Abstract
Key message
A major QTL controlling ovule abortion and SN was fine-mapped to a 80.1-kb region on A8 in rapeseed, and BnaA08g07940D and BnaA08g07950D are the most likely candidate genes.
Abstract
The seed number per silique (SN), an important yield determining trait of rapeseed, is the final consequence of a complex developmental process including ovule initiation and the subsequent ovule/seed development. To explore the genetic mechanism regulating the natural variation of SN and its related components, quantitative trait locus (QTL) mapping was conducted using a doubled haploid (DH) population derived from the cross between C4-146 and C4-58B, which showed significant differences in SN and aborted ovule number (AON), but no obvious differences in ovule number (ON). QTL analysis identified 19 consensus QTLs for six SN-related traits across three environments. A novel QTL on chromosome A8, un.A8, which associates with multiple traits, except for ON, was stably detected across the three environments. This QTL explained more than 50% of the SN, AON and percentage of aborted ovules (PAO) variations as well as a moderate contribution on silique length (SL) and thousand seed weight (TSW). The C4-146 allele at the locus increases SN and SL but decreases AON, PAO and TSW. Further fine mapping narrowed down this locus into an 80.1-kb interval flanked by markers BM1668 and BM1672, and six predicted genes were annotated in the delimited region. Expression analyses and DNA sequencing showed that two homologs of Arabidopsis photosystem I subunit F (BnaA08g07940D) and zinc transporter 10 precursor (BnaA08g07950D) were the most promising candidate genes underlying this locus. These results provide a solid basis for cloning un.A8 to reduce the ovule abortion and increase SN in the yield improvement of rapeseed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rapeseed (Brassica napus L., 2n = 38, AACC) is one of the most important oil crops worldwide that provide more than 15% of the global supply of edible vegetable oil (USDA ERS 2019) and also used as an important raw material for biofuel and feeding livestock. Developing high-yielding varieties has always been one of the most important goals of rapeseed breeding because of the growing demands for vegetable oil as well as the declining hectarage used for rapeseed production (Sun et al. 2018). The seed number per silique (SN) is one of the three direct yield determining traits (including silique number and seed weight) of rapeseed and has always received much attention (Fan et al. 2010). In general, a highly significant positive correlation was observed between SN and seed yield, indicating that increasing SN should be effective for improving rapeseed production. Although there is a rich variation for SN in rapeseed, the SN of the current cultivars (~ 20) is far below its maximum (> 30) in the germplasm resources (Chen et al. 2012), indicating that the potential of SN is yet to be maximized effectively in rapeseed yield improvement. Additionally, higher SN plants are thought to have superior colonization abilities because they produce more offsprings/seeds, which were more likely to be selected during crop domestication (Moles et al. 2005). Therefore, understanding the genetic basis of SN is of great importance not only for yield improvement through the utilization of advantageous variations in rapeseed breeding but also for plant evolution and crop domestication.
In the family Brassicaceae, the ovules upon fertilization give rise to the seeds (Van Went and Cresti 1988). The ovule number decides the putative maximal number of seeds in a silique. However, in many plants including rapeseed, the ovule number is much larger than the final seed number in a fruit (Mendham et al. 1981; Charlesworth 1989; Wang et al. 2011). Previous studies indicated that the formation of seeds, a complex process, is often determined by its components, including the ovule number per ovary (ON), the ratio of fertile/unfertile ovule and the proportion of fertilized ovules that developed into seeds (Yang et al. 2016; Khan et al. 2019). Thus, it is essential to understand the factors controlling ovule initiation and development from an agricultural and economical point of view, as they ultimately determine the final number of seeds and thereby influence the crop yield. Li et al. (2014) showed that SN has a highly significant correlation with the number of fertile ovules (r = 0.987), but not significantly correlated with the ovule number per ovary (ON), indicating that the normal development of ovule is an important factor affecting SN. Further cytological analyses revealed that part of the ovule’s primordium could not develop into mature seeds and the abortion of ovule occurred between the stages of megaspore degradation and uninucleate gametophyte (Li et al. 2014). Li et al. (2015) reported the map-based cloning of BnaC9.SMG7b on chromosome C9, a major positive regulator of SN in B. napus. BnaC9.SMG7b plays a role in regulating the formation of functional female gametophyte, thus determining the formation of fertile ovules (Li et al. 2015). Similarly, the cytological mechanism of a major QTL for SN (qSN.A6) was due to ovule abortion caused by incomplete cellularization of the embryo sac (Yang et al. 2016). These reports strongly showed that ovule abortion is most likely a key factor responsible for the natural variation of SN in rapeseed. Therefore, a deep understanding of the genetic basis of SN and its related components is of great importance for yield improvement in rapeseed breeding.
Typically, SN and its related components are complex quantitative traits controlled by polygenes and highly influenced by environmental conditions. As one of the important yield-related traits, SN has been extensively studied using quantitative trait locus (QTL) and genome-wide association study (GWAS) analysis in rapeseed. More than 100 QTLs distributed on all the 19 chromosomes of rapeseed genome had been identified for SN, with only 4 of them (located on A1, A6, C1 and C9 chromosomes, respectively) showing major effects and exhibiting phenotypic variances ranging from 23.5% to 57.8% individually (Radoev et al. 2008; Shi et al. 2009, 2015; Zhang et al. 2011, 2012; Chen et al. 2011; Cai et al. 2014a; Qi et al. 2014; Yang et al. 2016). The complex genetic basis of SN in rapeseed indicates that many genes are involved in the developmental/biological processes influencing seeds formation. Although many QTLs have been identified, only one gene underlying a major QTL controlling SN on C9, BnaC9.SMG7b, has been cloned by map-based strategy in B. napus (Li et al. 2015). qSN.A6, another major QTL for SN in rapeseed, was successfully fine mapped to a 267-kb region on chromosome A6 (Yang et al. 2016). Due to their increasing effect on SN, both BnaC9.SMG7b and qSN.A6 could increase the final seed yield by a proportion of 58.7% and 17.4%, respectively, which demonstrated their great potential and utilization in the yield improvement of rapeseed (Li et al. 2015; Yang et al. 2016). Recently, Khan et al. (2020) reported the functional characterization of BnaEOD3 gene by CRISPR/Cas9-mediated targeted mutations in rapeseed with high efficiency. SN and seed yield per plant of the mutants with combined loss-of-function alleles across four BnaEOD3 copies (including BnaA04g27100D, BnA05g01200D, BnaC04g00760D and BnaC04g50960D) showed a significant increase in varying degrees compared with that of wild-type, indicating that the quantitative involvement of the different BnaEOD3 copies affects SN (Khan et al. 2020). To date, no QTL mapping study on other SN-related traits has been reported yet in rapeseed, except for the GWAS study on ON reported by our group (Khan et al. 2019). As expected, ON was significantly positively correlated with SN, but not correlated with TSW, indicating that higher ON plants have the potential to increase SN (Khan et al. 2019). A total of 19 significant associated loci with minor effects were detected for ON, indicating a complex genetic basis of ON in rapeseed (Khan et al. 2019). Thus, the genetic basis for the natural variation of SN and its related components is still poorly understood.
In our previous study, we revealed that ovule abortion was the most important determinant of variation in SN through the cytological analysis of a series of doubled haploid (DH) lines and cultivars in Brassica napus (Li et al. 2014). To remove the effect of ON variation, two lines C4-146 and C4-58B from a HJ-DH population, which showed significant differences in SN and aborted ovule number (AON), but no obvious differences in ON (Li et al. 2014), were selected as the parents to develop segregating population and identify QTLs controlling the natural variation of ovule abortion and SN. We identified a novel QTL on chromosome A8, which pleiotropically controls ovule abortion and SN with a major effect. Furthermore, we determined the genomic location of this locus via fine mapping using an F2 population and identified the candidate genes. These results provide a solid basis for cloning this locus to further elucidate the genetic basis and molecular mechanisms underlying SN determination in rapeseed.
Material and methods
Plant materials
The DH population consisting of 290 individual lines was generated from a microspore culture of the F1 cross between C4-146 × C4-58B. A random subset including 190 lines was used for the linkage map construction and QTL mapping. The F1 plants generated from the reciprocal crosses between the two parental lines (C4-146 and C4-58B) were used for genetic analysis. Subsequently, the F1 plant's was self-pollinated to produce the F2 population used for the QTL fine mapping.
Field trials and trait evaluation
The DH population together with the two parents were planted in the winter-growing seasons of 2015–2016, 2016–2017 and 2017–2018. F1 hybrids, together with their parental lines, were grown in the winter-growing seasons of 2015–2016, 2017–2018, 2018–2019 and 2019–2020. The field experiment followed a randomized complete block design with three replicates. Each line contains 11–12 plants per row, with a distance of 21 cm between each plant in a row and 30 cm between rows. Five plants growing uniformly from each plot were chosen for trait evaluation. The mean of each trait calculated from the 5 sampled plants in one plot was used to determine the plot performance.
For the F2 population, a total of 16,421 individuals were grown in the winter-growing seasons of 2016–2017 and 2017–2018 for recombinant screening. The recombinants for the target QTL region were self-pollinated, and approximately 20 progenies from each recombinant were grown in the next winter-growing seasons to evaluate their phenotypes. There were 11–12 plants per row, and the rows were 30 cm apart.
All materials were grown in winter-type rapeseed growing season on the experimental farm of Huazhong Agricultural University, Wuhan, China. The field management was conducted according to standard breeding practice.
SN-related traits, including ON, AON, PAO, SN, SL and TSW, were measured essentially as previously described (Khan et al. 2019).
To investigate the pollen fertility, the pollen viability of parents and F1 were investigated using acetic red dyeing, as described by Yang et al. (2017).
Molecular marker development and genetic map construction
Molecular markers, including insertion/deletion (INDEL) and SNP markers in the whole-genome region, were newly developed (Table S1) based on the re-sequencing data of the parental lines. All primer pairs were designed using Primer5.0 software and synthesized by Sangon Biological and Engineering Co. (Shanghai, China).
Linkage analysis was performed using Mapmaker/Exp3.0 (Lincoln et al. 1992), and the genetic distance was calculated using the Kosambi function.
QTL mapping and statistical analysis
QTLs were detected using the composite interval mapping procedure with the program QTL Cartographer V2.5 (Wang et al. 2004) at a threshold of LOD = 2.5.
Identified QTLs were integrated using QTL meta-analysis with BioMercator2.1 (Goffinet and Gerber 2000). The identified QTLs associated with each trait that was repeatedly detected in different environments were integrated into a consensus QTL and overlapping consensus QTLs of different traits were further integrated into unique QTL.
The broad-sense heritability (h2B) of each trait in DH population was calculated as h2B = σ2g/(σ2g + σ2gl/n + σ2e/nr), where σ2g is genotypic variance; σ2gl variance due to genotype by environment interaction; σ2e error variance; n number of environments; and r number of replications. The estimates of σ2g, σ2gl and σ2e were obtained from an analysis of variance with the environment considered as a random effect.
The statistical analysis of the phenotypes (correlation coefficient, multiple comparisons) was conducted using Excel 2013 and Statistix 8.0 software.
Sequence analysis of candidate genes
Sequencing primer pairs covering the promoter sequences, full-length genomic DNA, and coding sequences of the candidate genes were designed according to the re-sequencing data of the parental lines (Table S2). The PCR products were cloned into the pEASY-T (TransGen Biotech, Beijing, China) vector and then sequenced using Sanger method.
Expression analysis of candidate genes
Different plant tissues were collected from the parental lines, across five stages during ovary and silique development (1–2 mm, 2–4 mm and 4-5 mm pistil, 3d and 5d ovary after pollination). Pistils of different lengths were collected from the buds before flowering. After flowering, individual flower was tagged on the day of flowering; then, siliques were collected at 3d and 5d after pollination (DAP). RNA extraction, cDNA preparation, real-time PCR and relative quantification were performed following the approach previously described by Hu et al. (2018). The relative amount of PCR product that was amplified using the designed primer sets (listed in Table S2) was normalized to the reference gene BnaA10g06670D (BnaUBC10), as described by Hu et al. (2018).
Results
Phenotypic variation for SN-related traits in parents, F1 and DH population
The DH lines and their corresponding parents as well as F1 were grown in the winter-type growing seasons. The phenotypic performances of six SN-related traits across the years were investigated (Table 1; Fig. 1). The average SN of C4-146 (18.34 ± 0.96) was nearly two times that of C4-58B (9.12 ± 0.65) across all investigated environments. However, the ON of the two parents was not significant in the three environments (Table 1), suggesting that the decrease of SN in C4-58B mainly result from defects after ovule initiation. Also, C4-146 had higher SL, lower AON, PAO and TSW than C4-58B in different environments (Table 1). The SN of F1 plants from the reciprocal crosses was all significantly lower than C4-146 but similar to C4-58B (Table 1), indicating the dominant nature of low SN over high SN in the cross between C4-146 and C4-58B. The other SN-related traits of F1 plants from the reciprocal crosses also displayed a similar trend with SN (Table 1). The pollen fertility of parents and F1 were investigated by acetic red dyeing. The results showed that the pollen viability of both C4-146 (94.90 ± 1.75%) and C4-58B (97.98 ± 1.47%) was high with no significant differences, whereas the pollen viability of the F1 obtained from the cross between C4-146 × C4-58B was significantly lower than those of their parents (Fig. S1; Table S3). Considering the similar phenotype of SN observed in both C4-58B and F1 (C4-146 × C4-58B), the variation of the pollen viability may not be the major factor affecting SN in the present study.
Trait performance of the parents and their reciprocal crossing F1 hybrids. Observation of the ovule number a, the seeds of five siliques b and the mature siliques c of C4-146, C4-58B, F1 hybrid of C4-146 × C4-58B and F1 hybrid of C4-58B × C4-146 (from left to right). Scale bars = 1 mm a, 1 cm b, and 1 cm c, respectively
The phenotypic performances of SN-related traits in the DH population are shown in Table 2 and Fig. 2. Continuous distributions and transgressive segregations in the DH population suggested a quantitative inheritance pattern for these traits. The distributions of SN, AON and PAO deviated from normality and appeared to have a bimodal pattern, indicating the involvement of a major gene and some minor genes for each of these traits. SL, TSW and ON showed nearly normal distribution across all environments, indicating the quantitative inheritance of these traits in the present population. Two-way ANOVA of these traits across environments indicated that the genotype of each line (G), environment (E) and genotype–environment interactions (G × E) had significant effects , respectively, except for the environmental effect on PAO (Table S4). In this population, SN showed the highest broad-sense heritability (0.97), followed by TSW/PAO (0.91) and AON (0.85), suggesting that these traits are stably inherited (Table S4). While the SL and ON have broad-sense heritability of 0.69 and 0.66, respectively, indicating that SL and ON are relatively sensitive and plastic to environmental variations (Table S4).
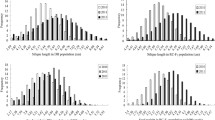
Frequency distribution of the seed number per silique(SN), the thousand seed weight (TSW), the silique length (SL), the ovule number per ovary (ON), the aborted ovule number per ovary (AON) as well as the percentage of aborted ovules (PAO) in the DH population at the indicated environment. DH lines carrying the C4-146 allele (AA) or the C4-58B allele (BB) were classified based on the genotype of two flanking markers BM1355 and BM1360
Significant phenotypic correlations were observed for most of these traits across three years (Table 3). The only exception is the ON trait which shows no significant correlations with SN, SL and TSW, indicating that ON is not influencing these three traits in this population. SN showed a highly negative correlation with PAO, AON and TSW, but had a highly positive correlation with SL, suggesting that the ovule abortion is a key negative regulator controlling SN and longer silique will produce more but smaller seeds.
Primary mapping of SN‑related QTLs in the DH population
Because the two parents C4-146 and C4-58B were selected from our previously reported HJ DH population which was used to construct a high-density genetic map (Cai et al. 2014b), we first analyzed the polymorphism distribution between C4-146 and C4-58B at the whole-genome scale using more than 2000 markers (Fig. S2). Subsequently, 179 molecular markers that covered all the polymorphic regions between the two parents were selected to genotype the DH population. A genetic map containing 17 chromosomes spanning a total of 684 cM of the B. napus genome was constructed, with the lack of chromosomes A6 and C1 because of lack of polymorphism between the parents (Fig. 3).
The genetic linkage map and QTLs for six SN-related traits identified in the DH population in three environments. The red, blue, black, orange, purple and green lines, respectively, indicate the confidence intervals of the consensus QTLs related to SN, TSW, SL, ON, AON and PAO in the DH population. The solid circles represent the positions of the QTL peak (color figure online)
A total of 40 QTLs for six examined traits were identified in the three environments with LOD values ranging from 2.78 to 48.48, which were distributed on 12 chromosomes (Table S5; Fig. 3). The phenotypic variation explained by individual QTL varied between 2.43% and 56.57% (Table S5). These identified QTLs were integrated into 19 reproducible consensus QTLs for each trait by QTL meta-analysis (Table 4). Among the identified QTLs, 6, 3, 4, 2, 1 and 3 consensus QTLs were detected for SL, SN, TSW, ON, AON and PAO, respectively. Eight consensus QTLs, including cqSL.A8, cqSN.A8, cqTSW.A7-1, cqTSW.A7-2, cqTSW.A8, cqON.C6-2, cqAON.A8 and cqPAO.A8, showed main effects and the others exhibited minor effects. Positive alleles for SN and ON and negative alleles for TSW, AON and PAO both originated from the C4-146 parental line while positive alleles for SL were dispersed between the two parents, in consistency with the trait performance of the two parents (Table S5; Table 4).
It is worth noting that cqSN.A8, flanked by BM1355 and BM1360 on chromosome A8, was a newly identified QTL and had the largest LOD scores (from 35.60 to 48.48) and explained as much as 56.57% of the SN variation in all environments. To evaluate the effect of cqSN.A8, the DH lines were classified into two groups (AA and BB) based on the genotype of the two flanking markers BM1355 and BM1360. Lines carrying the C4-146 alleles (AA) had an average SN of 19.20 ± 2.04, 18.45 ± 2.46 and 20.42 ± 2.26 in 2016, 2017 and 2018 environments, respectively, which were significantly higher than those carrying the C4-58B alleles (BB) with an average SN of 11.99 ± 2.19, 12.5 ± 2.25 and 12.88 ± 2.25 across the three environments. Clearly, the two genotypes of the DH lines at the cqSN.A8 locus corresponded exactly to the two distinct phenotypic classes, exhibiting bimodal distribution with 15–18 SN as the boundary (Fig. 2). Similarly, two consensus QTLs on chromosome A8 (cqAON.A8 and cqPAO.A8) also displayed a large effect (which explained about half of the trait variation in all environments), justifying the bimodal distribution exhibited by the two traits in the DH population (Fig. 2). The remaining two major consensus QTLs on chromosome A8 (cqSL.A8 and cqTSW.A8), which explained 36.74% and 15.82% of the phenotypic variance of SL and TSW on average, exhibited relatively minor effects, consistent with the nearly normal distribution of these traits in the DH population (Fig. 2). Interestingly, the confidence intervals of the five consensus QTLs on chromosome A8 for SN, AON, PAO, SL and TSW overlapped perfectly (Table 4).
Further meta-QTL analyses integrated these overlapped QTL into a unique QTL un.A8 (Table 5), which has a major pleiotropic effect on all these traits except for ON. The C4-146 alleles at un.A8 causes an increase in SN and SL and a simultaneous decrease in AON, PAO and TSW, respectively, justifying the significant correlations observed among these traits. Additionally, another two other unique QTLs were identified on chromosomes A4 and A7 (Table 5). The unique QTL un.A7 showed a major effect on TSW but had a minor effect on SN, SL and PAO. Also, the C4-146 alleles at un.A7 causes an increase in SN and SL and a simultaneous decrease in PAO and TSW, which is consistent with the correlations observed among these traits. For the minor unique QTL un.A4, the increasing allele for SN was inherited from C4-146 and the increasing allele for PAO was derived from C4-58B, reflecting the significant negative correlations between SN and PAO.
Fine mapping of the un.A8 locus
Due to the ease of assessment, SN was selected as the target trait for the fine mapping of un.A8 locus. To narrow down the genomic region containing un.A8, 24 polymorphic INDEL markers from the candidate region were developed and 18 recombinants between the BM1355 and BM1360 intervals in the DH population were screened. Based on the allelic composition and recombination breakpoints, the recombinants were grouped into three genotypes (Fig. 4a). The mean phenotypic value for each recombinant genotype was compared with the two parents to locate un.A8. Using these data, un.A8 was narrowed down to a region between BM1663 and BM1646, corresponding to an 820-kb region on chromosome A8 in the reference genome of B. napus cultivar Darmor-bzh (Chalhoub et al. 2014; Fig. 4a).
Fine mapping of the un.A8 locus. a Fine mapping of the un.A8 region with 18 recombinants between the BM1355 and BM1360 intervals in the DH population. The recombinants were grouped into three genotypes based on their genotypes at the target region, and the SN phenotype of each recombinant genotype was compared with the two parents. b Fine mapping of the qSN.A8 region with 16,421 F2 plants. The number under the line represents the number of recombinants at the corresponding markers. A total of 114 recombinants were grouped into twelve genotypes based on their genotypes at the target region, and the SN phenotype of each recombinant genotype was compared with their parents and F1 hybrid. c Gene distribution in the candidate region of different reference genomes. Dash lines connect homologous genes from the region showing perfect microsynteny between different genomes. AA, homozygous genotype for the C4-146 allele; BB, homozygous genotype for the C4-58B allele; H, heterozygous genotype
To further narrow down the position of QTL un.A8 and identify the candidate genes responsible for SN, a large F2 population (containing 16,421 individuals obtained from the cross between C4-146 and C4-58B) was constructed and surveyed using the two flanking markers BM1663 and BM1646. Considering the dominant nature of low SN over high SN in the cross between C4-146 and C4-58B, only recombinants between the C4-146 homozygous genotype (AA) and the other two genotypes (BB or Heterozygous) are informative for fine mapping. Then, 114 recombinants were identified and used for further genotyping with 10 markers within the candidate region. These recombinants were grouped into 12 genotypes according to the allelic composition and recombination breakpoints (Fig. 4b). The recombinants information indicated that the BM1638, BM1641, BM1642, BM1357, BM1664, BM1667 and BM1668 markers were located on one side of un.A8, while the BM1672 marker was located on the other side. Also, BM1669 and BM1670 were found to co-segregate with the un.A8 locus (Fig. 4b). To further confirm the genomic region containing un.A8, the progenies of 3 key recombinants (S5, S14 and S15) at the locus were genotyped using the flanking markers (BM1668 and BM1672) and the co-segregated markers (BM1669 and BM1670). All recombinants from these progenies were selected and were grouped into 6 genotypes for phenotyping (Fig. S3). Using these data, we confirmed that the genomic region containing the un.A8 locus was narrowed down to the DNA fragment bounded by BM1668 and BM1672, with an approximate length of 80.1-kb in the Darmor-bzh genome (Fig. 4c).
Candidate gene analysis of un.A8
Based on the released rapeseed genome information, the candidate interval contained six predicted genes in the Darmor-bzh genome (designated BnaA08g07900D through BnaA08g07950D) corresponding to 3 and 4 genes in the B. napus ZS11 and Shengli genomes, respectively (Song et al. 2020; Fig. 4c; Table 6). Among these putative genes, one gene encodes expressed protein without annotation while the other genes encode homologs of Arabidopsis ATPase family AAA domain-containing protein, F-box/kelch-repeat protein, related to ubiquitin 1, photosystem I subunit F and zinc transporter 10 precursor (Table 6). None of the six genes has been previously reported to be involved in SN regulation. Therefore, un.A8 might be a novel QTL responsible for SN control.
To identify the candidate gene, the transcript levels of the six genes in both parents across five stages during ovary and silique development (1–2 mm, 2–4 mm and 4-5 mm pistil, 3DAP and 5DAP ovary) were quantified using qRT-PCR. The transcript levels of BnaA08g07940D and BnaA08g07950D were significantly higher in C4-58B than C4-146 during ovary and silique development (Fig. 5d), which supports the dominant nature of the low SN phenotype. However, BnaA08g07930D was expressed in C4-146 but not in C4-58B during ovary and silique development. Through sequence comparisons, we identified a 1,445-bp deletion starting from 1,015-bp upstream of the transcription start site to the mid-region of the second exon of BnaA08g07930D in C4-58B compared with the C4-146, which resulted in the formation of a pseudogene gene in C4-58B (Fig. 5a; Fig. S4). The remaining three genes, BnaA08g07900D through BnaA08g07920D, were not expressed in the developing ovary and siliques. Analysis of mRNA accumulation patterns for these predicted genes based on recent public expression databases in the B. napus ZS11 showed their expression profiles in various tissues (Fig. S5). There are three genes expressed in the candidate interval of ZS11 genome, i.e., BnaA08G0085000ZS, BnaA08G0085100ZS and BnaA08G0085200ZS, which are corresponding to BnaA08g07920D, BnaA08g07940D and BnaA08g07950D in the Darmor-bzh genome (Fig. 4c). The BnaA08G0085100ZS and BnaA08G0085200ZS were expressed in various tissues, with the higher expression detected in the developing siliques, but the BnaA08G0085000ZS transcripts were almost undetectable at the developing siliques (Fig. S5). These results suggested that both BnaA08g07940D and BnaA08g07950D may contribute to the observed variations in SN, hence we focused on them for further analysis.
Candidate gene analysis of the target region. a The gene structure, natural variations between the parents and protein alignment of BnaA08g07930D are shown. b The gene structure, natural variations between the parents and protein alignment of BnaA08g07940D are shown. The amino acid with variation was highlighted with bold. c The gene structure, natural variations between the parents and protein alignment of BnaA08g07950D are shown. The amino acid with variation was highlighted with bold. d The expression patterns of BnaA08g07940D and BnaA08g07950D in various tissues of C4-58B and C4-146, as determined by qRT-PCR with normalization to BnaUBC10. Values are the mean ± SD of three biological replicates. ***Significantly different at P < 0.001; DAP, days after pollination
To characterize the relationship between the candidate genes and the variations in SN, the promoter sequences, full-length genomic DNA, and coding sequences of BnaA08g07940D and BnaA08g07950D in the parental lines were amplified. A total of 10 exonic SNPs were detected in the open reading frame of BnaA08g07940D with 2 non-synonymous SNPs which resulted in amino acid variations in the signal peptide (Fig. 5b; Figs. S6 and S7). In the promoter region of BnaA08g07940D, abundant polymorphisms were also identified between the parental lines, including a 6159-bp insertion (starting from 1019-bp upstream of the transcription start site), 7 small INDELs and 7 SNPs in C4-58B (Fig. 5b; Fig. S8). For another candidate gene BnaA08g07950D, a 3-bp insertion and an exonic SNP were detected in the first exon of C4-58B, which resulted in an addition of Valine acid residue at the 21st amino acid (aa) and substitution of phenylalanine to leucine at the 28th amino acid (Fig. 5c; Figs. S9-S11). The BnaA08g07950D allele also showed alternative splicing, which terminated at the first intron and resulted in 8-aa insertion and 150-aa truncation in the C-terminus of the predicted protein in C4-58B (Fig. 5c; Figs. S9-S11). Additionally, 11 SNPs and 2 small INDELs were identified in the promoter interval (~ 3.0 kb before the start codon) between the two parents (Fig. 5c; Fig. S12). Thus, these results further supported the idea that BnaA08g07940D and BnaA08g07950D were the most promising candidate genes underlying the un.A8 locus.
Discussion
In rapeseed, SN and its related components are of great importance for crop yield. SN is the final consequence of a complex developmental process including ovule initiation and subsequent ovule/seed development. Thus, factors controlling each of these developmental processes will ultimately affect the natural variations of SN. However, almost all previously reported studies focused more on SN but not its related components. To date, only a few SN-related QTLs have been cloned, and the genetic mechanisms regulating the natural variation of SN and its related components are largely unknown.
Previously, we conducted a genetic dissection of plant architecture and yield-related traits using the HJ DH population in rapeseed with a high-density genetic map (Cai et al. 2016). However, only several QTLs with minor effects were identified for SN. Considering PAO is a key factor responsible for the natural variation of SN in rapeseed (Li et al. 2014, 2015; Yang et al. 2016), two lines (C4-146 and C4-58B) with significant differences in SN and AON without any obvious differences in ON were selected from the HJ DH population as the parental lines in the present study. It is expected to reduce the interference caused by factors controlling ovule initiation in the present segregation population and then make the QTL mapping simpler and more effective. Indeed, genetic analysis revealed that variations of SN, AON and PAO were all controlled by a single Mendelian factor in the DH mapping population derived from the cross of C4-146 × C4-58B. QTL analysis identified 19 consensus QTLs for six SN-related traits across environments. It is worth noting that a novel QTL on chromosome A8, un.A8, which pleiotropically controls all these traits except for ON, was stably detected across environments. This QTL explained more than 50% of the SN, AON and PAO variations and also exhibit a moderate contribution on SL and TSW. The C4-146 alleles at the un.A8 locus causes an increase in SN and SL with a simultaneous decrease in AON, PAO and TSW, in consistency with the trait performance of the two parents (Table 1). Eventually, it was fine mapped to an 80.1-kb region, which provides a solid basis for cloning this locus to uncover the molecular mechanisms of SN determination in rapeseed. Thus, our present study provides an effective and highly efficient strategy for conducting QTL mapping of complicated traits.
The SN in rapeseed shows great variation, ranging from 5 to 35 in its germplasm resources, which is invaluable for the genetic dissection of natural variation as well as genetic improvement of the trait (Yang et al. 2016). SN is also a highly complex trait involved in many developmental processes from the ovule initiation to seed development, which is affected by both internal factors from the mother plant and their interactions with external environmental conditions (Li et al. 2014; Xu et al. 2014; Yang et al. 2016). To understand the relative importance of different biological processes for its natural variation in germplasm resources, we previously conducted cytological analyses of some cultivars and a series of DH lines including C4-146 and C4-58B which shows great SN variation (Li et al. 2014). The results showed that the percentage of fertile ovules had the highest correlation with SN (r = 0.987), while no significant correlation (r = 0.538) was observed between ON and SN. These results indicated that the process of ovule development is very important for the natural variation of SN in rapeseed. Further cytological analyses revealed that ovule development between the stages of megaspore degradation and uninucleate gametophyte is the key process affecting ovule abortion, which in turn affects SN in rapeseed (Li et al. 2014). Similarly, BnaC9.SMG7b, the only gene map-based cloned for SN, also plays a role in regulating the formation of functional female gametophyte, thus determining the formation of functional megaspores and subsequent development of mature ovules (Li et al., 2015). Thus, the identified pleiotropic QTL un.A8 in the present study should be one of the key genes involved in the regulation of ovule abortion during the ovule development stages between megaspore degradation and uninucleate gametophyte. Unlike BnaC9.SMG7b which is a positive regulator of SN in rapeseed, the un.A8 locus should be a negative regulator of the ovule abortion and SN because of higher AON and lower SN are dominant traits in this study. Therefore, the un.A8 locus determines the generation of unfertile ovules with an unknown mechanism other than the reported BnaC9.SMG7b in rapeseed.
In the present study, we fine-mapped this major locus to an 80.1-kb region, where six genes were predicted (Table 6). Expression analyses and DNA sequencing showed that two homologs of Arabidopsis photosystem I subunit F (BnaA08g07940D) and zinc transporter 10 precursor (BnaA08g07950D) may contribute to the observed variations in ovule abortion and SN (Fig. 5). There are no reports on the regulatory functions of these genes in the control of SN-related traits in plants. Thus, further functional analysis of un.A8 will shed more light on the regulatory mechanism of SN natural variations in rapeseed. Moreover, the unique QTL un.A8 showed a major effect on the ovule abortion and SN, and a minor effect on TSW, with a decrease in AON and TSW by 4.04 and 0.17 g, respectively, as well as a simultaneous increase in SN by 3.58 at the C4-146 allele of the locus. These results suggested that un.A8 is of great potential in the improvement of SN in B. napus. Specific markers developed for this locus in the present study will be valuable for marker-assisted selection breeding for higher yield. Moreover, lower SN at this locus is fully dominant, which indicated that loss-of-function mutants in the un.A8 locus will reduce the ovule abortion and increase SN. With the rapid application of gene-editing technology in rapeseed, the creation of CRISPR/Cas9-mediated gene disruption at the un.A8 locus looks promising for yield improvement of the elite rapeseed cultivars.
Apart from the major QTLs on chromosome A8, we also identified three other consensus QTLs, cqTSW.A7-1, cqTSW.A7-2 and cqON.C6-2, which showed major effects on TSW and ON, respectively (Table 4). Two QTLs on the A07 chromosome, cqTSW.A7-1 and cqTSW.A7-2, were overlapped with previously identified TSWA7a/cqSW.A07-1 and TSWA7b/cqSW.A07-2, respectively (Fan et al. 2010; Wang et al. 2020). Thus, these two QTLs are stably expressed across different genetic backgrounds and environments, making them suitable targets for molecular cloning and breeding for seed weight improvement. For the ON trait, there was no QTL mapping study reported yet in rapeseed. Thus, the cqON.C6-2 locus represents the first major QTL controlling ON, which provides a target for dissecting the molecular basis of ovule initiation in the future.
References
Cai DF, Xiao YJ, Yang W, Ye W, Wang B, Younas M, Wu JS, Liu KD (2014a) Association mapping of six yield-related traits in rapeseed (Brassica napus L.). Theor Appl Genet 127:85–96
Cai GQ, Yang QY, Yi B, Fan CC, Edwards D, Batleyet J, Zhou YM (2014b) A complex recombination pattern in the genome of allotetraploid Brassica napus as revealed by a high-density genetic map. PLoS ONE 9(10):e109910
Cai GQ, Yang QY, Chen H, Yang Q, Zhang CY, Fan CC, Zhou YM (2016) Genetic dissection of plant architecture and yield-related traits in Brassica napus. Sci Rep 6:21625
Chalhoub B, Denoeud F, Liu SY, Parkin IAP, Tang HB, Wang XY, Chiquet J, Belcram H, Tong CB, Samans B et al (2014) Early allopolyploid evolution in the post-neolithic Brassica napus oilseed genome. Science 345:950–953
Charlesworth D (1989) Why do plants produce so many more ovules than seeds. Nature 338:21–22
Chen W, Zhang YS, Yao JB, Ma CZ, Tu JX, Fu TD (2011) Quantitative trait loci mapping for two seed yield component traits in an oilseed rape (Brassica napus) cross. Plant Breed 130:640–646
Chen F, Zhang JF, Qi KC, Pu HM, Chen S (2012) The analysis on diversity of germplasm resource in Brassica napus L. Jiangsu Agric Sci 11:98–99
Fan CC, Cai GQ, Qin J, Li QY, Yang MG, Wu JZ, Fu TD, Liu KD, Zhou YM (2010) Mapping of quantitative trait loci and development of allele-specific markers for seed weight in Brassica napus. Theor Appl Genet 121:1289–1301
Goffinet B, Gerber S (2000) Quantitative trait loci: a meta-analysis. Genetics 155:463–473
Hu LM, Zhang H, Yang QY, Meng QW, Han SQ, Nwafor CC, Khan MHU, Fan CC, Zhou YM (2018) Promoter variations in a homeobox gene, BnA10.LMI1, determine lobed leaves in rapeseed (Brassica napus L.). Theor Appl Genet 131:2699–2708
Khan SU, Jiao YM, Liu S, Zhang KP, Khan MHU, Zhai YG, Olalekan A, Fan CC, Zhou YM (2019) Genome-wide association studies in the genetic dissection of ovule number, seed number, and seed weight in Brassica napus L. Ind crop prod 142:111877
Khan MHU, Hu LM, Zhu MS, Zhai YG, Khan SU, Ahmar S, Olalekan A, Zhang KP, Fan CC, Zhou YM (2020) Targeted mutagenesis of EOD3 gene in B. napus L. regulates seed production. J Cell Physiol 236:1996–2007
Li YP, Cheng Y, Cai GQF, CC, Zhou, YM, (2014) Cytological basis and molecular mechanism of variation in number of seeds per pod in Brassica napus. Sci Agric Sin 8:822–831
Li SP, Chen L, Zhang LW, Li X, Liu Y, Wu ZK, Dong FM, Wan LL, Liu KD, Hong DF, Yang GS (2015) BnaC9.SMG7b Functions as a positive regulator of the number of seeds per silique in Brassica napus by regulating the formation of functional female gametophytes. Plant Physiol 169:2744–2760
Lincoln S, Daly M, Lander E (1992) Constructing genetic maps with MAPMAKER/EXP 3.0, 3rd edn. Whitehead Institute Technical report, Whitehead Institute, Cambridge
Mendham NJ, Shipway PA, Scott RK (1981) The effects of delayed sowing and weather on growth, development and yield of winter oil-seed rape (Brassica napus). J Agric Sci 96:389–416
Moles AT, Ackerly DD, Webb CO, Tweddle JC, Dickie JB, Westoby M (2005) A brief history of seed size. Science 307:576–580
Qi LP, Mao L, Sun CM, Pu YY, Fu TD, Ma CZ, Shen JX, Tu JX, Yi B, Wen J (2014) Interpreting the genetic basis of silique traits in Brassica napus using a joint QTL network. Plant Breed 133:52–60
Radoev M, Becker HC, Ecke W (2008) Genetic analysis of heterosis for yield and yield components in rapeseed (Brassica napus L.) by quantitative trait locus mapping. Genetics 179:1547–1558
Shi JQ, Li RY, Qiu D, Jiang CC, Long Y, Morgan C, Bancroft I, Zhao JY, Meng JL (2009) Unraveling the complex trait of crop yield with quantitative trait loci mapping in Brassica napus. Genetics 182:851–861
Shi JQ, Zhan JP, Yang YH, Ye J, Huang SM, Li RY, Wang XF, Liu GH, Wang HZ (2015) Linkage and regional association analysis reveal two new tightly-linked major-QTLs for pod number and seed number per pod in rapeseed (Brassica napus L.). Sci Rep. https://doi.org/10.1038/srep14481
Song JM, Guan ZL, Hu JL, Guo CC, Yang ZQ, Wang S, Liu DX, Wang B, Lu SP, Zhou R, Xie WZ, Cheng YF, Zhang YT, Liu KD, Yang QY, Chen LL, Guo L (2020) Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat Plants 6:34–45
Sun LJ, Wang XD, Yu KJ, Li WJ, Peng Q, Chen F, Zhang W, Fu SX, Xiong DQ, Chu P, Guan RZ, Zhang JF (2018) Mapping of QTLs controlling seed weight and seed-shape traits in Brassica napus L. using a high-density SNP map. Euphytica. https://doi.org/10.1007/s10681-018-2303-3
Van Went J, Cresti M (1988) Pre-fertilization degeneration of both synergids in Brassica campestris ovules. Sex Plant Reprod 1:208–216
Wang S, Basten CJ, Zeng ZB (2004) Windows QTL Cartographer 2.0. Department of Statistics, North Carolina State University, Raleigh
Wang XJ, Mathieu A, Cournède PH, Allirand JM, Jullien A, de Reffye P, Zhang BG (2011) Variability and regulation of the number of ovules, seeds and pods according to assimilate availability in winter oilseed rape (Brassica napus L.). Field Crops Res 122:60–69
Wang H, Yan M, Xiong M, Wang PF, Liu Y, Xin Q, Wan LL, Yang GS, Hong DF (2020) Genetic dissection of thousand-seed weight and fine mapping of cqSW.A03-2 via linkage and association analysis in rapeseed (Brassica napus L.). Theor Appl Genet 133:1321–1335
Xu WJ, Fu Y, Dong HL, Chen ZF, Zhang QY, Mao SS, He YJ, Qian W (2014) Morphologic and physiologic characterization of an ovule abortion mutant in Brassica napus. Sci Agric Sin 47:2062–2068
Yang YH, Shi JQ, Wang XF, Liu GH, Wang HZ (2016) Genetic architecture and mechanism of seed number per pod in rapeseed: elucidated through linkage and near-isogenic line analysis. Sci Rep 6:24124
Yang YH, Wang Y, Zhan JP, Shi JQ, Wang XF, Liu GH, Wang HZ (2017) Genetic and cytological analyses of the natural variation of seed number per pod in rapeseed (Brassica napus L.). Front Plant Sci. https://doi.org/10.3389/fpls.2017.01890
Zhang LW, Yang GS, Liu PW, Hong DF, Li SP, He QB (2011) Genetic and correlation analysis of silique-traits in Brassica napus L. by quantitative trait locus mapping. Theor Appl Genet 122:21–31
Zhang LW, Li SP, Chen L, Yang GS (2012) Identification and mapping of a major dominant quantitative trait locus controlling seeds per silique as a single Mendelian factor in Brassica napus L. Theor Appl Genet 125:695–705
Acknowledgements
This study was supported by Grants 2015CB150200 from the National Key Basic Research Program of China. This study was also supported by the National Natural Science Foundation of China (No. 31671279, 31371240 and 31971976).
Author information
Authors and Affiliations
Contributions
C.C.F. and Y.M.Z conceived and designed the experiments. Y.M.J., K.P.Z., G.Q.C., X.Z., H.Z. and L.M.H performed the experiments. C.C.F., Y.M.J., O.A., B.R.W. and Y.M.Z wrote the manuscript. K.D.Y. and S.Q.H analyzed the bioinformatics.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Annaliese S Mason.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiao, Y., Zhang, K., Cai, G. et al. Fine mapping and candidate gene analysis of a major locus controlling ovule abortion and seed number per silique in Brassica napus L.. Theor Appl Genet 134, 2517–2530 (2021). https://doi.org/10.1007/s00122-021-03839-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-021-03839-6