-
Human scEMC10
- IGF-1R:
-
IGF-1 receptor
- ISM1:
-
Isthmin-1
- LCN2:
-
Lipocalin-2
- MASLD:
-
Metabolic dysfunction-associated steatotic liver disease
- mEMC10:
-
Membrane-bound EMC10
- NRG:
-
Neuregulin
- NRG1-Fc:
-
Fusion protein of human Nrg1 and the Fc domain of human IgG1
- oxLDL:
-
Oxidised LDL
- PI3K:
-
Phosphoinositide 3-kinase
- PKA:
-
Protein kinase A
- PPARγ:
-
Peroxisome proliferator-activated receptor γ
- scEMC10:
-
Secreted isoform of EMC10
- SREBP1:
-
Sterol regulatory element-binding protein 1
References
Banting FG, Best CH (1987) The Journal of Laboratory and Clinical Medicine: the internal secretion of the pancreas. Nutr Rev 45(4):55–57. https://doi.org/10.1111/j.1753-4887.1987.tb07442.x
Rahman MS, Hossain KS, Das S et al (2021) Role of insulin in health and disease: an update. Int J Mol Sci 22(12):6403. https://doi.org/10.3390/ijms22126403
Banting FG, Best CH, Collip JB, Campbell WR, Fletcher AA (1922) Pancreatic extracts in the treatment of diabetes mellitus. Can Med Assoc J 12(3):141–146
Kimball CP, Murlin JR (1923) Aqueous extracts of pancreas: iii. Some precipitation reactions of insulin. J Biol Chem 58(1):337–346. https://doi.org/10.1016/S0021-9258(18)85474-6
Sutherland EW, Cori CF (1949) Purification of the hyperglycemic-glycogenolytic factor from insulin and from gastric mucosa. J Biol Chem 180(2):825–837. https://doi.org/10.1016/S0021-9258(18)56702-8
Hædersdal S, Andersen A, Knop FK, Vilsbøll T (2023) Revisiting the role of glucagon in health, diabetes mellitus and other metabolic diseases. Nat Rev Endocrinol 19(6):321–335. https://doi.org/10.1038/s41574-023-00817-4
Drucker DJ, Habener JF, Holst JJ (2017) Discovery, characterization, and clinical development of the glucagon-like peptides. J Clin Invest 127(12):4217–4227. https://doi.org/10.1172/JCI97233
Baggio LL, Drucker DJ (2007) Biology of incretins: GLP-1 and GIP. Gastroenterology 132(6):2131–2157. https://doi.org/10.1053/j.gastro.2007.03.054
Obradovic M, Sudar-Milovanovic E, Soskic S et al (2021) Leptin and obesity: role and clinical implication. Front Endocrinol 12:585887. https://doi.org/10.3389/fendo.2021.585887
Berg AH, Combs TP, Du X, Brownlee M, Scherer PE (2001) The adipocyte-secreted protein Acrp30 enhances hepatic insulin action. Nat Med 7(8):947–953. https://doi.org/10.1038/90992
Yamauchi T, Kamon J, Waki H et al (2001) The fat-derived hormone adiponectin reverses insulin resistance associated with both lipoatrophy and obesity. Nat Med 7(8):941–946. https://doi.org/10.1038/90984
Combs TP, Berg AH, Obici S, Scherer PE, Rossetti L (2001) Endogenous glucose production is inhibited by the adipose-derived protein Acrp30. J Clin Invest 108(12):1875–1881. https://doi.org/10.1172/JCI14120
Wang ZV, Scherer PE (2016) Adiponectin, the past two decades. J Mol Cell Biol 8(2):93–100. https://doi.org/10.1093/jmcb/mjw011
Uhlén M, Fagerberg L, Hallström BM et al (2015) Tissue-based map of the human proteome. Science 347(6220):1260419. https://doi.org/10.1126/science.1260419
Reghupaty SC, Dall NR, Svensson KJ (2024) Hallmarks of the metabolic secretome. Trends Endocrinol Metab 35(1):49–61. https://doi.org/10.1016/j.tem.2023.09.006
Chen Z-Z, Gerszten RE (2020) Metabolomics and proteomics in type 2 diabetes. Circ Res 126(11):1613–1627. https://doi.org/10.1161/CIRCRESAHA.120.315898
Rodriguez-Muñoz A, Motahari-Rad H, Martin-Chaves L et al (2024) A systematic review of proteomics in obesity: unpacking the molecular puzzle. Curr Obes Rep. https://doi.org/10.1007/s13679-024-00561-4
Eldjarn GH, Ferkingstad E, Lund SH et al (2023) Large-scale plasma proteomics comparisons through genetics and disease associations. Nature 622(7982):348–358. https://doi.org/10.1038/s41586-023-06563-x
Thul PJ, Åkesson L, Wiking M et al (2017) A subcellular map of the human proteome. Science 356(6340):eaal3321. https://doi.org/10.1126/science.aal3321
Reboll MR, Korf-Klingebiel M, Klede S et al (2017) EMC10 (Endoplasmic reticulum membrane protein complex subunit 10) is a bone marrow-derived angiogenic growth factor promoting tissue repair after myocardial infarction. Circulation 136(19):1809–1823. https://doi.org/10.1161/CIRCULATIONAHA.117.029980
Zhou Y, Wu F, Zhang M et al (2018) EMC10 governs male fertility via maintaining sperm ion balance. J Mol Cell Biol 10(6):503–514. https://doi.org/10.1093/jmcb/mjy024
Liu L, Mao S, Chen K et al (2022) Membrane-bound EMC10 is required for sperm motility via maintaining the homeostasis of cytoplasm sodium in sperm. Int J Mol Sci 23(17):10069. https://doi.org/10.3390/ijms231710069
Umair M, Alfadhel M (1993) EMC10-Related Neurodevelopmental Disorder. In: Adam MP, Feldman J, Mirzaa GM, et al (eds) GeneReviews®. University of Washington, Seattle, Seattle (WA)
Kaiyrzhanov R, Rocca C, Suri M et al (2022) Biallelic loss of EMC10 leads to mild to severe intellectual disability. Ann Clin Transl Neurol 9(7):1080–1089. https://doi.org/10.1002/acn3.51602
Haddad-Eid E, Gur N, Eid S, Pilowsky-Peleg T, Straussberg R (2022) The phenotype of homozygous EMC10 variant: A new syndrome with intellectual disability and language impairment. Eur J Paediatr Neurol 37:56–61. https://doi.org/10.1016/j.ejpn.2022.01.012
Shao DD, Straussberg R, Ahmed H et al (2021) A recurrent, homozygous EMC10 frameshift variant is associated with a syndrome of developmental delay with variable seizures and dysmorphic features. Genet Med 23(6):1158–1162. https://doi.org/10.1038/s41436-021-01097-x
The UniProt Consortium (2023) UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Research 51(D1):D523–D531. https://doi.org/10.1093/nar/gkac1052
Wang X, Li Y, Qiang G et al (2022) Secreted EMC10 is upregulated in human obesity and its neutralizing antibody prevents diet-induced obesity in mice. Nat Commun 13(1):7323. https://doi.org/10.1038/s41467-022-34259-9
Wang X, Gong W, Liu Y et al (2009) Molecular cloning of a novel secreted peptide, INM02, and regulation of its expression by glucose. J Endocrinol 202(3):355–364. https://doi.org/10.1677/JOE-09-0086
Chen K, Dai J, Klöting N et al (2023) Serum scEMC10 levels are negatively associated with resting metabolic rate and age in humans. J Clin Endocrinol Metab 108(10):e1074–e1081. https://doi.org/10.1210/clinem/dgad214
Ridge KM, Eriksson JE, Pekny M, Goldman RD (2022) Roles of vimentin in health and disease. Genes Dev 36(7–8):391–407. https://doi.org/10.1101/gad.349358.122
Satelli A, Li S (2011) Vimentin as a potential molecular target in cancer therapy Or Vimentin, an overview and its potential as a molecular target for cancer therapy. Cell Mol Life Sci 68(18):3033–3046. https://doi.org/10.1007/s00018-011-0735-1
Mor-Vaknin N, Punturieri A, Sitwala K, Markovitz DM (2003) Vimentin is secreted by activated macrophages. Nat Cell Biol 5(1):59–63. https://doi.org/10.1038/ncb898
Park J-H, Kwon S, Park YM (2023) Extracellular vimentin alters energy metabolism and induces adipocyte hypertrophy. Diabetes Metab J 48(2):215–230. https://doi.org/10.4093/dmj.2022.0332
van Beijnum JR, Huijbers EJM, van Loon K et al (2022) Extracellular vimentin mimics VEGF and is a target for anti-angiogenic immunotherapy. Nat Commun 13:2842. https://doi.org/10.1038/s41467-022-30063-7
Shigyo M, Kuboyama T, Sawai Y, Tada-Umezaki M, Tohda C (2015) Extracellular vimentin interacts with insulin-like growth factor 1 receptor to promote axonal growth. Sci Rep 5:12055. https://doi.org/10.1038/srep12055
Gong DH, Dai Y, Chen S et al (2019) Secretory vimentin is associated with coronary artery disease in patients and induces atherogenesis in ApoE−/− mice. Int J Cardiol 283:9–16. https://doi.org/10.1016/j.ijcard.2019.02.032
Johnson K, Farley D, Hu S-I, Terkeltaub R (2003) One of two chondrocyte-expressed isoforms of cartilage intermediate-layer protein functions as an insulin-like growth factor 1 antagonist. Arthritis Rheum 48(5):1302–1314. https://doi.org/10.1002/art.10927
Bernardo BC, Belluoccio D, Rowley L, Little CB, Hansen U, Bateman JF (2011) Cartilage Intermediate Layer Protein 2 (CILP-2) is expressed in articular and meniscal cartilage and down-regulated in experimental osteoarthritis. J Biol Chem 286(43):37758–37767. https://doi.org/10.1074/jbc.M111.248039
Wu T, Zhang Q, Wu S et al (2019) CILP-2 is a novel secreted protein and associated with insulin resistance. J Mol Cell Biol 11(12):1083–1094. https://doi.org/10.1093/jmcb/mjz016
Hu W, Li K, Han H et al (2020) Circulating levels of CILP2 are elevated in coronary heart disease and associated with atherosclerosis. Oxid Med Cell Longev 2020:1871984. https://doi.org/10.1155/2020/1871984
Li Q, Pu D, Xia X, Liu H, Li L (2022) Serum concentrations of cartilage intermediate layer protein 2 were higher in overweight and obese subjects. Biomed Res Int 2022:6290064. https://doi.org/10.1155/2022/6290064
Osório L, Wu X, Zhou Z (2014) Distinct spatiotemporal expression of ISM1 during mouse and chick development. Cell Cycle 13(10):1571–1582. https://doi.org/10.4161/cc.28494
Pera EM, Kim JI, Martinez SL et al (2002) Isthmin is a novel secreted protein expressed as part of the Fgf-8 synexpression group in the Xenopus midbrain-hindbrain organizer. Mech Dev 116(1–2):169–172. https://doi.org/10.1016/s0925-4773(02)00123-5
Jiang Z, Zhao M, Voilquin L et al (2021) Isthmin-1 is an adipokine that promotes glucose uptake and improves glucose tolerance and hepatic steatosis. Cell Metabolism 33(9):1836–1852. https://doi.org/10.1016/j.cmet.2021.07.010
Zhao M, BanhosDanneskiold-Samsøe N, Ulicna L et al (2022) Phosphoproteomic mapping reveals distinct signaling actions and activation of muscle protein synthesis by Isthmin-1. Elife 11:e80014. https://doi.org/10.7554/eLife.80014
Wang J, Du J, Ge X, Peng W, Li W (2022) Circulating Ism1 reduces the risk of type 2 diabetes but not diabetes-associated NAFLD. Front Endocrinol 13:890332. https://doi.org/10.3389/fendo.2022.890332
Ruiz-Ojeda FJ, Anguita-Ruiz A, Rico MC et al (2023) Serum levels of the novel adipokine isthmin-1 are associated with obesity in pubertal boys. World J Pediatr 19(9):864–872. https://doi.org/10.1007/s12519-022-00665-8
Jaberi SA, Cohen A, D’Souza C et al (2021) Lipocalin-2: Structure, function, distribution and role in metabolic disorders. Biomed Pharmacother 142:112002. https://doi.org/10.1016/j.biopha.2021.112002
Yan Q-W, Yang Q, Mody N et al (2007) The adipokine lipocalin 2 is regulated by obesity and promotes insulin resistance. Diabetes 56(10):2533–2540. https://doi.org/10.2337/db07-0007
Mosialou I, Shikhel S, Liu J-M et al (2017) MC4R-dependent suppression of appetite by bone-derived lipocalin 2. Nature 543(7645):385–390. https://doi.org/10.1038/nature21697
Petropoulou P-I, Mosialou I, Shikhel S et al (2020) Lipocalin-2 is an anorexigenic signal in primates. eLife 9:e58949. https://doi.org/10.7554/eLife.58949
Zheng Y, Rajcsanyi LS, Kowalczyk M et al (2023) Lipocalin 2 – mutation screen and serum levels in patients with anorexia nervosa or obesity and in lean individuals. Front Endocrinol 14:1137308. https://doi.org/10.3389/fendo.2023.1137308
Ni J, Ma X, Zhou M et al (2013) Serum lipocalin-2 levels positively correlate with coronary artery disease and metabolic syndrome. Cardiovasc Diabetol 12(1):176. https://doi.org/10.1186/1475-2840-12-176
Tramunt B, Smati S, Grandgeorge N et al (2020) Sex differences in metabolic regulation and diabetes susceptibility. Diabetologia 63(3):453–461. https://doi.org/10.1007/s00125-019-05040-3
Gannon M, Kulkarni RN, Tse HM, Mauvais-Jarvis F (2018) Sex differences underlying pancreatic islet biology and its dysfunction. Mol Metab 15:82–91. https://doi.org/10.1016/j.molmet.2018.05.017
Cespedes JC, Liu M, Harbuzariu A et al (2018) Neuregulin in health and disease. Int J Brain Disord Treat 4(1):024. https://doi.org/10.23937/2469-5866/1410024
Suárez E, Bach D, Cadefau J, Palacin M, Zorzano A, Gumá A (2001) A novel role of neuregulin in skeletal muscle. Neuregulin stimulates glucose uptake, glucose transporter translocation, and transporter expression in muscle cells. J Biol Chem 276(21):18257–18264. https://doi.org/10.1074/jbc.M008100200
Cote GM, Miller TA, Lebrasseur NK, Kuramochi Y, Sawyer DB (2005) Neuregulin-1alpha and beta isoform expression in cardiac microvascular endothelial cells and function in cardiac myocytes in vitro. Exp Cell Res 311(1):135–146. https://doi.org/10.1016/j.yexcr.2005.08.017
Cantó C, Suárez E, Lizcano JM et al (2004) Neuregulin signaling on glucose transport in muscle cells. J Biol Chem 279(13):12260–12268. https://doi.org/10.1074/jbc.M308554200
Yu M, Wu S, Gong C, Chen L (2023) Neuregulin-1β increases glucose uptake and promotes GLUT4 translocation in palmitate-treated C2C12 myotubes by activating PI3K/AKT signaling pathway. Front Pharmacol 13:1066279. https://doi.org/10.3389/fphar.2022.1066279
Ennequin G, Boisseau N, Caillaud K et al (2015) Neuregulin 1 improves glucose tolerance in db/db mice. PLOS ONE 10(7):e0130568. https://doi.org/10.1371/journal.pone.0130568
Cantó C, Pich S, Paz JC et al (2007) Neuregulins increase mitochondrial oxidative capacity and insulin sensitivity in skeletal muscle cells. Diabetes 56(9):2185–2193. https://doi.org/10.2337/db06-1726
Gumà A, Díaz-Sáez F, Camps M, Zorzano A (2020) Neuregulin, an effector on mitochondria metabolism that preserves insulin sensitivity. Front Physiol 11:696. https://doi.org/10.3389/fphys.2020.00696
Ennequin G, Capel F, Caillaud K et al (2017) Neuregulin 1 improves complex 2-mediated mitochondrial respiration in skeletal muscle of healthy and diabetic mice. Sci Rep 7(1):1742. https://doi.org/10.1038/s41598-017-02029-z
Eldin AS, Fawzy O, Mahmoud E, Elaziz OHA, Enayet AEA, Khidr EG (2023) Serum neuregulin 1 in relation to ventricular function and subclinical atherosclerosis in type 2 diabetes patients. Endocrinol Diabetes Nutr (Engl Ed) 70(10):619–627. https://doi.org/10.1016/j.endien.2023.11.010
Shibuya M, Komi E, Wang R et al (2010) Measurement and comparison of serum neuregulin 1 immunoreactivity in control subjects and patients with schizophrenia: an influence of its genetic polymorphism. J Neural Transm 117(7):887–895. https://doi.org/10.1007/s00702-010-0418-3
Liu X, Gu X, Li Z et al (2006) Neuregulin-1/erbB-activation improves cardiac function and survival in models of ischemic, dilated, and viral cardiomyopathy. J Am Coll Cardiol 48(7):1438–1447. https://doi.org/10.1016/j.jacc.2006.05.057
Zhang P, Kuang H, He Y et al (2018) NRG1-Fc improves metabolic health via dual hepatic and central action. JCI Insight 3(5):e98522. https://doi.org/10.1172/jci.insight.98522
Harari D, Tzahar E, Romano J et al (1999) Neuregulin-4: a novel growth factor that acts through the ErbB-4 receptor tyrosine kinase. Oncogene 18(17):2681–2689. https://doi.org/10.1038/sj.onc.1202631
South JCM, Blackburn E, Brown IR, Gullick WJ (2013) The neuregulin system of ligands and their receptors in rat islets of Langerhans. Endocrinology 154(7):2385–2392. https://doi.org/10.1210/en.2012-2133
Wang G-X, Zhao X-Y, Meng Z-X et al (2014) The brown fat–enriched secreted factor Nrg4 preserves metabolic homeostasis through attenuation of hepatic lipogenesis. Nat Med 20(12):1436–1443. https://doi.org/10.1038/nm.3713
Tutunchi H, Ostadrahimi A, Hosseinzadeh-Attar M-J, Miryan M, Mobasseri M, Ebrahimi-Mameghani M (2020) A systematic review of the association of neuregulin 4, a brown fat-enriched secreted factor, with obesity and related metabolic disturbances. Obes Rev 21(2):e12952. https://doi.org/10.1111/obr.12952
Ma Y, Gao M, Liu D (2016) Preventing high fat diet-induced obesity and improving insulin sensitivity through neuregulin 4 gene transfer. Sci Rep 6:26242. https://doi.org/10.1038/srep26242
Chen Z, Wang G-X, Ma SL et al (2017) Nrg4 promotes fuel oxidation and a healthy adipokine profile to ameliorate diet-induced metabolic disorders. Mol Metab 6(8):863–872. https://doi.org/10.1016/j.molmet.2017.03.016
Díaz-Sáez F, Blanco-Sinfreu C, Archilla-Ortega A et al (2021) Neuregulin 4 downregulation induces insulin resistance in 3T3-L1 adipocytes through inflammation and autophagic degradation of GLUT4 vesicles. Int J Mol Sci 22(23):12960. https://doi.org/10.3390/ijms222312960
Guo L, Zhang P, Chen Z et al (2017) Hepatic neuregulin 4 signaling defines an endocrine checkpoint for steatosis-to-NASH progression. J Clin Invest 127(12):4449–4461. https://doi.org/10.1172/JCI96324
Li Y, Jin L, Jiang F et al (2021) Mutations of NRG4 contribute to the pathogenesis of nonalcoholic fatty liver disease and related metabolic disorders. Diabetes 70(10):2213–2224. https://doi.org/10.2337/db21-0064
Zhang L, Lu B, Wang W et al (2021) Alteration of serum neuregulin 4 and neuregulin 1 in gestational diabetes mellitus. Ther Adv Endocrinol Metab 12:20420188211049616. https://doi.org/10.1177/20420188211049614
Dai Y-N, Zhu J-Z, Fang Z-Y et al (2015) A case-control study: association between serum neuregulin 4 level and non-alcoholic fatty liver disease. Metabolism 64(12):1667–1673. https://doi.org/10.1016/j.metabol.2015.08.013
Wang R, Yang F, Qing L, Huang R, Liu Q, Li X (2019) Decreased serum neuregulin 4 levels associated with non-alcoholic fatty liver disease in children with obesity. Clin Obes 9(1):e12289. https://doi.org/10.1111/cob.12289
Tutunchi H, Mobasseri M, Aghamohammadzadeh N, Hooshyar J, Naeini F, Najafipour F (2021) Serum neuregulin 4 (NRG-4) level and non-alcoholic fatty liver disease (NAFLD): a case-control study. Int J Clin Pract 75(10):e14555. https://doi.org/10.1111/ijcp.14555
De Munck TJI, Boesch M, Verhaegh P et al (2021) Is there a role for neuregulin 4 in human nonalcoholic fatty liver disease? PLoS One 16(5):e0251822. https://doi.org/10.1371/journal.pone.0251822
Martínez C, Latorre J, Ortega F et al (2022) Serum neuregulin 4 is negatively correlated with insulin sensitivity in humans and impairs mitochondrial respiration in HepG2 cells. Front Physiol 13:950791. https://doi.org/10.3389/fphys.2022.950791
Cohen S (1962) Isolation of a mouse submaxillary gland protein accelerating incisor eruption and eyelid opening in the new-born animal. J Biol Chem 237:1555–1562. https://doi.org/10.1016/S0021-9258(19)83739-0
Edwin F, Wiepz GJ, Singh R et al (2006) A historical perspective of the EGF receptor and related systems. In: Patel TB, Bertics PJ (eds) Epidermal growth factor: methods and protocols. Humana Press, Totowa, NJ, pp 1–24
Cohen S, Carpenter G, King L (1980) Epidermal growth factor-receptor-protein kinase interactions. Co-purification of receptor and epidermal growth factor-enhanced phosphorylation activity. J Biol Chem 255(10):4834–4842
Mullican SE, Lin-Schmidt X, Chin C-N et al (2017) GFRAL is the receptor for GDF15 and the ligand promotes weight loss in mice and nonhuman primates. Nat Med 23(10):1150–1157. https://doi.org/10.1038/nm.4392
Freeth J, Soden J (2020) New advances in cell microarray technology to expand applications in target deconvolution and off-target screening. SLAS Discov 25(2):223–230. https://doi.org/10.1177/2472555219897567
Li Y, Ozment T, Wright GL, Peterson JM (2016) Identification of putative receptors for the novel adipokine CTRP3 using ligand-receptor capture technology. PLoS One 11(10):e0164593. https://doi.org/10.1371/journal.pone.0164593
Wan HT, Ng AH, Lee WK, Shi F, Wong CK-C (2022) Identification and characterization of a membrane receptor that binds to human STC1. Life Sci Alliance 5(11):e202201497. https://doi.org/10.26508/lsa.202201497
Frei AP, Moest H, Novy K, Wollscheid B (2013) Ligand-based receptor identification on living cells and tissues using TRICEPS. Nat Protoc 8(7):1321–1336. https://doi.org/10.1038/nprot.2013.072
Lopez-Garcia LA, Demiray L, Ruch-Marder S et al (2018) Validation of extracellular ligand-receptor interactions by Flow-TriCEPS. BMC Res Notes 11(1):863. https://doi.org/10.1186/s13104-018-3974-5
Siepe DH, Henneberg LT, Wilson SC et al (2022) Identification of orphan ligand-receptor relationships using a cell-based CRISPRa enrichment screening platform. eLife 11:e81398. https://doi.org/10.7554/eLife.81398
Chong Z-S, Ohnishi S, Yusa K, Wright GJ (2018) Pooled extracellular receptor-ligand interaction screening using CRISPR activation. Genome Biol 19:205. https://doi.org/10.1186/s13059-018-1581-3
Leong AZ-X, Lee PY, Mohtar MA, Syafruddin SE, Pung Y-F, Low TY (2022) Short open reading frames (sORFs) and microproteins: an update on their identification and validation measures. J Biomed Sci 29:19. https://doi.org/10.1186/s12929-022-00802-5
Hu F, Lu J, Matheson LS et al (2021) ORFLine: a bioinformatic pipeline to prioritize small open reading frames identifies candidate secreted small proteins from lymphocytes. Bioinformatics 37(19):3152–3159. https://doi.org/10.1093/bioinformatics/btab339
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Acknowledgements
We thank the Svensson lab (Stanford University School of Medicine) for feedback and discussions. Illustrations were created in Adobe Illustrator.
Funding
Work in the Svensson lab was funded by the NIH R01DK125260 and AHA 23IPA1042031 (KJS). LWW is supported by an American Heart Association Postdoctoral Fellowship. KJS has received research support from Merck.
Authors’ relationships and activities
The authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work.
Contribution statement
LWW and KJS conceptualised, wrote and edited the manuscript. Both authors approved the final manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wat, L.W., Svensson, K.J. Novel secreted regulators of glucose and lipid metabolism in the development of metabolic diseases. Diabetologia (2024). https://doi.org/10.1007/s00125-024-06253-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00125-024-06253-x