Abstract
Aims/hypothesis
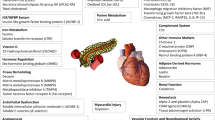
We selected the most informative protein biomarkers for the prediction of incident cardiovascular disease (CVD) in people with type 2 diabetes.
Methods
In this nested case–control study we measured 42 candidate CVD biomarkers in 1,123 incident CVD cases and 1,187 controls with type 2 diabetes selected from five European centres. Combinations of biomarkers were selected using cross-validated logistic regression models. Model prediction was assessed using the area under the receiver operating characteristic curve (AUROC).
Results
Sixteen biomarkers showed univariate associations with incident CVD. The most predictive subset selected by forward selection methods contained six biomarkers: N-terminal pro-B-type natriuretic peptide (OR 1.69 per 1 SD, 95% CI 1.47, 1.95), high-sensitivity troponin T (OR 1.29, 95% CI 1.11, 1.51), IL-6 (OR 1.13, 95% CI 1.02, 1.25), IL-15 (OR 1.15, 95% CI 1.01, 1.31), apolipoprotein C-III (OR 0.79, 95% CI 0.70, 0.88) and soluble receptor for AGE (OR 0.84, 95% CI 0.76, 0.94). The prediction of CVD beyond clinical covariates improved from an AUROC of 0.66 to 0.72 (AUROC for Framingham Risk Score covariates 0.59). In addition to the biomarkers, the most important clinical covariates for improving prediction beyond the Framingham covariates were estimated GFR, insulin therapy and HbA1c.
Conclusions/interpretation
We identified six protein biomarkers that in combination with clinical covariates improved the prediction of our model beyond the Framingham Score covariates. Biomarkers can contribute to improved prediction of CVD in diabetes but clinical data including measures of renal function and diabetes-specific factors not included in the Framingham Risk Score are also needed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cardiovascular disease (CVD) is a major cause of morbidity and mortality in people with diabetes. Prediction of CVD risk among people with diabetes is important not only to tailor clinical care but also to stratify higher risk patients in clinical trials to maximise trial power [1]. Biomarkers may help improve prediction, but whilst there are many reports on associations of single biomarkers with CVD risk in diabetes, there are few joint assessments of large numbers of biomarkers. Thus, there is little consensus on which subset of biomarkers in the literature is most informative for CVD risk prediction in diabetes.
The SUrrogate markers for Micro- and Macro-vascular hard endpoints for Innovative diabetes Tools (SUMMIT) consortium, funded by the European Union Innovative Medicines Initiative, is a collaboration involving 19 academic centres and six European Federation of Pharmaceutical Industries and Associations (EFPIA) partners across Europe to identify and characterise biomarkers for complications of diabetes [2]. We measured a panel of 42 candidate biomarkers with existing evidence for association with CVD risk in 1,123 incident CVD cases and 1,187 controls with type 2 diabetes from five European cohorts. The aims of the analysis were to select biomarkers predictive of CVD and assess the added value of the biomarkers beyond Framingham and clinical factors to prediction of CVD.
Methods
Data and sample sources
This study was a nested case–control design matched for sex and age. Cases and controls were identified within five cohorts: Genetics of Diabetes Audit and Research Tayside Study (Go-DARTS), Scania Diabetes Registry (SDR), MONItoring of trends and determinants in CArdiovascular disease/Cooperative Health Research in the Region of Augsburg (MONICA/KORA) study, Carotid Intima Media Thickness [IMT] and IMT-Progression as Predictors of Vascular Events in a High Risk European Population (IMPROVE) study and Stockholm 60-year Old Study [3–7] (electronic supplementary material [ESM] Table 1). For each cohort, participants with type 2 diabetes and no clinical history of CVD when the blood samples were taken were eligible for inclusion. A small number (n = 65) of participants who were prediabetic (i.e. had non-diabetic hyperglycaemia at the time of sampling and subsequently progressed to type 2 diabetes) were also eligible. Incident major CVD cases were defined as having acute CHD or an ischaemic stroke event subsequent to the baseline sample being taken. Detailed criteria are included in the ESM (Methods: Data and sample source).
Controls were free of CVD at the end of follow-up and matched to cases within each cohort by sex and were stratum-matched for age at the time of blood sampling (within 5 years). Overall 2,318 individuals were sampled, of whom eight had insufficient sample volume, leaving serum samples from 1,123 cases and 1,187 controls.
Biomarker selection process and laboratory analysis
Biomarker selection was based on existing evidence from the literature (incorporating a text mining process [8] and a manual literature review) and the availability of reliable validated assays to measure biomarker concentrations in small volumes of serum. We prioritised biomarkers of cardiac damage (high-sensitivity troponin T [hsTnT] and N-terminal pro-B-type natriuretic peptide [NT-proBNP]), renal disease (cystatin C, fetuin A), glucose-mediated damage in diabetes (soluble receptor for AGE [sRAGE]), matrix remodelling (matrix metalloproteinases and osteopontin) and coagulation (factor VII), as well as a lipoprotein of particular interest in diabetes (apolipoprotein C-III [apoCIII]) [9, 10], and a large set of immune- and inflammation-related proteins including interleukins and interferons. We measured 33 protein biomarkers (ESM Table 2) by multiplex immunoassay on the Human DiscoveryMAP customised panel using the Luminex 100/200 instrument by RBM (Myriad Rules Based Medicine, Austin, TX, USA) [11] and standard commercially available ELISAs and kits for nine biomarkers. The included biomarkers are listed in ESM Table 2 along with details of assays and assay metrics. For most biomarkers the inter-assay CV was <10% with the exception of apoCIII (16%), cathepsin S (17%), granulocyte-macrophage colony-stimulating factor (23%) and TNF-α (18%). (For more details on laboratory methods see ESM Methods: Biomarker selection process and laboratory analysis and ESM Table 2).
Statistical methods
Choice of covariates in the baseline model
Clinical covariates included in the models for biomarker selection were selected based upon their inclusion in existing CVD risk engines [12, 13] or known associations with CVD in type 2 diabetes and their availability for all cohorts. We also included baseline medication data in case any biomarker measurements were affected by drug use. All clinical covariates were measured at or close to the time of sampling. The chosen covariates were age, sex, smoking, systolic and diastolic blood pressure, LDL-cholesterol (LDL-C), HDL-cholesterol (HDL-C), triacylglycerol, diabetes duration, HbA1C, BMI, height, estimated GFR (eGFR) calculated using the Modification of Diet in Renal Disease 4-variable (MDRD4) equation, cohort, and current medication (including antihypertensive agents, aspirin, lipid-lowering agents and insulin).
Pre-processing and imputation
All continuous variables were Gaussianised using the LambertW package version 0.5 in R (http://cran.r-project.org/web/packages/LambertW) [14, 15]. Missing values of biomarkers and covariates were imputed using a sparse iterative regression model. For six biomarkers, >96% of participants had below detectable or missing levels and these were not analysed further (ESM Methods: Pre-processing and imputation). The impact of imputation on the analyses was explored.
Selection of predictive sets of biomarkers
We applied two complementary approaches for selecting which subsets of biomarkers contributed most to prediction: forward selection using logistic regression, and a top-down method based on logistic regression with an L1 (least absolute shrinkage and selection operator [LASSO]) regularisation penalty [16]. See ESM for details (Methods: Selection of predictive sets of biomarkers).
Predictive performance evaluation
We evaluated predictive performance of models by computing the area under the receiver operating characteristic curve (AUROC) on 50 folds of test data, where the test folds were not used either for model fitting or for biomarker selection. We used 30 inner folds to iteratively select biomarkers (forward selection) or to learn the penalty parameter (top-down approach). This nested cross-validation procedure provides an unbiased estimate of the predictive performance of the considered models. We assessed four models, two including only clinical covariates and two including clinical covariates and biomarkers. The first model considered is based on the Framingham Risk Score and was limited to the clinical factors in that score along with the relevant interaction terms for variables that have stratum-specific coefficients in that equation; the main effects were age, sex, total cholesterol, HDL-C, systolic blood pressure, antihypertensive treatment and current smoking, and interaction terms (sex × other variables, systolic blood pressure × antihypertensive treatment) were also allowed to enter the model [12]. We also considered a model including an extended set of clinical factors that included clinical measures beyond the Framingham Risk Score factors, including important diabetes measures such as diabetes duration, HbA1c, eGFR and insulin use. To the extended clinical covariate model we then added selected biomarkers from either the forward selection or the top-down selection models.
Net reclassification improvement
To assess the clinical relevance of the increment in predictive performance and for consistency with other reports on biomarker performance we estimated the net reclassification improvement (NRI) with a classification threshold of 10% risk at 5 years. Calculation of the NRI from a nested case–control study requires the reconstruction of the original cohort from the cases and controls that were sampled from this cohort [17]. This reconstruction was done by weighting the observations on cases and controls so that the proportion of cases equated to the risk at 5 years computed from the annual event rate, and by using the logistic regression model from the nested case–control study to calculate the predictive probability of disease for individuals in this reconstructed cohort. The event rates for the Go-DARTS cohort were used as a proxy for the rates in all cohorts.
Results
The final dataset consisted of 36 biomarkers in 2,310 individuals (1,123 cases, of which 755 were acute CHD and 368 strokes, and 1,187 controls). Median time to event was 3.2 years (interquartile range [IQR] 1.5–4.9) for cases and median follow-up for controls was 6.5 years (IQR 3.9–7.9). Baseline clinical data are summarised in Table 1.
Forward selection of biomarkers using logistic regression
The distributions of biomarkers in cases and controls are summarised in ESM Table 3. After adjustment for all clinical covariates, 16 biomarkers showed association with CVD at p < 0.05 (shown in the first two data columns in Table 2). The first iteration of the forward selection process fits a cross-validated logistic regression model for each biomarker singly, and evidence for association independently of all clinical covariate data is given by the magnitude of the increment in validation log-likelihood and by the false discovery rate (FDR)—the lower the FDR the stronger the association (Table 2). The strongest associations with CVD were for NT-proBNP, hsTnT, IL-6, apoCIII, cystatin C and IL-15 (p < 0.001, and with increments in test log-likelihood ≥5 natural log units and FDR <0.1 in the cross-validated forward selection models). Of these biomarkers, other than apoCIII, levels were higher in cases than controls.
Many of the biomarkers studied showed strong correlations with each other (ESM Fig. 1). Table 3 shows the selection of biomarkers in the forward selection nested procedure and their rank in the models (after all iterations) summarised over the 50 outer test/training folds. Six biomarkers were selected in 100% of the 50 outer training folds as improving prediction of CVD beyond clinical covariates. The ORs for the final model including these six biomarkers are also shown in Table 3.
Five of the six selected biomarkers were those that showed the strongest associations examined singly. Cystatin C, which showed a strong univariate association, was not retained by the forward regression due to its strong correlation with other biomarkers including NT-proBNP. Conversely, sRAGE was retained in the forward selection although its association with CVD before adjustment for the other biomarkers was weak. The biomarkers selected by forward selection were also retained as the best predictive biomarkers by the top-down method (data not shown). We used forward selection to identify which clinical covariates, beyond those included in the Framingham Risk Score and the already selected biomarkers, were important predictors of CVD. Three additional clinical covariates were selected: (in order of selection in the forward selection model) insulin therapy status, eGFR and HbA1c. See ESM (Results: Forward selection of biomarkers using logistic regression) for evaluation of the impact of imputation, Gaussianisation and the deletion of outliers on the biomarker selections.
Predictive value of models evaluated using AUROC
The impact of adding clinical covariates and biomarkers to the Framingham Risk Score model is shown in Table 4. The addition of the extended clinical covariates to the Framingham Risk Score model significantly increased the test log-likelihood as did addition of the selected biomarkers. There was also a significant impact of the addition of the biomarkers to the full clinical covariate model. The Framingham Risk Score model had the poorest predictive performance (AUROC = 0.59). However, in an age and sex stratum matched design the predictive performance attributable to the age and sex covariates is constrained by the matched design so the AUROCs for all the models will be less than if age and sex are not matched. The AUROCs were 0.66 for models including clinical covariates, only 0.72 for models with addition of biomarkers chosen by forward selection, and 0.71 for models learned from all biomarkers by the top-down approach. The greatest increase in AUROC was seen on addition of the first biomarker, which was always NT-proBNP (ESM Fig. 2).
NRI
By use of a 5 year CVD risk threshold of 10% to define high risk, adding the six biomarkers chosen by forward selection to a model based on clinical covariates only resulted in 3.67% of cases and 4.40% of non-cases being reclassified with respect to this threshold, giving a NRI of 8.07%.
Discussion
In this study of 42 potential CVD biomarkers we confirmed associations with CVD independently of clinical covariates for 16 biomarkers, and use of these biomarkers modestly improved the prediction of CVD beyond that obtained by clinical covariates with an increase in the AUROC from 0.66 to 0.72. Based on the conventional decision threshold of a 5-year CVD risk of 10%, we estimated that 8% of individuals would be reclassified. This improvement was generated by six biomarkers: NT-proBNP, hsTnT, IL-6, IL-15, apoCIII and sRAGE.
Our analysis has confirmed previously reported associations as well as novel associations with CVD in diabetes. There is increasing data on the importance of NT-proBNP [18] and hsTnT [19] for CVD prediction in the general population. Our data confirm their importance as predictive biomarkers in diabetes independently of eGFR and the other biomarkers. ApoCIII inhibits the catabolism of triacyglycerol-rich lipoproteins, partly through inhibition of their hepatic uptake and partly through inhibition of lipoprotein lipase activity. ApoCIII non-specific gene deletion in animal models is associated with premature atherosclerosis [20], and most studies suggest a likely pro-atherogenic role [21]. ApoCIII is known to be an important link between glycaemia and dyslipidaemia [22] as its expression is induced by high glucose levels and is responsive to insulin. However, whilst several studies have shown that higher apoCIII content on lipoproteins is associated with their atherogenicity [23, 24], the association of total circulating apoCIII itself with CVD has had less investigation [21]. One of the largest studies is the Hoorn study [25]. In this study, participants who died of CVD had higher baseline levels of apoCIII, and the association was mediated in part by the positive correlation of apoCIII levels with dysglycaemia and triacylglycerol levels [25]. In another analysis of two sex-specific cohorts, after adjustment for triacylglycerol levels, no association was found between total plasma apoCIII levels and incident CVD [24]. Our finding of an inverse association with CVD that became even more apparent on adjustment for HDL-C and triacylglycerol levels is therefore unexpected. As the direction of association with apoCIII was unexpected, we confirmed that apoCIII levels correlated positively with triacylglycerol levels (r = 0.25) but had little correlation with LDL-C (r = −0.02) or HDL-C (r = 0.06). The intra- and inter-assay CVs were quite high at 14.56% and 15.84%, respectively (ESM Table 2), but we confirmed that the lower apoCIII levels in cases than controls was found in all participating cohorts (data not shown). Thus, we are confident that the assay performed as expected and that this is a true finding that warrants further study. We note that our analysis is in non-fasting frozen samples; however, apoCIII has been shown to be unaltered by fasting [26] and by freezing [27]. One small study in patients with type 2 diabetes also found a lower level of apoCIII in cases than controls of CVD [27]. In view of the unexpected association in our study and the elevated inter-run CV for the assay, we re-ran all analyses excluding apoCIII and found that the selection of the other five biomarkers was not affected, although the total AUROC was somewhat lower (data not shown).
IL-15 is a proinflammatory cytokine for which there is substantial cell and animal model evidence for a role in atherosclerosis [28, 29], but its characteristics as a predictive biomarker for CVD in diabetes have not been reported previously. IL-6 has been characterised as a risk factor for CVD in several cohorts [30] and we confirm its importance here. In a subanalysis of the MONICA/KORA samples that also included C-reactive protein (CRP) measurement, the association between IL-6 and CVD was attenuated but the association of IL-15 with CVD remained (data not shown). Thus, it is likely that IL-6 will not add a great amount of information to a model in the presence of CRP. sRAGE has not been studied in many prospective studies of type 2 diabetes; we previously showed that higher levels were associated with an increased risk of CHD but little difference in stroke risk in clinical trials [31]. The inverse association between sRAGE and CVD found here, which is only apparent once it has been adjusted for eGFR and other biomarkers, is contradictory and requires further exploration in other cohorts.
We have used a candidate biomarker approach in this study rather than a global discovery approach using proteomics or metabolomics platforms. Of the biomarkers included in our study, osteopontin has also been identified as a biomarker in a proteomics study of atherosclerotic plaques and then validated in serum [32]. Other potential biomarkers, such as heat shock protein 27, cathepsin D and transthyretin [33], which have been identified by proteomics of atherosclerotic plaques, or the cytoskeletal protein vinculin from plasma proteomic profiling [34], were not included in our study.
To compare biomarkers we used an approach that is common in the field of machine learning, i.e. N-fold cross-validation, in which the entire dataset from across cohorts is partitioned into N test folds, models are fitted to each of the corresponding training folds and the performance of these models is evaluated by their ability to predict outcome in the test folds. We consider that the advantages of this approach in comparison with model fitting, for example on one half of the available cohorts and performance assessment in the other half are: (1) with N-fold cross-validation, model fitting uses a fraction (1–1/N) of the observations rather than only half the observations so it is more powerful if N is large; (2) evaluation of predictive performance on test data uses all the observations because every observation appears once in a test fold; and (3) all cohorts are represented in each training dataset and in each test dataset, so we can expect that when the existing cohorts are representative of new test patients the results should be more widely generalisable than the conventional approach.
A recent systematic review examined 12 CVD-related models developed for CVD prediction in diabetes and 33 that included diabetes as a factor in the model [35]. Of those developed in patients with diabetes, four examined total CVD risk, with two—one developed in the ADVANCE (Action in Diabetes and Vascular Disease: Preterax and Diamicron Modified Release Controlled Evaluation) study [36] and a Swedish study [37]—reporting the AUROC without any correction for overfitting (0.70 in both). Both of these models included age, sex, diabetes duration and HbA1c. The ADVANCE model also included albuminuria, retinopathy, non-HDL-cholesterol, atrial fibrillation and pulse pressure. The Swedish model included smoking, BMI, lipid-lowering drug use and systolic blood pressure but no measure of renal function. The United Kingdom Prospective Diabetes Study (UKPDS) risk engine for CHD and the Framingham Risk Score for CVD are the models most commonly recommended, but neither includes any measure of renal function, glycaemic control or stage or severity of diabetes. Our analysis shows that inclusion of eGFR, HbA1c and insulin therapy status is also important for CVD prediction in diabetes.
The clinical utility of this work is somewhat limited at present, in that the overall improvement in prediction due to addition of biomarkers is modest over and above clinical covariates. However, this modest improvement may still be of use when designing CVD endpoint trials as another means of enriching trials for high-risk participants. We have also demonstrated that inclusion of diabetes-specific CVD risk factors does improve the prediction of the models and should be considered when assessing patient risk.
A limitation of our study was that all the cohorts included were European and the extent to which our conclusions generalise to other populations remains to be established. A further issue was that low sample volumes precluded study of a more comprehensive set of all candidate biomarkers. This issue is a practical constraint for many epidemiologic cohorts. Larger gains in prediction might be demonstrable through measuring high dimensional metabolomic and lipidomic panels evaluable on small volumes of serum and this approach will be evaluated further. Finally, predictive performance is also a function of assay method: where more sensitive or more accurate assay methods are used these would be expected to improve the performance of the biomarker. Whilst we found that several of the cytokines on the Human DiscoveryMAP panel were undetectable in 70% or more of samples, a high level of missingness is to be expected for these low abundance cytokines when acute inflammation is not present.
In conclusion, six biomarkers—NT-proBNP, hsTnT, IL-6, IL-15, apoCIII and sRAGE—captured most of the predictive information about CVD from a panel of 42 biomarkers. By use of these biomarkers and an extensive set of clinical covariates, prediction was considerably improved compared with using only the covariates included in the Framingham Risk Score. For optimised prediction of CVD in diabetes, novel biomarkers can contribute; however, measures of renal function and diabetes-specific factors, not currently commonly used in risk scores, need to be included.
Abbreviations
- ADVANCE:
-
Action in Diabetes and Vascular Disease: Preterax and Diamicron Modified Release Controlled Evaluation
- apoCIII:
-
Apolipoprotein C-III
- AUROC:
-
Area under the receiver operating characteristic curve
- CRP:
-
C-reactive protein
- CVD:
-
Cardiovascular disease
- EFPIA:
-
European Federation of Pharmaceutical Industries and Associations
- eGFR:
-
Estimated GFR
- FDR:
-
False discovery rate
- Go-DARTS:
-
Genetics of Diabetes Audit and Research Tayside Study
- HDL-C:
-
HDL cholesterol
- hsTnT:
-
High-sensitivity troponin T
- IMPROVE:
-
Carotid Intima–Media Thickness [IMT] and IMT-Progression as Predictors of Vascular Events in a High Risk European Population
- IQR:
-
Interquartile range
- LASSO:
-
Least absolute shrinkage and selection operator
- LDL-C:
-
LDL cholesterol
- MONICA/KORA:
-
MONItoring of trends and determinants in CArdiovascular disease/Cooperative Health Research in the Region of Augsburg
- NT-ProBNP:
-
N-terminal pro-B-type natriuretic peptide
- NRI:
-
Net reclassification improvement
- SDR:
-
Scania Diabetes Registry
- sRAGE:
-
Soluble receptor for AGE
- SUMMIT:
-
SUrrogate markers for Micro- and Macro-vascular hard endpoints for Innovative diabetes Tools
References
U.S. Food and Drug Administration (2008) Guidance for industry: diabetes mellitus—evaluating cardiovascular risk in new antidiabetic therapies to treat type 2 diabetes. U.S. Food and Drug Administration, Silver Spring
Innovative Medicines Initiative (2010) Summit. Available from www.imi.europa.eu/content/summit. Accessed 30 Oct 2014
Pearson ER, Donnelly LA, Kimber C et al (2011) Variation in TCF7l2 influences therapeutic response to sulfonylureas—a GoDARTs study. Diabetes 56:2178–2182
Ahluwalia TS, Lindholm E, Groop LC (2011) Common variants in CNDP1 and CNDP2, and risk of nephropathy in type 2 diabetes. Diabetologia 54:2295–2302
Malarstig A, Silveira A, Wagsater D et al (2011) Plasma CD93 concentration is a potential novel biomarker for coronary artery disease. J Intern Med 270:229–236
Baldassarre D, Nyyssönen K, Rauramaa R et al (2010) Cross-sectional analysis of baseline data to identify the major determinants of carotid intima–media thickness in a European population: the IMPROVE study. Eur Heart J 31:614–622
Thorand B, Kolb H, Baumert J et al (2005) Elevated levels of interleukin-18 predict the development of type 2 diabetes: results from the MONICA/KORA Augsburg study, 1984–2002. Diabetes 54:2932–2938
Nuzzo A, Mulas F, Gabetta M et al (2010) Text mining approaches for automated literature knowledge extraction and representation. Stud Health Technol Inform 160:954–958
Hiukka A, Fruchart-Najib J, Leinonen E, Hilden H, Fruchart JC, Taskinen MR (2005) Alterations of lipids and apolipoprotein CIII in very low density lipoprotein subspecies in type 2 diabetes. Diabetologia 48:1207–1215
Gervaise N, Garrigue MA, Lasfargues G, Lecomte P (2000) Triglycerides, apo C3 and Lp B:C3 and cardiovascular risk in type II diabetes. Diabetologia 43:703–708
Myriad RBM (2011) HumanMAPs. Available from https://rbm.myriad.com/products-services/humanmap-services/. Accessed 30 Oct 2014
D’Agostino RB Sr, Vasan RS, Pencina MJ et al (2008) General cardiovascular risk profile for use in primary care: the Framingham Heart Study. Circulation 117:743–753
Stevens RJ, Kothari V, Adler AI, Stratton IM (2001) The UKPDS risk engine: a model for the risk of coronary heart disease in type II diabetes (UKPDS 56). Clin Sci 101:671–679
R Development Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Chen SSB, Gopinath RA (2001) Gaussianization. Adv Neural Inf Process Syst 13:423–429
Tibshirani R (1996) Regression shrinkage and selection via the Lasso. J R Stat Soc Ser B Methodol 58:267–288
Bansal A, Pepe MS (2013) Estimating improvement in prediction with matched case-control designs. Lifetime Data Anal 19:170–201
Battistoni A, Rubattu S, Volpe M (2012) Circulating biomarkers with preventive, diagnostic and prognostic implications in cardiovascular diseases. Int J Cardiol 157:160–168
Saunders JT, Nambi V, de Lemos JA et al (2011) Cardiac troponin T measured by a highly sensitive assay predicts coronary heart disease, heart failure, and mortality in the Atherosclerosis Risk in Communities Study. Circulation 123:1367–1376
Ordovas JM, Cassidy DK, Civeira F, Bisgaier CL, Schaefer EJ (1989) Familial apolipoprotein A-I, C-III, and A-IV deficiency and premature atherosclerosis due to deletion of a gene complex on chromosome 11. J Biol Chem 264:16339–16342
Chan DC, Chen MM, Ooi EM, Watts GF (2008) An ABC of apolipoprotein C-III: a clinically useful new cardiovascular risk factor? Int J Clin Pract 62:799–809
Caron S, Verrijken A, Mertens I et al (2011) Transcriptional activation of apolipoprotein CIII expression by glucose may contribute to diabetic dyslipidemia. Arterioscler Thromb Vasc Biol 31:513–519
Hiukka A, Stahlman M, Pettersson C et al (2009) ApoCIII-enriched LDL in type 2 diabetes displays altered lipid composition, increased susceptibility for sphingomyelinase, and increased binding to biglycan. Diabetes 58:2018–2026
Mendivil CO, Rimm EB, Furtado J, Chiuve SE, Sacks FM (2011) Low-density lipoproteins containing apolipoprotein C-III and the risk of coronary heart disease. Circulation 124:2065–2072
Scheffer PG, Teerlink T, Dekker JM et al (2008) Increased plasma apolipoprotein C-III concentration independently predicts cardiovascular mortality: the Hoorn Study. Clin Chem 54:1325–1330
Tentolouris N, Stylianou A, Lourida E et al (2007) High postprandial triglyceridemia in patients with type 2 diabetes and microalbuminuria. J Lipid Res 48:218–225
Roselli della Rovere G, Lapolla A, Sartore G et al (2003) Plasma lipoproteins, apoproteins and cardiovascular disease in type 2 diabetic patients. A nine-year follow-up study. Nutr Metab Cardiovasc Dis 13:46–51
Gokkusu C, Aydin M, Ozkok E et al (2010) Influences of genetic variants in interleukin-15 gene and serum interleukin-15 levels on coronary heart disease. Cytokine 49:58–63
van Es T, van Puijvelde GH, Michon IN et al (2011) IL-15 aggravates atherosclerotic lesion development in LDL receptor deficient mice. Vaccine 29:976–983
Lee JK, Bettencourt R, Brenner D, Le TA, Barrett-Connor E, Loomba R (2012) Association between serum interleukin-6 concentrations and mortality in older adults: the Rancho Bernardo study. PLoS ONE 7:e34218
Colhoun HM, Betteridge DJ, Durrington P et al (2011) Total soluble and endogenous secretory receptor for advanced glycation end products as predictive biomarkers of coronary heart disease risk in patients with type 2 diabetes: an analysis from the CARDS trial. Diabetes 60:2379–2385
de Kleijn DP, Moll FL, Hellings WE et al (2010) Local atherosclerotic plaques are a source of prognostic biomarkers for adverse cardiovascular events. Arterioscler Thromb Vasc Biol 30:612–619
Bleijerveld OB, Zhang YN, Beldar S et al (2013) Proteomics of plaques and novel sources of potential biomarkers for atherosclerosis. Proteomics Clin Appl 7:490–503
Kristensen LP, Larsen MR, Mickley H et al (2014) Plasma proteome profiling of atherosclerotic disease manifestations reveals elevated levels of the cytoskeletal protein vinculin. J Proteome 101:141–153
van Dieren S, Beulens JW, Kengne AP et al (2012) Prediction models for the risk of cardiovascular disease in patients with type 2 diabetes: a systematic review. Heart 98:360–369
Kengne AP, Patel A, Marre M et al (2011) Contemporary model for cardiovascular risk prediction in people with type 2 diabetes. Eur J Cardiovasc Prev Rehabil 18:393–398
Cederholm J, Eeg-Olofsson K, Eliasson B, Zethelius B, Nilsson PM, Gudbjornsdottir S (2008) Risk prediction of cardiovascular disease in type 2 diabetes: a risk equation from the Swedish National Diabetes Register. Diabetes Care 31:2038–2043
Acknowledgements
We wish to acknowledge all the SUMMIT partners (www.imi-summit.eu/). Reagents for the measurement of NT-proBNP, hsTnT and cystatin C were a donation from Roche Diagnostics AG (Penzberg, Germany).
Funding
The research leading to these results has received support from the Innovative Medicines Initiative Joint Undertaking under grant agreement no. 115006, resources of which are composed of financial contribution from the European Union’s Seventh Framework Programme (FP7/2007-2013) and in-kind contribution from EFPIA companies.
Duality of interest
FA is a director of the data analysis company Pharmatics Limited; he declares that there is no duality of interest in relation to the work described. EN is an employee with F. Hoffmann-La Roche Ltd, Switzerland. PMM is a stakeholder in Pharmatics Limited; he declares that there is no duality of interest in relation to the work described. HMC reports grants and personal fees from Pfizer Inc., grants and institutional consultancy fees from Sanofi Aventis, Regeneron and Novartis Pharmaceuticals. HCL, MC, TZ, LG, BT, CNP, AH, UdF, SJL, VS, NB, RB, NvZ and KL declare that there is no duality of interest associated with their contribution to this manuscript.
Contribution statement
HCL contributed to the interpretation of the data and drafted the manuscript. MC and FA contributed to the analysis and interpretation of the data and critically revised the manuscript. TZ, LG, BT, CNP, AH, UdF, KL and NVZ contributed to the acquisition of data and critically revised the manuscript. EN and VS contributed to the interpretation of the data and critically revised the manuscript. SJL contributed to the data analysis and critically revised the manuscript. NB and RB contributed to the study design and critically revised the manuscript. PMM contributed to the study design, analysis and interpretation of data and critically revised the manuscript. HMC contributed to the study conception and design, analysis and interpretation of data and critically revised the manuscript. All authors approved the version of the manuscript for publication. HMC is the guarantor of this work.
Author information
Authors and Affiliations
Consortia
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM Methods
(PDF 244 kb)
ESM Table 1
(PDF 195 kb)
ESM Table 2
(PDF 210 kb)
ESM Table 3
(PDF 160 kb)
ESM Fig. 1
(PDF 540 kb)
ESM Fig. 2
(PDF 260 kb)
Rights and permissions
About this article
Cite this article
Looker, H.C., Colombo, M., Agakov, F. et al. Protein biomarkers for the prediction of cardiovascular disease in type 2 diabetes. Diabetologia 58, 1363–1371 (2015). https://doi.org/10.1007/s00125-015-3535-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-015-3535-6