Abstract
Key message
A new resistance gene against Rice yellow mottle virus was identified and mapped in a 15-kb interval. The best candidate is a CC-NBS-LRR gene.
Abstract
Rice yellow mottle virus (RYMV) disease is a serious constraint to the cultivation of rice in Africa and selection for resistance is considered to be the most effective management strategy. The aim of this study was to characterize the resistance of Tog5307, a highly resistant accession belonging to the African cultivated rice species (Oryza glaberrima), that has none of the previously identified resistance genes to RYMV. The specificity of Tog5307 resistance was analyzed using 18 RYMV isolates. While three of them were able to infect Tog5307 very rapidly, resistance against the others was effective despite infection events attributed to resistance-breakdown or incomplete penetrance of the resistance. Segregation of resistance in an interspecific backcross population derived from a cross between Tog5307 and the susceptible Oryza sativa variety IR64 showed that resistance is dominant and is controlled by a single gene, named RYMV3. RYMV3 was mapped in an approximately 15-kb interval in which two candidate genes, coding for a putative transmembrane protein and a CC-NBS-LRR domain-containing protein, were annotated. Sequencing revealed non-synonymous polymorphisms between Tog5307 and the O. glaberrima susceptible accession CG14 in both candidate genes. An additional resistant O. glaberrima accession, Tog5672, was found to have the Tog5307 genotype for the CC-NBS-LRR gene but not for the putative transmembrane protein gene. Analysis of the cosegregation of Tog5672 resistance with the RYMV3 locus suggests that RYMV3 is also involved in Tog5672 resistance, thereby supporting the CC-NBS-LRR gene as the best candidate for RYMV3.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice is the staple food of more than half of the world’s population. Africa produced 24 million tons between 2004 and 2013, which is only 3.5% of world rice production but Africa accounts for 29.6% of world imports (http://faostat3.fao.org/). Despite regular improvement, yields remain low, and are not sufficient to feed the growing population.
Among various abiotic and biotic stresses that reduce rice yield, Rice yellow mottle virus (RYMV) disease is a major cause of losses in sub-Saharan Africa (Ochola and Tusiime 2011; Issaka et al. 2012; Kam et al. 2013). RYMV is a Sobemovirus endemic to Africa, and is prevalent in all the sub-Saharan countries where rice is cultivated. Extensive characterization of its diversity has revealed different viral strains, i.e. genetic groups, with gradual differentiation from East Africa, where it originated, to West Africa (Pinel-Galzi et al. 2015). The epidemiology of RYMV is complex (Traoré et al. 2009) and the efforts to manage the disease has, thus, been directed towards discovery of resistance genes.
Resistance to RYMV has been investigated in the two cultivated species of rice: Oryza sativa, originating from Asia and cultivated worldwide, and O. glaberrima, more recently domesticated and only cultivated locally in West Africa (Linares 2002). Several accessions, mostly of O. glaberrima species, were found to show no symptoms after RYMV inoculation, nor detectable virus content by an enzyme-linked immunosorbent assay (ELISA), and were, thus, considered highly resistant (Thiémélé et al. 2010). So far, two genes conferring high resistance to RYMV have been identified. The first one, RYMV1, encodes a translation initiation factor, eIF(iso)4G1 (Albar et al. 2006). Four recessive resistance alleles have been described: rymv1-2 in two resistant O. sativa accessions, and rymv1-3, -4 and -5 in O. glaberrima accessions. As observed for eIF4E or eIF(iso)4E translation initiation factors involved in resistance against plant viruses (Sanfaçon 2015), the eIF(iso)4G1 factor of susceptible plants interacts with theVPg (viral protein genome-linked) of RYMV, a viral protein covalently linked to the 5′ end of the viral RNA, while resistance is gained by the rupture of the interaction (Hébrard et al. 2010). The second resistance gene, RYMV2, also named CPR5-1, is homologous to the Arabidopsis thaliana gene for Constitutive expressor of Pathogenesis-Related genes-5 (CPR5), a nucleoporin controlling defense mechanisms (Bowling et al. 1997; Wang et al. 2014a; Gu et al. 2016). RYMV2-mediated resistance is recessive and is conferred by a null allele identified in seven O. glaberrima accessions (Orjuela et al. 2013).
Even if using genetic resistance is efficient and environment-friendly, the emergence of resistance-breaking virus variants is frequent and the durability of resistance genes is still a challenge (Kobayashi et al. 2014). Among resistance genes and alleles involved in high resistance to RYMV, only rymv1-2 allele has been transferred to high-yielding varieties, and very recently deployed in the fields (Bouet et al. 2013; Ndjiondjop et al. 2013); so, its durability is unknown. However, the resistance breakdown of accessions carrying resistance alleles at RYMV1 and RYMV2 genes has already been observed and characterized in experimental conditions. Mutations in RYMV associated with RYMV1 and RYMV2 resistance breakdown were found in the central domain of the VPg (Pinel-Galzi et al. 2007; Traoré et al. 2010) and the membrane anchor domain of polyprotein P2a (Pinel-Galzi et al. 2016), respectively. In addition, regardless of the resistance gene, the ability of the different isolates to overcome O. glaberrima or O. sativa resistant accessions may be associated with a trade-off among host species involving a polymorphism at amino acid 49 of the VPg (Poulicard et al. 2012; Pinel-Galzi et al. 2016). Although experimental conditions are not fully representative of field conditions, these results suggest that RYMV1 and RYMV2 resistance genes may not be sufficient to achieve durable resistance against RYMV.
Of the resistant accessions identified in O. glaberrima species by Thiémélé et al. (2010), some do not have a known resistant allele on either RYMV1 or RYMV2 and, therefore, may represent genetic diversity to be exploited against RYMV. We characterized, here, the specificity of the resistance of the O. glaberrima Tog5307 accession using isolates representative of RYMV diversity and report the fine mapping of the RYMV3 gene, which is involved in the resistance of Tog5307 and Tog5672 accessions.
Materials and methods
Plant material
Four rice accessions were used in this study. Tog5307 and Tog5672 are O. glaberrima accessions resistant to RYMV. Tog5672 has the resistance allele rymv1-4 for RYMV1 (Albar et al. 2006) but is suspected to carry an additional unknown resistance gene (Thiémélé et al. 2010). O. sativa spp. indica IR64 variety and O. glaberrima Tog6206 accession are highly susceptible to RYMV. An interspecific population derived from the Tog5307 × IR64 cross was developed to map Tog5307 resistance. Successive backcrosses, with IR64 as the recurrent parent and phenotypic selection for RYMV resistance at each step, were performed to introgress Tog5307 resistance into the IR64 genetic background. Independent lineages were derived from four resistant BC1F1 progenitors. The BC3F2 and BC3F3 generations were used for segregation analysis and fine genetic mapping of the resistance gene (Supplemental Fig. 1). The genetic basis of Tog5672 resistance was analyzed in an F2 population derived from the Tog5672 × Tog6206 cross.
Viral material
Eighteen RYMV isolates were used in this study. Their characteristics are listed in Table 1. Isolates were classified according to their country of origin, their viral strain, i.e. genetic group, and the presence of a glutamic acid (E49) or a threonine (T49) at position 49 of their VPg. The sample included nine T49 isolates from West Africa representing strains S1 and S2, four E49 isolates from West Africa representing strain S1, and five E49 isolates from East Africa, representing strains S4, S5 and S6. Isolates were propagated on IR64 susceptible plants and stored at −20 °C.
Four new isolates, collected on cultivated rice in Togo and Niger, were included in the viral corpus. For their characterization, total RNA was extracted with the RNeasy Plant Mini kit (Qiagen, Germany).The coat protein (CP) and the VPg genes of the RYMV genome were amplified by reverse transcription-polymerase chain reaction (RT-PCR) (Fargette et al. 2002; Hébrard et al. 2006). The CP sequences were compared to a set of CP sequences for phylogenetic reconstruction using the maximum-likelihood method (Traoré et al. 2010). All four isolates belonged to strain S1.
Phenotypic evaluation
The resistance spectrum was characterized in four different trials conducted in similar conditions. The plants were cultivated in greenhouse conditions in trays holding 28 individual 350-ml pots. Between 40 and 56 Tog5307 plants and eight IR64 control plants were inoculated with each isolate. Inoculum was prepared by grinding IR64-infected leaves in 0.1 M phosphate buffer (0.1 g of plant material per ml of buffer) and adding 600-mesh carborundum. Plants were mechanically inoculated at approximately 2 weeks after sowing by rubbing the inoculum on all the leaves. Resistance was assessed based on the presence/absence of symptoms and eventually by ELISA. Symptoms were observed on the leaves emerged after inoculation, at 2, 4 and 6 weeks after inoculation. Samples from the last emerged leaf were collected at 4 and 6 weeks after inoculation and virus content was estimated by direct double-antibody sandwich (DAS-) ELISA, as described by Ndjiondjop et al. (1999), using a polyclonal antiserum directed against an RYMV isolate from Madagascar (N’guessan et al. 2000). About 1 mg of fresh leaves in 1 ml buffer was used. Samples were considered negative if the optical density at 405 nm was less than three times the negative control. Since recovery from the infected state is highly unlikely, ELISA was first performed on samples collected at 4 weeks after inoculation on symptomatic plants, and, if negative, on samples collected at 6 weeks after inoculation; in the case of non-symptomatic plants, ELISA was first performed on samples collected at 6 weeks and, if positive, on samples collected at 4 weeks after inoculation.
For the genetic analysis of Tog5307 and Tog5672 resistance, BF1 isolate was used. The conditions for culture and RYMV inoculation were the same as described above. Resistance was assessed based on the presence/absence of symptoms and eventually confirmed by ELISA. Susceptible plants developed clear yellow mottles at 10–15 days post-inoculation, whereas resistant plants had no symptoms at all. ELISA was performed as described above on samples collected between 2 and 3 weeks after inoculation.
Genomic resources
Three different genome sequences were used in this study: the O. sativa spp. japonica Nipponbare release Os-Nipponbare-Reference-IRGSP-1.0 (Kawahara et al. 2013), the O. sativa spp. indica IR64 sequence Os-IR64-Draft-CSHL-1.0 (Schatz et al. 2014) and the O. glaberrima CG14 genome assembly Oryza_glaberrima_AGI1.1 (Wang et al. 2014b). The IR64 scaffold corresponding to the resistance gene region was identified with Blast 2.2.31+ (Camacho et al. 2009). Annotations on CG14 refer to the 2011-05-AGI conducted by Munich Information Center for Protein Sequences and browsed through Gramene release 49 (http://www.gramene.org/). Sequence comparison in the candidate region was visualized with Artemis software (Rutherford et al. 2000). Tog5307 and Tog5672 sequences at candidate open reading frames (ORFs) were obtained by Sanger sequencing of PCR amplification fragments at Beckman Coulter Genomics (Takeley, UK).
Development of molecular markers and PCR procedure
The 384 single nucleotide polymorphism (SNP) chip described in Orjuela et al. (2014) was used for the first step of Tog5307 resistance mapping.
PCR markers based on polymorphisms between CG14 and Nipponbare or IR64 sequences were developed for the fine mapping of Tog5307 resistance. A marker specific to the rymv1-4 allele was also developed to screen the Tog5672 × Tog6206 F2 population. Primers were designed using Primer3 (Untergasser et al. 2007) and checked for specificity in the Nipponbare genome using Primer blaster tool (Droc et al. 2009). Most of the markers were either indel markers or cleaved amplified polymorphic sequence (CAPS) markers. One derived CAPS (dCAPS) marker was developed based on the dCAPS Finder 2.0 software (Neff et al. 2002). The complete characteristics of the markers are listed in Table 2.
Genotyping was performed by direct PCR amplifications using leaf extracts or by standard PCR amplifications using extracted DNA. PCR conditions are described in detail in Orjuela et al. (2013). After amplification of CAPS or dCAPS markers, 10 µl of buffer–restriction enzyme mixture (2 µl of 10× reaction buffer, 0.4 µl of appropriate restriction enzyme, 0.2 µl of bovine serum albumin, and 7.4 µl of double-distilled H2O) was added to 10 µl of PCR product and incubated at 37 °C for 2 h. Electrophoresis of PCR or digestion products was performed in agarose, and DNA bands were visualized with ethidium bromide.
Results
Tog5307 resistance spectrum
Tog5307 accession was inoculated with 18 isolates of RYMV, selected as representative of the genetic diversity of the virus in Africa.
Clear symptoms were observed at 12 days after inoculation on all IR64 plants that had been inoculated with any of the isolates tested and on all the Tog5307 plants inoculated by Ng119 and Tz211 isolates. After Ni1 inoculation, symptoms were also observed on 60% of the plants at 12 days after inoculation, and on all the plants but one, at 4 weeks after inoculation. ELISA on randomly selected Tog5307 plants confirmed the presence of the virus. For the other isolates, almost all the Tog5307 inoculated plants were symptom-free at 2 weeks after inoculation and the percentage of diseased plants was assessed by ELISA performed on all the plants (Fig. 1). It ranged from 0 to 25% at 4 weeks after inoculation and from 0 to 45% at 6 weeks after inoculation. Infection rates at these two dates were highly correlated (R 2 = 0.95).
Percentage of diseased plants on Tog5307 accessions for 18 RYMV isolates at 4 weeks after inoculation. Plants were inoculated with RYMV isolates representing different strains of the virus (S1, S2, S4, S5, S6) and distinguished by a T or a E amino acid at position 49 of the VPg. The error bars represent 95% confidence intervals
Infection rates with Ni1, Ng119 and Tz211 isolates ranged between 98 or 100% at 4 weeks after inoculation and were significantly different from those obtained with other isolates (Fisher’s exact test, p < 10−3), revealing a strong specificity of interactions between Tog5307 resistance and RYMV isolates. At 4 weeks after inoculation, no infection cases were observed with isolates CIa, Ng111, Tz201 and Tz11, and a single case of infection each with isolates CI4, Ng113 and Mg1. These isolates differed significantly from BF1, CI2, CI8, and Ng106, which showed over 18% of infected plants, while the infection rate of CI3, Ng109, Tg274 and Tz5 was between the two.
As the T/E polymorphism at codon 49 of the VPg is associated with RYMV1 and RYMV2 resistance-breakdown ability and with a selective advantage of T49 isolates on O. glaberrima susceptible accessions (Traoré et al. 2010; Poulicard et al. 2012; Pinel-Galzi et al. 2016), its association with the infection rate on Tog5307 was analyzed. Considering all the isolates except Tz211, Ng119 and Ni1, significantly different infection rates (Fisher exact test, p < 10−3) were observed between T49 isolates (average infection rate of 16.3%) and E49 isolates (2.5%), suggesting that E/T polymorphism also plays a role in the interaction between RYMV isolates and Tog5307 resistance.
Inheritance of Tog5307 resistance
The inheritance of Tog5307 resistance was analyzed in populations derived from the Tog5307 × IR64 cross and subsequent backcrosses using IR64 as the recurrent parent (Supplemental Table 1). Resistance phenotype was assessed by inoculation with isolate BF1.
Preliminary observations of the presence/absence of symptoms were made on BC1F1, BC2F1 and BC3F1 hybrids in the course of the introgression process. Two contrasted phenotypes were observed. IR64 and some hybrids exhibited very clear symptoms as early as 10–14 days after inoculation, while Tog5307 and other hybrids showed no symptoms even several weeks after inoculation.
The segregation of resistance was evaluated in five BC3F3 families derived from four independent BC1F1. Between 16 and 24 plants per family were phenotyped. Resistance was assessed by the presence/absence of symptoms and ELISA. The presence of symptoms was perfectly correlated with the detection of virus in ELISA. A total of 82 resistant and 32 susceptible plants were identified, which fit a 3:1 resistant/susceptible segregation pattern (χ² = 0.57; P = 0.45). This segregation pattern was also supported by each individual family (Table 3). This result is in agreement with monogenic and dominant inheritance of resistance.
Identification of a candidate locus for Tog5307 resistance on chromosome 11
To identify the genomic region carrying the Tog5307 resistance gene, 18 BC1F1 plants which did not show any symptoms after RYMV inoculation were genotyped with the SNP chip described in Orjuela et al. (2014). Among the 384 SNP markers on the chip, 119 were polymorphic in the target population. As expected, BC1F1 were either heterozygous, or homozygous for IR64 alleles. Two markers showed a strong segregation distortion towards the heterozygous state (17 out of 18 individuals) (Fig. 2). This segregation strongly diverged from the 1:1 ratio (χ² = 14.2; p = 1.6 × 10−4), as expected for markers associated with resistance. These markers were on chromosome 6 at position 2,346,023 on the O. sativa Nipponbare reference genome and on chromosome 11 at position 26,619,446. On chromosome 6, the marker was located on the interspecific sterility locus S1 (Garavito et al. 2010), which is known to induce major segregation distortions toward the O. glaberrima allele in interspecific crosses between O. sativa and O. glaberrima (Lorieux et al. 2000). Thus, the distortion observed on chromosome 6 is likely to be due to S1. As no sterility locus has ever been reported in this region of chromosome 11 (Garavito et al. 2010), the second distortion is assumed to result from the resistance gene.
Genomic regions in linkage disequilibrium with resistance in a population of 18 resistant BC1F1 plants from the cross (IR64 × Tog5307) × IR64. Plants were challenged with BF1 isolate and genotyped with the SNP chip described in Orjuela et al. (2014). The number of heterozygous BC1F1 plants at each marker over the genome is indicated in ordinates. The two encircled regions on chromosomes 6 and 11 show segregation distortion toward the heterozygous state
Since none of the previously identified resistance genes (Albar et al. 2003; Orjuela et al. 2013) or QTLs (Boisnard et al. 2007) against RYMV cosegregated with resistance in this population, the resistance gene of Tog5307 was considered to be new and was named RYMV3, for Resistance-to-yellow-mottle-virus-3.
Fine mapping of RYMV3
Mapping of RYMV3 was confirmed and refined on BC3F2 and BC3F3 plants. In the first step, a BC3F2 population was phenotyped based on the presence/absence of symptoms. Thirty-seven symptom-free plants were genotyped with 13 markers designed in the candidate region on chromosome 11. The cosegregation of resistance with a region comprised between markers C5 and M4, corresponding to an 880-kb-region on the Nipponbare genomic sequence was confirmed.
In the second step, 1363 BC3F3 plants were genotyped at markers C5 and M4 (Fig. 3). Fifty-eight recombinant individuals were identified and genotyped with additional markers in the interval. The phenotyping of the most informative recombinants was performed based on symptom observation on their progenies, with 10 to 16 plants tested per progeny. The interval containing RYMV3 was finally bounded by markers M6 and C16. Two recombination events were located between marker M6 and RYMV3, and one event between RYMV3 and C16.
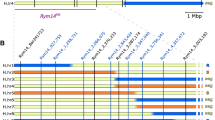
Physical map of the RYMV3 locus. a Fine mapping of the RYMV3 locus in BC3F3 families. The number of recombinants between the corresponding marker and the gene is in parentheses. b Genotypes of recombinants and phenotype in their selfing progenies (R resistant, seg segregation of resistance, S susceptible). Tog5307 homozygous, IR64 homozygous and heterozygous states are represented by black, white and gray bars, respectively. The number of recombinants corresponding to each genotype/phenotype pattern is indicated on the left. Recombination events indicate that RYMV3 is bounded by M6 and C16 markers
RYMV3 candidate genes
The length of the M6-C16 region containing RYMV3 is 15, 17 and 42 kb in the CG14, IR64 and Nipponbare genomic sequences, respectively. The differences in size were mainly explained by 2.1 and 3.5 kb insertions/deletions between the CG14 and IR64 genomes, and insertions of 2.4, 6.7 and 16 kb in the Nipponbare genome that were absent in the IR64 genome (Fig. 4).
Collinearity between the CG14, IR64 and Nipponbare genomes in the genomic region containing RYMV3, visualized with Artemis software (Rutherford et al. 2000). Collinear regions are in gray. Genes annotated in the different genomes are indicated by boxes representing exons. Annotations in the IR64 genome were obtained from Nipponbare ones. The white gene is ORGLA11g0175700. The hatched gene is the CC-NBS-LRR gene (ORGLA11G75800 on the CG14 genome and LOC_Os11g43700 on the Nipponbare genome). It is only partially included in the region M6-C16 containing RYMV3. Markers used for mapping are shown with dashed lines
Two ORFs had been annotated in the CG14 genome in the M6-C16 interval (Fig. 5). The first one (ORGLA11g0175700) was labeled as a gene coding for a hypothetical protein. Protein sequence analysis (http://www.ebi.ac.uk/interpro/) revealed a transmembrane structure with a 69-amino acid (aa) cytoplasmic domain, a 28-aa transmembrane domain and a 129-aa non-cytoplasmic domain, but no known conserved functional domains. Any ORF orthologous to the ORF of ORGLA11g0175700 was absent in the Nipponbare and IR64 sequences. However, two homologs of ORGLA11g0175700 were identified in the CG14 genome (ORGLA11g0103500 and ORGLA11g0189200) and in the Nipponbare genome (LOC_Os11g25530 and LOC_Os11g47750) on chromosome 11, 8.3 Mb upstream and 1.5 Mb downstream of locus RYMV3 in the CG14 genome.
Annotation of the RYMV3 candidate region and polymorphisms detected between CG14 and Tog5307. The regions sequenced in Tog5307 are indicated by thick black bars. Black arrows represent non-synonymous SNPs, the white arrow a synonymous SNP, and the white star a deletion of 33 bp. ORGLA11G0175700 is a putative transmembrane gene. ORGLA11G0175800 is a typical CC-NBS-LRR resistance gene
The second ORF (ORGLA11g0175800) was only partially located within the M6-C16 region, as C16 is located on the first intron. The orthologous ORF in the Nipponbare sequence is named LOC_Os11g43700. The encoded protein showed a coiled-coil (CC), a nucleotide-binding site (NBS) and a leucine-rich repeat (LRR) domain, characteristic of the large family of resistance genes, and consequently named CC-NBS-LRR genes (Takken and Goverse 2012).
These ORFs were sequenced by Sanger methodology in the Tog5307 accession. Three SNPs were detected between Tog5307 and the susceptible accession CG14 on gene ORGLA11g0175700. One of them was intronic, and the other two resulted in a glycine-to-arginine change at aa position 28 in the cytoplasmic domain, and a proline-to-leucine change at aa position 212 in the non-cytoplasmic domain. Two polymorphisms were identified in the part of the ORGLA11g0175800 gene located within the M6-C16 region. The first one was a 33-bp deletion in the first intron, and the second one a non-synonymous SNP in the LRR domain inducing a conversion of a lysine into an arginine at aa position 779.
Tog5672 second resistance gene is linked to RYMV3
Tog5672 has the rymv1-4 resistance allele of the RYMV1 gene (Albar et al. 2006) but segregation analysis suggested that this accession contains another resistance gene (Thiemele et al. 2010). As no evidence for a resistance allele on the RYMV2 gene was reported by Orjuela et al. (2013), we investigated if Tog5672 carries RYMV3. The sequence of the RYMV3 candidate genes in Tog5672 was assessed by Sanger sequencing. Out of the five polymorphisms identified between Tog5307 and CG14, Tog5672 displayed the Tog5307 genotype only for the SNP corresponding to aa position 779 in the LRR domain of the CC-NBS-LRR gene.
A cross was performed between Tog5672 and Tog6206, a susceptible O. glaberrima accession. F2 plants were inoculated with the BF1 isolate and resistance was assessed by the presence/absence of symptoms and ELISA. Plants were genotyped using the CAPS marker SNP_rymv1-4, targeting a polymorphism characterizing rymv1-4 allele, and the dCAPS marker LRR_RYMV3, targeting the polymorphism in the LRR domain of the candidate gene for RYMV3. As expected, plants homozygous for rymv1-4 turned out to be resistant (Table 4). Of the others, the seven plants homozygous for the Tog6206 allele at the LRR_RYMV3 marker were susceptible, whereas the 12 plants homozygous for the Tog5672 allele were resistant, clearly demonstrating that the second resistance determinant of Tog5672 is linked to RYMV3. However, regardless of being homozygous or heterozygous for the susceptibility allele at RYMV1, 12 out of 20 RYMV3 heterozygous plants had a clearly resistant phenotype, but the eight others showed symptoms and/or had low but positive ELISA scores. This may suggest that resistance expression may be influenced by additional factors.
Some F2 plants were also inoculated with Ng119 isolate. All the plants that were non-homozygous for rymv1-4 and heterozygous (eight plants) or homozygous for the Tog5672 allele (two plants) for RYMV3 were homogeneously infected at 2 weeks after inoculation. The second resistance determinant of Tog5672, thus, showed the same specificity of interaction as Tog5307 with BF1 and Ng119 isolates, which is in agreement with RYMV3 being involved in Tog5672 resistance.
Discussion
RYMV is one of the main causes of losses in rice crops in Africa and Madagascar, and breeding resistant cultivars is likely the best strategy to control this disease. However, understanding the specificity of interactions between RYMV isolates, resistance genes or alleles, and the genetic background is crucial to achieving durable resistance. The analysis of the Tog5307 resistance spectrum revealed contrasted patterns of interaction with RYMV isolates. Tog5307 behaved like a susceptible accession when challenged with Tz211, Ng119 and Ni1 isolates, that are, thus, presumed to be virulent on Tog5307, according to the definition of Vanderplank (1968). With the other isolates, infection occurred later and in a limited number of plants. These observations may reflect incomplete penetrance of resistance, i.e. the occurrence of plants with the resistant genotype that develop the disease, as previously described in other plant/virus pathosystems (Ouibrahim et al. 2014; Chandra-Shekara et al. 2008; Poque et al. 2015). However, they may also result from the emergence, in some plants, of a viral variant able to overcome resistance, as frequently observed in host/pathogen interactions (Lecoq et al. 2004; Quenouille et al. 2013 for plant/virus interactions) and well described for RYMV1- and RYMV2-mediated resistance (Hébrard et al. 2006; Pinel-Galzi et al. 2016). The two hypotheses may also coexist. Analysis of infected Tog5307 samples based on back-inoculation on Tog5307 plants and Illumina sequencing, as performed in Pinel-Galzi et al. (2016), is planned to determine which hypothesis applies. Nevertheless, in most of the cases, RYMV is unable to multiply in Tog5307 plants, which confirmed that Tog5307 resistance is as efficient as the resistance conferred by RYMV1 or RYMV2 genes.
Poulicard et al. (2012) reported a selective advantage of isolates with a T at position 49 of the VPg in susceptible O. glaberrima accessions when T49 and E49 isolates were inoculated in competition, and interpreted it as an adaptation to O. glaberrima species. This T/E polymorphism has also been previously identified as a key determinant of the ability to overcome resistance of Gigante (O. sativa accession carrying rymv1-2), Tog5681 (O. glaberrima accession carrying rymv1-3) and Tog7291 (O. glaberrima accession carrying a resistance allele at RYMV2 gene): E49 isolates rarely overcome the resistance of Tog5681 or Tog7291, while T49 isolates rarely overcome the resistance of Gigante (Traoré et al. 2010; Pinel-Galzi et al. 2016). Here, we observed that, excluding the isolates that infected all Tog5307 plants, infection rate was significantly higher for T49 isolates than for E49 isolates. This result also supports the association of T/E polymorphism with the frequency of infection cases in O. glaberrima resistant accessions regardless of the resistance gene. We are currently investigating the pattern of interaction between T49 and E49 isolates and near-isogenic lines of IR64 (O. sativa spp. indica) introgressed with different resistance genes to confirm these hypotheses. However, the E/T polymorphism is not the only determinant of the specificity of interaction between Tog5307 and RYMV isolates. In particular, this polymorphism does not explain why the T49 CIa isolate did not infect Tog5307 and why 100% infection rate was observed with the E49 Tz211 isolate.
Tog5307 resistance is under the control of RYMV3, the third major resistance gene to RYMV identified in O. glaberrima. Our mapping, sequencing and isolate specificity results suggest that it is also involved in Tog5672 resistance. We think that RYMV3 is sufficient to explain the resistance of Tog5307, but we cannot totally exclude that an interaction between RYMV3 and a gene closely linked to the S1 locus, on chromosome 6, is involved. In O. sativa × O. glaberrima crosses, the S1 locus is the main interspecific sterility locus and was shown to undergo a strong segregation distortion toward the O. glaberrima allele in progenies (Lorieux et al. 2000). This locus explains why such distortion was observed in BC1F1. In the same way, BC3F2 and BC3F3 used for RYMV3 fine mapping are expected to be homozygous for the O. glaberrima allele at locus S1. This lack of segregation would prevent the detection of a gene linked to locus S1 that may be involved in resistance in interaction with RYMV3. A small fraction of plants with the O. sativa allele at locus S1 would be sufficient to exclude the implication of such a gene. To date, backcrosses for introgression of RYMV3 in IR64 are as advanced as BC6. We confirmed the association of RYMV3 with resistance in BC6F2 but, despite screening more than one hundred BC6F2 plants, no plants with the O. sativa S1 allele were identified (data not shown).
The segregation of resistance in the Tog5672 × Tog6206 population does not exclude the involvement of another resistance gene, which could have been masked by S1 distortion or by the absence of polymorphism in the Tog5307 × IR64 population. Some plants belonging to the Tog5672 × Tog6206 population that were heterozygous for RYMV3 were only partially resistant, thereby supporting the involvement of such a gene. However, incomplete penetrance of RYMV3-mediated resistance could also explain the observed segregation. Penetrance may be affected by environmental factors and by a dosage effect, as previously described for different NB-LRR genes (Collmer et al. 2000; Chandra-Shekara et al. 2006; Zhu et al. 2010) or other resistance genes (Ouibrahim et al. 2014). In the Tog5307 × IR64 population, phenotyping was mostly performed on progenies and a small bias in the ratio between resistant and susceptible plants, that may be due to an incomplete penetrance of the resistance in heterozygous plants, would not have been detected. We checked the penetrance of the resistance in 84 BC5F3 plants from three segregating families developed during the introgression process of RYMV3 in IR64 and no heterozygous plants developed the disease (data not shown) which do not support a strong dosage effect in this population. All these results highlight the overriding role of RYMV3 in resistance but suggest that complex interactions between the gene, the genetic background and the features of the RYMV isolates may affect the expression of resistance.
We mapped RYMV3 in a 15-kb-long interval on the CG14 reference genome. Two candidate genes are annotated in this interval. ORGLA11g0175700, encodes a hypothetical protein with a probable transmembrane domain but this gene has no homologs outside the Oryza genus nor has any known conserved domain. However, resistance genes with no significant homology to any characterized proteins have already been described (Xiao et al. 2001; Wang et al. 2015). If ORGLA11g0175700 is involved in resistance, two non-synonymous polymorphisms between Tog5307 and CG14, one in the probable cytoplasmic domain and the other outside, are candidates to explain the gain in resistance. The other gene, ORGLA11g0175800, is a typical NB-LRR gene belonging to the CC-NBS-LRR subfamily. Genes in this class are the most common dominant resistance genes in all types of pathosystems. Resistance to plant viruses is not an exception to the rule, as 20 NB-LRR genes against plant viruses have been cloned so far (Boualem et al. 2016). NB-LRR proteins are cytoplasmic. They are composed of a central LRR domain involved in pathogen specificity, linked to a C-terminal NB-ARC (nucleotide-binding adaptor shared by APAF-1, certain R gene products and CED-4) domain required for signal transduction, and either a TIR (toll and interleukin-1 receptor) or a CC (coiled coil) domain in the N terminal, which play a role in the auto-inhibition of the protein and in protein–protein interactions (Takken and Goverse 2012). NB-LRR genes directly or indirectly specifically recognize a strain or isolate specific avirulence factor and trigger the resistance signaling cascade, which, in the majority of cases, leads to local cell death, named hypersensitive response. In the plant/virus pathosystem, an extreme resistance, characterized by no detectable viral accumulation and no visible necrosis, can also be observed (Bendahmane et al. 1999). In the ORGLA11g0175800 gene, we detected a single non-synonymous SNP between Tog5307 from CG14, leading to a conversion from a lysine to an arginine in the LRR domain. Although both are positively charged amino acids, they may confer different specificities to the protein. The LRR domain of NB-LRR proteins has been shown to play a central role in virus recognition (Farnham and Baulcombe 2006; Tomita et al. 2011; Sekine et al. 2012) and such polymorphism can explain the difference between a resistant and a susceptible plant, as previously demonstrated in rice/blast interactions (Bryan et al. 2000; Liu et al. 2007). The involvement of a NB-LRR gene would be in agreement with the observed isolate-specific resistance mediated by RYMV3 gene. Therefore, we believe that ORGLA11g0175800 is the best candidate for Tog5307 resistance. The functional complementation of a susceptible O. sativa variety with the Tog5307 allele of the candidate genes will confirm the role of the candidates, and exclude the involvement of additional genes in Tog5307 resistance.
In conclusion, we mapped RYMV3, a new resistance gene to RYMV, and identified a NB-LRR as a very convincing candidate gene. We showed that RYMV3-associated resistance can be codominant or incompletely penetrant, depending on the genetic background or the environmental conditions. The three known resistance genes to RYMV are presumed to be involved in different mechanisms of interaction with the virus. Their pyramiding will probably render the emergence of resistance-breaking variants difficult. This, is thus, a very promising way to achieve durable resistance, in particular in the regions of Africa where specific interactions between the local isolates, the resistance genes and the genetic background may rapidly impair the efficiency of varieties with a single resistance gene.
Author contribution statement
AG, LA, SC developed the mapping populations. IS, DS, EH provided and characterized the viral material. HP, LA, AG conceived and designed the experiments. HP, SC, OK performed phenotyping and genotyping of the mapping population. HP, FS performed the sequence analysis. HP, LA analyzed the data and wrote the paper. FS, EH revised the paper.
References
Abubakar Z, Ali F, Pinel A, Traoré O, N‘Guessan P, Notteghem J-L, Kimmins F, Konaté G, Fargette D (2003) Phylogeography of Rice yellow mottle virus in Africa. J Gen Virol 84:733–743. doi:10.1099/vir.0.18759-0
Albar L, Bangratz-Reyser M, Hébrard E, Ndjiondjop MN, Jones M, Ghesquière A (2006) Mutations in the eIF(iso)4G translation initiation factor confer high resistance of rice to Rice yellow mottle virus. Plant J 47:417–426. doi:10.1111/j.1365-313X.2006.02792.x
Bendahmane A, Kanyuka K, Baulcombe DC (1999) The Rx gene from potato controls separate virus resistance and cell death responses. Plant Cell 11:781–792. doi:10.1105/tpc.11.5.781
Boisnard A, Albar L, Thiémélé DE, Rondeau M, Ghesquière A (2007) Evaluation of genes from eIF4E and eIF4G multigenic families as potential candidates for partial resistance QTLs to Rice yellow mottle virus in rice. Theor Appl Genet 116:53–62. doi:10.1007/s00122-007-0646-6
Boualem A, Dogimont C, Bendahmane A (2016) The battle for survival between viruses and their host plants. Curr Opin Virol 17:32–38. doi:10.1016/j.coviro.2015.12.001
Bouet A, Amancho AN, Kouassi N, Anguete K (2013) Comportement de nouvelles lignées isogéniques de riz irrigué dotées du gène de résistance (rymv1) au RYMV en Afrique de l’ouest: situation en Côte d’Ivoire. Int J Biol Chem Sci 7:1221–1233. doi:10.4314/ijbcs.v7i3.28
Bowling SA, Clarke JD, Liu Y, Klessig DF, Dong X (1997) The cpr5 mutant of Arabidopsis expresses both NPR1-dependent and NPR1-independent resistance. Plant Cell 9:1573–1584. doi:10.1105/tpc.9.9.1573
Bryan GT, Wu KS, Farrall L, Jia Y, Hershey HP, McAdams SA, Faulk KN, Donaldson GK, Tarchini R, Valent B (2000) A single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta. Plant Cell 12:2033–2046. doi:10.1105/tpc.12.11.2033
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:421. doi:10.1186/1471-2105-10-421
Chandra-Shekara AC, Gupte M, Navarre D, Raina S, Raina R, Klessig D, Kachroo P (2006) Light-dependent hypersensitive response and resistance signaling against Turnip crinkle virus in Arabidopsis. Plant J 45:320–334. doi:10.1111/j.1365-313X.2005.02618.x
Collmer CW, Marston MF, Taylor JC, Jahn M (2000) The I gene of bean: a dosage-dependent allele conferring extreme resistance, hypersensitive resistance, or spreading vascular necrosis in response to the potyvirus Bean common mosaic virus. Mol Plant Microbe In 13:1266–1270. doi:10.1094/MPMI.2000.13.11.1266
Droc G, Périn C, Fromentin S, Larmande P (2009) OryGenesDB 2008 update: database interoperability for functional genomics of rice. Nucleic Acids Res 37:D992–D995. doi:10.1093/nar/gkn821
Fargette D, Pinel A, Halimi H, Brugidou C, Fauquet C, Van RM (2002) Comparison of molecular and immunological typing of isolates of Rice yellow mottle virus. Arch Virol 147:583–596. doi:10.1007/s007050200008
Fargette D, Pinel A, Abubakar Z, Traoré O, Brugidou C, Fatogoma S, Hébrard E, Choisy M, Séré Y, Fauquet C, Konaté G (2004) Inferring the evolutionary history of Rice yellow mottle virus from genomic, phylogenetic, and phylogeographic studies. J Virol 78:3252–3261. doi:10.1128/JVI.78.7.3252
Fargette D, Pinel A, Rakotomalala M, Sangu E, Traoré O, Sérémé D, Sorho F, Issaka S, Hébrard E, Séré Y, Kanyeka Z, Konaté G (2008) Rice yellow mottle virus, an RNA plant virus, evolves as rapidly as most RNA animal viruses. J Virol 82:3584–3589. doi:10.1128/JVI.02506-07
Farnham G, Baulcombe DC (2006) Artificial evolution extends the spectrum of viruses that are targeted by a disease-resistance gene from potato. Proc Natl Acad Sci USA 103:18828–18833. doi:10.1073/pnas.0605777103
Garavito A, Guyot R, Lozano J, Gavory F, Samain S, Panaud O, Tohme J, Ghesquière A, Lorieux M (2010) A genetic model for the female sterility barrier between Asian and African cultivated rice species. Genetics 185:1425–1440. doi:10.1534/genetics.110.116772
Gu Y, Zebell SG, Liang Z, Wang S, Kang B-H, Dong X (2016) Nuclear pore permeabilization is a convergent signaling event in effector-triggered immunity. Cell 166:1–13. doi:10.1016/j.cell.2016.07.042
Hébrard E, Pinel-Galzi A, Bersoult A, Siré C, Fargette D (2006) Emergence of a resistance-breaking isolate of Rice yellow mottle virus during serial inoculations is due to a single substitution in the genome-linked viral protein VPg. J Gen Virol 87:1369–1373. doi:10.1099/vir.0.81659-0
Hébrard E, Poulicard N, Gérard C, Traoré O, Wu HC, Albar L, Fargette D, Bessin Y, Vignols F (2010) Direct interaction between the Rice yellow mottle virus (RYMV) VPg and the central domain of the rice eIF(iso)4G1 factor correlates with rice susceptibility and RYMV virulence. Mol Plant Microbe In 23:1506–1513. doi:10.1094/MPMI-03-10-0073
Issaka S, Basso A, Sorho F, Onasanya A, Haougui A, Sido AY, Aké S, Fargette D, Séré Y (2012) Diagnosis and importance of rice yellow mottle disease epidemics in Niger republic. J Appli Biosci 50:3501–3511
Kam H, Laing M, Ouoba J (2013) Rice traits preferred by farmers and their perceptions of Rice yellow mottle virus (RYMV) disease in Cascades Region of Burkina Faso. Afr J Agric Res 8:2703–2712. doi:10.5897/AJAR12.1723
Kawahara Y, de la Bastide M, Hamilton JP, Kanamori H, McCombie WR, Ouyang S, Schwartz DC, Tanaka T, Wu J, Zhou S, Childs KL, Davidson RM, Lin H, Quesada-Ocampo L, Vaillancourt B, Sakai H, Lee SS, Kim J, Numa H, Itoh T, Buell CR, Matsumoto T (2013) Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice (N Y) 6:4. doi:10.1186/1939-8433-6-4
Kobayashi K, Sekine KT, Nishiguchi M (2014) Breakdown of plant virus resistance: can we predict and extend the durability of virus resistance? J Gen Plant Pathol 80:327–336. doi:10.1007/s10327-014-0527-1
Lecoq H, Moury B, Desbiez C, Palloix A, Pitrat M (2004) Durable virus resistance in plants through conventional approaches: a challenge. Virus Res 100:31–39. doi:10.1016/j.virusres.2003.12.012
Linares OF (2002) African rice (Oryza glaberrima): history and future potential. P Natl Acad Sci USA 99:16360–16365. doi:10.1073/pnas.252604599
Liu X, Lin F, Wang L, Pan Q (2007) The in Silico Map-Based Cloning of Pi36, a rice coiled-coil nucleotide-binding site leucine-rich repeat gene that confers race-specific resistance to the blast fungus. Genetics 176:2541–2549. doi:10.1534/genetics.107.075465
Lorieux M, Ndjiondjop MN, Ghesquière A (2000) A first interspecific Oryza sativa x Oryza glaberrima microsatellite-based genetic linkage map. Theor Appl Genet 100:593–601. doi:10.1007/s001229900061
N’guessan P, Pinel A, Caruana ML, Frutos R, Sy A, Ghesquiere A, Fargette D (2000) Evidence of the presence of two serotypes of Rice yellow mottle sobemovirus in Côte d’Ivoire. Eur J Plant Pathol 106:167–178. doi:10.1023/A:1008792109954
Ndjiondjop M-N, Albar L, Fargette D, Fauquet C, Ghesquière A (1999) The genetic basis of high resistance to Rice yellow mottle virus (RYMV) in cultivars of two cultivated rice species. Plant Dis 83:931–935. doi:10.1007/s001229900061
Ndjiondjop MN, Albar L, Sow M, Yao N, Djedatin G, Thiémélé D, Ghesquière A (2013) Integration of molecular markers in rice improvement: a case study on resistance to Rice yellow mottle virus. In: Wopereis M, Johnson D, Ahmadi N, et al. (eds) Realizing Africa’s Rice Promise. CABI, pp 161–172
Neff MM, Turk E, Kalishman M (2002) Web-based primer design for single nucleotide polymorphism analysis. Trends Genet 18:613–615. doi:10.1016/S0168-9525(02)02820-2
Ochola D, Tusiime G (2011) Survey on incidences and severity of Rice yellow mottle virus disease in Eastern Uganda. International Journal of Plant Pathology 2:15–25. doi:10.3923/ijpp.2011.15.25
Orjuela J, Thiémélé D, Kolade O, Chéron S, Ghesquière A, Albar L (2013) A recessive resistance to Rice yellow mottle virus is associated with a rice homolog of the CPR5 gene, a regulator of active defense mechanisms. Mol Plant Microbe In 26:1455–1463. doi:10.1094/MPMI-05-13-0127-R
Orjuela J, Sabot F, Chéron S, Vigouroux Y, Adam H, Chrestin H, Sanni K, Lorieux M, Ghesquière A (2014) An extensive analysis of the African rice genetic diversity through a global genotyping. Theor Appl Genet 127:2211–2223. doi:10.1007/s00122-014-2374-z
Ouibrahim L, Mazier M, Estevan J, Pagny G, Decroocq V, Desbiez C, Moretti A, Gallois JL, Caranta C (2014) Cloning of the Arabidopsis rwm1 gene for resistance to Watermelon mosaic virus points to a new function for natural virus resistance genes. Plant J 79:705–716. doi:10.1111/tpj.12586
Pinel A, N’Guessan P, Bousalem M, Fargette D (2000) Molecular variability of geographically distinct isolates of Rice yellow mottle virus in Africa. Arch Virol 145:1621–1638. doi:10.1007/s007050070080
Pinel-Galzi A, Rakotomalala M, Sangu E, Sorho F, Kanyeka Z, Traoré O, Sérémé D, Poulicard N, Rabenantoandro Y, Séré Y, Konaté G, Ghesquière A, Hébrard E, Fargette D (2007) Theme and variations in the evolutionary pathways to virulence of an RNA plant virus species. PLOS Pathog 3:e180. doi:10.1371/journal.ppat.0030180
Pinel-Galzi A, Traoré O, Séré Y, Hébrard E, Fargette D (2015) The biogeography of viral emergence: Rice yellow mottle virus as a case study. Curr Opin Virol 10:7–13. doi:10.1016/j.coviro.2014.12.002
Pinel-Galzi A, Dubreuil-Tranchant C, Hébrard E, Mariac C, Ghesquière A Albar L (2016) Mutations in Rice yellow mottle virus polyprotein P2a involved in RYMV2 gene resistance breakdown. Front Plant Sci. 71779 10.3389/fpls.2016.01779
Poque S, Pagny G, Ouibrahim L, Chague A, Eyquard JP, Caballero M, Candresse T, Caranta C, Mariette S, Decroocq V (2015) Allelic variation at the rpv1 locus controls partial resistance to Plum pox virus infection in Arabidopsis thaliana. BMC Plant Biol 15:159. doi:10.1186/s12870-015-0559-5
Poulicard N, Pinel-Galzi A, Traoré O, Vignols F, Ghesquière A, Konaté G, Hébrard E, Fargette D (2012) Historical contingencies modulate the adaptability of Rice yellow mottle virus. PLOS Pathog 8:e1002482. doi:10.1371/journal.ppat.1002482
Quenouille J, Montarry J, Palloix A, Moury B (2013) Farther, slower, stronger: how the plant genetic background protects a major resistance gene from breakdown. Mol Plant Pathol 14:109–118. doi:10.1111/j.1364-3703.2012.00834.x
Rutherford K, Parkhill J, Crook J, Horsnell T, Rice P, Rajandream MA, Barrell B (2000) Artemis: sequence visualization and annotation. Bioinformatics 16:944–945. doi:10.1093/bioinformatics/16.10.944
Sanfaçon H (2015) Plant translation factors and virus resistance. Viruses 7:3392–3419. doi:10.3390/v7072778
Schatz MC, Maron LG, Stein JC, Wences AH, Gurtowski J, Biggers E, Lee H, Kramer M, Antoniou E, Ghiban E, Wright MH, Chia JM, Ware D, McCouch SR, McCombie WR (2014) Whole genome de novo assemblies of three divergent strains of rice, Oryza sativa, document novel gene space of aus and indica. Genom Biol 15:506. doi:10.1186/s13059-014-0506-z
Sekine KT, Tomita R, Takeuchi S, Atsumi G, Saitoh H, Mizumoto H, Kiba A, Yamaoka N, Nishiguchi M, Hikichi Y, Kobayashi K (2012) Functional differentiation in the leucine-rich repeat domains of closely related plant virus-resistance proteins that recognize common avr proteins. Mol Plant Microbe In 25:1219–1229. doi:10.1094/MPMI-11-11-0289
Takken FLW, Goverse A (2012) How to build a pathogen detector: structural basis of NB-LRR function. Cur Opin Plant Biol 15:375–384. doi:10.1016/j.pbi.2012.05.001
Thiémélé DE, Boisnard A, Ndjiondjop MN, Chéron S, Séré Y, Aké S, Ghesquière A, Albar L (2010) Identification of a second major resistance gene to Rice yellow mottle virus, RYMV2, in the African cultivated rice species, O. glaberrima. Theor Appl Genet 121:169–179. doi:10.1007/s00122-010-1300-2
Tomita R, Sekine KT, Mizumoto H, Sakamoto M, Murai J, Kiba A., Hikichi Y, Suzuki K, Kobayashi K (2011) Genetic basis for the hierarchical interaction between Tobamovirus spp. and L resistance gene alleles from different pepper species. Mol Plant Microbe In 24:108–117. doi:10.1094/MPMI-06-10-0127
Traoré O, Pinel-Galzi A, Sorho F, Sarra S, Rakotomalala M, Sangu E, Kanyeka Z, Séré Y, Konaté G, Fargette D (2009) A reassessment of the epidemiology of Rice yellow mottle virus following recent advances in field and molecular studies. Virus Res 141:258–267. doi:10.1016/j.virusres.2009.01.011
Traoré O, Pinel-Galzi A, Issaka S, Poulicard N, Aribi J, Aké S, Ghesquière A, Séré Y, Konaté G, Hébrard E, Fargette D (2010) The adaptation of Rice yellow mottle virus to the eIF(iso)4G-mediated rice resistance. Virology 408:103–108. doi:10.1016/j.virol.2010.09.007
Untergasser A, Nijveen H, Rao X, Bisseling T, Geurts R, Leunissen JA (2007) Primer3Plus, an enhanced web interface to Primer3. Nucleic Acids Res 35:W71–W74. doi:10.1093/nar/gkm30
VanderPlank JE (1968) Disease resistance in plants. Academic, New York
Wang S, Gu Y, Zebell SG, Anderson LK, Wang W, Mohan R, Dong X (2014a) A noncanonical role for the CKI-RB-E2F cell-cycle signaling pathway in plant effector-triggered immunity. Cell Host Microbe 16:787–794. doi:10.1016/j.chom.2014.10.005
Wang M, Yu Y, Haberer G, Marri PR, Fan C, Goicoechea JL, Zuccolo A, Song X, Kudrna D, Ammiraju JSS, Cossu RM, Maldonado C, Chen J, Lee S, Sisneros N, de Baynast K, Golser W, Wissotski M, Kim W, Sanchez P, Ndjiondjop MN, Sanni K, Long M, Carney J, Panaud O, Wicker T, Machado CA, Chen M, Mayer KFX, Rounsley S, Wing RA (2014b) The genome sequence of African rice (Oryza glaberrima) and evidence for independent domestication. Nat Genet 46:982–988. doi:10.1038/ng.3044
Wang C, Zhang X, Fan Y, Gao Y, Zhu Q, Zheng C, Qin T, Li Y, Che J, Zhang M, Yang B, Liu Y, Zhao K (2015) XA23 is an executor R protein and confers broad-spectrum disease resistance in rice. Mol Plant 8:290–302. doi:10.1016/j.molp.2014.10.010
Xiao S, Ellwood S, Calis O, Patrick E, Li T, Coleman M, Turner JG (2001) Broad-spectrum mildew resistance in Arabidopsis thaliana mediated by RPW8. Science 291:118–120. doi:10.1126/science.291.5501.118
Zhu Y, Qian W, Hua J (2010) Temperature modulates plant defense responses through NB-LRR proteins. PLOS Pathog 6:e1000844. doi:10.1371/journal.ppat.1000844
Acknowledgements
This work was financially supported by the Agropolis Foundation (project N°1403-066, SURVEY) and the Global Rice Science Partnership (GriSP). The French Ministère de l’Enseignement Supérieur et de la Recherche provided a PhD grant to H. Pidon. We thank Christine Tranchant-Dubreuil, Agnès Pinel-Galzi and Harold Chrestin for technical help and Yacouba Séré for giving access to some RYMV isolates. We acknowledge François Anthony and Mathias Lorieux for helpful discussions. We would also like to thank our reviewers for their helpful comments in improving the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical standards
The authors declare that the experiments comply with the current laws of the country in which they were performed.
Additional information
Communicated by Michael Thomson.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2017_2853_MOESM1_ESM.pdf
Populations derived from the IR64 x Tog5307 cross and used for the mapping of the RYMV3 gene. Backcrosses were performed using resistant F1 plants as donors and the susceptible O. sativa variety IR64 as recurrent parent. The generations used for the analysis of resistance segregation and RYMV3 mapping and fine mapping are indicated on the right. For resistance segregation analysis, the number of resistant (R) and susceptible (S) plants is indicated for each tested progeny (PDF 467 KB)
Rights and permissions
About this article
Cite this article
Pidon, H., Ghesquière, A., Chéron, S. et al. Fine mapping of RYMV3: a new resistance gene to Rice yellow mottle virus from Oryza glaberrima . Theor Appl Genet 130, 807–818 (2017). https://doi.org/10.1007/s00122-017-2853-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-017-2853-0