Abstract
Tyrosyl-DNA phosphodiesterase 1 (TDP1) is a DNA repair enzyme that can reduce the efficacy of some anticancer drugs targeting topoisomerase 1 (TOP1) making it a promising target for antitumor therapy when combined with TOP1 poisons. Here we describe the synthesis of a number of adamantane-monoterpene conjugates 20a–g and 21a–g connected through a 1,3,4-thiadiazol-2(3H)-imine linker, where acyclic, monocyclic, and bicyclic structural types of monoterpenes were used. All the synthesized compounds demonstrated activity against TDP1 in micromolar range, with the most potent inhibitor being compound 21a (IC50 1.2 μM). The cytotoxic effects of these compounds determined in the HEK293A and HeLa cell lines were low to moderate. These findings imply that such compounds are promising for further development of new TDP1 inhibitors with favorable physicochemical properties.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
One of the enzymes involved in the control and modification of topological states of DNA is Topoisomerase 1 (TOP1). It plays a crucial role in processes like transcription, DNA replication, recombination, and chromatin remodeling [1, 2]. The catalytic mechanism of this enzyme involves the introduction of a temporary single-stranded break in DNA, which results in the rotation of the free DNA strand and the release of topological stress. Once DNA is relaxed, TOP1 religates the breaks by reversing its covalent binding.
Inhibition of the reverse cleavage of TOP1 is an underlying mechanism of action of some anticancer drugs. Camptothecin, isolated from the bark of the Camptotheca acuminata (Nyssaceae), was the first known TOP1 inhibitor. Antitumor drugs such as topotecan, irinotecan, and belotecan are semi-synthetic derivatives of camptothecin and are used in the treatment of a wide range of cancers, including leukemia, breast cancer, ovarian cancer, lung cancer etc. [3].
Chemotherapeutics, including camptothecin derivatives, have a number of disadvantages that can reduce its effectiveness and cause side effects such as diarrhea, kidney failure, neutropenia and thrombocytopenia. In addition, some tumors may become resistant to chemotherapy drugs, which makes treatment more difficult. Tyrosyl-DNA-phosphodiesterase 1 (TDP1) is a DNA repair enzyme, responsible for the cleavage of covalent TOP1-DNA complexes, including those resulting from the action of camptothecin-derived drugs [4]. Several studies have shown that the TDP1 enzyme mediates the drug resistance of some cancers [5]. Cell cultures with increased TDP1 expression are known to show resistance to TOP1 inhibitors [6], while cells with a low content of this enzyme are usually sensitive to TOP1 inhibitors [7]. A similar effect is also observed in animal models, a line of genetically modified TDP1 knockout mice showed hypersensitivity to camptothecin [8, 9]. Thus, the development of TDP1 inhibitors could lead to drugs that effectively enhance antitumor therapy.
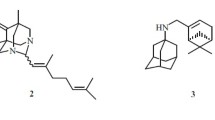
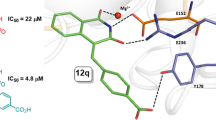
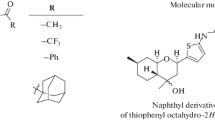
Currently, a number of compounds demonstrating inhibitory properties against TDP1 are known, such as thiazolidine-2,4-dione derivatives 1 and 2 containing aromatic and monoterpene moieties [10], compounds with usnic acid backbone 3 [11], deoxycholic acid derivatives 4 [12], isoquinolines 5 [13], benzophenanthridinones 6 [14] and a number of others [15,16,17] (Fig. 1).
Substances (for instance 7–11 in Fig. 2.) combining monoterpene and adamantane fragments through linkers of various types have also shown their effectiveness in inhibiting TDP1 [18, 19]. Furthermore, adamantane derivatives 12 and 14 having 1,2,4-triazole and 1,3,4-thiadiazole fragments not only exerted anti-TDP1 properties, but also demonstrated a synergistic effect on cancer cells when combined with topotecan [20, 21] (Fig. 2).
In this paper, we continued our research on the synthesis of adamantane-monoterpene conjugates connected via a heterocyclic core and the study of their inhibitory activity against TDP1; i.e., we synthesized a number of (1,3,4-thiadiazole-2(3H)-ylidene)-2,2,2-trifluoroacetamides containing an adamantane scaffold in the fifth position of the heterocyclic nucleus and having a monoterpene substituent of different nature in the third position, as well as their analogs without the trifluoroacetyl group. There are some studies showing that compounds containing a 1,3,4-thiadiazol-2(3H)-imine core exhibit valuable pharmacological activities (antitubercular [22], antiradical [23] etc.), to the best of our knowledge, there are no data regarding inhibitory properties of substances with 1,3,4-thiadiazol-2(3H)-imine core against TDP1.
Chemistry
For the synthesis of target products acyl chloride 15 was converted into compound 16 by its interaction with thiosemicarbazide in THF with 90% yield at first. Compound 16 then underwent intramolecular cyclization in concentrated sulfuric acid at room temperature to form corresponding 2-amino-1,3,4-thiadiazole 17 as described in [20]. Subsequent treatment of compound 17 with trifluoroacetic anhydride in DCM under cooling conditions afforded trifluoroacetamide 18 with 87% yield. Reaction of compound 18 with bromides 19a–f, obtained by the treatment of corresponding monoterpene alcohols with PBr3 or the NBS/PPh3 mixture, in DMF in the presence of potassium carbonate and sodium iodide at room temperature gave N-alkylated products 20a–f with good to excellent yields. Trifluoroacetamides 20a–f were then transformed into corresponding imines 21a–f by alkaline hydrolysis using LiOH in the THF/H2O biphasic medium at 60 °C.
Biology
The compounds synthesized were tested against TDP1 using a fluorescent biosensor [24]. Furamidine, a commercially available TDP1 inhibitor (IC50 1.23 µM), was used as a reference. All the compounds were shown to exhibit activity in the low micromolar concentration range. The most effective inhibitor is compound 21a containing a 3,7-dimethyloctyl substituent (Table 1). It is interesting to note that there is no clear trend between trifluoroacetamide derivatives and imines in the values of semi-inhibitory concentrations. On one hand, compound 21a turned out to be a more effective inhibitor compared to its precursor 20a; a similar trend of 21 being more potent TDP1 inhibitor than 20 is observed in pairs 21d–20d, 21e–20e, 21f–20f. On the other hand, when comparing the values of semi-inhibitory concentrations in pairs 21b–20b and 21c–20c, one can conclude that trifluoroacetyl group has slight positive effect on the ability to inhibit TDP1.
When comparing cytotoxic properties of compounds synthesized, one can conclude that the presence of trifluoroacetyl group significantly reduces CC50 values of a substance (Table 1, Fig. S1 in Supplementary). This correlation can be seen in almost all pairs of compounds such as 21b–20b, 21d–20d, 21e–20e, 21f–20f and is observed on both non-tumor cell lines: HEK293A (human embryonic kidney) and MRC-5 (human lung fibroblasts) and cancer cell line HeLa (cervical carcinoma).
Knowing the intrinsic cytotoxicity of the compounds is important in assessing the potential side effects of the concomitant therapy. TDP1 inhibitors should have minimal intrinsic toxicity as they are envisaged to be topotecan sensitizers. The cytotoxic effect of the compounds had moderate or low toxicity (20a–f, 21a–c) on HEK293A, MRC-5, and HeLa cells. None of the compounds potentiated the effect of topotecan, regardless of the inhibitory potency and intrinsic cytotoxicity, neither on cancer cells (HeLa) nor on non-tumor cells (HEK293A, MRC-5) (see Fig. S2 in supplementary).
Modeling
Twelve 1,3,4-thiadiazole derivatives were docked into the binding site of the TDP1 (PDB ID: 6W7K, resolution 1.70 Å) [25] enzyme. The TDP1 docking scaffold has been previously found to be robust [26]. The scoring functions GoldScore(GS) [27], ChemScore(CS) [28, 29], ChemPLP(Piecewise Linear Potential) [30] and ASP(Astex Statistical Potential) [31] in the GOLD (v2020.2.0) docking algorithm were used. The GOLD docking algorithm is known to be an excellent molecular modeling tool [32, 33].
The binding scores, for the TDP1 catalytic pocket, are given in Table S1; all the ligands have reasonable values. When the scores of the active ligands were correlated against their IC50 values, only weak trends were seen for ASP (R2 – 0.144), ChemPLP (R2 – 0.133), GS (R2 – 0.080) and CS (R2 – 0.263, see Fig. S3).
The predicted binding poses of the thiadiazole derivatives were examined; no dominant binding poses were found by the four scoring functions. ChemPLP and GS predicted similar poses for 20a,b,f, 21a–c. All the ligands overlapped with the co-crystallized ligand as well as occupying the catalytic pocket including the His263 and His493 amino acid residues. The adamantane group was predicted to bind to the same pocket as the phthalic acid of the co-crystallized structure by all four scoring functions. The binding mode, as predicted by ChemPLP, of the ligand with the best overall properties 20f is shown in Fig. 3. The aliphatic ring docked into a pocket below the adamantane binding pocket and the trifluoroacetimide extends towards the allosteric site but does not reach it (see Fig. 3A). The sulfur of the thiadiazole ring forms a hydrogen bond with the OH of Tyr204. His263 is predicted to form a π-alkyl interaction with the adamantane group (see Fig. 3B).
The docked pose of 20f in the catalytic site of TDP1 as predicted by the ChemPLP scoring function. A The ligand is shown in the ball-and-stick format. The catalytic pocket is to the right-hand site, the co-crystallized ligand is depicted as green sticks. The allosteric pocket is not predicted to be occupied (see circled area) although the trifluoroacetamide reaches towards it. The protein surface is rendered; blue depicts regions with a partial positive charge on the surface; red depicts regions with a partial negative charge and gray shows neutral areas. B The predicted configuration depicted as balls-and-sticks, H-boding is seen between the sulfur atom of the thiadiazole ring and the OH of the side chain of Tyr204 (stick format). The catalytic His263 amino acid residue is shown as sticks. The adjacent amino acids (<5 Å), buttressing the ligand, are shown as lines. The amino acids’ hydrogens on are not shown for clarity
Molecular dynamics simulations have suggested that the TDP1 inhibitors occupy an allosteric binding pocket next to the catalytic site as shown in Fig. 3A [34]. Molecular modeling and structural activity relations studies of usnic acid derivatives confirm the existence of this allosteric site and its occupancy having a beneficial effect on the overall binding efficacy [35]. None of the ligands were predicted to occupy the allosteric pocket explaining the relatively modest binding affinity observed.
Chemical space
The calculated molecular descriptors MW (molecular weight), log P (water-octanol partition coefficient), HD (hydrogen bond donors), HA (hydrogen bond acceptors), PSA (polar surface area) and RB (rotatable bonds) are given in Table S2. The values of the molecular descriptors lie in lead-like chemical space for PSA and HD, for HA in both lead- and drug-like space and in drug-like space for RB and MW. Finally, log P spans between drug-like space and beyond Known Drug Space (for the definition of lead-like, drug-like and Known Drug Space (KDS) regions see ref. [36] and Table S3). No correlations were found between the IC50 values of the active ligands and their descriptors.
The Known Drug Indexes (KDIs) for the ligands were calculated to gauge the balance of the molecular descriptors (MW, log P, HD, HA, PSA and RB) (for details see supporting materials). The KDI2a values for the ligands range from 3.43 to 4.36 with a theoretical maximum of 6 and the average of 4.08 (± 1.27) for known drugs and the values are quite good.
Both chemical series 20 and 21 contain electron rich imine moiety rendering them susceptible to an electrophilic attack. To check the overall redox stability of the ligands 20f and 21f were chosen and their ionization potentials (one-electron oxidation) and electron affinity (one-electron reduction) were calculated using density functional theory (DFT) and compared to the statistical distribution of known drugs [37]. The ionization potentials are 7.9 and 7.2 eV, respectively and 95% of drugs lie in the 6.0–9.0 eV range; the electron affinities are −0.7 and 0.3 eV with known drugs in the −1.5–2.0 eV range [37]. Thus, 20f and 21f are within the ranges of known drugs and are not expected to be excessively reactive.
Methodology
Molecular modeling
The compounds were docked against the crystal structure of TDP1 (PDB ID: 6W7K, resolution 1.70 Å) [25], which were obtained from the Protein Data Bank (PDB) [38, 39]. The GOLD (v2020.2.0) software suite was used to prepare the crystal structures for docking, i.e., the hydrogen atoms were added, water molecules deleted and the co-crystallized ligands identified: 4-[(2-phenylimidazo[1,2-a]pyridin-3-yl)amino]benzene-1,2-dicarboxylic acid (TG7). The Scigress version FQ 3.4.4 program [40] software suite was used to build the ligands and the MM3 [41,42,43] force field was used to identify the global minimum using the CONFLEX method [44] followed by structural optimization. The docking center for the TDP1 catalytic pocket was defined as the position of the co-crystallized ligand TG7 with 10 Å radius. Fifty docking runs were allowed for each ligand with default search efficiency (100%). The basic amino acids lysine and arginine were defined as protonated. Furthermore, aspartic and glutamic acids were assumed to be deprotonated. The GoldScore(GS) [27] and ChemScore(CS) [28, 29] ChemPLP(Piecewise Linear Potential) [30] and ASP(Astex Statistical Potential) [31] scoring functions were used to predict the binding modes and relative binding energies of the ligands using the GOLD v2020.2.0 software suite.
The QikProp 6.2 [45] software package was used to calculate the molecular descriptors of the molecules. The reliability of QikProp for the calculated descriptors is established [46]. The Known Drug Indexes (KDI) were calculated from the molecular descriptors as described by Eurtivong and Reynisson [47]. For application in Excel, columns for each property were created and the following equations used do derive the KDI numbers for each descriptor: KDI MW: = EXP(-((MW-371.76)^2)/(2*(112.76^2))), KDI Log P: =EXP(-((LogP-2.82)^2)/(2*(2.21^2))), KDI HD: =EXP(-((HD-1.88)^2)/(2*(1.7^2))), KDI HA: =EXP(-((HA-5.72)^2)/(2*(2.86^2))), KDI RB = EXP(-((RB-4.44)^2)/(2*(3.55^2))), and KDI PSA: =EXP(-((PSA-79.4)^2)/(2*(54.16^2))). These equations could simply be copied into Excel and the descriptor name (e.g., MW) substituted with the value in the relevant column. To derive KDI2A, this equation was used: = (KDI MW + KDI LogP + KDI HD + KDI HA + KDI RB + KDI PSA) and for KDI2B: = (KDI MW × KDI LogP × KDI HD × KDI HA × KDI RB × KDI PSA).
The Gaussian 16 software suite [48] was used with unrestricted DFT. The B3LYP functional hybrid approach was employed [49,50,51] and standard 6–31+G(d,p) diffused basis set [52, 53] was used for geometry optimization and frequency analysis (keywords: opt freq). The zero-point vibrational energies (ZPE) were scaled according to Wong (0.9804) [54]. In all cases, normal modes revealed no imaginary frequencies indicating that they represent minima on the potential energy surface. The subsequent energy calculations were then performed with the larger 6–311+G(2df, p) basis set. Adiabatic ionization potentials (IP) and adiabatic electron affinities (EA) were calculated as described in Forseman and Frisch [55]. The energies and ZPA are given in Table S4 Scheme 1.
Chemistry
All chemicals were purchased from commercial sources (Sigma Aldrich (St. Louis, MO, USA)), Acros Organics (Geel, Belgium) and used without further purification. 1H and 13C NMR spectra were recorded on Bruker AV-400 (Bruker Corporation, Billerica, MA, USA) (400.13 MHz and 100.61 MHz) and Bruker DRX-500 (Bruker Corporation, Billerica, MA, USA) (500.13 MHz and 125.76 MHz). Mass spectra (70 eV) were recorded on a DFS Thermo Scientific high-resolution mass spectrometer. A PolAAr 3005 polarimeter (Optical Activity, Ramsey, UK) was used to measure optical rotations [α]D. Melting points were measured on a Mettler Toledo FP900 Thermosystem apparatus (Mettler Toledo, Cornellà de Llobregat, Spain). Merck silica gel (Merck, Darmstadt, Germany, 63–200 µm) was used for column chromatography. Conversion of the starting materials was detected by Thin Layer Chromatography (ALUGRAM SIL G/UV254, Duren, Germany) using hexane-EtOAc (3:1) solvent mixture. Spectral and analytical measurements were carried out at the Multi-Access Chemical Service Center of Siberian Branch of Russian Academy of Sciences (SB RAS).
Synthesis of 2-(adamantane-1-carbonyl)hydrazine-1-carbothioamide 16
To a suspension of thiosemicarbazide (10.1 g, 110.8 mmol) in 200 ml of THF cooled to 0 °C was added a solution of 1-adamantanecarboxylic acid chloride 15 (10.0 g, 50.4 mmol) in 30 ml THF. The mixture was stirred at room temperature overnight; the solvent was evaporated under reduced pressure. To the residue was added water, the precipitate was filtered off, washed with water several times and dried, which afforded 11.5 g (90%) of compound 16 as a white solid. 1H NMR (400 MHz, DMSO-d6) δ 1.58–1.72 (m, 6H), 1.76–1.87 (m, 6H), 1.91–2.05 (m, 3H), 6.99 (s, 1H), 7.82 (s, 1H), 9.09 (s, 1H), 9.39 (s, 1H). 13C NMR (101 MHz, DMSO) δ 181.8, 176.3, 39.6, 38.0, 36.1, 27.5.
Synthesis of 5-(adamantan-1-yl)-1,3,4-thiadiazol-2-amine 17
Compound 16 (1.1 g, 4.7 mmol) was added to 20 ml of concentrated sulfuric acid. The solution was stirred overnight at room temperature. The mixture was poured into ice and then neutralized carefully with aqueous ammonia until pH 7-8. The resulting solid was filtered off, washed with water and dried. Recrystallization from ethanol gave the title compound as a pale yellow solid (0.8 g, 78%). 1H NMR (300 MHz, DMSO-d6) δ 1.66–1.79 (m, 6H), 1.84–1.96 (m, 6H), 1.97–2.09 (m, 3H), 6.95 (s, 2H). 13C NMR (126 MHz, CDCl3) δ 168.6, 167.7, 42.9, 37.4, 36.1, 28.0.
Synthesis of N-(5-(adamantan-1-yl)-1,3,4-thiadiazol-2-yl)-2,2,2-trifluoroacetamide 18
To a suspension of amine (1.00 g, 4.26 mmol) and triethylamine (0.66 ml, 4.74 mmol) in 20 ml of anhydrous DCM cooled to 0 °C was slowly added trifluoroacetic acid anhydride (0.66 ml, 4.74 mmol). The resulting solution was stirred at room temperature for 1 h, then the solvent was removed under reduced pressure and a saturated solution of NaHCO3 was added. The solid was filtered off, washed with water and dried. The product was isolated as a white solid (1.23 g, 87%). 1H NMR (300 MHz, Chloroform-d) δ 1.69–1.86 (m, 6H), 1.99–2.08 (m, 6H), 2.08–2.20 (m, 3H). 13C NMR (126 MHz, CDCl3) δ 173.6, 167.1, 162.1 (q, J = 38.4 Hz), 116.3 (q, J = 286.1 Hz), 42.5, 38.2, 36.1, 28.1. HRMS: m/z 331.0957 (M+ C14H16O1N3F3S1+, calc. 331.0961).
General procedure for the synthesis of compounds 20a–f
A mixture containing trifluoroacetamide 18 (0.90 g, 2.72 mmol), appropriate bromide (2.74 mmol), potassium carbonate (0.45 g, 3.26 mmol) and sodium iodide (0.05 g, 0.33 mmol) in 9 ml of DMF was stirred at room temperature overnight. The mixture was then diluted with water and the product was extracted with ethyl acetate. The combined organic phase was washed with brine and dried over sodium sulfate. The product was isolated by column chromatography on silica gel (eluent hexane/ethylacetate).
N-((Z)-5-(Adamantan-1-yl)-3-(3,7-dimethyloctyl)-1,3,4-thiadiazol-2(3H)-ylidene)-2,2,2-trifluoroacetamide 20a
Colorless oil, yield 90%
1H NMR (300 MHz, Chloroform-d) δ 0.84 (d, J = 6.6 Hz, 6H), 0.94 (d, J = 6.4 Hz, 3H), 1.05–1.17 (m, 3H), 1.17–1.24 (m, 1H), 1.24–1.40 (m, 3H), 1.48 (dq, J = 13.3, 6.7 Hz, 1H), 1.59–1.70 (m, 1H), 1.70–1.84 (m, 6H), 1.84–1.94 (m, 1H), 1.94–2.04 (m, 6H), 2.10 (q, J = 3.1 Hz, 3H), 4.30–4.50 (m, 2H). 13C NMR (101 MHz, CDCl3) δ 170.4, 166.9, 165.3 (q, J = 36.8 Hz), 117.1 (q, J = 285.7 Hz), 49.5, 42.3, 39.1, 38.3, 36.7, 36.2, 35.2, 30.2, 28.2, 28.0, 24.6, 22.7, 22.6, 19.4. HRMS: m/z 471.2526 (M+ C24H36O1N3F3S1+, calc. 471.2523).
N-((Z)-5-(Adamantan-1-yl)-3-((S)-3,7-dimethyloct-6-en-1-yl)-1,3,4-thiadiazol-2(3H)-ylidene)-2,2,2-trifluoroacetamide 20b
Colorless oil, yield 88%
1H NMR (300 MHz, Chloroform-d) δ 0.95 (d, J = 6.3 Hz, 3H), 1.12–1.29 (m, 1H), 1.30–1.49 (m, 2H), 1.57 (s, 3H), 1.62–1.69 (m, 3H), 1.69–1.84 (m, 7H), 1.84–1.96 (m, 2H), 1.96–2.02 (m, 7H), 2.07–2.16 (m, 3H), 4.30–4.50 (m, 2H), 5.04 (t, J = 7.1 Hz, 1H). 13C NMR (126 MHz, CDCl3) δ 170.15, 166.71, 165.14 (q, J = 36.8 Hz), 131.29, 124.19, 116.93 (d, J = 285.8 Hz), 49.30, 42.14, 38.19, 36.46, 36.04, 34.91, 29.70, 28.04, 25.51, 25.13, 19.09, 17.43. HRMS: m/z 469.2369 (M+ C24H34O1N3F3S1+, calc. 469.2366). \(\left[ \alpha \right]_D^{22.7}\) = +4.5 (c 1.65 in CHCl3).
N-((Z)-5-(Adamantan-1-yl)-3-((E)-3,7-dimethylocta-2,6-dien-1-yl)-1,3,4-thiadiazol-2(3H)-ylidene)-2,2,2-trifluoroacetamide 20c
Yellow oil, yield 80%
1H NMR (500 MHz, Chloroform-d) δ 1.54 (s, 3H), 1.61 (s, 3H), 1.70–1.81 (m, 6H), 1.81–1.84 (m, 3H), 1.95–1.99 (m, 6H), 2.00–2.07 (m, 4H), 2.07–2.12 (m, 3H), 4.94 (d, J = 7.4 Hz, 2H), 4.98–5.03 (m, 1H), 5.36 (t, J = 7.4 Hz, 1H). 13C NMR (126 MHz, CDCl3) δ 170.2, 166.3, 165.0 (q, J = 36.7 Hz), 143.1, 131.7, 123.4, 116.9 (q, J = 285.8 Hz), 116.1, 48.8, 42.0, 39.3, 38.1, 36.0, 28.0, 26.0, 25.5, 17.4, 16.4. HRMS: m/z 467.2213 (M+ C24H32O1N3F3S1+, calc. 467.2210).
N-((Z)-5-(Adamantan-1-yl)-3-(2-((S)-2,2,3-trimethylcyclopent-3-en-1-yl)ethyl)-1,3,4-thiadiazol-2(3H)-ylidene)-2,2,2-trifluoroacetamide 20d
White solid, mp 61.6–62.3 °C, yield 86%
1H NMR (400 MHz, Chloroform-d) δ 0.77 (s, 3H), 0.93 (s, 3H), 1.54–1.60 (m, 3H), 1.60–1.69 (m, 1H), 1.70–1.84 (m, 7H), 1.84–1.94 (m, 1H), 1.96–2.01 (m, 6H), 2.02–2.08 (m, 1H), 2.09–2.16 (m, 3H), 2.34 (dddd, J = 15.2, 7.6, 3.0, 1.6 Hz, 1H), 4.35 (ddd, J = 13.2, 8.6, 4.8 Hz, 1H), 4.46 (dt, J = 13.1, 7.8 Hz, 1H), 5.15–5.26 (m, 1H). 13C NMR (75 MHz, CDCl3) δ 170.3, 166.8, 165.2 (q, J = 36.8 Hz), 148.2, 121.4, 116.9 (q, J = 285.6 Hz), 50.6, 47.1, 46.8, 42.2, 38.2, 36.0, 35.0, 28.7, 28.0, 25.6, 19.5, 12.5. HRMS: m/z 467.2213 (M+ C24H32O1N3F3S1+, calc. 467.2215). \(\left[ \alpha \right]_D^{23.7}\) = −5.5 (c 0.61 in CHCl3).
N-((Z)-5-(Adamantan-1-yl)-3-(((1 R)-6,6-dimethylbicyclo[3.1.1]hept-2-en-2-yl)methyl)-1,3,4-thiadiazol-2(3H)-ylidene)-2,2,2-trifluoroacetamide 20e
Yellowish oil, yield 84%
1H NMR (500 MHz, Chloroform-d) δ 0.60 (s, 3H), 1.14 (d, J = 8.7 Hz, 1H), 1.18 (s, 3H), 1.68–1.82 (m, 6H), 1.89–2.00 (m, 6H), 2.00–2.06 (m, 1H), 2.07–2.14 (m, 4H), 2.16–2.31 (m, 2H), 2.34 (dt, J = 8.7, 5.7 Hz, 1H), 4.80–4.89 (m, 2H), 5.48–5.56 (m, 1H). 13C NMR (126 MHz, CDCl3) δ 169.8, 166.9, 165.1 (q, J = 36.8 Hz), 141.1, 123.2, 116.8 (q, J = 285.7 Hz), 55.3, 43.7, 42.0, 40.2, 38.1, 37.9, 35.9, 31.2, 31.1, 27.9, 25.8, 20.7. HRMS: m/z 465.2056 (M+ C24H30O1N3F3S1+, calc. 465.2053). \(\left[ \alpha \right]_D^{22.7}\) = −9.9 (c 2.27 in CHCl3).
N-((Z)-5-(Adamantan-1-yl)-3-(2-((1 R)-6,6-dimethylbicyclo[3.1.1]hept-2-en-2-yl)ethyl)-1,3,4-thiadiazol-2(3H)-ylidene)-2,2,2-trifluoroacetamide 20f
Colorless oil, yield 81%
1H NMR (400 MHz, Chloroform-d) δ 0.74 (s, 3H), 0.98 (d, J = 8.6 Hz, 1H), 1.24 (s, 3H), 1.68–1.83 (m, 6H), 1.93–1.99 (m, 6H), 1.99–2.04 (m, 1H), 2.05–2.15 (m, 6H), 2.31 (dt, J = 8.6, 5.6 Hz, 1H), 2.38–2.51 (m, 1H), 2.53–2.65 (m, 1H), 4.32 (ddd, J = 13.2, 8.0, 5.5 Hz, 1H), 4.45 (dt, J = 13.2, 7.6 Hz, 1H), 5.22 (tt, J = 3.1, 1.5 Hz, 1H). 13C NMR (126 MHz, CDCl3) δ 170.0, 166.7, 165.0 (q, J = 36.7 Hz), 143.4, 119.4, 116.9 (q, J = 285.6 Hz), 49.1, 45.1, 42.1, 40.3, 38.1, 37.7, 36.0, 35.1, 31.6, 31.2, 27.9, 26.0, 20.8. HRMS: m/z 479.2223 (M+ C25H32O1N3F3S1+, calc. 479.2209). \(\left[ \alpha \right]_D^{22.7}\) = −2.9 (c 1.60 in CHCl3).
General procedure for the synthesis of compounds 21a-f
To a solution of appropriate compound 20 in 10 ml of THF was added an aqueous solution of LiOH (prepared by dissolving 0.84 g (20 mmol) of LiOH*H2O in 10 ml of water). The mixture was stirred at 60 °C for 10 h. After completion of the reaction (detected by gas chromatography), the organic phase was separated and the product was extracted with diethyl ether. The combined organic phase was washed with brine and dried over sodium sulfate. The product was isolated by column chromatography.
5-(Adamantan-1-yl)-3-(3,7-dimethyloctyl)-1,3,4-thiadiazol-2(3H)-imine 21a
White solid, mp 92.1–92.9 °C, yield 71%
1H NMR (400 MHz, Chloroform-d) δ 0.83 (d, J = 6.6, 6H), 0.90 (d, J = 6.4 Hz, 3H), 1.06–1.15 (m, 3H), 1.16–1.35 (m, 2H), 1.40–1.57 (m, 3H), 1.64–1.80 (m, 8H), 1.81–1.86 (m, 6H), 2.00–2.06 (m, 3H), 3.83 (t, J = 7.3 Hz, 2H). 13C NMR (126 MHz, CDCl3) δ 163.4, 156.8, 45.4, 41.7, 39.0, 37.8, 36.9, 36.2, 34.7, 30.4, 28.1, 27.8, 24.5, 22.6, 22.5, 19.4. HRMS: m/z 375.2703 (M+ C22H37N3S1+, calc. 375.2697).
5-(Adamantan-1-yl)-3-((S)-3,7-dimethyloct-6-en-1-yl)-1,3,4-thiadiazol-2(3H)-imine 21b
Colorless oil, yield 67%
1H NMR (400 MHz, Chloroform-d) δ 0.92 (d, J = 6.3 Hz, 3H), 1.16 (dddd, J = 13.4, 9.3, 7.5, 6.0 Hz, 1H), 1.31–1.42 (m, 1H), 1.42–1.55 (m, 2H), 1.57 (d, J = 1.3 Hz, 3H), 1.65 (t, J = 1.3 Hz, 3H), 1.66–1.80 (m, 7H), 1.80–1.88 (m, 6H), 1.88–2.00 (m, 2H), 2.01–2.07 (m, 3H), 3.75–3.92 (m, 2H), 5.01–5.12 (m, 1H). 13C NMR (126 MHz, CDCl3) δ 163.3, 156.7, 131.0, 124.6, 45.3, 41.7, 37.8, 36.8, 36.2, 34.6, 30.0, 28.1, 25.6, 25.3, 19.2, 17.5. HRMS: m/z 373.2546 (M+ C22H35N3S1+, calc. 373.2543). \(\left[ \alpha \right]_D^{24.7}\) = +2.5 (c 0.40 in CHCl3).
5-(Adamantan-1-yl)-3-((E)-3,7-dimethylocta-2,6-dien-1-yl)-1,3,4-thiadiazol-2(3H)-imine 21c
Colorless oil, yield 69%
1H NMR (400 MHz, Chloroform-d) δ 1.57 (s, 3H), 1.64–1.66 (m, 3H), 1.66–1.71 (m, 3H), 1.71–1.74 (m, 5H), 1.74–1.77 (m, 1H), 1.82–1.85 (m, 6H), 1.98–2.12 (m, 7H), 4.43 (d, J = 6.6 Hz, 2H), 5.06 (ddt, J = 6.9, 5.3, 1.5 Hz, 1H), 5.34 (tq, J = 6.7, 1.3 Hz, 1H). 13C NMR (126 MHz, CDCl3) δ 163.4, 157.0, 139.6, 131.4, 123.8, 118.5, 45.1, 41.7, 39.3, 37.8, 36.2, 36.1, 28.1, 26.2, 17.6, 16.4. HRMS: m/z 371.2390 (M+ C22H33N3S1+, calc. 371.2392).
5-(Adamantan-1-yl)-3-(2-((S)-2,2,3-trimethylcyclopent-3-en-1-yl)ethyl)-1,3,4-thiadiazol-2(3H)-imine 21d
White solid, mp 65.2–67.1 °C, yield 72%
1H NMR (400 MHz, Chloroform-d) δ 0.76 (s, 3H), 0.95 (s, 3H), 1.58 (dd, J = 2.8, 1.4 Hz, 3H), 1.60–1.80 (m, 8H), 1.81–1.85 (m, 6H), 1.86–1.97 (m, 2H), 1.99–2.08 (m, 3H), 2.34 (dddd, J = 15.2, 7.4, 2.9, 1.5 Hz, 1H), 3.84 (dd, J = 8.1, 6.4 Hz, 2H), 5.18–5.27 (m, 1H). 13C NMR (151 MHz, CDCl3) δ 163.4, 156.8, 148.3, 121.6, 47.5, 46.8, 46.6, 41.8, 37.9, 36.3, 35.2, 28.2, 28.1, 25.7, 19.7, 12.5. HRMS: m/z 371.2390 (M+ C22H33N3S1+, calc. 371.2388). \(\left[ \alpha \right]_D^{23.7}\) = −3.3 (c 0.31 in CHCl3).
5-(Adamantan-1-yl)-3-(((1 R)-6,6-dimethylbicyclo[3.1.1]hept-2-en-2-yl)methyl)-1,3,4-thiadiazol-2(3H)-imine 21e
Colorless oil, yield 61%
1H NMR (400 MHz, Chloroform-d) δ 0.74 (s, 3H), 1.15 (d, J = 8.6 Hz, 1H), 1.20 (s, 3H), 1.61 – 1.74 (m, 6H), 1.75–1.85 (m, 6H), 1.97–2.05 (m, 4H), 2.11 (td, J = 5.6, 1.5 Hz, 1H), 2.14–2.29 (m, 2H), 2.29–2.34 (m, 1H), 4.25–4.34 (m, 2H), 5.38–5.46 (m, 1H). 13C NMR (126 MHz, CDCl3) δ 163.7, 156.5, 143.0, 120.3, 51.3, 43.4, 41.6, 40.4, 37.9, 37.7, 36.2, 31.2, 31.0, 28.0, 26.1, 20.9. HRMS: m/z 369.2233 (M+ C22H31N3S1+, calc. 369.2230). \(\left[ \alpha \right]_D^{24.7}\) = −16.8 (c 0.46 in CHCl3).
5-(Adamantan-1-yl)-3-(2-((1 R)-6,6-dimethylbicyclo[3.1.1]hept-2-en-2-yl)ethyl)-1,3,4-thiadiazol-2(3H)-imine 21f
White solid, mp 49.5–49.6 °C, yield 70%
1H NMR (400 MHz, Chloroform-d) δ 0.79 (s, 3H), 1.13 (d, J = 8.5 Hz, 1H), 1.23 (s, 3H), 1.62–1.76 (m, 6H), 1.77–1.88 (m, 6H), 1.97–2.09 (m, 5H), 2.10–2.25 (m, 2H), 2.28–2.47 (m, 3H), 3.82 (ddd, J = 8.8, 6.7, 2.3 Hz, 2H), 5.27 (tt, J = 3.0, 1.5 Hz, 1H). 13C NMR (126 MHz, CDCl3) δ 163.1, 156.7, 144.6, 118.1, 77.1, 45.4, 45.4, 41.7, 40.5, 37.8, 36.2, 34.7, 31.5, 31.2, 28.0, 26.1, 21.0. HRMS: m/z 383.2390 (M+ C23H33N3S1+, calc. 383.2387). \(\left[ \alpha \right]_D^{23.7}\) = −11.5 (c 0.26 in CHCl3).
Biological assays
TDP1 activity
TDP1 activity was determined as described in the work [24]. Briefly, we used a DNA biosensor, which is a 16-mer single-stranded DNA oligonucleotide carrying a fluorophore (FAM) at the 5′-end and a quencher (BHQ1) at the 3′-end. The biosensor (5′-[FAM] AAC GTC AGGGTC TTC C [BHQ]-3′) was synthesized in the Laboratory of Nucleic Acids Chemistry at the Institute of Chemical Biology and Fundamental Medicine (Novosibirsk, Russia). Due to the activity of the enzyme, the quencher is removed, which leads to an increase in the fluorescence intensity. The reaction was carried out at different concentrations of inhibitors (1.5% of DMSO, Sigma, St. Louis, MO, USA, in the control samples). The reaction mixtures contained TDP1 buffer (50 mM Tris-HCl pH 8.0, 50 mM NaCl, and 7 mM β-mercaptoethanol), 50 nM biosensor, and an inhibitor being tested. Purified TDP1 (1.5 nM) triggered the reaction.
The reactions were incubated on a POLARstar OPTIMA fluorimeter (BMG LABTECH, GmbH, Ortenberg, Germany) to measure fluorescence. The values of IC50 were determined in minimum three independent experiments and were calculated using embedded software MARS Data Analysis 2.0 (BMG LABTECH, GmbH, Ortenberg, Germany).
Cytotoxicity assays
Cytotoxicity of the compounds against HeLa (human cervical cancer), MRC-5 (human lung fibroblasts), and HEK293A (human embryonic kidney) cell lines was examined using the EZ4U Cell Proliferation and Cytotoxicity Assay (Biomedica, Vienna, Austria), according to the manufacturer’s protocols. The cells were grown in DMEM with 50 IU/mL penicillin, 50 μg/mL streptomycin (MP Biomedicals, Santa Ana, CA, USA), and 10% of fetal bovine serum (Biolot, St. Petersburg, Russia) in a 5% CO2 atmosphere. After reaching 30–50% confluence, the tested compounds were added to the medium. The volume of the added reagents was 1/100 of the total volume of the culture medium, and the amount of DMSO (Sigma, St. Louis, MO, USA) was 1% of the final volume. A minimum of two independent tests were performed with each inhibitor. To evaluate the influence of the inhibitors on the cytotoxic effect of topotecan (Selleck Chemicals, Houston, TX, USA), 50% cytotoxic concentrations (CC50) of topotecan were determined either in the presence or in the absence of 10 µM TDP1 inhibitors. Minimum two independent tests were performed with each inhibitor in combination with topotecan.
References
Takahashi DT, Gadelle D, Agama K, Kiselev E, Zhang H, Yab E. et al. Topoisomerase I (TOP1) dynamics: conformational transition from open to closed states. Nat Commun. 2022;13:59 https://doi.org/10.1038/s41467-021-27686-7.
Pommier Y, Pourquier P, Fan Y, Strumberg D. Mechanism of action of eukaryotic DNA topoisomerase I and drugs targeted to the enzyme. Biochim Biophys Acta. 1998;1400:83–106. 10.1016/S0167-4781(98)00129-8.
Xu Y, Her C. Inhibition of topoisomerase (DNA) I (TOP1): DNA damage repair and anticancer therapy. Biomolecules. 2015;5:1652–70. https://doi.org/10.3390/biom5031652
Comeaux EQ, van Waardenburg RCAM. Tyrosyl-DNA phosphodiesterase I resolves both naturally and chemically induced DNA adducts and its potential as a therapeutic target. Drug Metab Rev 2014;46:494–507. https://doi.org/10.3109/03602532.2014.971957
Leung E, Patel J, Hollywood JA, Zafar A, Tomek P, Barker D, et al. Validating TDP1 as an inhibition target for the development of chemosensitizers for camptothecin-based chemotherapy drugs. Oncol Ther 2021;9:541–56. https://doi.org/10.1007/s40487-021-00158-0
Barthelmes HU, Habermeyer M, Christensen MO, Mielke C, Interthal H, Pouliot JJ, et al. TDP1 overexpression in human cells counteracts DNA damage mediated by topoisomerases I and II. J Biol Chem 2004;279:55618–25. https://doi.org/10.1074/jbc.M405042200
Meisenberg C. TDP1/TOP1 ratio as a promising indicator for the response of small cell lung cancer to topotecan. J Cancer Sci Ther 2014;06:258–67. https://doi.org/10.4172/1948-5956.1000280
Katyal S, El-Khamisy SF, Russell HR, Li Y, Ju L, Caldecott KW, et al. TDP1 facilitates chromosomal single-strand break repair in neurons and is neuroprotective in vivo. EMBO J. 2007;26:4720–31. https://doi.org/10.1038/sj.emboj.7601869
Das BB, Antony S, Gupta S, Dexheimer TS, Redon CE, Garfield S, et al. Optimal function of the DNA repair enzyme TDP1 requires its phosphorylation by ATM and/or DNA-PK. EMBO J. 2009;28:3667–80. https://doi.org/10.1038/emboj.2009.302
Ivankin DI, Kornienko TE, Mikhailova MA, Dyrkheeva NS, Zakharenko AL, Achara C, et al. Novel TDP1 inhibitors: disubstituted thiazolidine-2,4-diones containing monoterpene moieties. Int J Mol Sci 2023;24:3834. https://doi.org/10.3390/ijms24043834
Chepanova AA, Zakharenko AL, Dyrkheeva NS, Chernyshova IA, Zakharova OD, Ilina ES, et al. Influence of tyrosyl-DNA phosphodiesterase 1 inhibitor on the proapoptotic and genotoxic effects of anticancer agent topotecan. Dokl Biochem Biophys 2023;508:25–30. https://doi.org/10.1134/S1607672922700077
Salomatina OV, Popadyuk II, Zakharenko AL, Zakharova OD, Chepanova AA, Dyrkheeva NS, et al. Deoxycholic acid as a molecular scaffold for tyrosyl-DNA phosphodiesterase 1 inhibition: a synthesis, structure–activity relationship and molecular modeling study. Steroids. 2021;165:108771. https://doi.org/10.1016/j.steroids.2020.108771
Conda-Sheridan M, Reddy PVN, Morrell A, Cobb BT, Marchand C, Agama K, et al. Synthesis and biological evaluation of indenoisoquinolines that inhibit both tyrosyl-DNA phosphodiesterase I (Tdp1) and topoisomerase I (Top1). J Med Chem 2013;56:182–200. https://doi.org/10.1021/jm3014458
Yang H, Qin C, Wu M, Wang F, Wang W, Agama K, et al. Synthesis and biological activities of 11‐ and 12‐substituted benzophenanthridinone derivatives as DNA topoisomerase IB and tyrosyl‐DNA phosphodiesterase 1 inhibitors. ChemMedChem. 2023;18:1–16. https://doi.org/10.1002/cmdc.202200593
Zhao XZ, Wang W, Lountos GT, Kiselev E, Tropea JE, Needle D, et al. Identification of multidentate tyrosyl-DNA phosphodiesterase 1 (TDP1) inhibitors that simultaneously access the DNA, protein and catalytic-binding sites by oxime diversification. RSC Chem Biol 2023;4:334–43. https://doi.org/10.1039/D2CB00230B
Huang SN, Pommier Y, Marchand C. Tyrosyl-DNA phosphodiesterase 1 (Tdp1) inhibitors. Expert Opin Ther Pat 2011;21:1285–92. https://doi.org/10.1517/13543776.2011.604314
Zakharenko AL, Luzina OA, Chepanova AA, Dyrkheeva NS, Salakhutdinov NF, Lavrik OI. Natural products and their derivatives as inhibitors of the DNA repair enzyme tyrosyl-DNA phosphodiesterase 1. Int J Mol Sci 2023;24:5781. https://doi.org/10.3390/ijms24065781
Chepanova AA, Mozhaitsev ES, Munkuev AA, Suslov EV, Korchagina DV, Zakharova OD, et al. The development of tyrosyl-DNA phosphodiesterase 1 inhibitors. combination of monoterpene and adamantine moieties via amide or thioamide bridges. Appl Sci 2019;9:2767. https://doi.org/10.3390/app9132767
Mozhaitsev ES, Zakharenko AL, Suslov EV, Korchagina DV, Zakharova OD, Vasil’eva IA, et al. Novel inhibitors of DNA repair enzyme TDP1 combining monoterpenoid and adamantane fragments. Anticancer Agents Med Chem 2019;19:463–72. https://doi.org/10.2174/1871520619666181207094243
Munkuev AA, Mozhaitsev ES, Chepanova AA, Suslov EV, Korchagina DV, Zakharova OD, et al. Novel Tdp1 inhibitors based on adamantane connected with monoterpene moieties via heterocyclic fragments. Molecules. 2021;26:3128. https://doi.org/10.3390/molecules26113128
Munkuev AA, Dyrkheeva NS, Kornienko TE, Ilina ES, Ivankin DI, Suslov EV, et al. Adamantane-monoterpenoid conjugates linked via heterocyclic linkers enhance the cytotoxic effect of topotecan. Molecules. 2022;27:3374. https://doi.org/10.3390/molecules27113374
Ramprasad J, Nayak N, Dalimba U. Design of new phenothiazine-thiadiazole hybrids via molecular hybridization approach for the development of potent antitubercular agents. Eur J Med Chem 2015;106:75–84. https://doi.org/10.1016/j.ejmech.2015.10.035
Cressier D, Prouillac C, Hernandez P, Amourette C, Diserbo M, Lion C, et al. Synthesis, antioxidant properties and radioprotective effects of new benzothiazoles and thiadiazoles. Bioorg Med Chem 2009;17:5275–84. https://doi.org/10.1016/j.bmc.2009.05.039
Zakharenko A, Khomenko T, Zhukova S, Koval O, Zakharova O, Anarbaev R, et al. Synthesis and biological evaluation of novel tyrosyl-DNA phosphodiesterase 1 inhibitors with a benzopentathiepine moiety. Bioorg Med Chem 2015;23:2044–52. https://doi.org/10.1016/j.bmc.2015.03.020
Zhao XZ, Kiselev E, Lountos GT, Wang W, Tropea JE, Needle D, et al. Small molecule microarray identifies inhibitors of tyrosyl-DNA phosphodiesterase 1 that simultaneously access the catalytic pocket and two substrate binding sites. Chem Sci 2021;12:3876–84. https://doi.org/10.1039/D0SC05411A
Dyrkheeva NS, Filimonov AS, Luzina OA, Zakharenko AL, Ilina ES, Malakhova AA, et al. New hybrid compounds combining fragments of usnic acid and monoterpenoids for effective tyrosyl-DNA phosphodiesterase 1 inhibition. Biomolecules. 2021;11:973. https://doi.org/10.3390/biom11070973
Jones G, Willett P, Glen RC, Leach AR, Taylor R. Development and validation of a genetic algorithm for flexible docking 1. J Mol Biol 1997;267:727–48. https://doi.org/10.1006/jmbi.1996.0897
Eldridge MD, Murray CW, Auton TR, Paolini GV, Mee RP. Empirical scoring functions: I. The development of a fast empirical scoring function to estimate the binding affinity of ligands in receptor complexes. J Comput Aided Mol Des 1997;11:425–45. https://doi.org/10.1023/a:1007996124545
Verdonk ML, Cole JC, Hartshorn MJ, Murray CW, Taylor RD. Improved protein-ligand docking using GOLD. Proteins 2003;52:609–23. https://doi.org/10.1002/prot.10465
Korb O, Stützle T, Exner TE. Empirical scoring functions for advanced protein−ligand docking with PLANTS. J Chem Inf Model 2009;49:84–96. https://doi.org/10.1021/ci800298z
Mooij WTM, Verdonk ML. General and targeted statistical potentials for protein-ligand interactions. Proteins 2005;61:272–87. https://doi.org/10.1002/prot.20588
Wang Z, Sun H, Yao X, Li D, Xu L, Li Y, et al. Comprehensive evaluation of ten docking programs on a diverse set of protein–ligand complexes: the prediction accuracy of sampling power and scoring power. Phys Chem Chem Phys 2016;18:12964–75. https://doi.org/10.1039/C6CP01555G
Bissantz C, Folkers G, Rognan D. Protein-based virtual screening of chemical databases. 1. evaluation of different docking/scoring combinations. J Med Chem 2000;43:4759–67. https://doi.org/10.1021/jm001044l
Zakharenko A, Luzina O, Koval O, Nilov D, Gushchina I, Dyrkheeva N, et al. Tyrosyl-DNA Phosphodiesterase 1 Inhibitors: usnic acid enamines enhance the cytotoxic effect of camptothecin. J Nat Prod 2016;79:2961–7. https://doi.org/10.1021/acs.jnatprod.6b00979
Dyrkheeva NS, Filimonov AS, Luzina OA, Orlova KA, Chernyshova IA, Kornienko TE, et al. New hybrid compounds combining fragments of usnic acid and thioether are inhibitors of human enzymes TDP1, TDP2 and PARP1. Int J Mol Sci 2021;22:11336. https://doi.org/10.3390/ijms222111336
Zhu F, Logan G, Reynisson J. Wine compounds as a source for HTS screening collections. A feasibility study. Mol Inform 2012;31:847–55. https://doi.org/10.1002/minf.201200103
Matuszek AM, Reynisson J. Defining known drug space using DFT. Mol Inform 2016;35:46–53. https://doi.org/10.1002/minf.201500105
Berman HM. The protein data bank. Nucleic Acids Res. 2000;28:235–42. https://doi.org/10.1093/nar/28.1.235
Berman H, Henrick K, Nakamura H. Announcing the worldwide Protein Data Bank. Nat Struct Mol Biol 2003;10:980–80. https://doi.org/10.1038/nsb1203-980
Scigress, Vers. F.Q 3.4.4. Kyushu Systems Ltd. 2008–2023. http://www.scigress.com
Allinger NL, Yuh YH, Lii JH. Molecular mechanics. The MM3 force field for hydrocarbons. 1. J Am Chem Soc 1989;111:8551–66. https://doi.org/10.1021/ja00205a001
Lii JH, Allinger NL. Molecular mechanics. The MM3 force field for hydrocarbons. 2. Vibrational frequencies and thermodynamics. J Am Chem Soc 1989;111:8566–75. https://doi.org/10.1021/ja00205a002
Lii JH, Allinger NL. Molecular mechanics. The MM3 force field for hydrocarbons. 3. The van der Waals’ potentials and crystal data for aliphatic and aromatic hydrocarbons. J Am Chem Soc 1989;111:8576–82. https://doi.org/10.1021/ja00205a003
Gotō H, Ōsawa E. An efficient algorithm for searching low-energy conformers of cyclic and acyclic molecules. J Chem Soc Perkin Trans. 1993;2:187–98. https://doi.org/10.1039/P29930000187
QikProp, version 6.2; Schrödinger: New York, NY, USA, 2021.
Ioakimidis L, Thoukydidis L, Mirza A, Naeem S, Reynisson J. Benchmarking the reliability of QikProp. Correlation between experimental and predicted values. QSAR Comb Sci 2008;27:445–56. https://doi.org/10.1002/qsar.200730051
Eurtivong C, Reynisson J. The development of a weighted index to optimise compound libraries for high throughput screening. Mol Inform 2019;38:1800068. https://doi.org/10.1002/minf.201800068
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, et al. Gaussian 16, Revision C.01, 2019.
Becke AD. Density-functional exchange-energy approximation with correct asymptotic behavior. Phys Rev A. 1988;38:3098–3100. https://doi.org/10.1103/PhysRevA.38.3098
Becke AD. Density-functional thermochemistry. III. The role of exact exchange. J Chem Phys 1993;98:5648–52. https://doi.org/10.1063/1.464913
Lee C, Yang W, Parr RG. Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B. 1988;37:785–9. https://doi.org/10.1103/PhysRevB.37.785
Frisch MJ, Pople JA, Binkley JS. Self-consistent molecular orbital methods 25. Supplementary functions for Gaussian basis sets. J Chem Phys 1984;80:3265–9. https://doi.org/10.1063/1.447079
Hariharan PC, Pople JA. The influence of polarization functions on molecular orbital hydrogenation energies. Theor Chim Acta. 1973;28:213–22. https://doi.org/10.1007/BF00533485
Wong MW. Vibrational frequency prediction using density functional theory. Chem Phys Lett 1996;256:391–9. https://doi.org/10.1016/0009-2614(96)00483-6
Foresman JB, Frisch Æ, Exploring chemistry with electronic structure methods. Gaussian Inc: Wallingford, CT, 1996.
Acknowledgements
Authors would like to acknowledge the Multi-Access Chemical Research Center SB RAS for spectral and analytical measurements.
Funding
This research was supported by the Russian Science Foundation (grant 19-13-00040).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Munkuev, A.A., Zakharenko, A.L., Kornienko, T.E. et al. Synthesis of adamantane-monoterpene conjugates with 1,3,4-thiadiazol-2(3H)-imine linker and evaluation of their inhibitory activity against TDP1. Med Chem Res 33, 324–335 (2024). https://doi.org/10.1007/s00044-023-03184-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-023-03184-x