Abstract
Key message
A novel light-dependent dominant lesion mimic mutant with enhanced multiple disease resistance was physiologically, biochemically, and genetically characterized; the causal gene was fine mapped to a 909 kb interval containing 38 genes.
Abstract
Identification of genes that confer multiple disease resistance (MDR) is crucial for the improvement of maize disease resistance. However, very limited genes are identified as MDR genes in maize. In this study, we characterized a dominant disease lesion mimics 8 (Les8) mutant that had chlorotic lesions on the leaves and showed enhanced resistance to both curvularia leaf spot and southern leaf blight. Major agronomic traits were not obviously altered, while decreased chlorophyll content was observed in the mutant, and the genetic effect of the Les8 mutation was stable in different genetic backgrounds. By BSR-seq analysis and map-based cloning, the LES8 gene was mapped into a 909 kb region containing 38 candidate genes on chromosome 9 wherein no lesion mimic or disease-resistance genes were previously reported. Using transcriptomics analysis, we found that genes involved in defense responses and secondary metabolite biosynthesis were enriched in the significantly up-regulated genes, while genes involved in photosynthesis and carbohydrate-related pathways were enriched in the significantly down-regulated genes in Les8. In addition, there was an overaccumulation of jasmonic acid and lignin but not salicylic acid in Les8. Taken together, this study revealed candidate genes and potential mechanism underlying Les8-conferred MDR in maize.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Multiple disease resistance (MDR) has been attracting increasing attention due to its importance in crop improvement. However, very few genes have been definitively identified as MDR genes in maize (Simmons et al. 1998; Yang et al. 2017; Li et al. 2022b; Wang et al. 2021). Disease lesion mimics mutants are a class of mutants that spontaneously display disease-like lesions without pathogen infection or obvious stimulus of injury. Since a large portion of such mutants showed enhanced disease resistance to various diseases, they have been considered as valuable resource for MDR studies (Walbot et al. 1983; Johal et al. 1995; Lorrain et al. 2003; Zhu et al. 2020; Li et al 2022a).
Disease lesion mimic mutants are abbreviated as Les for dominant and les for recessive mutations and have been studied in a few plant species. In arabidopsis, mutations of genes encoding NB-LRR proteins, RLKs/RLPs, and other types of proteins results in spontaneous leaf lesion and enhanced disease resistance (Lorrain et al. 2003; Wersch et al. 2016). For example, a gain-of-function mutation in TIR-NBS-LRR protein SNC1 resulted in constitutive expression of pathogenesis-related (PR) genes and significantly enhanced resistance against bacterial and fungal pathogens (Zhang et al. 2003). In rice, recent studies showed at least 30 Les or les mutants are implicated in immune response and cell death (Zhu et al. 2020). For instance, loss-of-function of CRL3 leads to increased accumulation of flg22- and chitin-induced reactive oxygen species, up-regulated PR gene expression, and enhanced resistance to Magnaporthe oryzae and Xanthomonas oryzae pv oryzae (Liu et al. 2017). Besides, genome editing of rice les gene RBL1 encoding a cytidine diphosphate diacylglycerol synthase confers broad-spectrum disease resistance without fitness penalty (Sha et al. 2023). In barley, loss-of-function of Les genes MLO confers broad-spectrum resistance to powdery mildew (Büschges et al. 1997; Piffanelli et al. 2004). Recently, an elite wheat variety possessing robust powdery mildew resistance was generated by genome editing of the wheat MLO (Li et al. 2022c). In rapeseed, the loss-of-function of Les gene LMM1 confers basal resistance to Sclerotinia sclerotiorum (Yu et al. 2023). Taken together, previous studies in various species highlight the value of Les or les mutants in dissecting the core plant immune factors and improving disease resistance of crops.
It was estimated that more than 200 lesion mimic loci might exist in maize (Neuffer and Calvert 1975), among which, 32 mutants have been genetically characterized, and 23 are dominant mutants, representing the largest class of gain-of-function mutants in maize (Johal et al. 1995). However, thus far, only 5 genes have been cloned and characterized, including Rp1-D21, LLS1/LES30, LES22, ZMM1, and NEC-T. The dominant Les gene Rp1-D21 was derived from an intergenic recombination event between two NLR genes, Rp1-D and Rp1-dp2 (Hu et al. 1996). Using the lesion phenotype of Rp1-D21 as a reporter, the modifiers of Rp1-D21-mediated hypersensitive defense response, including ZmCCoAOMT, ZmMIEL, were identified by genome-wide association analysis and quantitative trait loci mapping (Chintamanani et al. 2010; Chaikam et al. 2011; Olukolu et al. 2014). The biological function in defense response of those genes were further validated using a transient overexpression system in Nicotiana benthamiana, virus-induced gene silencing, and gene editing technologies in maize (Wang and Balint-Kurti 2016; Luan et al. 2020; Karre et al. 2021; Li et al. 2023). LLS1/LES30 encodes a pheophorbide a oxidase (PAO) that mediates chlorophyll degradation and triggers MDR (Gray et al. 1997; Simmons et al. 1998; Li et al. 2022b). Similarly, LES22 encodes uroporphyrinogen decarboxylase (UROD), a key enzyme in chlorophyll and heme biosynthesis (Hu et al. 1998). ZMM1 is a teosinte-derived allele of MYB transcription factor, which confers MDR to northern leaf blight, gray leaf spot, and southern corn rust (Wang et al. 2021). Besides, loss of NEC-T function disturbs the tetrapyrrole pathway, thus forming lesions in the mutant leaves (Zhao et al. 2022).
In this study, we characterized a dominant disease lesion mimics 8 (Les8), which showed elevated resistance to curvularia leaf spot (CLS) and southern leaf blight (SLB). Genetic analysis indicated the phenotype of Les8 was controlled by a dominant monogenic mutation. By BSR-seq analysis and map-based cloning, we mapped the LES8 gene to a 909 kb region containing 9 high-confidence candidate genes wherein no lesion mimic or disease-resistance genes were previously reported. Underlying mechanisms of Les8-conferred MDR were also revealed by transcriptomic and biochemical assay in this study.
Materials and methods
Plant material and growth conditions
The maize lesion mimic mutant Les8 (927D, Les8-N2005) was obtained from the Maize Genetics Cooperation Stock Center (http://maizecoop.cropsci.uiuc.edu/). We successively self-crossed the heterozygous Les8 plants to the F7 generation to purify the genetic background. All the plants were grown in the field at the experimental station of the Henan Agricultural University in Zhengzhou in summer, Henan Province, China, and the experimental station of the Henan Agricultural University in Sanya in winter, Hainan Province.
Physiological and biochemical analyses
To measure the content of chlorophyll, uniform leaves of 40-days-old WT and Les8 plant grown in the field were sampled. Six biological replicates were used, and each replicate consisted of leaves mixed from three independent plants. Extraction and measurement of chlorophyll were performed according to the previous method described by Qiu et al. (2016).
To detect H2O2, fresh leaf samples of three biological replicates from three different plants were cut into 2 × 3 cm rectangles and put into 1 mg ml–1 diaminobenzidine (DAB) solution (pH 3.8). After 8 h in the dark, the previous DAB was replaced by 90% ethanol to remove chlorophyll according to our previous study (Li et al. 2022b). Images were taken using a stereomicroscope (Olympus SZX7).
Evaluation of resistance to Curvularia lunata, Bipolaris maydis, and the growth assay of CX-3 under the supplement of the fresh leaf extraction
The CX-3, a dominant strain of Curvularia lunata (Wakker) Boed in China, was cultured on potato dextrose agar medium at 28 °C for 1 week in the dark. A suspension containing 1 × 106 spores ml–1 in distilled water with 0.02% Tween 20 was used to spray 40-days-old leaves of WT and Les8, and control leaves were sprayed with distilled water containing 0.02% Tween 20 only. For quantification of DNA of C. lunata and B. maydis, DNA was extracted from leaves of five biological replicates (three plants per replicate) from CX-3 inoculated and naturally infected leaves using the hexadecyltrimethyl-ammonium bromide (CTAB) method (Chen and Ronald 1999) and subjected to quantitative real-time (qRT)-PCR using CX-3 specific primers of the Clg2p gene (CLF-4, CLR-4, Table S1) and B. maydis specific primers of the ITS gene (BMF-3, BMR-3, Table S1). Plant DNA quantification was performed using primers specific for ZmGAPC1 (ZmGAPC1-DNA-F1, ZmGAPC1- DNA-R1, Table S1).
For the CX-3 culture assay, 20–50 g leaves of WT and Les8 were ground into powder using liquid nitrogen. After a supplement of 5 mL distilled water, the mixtures were filtered using a 0.22 μm filter membrane, and 2 ml of the fresh leaf juice obtained was then spread onto the surface of the PDA plates and dried in the clean bench. The 10 μL of 1:1 mixed CX-3 suspension (1 × 105 spores ml–1) and the fresh leaf juice was then dropped on the center of pretreated PDA plates. After cultured in the 28 °C for some time, the colony diameter and spore number of 10 replicates were measured and counted.
RNA library construction, sequencing, and bioinformatics analysis
Uniform leaves of three biological replicates (three independent plants per replicate) were sampled from 40-days-old WT and Les8. Total RNAs were extracted using an RNA extraction kit (Tiangen, DP432). Sequencing libraries were generated using NEBNext® Ultra™ RNA Library Prep Kit (NEB, E7770) according to the manufacturer’s instruction. Before sequencing, index codes were added to attribute sequences to each sample. Sequencing was performed on an Illumina Novaseq platform, and 150-bp paired-end reads were generated. The clean reads were then generated after removing low-quality sequencing reads and sequencing adapters.
Bioinformatics analysis of RNA-seq data
To obtain differently expressed genes (DEGs) between WT and Les8. Clean reads were aligned to the B73 RefGen_v4 maize reference genome using HISAT2 (Kim et al. 2015). The expression of each gene was normalized to fragments per kilobase of transcript per million reads (RPKM), and the R package DESeq2 (Love et al. 2014) was used to identify DEGs with fold-change (FC) > 2 and a false discovery rate (FDR) of < 0.01. Pearson’s correlation was performed to calculate the association between samples using the cor function in the R base package. To investigate the putative biological pathways regulated by the DEGs of WT and Les8, Gene Ontology (GO) enrichment analysis of the up-DEGs and down-DEGs was implemented by the GOseq R package-based Wallenius non-central hyper-geometric distribution (Young et al 2010).
Generation of the mapping population, bulked segregant RNA sequencing (BSR-seq), and fine mapping
To clone the candidate gene responsible for the phenotype of Les8, a heterozygous Les8 plant was crossed to the B73 plants, and the F2 population of B73 × Les8 was generated by selfing the F1 plants with lesion mimic phenotype. For BSR-seq, 30 F2 plants each with the mutant phenotype, and 30 F2 plants each with the WT phenotype were identified and collected as two bulk samples. Total RNAs were then extracted and sequenced as described above. BSR analysis was performed as a previously reported method (Liu et al. 2012). B73 RefGen_v4 genome was used as the reference genome for designing mapping primers and candidate gene prediction. The primer sequences of the molecular markers are listed in Table S1.
Cell wall preparation and quantification of lignin
Uniform leaves of four biological replicates (three independent plants per replicate) were sampled from 60-days-old WT and Les8 plants. After drying at 37 °C, these leaf materials were ground into power. 60–70 mg powder was incubated with 1.5 ml 70% ethanol in a 2 ml centrifuge tube and centrifuged at 10,000 rpm min−1 for 10 min. After 2 times, the pellet was resuspended and incubated with chloroform: methanol (1:1) three times; then, the residues were resuspended and extracted with acetone until colorless supernatant was observed. The resulting pellet was dried at 35 ℃. The G lignin and S lignin monomers were measured using the thioacidolysis method, as previously described (Gou et al. 2019).
Quantification of JA, SA, and their derivatives
To detect the content of hormones in WT and Les8, we collected uniform leaves from 40-days-old plants. SA and JA were extracted and quantitatively analyzed by MetWare (http://www.metware.cn/) using the AB Sciex 20QTRAP 6500 LC–MS/MS platform as previously described by Guo et al. (2021).
Statistical analysis
Significant differences between means were determined using Student’s t test. A Chi-square test (χ2) was used for genetic analysis.
Results
Our previous studies indicated that lesion mimic mutants are valuable resources for dissecting the mechanism of disease resistance in maize (Mu et al. 2021; Li et al. 2022b). We thus screened a series of Les mutants to obtain mutants that confer MDR. Among which, Les8 showed elevated resistance to curvularia leaf spot (CLS) and southern leaf blight (SLB) and was thus characterized in detail in this study.
Les8 is a typical light-dependent lesion mimic mutant
Under field conditions, Les8 mutant plants showed nearly normal growth and development in addition to slight chlorotic but non-necrotic lesions on the leaves (Fig. 1a–c). Chlorotic lesions of Les8 first initiate on the tips of the field-grown mutant leaves at ~ 4 weeks old, then expand toward the base of younger tissue (Fig. 1a–c). Consistent with the formation of the lesions in the leaves, the content of chlorophyll a and chlorophyll b was significantly lower in the Les8 mutant compared to that of WT (Fig. 1d). Previous studies showed the formation of lesions is usually subjected to exposure to light (Gray et al. 2002; Li et al. 2022b). We therefore covered the newly emerged leaves of Les8 and WT to study the effect of light on lesion formation. We found that lesions on the covered Les8 leaf surface were absent, whereas lesions developed normally on the adjacent uncovered leaf areas, suggesting the formation of lesions in this mutant was light-dependent (Fig. 1e). We examined the accumulation of hydrogen peroxide, one of the major signaling molecules that trigger defense response by diaminobenzidine (DAB) staining, and a higher accumulation of H2O2 was observed in Les8 leaves as compared to that in WT leaves (Fig. 1f), implying activated defense response in Les8.
Phenotypic and physiological characterization of Les8 mutant. a Phenotype of 30-day-old WT and Les8. The scale bar is 5 cm. b, c Representative images of natural growth leaves of WT (b) and Les8 (c). d Chlorophyll a and Chlorophyll b concentrations of the WT and Les8 leaves. e Phenotype of WT and Les8 mutant leaves covered with aluminum foil for 7 days. The scale bar is 3 cm. f Representative images of DAB staining for H2O2 in uninfected WT and Les8 leaves. The scale bar is 5 mm
Les8 confers enhanced resistance to CLS and SLB
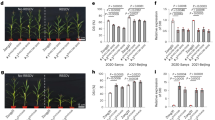
We investigated the disease resistance of Les8 to CLS and SLB, two major maize foliar diseases in China. Seven days after inoculation (DAI) with the C. lunata strain CX-3 under field conditions; a large number of fungal colonies were clearly observed on the leaves of WT, whereas very few disease lesions were observed on Les8 leaves harboring spontaneously formed chlorotic lesions (Fig. 2a–b). The resistance to CX-3 was further confirmed by lower amount of C. lunata DNA in the inoculated leaves of Les8 compared to the WT as detected by quantitative real-time PCR (Fig. 2c). In addition, Les8 showed enhanced resistance to SLB after natural infection under field condition (Fig. 2d). The disease lesion size and DNA amounts of B. maydis were both significantly reduced in the Les8 mutant compared to the WT (Fig. 2e–f). To further assess the resistance effect conferred by Les8 mutation, we generated the F2 population by crossing Les8 with the inbred line B73, and the plant height, disease resistance, and biomass were surveyed for plants with the WT and Les8 phenotype separately. While the plant height was not significantly changed (Figs. S1a, b), the resistance to CX-3 was significantly enhanced in plants of Les8 phenotype compared to that of WT phenotype (Fig. S1c). In addition, the biomass of CX-3 inoculated WT-looking plants was significantly lower than that of non-inoculated WT-looking plants. In contrast, the biomass of Les8-looking plants did not change significantly before and after CX-3 inoculation (Fig. S1d), indicating the potential value of Les8 mutation in the improvement of disease resistance.
Les8 mutant showed enhanced disease resistance to curvularia leaf spot and southern leaf blight in maize. a Representative images of WT and Les8 leaves at 7 days after inoculation with C. lunata. The scale bar is 3 cm. b Quantification of C. lunata colonies in WT and Les8 leaves 7 days after inoculation. Cfu, colony forming unit. (cfu) c Quantification of C. lunata DNA in WT and Les8 leaves 7 d after inoculation with C. lunata. d Representative images of WT and Les8 leaves after natural infection with B. maydis. The scale bar is 3 cm. e Percentage of B. maydis colonies in WT and Les8 leaves. f Quantification of B. maydis DNA in WT and Les8 leaves after natural infection. All data are means (± SD). Significant differences were determined using Student’s t test: *P < 0.05, **P < 0.01, ***P < 0.001
Fine mapping of LES8 gene
To investigate the genetic effect of Les8 mutation, we constructed five backcross populations by crossing Les8 to the maize inbred lines B73, Mo17, Zheng58, Chang7-2, and Qi319, respectively. All the BC1F1 mutant plants derived from the backcross populations have similar lesion mimic leaves as the Les8 mutant, demonstrating stable genetic effect of this mutation (Fig. S2). The F2 population of the B73 × Les8 displayed a 3:1 segregation, indicating that the lesion mimic phenotype of Les8 was controlled by a dominant monogenic mutation (Table S2). To determine the causal gene of Les8, BSR-seq analysis was conducted using the WT and mutant pools from the F2 population of B73 × Les8, and a causal locus was identified on chromosome 9 (Fig. 3a). Using 7735 F2 plants, the LES8 gene was mapped to a 909 kb region (Fig. 3b), and 38 protein-coding genes were found in this region according to the B73 RefGen_V4 reference genome (Table S3). We further investigated the expression level of these 38 genes according to our transcriptome analysis of Les8, and only 9 genes expressed highly in the leaves of Les8 (Table S3). We then considered these genes as high-confidence candidate genes of Les8. Among them, Zm00001d045335 and Zm00001d045359, encoding a putative NBS disease-resistance protein and mitogen-activated protein kinase, respectively, are probably related to maize disease resistance according to previous studies in arabidopsis (Meng and Zhang 2013; Wersch et al. 2020; Sun and Zhang 2022). Zm00001d045366 and Zm00001d045372 are potentially involved in the regulation of the redox state of proteins. Besides, Zm00001d045336, Zm00001d045339, Zm00001d045352, Zm00001d045358, and Zm00001d045361 were the other 5 genes that were highly expressed in the Les8. Interestingly, none of the genes were previously experimentally characterized as disease-resistance genes in maize, indicating that LES8 is a new gene controlling lesion mimic phenotype and MDR in maize.
Map-based cloning of Les8. a Bulked-segregant RNA-seq analysis of the causal locus in Les8. The dashed box indicates the causal locus on chromosome 9. b Fine mapping of the Les8 locus using 7735 F2 plants derived from B73 × Les8. The molecular markers and recombinants were listed above and below the black line, and the number of F2 plants was listed on the right of the black lines
Transcriptome analysis of Les8
To dissect the underlying mechanism of lesion formation and broad-spectrum resistance mediated by Les8 mutation, we performed transcriptome analysis of WT and Les8. After sequencing, a total of 143,889,995 clean reads were obtained from the resulting 6 RNA-seq libraries (Table S4). Pearson correlation coefficients varied from 0.888 to 0.982 among the six libraries indicating the high consistency among the biological replicates (Fig. S4). A total of 4571 differentially expressed genes (DEGs) were identified. Among them, 2636 genes were up-regulated, and 1935 genes were down-regulated in Les8 (Dataset S1). To confirm the reliability of our sequencing data, we performed qRT-PCR on 10 randomly selected genes; the results matched well with the RNA-seq data (Fig. S4).
We then performed GO analysis of the DEGs to dissect the possible molecular pathways affected by the Les8 mutation (Dataset S2). Consistent with the activated disease resistance in Les8, many defense-related GO terms including ‘defense to oomycetes,’ ‘cell surface receptor signaling,’ and ‘positive regulation to innate immune,’ were enriched in the up-regulated DEGs. In addition, genes related to defensive hormone biosynthesis and signaling, reactive oxygen species (ROS) metabolism as well as secondary metabolism were also markedly enriched, implying broadly activated defense response in Les8 (Fig. 4). In contrast, genes involved in photosynthesis and carbohydrate metabolisms were significantly enriched in down-regulated DEGs (Fig. 4), which were in line with the significantly decreased content of chlorophyll in the Les8 mutant (Fig. 1D).
Given the MDR of Les8, we looked in detail at the defense-related genes of the DEGs. Based on the enriched GO terms, 84 DEGs were identified as defense-related genes (Fig. 4, Dateset S3). We compared these genes to previously published pathogen-treated transcriptomics data using Plant Regulomics (Ran et al. 2020), and 48 genes were shared with genes previously reported as being responsive to pathogen infection (Dataset S4), demonstrating that defense responses were similarly activated in uninfected Les8 mutant as in pathogen-infected plants. Transcription factors (TFs) have previously been reported to play a vital role in regulating plant disease resistance, especially WRKYs and MYBs (Erpen et al. 2018). According to the prediction of the Plant Transcription Factor Database (Jin et al. 2016), 35 and 34 genes encoding MYB and WRKY TFs, respectively, were found differentially expressed in Les8 (Dataset S5). Intriguingly, we found all the 34 WRKYs were up-regulated Les8. Among them, three WRKYs (Zm00001d038451, Zm00001d012482, Zm00001d043025) were reported to be responsive to Fusarium ear rot infection (Lanubile et al. 2017; Liao et al. 2023).
Differential accumulation of jasmonic acid and salicylic acid in Les8 mutant
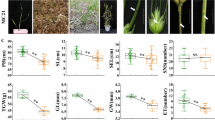
Given the enhanced disease resistance and significantly enriched GO terms of hormones, we quantified the major hormones involved in plant defense: jasmonic acid (JA), salicylic acid (SA), and their derivatives (Dataset S6). Intriguingly, the content of JA and its derivatives JA-Ile, H2JA, JA-Val, and 12-OH-JA was significantly increased in Les8 mutant compared to that of WT (Fig. 5a), whereas the content of free SA and its conjugated form (SAG) was significantly decreased in Les8 compared to that of WT (Fig. 5b). Consistently, genes involved in JA biosynthesis and signaling (ZmLOX, ZmAOS, ZmAOC, ZmOPR and ZmMYC7) were obviously up-regulated, and the SA biosynthesis-related gene ZmICS1 was significantly down-regulated (Fig. S5).
Quantification of JA, SA, and their derivatives in WT and Les8. a The concentration of JA, its derivatives JA-Ile, H2JA, JA-Val, and 12-OH-JA in WT and Les8. The red asterisks indicate that the compounds could not be detected in WT. b The SA and conjugated SA (SA β-glucoside, SAG) concentration in WT and Les8. JA-Ile, Jasmonoyl-L-isoleucine; H2JA, Dihydrojasmonic acid; JA-Val, N-[(-)-Jasmonoyl]-(L)-valine, 12-OH-JA, 12-Hydroxyjasmonic acid. All data are means (± SD). Significant differences were determined using Student’s t test: *P < 0.05, **P < 0.01, ***P < 0.001 (color figure online)
Transcriptomics analysis of genes involved in the biosynthesis of lignin, zealexins and kauralexins
Based on the GO enrichment analysis of DEGs, we wonder if Les8-conferred MDR is attributed to the biosynthesis of defense-related secondary metabolites, such as lignin, zealexins, and kauralexins, which were reported as the major phytoalexins in maize (Yang et al. 2017; Ding et al. 2020). According to the transcriptomic data, most genes responsible for lignin biosynthesis were up-regulated in Les8 compared to the WT (Fig. 6a, Dataset S7). In general, genes encoding lignin biosynthetic enzymes, including phenylalanine ammonialyase (PAL), cinnamoyl CoA reductase (CCR), caffeic acid O-methyltransferase (COMT), and laccase (LAC), were significantly up-regulated. In addition, MYB152 (Zm00001d021296), which was proposed to control the expression of maize PAL genes in a previous study (Zhang et al. 2016), was up-regulated in Les8. Given the general up-regulation of the lignin biosynthetic genes, we then quantified the content of G and S lignin monomer in Les8 and WT. While G lignin was not significantly changed, S lignin level was dramatically increased in Les8 leaves compared to that in WT (Fig. 6). As the major antibiotic sesquiterpenoids and diterpenoids in maize, zealexins and kauralexins' important role in plant disease resistance has been reported (Ding et al. 2020). We found the expressions of all the biosynthetic genes of zealexins and kauralexins were highly up-regulated in Les8 mutant compared to the WT, implying activated production of the two important antibiotics in Les8 (Fig. 6b–c).
Transcriptomic analysis of genes involved in lignin, zealexin, and kauralexin biosynthesis in Les8. a Transcriptomic analysis of gene expression in the lignin biosynthesis pathway. b Transcriptomic analysis of gene expression in the zealexins biosynthesis pathway. c Transcriptomic analysis of gene expression in the kauralexins biosynthesis pathway. The heat maps show the log2(fold-change) expression of the different homologous genes. PAL, phenylalanine ammonialyase; C4H, cinnamate 4-hydroxylase; 4CL, 4-coumarate CoA ligase; HCT, hydroxycinnamoyl; C3H, p-coumaroyl shikimate 3-hydroxylase; CSE, caffeoyl shikimate esterase; CCoAOMT, caffeoyl CoA O-methyltransferase; CCR, cinnamoyl CoA reductase; CAD, cinnamyl alcohol dehydrogenase; F5H, ferulate 5-hydroxylase; COMT, caffeic acid O-methyltransferase; LAC, laccase; ZX, zealexin; AN2, anther ear 2; KSL, kaurene synthase-like, CYP, cytochrome P450, KO2, kaurene oxidase 2
Since genes involved in antibiotic biosynthesis were up-regulated in Les8, we wondered if the enhanced disease resistance of Les8 was due to the antibiotic compounds generated in the mutant. We thus checked the effect of the leaf extracts on the growth of CX-3 strain. The diameter of the CX-3 colonies incubated on PDA plates supplemented with the Les8 extracts was significantly smaller than that with WT extracts at both 48 and 96 h after incubation (Fig. 7a–c). In addition, significantly decreased number of fungal spores was observed when CX-3 was incubated on PDA plates supplemented with Les8 extracts as compared to that with WT extracts, confirming the inhibitory effect of antibiotic compounds generated in Les8 (Fig. 7d).
Growth assay of CX-3 strain on PDA plate supplemented with leaf extraction of WT and Les8. a, b Phenotype of CX-3 strain grown on PDA plate supplemented with the fresh leaf extraction of WT (left) and Les8 (right) at 48 h a and 90 h b after incubation. c Quantification of the diameter of the CX-3 colony shown in A and B. d Quantification of the spore number of the CX-3 strain shown in a and b. Significant differences were determined using Student’s t test: ***P < 0.001
Discussion
Les mutants are valuable genetic resources for crop improvement to generate MDR varieties. In this study, we characterized a novel Les mutant (Les8) that confers MDR to both CLS and SLB in maize. When Les8 was crossed to the inbred line B73, enhanced disease resistance was observed in the Les8-looking plants of the F2 progeny. The Les8 phenotype is also stable in different genetic backgrounds including Mo17, Zheng58, Chang7-2, and Qi319 (Supplementary Fig. S1), indicating that Les8 is a valuable germplasm to be used for MDR improvement in maize. By BSR-seq and map-based cloning, the LES8 gene was mapped to a 909 kb interval and 9 high-confidence genes were identified. Transcriptomic analysis showed that defense-related genes were significantly up-regulated, while photosynthesis biosynthesis-related genes were significantly down-regulated in Les8. Our data support that differential accumulation of JA, SA and their derivatives as well as overaccumulation of S lignin and other phytoalexins in Les8 are responsible for Les-conferred MDR.
Les8 is a novel dominant MDR mutant valuable for maize breeding
According to our data, there are 38 protein-coding genes located in the candidate region. Among them, 9 high-confident genes were preliminarily identified based on the transcriptomics data (Table. S3). Based on current gene annotation, we found two genes are likely involved in Les8-conferred disease resistance. Zm00001d045335 encodes a putative NBS disease-resistance protein, which was known as the most common disease-resistance protein in plants (Wersch et al. 2020). Zm00001d045359 encodes the mitogen-activated protein kinase, which works as an important player in the three-kinase cascades and is responsible for converting membrane receptor signals and activating downstream immunity response when pathogens invade (Meng and Zhang 2013; Sun and Zhang 2022). It is worth noting that, although only 9 highly expressed genes were identified, the other 29 genes cannot be overlooked due to the limited ability of RNA-seq to detect the expressed genes. Besides, because our present prediction of candidate genes completely relies on the B73 reference genome, further refining and sequencing the region in the WT and Les8 is necessary, and the different haplotypes from the mutant also need to be considered to ultimately confirm the responsible gene.
Overaccumulation of JA and secondary metabolites are likely responsible for Les8-conferred MDR
In this study, we observed an overaccumulation of JA and its derivatives, but a dramatic decrease in SA and SAG. It was known that JA is implicated in resistance to necrotrophic pathogens, while SA is responsible for resistance to biotrophic pathogens (Yang et al. 2015; Andersen et al. 2018; Howe et al. 2018). These data are consistent with our observation that Les8 is resistant to the necrotrophic pathogen Bipolaris maydis and the hemibiotrophic pathogen Curvularia lunata which also has a necrotrophic life phase (Fig. S5). Therefore, Les8-conferred resistance to CLS and SLB is largely due to activation of JA biosynthesis and inhibition of SA biosynthesis. Besides, GO analysis of the up-regulated genes in Les8 revealed that genes involved in second metabolites were highly enriched (Fig. 4). Specially, genes involved in phenylpropanoid and lignin biosynthesis, such as PALs, 4CLs, CADs, LACs, were generally up-regulated (Fig. 6a). Consistently, S lignin accumulation was dramatically increased in Les8 (Fig. S6). Our study further highlights previous report that phenylpropanoid and lignin biosynthesis are critical for in disease resistance of maize (Wang et al. 2015; Wang and Balint-Kurti 2016; Yang et al. 2017; Li et al. 2019). Besides, genes involved in the biosynthesis of zealexins and kauralexins were also mostly up-regulated in Les8 (Fig. 6b–d). Moreover, we observed a significant inhibition of CX-3 growth by co-incubation with leaf extracts of Les8 (Fig. 7), confirming that Les8-conferred resistance to CLS and SLB is largely due to overaccumulation of the antibiotic secondary metabolites.
Data availability
All data generated or analyzed during this study are available within the article/supplementary files. The plant materials and datasets are available from the corresponding authors upon reasonable request. The sequencing data from the article can be found in the national genomic data center (https://ngdc.cncb.ac.cn/) under the following accession number: CRA013433.
References
Andersen E, Ali S, Byamukama E, Yen Y, Nepal M (2018) Disease resistance mechanisms in plants. Genes 9:339
Büschges R, Hollricher K, Panstruga R, Simons G, Wolter M, Frijters A, van Daelen R, van der Lee T, Diergaarde P, Groenendijk J et al (1997) The barley Mlo gene: a novel control element of plant pathogen resistance. Cell 88:695–705
Chaikam V, Negeri A, Dhawan R, Puchaka B, Ji J, Chintamanani S, Gachomo E, Zillmer A, Doran T, Weil C et al (2011) Use of mutant-assisted gene identification and characterization (MAGIC) to identify novel genetic loci that modify the maize hypersensitive response. Theor Appl Genet 123:985–997
Chen D, Ronald P (1999) A rapid DNA minipreparation method suitable for AFLP and other PCR applications. Plant Mol Biol Report 17:53–57
Chintamanani S, Hulbert S, Johal G, Balint-Kurti P (2010) Identification of a maize locus that modulates the hypersensitive defense response, using mutant-assisted gene identification and characterization. Genetics 184:813–825
Ding Y, Weckwerth P, Poretsky E, Murphy K, Sims J, Saldivar E, Christensen S, Char SN, Yang B, Tong A (2020) Genetic elucidation of interconnected antibiotic pathways mediating maize innate immunity. Nature Plants 6:1375–1388
Erpen L, Devi H, Grosser J, Dutt M (2018) Potential use of the DREB/ERF, MYB, NAC and WRKY transcription factors to improve abiotic and biotic stress in transgenic plants. Plant Cell, Tissue Organ Cult 132:1–25
Gou M, Yang X, Zhao Y, Ran X, Song Y, Liu CJ (2019) Cytochrome b5 is an obligate electron shuttle protein for syringyl lignin biosynthesis in arabidopsis. Plant Cell 31:1344–1366
Gray J, Close P, Briggs S, Johal G (1997) A novel suppressor of cell death in plants encoded by the Lls1 gene of maize. Cell 89:25–31
Gray J, Janick-Buckner D, Buckner B, Close PS, Johal GS (2002) Light-dependent death of maize lls1 cells is mediated by mature chloroplasts. Plant Physiol 130:1894–1907
Guo Q, Li X, Niu L, Jameson P, Zhou W (2021) Transcription-associated metabolomic adjustments in maize occur during combined drought and cold stress. Plant Physiol 186:677–695
Howe G, Major I, Koo A (2018) Modularity in jasmonate signaling for multistress resilience. Annu Rev Plant Biol 69:387–415
Hu G, Richter T, Hulbert S, Pryor T (1996) Disease lesion mimicry caused by mutations in the rust resistance gene rp1. Plant Cell 8:1367–1376
Hu G, Yalpani N, Briggs SP, Johal GS (1998) A porphyrin pathway impairment is responsible for the phenotype of a dominant disease lesion mimic mutant of maize. Plant Cell 10:1095–1105
Jin J, Tian F, Yang D, Meng Y, Kong L, Luo J, Gao G (2016) PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res 45:D1040–D1045
Johal G, Hulbert S, Briggs S (1995) Disease lesion mimics of maize: a model for cell death in plants. BioEssays 17:685–692
Karre S, Kim S, Samira R, Balint-Kurti P (2021) The maize ZmMIEL1 E3 ligase and ZmMYB83 transcription factor proteins interact and regulate the hypersensitive defence response. Mol Plant Pathol 22:694–709
Kim D, Langmead B, Salzberg S (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360
Lanubile A, Maschietto V, Borrelli V, Stagnati L, Logrieco A, Marocco A (2017) Molecular basis of resistance to Fusarium ear rot in maize. Front Plant Sci 8:1774
Li N, Lin B, Wang H, Li X, Yang F, Ding X, Yan J, Chu Z (2019) Natural variation in ZmFBL41 confers banded leaf and sheath blight resistance in maize. Nat Genet 51:1540–1548
Li J, Chen M, Fan T, Mu X, Gao J, Wang Y, Jing T, Shi C, Niu H, Zhen S et al (2022b) Underlying mechanism of accelerated cell death and multiple disease resistance in a maize lethal leaf spot 1 allele. J Exp Bot 73:3991–4007
Li Y, Gu J, Ma S, Xu Y, Liu M, Zhang C, Liu X, Wang G (2023) Genome editing of the susceptibility gene ZmNANMT confers multiple disease resistance without agronomic penalty in maize. Plant Biotechnol J 21:1525
Li C, Liu H, et al (2022a) Characterization and fine mapping of a lesion mimic mutant (Lm5) with enhanced stripe rust and powdery mildew resistance in bread wheat (Triticum aestivum L.). Theor Appl Genet pp 1–18
Li S, Lin D, et al. (2022c) Genome-edited powdery mildew resistance in wheat without growth penalties. Nature, pp 1–6
Liao X, Sun J, Li Q, Ding W, Zhao B, Wang B, Zhou S, Wang H (2023) ZmSIZ1a and ZmSIZ1b play an indispensable role in resistance against Fusarium ear rot in maize. Mol Plant Pathol 24:711–724
Liu S, Yeh CT, Tang HM, Nettleton D, Schnable PS (2012) Gene mapping via bulked segregant RNA-Seq (BSR-Seq). PLoS ONE 7:e36406
Liu Q, Ning Y, Zhang Y, Yu N, Zhao C, Zhan X, Wu W, Chen D, Wei X, Wang G (2017) OsCUL3a negatively regulates cell death and immunity by degrading OsNPR1 in rice. Plant Cell 29:345–359
Lorrain S, Vailleau F, Balagué C, Roby D (2003) Lesion mimic mutants: keys for deciphering cell death and defense pathways in plants? Trends Plant Sci 8:263–271
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:1–21
Luan Q, Zhu Y, Ma S, Sun Y, Liu X, Liu M, Balint-Kurti P, Wang G (2020) Maize metacaspases modulate the defense response mediated by the NLR protein Rp1-D21 likely by affecting its subcellular localization. Plant J 105:151–166
Meng X, Zhang S (2013) MAPK cascades in plant disease resistance signaling. Annu Rev Phytopathol 51:245–266
Mu X, Li J, Dai Z, Xu L, Fan T, Jing T, Chen M, Gou M (2021) Commonly and specifically activated defense responses in maize disease lesion mimic mutants revealed by integrated transcriptomics and metabolomics analysis. Front Plant Sci 12:690
Neuffer M, Calvert O (1975) Dominant disease lesion mimics in maize. J Hered 66:265–270
Olukolu B, Wang G, Vontimitta V, Venkata B, Marla S, Ji J, Gachomo E, Chu K, Negeri A, Benson J, Nelson R et al (2014) A genome-wide association study of the maize hypersensitive defense response identifies genes that cluster in related pathways. PLoS Genet 10:e1004562
Piffanelli P, Ramsay L, Waugh R, Benabdelmouna A, D’Hont A, Hollricher K, Jørgensen JH, Schulze-Lefert P, Panstruga R (2004) A barley cultivation-associated polymorphism conveys resistance to powdery mildew. Nature 430:887–891
Qiu N, Wang X, Yang F, Yang X, Yang W, Diao R, Wang X, Cui J, Zhou F (2016) Fast extraction and precise determination of chlorophyll. Chin Bull Botany 51:667
Ran X, Zhao F, Wang Y, Liu J, Zhuang Y, Ye L, Qi M, Cheng J, Zhang Y (2020) (2020) Plant Regulomics: a data-driven interface for retrieving upstream regulators from plant multi-omics data. Plant J 101:237–248
Sha G, Sun P, et al. (2023) Genome editing of a rice CDP-DAG synthase confers multipathogen resistance. Nature, pp 1–7
Simmons C, Hantke S, Grant S, Johal G, Briggs S (1998) The maize lethal leaf spot 1 mutant has elevated resistance to fungal infection at the leaf epidermis. Mol Plant Microbe Interact 11:1110–1118
Sun T, Zhang Y (2022) MAP kinase cascades in plant development and immune signaling. EMBO Rep 23:e53817
Walbot V, Hoisington DA, Neuffer M (1983) Disease lesion mimic mutations. Genetic Eng Plants, pp 431–442
Wang GF, Balint-Kurti P (2016) Maize homologs of CCoAOMT and HCT, two key enzymes in lignin biosynthesis, form complexes with the NLR Rp1 protein to modulate the defense response. Plant Physiol 171:2166–2177
Wang GF, He Y, Strauch R, Olukolu BA, Nielsen D, Li X, Balint-Kurti PJ (2015) Maize homologs of hydroxycinnamoyltransferase, a key enzyme in lignin biosynthesis, bind the nucleotide binding leucine-rich repeat Rp1 proteins to modulate the defense response. Plant Physiol 169:2230–2243
Wang H, Hou J, Ye P, Hu L, Huang J, Dai Z, Zhang B, Dai S, Que J, Min H et al (2021) A teosinte-derived allele of a MYB transcription repressor confers multiple disease resistance in maize. Mol Plant 14:1846–1863
Wersch R, Li X, Zhang Y (2016) Mighty dwarfs: arabidopsis autoimmune mutants and their usages in genetic dissection of plant immunity. Front Plant Sci 7:1717
Wersch S, Tian L, Hoy R, Li X (2020) Plant NLRs: the whistleblowers of plant immunity. Plant Commun 1:100016
Yang L, Li B, Zheng X, Li J, Yang M, Dong X, He G, An C, Deng X (2015) Salicylic acid biosynthesis is enhanced and contributes to increased biotrophic pathogen resistance in Arabidopsis hybrids. Nat Commun 6:7309
Yang Q, He Y, Kabahuma M, Chaya T, Kelly A, Borrego E, Bian Y, El Kasmi F, Yang L, Teixeira P et al (2017) A gene encoding maize caffeoyl-CoA O-methyltransferase confers quantitative resistance to multiple pathogens. Nat Genet 49:1364–1372
Young M, Wakefield M, Smyth G, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11:R14
Yu M, Fan Y, Li X, Chen X, Yu S, Wei S, Lu K (2023) LESION MIMIC MUTANT 1 confers basal resistance to Sclerotinia sclerotiorum in rapeseed via a salicylic acid-dependent pathway. J Exp Botany 74(18):5620–5634
Zhang Y, Goritschnig S, Dong X, Li X (2003) A gain-of-function mutation in a plant disease resistance gene leads to constitutive activation of downstream signal transduction pathways in suppressor of npr1-1, constitutive 1. Plant Cell 15:2636–2646
Zhang J, Zhang S, Li H, Du H, Huang H, Li Y, Hu Y, Liu H, Liu Y, Yu G et al (2016) Identification of transcription factors ZmMYB111 and ZmMYB148 involved in phenylpropanoid metabolism. Front Plant Sci 7:148
Zhao Y, Xu W, Wang L, Han S, Zhang Y, Liu Q, Liu B, Zhao X (2022) A maize necrotic leaf mutant caused by defect of coproporphyrinogen III oxidase in the porphyrin pathway. Genes 13:272
Zhu X, Zhu Ze M, Chern M, Chen X, Wang J (2020) Deciphering rice lesion mimic mutants to understand molecular network governing plant immunity and growth. Rice Sci 27:278–288
Acknowledgements
We thank the Maize Genetics Cooperation Stock Center for kindly supplying the Les8-N2005 mutant for this study. We thank Dr. Chunsheng Xue of Shenyang Agricultural University for providing the Curvularia lunata strain CX-3. We thank the National Natural Science Foundation of China (U2004207 to MG), National Key Research and Development Program of China (2022YFD1201801, to MG), the Fund for Distinguished Young Scholars in Henan (212300410007, to MG), Henan Province Joint Fund for Science and Technology Research (222103810003, to JL), Henan Province Major Science and Technology Project (221100110300, to JT), for the funding for this study.
Funding
This work was supported by the National Natural Science Foundation of China (U2004207, to MG), National key research and development program of China (2022YFD1201801, to MG), the Fund for Distinguished Young Scholars in Henan (212300410007, to MG), Henan Province Joint Fund for Science and Technology Research (222103810003, to JL), Henan Province Major Science and Technology Project (221100110300, to JT).
Author information
Authors and Affiliations
Contributions
MG, JK and HN designed the experiments; JL and TF performed most of the experiments and analyzed the data, YZ and MC worked on the RNA extraction and qRT-PCR; YW, JG, NZ, and JT worked on the map-based cloning; JT evaluated the phenotype of F2 population and provided critical advice on gene mapping; CZ and HN performed the pathogen test and phenotypic identification; YZ and XM measured the lignin content. JL and TF analyzed the transcriptional data; SZ and JF performed the BSR-seq analysis; JL and MG wrote the manuscript; all authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests about this work.
Ethical approval
All experiments and data analyses were conducted according to the current laws of the country. The manuscript has not been submitted to any other journal.
Additional information
Communicated by Mingliang Xu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Fan, T., Zhang, Y. et al. Characterization and fine mapping of a maize lesion mimic mutant (Les8) with enhanced resistance to Curvularia leaf spot and southern leaf blight. Theor Appl Genet 137, 7 (2024). https://doi.org/10.1007/s00122-023-04511-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-023-04511-x