Abstract
Aims/hypothesis
Diabetic retinopathy is a common complication of diabetes and a leading cause of visual impairment and blindness. Despite recent advances, our understanding of its pathophysiology remains incomplete. The aim of this study was to provide deeper insight into the complex network of molecular and cellular changes that underlie diabetic retinopathy by systematically mapping the transcriptional changes that occur in the different cellular compartments of the degenerating diabetic mouse retina.
Methods
Single-cell RNA sequencing was performed on retinal tissue from 12-week-old wild-type and Akimba (Ins2Akita×Vegfa+/–) mice, which are known to replicate features of clinical diabetic retinopathy. This resulted in transcriptome data for 9474 retinal cells, which could be annotated to eight distinct retinal cell types. Using STRING analysis, we studied differentially expressed gene networks in neuronal, glial and immune cell compartments to create a comprehensive view on the pathological changes that occur in the Akimba retina. Using subclustering analysis, we further characterised macroglial and inflammatory cell subpopulations. Prominent findings were confirmed at the protein level using immunohistochemistry, western blotting and ELISA.
Results
At 12 weeks, the Akimba retina was found to display degeneration of rod photoreceptors and presence of inflammatory cells, identified by subclustering analysis as monocyte, macrophage and microglial populations. Analysis of differentially expressed genes in the rod, cone, bipolar cell and macroglial compartments indicated changes in cell metabolism and ribosomal gene expression, gliosis, activation of immune system pathways and redox and metal ion dyshomeostasis. Experiments at the protein level supported a metabolic shift from glycolysis to oxidative phosphorylation (glyceraldehyde 3-phosphate dehydrogenase), activation of microglia/macrophages (isolectin-B4), metal ion and oxidative stress response (metallothionein and haem oxygenase-1) and reactive macroglia (glial fibrillary acidic protein and S100) in the Akimba retina, compared with wild-type mice. Our single-cell approach also indicates macroglial subpopulations with distinct fibrotic, inflammatory and gliotic profiles.
Conclusions/interpretation
Our study identifies molecular pathways underlying inflammatory, metabolic and oxidative stress-mediated changes in the Akimba mouse model of diabetic retinopathy and distinguishes distinct functional subtypes of inflammatory and macroglial cells.
Data availability
RNA-seq data have been deposited in the ArrayExpress database at EMBL-EBI (www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-9061.

Graphical abstract
Similar content being viewed by others
Avoid common mistakes on your manuscript.

Introduction
Diabetic retinopathy is a common microvascular complication of diabetes and the leading cause of sight loss in the working-age population [1, 2]. Based on the extent of microvascular degeneration and ischaemic damage, it is staged into non-proliferative diabetic retinopathy or advanced, proliferative diabetic retinopathy. Diabetic retinopathy progression goes hand in hand with increasing abnormalities of the vasculature, including microaneurysms, haemorrhages, capillary occlusion, blood–retinal barrier dysfunction, thickening of vascular basement membrane and neovascularisation [1,2,3]. These complications may lead to diabetic macular oedema, which is characterised by vascular leakage and concomitant macular swelling and is the most common cause of blindness in diabetic retinopathy patients [1, 2].
It has become clear that diabetic retinopathy comprises neurodegenerative [4, 5] and neuroinflammatory [6, 7] disease components, with glial, neuronal and endothelial dysfunction being interdependent and essential for disease development. However, a lot remains unknown about which changes occur in these individual cellular compartments and how they interplay. The advent of single-cell RNA sequencing (scRNAseq) technology now provides the opportunity to dissect complex biological systems, such as the retina, at single-cell resolution.
The Akimba mouse model is an attractive animal model to study the pathological processes that underlie diabetic retinopathy. Akimba mice are generated by crossbreeding Akita (Ins2Akita) mice, carrying a mutation in the Ins2 gene, with Kimba (trVEGF029) mice, which have transient photoreceptor-specific overexpression of Vegfa, encoding human vascular endothelial growth factor (VEGF) [8]. The Akimba mouse displays several pathogenic characteristics of clinical diabetic retinopathy, including ischaemia, angiogenesis, oedema, neurodegeneration, inflammation and fibrosis. In addition, these mice exhibit vascular complications comparable to clinical diabetic retinopathy, such as microaneurysms, capillary nonperfusion, vessel constriction, venous beading, tortuous vessels, neovascularisation and leaky capillaries [8, 9].
Employing the Akimba mouse as a model of diabetic retinopathy and making use of scRNAseq technology, the goal of this study was to obtain a systematic and comprehensive view on the cell-type-specific changes that occur in retina affected by diabetic retinopathy.
Methods
Animals
Animal procedures were approved by the Institutional Ethical Committee of KU Leuven, according to the 2010/63/EU Directive. Twelve-week-old male Akimba and wild-type (C57Bl/6 J) were obtained from Charles River Laboratories (France) as previously described [10] and housed under standard laboratory conditions. Animals with blood glucose levels >13.87 mmol/l were considered diabetic. Optical coherence tomography and fluorescein angiography (Spectralis, Heidelberg Engineering, Germany) were performed to evaluate retinopathy and retinal vascular leakage. Mice were anaesthetised with ketamine (Anesketin, 75 mg/kg, Eurovet, the Netherlands) and medetomidine (Domitor, 1 mg/kg, Orion Pharma, Finland). Pupils were dilated with tropicamide (Tropicol, 5 mg/ml, Théa Pharma, Belgium). Fluorescein angiography scans were taken after subcutaneous injection of 100 μl sodium fluorescein (5% wt/vol.). Anaesthesia was reversed with atipamezole (Antisedan, 1 mg/kg, Orion Pharma, Finland). Randomisation and blinding were not performed during the course of the study, except during immunohistochemical analysis.
Retina dissociation, single-cell preparation and sequencing
Retinas (n = 4 for Akimba, n = 2 for wild-type) were isolated in ice-cold DMEM. After rinsing with Dulbecco’s PBS containing 2% FBS, each retina was incubated with 1 ml digestion buffer (2 mg/ml collagenase-P, 200 U/ml DNAse-I, Sigma-Aldrich, Belgium) in M199 medium (Life Technologies, Belgium) at 37°C for 10 min. Retinal tissue was further dissociated by trituration and the suspension was filtered through a 40 μm cell strainer and centrifuged for 5 min at 300 g (4°C). Pooled retinal single-cell suspensions from wild-type and Akimba mice were counted on a Luna-FL Cell Counter (Logos Biosystems, France) and libraries were prepared with the Chromium Single-cell 3’ V2 Chemistry Library Kit, Gel Bead & Multiplex Kit and Chip Kit (10x Genomics, the Netherlands) aiming for 5000 cells per library. Barcoded libraries were sequenced on an Illumina (Illumina, USA) HiSeq4000 in 25-8-98 paired-end configuration.
Single-cell analysis
Raw sequencing data were demultiplexed to generate fastq-files, which were aligned to the mouse reference genome build mm10, using STAR, which is implemented in Cell Ranger (version 2.0, 10x Genomics, USA). A single-cell expression matrix was generated by Cell Ranger. Further calculations were performed in R (version 3.6.0; https://cran.r-project.org/bin/windows/base/old/3.6.0/) using Seurat (version 2.2; https://github.com/satijalab/seurat). The gene expression matrices of both samples were further filtered, and we retained the cells with >200 genes and <8000 genes; >400 unique molecular identifiers (UMIs) and <20% mitochondrial RNA (mtRNA). These criteria resulted in a total set of 9474 cells, of which 5738 were Akimba and 3736 wild-type. We detected 2797 and 2677 variable genes in the wild-type and the Akimba sample, respectively. The union of these genes was used to perform canonical correlation (CC) analysis. We retained 20 CCs based on the MetageneBicorPlot. We visualised our data using t-Distributed Stochastic Neighbor Embedding (t-SNE) technique and clustered on a resolution of 1.4. The model-based analysis of single-cell transcriptomics (MAST) algorithm was used to identify differentially expressed genes between clusters. Clusters were annotated based on cell-specific markers (electronic supplementary material [ESM] Table 1). Immune cells and macroglia were further clustered using the same approach: 22 CCs based on 2777 and 2808 variable genes from wild-type and Akimba mice, respectively, and a resolution of 1.25 for the immune cells; 22 CCs based on 2710 (wild-type) and 2728 (Akimba) variable genes at a resolution of 0.5 for macroglia. Differentially expressed genes between wild-type and Akimba cells were first selected based on p value (adjusted p < 0.05) and next sorted according to log2 fold change. Contaminating rod-specific genes [11, 12] were filtered out where necessary. Analogous, artefactual genes, as described by van den Brink et al [13], were excluded from analysis in cell-type-specific datasets. The top 100 differentially expressed genes were analysed for statistically enriched pathways (Gene Ontology [GO], Kyoto Encyclopedia of Genes and Genomes [KEGG] and Reactome) and functional association networks were constructed and annotated using STRING version 11.0 (https://string-db.org/). The Mus musculus genome was used as background genome. Unsupervised clustering based on k-means was used to build related gene networks.
Immunohistochemistry and histological analysis
Paraffin sections were incubated overnight with primary antibodies for S100 (rabbit monoclonal, 1/200, 52642, Abcam, UK), haem oxygenase-1 (HO-1, rabbit polyclonal, 1/100, ADI-SPA-896, Enzo Life Sciences, Belgium), glial fibrillary acidic protein (GFAP, mouse monoclonal, 1/400, G3893, Sigma-Aldrich, Belgium) and metallothionein (MT, mouse monoclonal, 1/100, ab12228, Abcam, UK), all diluted in TNB blocking buffer (0.1 mmol/l Tris-HCl, pH 7.5, 0.15 mmol/l NaCl, 0.5% blocking reagent). S100 was visualised using an Alexa Fluor-conjugated secondary antibody (1/200, A-11008, ThermoFisher Scientific, Belgium). GFAP and MT were labelled via MOM biotinylated anti-mouse IgG (VEC.BMK-2202, Labconsult, Belgium) and HO-1 via biotin-conjugated secondary antibodies (1/300, 111-065-144, Jackson ImmunoResearch, UK). Secondary antibodies were all diluted in TNB blocking buffer. Next, the signal was developed by the TSA Cy3 system (NEL704A001KT, Perkin-Elmer, Belgium). All antibodies were first validated on samples from diabetic animal models.
Representative high-resolution images were made with an Olympus FV1000 confocal microscope (Belgium). Morphometric analyses on images of the entire neuroretina (taken using a Zeiss Axiocam MrC5 microscope) were performed using Zen 2012 software (Zeiss, Germany). The immunopositive area of S100, HO-1, GFAP and MT was analysed on six sections (three sections anterior and posterior of the optic nerve) and normalised to the total retinal area. These measurements were averaged for each individual eye.
Whole-mount retinas were incubated overnight with biotinylated isolectin-B4 (Bandeiraea simplicifolia, 1/25, L-2140, Sigma-Aldrich, Belgium), followed by incubation with a streptavidin-conjugated Alexa Fluor secondary antibody (1/100, S32355, ThermoFisher Scientific, Belgium). Mosaic pictures of entire retinal whole-mounts were made using a Zeiss Axiocam MrC5 microscope and analysis of the isolectin-B4 positive area was performed using Metamorph software 2.2.0 (Leica MM AF Offline Analysis, Leica, Germany) and normalised to the total retinal area. Statistically significant differences were determined using a two-tailed unpaired Student’s t test via GraphPad Prism 8 software (USA).
ELISA and western blotting
Pooled retinas from both eyes were homogenised in lysis buffer (1% NP-40, 20 mmol/l Tris, 137 mmol/l NaCl, 10% glycerol, 1 mmol/l EDTA, 1 mmol/l activated sodium orthovanadate, 10 μg/ml leupeptin, 10 μg/ml aprotinin, complete protease inhibitor). Protein concentrations were determined using bicinchoninic acid (BCA) protein assay (23225, ThermoFisher Scientific, Belgium). For ELISA, samples were run on the glyceraldehyde 3-phosphate dehydrogenase (GAPDH) ELISA (ab176642, Abcam, UK) following the manufacturer’s instructions. For western blotting, samples were run on 4–12% gradient gel in MES SDS buffer and blotted using the iBlot system (ThermoFisher Scientific). After blocking, membranes were probed with anti-MT (1/1000, Abcam, ab12228) or anti-β-tubulin (1/500, ab6046, Abcam, UK) antibodies, followed by secondary antibody (1/2000, 170-6515 & 170-6516, Bio-Rad, Belgium) diluted in 5% milk in TBST (0.1% Tween 20 in 19.8 mmol/l Tris base, 150.7 mmol/l NaCl, pH 7.6). Detections were performed on an Azure Biosystems device and data were analysed in ImageJ. ELISA and western blot antibodies were first validated on Kimba samples. Statistically significant differences were determined using a two-tailed unpaired Student’s t test via GraphPad Prism 8 software (USA).
Results
Single-cell RNA sequencing analysis yields 15 clusters corresponding to eight retinal cell types
Single-cell transcriptome analysis was performed on retinal tissue of 12-week-old wild-type and Akimba mice. Characteristic features of diabetic retinopathy were confirmed in Akimba mice at this age (Fig. 1a-f). The analysis pipeline of this study is depicted in Fig. 1g. A total of 9474 cells were analysed, with on average 1280 genes and 3068 unique molecular identifiers detected per cell. Of these, 5738 cells (61%) originated from Akimba retinas and 3736 cells (39%) from wild-type retinas. Based on expression of the most variable genes, 15 cell clusters were generated (Fig. 2a-e), which were annotated as follows (Fig. 2f, ESM Table 1): rod photoreceptors (clusters 0, 1, 3, 13), cone photoreceptors (cluster 9), bipolar cells (clusters 4 and 5; containing, respectively, mainly cone and rod subtypes), amacrine cells (clusters 7 and 12; containing, respectively, mainly GABAergic and glycinergic subtypes), macroglia (Müller glia and astrocytes; clusters 2, 10, 14), immune cells (cluster 6), endothelial cells (cluster 11), and a composite cluster containing pericytes and fibroblasts (cluster 8). Müller glia and astrocytes could not be resolved as separate clusters, probably due to the low relative number of astrocytes in the total retinal cell population. The abundance of wild-type vs Akimba cells per cluster is depicted in Fig. 2c-d, the relative abundance of cell types for each genotype is depicted in Fig. 2e. Clusters corresponding to horizontal and retinal ganglion cells were not detected in this study. For rod, cone, bipolar, macroglial and immune cells, the data allowed further differential gene expression and/or subclustering analysis, as we report below.
Characterisation of Akimba and wild-type mice used in this study, and schematic overview of the experimental workflow. (a) Akimba mouse retinas presented severe retinal vascular pathology including microaneurysm-like leaky spots and tortuous vessels, as demonstrated on this representative fluorescein angiogram. (d) Wild-type mice did not manifest retinal leakage. (b) A corresponding optical coherence tomography scan demonstrates reduction in retinal thickness and intraretinal oedema (arrow) in Akimba eyes; (e) a normal, well-organised retinal morphology is seen in wild-type retina. (c, f) Representative haematoxylin and eosin-stained retinal cross-section of 12-week-old Akimba retina (c) and age-matched wild-type retina (f). Outer retinal layers (OPL, ONL, PRL) are highly disorganised or absent in Akimba eyes, as a result of photoreceptor degeneration. Scale bars (b, e), 600 μm; scale bars (bʹ, eʹ), 100 μm; scale bars (c, f), 20 μm. (g) Schematic overview of the experimental workflow that was followed in this study. RNFL/GCL, retinal nerve fibre layer/ganglion cell layer; IPL, inner plexiform layer; INL, inner nuclear layer; OPL, outer plexiform layer; ONL, outer nuclear layer; PRL, photoreceptor layer
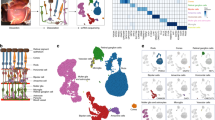
scRNAseq analysis of Akimba vs wild-type mouse retinas. (a) t-SNE plot of 9474 cells isolated from Akimba and wild-type retinas, showing 15 clusters that were identified. (b) t-SNE plot of 9474 cells isolated from the Akimba and wild-type retinas, showing cell clusters grouped together per cell type. P/F: pericytes/fibroblasts. (c) t-SNE plot of 9474 cells isolated from the Akimba and wild-type retinas, colour-coded according to genotype. (d) Number of Akimba (orange) and wild-type (blue) retinal cells per cluster. (e) Relative abundances of the different retinal cell types per genotype. Colour codes correspond to (b). Of note, rod photoreceptors were reduced in the Akimba retina, resulting in a higher proportion of other retinal cell types. (f) Violin plots showing the expression of selected cell-type-specific markers for all clusters. Expression data for all key marker genes for cluster annotation are available in ESM Table 1
Differential gene expression analysis in Akimba vs wild-type retinal neurons
Differential gene expression analysis was performed on rod photoreceptors (ESM Fig. 1, ESM Table 2, ESM Table 3), cone photoreceptors (ESM Fig. 2, ESM Table 4 and ESM Table 5) and bipolar cells (ESM Fig. 3, ESM Table 6 and ESM Table 7). Functional gene networks and pathways related to oxidative stress response and inflammation (antigen processing and presentation and/or interferon response) were found to be commonly upregulated in Akimba rods, cones and bipolar cells. Increase of oxidative phosphorylation (OXPHOS) transcripts was identified in cones and bipolar cells. In addition, increase of transcripts related to ATPase-mediated proton transport was observed in cones and pathways related to neuronal development in rods and bipolar cells. Functional networks that were commonly downregulated in Akimba rods, cones and bipolar cells were phototransduction, visual perception, ribosomes and glycolysis.
Differential gene expression analysis in Akimba vs wild-type macroglia
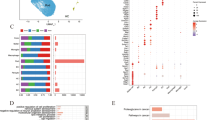
Based on differentially expressed genes, the following functional networks were found to be upregulated in Akimba macroglia (Fig. 3a, ESM Table 8): ribosome, cytoskeleton, immune system processes (antigen processing and presentation, interferon response), S100 proteins, glutathione metabolism, iron ion homeostasis, cell cycle regulation/apoptosis and OXPHOS. Notably, we observed several markers of reactive gliosis, including Gfap, Lcn2, Serpina3n, Serping1 and C4b [14]. Downregulated genes in Akimba macroglia associated into glycolysis, central nervous system (CNS) development and OXPHOS (Fig. 3b, ESM Table 9). Although OXPHOS-related transcripts were observed in both datasets, data indicating upregulation of these transcripts in Akimba mice was most prominent.
Subcluster analysis of macroglia
Next, we evaluated whether the macroglial cells in our dataset could be subdivided in subpopulations hallmarked by different activation states/biological pathways and identified four subclusters (0, 1, 2 and 4) with a macroglial transcriptional profile (Fig. 4). Analysis of the gene signature of these subclusters led us to identify different macroglia phenotypes characterised by transcripts related to cytoskeleton organisation, Rho GTPase signalling and S100 proteins (subcluster 0); antigen presentation (class I MHC), immune system, and endoplasmic reticulum (subcluster 1); metabolism and ribosomal proteins (subcluster 2) and extracellular matrix (ECM) deposition, collagens, (ion) transporters, glutathione metabolism and specific growth factor signalling pathways such as IGF binding proteins (IGFBPs) and TGFβ/bone morphogenetic protein (BMP) signalling (subcluster 4) (Table 1, ESM Table 10). Of note, in addition to macroglia, subcluster four was found to contain a small percentage of cells expressing retinal pigment epithelium (RPE) markers.
Subcluster analysis of immune cells
Differential gene expression analysis of the global Akimba retina vs wild-type retina was also evaluated. Although this analysis compares different cell populations, for which relative abundances differ per genotype (Fig. 2e), results indicated an increase in inflammatory pathways including antigen processing and presentation, complement and a heterogenous network comprising immune system and regulation of endocytosis in the Akimba retina (ESM Fig. 4, ESM Table 11 and ESM Table 12). In agreement with this, 196 out of 219 immune cells in our dataset were derived from Akimba retina. To obtain a better view on their diversity, immune cells were further subclustered. Based on the genes with greatest specificity for each subcluster (ESM Table 1), we identified three high quality immune cell subpopulations (Fig. 5): (1) Ccr2low non-inflammatory monocytes, (2) resting and activated microglia and (3) inflammatory monocyte-derived macrophages. The latter population was exclusively present in Akimba retinas.
Subclustering and annotation of immune cells. (a) Heat map of normalised expression of the top 10 genes with greatest specificity for each cell subcluster. (b) t-SNE plot depicting three immune cell subclusters. (c) Pie charts showing the proportion (absolute cell number and percentage) of the different immune cell populations in the wild-type and Akimba retina. Subclusters 3 and 4 could not be confidently annotated as immune cells and were excluded from further analysis. Colour codes are the same as in (b)
Protein markers of metabolic switch, gliosis, inflammation, metal ion and oxidative stress response confirm single-cell transcriptome findings
To confirm the observations from the scRNAseq analysis, expression of several key markers was verified at the protein level. Immunohistochemistry confirmed an increase of S100 proteins and GFAP in the 12-week-old Akimba retinas, localised in the macroglia (Fig. 6a–f). Second, confirming activation of immune responses, we observed an increase of the isolectin-B4+ signal (i.e. microglia activation and inflammatory cell influx) in the Akimba retina (Fig. 6g–i). Third, both in the global and cell-type-specific differential gene expression analyses we found an increase in transcripts related to metal ion and oxidative stress response. Therefore, we evaluated MT and HO-1 expression via western blot and immunohistochemistry. A significant increase of these proteins was observed in Akimba retinas, confirming the upregulation of these cellular processes, with MT and HO-1 localising mainly to neurons in the outer and inner retina (Fig. 7a–f). Finally, the apparent metabolic switch from glycolysis to OXPHOS was evaluated with an ELISA for the glycolytic enzyme GAPDH on retinal lysates of 6-, 8-, 10- and 12-week-old wild-type and Akimba mice. Our results confirmed a significant reduction of retinal GAPDH protein levels in Akimba mice at all time points evaluated, with no significant changes in the Akimba retina over this 6-week time frame (Fig. 7g).
Histology analysis supports reactive macroglia and immune responses in the Akimba retina. (a–c) GFAP immunostaining on wild-type (a) and Akimba (b) retinal sections from animals at 12 weeks of age, and morphometric quantification of GFAP immunolabelling (c; n = 9 wild-type, n = 6 Akimba). (d–f) Representative images of S100 immunostaining on retinal sections of 12-week-old wild-type (d) and Akimba (e) mice, and morphometric quantification of S100 immunolabelling (f; n = 4). (g–i) Isolectin-B4 (IB-4) staining on retinal whole-mounts of 12-week-old wild-type (g) and Akimba (h) mice, and quantification of IB-4+ area as a measure for activation/infiltration of macrophages and microglia (i; n = 7 wild-type, n = 17 Akimba). Scale bars, 50 μm. Data are shown as mean ± SEM. RNFL/GCL, retinal nerve fibre layer/ganglion cell layer; IPL, inner plexiform layer; INL, inner nuclear layer; OPL, outer plexiform layer; ONL, outer nuclear layer; PRL, photoreceptor layer
Protein analysis supports metal ion and redox dyshomeostasis and metabolic shift in the Akimba retina. (a–c) Immunostaining for the oxidative stress marker HO-1 on wild-type (a) and Akimba (b) retinal sections from animals at 12 weeks of age, and morphometric quantification of HO-1 immunolabelling (c; n = 9 wild-type, n = 6 Akimba). Scale bar, 50 μm. (d, e) Representative images (n = 5) of MT immunostaining on retinal sections of 12-week-old wild-type and Akimba mice. (f) Western blot for MT protein on retinal lysates of 12-week-old wild-type and Akimba mice (n = 5); relative expression was normalised to β-tubulin (TUBB). (g) Protein levels of the glycolytic enzyme GAPDH, measured via ELISA, in retinal lysates of 6- to 12-week-old wild-type and Akimba mice (n = 10). Scale bars, 50 μm. Data are shown as mean ± SEM. RNFL/GCL, retinal nerve fibre layer/ganglion cell layer; IPL, inner plexiform layer; INL, inner nuclear layer; OPL, outer plexiform layer; ONL, outer nuclear layer; PRL, photoreceptor layer
Discussion
Our results confirm that at 12 weeks of age, the Akimba retina is dysfunctional, with loss of rod photoreceptors and gliosis, as well as influx and activation of immune cells. In addition, we observe alterations of metabolism, ribosomal proteins, metal ion and redox homeostasis, and cytoskeletal components. Our findings are in line with previous bulk transcriptomics studies in streptozotocin (STZ)-induced diabetic retinopathy models [15,16,17]. In addition, enrichment of immune processes such as interferon signalling and antigen presentation have also been observed in models of retinal degeneration [15,16,17]. Our observations identify cell-type-specific changes and shed more light on subtypes of inflammatory and macroglial cells that are present in this diabetic retinopathy model. Below we discuss prominent observations in our dataset.
Reactive gliosis in Akimba macroglia
As a universal response to CNS injury, macroglia (Müller cells and astrocytes) undergo morphological and functional alterations that modulate, among others, neuron and synapse function, endothelial barrier integrity, and ECM and immune cell function. In line with published reports, our data in Akimba macroglia confirm upregulation of reactive gliosis markers associated with cytoskeleton, cell cycle, peptidase regulation and S100 proteins [14, 18, 19]. Upregulation of these functional networks has also been observed in other animal models of diabetic retinopathy and in human diabetic retina [20,21,22,23]. For S100 proteins and GFAP, results were confirmed on the protein level, and correspond to previous findings in Akimba retinas [9, 24]. Specifically for S100 proteins, upregulation has been reported in the retina of rodent diabetic retinopathy models, including Akimba mice, and in the vitreous and plasma of diabetic retinopathy patients, where it was found to be a marker for proliferative diabetic retinopathy [25,26,27]. Their pleiotropic role links S100 proteins to inflammation, angiogenesis and fibrosis—3 key processes in diabetic retinopathy. Furthermore, our findings underscore that macroglia contribute to the inflammatory response via antigen processing and presentation as well as interferon signalling, which enhance attraction/activation of immune cells [19, 28]. Together, these transcriptional changes may compromise Müller glia support functions and may contribute to the imbalanced retinal ion and water homeostasis, oedema, production of reactive oxygen species, proinflammatory molecules and neurodegeneration that have been observed in the Akimba retina.
Maladaptive metabolic changes in Akimba retina
Analysis of the transcriptional changes in the Akimba retina as a whole, as well as in individual neuronal and macroglial cell types, indicates a reduction of transcripts related to glycolysis and concomitant increase of transcripts related to OXPHOS. On the one hand, photoreceptors mainly take up glucose from the choroidal vasculature and metabolise it via aerobic glycolysis [29]. Given the high metabolic demand of phototransduction, the downregulation of glycolysis genes observed in rods and cones in this study may reflect the intimate connection between metabolism and function/survival in these cells and the loss of functional photoreceptors. Macroglia, on the other hand, mediate neuron–glia metabolic coupling. According to the lactate shuttle hypothesis, astrocytes take up glucose, metabolise it to lactate via glycolysis and transport this lactate to neighbouring neurons, where it serves as the primary metabolite [30, 31]. Given that in astrocytes, insulin signalling is required for efficient glucose uptake into the brain [32], it is likely that intracellular glucose availability is reduced in Akimba macroglia. Sustained glucose deprivation has been described to induce a metabolic shift in Müller cells from glycolysis to OXPHOS [33]. As a consequence, less lactate can be shuttled to neurons, leading to increased oxidative stress and energy depletion [30]. Given the intricate relationship of inner retinal neurons with Müller glia, this hypothesis may explain the loss of retinal ganglion cells and inner nuclear layer thinning reported in Akimba retinas [8]. Notably, western blotting confirmed that hyperglycaemia leads to decreased levels of the glycolysis checkpoint protein GAPDH, which may in turn result in the diversion of upstream glycolytic intermediates to alternative pathways, Müller glia dysfunction and cell death [34]. Accordingly, GAPDH was found to be reduced in the retina of diabetic rodents, and remains compromised even after glycaemic control is reinstituted, suggesting that it plays a key role in diabetic retinopathy progression [35,36,37].
Differential activation states illustrate macroglial diversity in the retina
In line with reported astrocyte and Müller glia heterogeneity [14, 18, 19, 38, 39], subcluster analysis of macroglia suggests the presence of distinct macroglial subpopulations. One subpopulation was characterised by immune processes such as antigen presentation (class I MHC). Elevated MHC transcripts have been reported in retinal Müller cells from STZ-induced diabetic rats [40]. A second macroglial subtype displayed a profibrotic profile related to ECM deposition, collagen synthesis and processing, cell-ECM interactions as well as growth factor signalling pathways such as IGFBPs and BMPs. BMPs have been described as important regulators of macroglia differentiation and reactive gliosis in retinal macroglia [41]. IGFBPs on the other hand have been demonstrated to be expressed by specific Müller cell phenotypes relating to fibrotic retinal disease in vitro [42]. It was interesting to observe that in this subcluster, macroglial cells clustered together with a small percentage of cells carrying RPE markers, another cell type which has been reported to undergo transition into a fibrotic phenotype and to be involved in retinal fibrotic processes. Taken together, these data are in line with prior reports identifying Müller cells in diabetic epiretinal membranes and describing their ability to transdifferentiate into myofibroblast-like cells associated with proliferative diabetic retinopathy. The third and fourth subpopulations were characterised by cytoskeleton organisation, S100 proteins and metabolic/ribosomal proteins respectively, connecting to our findings on gliosis and metabolism in Akimba macroglia.
Inflammatory processes in the Akimba retina
A recurring observation in this study, supported by the differential gene expression analysis of the global retina and macroglia, subclustering of inflammatory cells, and confirmatory staining for isolectin-B4, is the increase of immune system-related pathways in the Akimba retina. Our data indicate a prominent increase and activation of resident and invading immune cells in the Akimba retina, as previously observed in Akimba mice [8, 9], animal models of photoreceptor degeneration [43,44,45] and human diabetic retinopathy [46]. Microglia, macrophage, leucocyte and (although we did not detect these cells as an individual immune subcluster) lymphocyte activation, as well as monocyte and leucocyte differentiation, were identified as significantly enriched pathways, and are believed to contribute to diabetic retinopathy progression [47]. Inversely, the prominent photoreceptor cell death in the Akimba retina might also trigger microglia/macrophage migration in order to remove debris and dying cells. We found that Akimba retinas exhibit features of advanced diabetic retinopathy, with high expression of complement genes. Dysregulation of the complement system has previously been described in diabetic retinopathy patients and animal models [48, 49]. Furthermore, a role for the NLR family pyrin domain containing 3 (NLRP3) inflammasome during pathological neovascularisation in the Akimba mouse model was previously reported. Activation/upregulation of this inflammatory complex was only recently described in the vitreous humour of diabetic retinopathy patients [50, 51]. Although the NLRP3 inflammasome was not identified in the top 100 genes of our differential expression analysis, NLRP3 inflammasome-related genes, including Casp1, Pycard, Il1b and Il18, were significantly upregulated in Akimba retinas, confirming the findings by Chaurasia et al [9].
Metal ion and redox dyshomeostasis in the Akimba retina
Metal ions such as iron, copper and zinc, play vital roles in the structure and function of proteins involved in metabolism, homeostasis and oxidative stress response pathways [52]. In our dataset, transport, redox homeostasis (such as glutathione) and metal ion response pathways were upregulated in macroglia and all cell differential expression analyses, which was further confirmed on the protein level for the metal storage, transport and detoxification protein MT. Abnormal iron accumulation, increased levels of transferrin and ferritin and decreased zinc levels have been observed in the vitreous humour and/or retina of diabetic retinopathy, age-related macular degeneration, glaucoma, retinal vein occlusion and aged patients [52,53,54,55] and are believed to predispose the retina to oxidative stress [52, 53]. Furthermore, it has been described that iron toxicity activates the NLRP3 inflammasome [56]. Although reducing unbound free metal ions and serving antioxidant functions [52], our data suggest that this upregulation is insufficient to restore homeostasis. Indeed, we found oxidative stress response gene networks (e.g. Gpx1, Gpx3, Gpx4, Prdx1 and Prdx6) to be highly upregulated in Akimba total retina, cones and macroglia. These transcriptional observations were supported by immunostaining indicating increased levels of the oxidative stress marker HO-1 in Akimba vs wild-type retinas. Interestingly, ‘ferroptosis’ was found to be an enriched cellular pathway in several clusters. This form of regulated cell death is triggered by cellular iron overload and generation of lipid peroxidation products and has been associated with neurodegenerative pathologies such as age-related macular degeneration, Alzheimer’s disease and Parkinson’s disease [57]. Indeed, several pathway analyses in this study pointed out commonalities with these neurodegenerative diseases.
Limitations of the study
Despite the Akimba mouse being a translational model of diabetic retinopathy, it has certain limitations: Akimba mice lack pre-retinal neovascularisation and, whereas 12-week-old Akimba mice display mainly photoreceptor loss, diabetic retinopathy patients additionally display loss of retinal ganglion cells and amacrine cells [58]. In agreement with clinical diabetic retinopathy, the combination of hyperglycaemia and elevated VEGF levels drives progression into proliferative diabetic retinopathy. In Akimba mice, transient overexpression of human VEGF165 is localised in photoreceptors, but augmented endogenous mouse VEGF levels have also been observed in Akimba mice [8, 9, 59, 60]. Importantly, we confirmed that 12-week-old Akimba mice display features of diabetic retinopathy such as microaneurysms, vascular leakage, neurodegeneration and oedema. In addition, our choice of this age was based on the fact that in vivo efficacy of therapeutic intervention has been shown in 12-week-old Akimba mice [10, 61 and Hu TT, De Vriese A, Vanheukelom V, Feyen J (unpublished results)]. In terms of analysis, it was not possible to evaluate variability between individuals as retinal cells from individual mice have been pooled in this study. In addition, our approach did not have the resolution to separate clusters of Müller cells and astrocytes and could not identify horizontal or retinal ganglion cell clusters. Some resolution may have been lost due to the presence of rod-specific transcripts in all clusters, a widely acknowledged issue with retinal scRNAseq [11, 12]. We were unable to filter out this contamination in the earliest stages of analysis, but took great care in excluding these transcripts in downstream analyses. Nevertheless, using this approach it cannot be excluded that less known or less specific rod transcripts are retained in the analysis or vice versa.
Conclusion
Taken together, using single-cell RNA sequencing, we obtained a more in-depth view of the transcriptional signatures and diversity of neuronal, glial and inflammatory cell populations in the Akimba model of diabetic retinopathy. We observed: (1) shifts in retinal cell population abundance, most prominently rod photoreceptors and specific inflammatory cell populations; (2) metabolic and functional alterations of macroglia, which may compromise their support functions; (3) activation of immune system pathways, redox and metal ion dyshomeostasis; and (4) heterogeneity in macroglia phenotypes, including distinct profibrotic and proinflammatory populations. Our findings support the emerging idea that macroglia play a key role in retinal neurodegenerative diseases. The findings presented in this manuscript should be confirmed in additional early/intermediate stage diabetic retinopathy models and, to confirm their relevance in humans, in patient samples. Additionally, differentially expressed pathways need to be evaluated in functional experiments to evaluate whether they play a detrimental or protective role in diabetic retinopathy.
Data availability
RNA-seq data have been deposited in the ArrayExpress database at EMBL-EBI (www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-9061.
Abbreviations
- BMP:
-
Bone morphogenetic protein
- CC:
-
Canonical correlation
- CNS:
-
Central nervous system
- ECM:
-
Extracellular matrix
- GAPDH:
-
Glyceraldehyde 3-phosphate dehydrogenase
- GFAP:
-
Glial fibrillary acidic protein
- GO:
-
Gene Ontology
- HO-1:
-
Haem oxygenase-1
- IGFBP:
-
IGF binding proteins
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- MT:
-
Metallothionein
- OXPHOS:
-
Oxidative phosphorylation
- RPE:
-
Retinal pigment epithelium
- scRNAseq:
-
Single-cell RNA sequencing
- STZ:
-
Streptozotocin
- t-SNE:
-
t-Distributed Stochastic Neighbor Embedding
- VEGF:
-
Vascular endothelial growth factor
References
Antonetti DA, Klein R, Gardner TW (2012) Diabetic retinopathy. N Engl J Med 366(13):1227–1239. https://doi.org/10.1056/NEJMra1005073
Stitt AW, Curtis TM, Chen M et al (2016) The progress in understanding and treatment of diabetic retinopathy. Prog Retin Eye Res 51:156–186. https://doi.org/10.1016/j.preteyeres.2015.08.001
Sergeys J, Etienne I, Van Hove I et al (2019) Longitudinal in vivo characterization of the streptozotocin-induced diabetic mouse model: focus on early inner retinal responses. Invest Ophthalmol Vis Sci 60(2):807–822. https://doi.org/10.1167/iovs.18-25372
Lynch SK, Abramoff MD (2017) Diabetic retinopathy is a neurodegenerative disorder. Vis Res 139:101–107. https://doi.org/10.1016/j.visres.2017.03.003
Barber AJ, Baccouche B (2017) Neurodegeneration in diabetic retinopathy: Potential for novel therapies. Vis Res 139:82–92. https://doi.org/10.1016/j.visres.2017.06.014
Rubsam A, Parikh S, Fort PE (2018) Role of inflammation in diabetic retinopathy. Int J Mol Sci 19(4). https://doi.org/10.3390/ijms19040942
Tang J, Kern TS (2011) Inflammation in diabetic retinopathy. Prog Retin Eye Res 30(5):343–358. https://doi.org/10.1016/j.preteyeres.2011.05.002
Rakoczy EP, Ali Rahman IS, Binz N et al (2010) Characterization of a mouse model of hyperglycemia and retinal neovascularization. Am J Pathol 177(5):2659–2670. https://doi.org/10.2353/ajpath.2010.090883
Chaurasia SS, Lim RR, Parikh BH et al (2018) The NLRP3 inflammasome may contribute to pathologic neovascularization in the advanced stages of diabetic retinopathy. Sci Rep 8(1):2847. https://doi.org/10.1038/s41598-018-21198-z
Van Bergen T, Hu TT, Etienne I, Reyns GE, Moons L, Feyen JHM (2017) Neutralization of placental growth factor as a novel treatment option in diabetic retinopathy. Exp Eye Res 165:136–150. https://doi.org/10.1016/j.exer.2017.09.012
Macosko EZ, Basu A, Satija R et al (2015) Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161(5):1202–1214. https://doi.org/10.1016/j.cell.2015.05.002
Siegert S, Cabuy E, Scherf BG et al (2012) Transcriptional code and disease map for adult retinal cell types. Nat Neurosci 15(3):487–495. https://doi.org/10.1038/nn.3032
van den Brink SC, Sage F, Vertesy A et al (2017) Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat Methods 14(10):935–936. https://doi.org/10.1038/nmeth.4437
Zamanian JL, Xu L, Foo LC et al (2012) Genomic analysis of reactive astrogliosis. J Neurosci 32(18):6391–6410. https://doi.org/10.1523/jneurosci.6221-11.2012
Liu YJ, Lian ZY, Liu G, Zhou HY, Yang HJ (2016) RNA sequencing reveals retinal transcriptome changes in STZ-induced diabetic rats. Mol Med Rep 13(3):2101–2109. https://doi.org/10.3892/mmr.2016.4793
Kandpal RP, Rajasimha HK, Brooks MJ et al (2012) Transcriptome analysis using next generation sequencing reveals molecular signatures of diabetic retinopathy and efficacy of candidate drugs. Mol Vis 18:1123–1146
Freeman WM, Bixler GV, Brucklacher RM et al (2009) Transcriptomic comparison of the retina in two mouse models of diabetes. J Ocul Biol Dis Inform 2(4):202–213. https://doi.org/10.1007/s12177-009-9045-3
Roesch K, Jadhav AP, Trimarchi JM et al (2008) The transcriptome of retinal Müller glial cells. J Comp Neurol 509(2):225–238. https://doi.org/10.1002/cne.21730
Vecino E, Rodriguez FD, Ruzafa N, Pereiro X, Sharma SC (2016) Glia-neuron interactions in the mammalian retina. Prog Retin Eye Res 51:1–40. https://doi.org/10.1016/j.preteyeres.2015.06.003
Eastlake K, Heywood WE, Banerjee P et al (2018) Comparative proteomic analysis of normal and gliotic PVR retina and contribution of Müller glia to this profile. Exp Eye Res 177:197–207. https://doi.org/10.1016/j.exer.2018.08.016
Grosche A, Hauser A, Lepper MF et al (2016) The proteome of native adult müller glial cells from murine retina. Mol Cell Proteomics 15(2):462. https://doi.org/10.1074/mcp.M115.052183
Gu L, Xu H, Zhang C, Yang Q, Zhang L, Zhang J (2019) Time-dependent changes in hypoxia- and gliosis-related factors in experimental diabetic retinopathy. Eye 33(4):600–609. https://doi.org/10.1038/s41433-018-0268-z
Mizutani M, Gerhardinger C, Lorenzi M (1998) Muller cell changes in human diabetic retinopathy. Diabetes 47(3):445–449. https://doi.org/10.2337/diabetes.47.3.445
Danesh-Meyer HV, Zhang J, Acosta ML, Rupenthal ID, Green CR (2016) Connexin43 in retinal injury and disease. Prog Retin Eye Res 51:41–68. https://doi.org/10.1016/j.preteyeres.2015.09.004
Abu El-Asrar AM, Nawaz MI, De Hertogh G et al (2014) S100A4 is upregulated in proliferative diabetic retinopathy and correlates with markers of angiogenesis and fibrogenesis. Mol Vis 20:1209–1224
Dong N, Shi H, Xu B, Cai Y (2015) Increased plasma S100A12 levels are associated with diabetic retinopathy and prognostic biomarkers of macrovascular events in type 2 diabetic patients. Invest Ophthalmol Vis Sci 56(8):4177–4185. https://doi.org/10.1167/iovs.15-16470
Lim RR, Barathi VA, Mohan RR, Ghosh A, Chaurasia SS (2017) Expression of S100 proteins in Diabetic Retinopathy. Invest Ophthalmol Vis Sci 58(8):5222
Bombeiro AL, Hell RC, Simoes GF, Castro MV, Oliveira AL (2017) Importance of major histocompatibility complex of class I (MHC-I) expression for astroglial reactivity and stability of neural circuits in vitro. Neurosci Lett 647:97–103. https://doi.org/10.1016/j.neulet.2017.03.038
Narayan DS, Chidlow G, Wood JPM, Casson RJ (2017) Glucose metabolism in mammalian photoreceptor inner and outer segments. Clin Exp Ophthalmol 45(7):730–741. https://doi.org/10.1111/ceo.12952
Toft-Kehler AK, Skytt DM, Svare A et al (2017) Mitochondrial function in Muller cells - Does it matter? Mitochondrion 36:43–51. https://doi.org/10.1016/j.mito.2017.02.002
Vohra R, Kolko M (2018) Neuroprotection of the inner retina: Müller cells and lactate. Neural Regen Res 13(10):1741–1742. https://doi.org/10.4103/1673-5374.238612
Garcia-Caceres C, Quarta C, Varela L et al (2016) Astrocytic insulin signaling couples brain glucose uptake with nutrient availability. Cell 166(4):867–880. https://doi.org/10.1016/j.cell.2016.07.028
Toft-Kehler AK, Skytt DM, Kolko M (2018) A perspective on the muller cell-neuron metabolic partnership in the inner retina. Mol Neurobiol 55(6):5353–5361. https://doi.org/10.1007/s12035-017-0760-7
Jayaguru P, Mohr S (2011) Nuclear GAPDH: changing the fate of Müller cells in diabetes. J Ocul Biol Dis Inform 4(1–2):34–41. https://doi.org/10.1007/s12177-012-9085-y
Du X, Matsumura T, Edelstein D et al (2003) Inhibition of GAPDH activity by poly(ADP-ribose) polymerase activates three major pathways of hyperglycemic damage in endothelial cells. J Clin Invest 112(7):1049–1057. https://doi.org/10.1172/jci18127
Kanwar M, Kowluru RA (2009) Role of glyceraldehyde 3-phosphate dehydrogenase in the development and progression of diabetic retinopathy. Diabetes 58(1):227–234. https://doi.org/10.2337/db08-1025
Madsen-Bouterse S, Mohammad G, Kowluru RA (2010) Glyceraldehyde-3-phosphate dehydrogenase in retinal microvasculature: implications for the development and progression of diabetic retinopathy. Invest Ophthalmol Vis Sci 51(3):1765–1772. https://doi.org/10.1167/iovs.09-4171
Liddelow SA, Guttenplan KA, Clarke LE et al (2017) Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541:481. https://doi.org/10.1038/nature21029https://www.nature.com/articles/nature21029#supplementary-information
Pfeiffer RL, Marc RE, Kondo M, Terasaki H, Jones BW (2016) Muller cell metabolic chaos during retinal degeneration. Exp Eye Res 150:62–70. https://doi.org/10.1016/j.exer.2016.04.022
Gerhardinger C, Costa MB, Coulombe MC, Toth I, Hoehn T, Grosu P (2005) Expression of acute-phase response proteins in retinal Muller cells in diabetes. Invest Ophthalmol Vis Sci 46(1):349–357. https://doi.org/10.1167/iovs.04-0860
Dharmarajan S, Gurel Z, Wang S, Sorenson CM, Sheibani N, Belecky-Adams TL (2014) Bone morphogenetic protein 7 regulates reactive gliosis in retinal astrocytes and Muller glia. Mol Vis 20:1085–1108
King JL, Guidry C (2004) Muller cell production of insulin-like growth factor-binding proteins in vitro: modulation with phenotype and growth factor stimulation. Invest Ophthalmol Vis Sci 45(12):4535–4542. https://doi.org/10.1167/iovs.04-0447
Ronning KE, Karlen SJ, Miller EB, Burns ME (2019) Molecular profiling of resident and infiltrating mononuclear phagocytes during rapid adult retinal degeneration using single-cell RNA sequencing. Sci Rep 9(1):4858. https://doi.org/10.1038/s41598-019-41141-0
Sudharsan R, Beiting DP, Aguirre GD, Beltran WA (2017) Involvement of innate immune system in late stages of inherited photoreceptor degeneration. Sci Rep 7(1):17897. https://doi.org/10.1038/s41598-017-18236-7
Blank T, Goldmann T, Koch M et al (2017) Early microglia activation precedes photoreceptor degeneration in a mouse model of CNGB1-linked retinitis pigmentosa. Front Immunol 8:1930. https://doi.org/10.3389/fimmu.2017.01930
Zeng H-Y, Green WR, Tso MOM (2008) Microglial activation in human diabetic retinopathy. Arch Ophthalmol 126(2):227–232. https://doi.org/10.1001/archophthalmol.2007.65
Altmann C, Schmidt MHH (2018) The role of microglia in diabetic retinopathy: inflammation, microvasculature defects and neurodegeneration. Int J Mol Sci 19(1):110. https://doi.org/10.3390/ijms19010110
Clark SJ, Bishop PN (2018) The eye as a complement dysregulation hotspot. Semin Immunopathol 40(1):65–74. https://doi.org/10.1007/s00281-017-0649-6
Yanai R, Thanos A, Connor KM (2012) Complement involvement in neovascular ocular diseases. Adv Exp Med Biol 946:161–183. https://doi.org/10.1007/978-1-4614-0106-3_10
Loukovaara S, Piippo N, Kinnunen K, Hytti M, Kaarniranta K, Kauppinen A (2017) NLRP3 inflammasome activation is associated with proliferative diabetic retinopathy. Acta Ophthalmol 95(8):803–808. https://doi.org/10.1111/aos.13427
Chen H, Zhang X, Liao N et al (2018) Enhanced expression of NLRP3 inflammasome-related inflammation in diabetic retinopathy. Invest Ophthalmol Vis Sci 59(2):978–985. https://doi.org/10.1167/iovs.17-22816
Song D, Dunaief JL (2013) Retinal iron homeostasis in health and disease. Front Aging Neurosci 5:24. https://doi.org/10.3389/fnagi.2013.00024
Hahn P, Ying GS, Beard J, Dunaief JL (2006) Iron levels in human retina: sex difference and increase with age. Neuroreport 17(17):1803–1806. https://doi.org/10.1097/WNR.0b013e3280107776
Ciudin A, Hernández C, Simó R (2010) Iron overload in diabetic retinopathy: a cause or a consequence of impaired mechanisms? Exp Diabetes Res 2010:714108. https://doi.org/10.1155/2010/714108
Chaudhary K, Promsote W, Ananth S et al (2018) Iron overload accelerates the progression of diabetic retinopathy in association with increased retinal renin expression. Sci Rep 8(1):3025. https://doi.org/10.1038/s41598-018-21276-2
Gelfand BD, Wright CB, Kim Y et al (2015) Iron toxicity in the retina requires Alu RNA and the NLRP3 inflammasome. Cell Rep 11(11):1686–1693. https://doi.org/10.1016/j.celrep.2015.05.023
Doll S, Conrad M (2017) Iron and ferroptosis: A still ill-defined liaison. IUBMB Life 69(6):423–434. https://doi.org/10.1002/iub.1616
Simo R, Stitt AW, Gardner TW (2018) Neurodegeneration in diabetic retinopathy: does it really matter? Diabetologia 61(9):1902–1912. https://doi.org/10.1007/s00125-018-4692-1
Wisniewska-Kruk J, Klaassen I, Vogels IM et al (2014) Molecular analysis of blood-retinal barrier loss in the Akimba mouse, a model of advanced diabetic retinopathy. Exp Eye Res 122:123–131. https://doi.org/10.1016/j.exer.2014.03.005
Matsunaga N, Chikaraishi Y, Izuta H et al (2008) Role of soluble vascular endothelial growth factor receptor-1 in the vitreous in proliferative diabetic retinopathy. Ophthalmology 115(11):1916–1922. https://doi.org/10.1016/j.ophtha.2008.06.025
Hu TT, Vanheukelom V, De Vriese A, Feyen JHM (2016) Quantitative assessment of retinal permeability in the diabetic Akimba mouse: validation of a promising animal model for diabetic retinopathy. Acta Ophthalmologica 94(S256). https://doi.org/10.1111/j.1755-3768.2016.0550
Acknowledgements
The authors wish to thank A. De Vriese, V. Vanheukelom and H. Moreau from Oxurion NV, L. Noterdaeme from KU Leuven, and B. Tembuyser and T. Van Brussel from VIB Center for Cancer Biology for their technical support. Some of the data were presented as an abstract at the 2019 European Association for Vision and Eye Research and European Association for the Study of Diabetic Eye Complications meetings.
Authors’ relationships and activities
IVH, TTH, TVB, KB, IE and MP are current employees at Oxurion NV; JHMF serves as a consultant to Oxurion NV. The other authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work.
Funding
LDG is a postdoctoral fellow supported by the Flemish Research Foundation (FWO Vlaanderen). The computational resources used in this work were provided by the Flemish Supercomputer Center (VSC), funded by the Hercules Foundation and the Flemish Government, Department of Economy, Science and Innovation (EWI). The funding agencies were not involved in the design of the study; the collection, analysis, and interpretation of data; writing the report; or the decision to submit the report for publication.
Author information
Authors and Affiliations
Contributions
IVH designed the study, performed experiments, analysed data and wrote the manuscript. LDG performed experiments, analysed data and wrote the manuscript. TTH, TVB, KB, IE, BB and EM performed experiments, analysed data and revised the manuscript. DL, LM and JHMF designed the study and revised the manuscript. MP designed the study, analysed data and wrote the manuscript. All authors approve the final version of the manuscript. MP is responsible for the integrity of the work as a whole.
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM Table 1
(XLSX 22 kb)
ESM Table 2
(XLSX 92 kb)
ESM Table 3
(XLSX 58 kb)
ESM Table 4
(XLSX 61 kb)
ESM Table 5
(XLSX 54 kb)
ESM Table 6
(XLSX 76 kb)
ESM Table 7
(XLSX 62 kb)
ESM Table 8
(XLSX 70 kb)
ESM Table 9
(XLSX 87 kb)
ESM Table 10
(XLSX 108 kb)
ESM Table 11
(XLSX 95 kb)
ESM Table 12
(XLSX 56 kb)
ESM 13
(PPTX 30.2 mb)
Rights and permissions
About this article
Cite this article
Van Hove, I., De Groef, L., Boeckx, B. et al. Single-cell transcriptome analysis of the Akimba mouse retina reveals cell-type-specific insights into the pathobiology of diabetic retinopathy. Diabetologia 63, 2235–2248 (2020). https://doi.org/10.1007/s00125-020-05218-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-020-05218-0