Abstract
The objective of this study was to design new nitroimidazole-based derivatives as strong urease inhibitors for the treatment of H. pylori infections. New series of nitroimidazole derivatives, 4a–k, were synthesized by using TBTU as the catalyst and assayed as Jack bean urease inhibitors. The facile synthetic approach was employed for the preparation of targeted molecular designs in good to excellent yields, ranged from 65 to 92%. Accordingly, all the synthesized compounds, 4a–k (IC50 = 1.43–7.72 μM), were more potent than the standard urease inhibitors, thiourea, and hydroxyurea. Among the derivatives, 4d had the most urease inhibitor activity (IC50 = 1.43 μM), over 15-fold more potent than thiourea and 70-fold than hydroxyurea. True to form, the result of molecular docking studies was in good congruence with those obtained from in vitro tests.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Helicobacter pylori (H. pylori) is one of the most widespread bacterial infections in the human population. However, a large number of patients do not reveal clinical pathogenic symptoms. The pathogen is transmitted orally during childhood through feces, vomitus, saliva, and water [1, 2]. H. pylori are well-known for triggering gastritis, gastric ulcers, and gastric adenocarcinoma [3]. Although both innate and adaptive immunity cooperate to restrain H. pylori from colonization, the pathogen applies various strategies to run away from the immune system [4].H. pylori can exert pathogenicity by overwhelming the acidic pH of the stomach through its vital enzyme, urease, a nickel-containing enzyme that catalyzes the hydrolysis of urea to two molecules of ammonia and carbon dioxide [5,6,7]. It utilizes two types of urease; a surface urease, which works in neutral pH, and a cytoplasmic urease, which is activated in acidic pH (2.5–6.5). The cytoplasmic urease is responsible for bacteria’s pH homeostasis. It is regulated by a \({\mathrm{H}}^ +\)- gated transporter facilitates urea permeability through the membrane and thus plays an essential role in bacterial acid resistance [8, 9].
Crystallographic analysis of H. pylori urease illustrates the spatial complexity of the enzyme that protects it against environmental catalytic change [10, 11]. Reports indicate that the active site of this particular enzyme in H. pylori is different from other bacterial ureases, which leads to a high affinity to the substrate [12]. The ammonia produced by the hydrolysis activity of urease neutralizes gastric hydrochloric acid, hence, eases bacterial colonization in epithelial cells [13].
Due to the growing concern of drug-resistant strains of H. pylori to antibiotics, the application of antibiotics to eliminate H. pylori is becoming less frequent and new methods for eradication are considered. Resistant bacteria survive from the treatment regimen, propagate, and settle in the mucosal layer of the stomach [14]. Consequently, the resistant population triggers the inoculum effect, which means a higher “minimum inhibitory concentration” is required to eradicate the pathogen [15]. The focus is now on alternative paths, such as building an unstable acidic environment for the pathogen. Therefore, therapeutic achievements depend on abolishing the H. pylori urease enzyme.
A typical treatment for H. pylori infection involves the co-administration of a proton pump inhibitor and antibiotics [16,17,18]. Drug-resistant strains have been reported after the frequent use of macrolides (clarithromycin or azithromycin), tetracycline, amoxicillin, and imidazole (metronidazole or tinidazole) [19,20,21]. Moreover, the higher dose of these antibiotics could result in adverse effects such as nausea, vomiting, abdominal pain, diarrhea, and bloating [22,23,24].
Lately, the in vitro efficacy assessment of urease enzyme inhibitors using phosphoramidites has been conducted successfully [25, 26]. However, the in-vivo tests and clinical trials were not promising due to problems such as hydrolytic instability and toxicity [27, 28]. Acetohydroxamic acid is one of the few urease inhibitors that has gained FDA approval [29,30,31]. Nitroimidazole derivatives are commonly used in antibacterial and antiprotozoal therapies [32].
According to previous studies, a considerable number of H. pylori strains (6–27%) are resistant to metronidazole, a common nitroimidazole derivative [33], due to frequent medications with nitroimidazoles [34]. Then, the main objective of this study was to design new metronidazole-based derivatives with remarkable inhibitory activity that could preserve metronidazole’s role as an acceptable antibiotic and could be used as strong urease inhibitors in the treatment of H. pylori infections.
Results and discussion
Chemistry
Eleven novel compounds have been synthesized as urease inhibitors according to the pathway shown in Scheme 1. In this current work 2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetohydrazide (3) was synthesized according to the previously mentioned method [35].
At the first step, ethyl 2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetate (2) was synthesized from a reaction between 2-methyl-5-nitro-1H-imidazole (1) and ethyl bromoacetate in dry acetone as a solvent, then the excess amount of hydrazine hydrate was added dropwise to compound 2 in ethanol solvent to produce 2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetohydrazide (3) [35]. In the third step, our final compounds (4a–k) were synthesized through a condensation reaction between 2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetohydrazide (3) and different aromatic acid derivatives in the presence of 2-(1H-benzotriazole-1-yl)-1,1,3,3-tetramethyluronium tetrafluoroborate (TBTU) as a coupling reagent while there exist triethylamine (TEA) as base and dimethylformamide (DMF) as a solvent in our reaction medium at 0 °C. Our final compounds yield ranged from 65 to 92%. Final compound structures were confirmed by using IR and NMR spectra.
Biological activity
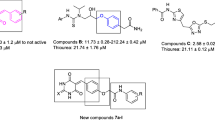
All eleven synthesized compounds (4a–k) were evaluated for their inhibitory activity against Jack bean urease. Thiourea and hydroxyurea were taken as the reference drug (Table 1). All of the tested compounds presented acceptable inhibitory activity mainly compounds 4d (IC50 = 1.43 µM) and 4k (IC50 = 1.70 µM). These two derivatives demonstrated potent in vitro inhibitory activity, which was comparable to standard inhibitor thiourea (IC50 = 22.01 µM) and hydroxyurea (IC50 = 100.00 µM) (Table 1). The comparison of the inhibitory activity of the synthesized compounds and metronidazole defined that the introduction of a hydrazine moiety instead of 2-hydroxy, and substitution on the benzene ring, play an important role in the urease inhibitory activity of synthetic compounds. Interestingly, methoxy-substituted compounds showed a different pattern.
Among them, compound 4k (IC50 = 1.70 µM), which bears a methoxy group at ortho-position, displayed an increased inhibitory action in comparison to compounds 4c (IC50 = 4.18 μM), with a methoxy group at para-position, and compound 4 h (IC50 = 2.74 μM), that bears both of them simultaneously.
Order of inhibitory activity stated the respective potency of 4k>4h > 4c . This outcome indicates that possibly the methoxy substitution at para-position leads to an increase in the steric hindrance or the ortho substitution interacts greatly with the active site of the enzyme. However, all of these compounds were more active as compared to the standard. Compound 4d (IC50 = 1.43 µM), holding an indole-3-acetic acid moiety, displayed fifteen-fold more activity in comparison to the standard thiourea and was the most potent derivative of the series. The desirable activity of this compound might be as a result of the appropriate binding interaction of the indole group with the active site of the enzyme. Subsequently, compound 4d exhibited a great inhibitory activity against Jack bean urease (Table 1), which fascinated our interest in more structure modification of them as lead compounds for new urease inhibitors or anti-H. pylori agents.
Docking study
To predict possible interactions between the synthesized compounds and the active site of the Jack bean urease enzyme, molecular docking simulations were performed. Docking results indicate that all of the compounds interact well with the active site of the enzyme with a binding energy range of −9.54 to −6.84 kcal/mol, which are stronger than the binding energy of the standard compounds thiourea and hydroxyurea, with a binding energy of −3.45 and −5.57 kcal/mol, respectively (Table 1). The superimposed position of docking poses of all compounds in the active site of the enzyme was shown in Fig. 1a. The highest binding energy, −9.54 kcal/mol, is related to compound 4d, representing the highest urease inhibitory activity with an IC50 of 1.432 µM in comparison to other compounds. The placement of the compound 4d in the active site suggests that the inhibitory effect is probably related to the conformational changes that occur at the entrance of the tunnel leading to the nickel atoms, and therefore, it could prevent the substrate from entering the tunnel (Fig. 1b). Compound 4d, with an indole ring, forms three hydrogen bonds with Gln 635, Gly 638, and Gly 641 residues and interacts hydrophobically with Glu 418, Arg 639, Val 640, and Glu 642 (Fig. 1c). The SwissADME webserver was used to determine the ADME properties [36]. So, compound 4d, with a molecular weight of 356.34 g/mol, log p −0.14, forms 3 and 5 hydrogen bonds, donor and acceptor, respectively, fulfill all the criteria declared in the Lipinski rule of five (Fig. 1d) and preset its potential drug ability [37]. It is expected that compound 4d, with the highest urease inhibitory activity, and also the highest binding interactions, and proper ADME properties, could be a promising candidate for further evaluation.
Conclusion
In this current work, eleven new nitroimidazole derivatives were synthesized and their inhibitory activity against the Jack bean urease enzyme was determined. The obtained results demonstrated that all the synthesized compounds were more potent than the standard inhibitors. The docking study showed that all of the compounds successfully occupied and interacted with the active site of the urease enzyme. Eventually, among all compounds, compound 4d indicates the lowest IC50 value of urease inhibitory activity and lowest docking binding energy and is known as the potent compound of this series. According to the urease inhibitory activity of our final compounds, it seems beneficial to consider nitroimidazole as an important moiety for design and synthesize potent urease inhibitors.
Experimental/material and methods
General
All chemicals used in this study were obtained from Merck (Germany) and used without further purification. Melting points were determined with a Kofler hot stage apparatus and were uncorrected. 1H and 13C NMR spectra were recorded with a Bruker FT-500, using TMS as an internal standard. Coupling constant (J) values are presented in Hertz (Hz), and spin multiples are given as s (singlet), d (doublet), t (triple), and m (multiple). Infrared spectra were acquired on a Nicolet Magna 550-FT spectrometer. IR spectra of solid were recorded in KBr, and the absorption band was given in wavenumbers in cm−1.
Urease inhibitory activity
Sodium nitroprusside and Jack bean urease (EC 3.5.1.5) were purchased from Sigma (St. Louis, MO, USA). Ultra-pure water (HPLC grade, Duksan, Korea) was utilized in all experiments. The compounds including, 4a–k and inhibitors, were dissolved in deionized water. The urease inhibitory activity of the synthesized compounds was assessed by a procedure explained previously [38, 39].
In brief, the modified Berthelot spectrophotometric method was used to evaluate urease inhibitory of the synthesized derivatives at the concentration range of 0–10 mg/ml and measuring the absorbance at 625 nm. The jack bean urease enzyme was activated after 30 min incubation at 37 °C, and then, the absorbance of the testing solutions was measured after adding a combination of phenol and alkali reagents. The thiourea and hydroxyurea were used as the reference standard inhibitors, and all the experiments were performed in triplicate.
Finally, the percentage of urease inhibition activity was calculated as
T is the absorbance of the test samples (synthesized compounds (4a–k) or positive control) in the presence of the enzyme, and C (control) is the absorbance of the reagents in the presence of the enzyme. The IC50 values were calculated using GraphPad Prism 5 software (GraphPad Software Inc., San Diego, CA) (Table 1).
Docking method
Preparation of the input files of the ligands and receptor was done by AutoDockTools 1.5.6 (ADT) [40]. Jack bean urease enzyme with PDB ID: 3LA4 was downloaded from the protein data bank. MarvinSketche version 15.2.2 was used to draw the structure of the ligands in 2D format. Chem3D ultra version 8.0 was used to minimized energy and converting the 2D structure to PDB format. Receptor preparation was performed as follows, all water molecules were deleted, and then polar hydrogens were added and, Kollman charges were assigned, and non-polar hydrogens were merged. Grid box with the size of 60 × 60 × 60 Å with grid center −43.931, −43.497, 77.423 (X, Y, and Z) and grid-point spacing of 0.375 Å was located near the Ni atom in the active site of the enzyme. The grid maps of each atom type were calculated by AutoGrid 4.2 [40]. The following parameter was set to calculate docking simulations by AutoDock 4.2: 2.5 × 106 maximum number of energy evaluations with 30 run jobs and Lamarckian genetic algorithm (LGA) with the population size of 150, and the default value for the rest of the parameters [39,40,41]. The docking poses with the lowest binding energy value of each compound were used to illustrate by LIGPLOT+ version v.2.2 [42] and PyMol version 1. Level [43].
General procedure for the synthesis of compounds 4a–k
Ethyl (2-methyl-5-nitro-1H-imidazol-1-yl)acetate (2)
A mixture of 2-methyl-5-nitro-1H-imidazole 1 (1 mmol), ethyl bromoacetate (1 mmol), and potassium carbonate (1.5 mmol) (5–10 mL) was refluxed in dry acetone for 50 h. The reaction mixture was cooled to room temperature, and the solvent was distilled off from the filtrate. The crude ester thus obtained was purified by recrystallization from ethanol. Yield 85% (213.07 mg); mp 96 °C [35].
2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetohydrazide (3)
Ethyl (2-methyl-5-nitro-1H-imidazol-1-yl)acetate 2 (2 mmol) was dissolved in ethanol as a solvent and, then, an excessive amount of hydrazine hydrate (3 mmol) was added dropwise to the mixture. The reaction mixture was refluxed for 10 h and then cooled to room temperature. The solid formed, filtered, dried, and recrystallized from ethanol to get pure compound 3. Yield 60% (238.8 mg); mp 189 °C [35].
Experimental procedure for the synthesis of nitroimidazole derivatives (4a–k)
A mixture of benzoic acid derivatives (1 mmol), 0.32 g (1 mmol) of TBTU, and 0.28 mL (2 mmol) of triethylamine (TEA) in 2 mL dimethylformamide (DMF) was stirred at 0 °C for 3 min. Then 2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetohydrazide 3 (1 mmol) was added to the reaction medium. The reaction mixture was stirred at 0 °C for 1 h and left overnight at room temperature. The reaction mixture was poured into ice water, filtered, washed with 5% aqueous citric acid solution (2 × 10 mL), saturated sodium bicarbonate solution (2 × 10 mL), and water. The crude product was recrystallized from dichloromethane to obtain pure final compounds (4a–k).
4-chloro-N′-(2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetyl)benzohydrazide (4a)
White solid; yield: 78% (263.4 mg), mp = 231–233 °C. IR (KBr): νmax 3318 (NH), 3055 (C–H aromatic), 2986 (C–H aliphatic), 1688 (C=O), 1563–1358 (NO2) cm−1. 1H NMR (DMSO-d6, 500 MHz): δ = 2.34 (s, 3H, CH3), 4.97 (s, 2H, CH2), 7.58–7.60 (d, J = 8.1 Hz, 2H, H-3, H-5), 7.88–7.90 (d, J = 8.1 Hz, 2H, H-2, H-6), 8.35 (s, 1H, CH, Imidazole), 10.55–10.67 (m, 2H, 2×NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.1(CH3), 48.2 (CH2), 129.3 (C-1, Ph), 129.9 (CH, Ph), 130.2 (CH, Ph), 131.4 (CH, Ph), 137.5 (C-4, Ph), 137.8 (C-4, imidazole), 145.7 (C-5, imidazole), 146.5 (C-2, imidazole), 165.1 (C=O), 165.9 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C13H12ClN5O4: 338.0650, found: 337.0578.
4-methyl-N′-(2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetyl)benzohydrazide (4b)
White solid; yield: 78% (247.5 mg), mp = 228–230 °C. IR (KBr): νmax 3375 (NH), 3089 (C–H aromatic), 2979 (C–H aliphatic), 1680 (C=O), 1558–1346 (NO2) cm−1. 1Hs NMR (DMSO-d6, 500 MHz): δ = 2.25 (s, 3H, CH3), 2.36 (s, 3H, CH3), 4.96 (s, 2H, CH2), 7.21–7.23 (d, J = 8.25 Hz, 2H, H-3, H-5), 7.86–7.88 (d, J = 8.25 Hz, 2H, H-2, H-6), 8.35 (s, 1H, CH, Imidazole’), 10.43 (s, 2H, 2×NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.1 (CH3), 21.6 (CH3), 48.2 (CH2), 123.9 (C-4, Ph), 128.0 (CH, Ph), 129.6 (CH, Ph), 129.7 (C–H, Ph), 129.9 (C-1, Ph), 142.6 (C-4, Imidazole), 145.7 (C-5, Imidazole), 146.5 (C-2, Imidazole), 165.9 (C=O), 166.0 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C14H15N5O4: 317.1124, found: 318.1199.
4-methoxy-N′-(2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetyl)benzohydrazide (4c)
White solid; yield: 72% (239.9 mg), mp = 249–251 °C. IR (KBr): νmax 3335 (NH), 3026 (C–H aromatic), 2991 (C–H aliphatic), 1680 (C=O), 1558–1354 (NO2) cm−1. 1H NMR (DMSO-d6, 500 MHz): δ = 2.36 (s, 3H, CH3), 4.96 (s, 2H, CH2), 7.02–7.04 (d, J = 8.35 Hz, 2H, H-3, H-5), 7.86–7.88 (d, J = 8.35 Hz, 2H, H-2, H-6), 8.35 (s, 1H, CH, Imidazole’), 10.43 (s, 2H, 2×NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.1(CH3), 48.2 (CH2), 56.0 (O–CH3), 114.3 (CH-2, Ph), 114.4 (C-6, Ph), 124.0 (CH-3, Ph), 124.3 (CH-5, Ph), 124.8, 130.0 (C-1, Ph), 141.3 (C-4, Imidazole) 145.7 (C-5, Imidazole), 146.5 (C-2, Imidazole), 163.0 (C-4, Ph), 165.6 (C=O), 165.9 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C14H15N5O5: 333.1073, found: 332.1003.
(1H-Indol-3-yl)-acetic acid N′-[2-(2-methyl-5-nitro-imidazol-1-yl)-acetyl]-hydrazide (4d)
White solid; yield: 65% (231.6 mg), mp = 234–236 °C. IR (KBr): νmax 3318 (NH), 3055 (C–H aromatic), 2986 (C–H aliphatic), 1688 (C=O), 1563–1358 (NO2) cm−1. 1H NMR (DMSO-d6, 500 MHz): δ = 2.29 (s, 3H, CH3), 3.59 (s, 2H, CH2), 4.87 (s, 2H, CH2), 6.96–6.99 (t, J = 7.75 Hz, 1H, H-4), 7.05–7.08 (t, J = 7.75 Hz, 1H, H-5), 7.24 (d, J = 2.5 Hz, 1H, H-2), 7.33–7.35 (d, J = 8.3 Hz, 1H, H-7), 7.57–7.59 (d, J = 8.3 Hz, 1H, H-4), 8.31 (s, 1H, CH, Imidazole’), 10.20 (s, 1H, NH), 10.37 (s, 1H, NH), 10.89 (s, 1H, NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.1 (CH3), 31.1 (CH2), 48.1 (CH2), 108.6 (C-3-indole), 111.9 (CH-7-indole), 118.9 (CH-4-indole), 119.3 (CH-5-indole), 121.6 (CH-6-indole), 124.0, (CH-2-indole), 124.5 (C-3a-indole), 127.7 (C-7a-indole), 136.6 (C-4, Imidazole), 145.7 (C-5, Imidazole), 146.4 (C-2, Imidazole), 165.4 (C=O), 170.2 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C16H16N6O4: 356.1233, found: 357.1307.
2,6-dichloro-N′-(2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetyl)benzohydrazide (4e)
White solid; yield: 65% (241.9 mg), mp = 221–223 °C. IR (KBr): 3348 (NH), 3029 (C–H aromatic), 2964 (C–H aliphatic), 1672 (C=O), 1545–1359 (NO2) cm−1. 1H NMR (DMSO-d6, 500 MHz): δ = 2.34 (s, 3H, CH3), 4.46 (s, 2H, CH2), 7.59–7.62 (t, J = 7.6 Hz, 1H, H-4), 7.79–7.81 (d, J = 7.65 Hz, 2H, H-3, H-5), 8.25 (s, 1H, CH, Imidazole’), 10.21 (s, 2H, 2×NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.4(CH3), 48.8 (CH2), 126.4 (CH-4, Ph), 129.1 (CH-3, Ph), 129.6 (CH-5, Ph), 131.8 (C-2, Ph), 132.3 (C-6, Ph), 133.7 (C-4, Imidazole), 135.1 (C-5, Imidazole), 141.3 (C-2, Imidazole), 164.0 (C=O), 165.4 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C13H12FN5O4: 321.0873, found: 322.0943.
N′-(2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetyl)benzohydrazide (4f)
White solid; yield: 84% (254.7 mg), mp = 234–236 °C. IR (KBr): νmax 3340 (NH), 3063 (C–H aromatic), 2978 (C–H aliphatic), 1679 (C=O), 1550–1352 (NO2) cm−1. 1H NMR (DMSO-d6, 500 MHz): δ = 2.35 (s, 3H, CH3), 4.97 (s, 2H, CH2), 7.49–7.52 (t, J = 7.6 Hz, 2H, H-3, H-5), 7.56–7.58 (t, J = 7.65 Hz, 1H, H-4), 7.87 (d, J = 7.6 Hz, 2H, H-2, H-6), 8.34–8.36 (s, 1H, CH, Imidazole’), 10.53 (s, 2H, 2×NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.1 (CH3), 48.2 (CH2), 124.0 (CH-4, Ph), 128.1(CH-3, Ph), 128.2 (CH-5, Ph), 129.1 (CH-2, Ph), 129.2 (CH-6, Ph), 132.6 (C-1, Ph), 136.7 (C-4, Imidazole), 145.7 (C-5, Imidazole), 146.5 (C-2, Imidazole), 165.9 (C=O), 166.1 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C13H13N5O4: 303.0968, found: 304.1042.
N′-(2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetyl)-4-nitrobenzohydrazide (4g)
White solid; yield: 92% (320.4 mg), mp = 272–274 °C. IR (KBr): νmax 3358 (NH), 3055 (C–H aromatic), 2983 (C–H aliphatic), 1692 (C=O), 1568–1360 (NO2) cm−1. 1H NMR (DMSO-d6, 500 MHz): δ = 2.35 (s, 3H, CH3), 4.99 (s, 2H, CH2), 8.09–8.11 (d, J = 8.25 Hz, 2H, H-2, H-6), 8.35 (s, 1H, CH, Imidazole), 8.36–8.38 (d, J = 8.25 Hz, 2H, H-3, H-5), 10.80 (s, 2H, 2×NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.1(CH3), 48.2 (CH2), 124.0 (C-1, Ph),124.3 (CH-3, Ph), 124.4 (CH-2, Ph), 138.3 (C-4, Imidazole), 145.7 (C-5, Imidazole), 146.5 (C-2, Imidazole), 150.1(C-4, Ph), 164.6 (C=O), 165.8 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C13H12N6O6: 348.0818, found: 347.0743.
2,4-dimethoxy-N′-(2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetyl)benzohydrazide (4 h)
White solid; yield: 75% (272.4 mg), mp = 237–239 °C. IR (KBr): νmax 3328 (NH), 3040 (C–H aromatic), 2975 (C–H aliphatic), 1680 (C=O), 1555–1355 (NO2) cm−1. 1H NMR (DMSO-d6, 500 MHz): δ = 2.35 (s, 3H, CH3), 3.83 (s, 3H, O–CH3), 3.90 (s, 3H, O–CH3), 4.92 (s, 2H, CH2), 6.79–6.80 (d, J = 1.5 Hz, 2H, H-3, H-5), 7.79 (s, 1H, H-6), 8.32 (s, 1H, CH, Imidazole’), 9.88 (s, 1H, NH), 10.69 (s, 1H, NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.1 (CH3), 48.1 (CH2), 56.1 (O–CH3), 56.7 (O– CH3), 99.1 (CH-2, Ph), 106.5 (CH-5, Ph), 124.0 (C-1, Ph), 133.1(C-4, Imidazole), 145.7 (C-5, Imidazole), 146.5 (C-2, Imidazole), 159.4 (C-2, Ph), 163.8 (C-4, Ph), 164.2 (C=O), 164.8 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C15H17N5O6: 363.1179, found: 364.1256.
N′-(2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetyl)-2, Phenoxybenzohydrazide (4i)
White solid; yield: 68% (268.8 mg), mp = 195–197 °C. IR (KBr): νmax 3346 (NH), 3052 (C–H aromatic), 2984 (C–H aliphatic), 1685 (C=O), 1548–1339 (NO2) cm−1. 1H NMR (DMSO-d6, 500 MHz): δ = 2.32 (s, 3H, CH3), 4.91 (s, 2H, CH2), 6.87–6.89 (d, J = 8.35 Hz, 1H, H3), 7.05–7.07 (d, J = 7.9 Hz, 2H, H-2, H-6), 7.16-7.19 (t, J = 8.3 Hz, 1H, H-4), 7.64–7.66 (d, J = 7.6 Hz, 1H, H-6), 8.32 (s, 1H, CH, Imidazole’), 10.33 (s, 1H, NH), 10.65 (s, 1H, NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.1 (CH3), 48.1 (CH2), 119.1 (CH-2ʹ,6ʹ, Ph), 119.2 (CH-4ʹ, Ph), 119.8 (CH-3, Ph), 123.9 (CH-5, Ph), 124.5 (C-1, Ph), 126.1 (CH-6, Ph), 130.6 (CH-3ʹ,5ʹ, Ph), 132.9 (C-4, Imidazole), 145.7 (C-5, Imidazole), 146.5 (C-2, Imidazole), 154.9 (C-2, Ph), 156.7 (CH-1ʹ, Ph), 164.9 (C=O), 165.3 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C19H17N5O5: 395.1229, found: 396.1302.
N′-(2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetyl)nicotinohydrazide (4j)
White solid; yield: 68% (206.9 mg), mp = 266–268 °C. IR (KBr): νmax 3338 (NH), 3045 (C–H aromatic), 2988 (C–H aliphatic), 1679 (C=O), 1610 (C = N), 1564–1342 (NO2) cm−1. 1H NMR (DMSO-d6, 500 MHz): δ = 2.35 (s, 3H, CH3), 4.97 (s, 2H, CH2), 7.52–7.54 (d, J = 6.5 Hz, 1H, H5), 8.20–8.22 (d, J = 7.7 Hz, 1H, H-6), 8.34 (s, 1H, CH, Imidazole), 8.73–8.75 (t, J = 3.5 Hz, 1H, H-5), 9.02 (s, 1H, H-2), 10.68 (s, 2H, 2×NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.1 (CH3), 48.3 (CH2), 123.7 (C-5-pyridin), 124.3 (C-1-pyridin), 135.0 (C-6-pyridin), 135.7 (C-4, Imidazole), 145.7 (C-5, Imidazole), 146.5 (C-2, Imidazole), 149.0 (C-4-pyridin), 152.9 (C-2-pyridin), 164.4 (C=O), 165.4 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C12H12N6O4: 304.0920, found: 305.0992.
2-methoxy-N′-(2-(2-methyl-5-nitro-1H-imidazol-1-yl)acetyl)benzohydrazide (4k)
White solid; yield: 70% (231.2 mg), mp = 251–253 °C. IR (KBr): νmax 3358 (NH), 3044 (C–H aromatic), 2989 (C–H aliphatic), 1680 (C=O), 1545–1360 (NO2) cm−1. 1H NMR (DMSO-d6, 500 MHz): δ = 2.35 (s, 3H, CH3), 3.88 (s, 3H, O–CH3), 4.94 (s, 2H, CH2), 7.04–7.07 (t, J = 7.7 Hz, 1H, H-5), 7.15–7.17 (d, J = 8.35 Hz, 1H, H-3), 7.50–7.53 (t, J = 7.85 Hz, 1H, H-4), 7.71–7.73 (d, J = 7.85 Hz, 1H, H-6), 8.34 (s, 1H, CH, Imidazole), 10.10 (s, 1H, NH), 10.72 (s, 1H, NH) ppm. 13C NMR (DMSO-d6, 500 MHz): δ = 13.1 (CH3), 48.1 (CH2), 56.5 (CH3), 112.7 (CH-3, Ph), 121.2 (CH-5, Ph), 122.0 (C-1, Ph), 124.2 (CH-4, Ph), 130.9 (CH-6, Ph), 133.6 (C-4, Imidazole), 145.7 (C-5, Imidazole), 146.5 (C-2, Imidazole), 157.6 (C-2, Ph), 165.0, 165.1 (C=O), 165.5 (C=O) ppm. HRMS: m/z [M+H]+ calcd for C14H15N5O5: 333.1073, found: 334.1146.
References
Cave DR. Transmission and epidemiology of Helicobacter pylori. Am J Med. 1996;100:12S–8S. https://doi.org/10.1016/S0002-9343(96)80224-5
Suerbaum S, Michetti P. Helicobacter pylori infection. N Engl J Med. 2002;347:1175–86. https://doi.org/10.1056/NEJMra020542
Krzyżek P, Gościniak G, Fijałkowski K, Migdał P, Dziadas M, Owczarek A, et al. Potential of bacterial cellulose chemisorbed with anti-metabolites, 3-bromopyruvate or sertraline, to fight against helicobacter pylori lawn biofilm. Int J Mol Sci. 2020;21:9507 https://doi.org/10.3390/ijms21249507
Portal-Celhay C, Perez-Perez GI. Immune responses to helicobacter pylori colonization: mechanisms and clinical outcomes. Clin Sci. 2006;110:305–14. https://doi.org/10.1042/CS20050232
Svane S, Sigurdarson JJ, Finkenwirth F, Eitinger T, Karring H. Inhibition of urease activity by different compounds provides insight into the modulation and association of bacterial nickel import and ureolysis. Sci Rep. 2020;10:1–4. https://doi.org/10.1021/ac60252a045
Kafarski P, Talma M. Recent advances in design of new urease inhibitors: A review. J Adv Res. 2018;13:101–12. https://doi.org/10.1016/j.jare.2018.01.007
Mazzei L, Musiani F, Ciurli S. The structure-based reaction mechanism of urease, a nickel dependent enzyme: tale of a long debate. J Bio Inorg Chem. 2020;18:1–7. https://doi.org/10.1007/s00775-021-01855-x
Graham DY, Miftahussurur M. Helicobacter pylori urease for diagnosis of Helicobacter pylori infection: a mini review. J Adv Res. 2018;13:51–7. https://doi.org/10.1016/j.jare.2018.01.006
Hassan ST, Šudomová M. The development of urease inhibitors: what opportunities exist for better treatment of Helicobacter pylori infection in children? Children. 2017;4(1):2 https://doi.org/10.3390/children4010002
Hu LT, Foxall PA, Russell R, Mobley H. Purification of recombinant Helicobacter pylori urease apoenzyme encoded by ureA and ureB. Infect Immun. 1992;60:2657–66. PubMed:1612736
Zerner B. Recent advances in the chemistry of an old enzyme, urease. Bioorg Chem. 1991;19:116–31. https://doi.org/10.1016/0045-2068(91)90048-T
Rektorschek M, Buhmann A, Weeks D, Schwan D, Bensch KW, Eskandari S, et al. Acid resistance of Helicobacter pylori depends on the UreI membrane protein and an inner membrane proton barrier. Mol Microbiol. 2000;36:141–52. https://doi.org/10.1046/j.1365-2958.2000.01835.x
Arora R, Issar U, Kakkar R. Identification of novel urease inhibitors: pharmacophore modeling, virtual screening and molecular docking studies. J Biomol Struct Dyn. 2019;37:4312–26. https://doi.org/10.1080/07391102.2018.1546620
Schmalstig AA, Benoit SL, Misra SK, Sharp JS, Maier RJ. Noncatalytic antioxidant role for Helicobacter pylori urease. J Bacteriol. 2018;200(17):e00124–18. https://doi.org/10.1128/JB.00124-18
Graham DY. Antibiotic resistance in Helicobacter pylori: implications for therapy. Gastroenterology. 1998;115:1272–7. https://doi.org/10.1016/S0016-5085(98)70100-3
Savarino V, Dulbecco P, De Bortoli N, Ottonello A, Savarino E. The appropriate use of proton pump inhibitors (PPIs): need for a reappraisal. Eur. J. Intern. Med. 2017;37:19–24. https://doi.org/10.1016/j.ejim.2016.10.007
Chen P, Li L, Wang H, Zhao J, Cheng Y, Xie J, et al. Omeprazole, an inhibitor of proton pump, suppresses De novo lipogenesis in gastric epithelial cells. Biomed Pharmacother. 2020;130:110472 https://doi.org/10.1016/j.biopha.2020.110472
Hassan S, Majerová M, Šudomová M, Berchová K. Antibacterial activity of natural compounds-essential oils. Ceska Slov Farm. 2015;64(Dec):243–53. (6)Czech. PMID: 26841699
Shmuely H, Topaz S, Berdinstein R, Yahav J, Melzer E. High metronidazole and clarithromycin resistance of helicobacter pylori isolated from previously treated and naïve patients. Isr Med Assoc J. 2020;22:562–6. PMID: 33070487
Gehlot V, Mahant S, Mukhopadhyay AK, Das K, De R, Kar P, et al. Antimicrobial susceptibility profiles of Helicobacter pylori isolated from patients in North India. J Glob Antimicrob Resist. 2016;5:51–6. PMID: 28224028
Chaabane NB, Al-Adhba HS. Ciprofloxacin-containing versus clarithromycin-containing sequential therapy for Helicobacter pylori eradication: A randomized trial. Indian J Gastroenterol. 2015;34:68–72. https://doi.org/10.1007/s12664-015-0535-x
Bartnik W. Clinical aspects of Helicobacter pylori infection. Pol Arch Med Wewn. 2008;118:426–30. PMID: 18714738
Wang Y, Zhao R, Wang B, Zhao Q, Li Z, Zhu-Ge L, et al. Sequential versus concomitant therapy for treatment of Helicobacter pylori infection: an updated systematic review and meta-analysis. Eur J Clin Pharmacol. 2018;74:1–13. https://doi.org/10.1007/s00228-017-2347-7
Hassan ST, Berchová K, Majerová M, Pokorná M, Švajdlenka E. In vitro synergistic effect of Hibiscus sabdariffa aqueous extract in combination with standard antibiotics against Helicobacter pylori clinical isolates. Pharm Biol. 2016;54:1736–40. https://doi.org/10.3109/13880209.2015.1126618
Hassan ST, Žemlička M. Plant‐derived urease inhibitors as alternative chemotherapeutic agents. Arch Pharm. 2016;349:507–22. https://doi.org/10.1002/ardp.201500019
Kosikowska P, Berlicki Ł. Urease inhibitors as potential drugs for gastric and urinary tract infections: a patent review. Expert opin Ther Pat. 2011;21:945–57. https://doi.org/10.1517/13543776.2011.574615
Modolo LV, de Souza AX, Horta LP, Araujo DP, de Fatima A. An overview on the potential of natural products as ureases inhibitors: A review. J Adv Res. 2015;6:35–44. https://doi.org/10.1016/j.jare.2014.09.001
Follmer C. Ureases as a target for the treatment of gastric and urinary infections. J Clin Pathol. 2010;63:424–30. https://doi.org/10.1136/jcp.2009.072595
Jain SK, Haider T, Kumar A, Jain A. Lectin-conjugated clarithromycin and acetohydroxamic acid-loaded PLGA nanoparticles: a novel approach for effective treatment of H. pylori. AAPS PharmSciTech. 2016;17:1131–40. https://doi.org/10.1208/s12249-015-0443-5
Umamaheshwari R, Jain N. Receptor-mediated targeting of lipobeads bearing acetohydroxamic acid for eradication of Helicobacter pylori. J control Release. 2004;99:27–40. https://doi.org/10.1016/j.jconrel.2004.06.006
Griffith DP. Urease stones. Urol Res. 1979;7:215–21. https://doi.org/10.1007/BF00257208
Naureen S, Chaudhry F, Asif N, Munawar MA, Ashraf M, Nasim FH, et al. Discovery of indole-based tetraarylimidazoles as potent inhibitors of urease with low antilipoxygenase activity. Eur J Med Chem. 2015;102:464–70. https://doi.org/10.1016/j.ejmech.2015.08.011
Cunha ES, Chen X, Sanz-Gaitero M, Mills DJ, Luecke H. Cryo-EM structure of Helicobacter pylori urease with an inhibitor in the active site at 2.0 Å resolution. Nat. Commun. 2021;12:1–8. https://doi.org/10.1038/s41467-020-20485-6
Der Wouden V, Zwet V. nitroimidazole resistance in Helicobacter pylori. Aliment Pharmacol Ther. 2000;14:7–14. https://doi.org/10.1046/j.1365-2036.2000.00675.x
Wani MY, Bhat AR, Azam A, Athar F. Nitroimidazolyl hydrazones are better amoebicides than their cyclized 1, 3, 4-oxadiazoline analogues: In vitro studies and Lipophilic efficiency analysis. Eur J Med Chem. 2013;64:190–9. https://doi.org/10.1016/j.ejmech.2013.03.034
Daina A, Michielin O, Zoete V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep. 2017;7:42717 https://doi.org/10.1038/srep42717
Benet LZ, Hosey CM, Ursu O, Oprea TI. BDDCS, the rule of 5 and drugability. Adv Drug Deliv Rev. 2016;101:89–98. https://doi.org/10.1016/j.addr.2016.05.007
Peytam F, Adib M, Mahernia S, Rahmanian-Jazi M, Jahani M, Masoudi B, et al. Isoindolin-1-one derivatives as urease inhibitors: Design, synthesis, biological evaluation, molecular docking and in-silico ADME evaluation. Bioorg Chem. 2019;87:1–11. https://doi.org/10.1016/j.bioorg.2019.02.051
Azizian H, Nabati F, Sharifi A, Siavoshi F, Mahdavi M, Amanlou M. Large-scale virtual screening for the identification of new Helicobacter pylori urease inhibitor scaffolds. J Mol Model. 2012;18:2917–27. https://doi.org/10.1007/s00894-011-1310-2
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem. 2009;30:2785–91. https://doi.org/10.1002/jcc.21256
Hosseini FS, Amanlou M. Anti-HCV and anti-malaria agent, potential candidates to repurpose for coronavirus infection: Virtual screening, molecular docking, and molecular dynamics simulation study. Life Sci. 2020;258:118205 https://doi.org/10.1016/j.lfs.2020.118205
Laskowski RA, Swindells MB. LigPlot+: multiple ligand–protein interaction diagrams for drug discovery. J Chem Inf Model. 2011;51:2778–86. https://doi.org/10.1021/ci200227u
Schrödinger LLC. The PyMol molecular graphics system. 2017.Available from: https://pymol.org
Acknowledgements
The research reported in this publication was supported by Elite Researcher Grant Committee under award number [996450] from the National Institute for Medical Research Development (NIMAD), Tehran, Iran.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Talebi, M., Hamidian, E., Niasari-Naslaji, F. et al. Synthesis, molecular docking, and biological evaluation of nitroimidazole derivatives as potent urease inhibitors. Med Chem Res 30, 1220–1229 (2021). https://doi.org/10.1007/s00044-021-02727-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-021-02727-4