Abstract
Activation of hepatic stellate cell (HSC), which is the main source of extracellular matrix, plays a pivotal role in liver fibrogenesis. Autophagy of hepatic stellate cell has been recently implicated in liver fibrosis, but the regulation of hepatic stellate cell autophagy during this process remains poorly understood. Here, we first identified miR-96-5p as an aberrantly expressed miRNA in fibrotic liver tissues. Next, we transfected miR-96-5p mimic into human hepatic stellate cell line LX-2 and observed decreased protein and mRNA levels of α-SMA and Col1A1. In addition, transfection of miR-96-5p mimic significantly reduced autophagy activity of LX-2 cells, while transfection of miR-96-5p inhibitor promoted LX-2 cell autophagy. Moreover, autophagy-related protein 7 (ATG7) was predicted as a potential target of miR-96-5p and luciferase assay confirmed its direct interaction with miR-96-5p. Finally, reintroduction of ATG7 into LX-2 cells reversed miR-96-5p-mediated inhibition of autophagy as well as α-SMA and Col1A1 expression. In conclusion, we demonstrated that miR-96-5p can inhibit hepatic stellate cell activation by blocking autophagy via ATG7. These findings provide new insight into the development of miRNA-based anti-fibrotic strategies.
Key messages
• Altered miRNA expression profile is observed in fibrotic liver tissues.
• miR-96-5p can inhibit HSC activation.
• Autophagy of HSC is repressed by miR-96-5p during activation.
• ATG7 is a direct target of miR-96-5p.
• ATG7 can rescue miR-96-5p-mediated inhibition of autophagy and HSC activation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatic stellate cells (HSCs) are mesenchymal cells of the liver that are located in the space of Disse and are quiescent in a healthy liver [1]. When liver injury such as viral infection occurs, HSCs become activated to produce extracellular matrix (ECM) to form a scar and prevent liver damage from exacerbating [2]. However, prolonged and repeated activation of HSCs in chronic liver disease results in excessive ECM accumulation, which distorts the hepatic architecture and leads to liver fibrosis and further cirrhosis [3, 4]. Therefore, reducing the number of activated HSCs or inhibiting HSC activation has been a focus of research for liver fibrosis prevention.
In recent years, studies have revealed that macroautophagy (hereafter referred to as “autophagy”) deregulation was involved in HSC activation and liver fibrosis [5,6,7,8]. Thoen et al. [9] observed an increased autophagic flux during HSC activation and showed that blockade of autophagy resulted in inhibition of in vitro HSC activation. In addition, it is reported that quiescent HSCs can upregulate autophagy upon liver injury and further liberate free fatty acids that can undergo β-oxidation to fuel and maintain the activated HSCs [10]. The same research group also confirmed that endoplasmic reticulum stress induces HSC activation in an autophagy-dependent way [11]. Taken together, autophagy may serve as a potential target for developing anti-fibrotic strategies.
MicroRNAs (miRNAs) are short non-coding RNAs that regulate gene expression post-transcriptionally through translational repression or mRNA degradation [12], and play roles in the pathogenesis of various human diseases including chronic liver disease [13]. A number of studies have demonstrated that miRNA-mediated autophagy modulation plays an important part in disease progression [14]. However, its involvement in HSC activation and liver fibrosis has not been fully characterized.
In the current study, we have identified miR-96-5p as an aberrant expressed microRNA in fibrotic human liver tissue and further investigation revealed that miR-96-5p prevents HSC activation by regulating autophagy via targeting autophagy-related (ATG) gene ATG7.
Materials and methods
Cell culture and treatment
LX-2 cells, the immortalized human HSC line and human embryonic kidney (HEK) 293T cell line, were maintained in Dulbecco’s modified eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and antibiotics (penicillin and streptomycin) at 37 °C in 5% CO2. For treatment, cells were incubated with 5 ng/ml recombinant human transforming growth factor beta 1 (TGF-β1) (Peprotech Inc., Rocky Hill, NJ), 5 μM rapamycin (Rapa) for the indicated times.
Plasmid constructs and miRNAs
The ATG7-expressing construct was generated by cloning full-length ATG7 into pCDH-puro vector using the following primers: 5′-acagctagctgcgctgaac atggcggcagctacggggga-3′ (forward), 5′-acgaattctcagatggtctcatcatcg-3′ (reverse). For luciferase assay, the miR-96-5p binding site found in the 3′-UTR region of the ATG7 mRNA and its mutated version were cloned into a modified pGL3-control vector; the linker sequences used are as follows: wild type (WT) sequences: 5′-cgcctgccaaaccc-3′, 5′-tcgagggtttggcaggcgagct-3′ and mutant sequences: 5′- cgccgctcaaaccc-3′, 5′-tcgagggtttgagcggcgagct-3′. The pRL-TK vector encoding Renilla luciferase was used as an internal control reporter. The miR-96-5p mimics, miRNA mimic negative control (MMNC), miR-96-5p inhibitor, and miRNA inhibitor negative control (MINC) were purchased from RiboBio (Guangzhou, China).
Transfection and luciferase activity assay
Transient transfection of LX-2 and 293T cells was performed using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions. Reporter constructs containing wild-type or mutant ATG7 3′-UTR were co-transfected with miR-96-5p mimic or MMNC (50 nM). Forty-eight hours after transfection, cells were harvested and lysed. Luciferase reporter activities were measured using the Dual-Glo Luciferase Assay System (Promega). Firefly luciferase activity was normalized to Renilla luciferase activity.
GFP-LC3 analyses
Thirty-six hours after co-transfection of miRNAs and GFP-LC3, LX-2 cells were incubated in DMEM containing 5 μM rapamycin for 24 h. Puncta expression of GFP was photographed using a digital camera and software (Leica). A threshold of GFP-LC3 puncta < 10 per LX-2 cell was determined as non-autophagic. Percentage of cells displaying autophagic GFP-LC3 puncta expression was quantified (mean ± SD).
Western blot analysis and antibodies
For Western blotting, protein samples were analyzed by SDS-PAGE and transferred onto nitrocellulose membranes, followed by blocking and probing with primary antibodies against ATG7 (Sigma-Aldrich, A2856, 1:1000), SQSTM1/p62 (Abcam, ab109012, 1:1000), ACTB (Cell Signaling, 3700S, 1:1000), α-SMA (Abcam, ab32575, 1:1000), Col1A1 (Abcam, ab166606, 1:1000), and LC3 (Abcam, ab192890, 1:1000) for detection.
Immunofluorescence
LX-2 cells were seeded onto 24-well plates containing 1-cm-diameter glass coverslips. The cells were transfected with 50 nM miR-96-5p mimic or inhibitor and incubated with 5 ng/ml TGF-β1 for 48 h. After treatments, cells were fixed with 4% paraformaldehyde for 15 min and permeabilized with 1% Triton X-100 in PBS for 10 min at room temperature. Cells were washed with PBS, blocked with 5% BSA in PBS for 1 h, and incubated with anti-α-SMA (Abcam, ab32575, 1:100) for 2 h at room temperature. Nuclei were stained with DAPI (Sigma-Aldrich). Cell fluorescence was visualized by a Leica microscope.
RNA extract and quantification of mRNA and miRNA expression
Total RNA was isolated from liver tissues and harvested LX-2 cells using TRIzol reagent (Life Technologies), and cDNA was obtained by reverse transcription with RT kit KR108 (for mRNA) and KR211 (for miRNA) (Tiangen, Beijing, China). qRT-PCR was performed with SYBR green real-time PCR kit FP205 (for mRNA) and FP401 (for miNRA) (Tiangen, Beijing, China). The primer sequences used for quantification are listed in Table 1. Relative mRNA or miRNA expression was determined using the 2-∆∆CT method. GAPDH or U6 was used as an endogenous control. Melting curve analysis was performed to verify the specificity of the products, and each sample was tested in triplicate.
Human fibrotic liver tissue samples
Fibrotic human liver tissue samples were obtained from biopsies of patients from Huashan Hospital with informed consents. RNA extraction and miR-96-5p expression level analysis were performed as described above. Experiments and procedures were in accordance with the Helsinki Declaration and approved by the Human Ethics Committee of Fudan University.
Statistical analysis
All data were expressed as mean ± standard deviation (SD). Differences between samples were analyzed by Student’s t test using SPSS V19.0 software (SPSS Inc., Chicago, IL, USA). Two-sided p < 0.05 was considered as statistically significant. Column bar graphs were performed with GraphPad Prism 5 (GraphPad Software Inc., San Diego, CA, USA).
Results
miR-96-5p is upregulated in fibrotic human liver tissue and can inhibit pro-fibrogenic signaling in human stellate cells activation
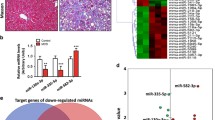
To determine miRNAs that are differently expressed during fibrosis, we compared the miRNA expression profiles of stage 0 fibrotic samples and stage 4 fibrotic samples using an Agilent miRNA microarray. The expression patterns of some miRNAs were distinct as shown in the heat map (Fig. 1a). The higher magnification view of the zone showing the differentially expressed miRNAs, among which miR-96-5p was consistently upregulated in stage 4 fibrotic samples when compared with stage 0 fibrotic samples. Quantitative reverse transcription-polymerase chain reaction (qRT-PCR) was engaged to confirm the expression alteration of miR-96-5p at different stages of fibrosis that had been revealed by miRNA expression profiling (Fig. 1b). In addition, in LX-2 cells (a human HSC cell line), increased mRNA levels of α-smooth muscle actin (α-SMA) and collagen type I alpha 1 (Col1α1) following TGF-β1 treatment were accompanied by elevated endogenous miR-96-5p levels (Fig. 1c). Thus, we speculate that increased miR-96-5p level is a response to pro-fibrogenic signaling and may prevent fibrogenesis. Activated HSCs are the main source of ECM proteins including Col1α1 and are generally characterized by the presence of α-SMA [2]. To evaluate if miR-96-5p was involved in the pro-fibrogenic activation of HSC, miR-96-5p mimic or miR-96-5p inhibitor was transfected into human LX-2 cells. Interestingly, our qRT-PCR analysis revealed that miR-96-5p mimic significantly decreased the mRNA levels of α-SMA and Col1α1 (Fig. 1d), while transfection with miR-96-5p inhibitor showed a marked elevation in the expression of α-SMA and Col1α1 mRNA levels (Fig. 1e). Subsequent Western blotting confirmed that miR-96-5p mimic resulted in a significant reduction of the protein levels of α-SMA and Col1α1 (Fig. 1f), while the a-SMA and Col1A1 protein levels were enhanced by miR-96-5p inhibitor (Fig.1g). Besides, immunofluorescence staining for α-SMA revealed that α-SMA accumulation decreased markedly under miR-96-5p mimic transfection (Fig. 1h). These results suggest that miR-96-5p inhibits pro-fibrogenic signaling during HSC activation. We next searched TargetScan database (http://www.targetscan.org/vert_71/) and confirmed that α-SMA and Col1α1 are not potential direct targets of miR-96-5p; thus, we focused our study on autophagy.
miR-96-5p is aberrantly expressed in fibrotic human liver tissues and can suppress LX-2 cell activation in vitro. a Heat map showing relative expression of miRNAs between stage 0 (S0) and stage 4 (S4) fibrotic tissues. Differentially expressed miRNAs are shown as magnified image on the right panel. b qRT-PCR analysis of miR-96-5p expression in liver tissues with different fibrotic stages. *p < 0.05, ***p < 0.001. c qRT-PCR analysis of α-SMA, Col1, and miR-96-5p expression in LX-2 cells treated with 5 ng/ml TGF-β1 for the indicated times. *p < 0.05, **p < 0.01, ***p < 0.001 compared to the previous time point. d qRT-PCR analysis of α-SMA and Col1 expression in LX-2 cells transfected with 50 nM miRNA mimic negative control (MMNC) or miR-96-5p mimic for 48 h. **p < 0.01, ***p < 0.001. e qRT-PCR analysis of α-SMA and Col1 expression in LX-2 cells transfected with 50 nM miRNA inhibitor negative control (MINC) or miR-96-5p inhibitor for 48 h. *p < 0.05, ***p < 0.001. f Western blot analysis and quantification of α-SMA and Col1 protein levels in LX-2 cells transfected as indicated in d. *p < 0.05, **p < 0.01. g Western blot analysis and quantification of α-SMA and Col1 protein levels in LX-2 cells transfected as indicated in e. **p < 0.01. h Fluorescence microscopy images of LX-2 cells stained with specific antibody against α-SMA (red) after transfection as indicated. Nuclei were stained with DAPI (blue). Scale bars 2 mm
miR-96-5p suppresses hepatic stellate cell autophagy
It was reported that miR-96 can regulate autophagy in prostate cancer cells under hypoxia [15], and given that autophagy has been identified as an important player in HSC activation, we hypothesized that miR-96-5p inhibits HSC activation by impairing autophagy. To experimentally verify this postulation, LX-2 cells were co-transfected with GFP-LC3 and miR-96-5p mimic or miRNA mimic negative control (MMNC) followed by rapamycin administration. As shown in Fig. 2, overexpression of miR-96-5p significantly repressed rapamycin-induced GFP-LC3 puncta formation (Fig. 2a, b). The degradation of autophagy receptor SQSTM1/p62 reflects autolysosomal lytic activity and autophagic flux and therefore serves as an autophagy marker [16]. In line with the above findings, Western blotting revealed increased protein levels of SQSTM1/p62 following miR-96-5p mimic transfection while decreased LC3-I to LC3-II conversion was detected in the same condition (Fig. 2c, d). Next, GFP-LC3 and miR-96-5p inhibitor or miRNA inhibitor negative control (MINC) were co-transfected into LX-2 cells, which received subsequent rapamycin treatment. As expected, we observed a significant increase of rapamycin-dependent GFP-LC3 puncta formation (Fig. 3a, b) and LC3-I to LC3-II conversion (Fig. 3c, d). In addition, miR-96-5p inhibitor significantly promoted SQSTM1/p62 degradation (Fig. 3c, d). Taken together, these findings demonstrate that HSC autophagy can be inhibited by miR-96-5p.
Overexpression of miR-96-5p inhibits rapamycin-induced autophagy in LX-2 cells. a Fluorescence images of GFP-LC3 puncta formation in LX-2 cells transfected with 50 nM miRNA mimic negative control (MMNC) or miR-96-5p mimic in response to DMSO (carrier) or rapamycin (Rapa, 2.5 μM, 24 h). b Quantitative analysis of the experiments in a. ***p < 0.001, NS not significant. c Western blot analysis of p62 and LC3 protein levels in LX-2 cells treated as indicated in a. d Quantitative analysis of p62 (left) and LC3 (right) protein levels in c. **p < 0.01, ***p < 0.001
Inhibition of endogenous miR-96-5p promotes rapamycin-induced autophagy in LX-2 cells. a Fluorescence images of GFP-LC3 puncta formation in LX-2 cells transfected with 50 nM miRNA inhibitor negative control (MINC) or miR-96-5p inhibitor in response to DMSO (carrier) or rapamycin (Rapa, 2.5 μM, 24 h). b Quantitative analysis of the experiments in a. ***p < 0.001, NS not significant. c Western blot analysis of p62 and LC3 protein levels in LX-2 cells treated as indicated in a. d Quantitative analysis of p62 (left) and LC3 (right) protein levels in c. ***p < 0.001
miR-96-5p represses ATG7 expression in human hepatic stellate cells
To unravel the mechanism of miR-96-5p-mediated autophagy inhibition, we searched the TargetScan database and ATG7 (GenBank accession number: NM_006395.2) was identified as a target with two putative miR-96-5p seed match sites (Fig. 4a). To confirm the bioinformatics-based predictions, LX-2 cells were transfected with miR-96-5p mimic or MMNC and followed by rapamycin administration; ATG7 protein and mRNA levels were measured by Western blotting and qRT-PCR, respectively. Indeed, the transcript levels of ATG7 were also decreased by miR-96-5p mimic (Fig. 4b). Besides, a significant reduction of ATG7 protein levels was detected in LX-2 cells transfected with miR-96-5p mimic rather than cells transfected with MMNC (Fig. 4c, d). Conversely, introduction of miR-96-5p inhibitor showed an increase in the expression of ATG7 protein levels compared with MINC (Fig. 4e, f). These results indicate that miR-96-5p exerts both transcriptional and translational regulation on ATG7 expression.
miR-96-5p suppresses ATG7 expression in LX-2 cells. a miR-96-5p target sequences in the 3′-UTR of the ATG7 mRNA. b qRT-PCR analysis of ATG7 expression in LX-2 cells transfected with 50 nM miRNA mimic negative control (MMNC) or miR-96-5p mimic in response to rapamycin (2.5 μM, 24 h). ***p < 0.001. c Western blot analysis of ATG7 protein levels in LX-2 cells transfected with 50 nM miRNA mimic negative control (MMNC) or miR-96-5p mimic in response to rapamycin (2.5 μM) at indicated times. d Quantitative analysis of the experiments in c. **p < 0.01, NS not significant. e Western blot analysis of ATG7 protein levels in LX-2 cells transfected with 50 nM miRNA inhibitor negative control (MINC) or miR-96-5p inhibitor in response to rapamycin (2.5 μM, 24 h). f Quantitative analysis of the experiments in e. **p < 0.01, NS not significant
ATG7 is a direct target of miR-96-5p
Based on bioinformatic prediction, 3′-UTR of ATG7 mRNA contains two putative binding sites (Fig. 5a). One of the predicted binding sites (1287–1293 nt) was highly conserved while the other site was relative poorly conserved (Fig. 5b). To further determine whether miR-96-5p directly targets the ATG7 mRNA, reporter constructs (pGL3-ATG7) with putative WT or mutated 3′-UTR downstream of the firefly luciferase were cloned (Fig. 5c). HEK 293T cells were co-transfected with pRL-TK, pGL3-ATG7 (3′-UTR WT or mutant) and miR-96-5p mimic or MMNC. Transfection with miR-96-5p mimic resulted in a significant decrease of the luciferase activity of the reporter vector containing WT 3′-UTR (Fig. 5d). In contrast, the reduction in the luciferase activity was completely abolished after point mutation in miR-96-5p binding site in the 3′-UTR of ATG7 (3′-UTR mutant) (Fig. 5d). These results are consistent with a direct interaction between miR-96-5p and 3′-UTR of ATG7 mRNA.
ATG7 is a direct target of miR-96-5p. a The ATG7 mRNA 3′-UTR contains two binding sites (red letters) for miR-96-5p. b One of the putative binding sites (upper panel) is highly conserved while the other one (lower panel) is less conserved. c Schematic representation of luciferase constructs contains ATG7 3′-UTR with wild type (WT) or mutant type (mutant) miR-96-5p binding site. d Luciferase activity in HEK 293T cells co-transfected with WT or mutant ATG7-luciferase constructs and 50 nM miRNA mimic negative control (MMNC) or miR-96-5p mimic. ***p < 0.001, NS not significant
Forced ATG7 expression restores miR-96-5p-mediated autophagy inhibition and pro-fibrogenic signaling
To testify that miR-96-5p-mediated ATG7 downregulation was responsible for the reduced autophagy activity and hepatic stellate cell activation, a DNA construct expressing ATG7 was used. In our experiments, rapamycin-induced GFP-LC3 puncta formation was suppressed by miR-96-5p mimic but reversed by forced ATG7 expression in LX-2 cells (Fig. 6a, b). In consistency with these results, Western blotting demonstrated increased ATG7 protein levels and LC3-I to LC3-II conversion while decreased SQSTM1/p62 protein expression under ATG7 overexpression (Fig. 6c). Moreover, overexpressed ATG7 promoted pro-fibrogenic signaling as indicated by significantly increased α-SMA and Col1α1 protein abundance (Fig. 6d). These data demonstrate that ATG7 is an important target of miR-96-5p for inhibiting autophagy and hepatic stellate cell activation.
Overexpression of ATG7 reverses miR-96-5p-mediated inhibition of autophagy and hepatic stellate activation. a Fluorescence images of GFP-LC3 puncta formation in LX-2 cells transfected and treated as indicated. b Quantitative analysis of the experiments in a. ***p < 0.001. c Western blot analysis of ATG7, p62, and LC3 protein levels in LX-2 cells treated as described in a. d Western blot analysis of α-SMA and Col1 protein levels in LX-2 cells treated as described in a
Discussion
Liver fibrosis is an integrated cellular response to chronic liver injuries and is characterized by the excessive deposition of ECM proteins [17]. The central event in liver fibrosis is believed to be HSC activation, which makes HSCs transdifferentiated into myofibroblasts that serve as the major source of ECM proteins [1]. Accumulating evidence suggests that liver fibrosis, even cirrhosis is reversible and studies have revealed that the number of activated HSCs was significantly decreased during the regression of hepatic fibrosis [2]. Therefore, preventing HSC activation and promoting the reduction of activated HSCs represent a promising strategy to resolve liver fibrosis. In the current study, we investigated the functional role of miR-96-5p-mediated autophagy regulation in HSC activation.
There is plenty of evidence that miRNAs play a role in liver pathology, and in animal models and human studies, a number of miRNAs have been implicated in the pathogenesis of liver fibrosis [13, 18]. Here, we first observed that miR-96-5p was upregulated during the development of liver fibrosis through microarray and qRT-PCR analyses of fibrotic tissues. In vitro induction of HSC activation by TGF-β1 was then performed and further qRT-PCR analyses revealed that the markers of HSC activation increased preceding miR-96-5p elevation. Western blotting showed that miR-96-5p suppressed the mRNA and proteins levels of α-SMA and Col1α1. miR-96-5p has been associated with cancer progression, auditory system formation, and neuroprotection [19,20,21,22,23]. In idiopathic pulmonary fibrosis, it has been reported that miR-96 contributes to the maintenance of pathological phenotype of fibroblasts and leads to fibrosis progression [24]. Our findings support that miR-96-5p may also play a role in liver fibrosis by inhibiting HSC activation.
Autophagy is a lysosomal degradation process critical for cell differentiation, survival, and homeostasis [25, 26]. Recent studies have highlighted its implication in various human diseases, including hepatic fibrosis [27, 28]. In mouse models, Lodder et al. [29] showed that, by limiting interleukin 1A and 1B production, Kuffer cell autophagy exerts protective effects against liver fibrosis. While in HSCs, evidence points towards autophagy playing a pro-fibrotic role [8,9,10], but how autophagy was regulated during HSC activation is poorly understood. We here introduced miR-96-5p as a regulator of autophagy during this process. Overexpression of miR-96-5p repressed GFP-LC3 puncta formation, LC3-I to LC3-II conversion as well as p62 degradation in LX-2 cells treated with the frequently used autophagy stimulus rapamycin. In addition, miR-96-5p-mediated autophagy suppression was abolished by miR-96-5p-specific inhibitor. Collectively, these data suggest that miR-96-5p is an important miRNA in autophagy regulation.
Autophagy depends on the coordinated interaction of autophagy-related proteins and ATG7 was identified as a target of miR-96-5p in the current study. We demonstrated that miR-96-5p overexpression reduced the protein and mRNA levels of ATG7. More importantly, in addition to bioinformatic analysis, we observed a decreased luciferase activity of reporter gene containing the wild-type ATG7 3′UTR, while introduction of mutations to the sequence abrogated the inhibitory effect. Consistent with our results, Ma et al. [15] also confirmed that miR-96-5p regulates autophagy by direct targeting ATG7 in prostate cancer cell line under hypoxia. Taken together, these results indicate that ATG7 is a direct target of miR-96-5p.
In our “rescue experiments,” miR-96-5P-mediated autophagy blockage was reversed by reintroduction of ATG7 in LX-2 cells. Forced expression of ATG7 restored GFP-LC3 puncta formation, LC3-I to LC3-II conversion, and p62 degradation. Besides, the inhibitory effect of miR-96-5p on HSC activation was also abolished, since an enhanced α-SMA and Col1α1 protein expression was detected with the reintroduction of ATG7.
In conclusion, our study revealed that miR-96-5p is a suppressor of HSC activation and this inhibitory effect mainly depends on miR-96-5p-mediated autophagy repression. Moreover, ATG7 was identified as a direct target of miR-96-5p (Fig. 7). However, since the present study was performed in vitro and may not reflect the actual conditions in vivo, further studies are required to expand these findings in primary HSCs and rodent models. If reproduced, then regulating miR-96-5p may represent a promising therapeutic strategy for liver fibrosis.
References
Friedman SL (2008) Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiol Rev 88:125–172
Yin C, Evason KJ, Asahina K, Stainier DY (2013) Hepatic stellate cells in liver development, regeneration, and cancer. J Clin Invest 123:1902–1910
Lee UE, Friedman SL (2011) Mechanisms of hepatic fibrogenesis. Best Pract Res Clin Gastroenterol 25:195–206
Bataller R, Brenner DA (2005) Liver fibrosis. J Clin Invest 115:209–218
Thoen LF, Guimaraes EL, Grunsven LA (2012) Autophagy: a new player in hepatic stellate cell activation. Autophagy 8:126–128
Mallat A, Lodder J, Teixeira-Clerc F, Moreau R, Codogno P, Lotersztajn S (2014) Autophagy: a multifaceted partner in liver fibrosis. Biomed Res Int 2014:869390
Hsu CC, Schwabe RF (2011) Autophagy and hepatic stellate cell activation—partners in crime? J Hepatol 55:1176–1177
Hernandez-Gea V, Friedman SL (2012) Autophagy fuels tissue fibrogenesis. Autophagy 8:849–850
Thoen LF, Guimaraes EL, Dolle L, Mannaerts I, Najimi M, Sokal E, van Grunsven LA (2011) A role for autophagy during hepatic stellate cell activation. J Hepatol 55:1353–1360
Hernandez-Gea V, Ghiassi-Nejad Z, Rozenfeld R, Gordon R, Fiel MI, Yue Z, Czaja MJ, Friedman SL (2012) Autophagy releases lipid that promotes fibrogenesis by activated hepatic stellate cells in mice and in human tissues. Gastroenterology 142:938–946
Hernandez-Gea V, Hilscher M, Rozenfeld R, Lim MP, Nieto N, Werner S, Devi LA, Friedman SL (2013) Endoplasmic reticulum stress induces fibrogenic activity in hepatic stellate cells through autophagy. J Hepatol 59:98–104
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Yu K, Shi G, Li N (2015) The function of microRNA in hepatitis B virus-related liver diseases: from dim to bright. Ann Hepatol 14:450–456
Xu J, Wang Y, Tan X, Jing H (2012) MicroRNAs in autophagy and their emerging roles in crosstalk with apoptosis. Autophagy 8:873–882
Ma Y, Yang HZ, Dong BJ, Zou HB, Zhou Y, Kong XM, Huang YR (2014) Biphasic regulation of autophagy by miR-96 in prostate cancer cells under hypoxia. Oncotarget 5:9169–9182
Bjorkoy G, Lamark T, Brech A, Outzen H, Perander M, Overvatn A, Stenmark H, Johansen T (2005) p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J Cell Biol 171:603–614
Friedman SL (2010) Evolving challenges in hepatic fibrosis. Nat Rev Gastroenterol Hepatol 7:425–436
Szabo G, Bala S (2013) MicroRNAs in liver disease. Nat Rev Gastroenterol Hepatol 10:542–552
Hong Y, Liang H, Uzair Ur R, Wang Y, Zhang W, Zhou Y, Chen S, Yu M, Cui S, Liu M et al (2016) miR-96 promotes cell proliferation, migration and invasion by targeting PTPN9 in breast cancer. Sci Rep 6:37421
Baik SH, Lee J, Lee YS, Jang JY, Kim CW (2016) ANT2 shRNA downregulates miR-19a and miR-96 through the PI3K/Akt pathway and suppresses tumor growth in hepatocellular carcinoma cells. Exp Mol Med 48:e222
Guo H, Li Q, Li W, Zheng T, Zhao S, Liu Z (2014) MiR-96 downregulates RECK to promote growth and motility of non-small cell lung cancer cells. Mol Cell Biochem 390:155–160
Kuhn S, Johnson SL, Furness DN, Chen J, Ingham N, Hilton JM, Steffes G, Lewis MA, Zampini V, Hackney CM et al (2011) miR-96 regulates the progression of differentiation in mammalian cochlear inner and outer hair cells. Proc Natl Acad Sci U S A 108:2355–2360
Kinoshita C, Aoyama K, Matsumura N, Kikuchi-Utsumi K, Watabe M, Nakaki T (2014) Rhythmic oscillations of the microRNA miR-96-5p play a neuroprotective role by indirectly regulating glutathione levels. Nat Commun 5:3823
Nho RS, Im J, Ho YY, Hergert P (2014) MicroRNA-96 inhibits FoxO3a function in IPF fibroblasts on type I collagen matrix. Am J Physiol Lung Cell Mol Physiol 307:L632–L642
Min L, Choy E, Pollock RE, Tu C, Hornicek F, Duan Z (2017) Autophagy as a potential target for sarcoma treatment. Biochim Biophys Acta 1868:40–50
Mizushima N, Komatsu M (2011) Autophagy: renovation of cells and tissues. Cell 147:728–741
Rautou PE, Mansouri A, Lebrec D, Durand F, Valla D, Moreau R (2010) Autophagy in liver diseases. J Hepatol 53:1123–1134
Hilscher M, Hernandez-Gea V, Friedman SL (2012) Autophagy and mesenchymal cell fibrogenesis. Biochim Biophys Acta 1831:972–978
Lodder J, Denaes T, Chobert MN, Wan J, El-Benna J, Pawlotsky JM, Lotersztajn S, Teixeira-Clerc F (2015) Macrophage autophagy protects against liver fibrosis in mice. Autophagy 11:1280–1292
Funding
This work was supported by the National Natural Science Foundation of China [grant numbers 81770565, 81371821].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors have nothing to disclose.
Rights and permissions
About this article
Cite this article
Yu, K., Li, N., Cheng, Q. et al. miR-96-5p prevents hepatic stellate cell activation by inhibiting autophagy via ATG7. J Mol Med 96, 65–74 (2018). https://doi.org/10.1007/s00109-017-1593-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-017-1593-6